Abstract
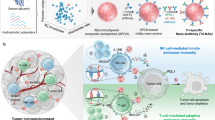
Colorectal cancer (CRC) is the second leading cause of cancer-related mortality, and the incidence of early-onset colon cancer has been increasing globally in recent years. The development of immunotherapies for colon cancer is critical for providing new treatment strategies to combat drug resistance. Here, a bispecific nanobody against PD-L1 (BsNb-PD-L1) constructed using genetically encoded noncanonical amino acids (ncAAs) is reported. A computational protocol was developed to identify appropriate sites in the nanobody for incorporating p-acetylphenylalanine (pAcF). Variants of nanobodies PV2 and PV3 with pAcF incorporated were conjugated with linkers containing an aminooxy functionality to enable oxime ligation. The resulting PV2-S71 + PV3-N77 bispecific nanobody (BsNb-ncAA) exhibited higher thermostability and binding affinity compared to the nanobody monomers and the BsNb constructed by simply fusing two proteins. Moreover, in an in vitro phagocytosis model, the BsNb-ncAA exhibited improved capability to inhibit immune evasion and showed stronger biological activity compared to the fusion protein PV2-PV3. Furthermore, the BsNb-ncAA resulted in a marked increase in the number of CD8+ T cells within tumor tissues and demonstrated efficient inhibitory effects against colon tumor growth in vivo. Our study provides a general strategy for constructing BsNbs, which has potential applications in other cancer immunotherapy.

Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the second leading cause of cancer-related mortality, and the incidence of early-onset colon cancer has been increasing worldwide globally in recent years1. The growing popularity of meat-sweet diets, environmental degradation from industrialization, and inadequate colonoscopy screening programs have led to a troubling rise in the incidence of CRC in developing countries. The global incidence is projected to reach 2.5 million new cases annually by 20352. Approximately 20% of patients suffer from metastatic colorectal cancer at first diagnosis, and up to 50% of patients with localized disease eventually develop metastases3. Unfortunately, the five-year survival rate of metastatic CRC is less than 20%. Systemic therapy is the primary treatment for metastatic CRC and the precise treatment based on the molecular pathogenesis of the tumor improves overall survival4,5. Remarkably, immune checkpoint inhibitor (ICI) treatment targeting the PD-1/PD-L1 pathway is highly effective for metastatic mismatch repair-deficient and microsatellite instability-high (d-MMR/MSI-H) CRC6. However, the overall response rate is limited by the poor permeability of conventional anti-PD-L1 monoclonal antibodies to tumor tissues and lamina propria7,8,9. It is imperative to develop miniaturized, high-affinity anti-PD-L1 antibody drugs to treat CRC.
Nanobodies, known as single-domain antibodies (sdAb), have only one heavy chain variable region domain (VHH). Nanobodies have the characteristics of small molecular weight, strong tissue permeability, stable structure and low immunogenicity, making them well-suited for humanization and affinity maturation10. Therefore, nanobodies are no longer seen as an alternative to conventional monoclonal antibodies and are rationally designed to deal with complex application scenarios and change the pattern of disease treatment11. Nanobody-based reagents were developed to diagnose and treat several cancer diseases. At the same time, multivalent and multi-specific nanobodies were constructed to improve their affinity and specificity12. However, our previous study revealed that fusion-expressed multivalent nanobodies exhibited reduced affinity, possibly due to the spatial structure blocking the binding epitope13. The loss of affinity can be mitigated by optimizing the type and length of linker, but the development of site-specific multivalent nanobodies construction methods has a wide range of applications.
A number of recombinant strategies have been developed to synthesize bispecific antibodies (BsAbs), which include single chain variable fragment (scFv)-derived formats such as diabodies, tandem diabodies, BiTEs (bispecific T-cell engager), and DARTs (Dual Affinity Re-Targeting), as well as immunoglo-bulin G (IgG)-based formats such as Triomab, DVD-Ig (Dual Variable Domain antibodies), and two-in-one antibodies14. One commonly employed approach for the production of bispecific antibodies involves the use of fusion proteins. In this method, two antibodies with distinct target specificities are linked together through a short linker of peptides. This strategy is favored for its inherent flexibility15,16. Although this method allows modular assembly of antibody fragments, the genetic fusion process is inherently restricted to a two-dimensional configuration, with paired genes constrained to either the 5’ or 3’ terminal of the construct. Such linear connectivity fails to account for the three-dimensional spatial organization critical to protein function17. Other studies have demonstrated that forced head-to-tail fusion can disrupt interdomain cooperativity, leading to reduced binding avidity or unintended steric clashes that compromise biological activity18,19. To overcome these constraints, surface-directed conjugation strategies have emerged as a promising alternative. By exploiting solvent-exposed amino acid side chains on antibody frameworks, these methods enable precise control over protein orientation during conjugation, thereby facilitating the rational design of bispecific antibodies with optimized spatial arrangements for enhanced activity and stability. The success of such surface-based conjugation hinges on two critical criteria: (i) the availability of orthogonal chemical handles that selectively target specific amino acid residues without cross-reacting with other surface-exposed side chains, and (ii) coupling chemistries that preserve native protein folding and function. Several chemical approaches have been developed that predominantly utilize the reactivity of lysine or cysteine residues within the antibody. However, lysine modification often yields heterogeneous products due to multiple reactive surface lysine residues in the antibody. While cysteine-based approaches are more selective, the reaction is more complicated due to presence of multiple disulfide bonds in the antibody.
Genetically encoded noncanonical amino acids (ncAAs) have been applied to generate BsAbs with defined geometries. Specifically, the p-acetylphenylalanine (pAcF) was incorporated into each antibody, and the ketone group of pAcF was linked using linkers with an aminooxy functionality to enable oxime ligation. This method synthesizes an anti-HER2/anti-CD3 BsAb, demonstrating excellent in vitro activity and efficiently cross-linked HER2+ and CD3+ cells7. A similar method has also been applied for the synthesis of a BsAb, aCLL1-aCD3, which recruits cytotoxic T cells to CLL1 positive cells. This BsAb demonstrates potent and selective cytotoxicity in vitro and effective treatment of tumors in the cell line xenograft model20. Additionally, a BsAb targeting B cell maturation antigen (BCMA) has been developed. This BsAb redirects T cells to lyse malignant multiple myeloma cells, which demonstrates a great potency in lysing BCMA-positive cell lines21. However, despite these advances, generating BsAb through the incorporation of ncAAs still presents challenges. Due to the low incorporation efficiency of ncAAs, the production yield of antibodies containing ncAAs is low, which limits the overall production of BsAbs. Additionally, the ncAA incorporation sites must be selected carefully, as the inappropriate incorporation could impact the antibody stability and binding affinity. Currently, no general methods have been developed to guide the selection of ncAA incorporation sites in antibodies.
In this study, we developed a strategy to synthesize BsNb against PD-L1 for colorectal cancer, utilizing the incorporation of ncAA, pAcF. We first established a computational protocol to evaluate the stability and binding affinity of all possible single point mutants with pAcF incorporated. Based on these computations, pAcF was incorporated into specific sites, and the BsNb-ncAA was constructed using two linkers that conjugate with pAcF through stable oxime bond. The resulting BsNb-ncAA demonstrated high affinity, stability, and specificity. It effectively blocked the PD-1/PD-L1 pathway and showed promising results in animal models.
Results and discussion
Computation-aided identification of ncAA incorporation site
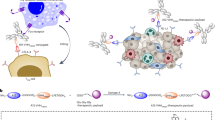
Two high-affinity anti-PD-L1 nanobodies, PV2 and PV3, sharing a sequence identity of 86.5%, were selected for constructing bispecific nanobodies (BsNbs) for CRC treatment (Fig. S1). The molecular docking results indicate that PV2 and PDL1, as well as PV3 and PDL1, bind to different epitopes. The amino acid contact sites involved in the interactions also differ between the two complexes, with PV3 engaging with a greater number of contact points on PDL1 compared to PV2 (Fig. S2), and their combination was expected to improve reactivity and reduce off-target effects compared to a single nanobody22. To synthesize an effective BsNb, the ncAA pAcF was first incorporated at defined sites in each nanobody. The mutant nanobodies were then selectively coupled by a stable oxime bond to bifunctional linkers, with an alkoxy-amine on one terminus and an azide or cyclooctyne group on the other (Fig. 1A). The two nanobody-linker conjugates were linked to obtain the BsNb through a copper-free [3 + 2] Huisgen cyclo-addition7,23. We selected pAcF as the handles for constructing conjugates instead of other chemistry such as azide mediated click chemistry as the cyclopropane-tetrazine ligation employed in the pAcF system demonstrates superior bioorthogonality compared to azide-alkyne cycloadditions, effectively minimizing off-target reactions in complex cellular environments. Additionally, the pAcF has become a widely applied ncAA in bioconjugate chemistry, particularly for antibody conjugation applications, owing to its well-established genetic encoding systems and proven biocompatibility profiles24,25,26.
