Abstract
Moyamoya disease (MMD) is a cerebrovascular disorder characterized by progressive intracranial arterial stenosis, which can lead to ischemic or hemorrhagic stroke. However, its pathogenesis remains poorly understood, limiting advances in the diagnosis and treatment of MMD. We perform data-independent acquisition (DIA) proteomics on serum samples from 40 MMD patients and 20 healthy controls, followed by ELISA validation of an additional 45 cohort. Vascular organoids are generated from induced pluripotent stem cells (iPSCs) derived from the peripheral blood, and analyzed using histological staining, transcriptomics, and single-cell RNA sequencing (scRNA-seq). Histological examination of the temporal superficial artery (STA) in MMD patients reveals abnormal accumulation of vascular smooth muscle cells (VSMCs). DIA proteomics identify significant upregulation of TUBA4A and TUBB4B in MMD serum. In vitro assays demonstrate that these tubulin proteins promoted VSMC proliferation, migration, and contractile-to-synthetic phenotypic switching through the GJA1/PI3K/AKT/KLF4 signaling pathway. Single-cell RNA sequencing of MMD vascular organoids shows an increased proportion of synthetic VSMCs with upregulated TUBA4A and TUBB4B expression. Our findings suggest that overexpression of TUBA4A and TUBB4B contributes to pathological vascular remodeling in MMD through the GJA1/PI3K/AKT/KLF4 pathway, as well as therapeutic targets for intervention in MMD vascular remodeling.

Similar content being viewed by others
Introduction
Moyamoya disease (MMD) is a chronic cerebrovascular disorder characterized by the progressive narrowing or occlusion of the terminal segments of the internal carotid arteries (ICAs) and abnormal neovascularization at the skull base1. It may cause severe hemorrhagic or ischemic stroke, cognitive impairment, motor and sensory dysfunction2,3 The pathogenesis of MMD is not fully understood currently. Due to this, no drug treatments have been developed to prevent the progression of MMD at the mechanistic level1,3. Surgical revascularization is worldwide considered the only treatment able to reduce the recurrence of ischemic and hemorrhagic stroke risk4. Therefore, it is essential to further elucidate the pathogenesis of MMD, to develop non-surgical, pharmacological treatment strategies and expand therapeutic options beyond revascularization.
Previous studies have demonstrated that the main pathological changes in MMD vessels are intimal thickening and abnormal abundance of smooth muscle cells (VSMC) in media; the mechanisms underlying these changes remain unclear5,6,7. As for causative genes of MMD, the Ring Finger Protein 213 (RNF213) was described as the first susceptibility gene for MMD, and has been reported to be associated with vascular defects, including pathological angiogenesis, endothelial dysfunction, and disruption of blood-brain barrier integrity8,9,10,11,12. These would be associated with intimal hyperplasia and the initiation of MMD. Several other genes were also identified as being linked to MMD, for example, Matrix Metalloproteinases (MMPs), Vascular Endothelial Growth Factor (VEGF), etc., but it remains difficult to interpret how the abnormal vessels formed in MMD through these genes alone13,14,15. Previous proteomic studies have also identified several proteins to be dysregulated in the serum or cerebrospinal fluid of MMD patients, suggesting their potential involvement in the mechanism of MMD, such as Cadherin 18 (CDH18), Serum apolipoprotein E (APOE), and Angiopoietin-2 (Ang-2)16,17,18. However, few studies have systematically investigated cytoskeletal proteins or used high-resolution methods like data-independent acquisition (DIA) to identify novel molecular signatures in MMD-related vascular pathology. This highlights the need for advanced proteomic profiling strategies to identify novel biomarkers and elucidate the molecular mechanisms underlying in MMD.
Appropriate in vivo research models can greatly assist in exploring the unknown etiology of MMD. An ideal in vivo model should reflect multiple aspects of MMD, including genetic susceptibility, vasculopathy, and features of chronic cerebral ischemia19. Current models for MMD have involved three general approaches: surgical, genetic and immunological, but these models reproduce the disparate aspects of MMD independently19,20. Surgical hypoperfusion models involving ligation of common carotid artery (CCA) or internal carotid artery (ICA) can mimic the ischemic environment of MMD, but it alone does not capture its genetic components, such as mutations in susceptibility genes like RNF213, MMP, and VEGF21,22,23. Genetic strategies that knock-out or knock-in susceptibility genes, such as RNF213, in mice can model the genetic background associated with MMD, but previous efforts have failed to reproduce the histopathological changes observed in MMD, such as abnormal abundance of VSMC, abnormal collateral networks or intimal thickening12,22,24,25. In immunological models, MMD-like features have been induced in rabbits and rats by injecting heterogeneous serum or Propionibacterium acnes, resulting in intimal thickening and disruption of the internal elastic lamina. However, these models fail to reproduce key features of MMD, such as progressive stenosis of ICA26,27. The lack of a stable and ideal model has greatly limited research on MMD. On the one hand, the lack of a stable and reliable animal model has significantly hindered mechanistic studies of MMD, as existing models fail to simultaneously replicate both the characteristic vascular pathology and the genetic background associated with MMD. On the other hand, the absence of such a model also limits the development and validation of potential therapeutic interventions, thereby impeding progress in the advancement of effective treatment strategies for MMD. Therefore, there is an urgent need for a reliable disease model to investigate the pathogenesis of MMD and to facilitate the development of effective therapeutic strategies.
Organoids, the in vitro 3D tissues obtained by stem cell culture and differentiation, have the capability to exhibit the anatomical and functional characteristics of real organs, and can present features relevant to human diseases, and thus be applied as models in the study of disease pathogenesis28. So far, a variety of organoid-based disease models have been developed for genetic diseases, host-pathogen interactions, or cancers, which appropriately demonstrate known pathological characteristics28,29,30,31. In the field of vascular organoids, Wimmer et al. first developed a vascular organoid model by differentiating pluripotent stem cells (PSCs), which successfully recapitulated vascular pathological changes under hyperglycemic conditions associated with diabetes32. Lan et al. modeled a human blood-brain barrier assembloid using brain and vascular organoids derived from human PSCs, and applied it for the study of cerebral cavernous malformations33. In addition, vascular organoid models have also been applied to the study of diseases such as Alzheimer’s disease, cerebrovascular malformations, and acute liver failure34,35,36. As for MMD research, although no vascular organoid models have been specifically developed for MMD currently, the vascular organoid model holds great potential for investigating the structural and functional alterations of affected vessels and exploring the pathophysiological mechanisms involved. Therefore, vascular organoid models can serve as a valuable complement to existing MMD research models.
Despite increasing insights into the genetic and pathological features of MMD, the molecular mechanisms underlying its vascular abnormalities remain unclear, and suitable human-based experimental models are still lacking. In this study, we aimed to establish a patient iPSCs-derived vascular organoid model recapitulating the pathological features and genetic background of MMD. In parallel, we aimed to identify novel molecular factors involved in MMD-associated vascular abnormalities through combining vascular organoid, single-cell transcriptomic analysis, and DIA-based proteomics. Finally, we established a vascular organoid model to enable mechanistic exploration and facilitate future therapeutic discovery. Through in vitro experiments and the organoid platform, we investigated how dysregulated cytoskeletal proteins, α-tubulin 4A (TUBA4A) and β-tubulin 4B (TUBB4B), contribute to VSMC migration, phenotypic switching, and proliferation in MMD.
Methods
Patient enrollment and subgrouping
Patients examined between January 2022 and June 2024 with MMD were clinically diagnosed by medical history and Digital Subtraction Angiography examination following the diagnostic guidelines37. All the patients had sporadic MMD with no family history of MMD among first-degree relatives. Clinical information, including age, sex, history of hypertension, diabetes, coronary artery disease, hyperlipidemia, smoking, alcohol intake, symptom duration, and Suzuki stage, was collected from electronic medical records and Picture Archiving and Communication System (PACS) reports. Medical history was also reviewed to exclude participants with autoimmune diseases, systemic infections, malignancies, or prior cerebrovascular surgical interventions that might confound the interpretation of vascular pathology. Patients diagnosed with MMD were classified into two subgroups: adult patients with hemorrhagic history (HEM) and adult patients with ischemic history (IS). Healthy adult volunteers without MMD or any underlying diseases were enrolled as healthy controls (HC).
Among the enrolled MMD patients, the selection criteria for individuals used in iPSC generation and vascular organoid construction were as follows: (1) age around 40 years, corresponding to the second peak in MMD’s two-peak age distribution (two peaks around age 10 and 401); (2) inclusion of both sexes; (3) clinical diversity, including both IS-type and HEM-typr cases; (4) symptom duration between 6 months and 2 years with moderate to moderately severe clinical presentations; (5) sporadic MMD with no family history; (6) Suzuki stage III to V on angiography of the affected hemisphere; and (7) absence of major comorbidities, including hypertension, diabetes, coronary artery disease, and hyperlipidemia, as well as no history of smoking or alcohol intake. These criteria were applied to ensure clinical representativeness and to avoid the potential influence of other disease conditions on model construction. All human tissue samples were obtained with written informed consent from the patients. All ethical regulations relevant to human research participants were followed. This study and the involved protocols were approved by the Institutional Ethics Committee of Beijing Tiantan Hospital, Beijing, China (KY2023-204-02).
