Abstract
Metabolic dysfunction-associated steatohepatitis (MASH) is an important phase in the progression of metabolic dysfunction-associated steatotic liver disease to end-stage liver diseases, posing an increasing threat to public health worldwide with limited treatment options. Here we show that GPR110 is a liver-selective G-protein-coupled receptor closely associated with MASH in a sex-specific manner. Hepatocyte-specific Gpr110 knockout protects against MASH in female, but not male mice. The GPR110 variant rs937057 T > C is associated with a higher prevalence of metabolic dysfunction-associated steatotic liver disease in women. The improved liver phenotypes in female mice are abrogated by knocking down the expression of hepatic oestrogen receptor alpha (Esr1). Mechanistically, GPR110 couples to Gαs and activates protein kinase A, thereby inducing phosphorylation of NFAT2, which inhibits its nuclear translocation and transcriptional activity, leading to suppressed Esr1 transcription in hepatocytes. Taken together, these results demonstrate a sex-specific role of GPR110 in MASH by regulating hepatic oestrogen sensitivity, suggesting inhibition of GPR110 as a potential sex-specific therapy for MASH.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data supporting the findings from this study are available within the manuscript and its Supplementary Information. RNA-seq data are available in the Genome Sequence Archive under accession number PRJCA049229. Source imaging data are available in figshare (https://doi.org/10.6084/m9.figshare.28532336)91. Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding author. Source data are provided with this paper.
Code availability
This paper does not report original code.
References
Younossi, Z. M. et al. The global epidemiology of nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH): a systematic review. Hepatology 77, 1335–1347 (2023).
Diehl, A. M. & Day, C. Cause, pathogenesis, and treatment of nonalcoholic steatohepatitis. N. Engl. J. Med. 377, 2063–2072 (2017).
Friedman, S. L., Neuschwander-Tetri, B. A., Rinella, M. & Sanyal, A. J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 24, 908–922 (2018).
Huang, D. Q. et al. Changing global epidemiology of liver cancer from 2010 to 2019: NASH is the fastest growing cause of liver cancer. Cell Metab. 34, 969–977 (2022).
Ren, Z., Simons, P., Wesselius, A., Stehouwer, C. D. A. & Brouwers, M. Relationship between NAFLD and coronary artery disease: a Mendelian randomization study. Hepatology 77, 230–238 (2023).
Liu, Z. et al. Metabolic dysfunction-associated fatty liver disease and the risk of 24 specific cancers. Metabolism 127, 154955 (2022).
Byrne, C. D. & Targher, G. NAFLD as a driver of chronic kidney disease. J. Hepatol. 72, 785–801 (2020).
Riazi, K. et al. The prevalence and incidence of NAFLD worldwide: a systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 7, 851–861 (2022).
Man, S. et al. Prevalence of liver steatosis and fibrosis in the general population and various high-risk populations: a nationwide study with 5.7 million adults in China. Gastroenterology 165, 1025–1040 (2023).
Ji, H. et al. Sex differences in prevalence and prognosis of steatotic liver disease phenotypes: biological sex matters. J. Hepatol. 80, e68–e69 (2024).
Kingwell, K. NASH field celebrates ‘hurrah moment’ with a first FDA drug approval for the liver disease. Nat. Rev. Drug Discov. 23, 235–237 (2024).
Harrison, S. A. et al. A phase 3, randomized, controlled trial of resmetirom in NASH with liver fibrosis. N. Engl. J. Med. 390, 497–509 (2024).
Stevens, R. C. et al. The GPCR Network: a large-scale collaboration to determine human GPCR structure and function. Nat. Rev. Drug Discov. 12, 25–34 (2013).
Chan, H. C. S., Li, Y., Dahoun, T., Vogel, H. & Yuan, S. New binding sites, new opportunities for GPCR drug discovery. Trends Biochem. Sci. 44, 312–330 (2019).
Dorsam, R. T. & Gutkind, J. S. G-protein-coupled receptors and cancer. Nat. Rev. Cancer 7, 79–94 (2007).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schiöth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Drucker, D. J. Mechanisms of action and therapeutic application of glucagon-like peptide-1. Cell Metab. 27, 740–756 (2018).
Garber, A. et al. Liraglutide versus glimepiride monotherapy for type 2 diabetes (LEAD-3 Mono): a randomised, 52-week, phase III, double-blind, parallel-treatment trial. Lancet 373, 473–481 (2009).
Astrup, A. et al. Effects of liraglutide in the treatment of obesity: a randomised, double-blind, placebo-controlled study. Lancet 374, 1606–1616 (2009).