A General scheme for generating BsNbs; B Flowchart for computationally selecting suitable ncAA incorporation sites; C Estimated Cartesian_ddG and Flex_ddG values for PV2 and PV3 using Rosetta Cartesian_ddG and Flex_ddG. Pink dots, amino acids predicted to be suitable pAcF incorporation sites; Orange dots, amino acids predicted to significantly influence stability. Amino acids in CDR region are removed; D, Structures of PV2 and PV3 showing the positions of pAcF incorporation.
Since the incorporation of pAcF at inappropriate sites could impact the stability and affinity of the nanobody, we developed computational protocol to aid in selecting ncAA incorporation sites (Fig. 1B). Previously, combinations of various computational approaches have successfully estimated the effects of mutations on protein stability and activity27. Here, we employed AlphaFold2, molecular dynamics (MD) simulations, ZDOCK and Rosetta to accurately estimate nanobody stability and affinity change upon ncAA mutation (Fig. 1B). For stability evaluation, the structures of nanobodies PV2 and PV3 were first obtained using AlphaFold2. To further assess conformational dynamics, MD simulations were performed for nanobodies. Subsequently, Rosetta Cartesian_ddG protocol was applied to calculate the ddG values of all the single point variants with pAcF incorporated at all possible sites. For evaluation of the binding affinity, the antigen structures were first obtained from RCSB protein data bank. To refine these structures and assess their dynamic stability, MD simulations of the antigen models were performed. ZDOCK was then applied to obtain the complex structures of the antigen and nanobodies. These structures were evaluated for binding affinity changes upon pAcF incorporation using the Rosetta Flex_ddG protocol. After obtaining the Cartesian_ddG and Flex_ddG data, variants showing improved stability and binding affinity were manually checked for pAcF orientation. Since the BsNbs were constructed through corresponding linkers, variants with pAcF facing outward would be more appropriate than those facing inward for synthesizing BsNbs. Additionally, tools including AbSRA and NovoPro were used to identify the complementary determining region (CDR) regions, which are not ideal sites for incorporating ncAA due to the risk of impacting binding affinity.
Stability and binding affinity changes relative to the wild type were calculated for all possible single variants with pAcF substitution in nanobodies PV2 and PV3 using Rosetta Cartesian_ddG and Flex_ddG (Fig. 1C). The mutants exhibited a significant effect on the nanobodies stability, as evidenced by the Cartesian ddG values ranging from -8.9 kcal/mol to 63.8 kcal/mol for PV2 and −9.4 kcal/mol to 73.3 kcal/mol for PV3. Moreover, the variants exhibited Flex ddG values ranging from −0.8 kcal/mol to 2.4 kcal/mol, implying a limited effect on binding affinity, which might be attributed to variants being distant from protein binding interfaces (Fig. 1C). According to the results of the Rosetta Cartesian_ddG and Flex_ddG, we manually checked the PV2 and PV3 variants predicted to be highly stable, exhibit low binding free energy and are located outside the CDR regions, and those having pAcF facing outward were selected for experimental characterization, including PV2 variants S8pAcF, S26pAcF, S71pAcF, S74pAcF, Q82pAcF, S85pAcF and PV3 variants S8pAcF, S26pAcF, S71pAcF, N77pAcF, S85pAcF, K87pAcF (Fig. 1D & Table S1). Variants predicted to significantly decrease stability, such as C23pAcF, I70pAcF, C96pAcF for PV2 and C23pAcF, I70pAcF, G119pAcF for PV3, were also included as controls. A total of 18 mutants were finally selected, with 9 variants each for PV2 and PV3.
Characteristics of nanobodies with pAcF incorporated
An evolved tRNA/aminoacyl-tRNA synthetase (pCNFRS) pair was selected to site-specifically incorporate pAcF at defined sites in each of two nanobodies in response to an amber nonsense codon27. Considering the low yield of expressed protein with pAcF incorporated, we engineered pCNFRS to improve the incorporation of pAcF by introducing a quadruple mutation H283T/P284S/M285D/D286V (TSDV) located at the anticodon recognition domain that was proved useful in pAzFRS for improving the incorporation of (p-azido-phenylalanine) pAzF28. We tested the performance of the mutation in incorporating pAcF at the second position of sfGFP. In the results, the cells containing pCNFRS-TSDV showed a significant improvement in the fluorescence intensity compared to the wild type pCNFRS, while the misincorporation of canonical amino acids was weakly decreased (Fig. 2A). The SDS-PAGE of purified sfGFP also showed that TSDV mutation in pCNFRS increased the yield of sfGFP with pAcF incorporated by around 2-fold, consistent with the fluorescence intensity observed for purified protein (Fig. 2B). The pCNFRS-TSDV was then applied to construct the nanobodies with pAcF incorporated at different sites.
A The effect of TSDV mutation on pAcF-incorporated sfGFP expression; Treated, sfGFP2TAG expression in presence of pAcF in cell growth medium; Control, sfGFP2TAG expression in absence of pAcF in cell growth medium; GFP(WT), wild-type sfGFP expression. B Expression of sfGFP with pAcF incorporated by pCNFRS and pCNFRS-TSDV; C The melting temperatures of PV2 and PV3 nanobody mutants. Melting temperatures of PV2 and PV3 variants were measured using differential scanning fluorimetry. (Note: The data in the figure are x ± SD, n = 5 independent experiment; a significant difference from the WT group, *: p < 0.05, **: p < 0.01, ***: p < 0.001, ****: p < 0.0001); D Binding affinity of PV2 and PV3 nanobody mutants determined by ELISA (n = 3 independent experiment); E, The binding constant of PV2 and PV3 nanobody mutants.
PV2 and PV3 nanobody mutants with pAcF incorporated were purified and verified by SDS-PAGE and mass spectrometry to ensure correct insertion of the ncAA (Figures S3&4). The yield of all ncAA containing nanobodies ranged from 0.2 to 3.9 mg per liter culture, lower than that of wild type nanobody (Table S2). This could be attributed to the low efficiency of ncAA incorporation system, although we have engineered the pCNFRS for improved incorporation of pAcF (Fig. 2A). ELISA and differential scanning fluorescence tests were then conducted to measure the thermostabilities and binding affinities of nanobody variants. As for the PV2 nanobody, mutants S26pAcF, S71pAcF, and Q82pAcF showed Tm values of 52.2 °C, 52.9 °C, and 53.1 °C, higher than 47.9 °C of wild type. Moreover, C96pAcF showed the lowest thermostability of 42.2 °C, among all the variants, 5°C lower than the wild type, consistent with the Cartesian ddG value (Fig. 2C). Other variants showed similar Tm values to those of the wild type. As for the PV3 variants, S71pAcF and N77pAcF had Tm values of 58.7 °C and 60.0 °C, 6.7 °C and 8 °C higher than that of wild-type PV3. G119pAcF showed lower Tm than the wild type, consistent with the Cartesian ddG value (Fig. 2C). However, unexpectedly, K87pAcF showed the lowest stability among all the variants with the Tm value of 44.4 °C, although it was predicted to be more stable than the wild type. Additionally, both C23pAcF and I70pAcF of PV2 and PV3, which were predicted to have low stability, demonstrated similar or even greater thermostability than the wild type. This suggests that there may be a false negative error in the Rosetta Cartesian ddG program.