Finally, A total of 73 patients diagnosed with MMD between January 2020 and August 2022 were enrolled in this study. Peripheral blood samples were taken from all 73 patients. Among them, DIA proteomics were performed on the peripheral blood of 40 MMD patients, including 20 ischemic (IS) and 20 hemorrhagic (HEM) MMD. We enrolled 20 age- and sex-matched HC as controls. Three MMD patients were selected according to the inclusion criteria described above, and 3 HC individuals were also selected whose peripheral blood was used to generate iPSCs and subsequently construct vascular organoids. The 3 patients included one male and two females, with two IS-type and one HEM-type case. All were sporadic cases without comorbidities, aged 31, 41, and 45 years, with symptom durations ranging from 6 months to 2 years and Suzuki stage III–IV on the affected side. Detailed clinical information of the 3 patients is provided in Supplementary Table 1. The remaining 30 patients (including 15 IS and 15 HEM) were selected as a validation group to validate the protein expression observed in the DIA discovery cohort via ELISA experiments from their serum. We also enrolled 15 age- and sex-matched HC as controls in the validation group. Detailed information on all patients can be found in the supplementary materials. (Supplementary Table 1).
Serum sample collection for DIA proteomics and ELISA
Peripheral blood samples for DIA proteomic and ELISA experiments were collected preoperatively via venipuncture, preferably from the cubital or forearm vein to minimize the risk of hemolysis. CPT Vacutainer containing sodium citrate as an anticoagulant were used for collection. Approximately 5 mL of blood was collected from each participant. After sampling, the blood was allowed to clot at room temperature for 30 to 60 minutes. The samples were then centrifuged at 3000 × g for 10 minutes at 4°C, and the upper serum layer was carefully transferred into new tubes using sterile pipettes, ensuring that no red blood cells or clot debris were collected. For short-term storage, serum samples were kept at 4°C and processed within 24 hours. For long-term storage, serum was aliquoted into nuclease-free cryovials and stored at −80°C to prevent protein degradation and maintain sample integrity. All samples were labeled with patient ID, collection time, and processing time, and hemolyzed samples were excluded to ensure reliable proteomic analysis and robust experimental results.
Peripheral blood sample collection for vascular organoid modeling
Peripheral blood samples for vascular organoid construction were collected via venipuncture, preferably from the cubital or forearm vein to minimize the risk of hemolysis. CPT Vacutainer containing sodium citrate as an anticoagulant was used for collection. Approximately 10 mL of blood was collected from each participant. To maintain the viability of cells in the blood, the sample should be stored at 37 °C, and iPSC induction must be initiated within 6 hours.
STA tissue sample collection for immunofluorescence
As for the collection of superficial temporal artery (STA) specimens, this process was performed by experienced surgeons during bypass surgery for MMD patients. During surgery for MMD patients, trimming of the superficial temporal artery (STA) is typically performed to ensure a precise anastomosis with the middle cerebral artery (MCA). This trimming is a common surgical measure aimed at improving the patient’s outcome and poses no harm to the patient. Each time, 1–3 mm segments of the STA are excised. Control samples of STA are obtained from patients undergoing craniotomy for conditions such as epilepsy or trauma, without cerebrovascular disease. These surgeries often require passing through the region near STA during the surgical approach. To ensure the procedure’s success, such as achieving adequate surgical exposure, the STA must be transected. Since the distal segment of the transected STA loses its blood supply and no longer serves a functional role for the patient, the surgeon excises a 1–3 mm segment of the vessel during the procedure for use as a sample. The samples are preserved in liquid nitrogen immediately after excision to maintain their structural and molecular integrity for subsequent staining and analysis. All the protocols of sample collection were approved by the Institutional Ethics Committee of Beijing Tiantan Hospital, Beijing, China (KY2023-204-02).
DIA proteomic profile sequencing and analysis
Data-independent acquisition (DIA) proteomics was used to quantify serum protein expression in patients across the HEM, IS, CMD, and HC groups. The DIA proteomics detecting methods were mentioned in our previous study38. Briefly, serum proteins were denatured and reduced using dithiothreitol (DTT), followed by alkylation with iodoacetamide. After enrichment using SPE C18 columns, samples were redissolved in ammonium bicarbonate buffer, and protein concentrations were determined by Bradford assay. Protein digestion was performed with trypsin at a 1:20 (w/w) enzyme-to-protein ratio for 14–16 hours at 37 °C. For high-pH reversed-phase (RP) separation, equal amounts of peptides were pooled and separated using a Shimadzu LC-20AB HPLC system with a Gemini C18 column. Peptides were then ionized via nanoESI and analyzed using an Orbitrap Exploris 480 mass spectrometer (Thermo Fisher Scientific) for DIA quantification and spectral library generation.
For bioinformatics analysis, DDA data were processed with MaxQuant using the Andromeda search engine, and DIA quality control was handled with the mProphet algorithm. GO, KEGG, and pathway enrichment analyses were conducted to identify functional annotations. Differential protein expression was explored using protein-protein interaction maps via STRING (v3.2.1). Bioinformatics analyses were performed in R (v3.4). DIA analysis services were provided by the Beijing Genomics Institute (BGI, Shenzhen, China).
Vascular organoids generation
Peripheral venous blood (8 mL) from MMD patients was collected into anticoagulant tubes and maintained at 37°C. PBMCs were isolated using Ficoll density gradient centrifugation. Whole blood was diluted 1:1 with saline, gently mixed, and layered onto the separation medium (separation medium: diluted blood = 1:2). After centrifugation at 800 g for 20–30 minutes at room temperature, the PBMC layer was carefully aspirated and transferred to a 15 mL centrifuge tube. Cells were resuspended in 10 mL of dilution buffer and centrifuged at 250 g for 10 minutes at room temperature. This washing step was repeated 1–2 times.
PBMC derived from peripheral blood were reprogrammed into iPSC lines using the CytoTune 2.0 iPS Sendai Virus Reprogramming Kit (ThermoFisher). On Day -4, peripheral blood mononuclear cells (PBMCs) were seeded at a density of 5 × 10⁵ cells/mL in the center of a 24-well plate with complete PBMC culture medium. From Day −3 to Day −1, half of the medium was replaced daily with 0.5 mL of fresh PBMC culture medium. On Day 0, cells were transduced with CytoTune™ 2.0 Sendai reprogramming vectors and incubated overnight. The following day, the medium was replaced to remove the vectors. On Day 3, the transduced cells were plated on rhVTN-N coated dishes in StemPro™-34 medium without cytokines. From Day 4 to Day 6, the medium was replaced every other day. On Day 7, the transition to iPSC medium began by replacing half of the StemPro™-34 medium with complete iPSC medium. On Day 8, the medium was fully replaced with iPSC medium, and the cells were cultured on Matrigel-coated dishes. From Day 9 to Day 28, the medium was replaced daily, and iPSC colonies were monitored. Once iPSC colonies were ready, live staining was performed, and undifferentiated iPSCs were picked and transferred to fresh Matrigel-coated dishes for expansion and purification.
IPSCs were differentiated into vascular organoids following the methods reported in previous study39. IPSCs were first cultured in mTeSR1 medium on Matrigel-coated plates until they reached 80–90% confluency. The cells were then dissociated into single cells using Accutase, resuspended in aggregation medium, and seeded at a density of 2 × 10^5 cells per well in low-attachment 6-well plates containing 50 μM Y-27632. The cells were incubated for 1 day to form smooth aggregates. On Day 0, the aggregates were transferred to a new medium (N2B27 Medium supplemented with 12 μM CHIR99021 and 30 ng/mL BMP-4) and cultured for 3 days to induce mesoderm differentiation. The medium was gently pipetted daily to prevent clumping. On Day 3, the aggregates were transferred to N2B27 Medium supplemented with 100 ng/mL VEGF-A and 2 µM forskolin and cultured for 2 days. The cell aggregates were then embedded in a two-layer Collagen I-Matrigel mixture in a 12-well plate (0.5 mL for layer 1 and 0.5 mL for layer 2) and incubated at 37 °C to solidify each layer. After embedding, StemPro-34 SFM medium with 15% FBS, 100 ng/mL VEGF-A, and 100 ng/mL FGF-2 was added. The organoids were cultured for 1-3 days, with medium changes every other day, to induce vascular differentiation and sprouting. At day 10, the vascular networks were formed and extracted from the Collagen I-Matrigel to a 96-well ultra-low-attachment plate to form VOs.
Transcriptomics sequencing
Total RNA was extracted from 10 organoids per sample using the RNAprep Pure Micro Kit (DP420, Tiangen Biotech), following the manufacturer’s instructions. On-column DNase I digestion was performed to eliminate residual genomic DNA contamination. The concentration and integrity of the extracted RNA were assessed using a NanoDrop spectrophotometer (Thermo Fisher) and Agilent 2100 Bioanalyzer (Agilent Technologies), with RNA Integrity Number (RIN) values > 7 deemed suitable for library construction. RNA libraries were prepared using the NEBNext Ultra Directional RNA Library Prep Kit for Illumina, following the standard protocol, which included mRNA enrichment, RNA fragmentation, cDNA synthesis, adapter ligation, and PCR amplification. Quality control was performed at each step, with the final libraries evaluated using a Qubit fluorometer (Thermo Fisher) and an Agilent 2100 Bioanalyzer. Sequencing was conducted on the Illumina NovaSeq 6000 platform using a paired-end 150 bp configuration, generating approximately 20 million reads per sample to ensure comprehensive transcriptome coverage. After sequencing, raw data underwent multiple quality control steps: (1) sequencing error rate distribution checks to assess accuracy, ensuring the average error rate was below 1%; (2) GC content distribution checks to detect potential biases or contamination, retaining only samples with normal distributions; and (3) raw data filtering to remove reads containing adapter sequences, over 10% unknown bases (N), or more than 50% low-quality bases (Q-score < 20), yielding high-quality clean reads for downstream analyses. Differentially expressed genes (DEGs) were identified using thresholds of FDR < 0.01 and log2FC > 1. All bioinformatics analyses, including alignment, normalization, and DEG identification, were conducted by Novogene Bioinformatics Technology Co., Ltd. (Beijing, China).