We further carried out 60-ns MD simulations with 3 replicas for PV2-C23pAcF and PV2-C96pAcF variants to understand the effect of pAcF incorporation on protein conformation, and the RMSD, RMSF, radius of gyration, the existence of hydrogen bonds and salt bridges were analyzed (Figs. S5-S8 and Tables S3-S4). PV2-C23pAcF and PV2-C96pAcF exhibited higher RMSD, RMSF and radius of gyration values. This is consistent with Cartesian_ddG calculations. A disulfide bond is formed between C23 and C96 in the nanobodies, and introducing pAcF at these positions would break the disulfide bond. However, the stability of C23pAcF increases after the disulfide bond is destroyed, which is the opposite of the stability change of C96pAcF. MD simulations demonstrated that, compared to the wild-type PV2 (WT), PV2-C23pAcF exhibited a significantly higher total number of hydrogen bonds in replica 1—approximately 80 more—whereas PV2-C96pAcF showed only a slight increase relative to WT (Fig. S8). In replica 1, the extra hydrogen bonds in C23pAcF were found to be distributed throughout various regions of the protein, which would contribute to the increased stability observed (Fig. S9A). Figure S9B further showed that the number of hydrogen bonds formed by Cys23 in WT was markedly lower than in C23pAcF, indicating increased local interactions at this site of C23pAcF. Conversely, Cys96 in WT formed similar hydrogen bonds with C96pAcF. The breakage of the disulfide bond at C96pAcF would hence lead to a decrease in the stability. Despite certain discrepancies between the predicted and the actual results, the computation tools successfully identified nanobody mutants with significantly improved thermal stability, demonstrating the feasibility of using this prediction scheme for identifying stable nanobody mutants with pAcF incorporated.
The binding affinity of wild-type and mutant PV2 and PV3 to the antigen PD-L1 was also characterized by ELISA (Fig. 2D). PD-L1 was prepared using the HEK-293 eukaryotic expression system and coated on a 96-well plate for antigen-antibody binding detection. Overall, considering the affinity results of the PV2 and PV3 nanobody mutants, all mutants maintained a high level of affinity. Only a few of the variants showed a decrease in affinity, which might be due to the selected mutations being distant from the CDR region.
The binding affinity of the wild-type PV2 was 15.1 nM. Most of the mutants showed increased binding affinity, with S71pAcF and S26pAcF having the most significant improvements at 1.7 nM and 1.6 nM, representing a 9-fold increase compared to the wild type. C96pAcF showed the lowest binding affinity, with a Kd value of 37.4 nM, 22.3 nM higher than that of the wild type, consistent with the Flex ddG calculation (Fig. 2E). N74pAcF, predicted to have the most significant improvement in binding affinity, indeed showed enhanced binding compared to the wild type. The affinity of the PV3 wild type was 51.0 nM, whereas the mutants K87pAcF, C23pAcF, S85pAcF and N77pAcF showed a significant increase in binding affinity. K87pAcF had the highest affinity (13.7 nM), a 3.7-fold increase compared to the wild type (Fig. 2D, E). Unexpectedly, the S71pAcF that was predicted to have an improvement in binding affinity showed a slightly higher Kd value than the wild type.
The computer-aided screening method narrowed the range of mutants for experimental testing. Most of the predicted optimal mutation sites showed improved thermal stability and affinity compared to the wild type, demonstrating the feasibility of this screening approach. Considering the Tm values and binding affinities of all PV2 and PV3 mutants, PV2-S26pAcF, PV2-S71PAcF, PV3-S85pAcF and PV3-N77pAcF which showed significantly improved stability and binding affinity, were selected for 2 times 2 combination, resulting in four pairs of BsNbs.
Synthesis of bispecific nanobodies
Nanobodies harboring pAcF were individually ligated with TET or BCN linkers. Purified PV2-TET was then conjugated with PV3-BCN to produce heterodimeric bispecific nanobodies containing ncAA (BsNbs-ncAA). Four pairs of BsNbs including PV2-S26 + PV3-S77, PV2-S26 + PV3-S85, PV2-S71 + PV3-S77 and PV2-S71 + PV3-S85 were constructed and purified by size-exclusion chromatography (Fig. S10). All four pairs of BsNbs exhibited the corresponding BsNb peak in chromatogram, and the conjugation efficiency was greater than 50% (Table S5). PV2-S71 + PV3-N77 exhibited the highest BsNb yield of 74.9%, while PV2-S26 + PV3-S85 showed a low coupling efficiency of 52.9% compared to other BsNbs (Fig. 3A). SDS-PAGE confirmed the correct formation of the BsNb PV2-S71 + PV3-N77, which was selected for further characterization of thermostability and binding affinity (Fig. 3B). The BsNb PV2-PV3 constructed by simply fusing two nanobodies PV2 and PV3 (BsNb-fusion) was also characterized for comparison.
A Conjugation efficiency of four different BsNbs. The peak area of BsNb and unbound nanobodies were analyzed when protein purification was carried out using the size-exclusion chromatography. The ratio was calculated as the percentage ratio of BsNbs peak area to the total integrated peak area; B Characterization of nanobody monomers and conjugates by SDS-PAGE. M, marker; Lane 1-7: PV2-S71, PV2-S71-Tet, PV3-N77, PV3-N77-BCN, PV2-S71 + PV3-N77, PV2-S71 + PV3-N77 purified, PV2-PV3 fusion protein; C Melting temperatures of PV2-PV3 and PV2-S71 + PV3-N77. The data in the figure are x ± SD, n = 5 independent experiment; a significant difference among the groups, ****: p < 0.0001); D EC50 values of nanobodies and BsNbs towards PD-L1 obtained by ELISA measurements. (n = 3 independent experiment) E Binding affinity of PV2, PV3, PV2-PV3 and PV2-S71 + PV3-N77; F SPR sensorgram binding profiles of PV2-PV3 and PV2-S71 + PV3-N77.
The measured Tm values were 56.8 oC and 50.5 oC for the PV2-S71 + PV3-N77 and fusion protein PV2-PV3, respectively, indicating the superior thermal stability of the BsNb constructed with ncAA (Fig. 3C). This can be attributed to the incorporation of pAcF at the mutation sites, as PV2-S71pAcF and PV3-N77pAcF enhanced the thermal stability by 5 oC and 8 oC, respectively (Fig. 2C). The EC50 values obtained fitting the ELISA values revealed that the apparent binding affinity of the PV2-PV3 fusion protein was 61.1 nM, while that of PV2-S71 + PV3-N77 was 6.1 nM, representing a 10-fold improvement in binding affinity for BsNb-ncAA compared to BsNb-fusion (Fig. 3D, E). BsNb-ncAA also showed a significant increase in binding efficiency relative with wild-type PV2 and PV3 (Fig. 3D, E). However, the EC50 of PV2-S71 + PV3-N77 was lower than PV2-S71pAcF and higher than PV2-N77pAcF, suggesting a compromise between two single variants (Fig. 2E). Thermostability and binding measurement suggest that introducing pAcF not only provides a chemical handle for conjugation but also facilitates in vitro affinity maturation and thermostability improvement, leading to the construction of BsNbs with enhanced binding efficacy and thermostability.
To confirm the binding affinities of the two bispecific nanobody constructs, we also performed surface plasmon resonance (SPR) measurements to determine kD value (Fig. 3F). Based on results of SPR, the PV2-PV3 fusion protein demonstrated a kD value of 63.5 nM, while the PV2-S71 + PV3-N77 combination exhibited a significantly enhanced binding affinity with a kD of 6.34 nM. This 10-fold improvement in binding affinity for the PV2-S71 + PV3-N77 complex aligns with previous ELISA results, validating the affinity advantage associated with this bispecific nanobody construction method.
Bispecific nanobody activity measured by macrophage phagocytosis
Previous studies have demonstrated that PD-L1 antibodies can significantly enhance the proliferation and activation of macrophages in vivo, leading to a proinflammatory phenotype29,30. In our study, we used the phagocytosis rate of activated macrophages against colon cancer cells with high PD-L1 expression as an indicator of their immune capacity. HT29 cells (red) and macrophages (blue) were stained with DiD dye and Hoechst 33342 dye, respectively (Fig. 4A). After antibody administration and co-culture, fluorescence was observed under a confocal microscope to detect the phagocytosis rate of macrophages. Over one hundred photos were taken, and one hundred randomly selected blue fluorescence-labeled macrophages were observed and recorded to count the number of macrophages exhibiting red and blue merged fluorescence, indicating phagocytosis. The phagocytosis rate for each group was then calculated (Fig. 4). Among the three antibody drug groups, the BsNb-ncAA and the Durvalumab, a FDA-approved immune checkpoint inhibitor drug, showed a significant difference (p < 0.05, n = 3) compared to the PBS control group, while the BsNb-fusion and wild-type PV2 did not show a significant difference (Fig. 4A, B). The phagocytosis rate of the PV2-PV3 was approximately 17.7%, while that of PV2-S71 + PV3-N77 was approximately 26.3%. These results indicated that in the in vitro phagocytosis model, the BsNb-ncAA exhibited improved capability to inhibit immune evasion and had stronger biological activity than the fusion protein PV2-PV3.
A PD-L1 blockade increases macrophage phagocytosis of tumor cells in vitro under laser confocal scanning microscopy; B In vitro phagocytosis of DiI-labeled HT-29 cells pretreated with PBS or anti-PD-L1 antibody. The data in the figure. (x ± SD, n = 3 biologically independent samples; a significant difference from the PBS group, *: p < 0.05).
In vivo antitumor activity of bispecific nanobody
To further validate the in vivo anti-tumor activity of the BsNb-ncAA developed, therapeutic experiments were conducted on mice. The anti-tumor effects of PV2-S71 + PV3-N77 were determined using an hPD-L1 MC38 PD-1 knock-in mice model (Fig. 5A). Tumor volumes and body weight were measured every two days and the results were as good as the ones obtained with the commercial antibody Durvalumab (Fig. 5B, C), although it must be underlined that bispecific nanobodies and Durvalumab were used at the same concentrations (10 μg/mL), namely the molar concentration of the bivalent nanobodies was six times higher. The results clearly showed that PV2-S71 + PV3-N77 treatment delayed tumor progression compared to the PV2-PV3 group, indicating stronger tumor inhibition capacity (Fig. 5D).
A Experimental design for in vivo tumor-challenge and PV2-S71 + PV3-N77 treatment plan. MC38 xenograft mice model was established. PV2-PV3, PV2-S71 + PV3-N77 and durvalumab (10 mg/kg, once every three days for six times) was treated for 15 days when the tumor grew to 80–100 mm3. (created with BioRender.com, with permission); B The tumor volume of mice treated with different treatment strategies; C Body weight of each group (n = 6 biologically independent samples); D The tumor weight and image of tumors isolated from tumor-bearing mice on day 18; E Effect of bispecific nanobodies on the cytokine concentration in serum; F H&E and IHC staining of tumor tissues; G The expression of Ki67 and CD8 were confirmed using an immunohistochemistry assay in tumor tissues. (x ± SD, n = 6 biologically independent samples; a significant difference from the Normal saline group, *: p < 0.05, **: p < 0.01, ***: p < 0.001).
During tumor suppression, CD8+ T cells are identified as one of the main effector cells blocking the PD-1/PD-L1 signaling pathway to activate an immune response. The activation of tumor-specific CD8+ T cells within tumors is a crucial indicator of immunotherapy, and IFN-γ serves as an important biomarker for producing CD8+ T cells31,32. Additionally, blocking the PD-1/PD-L1 signaling pathway through therapy can result in an increase in IFN-γ production33. Furthermore, IL-2, a T cell growth factor, is used as an indicator for T cell proliferation and activation. The concentration of IL-2 and IFN-γ in the serum of the four groups of mice were hence measured to reflect the blocking effect of treatments on the PD-1/PD-L1 pathway. The results indicated that the levels of IL-2 and IFN-γ in the PV2- S71 + PV3-N77 group were significantly increased compared to those in the PV2 + PV3 group (Fig. 5E). This suggests an enhancement in the secretion of cytokines involved in anti-tumor immune response in the PV2-S71 + PV3-N77 group. The IHC results also revealed a significant increase in the number of tumor-infiltrating CD8+ T cells in the PV2-S71 + PV3-N77 group compared to the PV2 + PV3 group, indicating stronger antigen-specific CD8+ T cell activation to combat cancer cells. Additionally, the expression of Ki67 was significantly reduced in the PV2-S71 + PV3-N77 group compared to the PV2 + PV3 group (Fig. 5F, G). Ki67 is present during all active phases of the cell cycle (G1, S, G2, and M) but absent in resting cells (G0), making it a key marker for assessing tumor cell proliferation.
These findings revealed a more pronounced effect on the tumor proliferation of the PV2-S71 + PV3-N77 group. We noticed that the PV2-S71 + PV3-N77 group exhibited a more pronounced tumor inhibition effect and enhanced secretion of immune response cytokines compared to the Durvalumab group. This finding contrasts with the results obtained from previous immunophagocytosis experiments. We consider this discrepancy stems from the unique in vivo pharmacological advantages of nanobodies with its small size enabling a deeper tumor penetration. Previous studies have also mentioned that tumour blood vessels are leaky, containing pores that can reach over 100 nm in size. This leads to a phenomenon that is referred to as the enhanced permeability and retention effect (EPR)34,35. Therefore, we posit that this differential pharmacological properties may account for the observed discrepancy in experimental outcomes between the in vitro phagocytosis assays and in vivo tumor suppression models.
To further illustrate the observed in vivo effects of the BsNb-ncAA treated group, we analyzed infiltrating immune cells in the tumors of four groups of mice. Flow cytometry results revealed that the proportion of CD45+ cell in tumor tissues was slightly higher in the PV2-S71 + PV3-N77 group compared to both the control group and the fusion protein group (Fig. 6A–C). Additionally, the proportion of CD3+, CD4+, and CD8+ T cells was significantly higher in the BsNb-ncAA group. Furthermore, we detected CD8+ T cell infiltration in each group of tumors by flow cytometry. As shown in Fig. 6D, there was a significant increase in the number of CD8+ T cells within tumor tissues in the PV2-S71 + PV3-N77 treatment group. It indicated that PV2-S71 + PV3-N77 could indirectly inhibit tumor growth by blocking the PD-1/PD-L1 pathway and further promoting the infiltration of CD8+ T cells into tumor tissues. In addition, we evaluated the proportion of CD8+ T cells producing IFN-γ within tumor tissues, and a significant increase was observed in this population within the BsNb-ncAA treated group as compared to the other treatment group (Fig. 6E). These findings suggested that improved tumor inhibition in the PV2-S71 + PV3-N77 group possibly resulted from increased infiltration of CD8+ T cells within the tumor as well as activation and proliferation of CD8+ T cells. Although the PV2-S71 + PV3-N77 obtained showed improved tumor growth inhibition efficiency, the clinical application of the bispecific nanobody requires deliberate optimization to minimize immunogenicity. Although their humanized nanobody framework inherently reduces anti-drug antibody (ADA) risks compared to camelid-derived precursors, additional humanization through CDR grafting onto human IgG scaffolds may prove beneficial for chronic therapies. Complementary approaches like surface charge neutralization, PEGylation modifications, or Fc-region fusion could further enhance serum half-life while mitigating ADA responses36,37.
A Single-cell suspensions from the excised MC38 tumors treated as indicated were stained for immune cell markers CD45, CD3, CD4, and CD8 for T cells according to the analysis method shown in (B); C CD8+ IFN-γ+ T cell were stained and analyzed; D The CD8+ T cells infiltration in each group; E The ratio of CD8+ IFN-γ+ T cells in the tumor. (x ± SD, n = 6 biologically independent samples; a significant difference from the Normal saline group, *: p < 0.05, **: p < 0.01, ***: p < 0.001).
Conclusion
In summary, we developed bispecific nanobodies (BsNbs) that could simultaneously target different epitopes of the PD-L1 for CRC treatment through the technology of gene codon expansion. A computational protocol was initially developed to identify the suitable sites for incorporating pAcF, which acts as a handle for linking two nanobodies through covalent bonds. We verified that the BsNb-ncAA exhibited enhanced thermostability and affinity in vitro compared to both the single nanobodies and the BsNbs constructed by fusion expression of two nanobodies (BsNb-fusion). Moreover, a macrophage phagocytosis study revealed that the BsNb-ncAA exhibited improved capability to inhibit immune evasion and demonstrated stronger biological activity than the BsNb-fusion. These superior properties of the BsNb-ncAA led to efficient tumor growth inhibition in hPD-L1 MC38 PD-1 knock-in mice. Flow cytometry results indicated that the improved tumor inhibition by BsNb-ncAA was likely due to their ability to stimulate stronger CD8+ cytotoxic T cells. We expect the proposed strategy for constructing BsNb will provide a robust and general platform useful for immunotherapy of other cancers.