For bioinformatics analysis, DEGs were analyzed using the clusterProfiler package in R (v4.1.2), which performed both GO and KEGG enrichment analyses with adjusted P-values (FDR < 0.05) to account for multiple comparisons. Significantly enriched terms and pathways were visualized using bar plots and bubble plots, highlighting key biological processes and pathways potentially relevant to the experimental conditions. For both GO and KEGG analyses, statistical significance was determined using adjusted P-values (FDR < 0.05) to control for multiple testing. Specifically, pathways or terms with P < 0.05 after adjustment were considered significantly enriched.
Single-cell RNA sequencing
For each sample, 20 organoids were collected and washed with cold PBS to remove any residual culture media. The organoids were dissociated into single cells using enzymatic digestion with collagenase and trypsin at optimized concentrations, followed by gentle pipetting to achieve a uniform single-cell suspension. The suspension was passed through a 40 µm cell strainer to remove undigested tissue. Cell viability was assessed using Trypan Blue exclusion, and only samples with viability exceeding 85% were processed further to ensure high-quality data. For each sample, 10,000 viable single cells were collected and suspended in PBS containing 0.04% BSA to prevent clumping. The single-cell suspension was then loaded onto the Chromium Single Cell Controller (10x Genomics) to encapsulate individual cells into gel bead-in-emulsions (GEMs), with each cell uniquely barcoded to enable single-cell gene expression tracking. cDNA libraries were prepared using the Single Cell 3’ Library & Gel Bead Kit v3 (10x Genomics) following the manufacturer’s instructions, including cell lysis, reverse transcription, cDNA amplification, and library preparation. Quality control was conducted using a Qubit fluorometer and Agilent 2100 Bioanalyzer to ensure proper concentration and fragment size distribution. Sequencing was performed on a 10x Genomics platform at Novogene Co., Ltd. (Beijing, China), generating high-throughput paired-end reads. The raw sequencing data were processed using the 10x Genomics bioinformatics pipeline, which included demultiplexing, barcode processing, alignment, and quantification of gene expression at the single-cell level. Raw sequencing data were processed using CellRanger (v6.1.2, 10x Genomics) with the hg38 reference genome. FASTQ files were demultiplexed and aligned using the default pipeline. Gene expression matrices were generated based on cell barcodes and unique molecular identifiers (UMIs).
For bioinformatics analysis, sc-RNAseq data were processed and analyzed using the Seurat package (v4.3.0) in R (v4.1.2). Cells were first filtered based on quality control metrics, including the number of detected genes, total UMI counts, and mitochondrial gene content, to remove low-quality cells or doublets. Cells were retained if they met all the following criteria: more than 200 detected genes and fewer than 6000 genes per cell, mitochondrial gene UMI proportion below 15%, and hemoglobin gene UMI proportion below 5%. Normalization and scaling were performed, followed by principal component analysis (PCA) to reduce dimensionality. Cell clustering was conducted using the Louvain algorithm based on the top principal components, and clusters were visualized using Uniform Manifold Approximation and Projection (UMAP). Identified clusters were annotated according to known marker genes. For differential gene expression analysis between the same cell type across different sample groups, the FindMarkers function in Seurat was used with the Wilcoxon rank-sum test. Genes with adjusted P-values (FDR) < 0.05 and log2 fold change (log2FC) > 0.25 were considered significantly differentially expressed. Cell-cell interaction analysis was conducted using the CellChat package (v1.5.0), which infers intercellular communication networks based on known ligand-receptor pairs. Significant interactions were identified with P-values < 0.05. Network visualization and pathway enrichment analysis provided insights into key signaling pathways involved in cellular communication across clusters and conditions. The custom codes used for data analysis are provided in the Supplementary Materials (Supplementary Data 3).
Plasmid, Small hairpin RNA and lentiviral construction
Specific primers were designed targeting the TUBA4A and TUBB4B sequences. Based on the multiple cloning sites of the pcDNA3.1 vector, Xho I (CTCGAG) and Xba I (TCTAGA) restriction sites were added to the upstream and downstream of the primers, respectively. The PCR product containing the Xho I and Xba I sites was then purified and recovered. The CDS sequences of human TUBA4A and TUBB4B were amplified by PCR using the following primers: TUBA4A forward, 5′- CCGCTCGAGTGAGACCTGTCACCCCGACT-3′; TUBA4A reverse, 5′- TGCTCTAGATAATGGCACAGCCCCAGCTC -3′; TUBB4B forward, 5′- CCGCTCGAGGTTTGCACCTCGCTGCTCCA -3′; and TUBB4B reverse, 5′- TGCTCTAGAGCTCTTGGGGCGATGTCATC -3′.
As for (Gap junction A1) GJA1 Lentivirus, based on the multiple cloning sites of the pLVX-Puro vector, Xho I (CTCGAG) and BamH I (GGATCC) restriction sites were added to the upstream and downstream of the primers, respectively. The PCR product containing the Xho I and BamH I sites was then purified and recovered. The CDS sequences of human GJA1 were amplified by PCR using the following primers: GJA1 forward, 5′- CCGCTCGAGTTTCATTAGGGGGAAGGCGT-3′; GJA1 reverse, 5′- CGCGGATCCCTCCAGAACACATGATCTGATGG -3′. The GJA1 plasmids were transfected into 293 T cells. The supernatants were collected and concentrated to construct the lentiviruses.
ShRNA plasmids for targeted silencing of TUBA4A (sh-TUBA4A), TUBB4B (sh-TUBB4B), and GJA1 (sh-GJA1), and the control non-targeting plasmid (sh-NC) were constructed by inserting the following short hairpin sequences into the PLKO.1-puro vector: 5′- CCGGGCTCTCTGTTGACTATGGCAACTCGAGTTGCCATAGTCAACAGAGAGCTTTTTG - 3′ for sh-TUBA4A, 5′- CCGGCGCATCTCTGTGTACTACAATCT CGAGATTGTAGTACACAGAGATGCGTTTTTG -3′ for sh-TUBB4B, 5′- CCGGGCCCAAACTGATGGTGTCAATCTCGAGATTGACACCATCAGTTTGGGCTTTTTG - 3′ for sh-GJA1, and 5′-CCGGCAACAAGATGAAGAGCACCAAC TCGAGTTGGTGCTCTTCATCTTGTTGTTTTTG -3′ for sh-NC.
All recombinant plasmids were confirmed by Sanger sequencing, and only plasmids with completely verified sequences were used in downstream transfection experiments.
Immunofluorescence staining
Immunofluorescence (IF) staining was performed on vascular organoids following the protocol previously described in the literature39. As for STA tissue samples, STA tissue samples were fixed in 4% paraformaldehyde at room temperature for 24 hours, followed by embedding in paraffin and sectioning into 4 µm thick slices. Sections were deparaffinized in xylene and rehydrated through graded alcohols. Antigen retrieval was performed by heating the sections in citrate buffer (pH 6.0) at 95 °C for 20 minutes. After cooling to room temperature, sections were blocked with 5% bovine serum albumin (BSA) in PBS for 1 hour to prevent non-specific binding. The tissue sections were then incubated with fluorophore-conjugated primary antibodies at 4 °C overnight. The following day, sections were washed three times with PBS. After washing, nuclei were counterstained with DAPI, and the sections were mounted with antifade mounting medium. Fluorescent images were captured using a confocal microscope at appropriate magnifications. The antibodies and their concentration used were listed in the supplementary materials (Table S2).
Enzyme-linked immunosorbent assay
Enzyme-linked immunosorbent assays (ELISA) were used to measure human TUBA4A and TUBB4B protein in serum samples using Human Tubulin Alpha-4A ELISA Kit (EKL61314, Biomatik, Canada) and Human Tubulin Beta-4B Chain ELISA Kit (Human) (abx548829, Abbexa, USA). The human VEGF, Ang-2, ApoE proteins in in serum samples were measured using Human VEGF ELISA Kit (DVE00, R&D Systems, USA), Human Ang-2 ELISA Kit (DANG20, R&D Systems, USA), Human ApoE ELISA Kit (HUES01675, AssayGenie, Ireland).
Cell culture and treatment
The human brain vascular smooth muscle cells (HBVSMCs) were purchased from ScienCell Research Laboratories (Hubei, China). The culture conditions for HBVSMCs were SMCM medium containing 2% FBS, 1% P/S double antibody and 1% smooth muscle cell growth supplement (SMCGS). The HBVSMC cells were incubated in DMEM containing 2% FBS for 24 h, followed by incubation in DMEM medium containing 2.5% heat-inactivated human serum from MMD patients and HC for an additional 24 h.
Plasmid, Small hairpin RNA and Lentiviral Construction
Plasmids encoding TUBA4A, TUBB4B and GJA1 were purchased from the Boen Company (Guangzhou, China). ShRNAs against TUBA4A, TUBB4B and GJA1, and the corresponding controls were obtained from Boen Company (Guangzhou, China). HBVSMCs cells were transfected with various types of lentiviruses at a multiplicity of infection of 10, and 5 µg/ml puromycin was used for selection for 2 weeks to obtain stably transfected cell lines. PCR and western blotting assays were used to verify the transfection efficiency of the lentiviruses. The detailed procedures for plasmid, shRNA, and lentivirus construction can be found in the supplementary materials.