Materials and methods
Calculation system setup
Nanobody structures PV2 and PV3 were calculated by AlphaFold238,39,40. The antigen structures of PD-L1 (PDB ID: 5GRJ)41 were taken from the RCSB Crystal Database. Crystallographic water molecules and unrelated residues in PDBs were removed. The CDR region was identified by both Chothia and Kabat methods from the online tool AbRSA [http://aligncdr.labshare.cn/aligncdr/abrsa.php] and Novo Pro [https://www.novopro.cn/tools/].
Molecular dynamics simulations
As shown in Fig. 1, the first batch of MD simulations was carried out for 2 wild type nanobody systems (PV2-WT and PV3-WT) and 1 antigen system (PD-L1) at 300 K. The second batch of MD simulations was performed on 2 nanobody systems PV2-C23pAcF and PV2-C96pAcF. The protonation states of titratable residues (histidine, glutamic acid and aspartic acid) were assigned at experimental pH using PROPKA342,43. MD simulations are conducted using CHARMM36 protein force fields44 within GROMACS 2022.3 package45. The force constant of the harmonic biasing potential is 1000 kJ mol-1 nm2. The TIP3P water model was used to solvate the protein-ligand complex in a rectangular box with periodic boundary conditions and a minimum distance between the solute and the box of 10 Å. Sodium and Chloride ions were added to the solution to maintain the system’s neutrality.
Initially, the system was minimized for 10,000 steps using the steepest descent algorithm. In the pre-equilibrium stage, the system was gradually heated to 300 K using 200 ps in the NVT ensemble and 200 ps in the NPT ensemble at 1 atm. Then, the NPT ensemble was further employed for 50 ns to equilibrate the system at 300 K and 1 atm, followed by a 10 ns production run for collecting equilibrated configurations at each 10 ps interval. All MD simulations were carried out with 3 replicas. The initial configurations of replica 2 and replica 3 were from the 50 ns and 60 ns of replica 1, respectively. The detailed MD system setup were summarised in Table S6.
The velocity-rescaling thermostat46 with a time constant equal to 0.1 ps was employed throughout the simulations to keep the temperature constant. To maintain the pressure, the Berendsen pressure coupling47 was used in the pre-equilibrium run, and the Parrinello-Rahman pressure coupling48,49 was used in the equilibrium and production run, with the pressure time constant and isothermal compressibilities set to 2 ps and 4.5 × 10−5 bar−1, respectively. A time step of 2 fs for integration of the equations of motion was used throughout the simulation. A cutoff of 12 Å is used for nonbonded interactions. The particle mesh Ewald algorithm50 was used to calculate long-range electrostatic interactions.
Rosetta calculations
Several specialized protocols within the Rosetta software suite were employed to evaluate the impact of pAcF mutations on protein affinity maturation and stability. The following steps outline the procedures used51,52.
pAcF parameterization
The initial structures of pAcF were generated using the molecular modeling software Avogadro53,54. These structures were then subjected to energy minimization using the Gaussian1655 software package, which optimized the geometries to ensure that the pAcF had the most stable conformations before integration into Rosetta. The minimized structures were converted into parameter files compatible with Rosetta using the MolfileToParams.py protocol. This step involved defining atom types, partial charges, bond lengths, bond angles, and torsional parameters specific to the pAcF. The accurate parameterization of pAcF was crucial for maintaining the integrity of their chemical properties during subsequent Rosetta simulations, allowing for reliable modeling of their behavior within the protein structure.
Generation of pAcF rotamer libraries
The conformational flexibility of the pAcF was modeled by generating a rotamer library, which includes possible side-chain conformations. The MakeRotLib protocol in Rosetta was employed to create the rotamer library files specific to the pAcF. The rotamer library file was then integrated into Rosetta to ensure that the side-chain conformational space of the pAcF is sampled appropriately during simulations. This process is critical for capturing the full range of interactions the pAcF side chains might engage in.
Calculation of unfolded energy for pAcF
To assess the impact of pAcF on protein stability, the Rosetta Unfolded State Energy Calculator protocol was employed. This begins with obtaining input PDB files through the PISCES server56. Subsequently, the Unfolded State Energy Calculator protocol generates a relaxed unfolded state model for the pAcF-containing protein variant. The energies of the folded and unfolded states are compared to quantify the stabilizing or destabilizing effects of pAcF mutations on protein folding.
Energy estimation with Cartesian_ddg protocol
The Cartesian_ddg protocol was utilised to estimate the energetic effects of pAcF mutations on protein stability52,57,58. Unlike traditional methods that rely on internal coordinate systems, the Cartesian_ddg protocol minimizes the energy in Cartesian space, allowing for finer adjustments to the protein’s structure. Initial structures for the Cartesian_ddg calculations were selected from frames at 50, 52, 54, 56, 58, and 60 ns of the wild-type PV2 and PV3 MD simulations. The protein structures were pre-relaxed to relieve steric clashes or strain, followed by energy minimization. ΔΔG values were computed by comparing the energy of the wild-type protein to that of the mutant in its most stable conformation. The final ΔΔG values were averaged and ranked to assess the impact of pAcF mutations on protein stability. The talaris2014_cart scoring function was employed to evaluate these energies59.
Binding free energy calculations with Flex_ddg protocol
To assess the impact of pAcF mutations on protein-protein interactions, the flex_ddg protocol60 was employed. This protocol evaluates binding free energy changes by accounting for backbone flexibility during mutations. It generates various conformations for both the wild-type and mutant complexes, calculating the binding energy for each. For these calculations, the PV2-PD-L1 and PV3-PD-L1 complexes were used. The protocol was configured with parameters including 5000 iterations for minimization, an absolute score convergence threshold of 1.0, 35000 backrub trials, and a backrub trajectory stride of 7000. The resulting ΔΔG values, reflecting the impact of the pAcF mutation on binding affinity, were derived from the average output of a generalized additive model, based on 35 independent runs of the protocol. The flexibility inherent in this protocol is particularly suited for examining mutations that might induce substantial conformational shifts in the protein structure.
Generation of recombinant human PD-L1 proteins
Recombinant human PD-L1 proteins were produced in HEK 293 cells (KaiRui Biotech) transformed with pcDNA3.1-N-GST-TEV vector. The HEK293 cells were cultured in KOP293 culture medium (KaiRui Biotech) at 37 °C, 5% CO2 and 120 r/min. Cells were transfected when the cell density reached 2 × 106/mL. The plasmid-carrier complex was prepared using KPM and TA-293 transfection reagents (KaiRui Biotech). After 24 h of culture, cell protein expression enhancer (KE 293) and nutritional additives (KT-FEED 50×) were added to increase the protein yields. After 7 days of culture, cells were centrifuged for 15 min at 4000 × g. Target GST-tagged proteins were purified from the resulting supernatants using a column packed with Glutathione Sepharose High Performance resin (Cytiva), followed by elution with a solution containing 15 mM glutathione. The final purified samples were desalted into phosphate buffer saline (PBS) for later tests and separated on a gel with a concentration of 12.5% SDS-PAGE.
Methanococcus Jannaschii tyrosyl tRNA synthetase mutation and validation
The pCNFRS-TSDV sequence was amplified utilizing the pULTRA-CNF plasmid (#48215) as a template, harboring the pCNFRS gene, with the TSDV-F/R primers (Table S7). It contains a mutation of the amino acid sequence TSDV at positions 283-286 of pCNFRS. The pET21a-sfGFP2TAG plasmid containing the sfGFP gene with a substitution TAG at the second position of amino acid. Furthermore, amplification of the pET21a-sfGFP was carried out using pET21a-sfGFP2TAG as a template along with the wtGFP-F/R primers (Table S7). The plasmids containing the gene of pCNFRS, pET21a-sfGFP and pET21a-sfGFP2TAG was extracted by SanPrep Column Plasmid Mini-Preps Kit. Subsequently, both pET21a-sfGFP2TAG and vectors expressing TSDVRS were co-transformed into competent E. coli BL21(DE3) cells. Three clones were picked and transferred into 5 mL LB liquid medium supplemented with 100 μM ampicillin sodium (Amp) and 50 μM spectinomycin dihydrochloride pentahydrate (Spe), and incubated at 37 °C, 220 rpm for 6 h in an orbital shaking incubator. Then the optical density at 600 nm (OD600) was determined, and the appropriate volume of bacterial culture (x μL, where x = 200 × 0.8/OD600) was collected by centrifugation at 4000 rpm for 2 min. The bacterial pellet was then resuspended in 800 μL of GMML liquid medium (M9 salts, 1% glycerol, 1 mM MgSO4 and 0.1 mM CaCl2 and 0.3 mM Leucine) containing Amp, Spe, IPTG (0.5 mM) and a specific ncAA (1 mM), This suspension was then transferred to 96-deep-well plates (2.2 mL) and incubated at 37 °C, 220 rpm for 11 h, with monitoring of fluorescence intensity (excitation 485 nm and emission 515 nm) as well as OD600 by Microplate Readers.