Cell proliferation assay
Cell proliferation abilities of HBVSMCs were evaluated using a Click-iT™ EdU imaging kit (C0085L, Beyotime biotechnology) according to the protocols of the manufacturer. The cells were observed and imaged under a fluorescence microscope using red (Ex/Em = 495/519 nm) filters.
Wound-healing assay
The Wound-Healing Assay was used to analyze the migration ability of HBVSMCs. The treated HBVSMC cells were seeded into 6-well plates (5 × 105 cells per well) in DMEM containing 10% FBS and maintained at 37°C in 5% CO2 overnight. Subsequently, a 200 μl pipette tip was used to create scratches across the cell monolayer, followed by washing the cells three times with PBS. Each well was then supplemented with 2 ml of DMEM and incubated for 48 h. Microscopic images were captured at 200× magnification.
Western blot
The treated cells were resuspended and seeded in 6-well culture plates (5 × 105 cells per well). After the cells were attached to the wall, whole-cell lysates were prepared using RIPA buffer. The protein concentration was determined using a Pierce BCA Protein Assay kit (Thermo Fisher Scientific). After electrophoresis, electrotransfer, and incubation with the primary and secondary antibodies, signals were detected using Novex ECL HRP chemiluminescent substrate reagent kit (WP20005, Thermo Fisher Scientific). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the loading control. The primary antibodies and their concentration used were listed in the supplementary materials (Table S2).
Flow cytometric cell cycle analysis
The treated HBVSMC cells were seeded into 6-well plates (5 × 105 cells per well) in DMEM containing 10% FBS and maintained at 37 °C in 5% CO2 overnight. The cells were collected, washed with PBS, and the concentration was adjusted to 1 × 106 cells/ml. Single-cell suspensions of cells were fixed in 70% ethanol at 4 °C overnight. The cell cycle stage of HBVSMC cells was determined using a cell cycle assay kit (KGA512, KeyGEN Biotech, NanJing, China). The samples were then analyzed using a flow cytometer (Beckman DxFlex, Beckman) at 488 nm excitation. The flow cytometry gating strategy is shown in Supplementary Fig. 9. Intact cells were first selected based on FSC-A vs. SSC-A parameters (R1), followed by exclusion of doublets using FSC-A vs. FSC-H (R2). Viable cells were then defined as PI-negative in the FL7-A channel, with the threshold set using unstained controls. The same gating strategy was applied to all samples. The data were analyzed using FlowJo software (FlowJo, LLC, Ashland, OR, USA). The percentages of cells in the G1, S, and G2 phases were calculated, respectively.
Statistics and reproducibility
All experiments were performed with three independent biological replicates for each group (n = 3) unless otherwise stated. For ELISA assays, fifteen independent biological replicates were performed for each group (n = 15). Experimental values are presented as means ± standard deviations (SDs). Statistical significance was assessed using two-tailed non-parametric tests: the independent sample Student’s t-test for two-group comparisons and the one-way analysis of variance (ANOVA) followed by Tukey’s test for three or more groups. A P-value < 0.05 was considered statistically significant. All analyses were conducted using GraphPad Prism 9 (Version 9.4.0) and organized with Adobe Illustrator 2022 (Version 2022).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
DIA proteomic analysis reveals the upregulation of TUBA4A and TUBB4B proteins in the serum of MMD
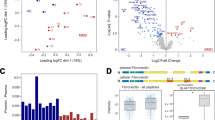
We systematically analyzed the DIA proteomics data of the serum from 40 MMD patients and 20 HC reported in our previous study38. Differential protein analysis revealed 5 members of the tubulin protein family upregulated in the MMD serum proteome, including TUBA1A (Log2FC (MMD/HC) = 1.71, p = 0.011), TUBA4A (Log2FC (MMD/HC) = 2.70, p = 0.001), TUBB5 (Log2FC (MMD/HC) = 1.16, p = 0.021), TUBB1 (Log2FC (MMD/HC) = 1.96, p = 0.006), and TUBB4B (Log2FC (MMD/HC) = 2.10, p = 0.003) (Supplementary Fig. 1A, B). The chordal graph shows that these proteins have a strong correlation with cytoskeleton-associated pathways, including cytoskeletal organization and cadherin binding (Supplementary Fig. 1C). Given the known relevance of cytoskeletal abnormalities in the pathogenesis of MMD, we selected the two most prominently upregulated tubulin proteins with the highest Log2FC values, TUBA4A and TUBB4B, for further validation and mechanistic investigation. To validate these proteomic findings, ELISA was performed on an independent validation cohort comprising 45 individuals (30 MMD and 15 HC), which confirmed the significant upregulation of proteins TUBA4A and TUBB4B in the serum of MMD patients (Fig. 1A–C). Meanwhile, several proteins associated with MMD mentioned in previous studies have also been detected in our proteomics database13,18, including CDH18 (Log2FC (MMD/HC) = 2.33, p = 0.001), MMP8 (Log2FC (MMD/HC) = 1.53, p = 0.001), and MMP9 (Log2FC (MMD/HC) =-2.70, p = 0.001). The Supplementary Data 1 listed all differently expressed proteins from the DIA proteomics database. In addition, we randomly selected serum from 3 MMD and 3 HC samples from the DIA cohort and performed ELISA for VEGF, ANG-2, and APOE as referenced in prior studies14,16,17. Compared to HC, the expression levels of VEGF, ANG-2, and APOE proteins in MMD serum were all elevated (Supplementary Fig. 1D–F). Quantitative results and source data for all experiments are provided in Supplementary Data 2.
A ELISA validation of differentially expressed proteins identified in serum proteomics. B The bar chart showing the expression levels of TUBA4A in the serum validation cohort (mean ± SD, n = 15 biologically independent samples). C The bar chart showing the expression levels of TUBB4B in the serum validation cohort (mean ± SD, n = 15 biologically independent samples). D Schematic diagram of the cell model construction for TUBA4A/TUBB4B overexpression and GJA1 knockdown in HBVSMCs, and the co-culture of smooth muscle cell (SMC) and endothelial cell (EC). E 5-Ethynyl-2′-deoxyuridine (EDU) staining of the TUBA4A/TUBB4B overexpression HBVSMC. Colors: EDU (red), 4′,6-diamidino-2-phenylindole (DAPI) (blue). Scale = 20 µm. The bar chart of H shows the proportion of proliferation HBVSMC (mean ± SD, n = 3 biologically independent samples). F Scratch assay showing the migration ability of GJA1 knockdown HBVSMC. Scale = 200 µm. The bar chart of I shows the proportion of migration HBVSMC (mean ± SD, n = 3 biologically independent samples). G Flow cytometric cell cycle analysis for the TUBA4B/TUBB4B overexpression HBVSMC after GJA1 knockdown. The bar chart of J shows the proportion of HBVSMC in G0/G1, S, and G2/M cell cycle phases (mean ± SD, n = 3 biologically independent samples).
VSMCs exhibit enhanced migration and proliferation, and a tendency of contractile-to-synthetic phenotypic switching after MMD-serum treatment
Immunofluorescence (IF) staining was performed on the superficial temporal artery (STA) from MMD patients and non-MMD controls. The walls of the STA in MMD patients exhibit a significant intimal thickening, in which an abundant presence of VSMCs was observed (Fig. 2A, B). It demonstrates the overproliferation and migration of VSMCs in the vessel wall of MMD.
A Schematic diagram illustrating the collection of STA specimens during surgery and the intimal thickening in MMD arteries. B Immunofluorescence staining of the STA in control subjects and MMD patients. Scale = 200 μm. αSMA, green; DAPI, blue. C Schematic diagram of the underlying mechanism for intimal thickening in MMD. STA = Superficial Temporal Artery.
Serum collected from patients in the MMD and HC groups was utilized to incubate HBVSMCs to explore the effect of MMD patient (IS and HEM patients) serum on VSMC (Supplementary Fig. 2A). After 24 hours of serum incubation, the MMD serum-treated VSMCs exhibited significantly promoted abilities of proliferation (Supplementary Fig. 2B, G). The scratch assay showed that VSMCs incubated with MMD serum had an enhanced migration capacity compared to those incubated with HC serum (Supplementary Fig. 2C, H). The western blot showed that the contractile phenotype biomarkers in MMD serum-treated VSMCs, CNN1, and SM22α, were downregulated compared to HC-serum-treated VSMCs, while the synthetic phenotype biomarkers, S100A4 and ZO-2, were upregulated (Supplementary Fig. 2D, I). It showed that the contractile-to-synthetic phenotypic switching occurred in VSMCs, followed by MMD serum treatment. Taken together, these findings demonstrated that factors present in MMD patient serum can induce increased migration, phenotypic switching, and proliferation of VSMCs under in vitro conditions. Considering our previous proteomic data identifying elevated levels of TUBA4A and TUBB4B in MMD serum, we next sought to determine whether these proteins may functionally contribute to the observed cellular changes of VSMCs.
TUBA4A and TUBB4B promote the abilities of migration and proliferation, and induce contractile-to-synthetic phenotypic switching of VSMCs in MMD
Consistent with the results of the DIA proteomic sequencing, increased expression of TUBA4A and TUBB4B was also observed in the MMD serum-treated HBVSMC (Supplementary Fig. 2E, J). To investigate their direct role in MMD-associated VSMC abnormalities, overexpression and knockdown for the two genes were performed using an overexpressing plasmid and small hairpin RNAs (shRNA) in VSMCs, respectively (Supplementary Fig. 3A, 4A, 4K, L). VSMCs overexpressing TUBA4A or TUBB4B showed significantly increased EdU incorporation rates and a higher proportion of S-phase cells compared to control vector-transfected cells, indicating enhanced proliferative capacity (Fig. 1E, Supplementary Fig. 3D–F, Supplementary Fig. 4B, C). Scratch assays demonstrated greater closure rates in VSMCs overexpressing the two genes, reflecting an enhanced migratory activity (Supplementary Fig. 3C). Western blot analysis revealed that overexpression of TUBA4A or TUBB4B resulted in decreased levels of contractile phenotype markers (CNN1, SM22α) and increased levels of synthetic markers (S100A4, ZO-2) (Supplementary Fig. 3B, 3H). Conversely, knockdown of either two gene reversed these effects in the MMD serum-treated VSMCs (Supplementary Fig. 3B–D, 3G, 3I, 4D, E). These results demonstrated that TUBA4A and TUBB4B contribute to key changes in VSMCs under MMD-related conditions, including migration, contractile-to-synthetic phenotypic switching and proliferation.
TUBA4A and TUBB4B induce VSMC changes via GJA1-related pathway
In HBVSMCs treated with MMD serum and overexpressing TUBA4A or TUBB4B, elevated expression of GJA1 was observed, suggesting potential involvement of this gene in the downstream signaling cascade (Supplementary Fig. 2F, 2K, 3E, 3J). To determine the functional role of GJA1 in mediating these cellular changes, we utilized overexpression plasmids and shRNA to upregulate or downregulate the level of GJA1, respectively (Supplementary Fig. 5A, 4F–G, 4M, N). Overexpression of GJA1 in VSMCs led to enhanced migration ability, increased proportion of proliferating cells, and promotion of the contractile-to-synthetic phenotypic switching (Supplementary Fig. 5C–F). Conversely, knockdown of GJA1 in VSMCs overexpressing TUBA4A or TUBB4B attenuated these phenotypic changes, including reversal of the elevated migration, proliferation, and contractile-to-synthetic phenotypic switching (Fig. 1D, F–J, Supplementary Fig. 4H, I, Supplementary Fig. 4O, P, Supplementary Fig. 4J, Supplementary Fig. 4Q). Collectively, these results demonstrated that TUBA4A and TUBB4B promote the migration, contractile-to-synthetic phenotypic switching, and proliferation of VSMCs in MMD through activation of GJA1.
MMD-derived vascular organoids exhibit an extensive and densely branched vascular network
We collected the peripheral blood from 3 MMD patients and 3 HC to generate a vascular organoid model, according to the previous protocols39,40,41. The overview of the process for developing the vascular organoids was shown in Fig. 3A. Figure 3B shows the morphological progression of the various stages from iPSC aggregates to vascular organoids under light microscopy. To evaluate the vascular morphology, immunofluorescence (IF) staining was performed on the differentiated vascular network at day 10 and the vascular organoid at day 25. Vascular networks at day 10 exhibit a highly branched vascular network (Fig. 3C). The 3D reconstruction of the vascular organoids demonstrates the spatial morphology of the vascular organoids, and an interweaving of ECs (CD31+) and SMCs (αSMA + ) within the organoids (Figs. 4A, 5A). Comparison of organoids from MMD patients and HCs showed that MMD-derived vascular organoids exhibited a denser and more extensive vascular network (Fig. 4A, Supplementary Fig. 6A). This demonstrated an excessive vascular proliferation in the vascular organoids derived from MMD patients.
A The flowchart of vascular organoid generation derived from HC and MMD patients. B Bright-field images of human induced pluripotent stem cells (iPSCs) differentiation into vascular organoids (day-3 – day25). Scale = 300 μm. MMD Moyamoya Disease, HC Healthy Controls. C Representative immunofluorescence of CD31 and αSMA expressing endothelial and smooth muscle cells, respectively, shows the establishment of vascular networks. CD31, red; αSMA, green; DAPI, blue; HC Healthy Controls; MMD, Moyamoya Disease. Scale 100 μm.
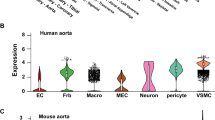
A Representative immunofluorescence and three-dimensional (3D) reconstruction of HC and MMD vascular organoids. CD31, green; αSMA, red; DAPI, blue. Scale = 100μm. n = 3. B Schematic representation of single-cell RNA sequencing. A total of 85,392 single cells were captured, including organoids derived from 2 MMD patients and 2 HC. C A UMAP plot displaying all cells colored according to the 9 major cell types. D The proportion of different cell clusters within each organoid. E The percentage of each cell type in vascular organoids derived from MMD and HC. SingleR Single-cell RNA sequencing.
A Immunofluorescence staining of smooth muscle cells during the vascular network phase in MMD organoids. B The weight and number of connections between smooth muscle cells and other cell types in the MMD organoids. C Immunofluorescence staining of TUBA4A in vascular organoids from MMD patients and HC. TUBA4A, red; DAPI, blue. n = 3. D Western blot showing the expression level of TUBA4A and TUBB4B in the vascular organoids. The bar chart shows the expression level of TUBA4A and TUBB4B (mean ± SD, n = 3 biologically independent samples). E GO enrichment analysis of smooth muscle cells in vascular organoids. GO Gene Ontology.
Transcriptomic profiling reveals the correlation of beta-tubulin and DEGs between MMD and HC vascular organoids
Bulk RNA sequencing was performed on organoid tissues to identify differentially expressed genes (DEGs) between organoids derived from MMD patients and HC individuals. A total of 112 genes were upregulated and 268 genes were downregulated in MMD-derived organoids compared to those from HCs (Supplementary Fig. 7A). GO enrichment analysis for these DEGs revealed significant enrichment in molecular functions (MF) related to beta tubulin binding, filamin protein binding, etc. (Supplementary Fig. 7B). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis identified enrichment in pathways such as Toll-like receptor signaling pathway, Cytokine-cytokine receptor interaction, etc. (Supplementary Fig. 7C). The alternative splicing analysis of six vascular organoid is shown in Supplementary Fig. 7D. The transcriptomic data collectively showed that DEGs distinguishing MMD from HC organoids are functionally associated with tubulin-related cytoskeletal processes.
Single-cell RNA sequencing reveals increased abundance and functional interactions of VSMCs in MMD organoid
To analyze the cell types of vascular organoids, we performed single-cell RNA sequencing (scRNA-seq) for 2 MMD organoids and 2 HC organoids. By digesting the vascular organoids in day 25, a total of 85,392 single cells were collected and followed by scRNA-seq (Fig. 4B). Following quality control filtering as described in the Methods section, a total of 16,507 low-quality cells were excluded, and 71,627 high-quality cells were retained for downstream analysis. A total of 9 cell types is annotated, including smooth muscle cells (SMC), endothelial cells, iPSCs, MSC, fibroblasts, DC, neurons, tissue stem cells and chondrocytes (Fig. 4C). The proportion of various cell type in each sample was shown in Fig. 4D. The proportion of VSMCs was significantly higher in MMD organoids compared to HC organoids (Fig. 4E). SMCs also exhibited a high number of predicted intercellular interactions and stronger interaction weights with other cell types, including ECs, chondrocytes, MSC and fibroblasts (Fig. 5B). We further analyzed the interactions between different cell clusters. The GO enrichment analysis of VSMCs in organoids is shown in Fig. 5E. Collectively, scRNA-seq analysis reveals an expansion of the VSMC population and increased functional interactions in MMD-derived vascular organoids, which are potentially related to MMD pathogenesis.
MMD vascular organoids exhibit an upregulation of TUBA4A and activation of the downstream GJA1/PI3K/AKT/KLF4 pathways
The IF staining was performed on the vascular organoids at day 25 and observed an enhanced fluorescence signal of TUBA4A protein in the MMD organoids compared with HC (Fig. 5C, Supplementary Fig. 6B). The western blot confirmed the upregulated expression of TUBA4A and TUBB4B in MMD vascular organoids (Fig. 5D). Meanwhile, the activation of the downstream GJA1/PI3K/AKT/KLF4 pathway of TUBA4A and TUBB4B was also observed in MMD organoids (Supplementary Fig. 8A, 8F-K). Additionally, in the vascular organoids, we observed a decrease in the expression levels of VSMCs' contractile markers (CNN1 and SM22α) and an increase in synthetic markers (S100A4 and ZO-2) (Supplementary Fig. 8A-E). These molecular changes demonstrated that SMCs derived from MMD organoids also have a tendency of contractile-to-synthetic phenotypic switching.
Meanwhile, following treatment with MMD serum, the expression of GJA1, p-AKT, p-PI3K, and KLF4 in HBVSMCs was found to be significantly upregulated (Supplementary Fig. 2F, 2K). In HBVSMCs overexpressing TUBA4A and TUBB4B, the expression levels of GJA1, p-AKT, p-PI3K, and KLF4 were also significantly upregulated (Supplementary Fig. 3E, 3J). Upon knockdown of TUBA4A or TUBB4B, the expression levels of GJA1, p-AKT, p-PI3K, and KLF4 were downregulated (Supplementary Fig. 3E, 3K). These results suggest that TUBA4A or TUBB4B modulates the changes of HBVSMCs phenotypes by activating these downstream molecules.