Formatting, expression and purification of nanobodies with ncaa incorporated
The insertion of ncAA is achieved by mutating the sequence to a stop codon UAG at a specific site in the protein. The wild-type nanobody PV2 and PV3 sequences (Fig. S1) were inserted between XbaI and NotI of pET-32a (+), with the His-tag fused on the C-terminal. The selected amino acid site in the nanobody was replaced with a UAG stop codon by site-directed mutagenesis for incorporating ncAA with corresponding primers (Table S5). The process of incorporating ncAA into the nanobody is achieved by transforming a plasmid expressing evolved orthogonal aminoacyl tRNA synthetase (pCNFRS-TSDV)/tRNA pairs into bacteria. The two plasmid vectors were transferred into E. coli TransB (DE3) (purchased from the Beijing TransGen Biotech, Beijing, China) and the bacteria were cultured in LB medium supplemented with ampicillin and spectinomycin. The temperature and IPTG concentration for protein expression were consistent with the above conditions, and ncAA p-acetylphenylalanine (pAcF) with a final concentration of 1 mM was added. After centrifugation, the bacterial supernatants were loaded onto a Ni Sepharose High-performance (Cytiva) column, and the proteins were eluted with 500 mM imidazole. The purified proteins were desalted into 50 mM sodium acetate buffer (pH 4.5).
Construction of bispecific nanobodies
To construct the PV2-PV3 recombinant fusion nanobody, a 15-amino acid flexible linker (GGGGS)3 was selected as the connecting peptide. The coding sequence was cloned into the pET-32a (+) plasmid vector through insertion between the XbaI and NotI. Subsequent protein expression and purification procedures were maintained under identical conditions to monovalent nanobody production.
The linkers used for constructing ncAA incorporated bispecific nanobodies were heterobifunctional chemical crosslinkers BCN (Bicyclononyne, MW 401.2287) and Tet (Tetrazine, MW 387.1), which were obtained from commercial sources and used without further purification. The concentration of the nanobody was adjusted to 1 mg/mL, and then the alkoxy group on pAcF was connected to azide group in linker (50 molar excess) through oxime ligation. After the reaction, the protein-linker compound was desalted into PBS, and then cyclooctyl group of BCN linker was connected to azide groups of TET linker through copper-free click reaction. Excess protein-linker compounds were removed by size-exclusion methods, yielding highly homogeneous conjugation products as determined by 12.5% SDS-PAGE and MALDI-TOF mass spectrometry. The chromatogram obtained from the size exclusion chromatography process revealed variations in the reaction efficiency of BsNbs constructed using different combinations of mutants. The ratio of the peak area of BsNbs to uncombined free monomers in the chromatogram was utilized as a metric for comparing the reaction efficiency of the dual antibody linkage across different groups.
Mass spectrometry characterization
Matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) was utilized for the accurate measurement of the molecular weight of mutant nanobodies following the incorporation of ncAAs. The protein sample was desalted into deionized water, and then 1 μL of the sample was mixed with 1 μL of sinapinic acid (SA) matrix solution before being applied to the sampling target. Subsequently, after drying and crystallization, MALDI-TOF mass spectrometry was employed for analysis. Data Analysis 4.1 software was used to interpret the data results.
Enzyme-linked immunosorbent assay of ncAA modified nanobody
The recombinant hPD-L1 were diluted in coating buffer (50 mM carbonate buffer, pH 9.6) to a concentration of 20 µg/mL. The diluted proteins were then applied to each well of a 96-well plate, and the plate was incubated at 4 °C overnight. Bovine serum albumin was used as negative control. The following day, after three washes with phosphate buffered saline with tween 20 (PBST), the plate was blocked for 2 hours at 37 °C in PBS containing 5% skim milk. After additional washing in PBST, nanobodies were diluted four times ten gradients starting at 400 μg/mL concentration and were added to the wells and incubated for 2.5 h at 37 °C. Following another round of washing, HRP-labelled anti-His mouse monoclonal secondary antibodies were added and incubated at 37 °C for an additional 2 h. The color reaction was initiated by adding 100 µL of 3,3’,5,5’-Tetramethylbenzidine (TMB) substrate per well. Following a 10 min dark incubation period, the reaction was terminated by adding 100 µL H2SO4 to each well, and the Varioskan LUX microplatereader (Thermo Scientific) was utilized to measure the absorbance at OD450. The obtained data were subtracted from the background values, and non-linear regression analysis was performed using Graphpad Prism software to calculate the IC50 of the curve, thereby determining the affinity of the nanobody.
Thermal stability measurements
Temperature-dependent fluorescence variation was measured using a LightCycler 480 II instrument (Roche). A total of twenty microliters was prepared, consisting of 2 µL of nanobody (20 µM), 0.4 µL of 250 mM TCEP, and 1 µL of 200×SYPRO Orange dye (Sigma-Aldrich) in PBS. The mixture was then subjected to an excitation wavelength of 465 nm and an emission wavelength of 580 nm, with a temperature gradient ranging from 25 °C to 95 °C. The acquisition mode was set to continuous with a rate of 0.01°C/s. The melting temperature (Tm) values were calculated using the LightCycler Thermal Shift Analysis software (Roche Applied Science) and Origin 8.0 software.
Surface plasmon resonance assay of ncAA modified nanobody
SPR analysis was performed using a Biacore T200 instrument (Cytiva) with the following experimental parameters. The running buffer for all SPR measurements consisted of phosphate-buffered saline (PBS, pH 7.4). Prior to analyte injection, recombinant hPD-L1 was covalently immobilized onto a CM5 sensor chip (Cytiva) through standard amine coupling chemistry. For ligand preparation, hPD-L1 protein was diluted in 10 mM sodium acetate buffer (pH 4.5) to achieve a working concentration of 5 μg/mL, resulting in a final immobilization level of 800 response units (RU) after surface activation. The biosensor surface was blocked with ethanolamine-HCL (1.0 M). Samples at concentrations ranging from 1.56 to 200 nM were injected over the chip at a 30 µL/min flow rate, with a 120 s contact time and a 900 s dissociation period. Sensorgram data were processed and analyzed using the Biacore T200 Evaluation Software (version 3.2.1) with a 1:1 Langmuir binding model to calculate apparent binding affinity (KD) values.
Cell culture
The human PD-L1 MC38 CRC cell line was obtained from Shanghai Model Organisms Center, Inc. (Shanghai, China). hPD-L1 MC38 cells were cultivated in Dulbecco’s Modified Eagle Medium containing 10% (v/v) fetal bovine serum (FBS), 50 µg/mL hygromycin B, and antibiotics (100 U/mL penicillin and 100 µg/mL streptomycin). HT29 cells (ATCC HTB-38) and THP-1 cells (ATCC TIB-202) were obtained from the American Type Culture Collection (ATCC). HT29 cells were cultured in McCoy’s 5 A medium, and THP-1 cells were maintained in RPMI-1640 medium. Both media were supplemented with 10% (v/v) FBS, 100 U/mL penicillin, and 100 µg/mL streptomycin. All cells were maintained in a humidified atmosphere at 37 °C with 5% CO2.