To further investigate whether the PI3K/AKT/KLF4 pathway mediates the GJA1 overexpression-caused changes of VSMCs, the overexpression plasmids and shRNA were utilized to upregulate or downregulate the level of GJA1, respectively (Supplementary Fig. 5A, 4F, G, 4M, N). We found that overexpression of GJA1 significantly upregulated the expression levels of p-AKT, p-PI3K, and KLF4 (Supplementary Fig. 5B, 5H). However, upon knockdown of GJA1, the upregulation of these molecules induced by TUBA4A or TUBB4B was blocked (Supplementary Fig. 5B, 5G). Then we used Kenpaullone, a KLF4 inhibitor, to downregulate the expression of the downstream KLF4 molecule in this pathway (Supplementary Fig. 5A). After the KLF4 inhibition for HBVSMCs, the enhanced proliferation, migration, and contractile-to-synthetic phenotypic switching induced by GJA1 overexpression were all blocked (Supplementary Fig. 5B–F, 5I). These results demonstrate that the GJA1/PI3K/AKT/KLF4 pathway contributes to the TUBA4A and TUBB4B induced changes of HBVSMC (Fig. 2C).
TUBA4A or TUBB4B overexpression in HBVSMCs promotes the proliferation of HBECs via KLF4 in a GJA1-dependent manner
To investigate the effects of the TUBA4A/TUBB4B overexpressed VSMC on ECs, a co-culture system of HBVSMCs and HBECs was established (Fig. 1D). Overexpression of TUBA4A/TUBB4B in HBVSMCs led to a significant increased proliferation of the co-cultured HBECs (Fig. 6A–C). Knocked down of GJA1 in HBEC attenuated the increased proliferative ability of HBECs induced by TUBA4A/TUBB4B overexpressed HBVSMCs (Fig. 6A–F). Similarly, pharmacological inhibition of KLF4 using Kenpaullone reduced HBEC proliferation in the SMC-EC co-culture system (Figs. 6A-B, G). In addition, western blot analysis showed that KLF4 expression levels in HBECs were elevated following co-culture with TUBA4A-overexpressing HBVSMCs (Fig. 6E, G). However, when GJA1 was knocked down in HBECs, the increase of KLF4 levels in the co-cultured HBECs was abolished, suggesting a GJA1-dependent intercellular signaling (Fig. 6E, G). These results demonstrate that KLF4 upregulation in HBECs is mediated by intercellular communication with TUBA4A/TUBB4B-overexpressing HBVSMCs through GJA1, which may contribute to the enhanced endothelial proliferation observed in the co-culture system (Fig. 2C).
A Ki67 staining of Human Brain Endothelial Cells (HBECs) after co-culture with treated HBVSMCs. The bar chart of B shows the proportion of proliferation HBEC (mean ± SD, n = 3 biologically independent samples). C Western blot showing the expression levels of TUBA4A, TUBB4B, and GJA1 in HBVSMCs under different treatment conditions in the SMC-EC co-culture system. The bar chart of D shows the expression level of GJA1 in HBVSMCs (mean ± SD, n = 3 biologically independent samples). E Western blot showing the expression levels of TUBA4A, TUBB4B, and GJA1 in HBECs under different treatment conditions in the SMC-EC co-culture system. The bar chart of F and G shows the expression level of GJA1 and KLF4 in HBECs, respectively (mean ± SD, n = 3 biologically independent samples). HBVSMC Human brain vascular smooth muscle cells, HBEC Human brain Endothelial cells, SMC-EC Smooth muscle cells-Endothelial cells.
Discussion
The pathogenesis of MMD remains unclear, resulting in a lack of available mechanism-based therapies42. There is an urgent need for suitable methodologies to further elucidate the underlying mechanism of MMD. In this study, we developed the first MMD vascular organoids derived from iPSCs of MMD patients, aiming to investigate its vascular changes and underlying mechanism in a controlled human-based system. Single-cell RNA sequencing of these organoids showed the increased proportion and functional abnormalities of VSCMs in the MMD vascular organoid. In parallel, DIA proteomics identified significant upregulation of cytoskeletal proteins TUBA4A and TUBB4B in patient serum. Functional experiments further demonstrated that the overexpression of these proteins enhanced the migration, contractile-to-synthetic phenotypic switching, and proliferation of VSMCs, as well as enhanced the proliferation of co-cultured ECs. These changes were mediated through the GJA1/PI3K/AKT/KLF4 signaling axis, which also promoted endothelial proliferation in a co-culture model. Taken together, this study provides a novel MMD-specific vascular organoid platform and uncovers VSMC changes in the mechanism of intimal thickening in MMD vessels.
The main histological characteristics of pathological arteries in MMD is the significant thickening of the intima in the vessel wall which may be the primary pathological change contributing to the progressive stenosis of vessels in MMD38,43. Previous histological studies have demonstrated that the main component of the thickening intima is over-proliferative VSMCs5,6. These findings demonstrate that VSMCs may play a significant role in intimal thickening of MMD vessels. However, it remains unclear why VSMCs, which typically only grow in the media of normal blood vessels, abnormally appear in large numbers in the intima of MMD arteries. This indicates that VSMCs acquire migratory capacity, enabling their translocation from the media to the intima; however, the underlying mechanism governing this acquisition remains unclear. VSMCs can be roughly classified into two phenotypes: contractile and synthetic phenotypes44. In response to vascular injury or alterations in local environmental cues, relatively quiescent contractile VSMCs can transform into a synthetic phenotype with increased migratory, proliferative, and synthetic properties45. VSMC phenotype switching has been widely studied in atherosclerosis and is considered as one of the initiating factors of atherosclerosis, but few studies have investigated VSMC phenotypic switching in MMD46. In our study, VSMC co-cultured with MMD patient serum exhibited enhanced migratory capacity accompanied by a phenotypic switch from the contractile to the synthetic type. This demonstrates that these changes are present in the VSMCs of MMD.
A reliable model is essential for investigating the pathogenesis of MMD and evaluating potential therapeutic interventions. Current methods developed for creating the MMD animal models generally involve surgical, immunological, genetic, or combinatory approaches19,20. However, none of these modeling approaches can simultaneously recapitulate the genetic background and vascular pathology of MMD47,48,49,50. The absence of reliable animal models for MMD greatly limited the research on the pathogenesis and drug development for MMD. Recently, organoids have provided new insights for studying organ-specific functions and disease mechanisms by simulating the complex three-dimensional structures of in vivo organs51. Unlike animal models, patient-derived vascular organoids provide a physiologically relevant in vitro system that retains the genetic background of patients and more accurately recapitulates human-specific cellular phenotypes and molecular signaling pathways51,52. In the field of vascular diseases, researchers have successfully developed 3D vascular organoids for several diseases, such as diabetes and cavernous malformation, advancing our understanding of vascular diseases32,33. The vascular organoid model represents a highly promising approach for MMD research53. However, in the field of MMD, a stable model of vascular organoids specific to MMD has not yet been established.
In this study, we established a vascular organoid model derived from iPSCs of MMD patients. This is, to our knowledge, the first in vitro 3D system that incorporates the genetic and cellular context of MMD, enabling the study of disease-relevant vascular changes in a controlled setting. Compared with previous in vivo models that often lack reproducibility or genetic specificity, our organoids recapitulate essential features such as increased vascular and VSMC proliferation47,48,49,50. Subsequently, histological staining for organoids demonstrates a significant expression and interweaving of ECs and VSMCs. Additionally, the MMD vascular organoids exhibited a denser vascular network and more pronounced fluorescence signals from VSMCs. It indicates the enhanced angiogenesis in the MMD vascular organoids, consistent with our previous study38,54. Single-cell RNA sequencing confirmed an increased proportion of VSMCs in the MMD vascular organoids compared to those from HC individuals, which indicates a proliferation of VSMCs in MMD organoids. The MMD vascular organoids we constructed recapitulated the previously reported abnormal increase of VSMCs observed in pathological vessels of MMD5,6. In summary, the vascular organoids with the genetic background of MMD patients exhibited enhanced angiogenic capacity, along with excessive proliferation and functional abnormalities of VSMCs.
Our DIA proteomic mass spectrometry analysis revealed widespread upregulation of five microtubule family proteins in the serum of MMD patients, including TUBA1A, TUBA4A, TUBB1, TUBB5, and TUBB4B. Bulk RNA transcriptomic analysis of the organoids also revealed enrichment of the DEGs in the β-tubulin pathway. This indicates that the upregulation of microtubule proteins is closely associated with the pathogenesis of MMD. Microtubules are essential components of the cytoskeleton, composing of repeating α-tubulin and β-tubulin heterodimers55. They are crucial for a variety of cellular functions, including cellular connectivity, intracellular transportation, cell motility, and division56,57,58. Several studies confirmed that microtubules can regulate the migration of SMCs by modulating the polarization process, lamellipodia formation, and the assembly of focal adhesions59,60. It indicates a potential link of microtubules to vascular growth and development61. Among the upregulated microtubule family proteins, we selected the two most prominently elevated members—TUBA4A and TUBB4B—for further investigation into their roles in the pathogenesis of MMD by in vitro experiments.
The following experiments revealed that the overexpression of TUBA4A and TUBB4B led to the upregulation of Gap junction A1 (GJA1). GJA1 encodes Connexin43 (Cx43/GJA1), which forms gap junction channels between cells, allowing the transmission of molecules, ions, and electrical signals, thereby facilitating intercellular communication62,63,64. Pericyte damage and the deficiency of GJA1 lead to impaired vasomotor function in diabetic retinopathy, suggesting a regulatory role of GJA1 in vascular function65. A previous study also reported the upregulation of GJA1 in the pathological cerebral arteries from MMD patients by conducting IF staining66. A previous study also reported the upregulation of GJA1 in the pathological cerebral arteries from MMD patients by conducting IF staining, suggesting a potential association between GJA1 and MMD66. Krüppel-like factor 4 (KLF4), a highly conserved transcription factor, has been widely reported as a key regulatory factor for the phenotypic switching of VSMCs67,68. In our study, we found that overexpression of TUBA4A and TUBB4B upregulates KLF4 expression by activating the PI3K/AKT signaling pathway. After the upregulation of KLF4, we observed contractile-to-synthetic phenotypic switching in VSMCs of MMD, accompanied by enhanced migratory and proliferative capacities. Meanwhile, the application of a KLF4 inhibitor reversed these TUBA4A or TUBB4B-induced changes of VSMCs. It suggests that KLF4 is a potential target for suppressing the functional abnormalities of VSMCs in MMD.
Taken together, based on our findings from vascular organoids, DIA-based proteomics, and in vitro experiments, we propose a novel hypothesis regarding the mechanism underlying intimal thickening in MMD: Aberrant upregulation of TUBA4A and TUBB4B in VSMC of MMD activates the crucial GJA1/PI3K/AKT/KLF4 signaling pathway, leading to a phenotypic switch from a contractile to a synthetic phenotype. The phenotypically transformed VSMCs acquire enhanced migratory and proliferative capacities, enabling their migration from the vessel media to the intima, where they undergo excessive proliferation. This pathological process leads to intimal thickening of the affected vessels in MMD and ultimately results in the progressive stenosis characteristic of MMD vasculopathy. Our graphical abstract illustrates the mechanistic pathway underlying this process.
Limitations of the study
Although organoid models have advantages in simulating the physiological and pathological features of blood vessels, they are still not fully representative of the complex pathological environment in vivo. Organoid models may lack certain systemic factors, such as the influence of the immune system, which may affect the interpretation of the results. In addition, a further exploration method can be to transplant vascular organoids into immunodeficient mice, which will allow human and mouse blood vessels to anastomose and reconstruct blood flow in human capillaries to further judge the condition of the model. Meanwhile, while our study demonstrates the functional role of TUBA4A and TUBB4B in VSMC remodeling, the upstream signaling mechanisms responsible for the upregulation of these cytoskeletal proteins in MMD remain unclear. Our serum–VSMC co-culture experiments suggest the presence of additional regulatory pathways, which warrant further investigation in future studies. Another limitation of this study is that DIA proteomics cannot detect genetic variants such as the RNF213 p.R4810K mutation, which is a major focus in MMD research, thereby restricting our analysis to protein-level dysregulation rather than underlying genomic changes15,69. In future work, we will expand genetic studies on MMD patient samples to investigate pathogenesis at the genetic level.
Conclusion
This study successfully established a vascular organoid model derived from MMD patients’ iPSCs, and through single-cell sequencing and proteomics analysis, revealed significant abnormalities in the quantity and function of VSMCs in MMD. The findings demonstrate that the abnormal overexpression of TUBA4A and TUBB4B may promote phenotypic switching, proliferation, and migration of VSMCs by regulating the GJA1 signaling pathway, ultimately contributing to the intimal thickening observed in MMD, a key pathological feature of the disease.
Our results provide new insights into the pathogenesis of MMD and suggest that TUBA4A and TUBB4B could serve as potential therapeutic targets. Future studies should further validate the functional roles of these genes and the underlying pathways and explore their impact in different populations and subtypes to advance the development of novel therapeutic strategies for MMD.
Ethics approval and consent to participate
Written informed consent was obtained from all participants. All human tissue samples were obtained with written informed consent from the patients All ethical regulations relevant to human research participants were followed. This study and the involved protocols were approved by the Institutional Ethics Committee of Beijing Tiantan Hospital, Beijing, China (KY2023-204-02).
Data availability
Uncropped and unedited blot/gel images accompanied by size markers for all relevant western blots figure panels are available in Supplementary Materials. The raw DIA proteomics data have been deposited in the ProteomeXchange Consortium via the iProX partner repository (ID PXD038468). The raw data for transcriptome sequencing have been deposited in the NCBI Sequence Read Archive (SRA) database (ID PRJNA1367356). The raw data for single-cell sequencing have been deposited in the NCBI Sequence Read Archive (SRA) database (PRJNA1367840). The newly generated plasmids have been deposited in NCBI GenBank with BankIt submission IDs PX673704, PX673705 and PX673706. All other source data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Custom computer code used to generate, process, and analyze the sequencing data in this study is available from the corresponding author upon reasonable request.
References
Gonzalez, N. R. et al. Adult Moyamoya Disease and Syndrome: Current Perspectives and Future Directions: A Scientific Statement From the American Heart Association/American Stroke Association. Stroke 54, e465–e479 (2023).
Research Committee on the P, Treatment of Spontaneous Occlusion of the Circle of W, Health Labour Sciences Research Grant for Research on Measures for Infractable D Guidelines for diagnosis and treatment of moyamoya disease (spontaneous occlusion of the circle of Willis). Neurol. Med Chir. 52, 245–266 (2012).
Kuroda, S. & Houkin, K. Moyamoya disease: current concepts and future perspectives. Lancet Neurol. 7, 1056–1066 (2008).
Rifino, N. et al. Diagnosis and management of adult Moyamoya angiopathy: An overview of guideline recommendations and identification of future research directions. Int J. Stroke 20, 512–523 (2025).
Takekawa, Y., Umezawa, T., Ueno, Y., Sawada, T. & Kobayashi, M. Pathological and immunohistochemical findings of an autopsy case of adult moyamoya disease. Neuropathology 24, 236–242 (2004).
Takagi, Y., Kikuta, K., Nozaki, K. & Hashimoto, N. Histological features of middle cerebral arteries from patients treated for Moyamoya disease. Neurol. Med Chir. 47, 1–4 (2007).
Takagi, Y. et al. Histopathological Characteristics of Distal Middle Cerebral Artery in Adult and Pediatric Patients with Moyamoya Disease. Neurol. Med Chir. 56, 345–349 (2016).
Ye, F. et al. RNF213 loss-of-function promotes pathological angiogenesis in moyamoya disease via the Hippo pathway. Brain 146, 4674–4689 (2023).
Roy, V. et al. Moyamoya disease susceptibility gene RNF213 regulates endothelial barrier function. Stroke 53, 1263–1275 (2022).
Morito D. et al. Moyamoya disease-associated protein mysterin/RNF213 is a novel AAA+ ATPase, which dynamically changes its oligomeric state. Sci. Rep. 4 https://doi.org/10.1038/srep04442 (2014).
Liu, W. et al. Identification of RNF213 as a susceptibility gene for moyamoya disease and its possible role in vascular development. PLoS One 6, e22542 (2011).
Sonobe, S. et al. Temporal profile of the vascular anatomy evaluated by 9.4-T magnetic resonance angiography and histopathological analysis in mice lacking RNF213: A susceptibility gene for moyamoya disease. Brain Res. 1552, 64–71 (2014).
Lu, J. et al. MMP-9 as a biomarker for predicting hemorrhagic strokes in Moyamoya Disease. Front. Neurol. 12, 721118 (2021).
Park, Y. S. et al. The role of VEGF and KDR polymorphisms in moyamoya disease and collateral revascularization. PLoS One 7, e47158 (2012).
Mertens, R. et al. PJTsr. The genetic basis of Moyamoya Disease. Transl. Stroke Res. 13, 25–45 (2022).
Wu, H. et al. APOE as potential biomarkers of moyamoya disease. Front. Neurol. 14, 1156894 (2023).
Gorla, G. et al. Angiopoietin-2 associates with poor prognosis in Moyamoya angiopathy. Ann. Clin. Transl. Neurol. 11, 1590–1603 (2024).
Guo, D., Dong, Y., Li, H., Li, H. & Yang, B. Proteomics and digital subtraction angiography approaches reveal CDH18 as a potential target for therapy of moyamoya disease. Biol. Direct 19, 76 (2024).
Rallo, M., Akel, O., Gurram, A. & Sun HJNf. Experimental animal models for moyamoya disease and treatment: a pathogenesis-oriented scoping review. Neurosurg. Focus 51, E5 (2021).
Vijay L. et al. Experimental animal models for the study of moyamoya disease. 51, https://doi.org/10.3171/2021.5.Focus21282 (2021).
Choi, S. A. et al. Impaired functional recovery of endothelial colony-forming cells from moyamoya disease in a chronic cerebral hypoperfusion rat model. J. Neurosurg. Pediatr. 23, 204–213 (2018).
Rallo, M. S., Akel, O., Gurram, A. & Sun, H. Experimental animal models for moyamoya disease and treatment: a pathogenesis-oriented scoping review. Neurosurg. Focus 51, E5 (2021).
Sato-Maeda, M. et al. Transient middle cerebral artery occlusion in mice induces neuronal expression of RNF213, a susceptibility gene for moyamoya disease. Brain Res. 1630, 50–55 (2016).
Morimoto T. et al. Dysregulation of RNF213 promotes cerebral hypoperfusion. Sci. Rep. 8, https://doi.org/10.1038/s41598-018-22064-8 (2018).
Kanoke, A. et al. Temporal profile of the vascular anatomy evaluated by 9.4-tesla magnetic resonance angiography and histological analysis in mice with the R4859K mutation of RNF213, the susceptibility gene for moyamoya disease. Brain Res. 1624, 497–505 (2015).