In vitro phagocytosis by immunofluorescence confocal microscopy
Immunofluorescence confocal microscopy was used to evaluate the efficiency of targeted blocking PD-L1/PD-1 pathway to improve the phagocytosis ability of macrophages to cancer cells. THP-1 cells in logarithmic growth phase were cultured in medium with a final concentration of 80 ng/mL PMA (Sigma-Aldrich, St. Louis, MO, USA) for 24 h, followed by transfer to PMA-free medium for another 24 h to obtain differentiated human macrophages. THP-1 cells were detached and seeded onto a transparent 12-well tissue culture plate at 5×104 cells per well. HT-29 tumor cells were incubated for 10 min with 4 µM DiD Perchlorate solution (Yeasen, China). Labeled HT-29 cells were pretreated with PBS, PV2, ncAA-nanobody, PV2-PV3 fusion nanobodies and Durvalumab served as positive controls and seeded onto 12-well culture plate at 2 × 105 cells per well (macrophage tumor cell ratio: 1:4). After 3.5 h co-incubation, residual HT-29 cells were washed and THP-1 cells were stained with Hoechst 33342 (Solarbio, China). Multiple random fields were imaged for each replicate using Laser Confocal Scanning Microscope (Leica Stellaris 8). The ratio of DiD+ THP-1 cells to total Hoechst 33342+ THP-1 cells was counted for each field to score phagocytosis.
Mice
hPD-1 mice (Cat. NO. NM-HU-00015) were purchased from Shanghai Model Organisms Center, Inc. subcutaneous graft tumor model of colorectal cancer was established by subcutaneously injecting 1.5 × 106 MC38-hPD-L1cells in a 0.2 mL PBS solution containing 10% FBS and 25% Matrigel into the right flank of hPD-1 knock-in mice (8 weeks, female). The mice bearing the MC38-hPD-L1 tumor model were acclimated at 25 °C and 55% of humidity under natural light/dark conditions, with standard mice chow and water available ad libitum. At 10 days after inoculation, tumors grew to an average of 100 mm3 (calculating formula is 1/2 × length × width2). At this time, 24 mice were randomly distributed into 4 groups. As shown in Figs. 5A, 4 groups received the corresponding drug 10 mg/kg (once every three days for six times).
The mice were fed normally after administration, and the survival status of the mice was observed, and the body weight and tumor volume of the mice were recorded as previously described. The maximal permitted tumour volume was set at 2000 mm³. Any mice that reached this limit were euthanized immediately, and this limit was not exceeded in the experiment. All animals were sacrificed 3 days after the last drug administration, followed by weighing of the tumors. The mice were bred at an animal care facility certified by the Tianjin Management Committee of Laboratory Animals in the Institute of Radiation Medicine Chinese Academy of Medical Sciences. All animal experiments were approved by the Animal Ethics Committee of the Tianjin University (Approval No. TJUE-2024-280). We have complied with all relevant ethical regulations for animal use.
Cytokine analysis
Enzyme-linked immunosorbent assay (ELISA) is used to detect the expression levels of cytokine in the serum. Mouse Interleukin 2 ELISA Kit (E-EL-M0042) and MS Mouse interferon gamma ELISA Kit (E-MSEL-M0007) (Elabscience, China) were used to detect the cytokine concentration and the absorbance was measured at OD450 using a Varioskan LUX microplate reader (Thermo Scientific).
H&E staining and immunohistochemistry analysis
The tumor tissues fixed in 4% paraformaldehyde solution were paraffinized and sectioned (5 mm thick). The paraffin sections were dewaxed in xylene and rehydrated using a graded series of ethanol. Ten, the sections were stained with hematoxylin and eosin (H&E). For immunohistochemistry, the sections were deparaffinized and rehydrated, processed with microwave antigen retrieval, and incubated overnight with primary antibodies against Ki67 (ab15580), and CD8 alpha (ab217344) (Abcam, Cambridge, UK) at 4 °C. They were further incubated with secondary antibodies and streptavidin–horseradish peroxidase with diaminobenzidine.The resulting images were captured using an Axio Observer 7 microscope (Zeiss).
Flow cytometry of tumor infiltrating immune cells
Mice were euthanized, and tumors were mechanically dissociated and digested in a mixture of enzymes (0.1% collagenase type IV, 0.01% hyaluronidase, 1% FBS, 20 U/mL DNase I dissolved in HBSS buffer) for 30 minutes at 37 °C. Primary cells were harvested, washed, and resuspended in HBSS containing 1% FBS to prepare a single-cell suspension. Separate lymphocytes using Ficoll density gradient centrifugation, stained cells with the indicated markers for flow cytometry (CytoFLEX, Beckman Coulter). The resultant single cells were stained with antibodies purchased from Biolegend (USA): PE/Cy7 anti-mouse CD45 (1:400, #103113), FITC anti-mouse CD3 (1:400, #100203), APC anti-mouse CD4 (1:400, #100411), PE anti-mouse CD8α (1:400, #100707), APC anti-mouse IFN-γ(1:100, #505809). Data were analyzed using FlowJo 10.8.1 software (TreeStar, Inc., Ashland, OR).
Statistical and reproducibility
The mean ± standard error of the mean was used to express the findings. Software called GraphPad Prism 8.4.3 was used to analyze the data, and it was calculated using an ordinary one-way ANOVA (and Nonparametric or mix). For dose-response curves, the IC50 values were determined by non-linear regression analysis. Sample size and numbers of replicates were described in detail in the figure legends. Replicates definitions were described in the legends accordingly. Statistical significance was defined as a p-value < 0.05.
Supporting information
The authors have cited additional references within the Supporting Information and provided detailed methods, protocols, additional information, results in Tables S1-S6 and supporting figures in Figs. S1-S11.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. The numerical data that make up the figures in the paper are shown in Supplementary Data 1. For MD simulations, the topology (pac.itp) and force field (pac.prm) of noncanonical amino acid pAcF are provided in the Supplementary Files. The initial coordinates for the simulation input files and the coordinate files for the final output of PDL1_WT, PV2_WT, PV2_C23pAcF, PV2_C96pAcF, and PV3_WT are provided as supplementary files. MS raw files and the corresponding results files, including databases, were deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD069482. Should any raw data files be needed in another format, they are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Eng, C. et al. A comprehensive framework for early-onset colorectal cancer research. Lancet Oncol. 23, e116–e128 (2022).
Dekker, E., Tanis, P. J., Vleugels, J. L. A., Kasi, P. M. & Wallace, M. B. Colorectal cancer. Lancet 394, 1467–1480 (2019).
Ciardiello, F. et al. Clinical management of metastatic colorectal cancer in the era of precision medicine. CA Cancer J. Clin. 72, 372–401 (2022).
Biller, L. H. & Schrag, D. Diagnosis and treatment of metastatic colorectal cancer: a review. JAMA 325, 669–685 (2021).
Song, Y., Teng, L., Chen, Y. & Dong, C. M. Glycopolypeptide coordinated nanovaccine: fabrication, characterization, and antitumor immune response. Chem. Bio Eng. 1, 633–646 (2024).
Li, J. et al. Remodeling of the immune and stromal cell compartment by PD-1 blockade in mismatch repair-deficient colorectal cancer. Cancer Cell 41, 1152–1169.e1157 (2023).
Kim, C. H. et al. Synthesis of bispecific antibodies using genetically encoded unnatural amino acids. J. Am. Chem. Soc. 134, 9918–9921 (2012).
Wu, Q. et al. Small molecule inhibitors targeting the PD-1/PD-L1 signaling pathway. Acta Pharm. Sin. 42, 1–9 (2021).
Wu, Y. et al. A highly stable human single-domain antibody-drug conjugate exhibits superior penetration and treatment of solid tumors. Mol. Ther. 30, 2785–2799 (2022).
Li, J. et al. Affinity maturation of antibody fragments: a review encompassing the development from random approaches to computational rational optimization. Int. J. Biol. Macromol. 247, 125733 (2023).
Wang, J. et al. Novel bispecific nanobody mitigates experimental intestinal inflammation in mice by targeting TNF-alpha and IL-23p19 bioactivities. Clin. Transl. Med. 14, e1636 (2024).
Wang, J. et al. Research progress and applications of multivalent, multispecific and modified nanobodies for disease treatment. Front. Immunol. 12, 838082 (2021).
Bai, Z. et al. Design of nanobody-based bispecific constructs by in silico affinity maturation and umbrella sampling simulations. Comput. Struct. Biotechnol. J. 21, 601–613 (2023).