Rao M. et al. Clinical and experimental pathology of Moyamoya disease. 116, 1845–1849 (2003).
Yamada H. et al. The relationship between moyamoya disease and bacterial infection. Clin. Neurol. Neurosurg. S221–S224 (1997).
Rossi, G., Manfrin, A. & Lutolf, M. P. Progress and potential in organoid research. Nat. Rev. Genet. 19, 671–687 (2018).
Kretzschmar, K. Cancer research using organoid technology. J. Mol. Med. 99, 501–515 (2021).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
McCracken, K. W. et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 516, 400–404 (2014).
Reiner A. W. et al. Human blood vessel organoids as a model of diabetic vasculopathy. 565, https://doi.org/10.1038/s41586-018-0858-8 (2019).
Dao, L. et al. Modeling blood-brain barrier formation and cerebral cavernous malformations in human PSC-derived organoids. Cell Stem Cell 31, 818–833.e811 (2024).
Ji Y. et al. Alzheimer’s disease patient brain extracts induce multiple pathologies in novel vascularized neuroimmune organoids for disease modeling and drug discovery. Mol. Psychiatry. https://doi.org/10.1038/s41380-025-03041-w 2025.
Arce, M. et al. KRIT1 heterozygous mutations are sufficient to induce a pathological phenotype in patient-derived iPSC models of cerebral cavernous malformation. Cell Rep. 44, 115576 (2025).
Zhu, L. et al. Construction of vascularized liver microtissues recapitulates angiocrine-mediated hepatocytes maturation and enhances therapeutic efficacy for acute liver failure. Bioact. Mater. 50, 525–539 (2025).
JNm-c. Guidelines for diagnosis and treatment of moyamoya disease (spontaneous occlusion of the circle of Willis). Neurol. Med. Chir. 52, 245-266 (2012).
He, S. et al. Upregulated cytoskeletal proteins promote pathological Angiogenesis in Moyamoya disease. Stroke 54, 3153–3164 (2023).
Wimmer, R. A., Leopoldi, A., Aichinger, M., Kerjaschki, D. & Penninger, J. M. Generation of blood vessel organoids from human pluripotent stem cells. Nat. Protoc. 14, 3082–3100 (2019).
Okita, K. et al. An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells 31, 458–466 (2013).
Fusaki, N., Ban, H., Nishiyama, A., Saeki, K. & Hasegawa, M. Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 85, 348–362 (2009).
Scott, R. M. & Smith, E. R. Moyamoya disease and moyamoya syndrome. N. Engl. J. Med. 360, 1226–1237 (2009).
Fox, B. M., Dorschel, K. B., Lawton, M. T. & Wanebo, J. E. Pathophysiology of Vascular Stenosis and Remodeling in Moyamoya Disease. Front Neurol. 12, 661578 (2021).
Frismantiene, A., Philippova, M., Erne, P. & Resink, T. J. Smooth muscle cell-driven vascular diseases and molecular mechanisms of VSMC plasticity. Cell Signal 52, 48–64 (2018).
Gary K. O., Meena S. K., Brian R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, https://doi.org/10.1152/physrev.00041.2003 (2004).
Basatemur, G. L., Jorgensen, H. F., Clarke, M. C. H., Bennett, M. R. & Mallat, Z. Vascular smooth muscle cells in atherosclerosis. Nat. Rev. Cardiol. 16, 727–744 (2019).
Wen, J. et al. Mutation of rnf213a by TALEN causes abnormal angiogenesis and circulation defects in zebrafish. Brain Res. 1644, 70–78 (2016).
Starosolski, Z. et al. Ultra High-Resolution In vivo Computed Tomography Imaging of Mouse Cerebrovasculature Using a Long Circulating Blood Pool Contrast Agent. Sci. Rep. 5, 10178 (2015).
Ren, X. et al. Linking cortical astrocytic neogenin deficiency to the development of Moyamoya disease-like vasculopathy. Neurobiol. Dis. 154, 105339 (2021).
Yamada, H. et al. The relationship between moyamoya disease and bacterial infection. Clin. Neurol. Neurosurg. 99, S221–S224 (1997).
Kim, J., Koo, B. K. & Knoblich, J. A. Human organoids: model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 21, 571–584 (2020).
Han, X. et al. Landscape of human organoids: Ideal model in clinics and research. Innovations 5, 100620 (2024).
Naderi-Meshkin, H. et al. Vascular organoids: unveiling advantages, applications, challenges, and disease modelling strategies. Stem Cell Res Ther. 14, 292 (2023).
He, S. et al. Metabolomic signatures associated with pathological angiogenesis in moyamoya disease. Clin. Transl. Med. 13, e1492 (2023).
Binarova P., Tuszynski J. Tubulin: Structure, functions and roles in disease. Cells. 8, https://doi.org/10.3390/cells8101294 (2019).
Janke, C. & Magiera, M. M. The tubulin code and its role in controlling microtubule properties and functions. Nat. Rev. Mol. Cell Biol. 21, 307–326 (2020).
Vicente, J. J. & Wordeman, L. The quantification and regulation of microtubule dynamics in the mitotic spindle. Curr. Opin. Cell Biol. 60, 36–43 (2019).
Guedes-Dias P., Holzbaur E. L. F. Axonal transport: Driving synaptic function. Science. 366, https://doi.org/10.1126/science.aaw9997 (2019).
Silverman-Gavrila, R. et al. MJTAjop. Rear polarization of the microtubule-organizing center in neointimal smooth muscle cells depends on PKCα, ARPC5, and RHAMM. Am. J. Pathol. 178, 895–910 (2011).
Zhu, X., Efimova, N., Arnette, C., Hanks, S. & Kaverina, I. J. C. Podosome dynamics and location in vascular smooth muscle cells require CLASP-dependent microtubule bending. Cytoskeleton 73, 300–315 (2016).
Pasquier, E., André, N. & Braguer DJCcdt. Targeting microtubules to inhibit angiogenesis and disrupt tumour vasculature: implications for cancer treatment. Curr. Cancer Drug Targets 7, 566–581 (2007).
Thévenin, A. et al. Proteins and mechanisms regulating gap-junction assembly, internalization, and degradation. Hysiology 28, 93–116 (2013).
Molica, F., Figueroa, X., Kwak, B., Isakson, B. & Gibbins, J. Connexins and Pannexins in Vascular Function and Disease. Int. J. Mol. Sci. 19, 1663 (2018).
Epifantseva, I. & Shaw, R. M. Intracellular trafficking pathways of Cx43 gap junction channels. Biochim. Biophys. Acta (BBA) - Biomembranes. 1860, 40–47 (2018).
Ivanova, E., Kovacs-Oller, T. & Sagdullaev, B. T. Vascular Pericyte Impairment and Connexin43 Gap Junction Deficit Contribute to Vasomotor Decline in Diabetic Retinopathy. J. Neurosci. 37, 7580–7594 (2017).
Liao, J. et al. Expression of Connexin43 in Cerebral Arteries of Patients with Moyamoya Disease. J. Stroke Cerebrovasc. Dis. 27, 1107–1114 (2018).
Yap, C., Mieremet, A., de Vries, C., Micha, D. & de Waard, V. J. A. thrombosis, biology v. Six Shades of Vascular Smooth Muscle Cells Illuminated by KLF4 (Krüppel-Like Factor 4). Arterioscler. Thromb. Vasc. Biol. 41, 2693–2707 (2021).
Zheng, B., Han, M. & Wen, J. K. Role of Krüppel-like factor 4 in phenotypic switching and proliferation of vascular smooth muscle cells. IUBMB Life 62, 132–139 (2010).
Kronenburg, A., Braun, K. P., van der Zwan, A. & Klijn, C. J. Recent advances in moyamoya disease: pathophysiology and treatment. Curr. Neurol. Neurosci. Rep. 14, 423 (2014).
Acknowledgements
Thanks to the health care workers in the department for their help with the project. Thanks to all participants for their support and cooperation. This study was supported by the National Natural Science Foundation of China (82171887 and 82371296 to RW, 82471337 to YL), National High Level Hospital Clinical Research Funding (2023-PUMCHE-011 to YL), and Beijing Natural Science Foundation (No. 7252042 to RW).
Author information
Authors and Affiliations
Contributions
Shihao He, Yuanli Zhao, Rong Wang provide ideas and make important contributions in various aspects. Shihao He, Junze Zhang, Xilong Wang, Zhenyu Zhou, Yanru Wang, Xun Ye, Ziqi Liu, Xiaokuan Hao, Yuanli Zhao, Rong Wang, collected clinical sample specimens and information. Shihao He, Junze Zhang Zhenyu Zhou, Zhenyu Zhou, Yanru Wang, completed the in vitro part of the experiment. Shihao He, Junze Zhang, and Shaoqi Xu did single-cell sequencing and analysis. Zhen Qi, Dandan Li assist in organoid culture. Shihao He, Junze Zhang, Zhenyu Zhou, and Shaoqi Xu completed the data integration and analysis. All the authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
Not applicable.
Peer review
Peer review information
Communications Biology thanks Yulius Hermanto and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Benjamin Bessieres.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, S., Zhang, J., Wang, X. et al. Organoid Modeling and Single-Cell Profiling Reveal Smooth Muscle Cell Migration in Moyamoya Disease. Commun Biol 9, 198 (2026). https://doi.org/10.1038/s42003-025-09476-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09476-9