Labrijn, A. F., Janmaat, M. L., Reichert, J. M. & Parren, P. Bispecific antibodies: a mechanistic review of the pipeline. Nat. Rev. Drug Discov. 18, 585–608 (2019).
Kontermann, R. E. Dual targeting strategies with bispecific antibodies. MAbs 4, 182–197 (2012).
Suurs, F. V., Lub-de Hooge, M. N., de Vries, E. G. E. & de Groot, D. J. A. A review of bispecific antibodies and antibody constructs in oncology and clinical challenges. Pharm. Ther. 201, 103–119 (2019).
Hutchins, B. M. et al. Site-specific coupling and sterically controlled formation of multimeric antibody Fab fragments with unnatural amino acids. J. Mol. Biol. 406, 595–603 (2011).
Schaefer, W. et al. Immunoglobulin domain crossover as a generic approach for the production of bispecific IgG antibodies. Proc. Natl. Acad. Sci. USA 108, 11187–11192 (2011).
Lewis, S. M. et al. Generation of bispecific IgG antibodies by structure-based design of an orthogonal Fab interface. Nat. Biotechnol. 32, 191–198 (2014).
Lu, H. et al. Targeting human C-type lectin-like molecule-1 (CLL1) with a bispecific antibody for immunotherapy of acute myeloid leukemia. Angew. Chem. Int. Ed. Engl. 53, 9841–9845 (2014).
Ramadoss, N. S. et al. An anti-B cell maturation antigen bispecific antibody for multiple myeloma. J. Am. Chem. Soc. 137, 5288–5291 (2015).
Li, S. et al. Nanobody against PDL1. Biotechnol. Lett. 42, 727–736 (2020).
Aldeghi, M., Gapsys, V. & de Groot, B. L. Accurate estimation of ligand binding affinity changes upon protein mutation. ACS Cent. Sci. 4, 1708–1718 (2018).
Blackman, M. L., Royzen, M. & Fox, J. M. Tetrazine ligation: fast bioconjugation based on inverse-electron-demand Diels-Alder reactivity. J. Am. Chem. Soc. 130, 13518–13519 (2008).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Adhikari, A. et al. Reprogramming natural proteins using unnatural amino acids. RSC Adv. 11, 38126–38145 (2021).
Young, D. D. et al. An evolved aminoacyl-tRNA synthetase with atypical polysubstrate specificity. Biochemistry 50, 1894–1900 (2011).
Gan, R. et al. Translation system engineering in Escherichia coli enhances non-canonical amino acid incorporation into proteins. Biotechnol. Bioeng. 114, 1074–1086 (2017).
Hartley, G. P., Chow, L., Ammons, D. T., Wheat, W. H. & Dow, S. W. Programmed cell death ligand 1 (PD-L1) signaling regulates macrophage proliferation and activation. Cancer Immunol. Res. 6, 1260–1273 (2018).
Fleetwood, A. J., Lawrence, T., Hamilton, J. A. & Cook, A. D. Granulocyte-macrophage colony-stimulating factor (CSF) and macrophage CSF-dependent macrophage phenotypes display differences in cytokine profiles and transcription factor activities: implications for CSF blockade in inflammation. J. Immunol. 178, 5245–5252 (2007).
Ayers, M. et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Investig. 127, 2930–2940 (2017).
Zhai, W. et al. Blocking of the PD-1/PD-L1 interaction by a novel cyclic peptide inhibitor for cancer immunotherapy. Sci. China Life Sci. 64, 548–562 (2021).
Peng, W. et al. PD-1 blockade enhances T-cell migration to tumors by elevating IFN-gamma inducible chemokines. Cancer Res. 72, 5209–5218 (2012).
Keyaerts, M. et al. Phase I study of 68Ga-HER2-nanobody for PET/CT assessment of HER2 expression in breast carcinoma. J. Nucl. Med. 57, 27–33 (2016).
Wu, J. The enhanced permeability and retention (EPR) effect: the significance of the concept and methods to enhance its application. J. Pers. Med. 11 (2021).
Muyldermans, S. A guide to: generation and design of nanobodies. FEBS J. 288, 2084–2102 (2020).
Kontermann, R. E. Strategies for extended serum half-life of protein therapeutics. Curr. Opin. Biotechnol. 22, 868–876 (2011).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M., Steinegger, M. & Söding, J. MMseqs2 desktop and local web server app for fast, interactive sequence searches. Bioinformatics 35, 2856–2858 (2019).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45, D170–D176 (2017).
Liu, K. et al. Structural basis of anti-PD-L1 monoclonal antibody avelumab for tumor therapy. Cell Res. 27, 151–153 (2017).
Sondergaard, C. R., Olsson, M. H., Rostkowski, M. & Jensen, J. H. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Olsson, M.H., Søndergaard, C.R., Rostkowski, M. & Jensen, J.H. PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537, (2011).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Nosé, S. & Klein, M. L. Constant pressure molecular dynamics for molecular systems. Mol. Phys. 50, 1055–1076 (2006).
Herce, H. D., Garcia, A. E. & Darden, T. The electrostatic surface term: (I) periodic systems. J. Chem. Phys. 126, 124106 (2007).
Renfrew, P. D., Choi, E. J., Bonneau, R. & Kuhlman, B. Incorporation of noncanonical amino acids into Rosetta and use in computational protein-peptide interface design. PLoS One 7, e32637 (2012).
Park, H. et al. Simultaneous optimization of biomolecular energy functions on features from small molecules and macromolecules. J. Chem. Theory Comput. 12, 6201–6212 (2016).
Hanwell, M. D. et al. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminf. 4, 17 (2012)
Avogadro: an open-source molecular builder and visualization tool. Version 1.2.0. http://avogadro.cc/.
Frisch, M. J. T. et al. 16 Rev. C.01 (Gaussian, Inc., 2016).
Wang, G. & Dunbrack, R. L. Jr. PISCESa:protein sequence culling server. Bioinformatics 19, 1589–1591 (2003).
Kellogg, E. H., Leaver-Fay, A. & Baker, D. Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins 79, 830–838 (2011).
Alford, R. F. et al. The Rosetta all-atom energy function for macromolecular modeling and design. J. Chem. Theory Comput. 13, 3031–3048 (2017).
Smith, P. & Lorenz, C. D. LiPyphilic: A Python toolkit for the analysis of lipid membrane simulations. J. Chem. Theory Comput. 17, 5907–5919 (2021).
Barlow, K. A. et al. Flex ddG: Rosetta ensemble-based estimation of changes in protein-protein binding affinity upon mutation. J. Phys. Chem. B 122, 5389–5399 (2018).
Acknowledgements
This work was supported by grants from the National Key Research and Development Project (Grant No. 2024YFA0918500), “Pioneer” and “Leading Goose” R&D Program of Zhejiang (Grant No. 2025C01097), the National Natural Science Foundation of China (Grant No. 22108245 and No. 22378351), National Key Research and Development Program of China (2025YFA0922702), the China-CEEC Joint Education Project (Grant No. 2022196), the Fundamental Research Funds for the Central Universities (Grant No. 226-2022-00214), the Science and Technology Program of Tianjin, China (Grant No. 22YFZCSN00090), and the Independent Innovation Foundation of Tianjin University (Grant No. 2023XQM-0014). The authors thank AI + High Performance Computing Center of ZJU-ICI for their support in molecular dynamics simulations.
Author information
Authors and Affiliations
Contributions
H.H., G.K., and H.Y. contributed to the initial design of the study, interpreted the data, and wrote the manuscript. X.H. and L.J. designed and conducted the research, collected and analyzed data, interpreted the results, and wrote the manuscript. L.J. contributed to the design of the computational experiments. H.Y. conducted the research and analyzed data. W.C., Y.G., M.G., Y.S., Z.W., Y.W., and M.Z. performed the research. A.D.M. refined the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Tao Liu, Qun Zhou, and the other anonymous reviewer for their contribution to the peer review of this work. Primary Handling Editors: Huan Bao and Johannes Stortz. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, X., Jiang, L., Yuan, H. et al. Noncanonical amino acid-aided synthesis of anti-PD-L1 bispecific nanobody for colon cancer immunotherapy. Commun Biol 8, 1844 (2025). https://doi.org/10.1038/s42003-025-09222-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09222-1