Sorli, C. et al. Efficacy and safety of once-weekly semaglutide monotherapy versus placebo in patients with type 2 diabetes (SUSTAIN 1): a double-blind, randomised, placebo-controlled, parallel-group, multinational, multicentre phase 3a trial. Lancet Diabetes Endocrinol. 5, 251–260 (2017).
Davies, M. et al. Semaglutide 2.4 mg once a week in adults with overweight or obesity, and type 2 diabetes (STEP 2): a randomised, double-blind, double-dummy, placebo-controlled, phase 3 trial. Lancet 397, 971–984 (2021).
Leeson-Payne, A. et al. Loss of GPR75 protects against non-alcoholic fatty liver disease and body fat accumulation. Cell Metab. 36, 1076–1087 (2024).
Sass, F. et al. NK2R control of energy expenditure and feeding to treat metabolic diseases. Nature https://doi.org/10.1038/s41586-024-08207-0 (2024).
Han, F. et al. Hypercholesterolemia risk-associated GPR146 is an orphan G-protein coupled receptor that regulates blood cholesterol levels in humans and mice. Cell Res. 30, 363–365 (2020).
Yu, H. et al. GPR146 deficiency protects against hypercholesterolemia and atherosclerosis. Cell 179, 1276–1288 (2019).
Hu, X. et al. A gut-derived hormone regulates cholesterol metabolism. Cell 187, 1685–1700 (2024).
Lee, J. W. et al. Orphan GPR110 (ADGRF1) targeted by N-docosahexaenoylethanolamine in development of neurons and cognitive function. Nat. Commun. 7, 13123 (2016).
Wu, M. et al. Amelioration of non-alcoholic fatty liver disease by targeting adhesion G protein-coupled receptor F1 (Adgrf1). Elife https://doi.org/10.7554/eLife.85131 (2023).
Marinović, S. et al. NKG2D-mediated detection of metabolically stressed hepatocytes by innate-like T cells is essential for initiation of NASH and fibrosis. Sci. Immunol. 8, eadd1599 (2023).
Xiong, X. et al. Mapping the molecular signatures of diet-induced NASH and its regulation by the hepatokine Tsukushi. Mol. Metab. 20, 128–137 (2019).
Luo, Y. et al. Role of liquid fructose/sucrose in regulating the hepatic transcriptome in a high-fat Western diet model of NAFLD. J. Nutr. Biochem. 112, 109174 (2023).
The Tabula Muris Consortium, Overall coordination, Logistical coordination et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature https://doi.org/10.1038/s41586-018-0590-4 (2018).
Matsumoto, M. et al. An improved mouse model that rapidly develops fibrosis in non-alcoholic steatohepatitis. Int J. Exp. Pathol. 94, 93–103 (2013).
Yang, J. et al. A CDAHFD-induced mouse model mimicking human NASH in the metabolism of hepatic phosphatidylcholines and acyl carnitines. Food Funct. 15, 2982–2995 (2024).
Camargo, F. N. et al. Western diet-fed ApoE knockout male mice as an experimental model of non-alcoholic steatohepatitis. Curr. Issues Mol. Biol. 44, 4692–4703 (2022).
Yan, Z., Yan, H. & Ou, H. Human thyroxine binding globulin (TBG) promoter directs efficient and sustaining transgene expression in liver-specific pattern. Gene 506, 289–294 (2012).
Lefebvre, P. & Staels, B. Hepatic sexual dimorphism - implications for non-alcoholic fatty liver disease. Nat. Rev. Endocrinol. 17, 662–670 (2021).
Wolf, M. J. et al. Metabolic activation of intrahepatic CD8+ T cells and NKT cells causes nonalcoholic steatohepatitis and liver cancer via cross-talk with hepatocytes. Cancer Cell 26, 549–564 (2014).
Lee, C. et al. Formyl peptide receptor 2 determines sex-specific differences in the progression of nonalcoholic fatty liver disease and steatohepatitis. Nat. Commun. 13, 578 (2022).
Cui, J., Shen, Y. & Li, R. Estrogen synthesis and signaling pathways during aging: from periphery to brain. Trends Mol. Med. 19, 197–209 (2013).
Stoveken, H. M., Hajduczok, A. G., Xu, L. & Tall, G. G. Adhesion G protein-coupled receptors are activated by exposure of a cryptic tethered agonist. Proc. Natl Acad. Sci. USA 112, 6194–6199 (2015).
Zhu, X. et al. Structural basis of adhesion GPCR GPR110 activation by stalk peptide and G-proteins coupling. Nat. Commun. 13, 5513 (2022).
Lee, C., Kim, J. & Jung, Y. Potential therapeutic application of estrogen in gender disparity of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. Cells https://doi.org/10.3390/cells8101259 (2019).
Besse-Patin, A. et al. Estrogen signals through peroxisome proliferator-activated receptor-γ coactivator 1α to reduce oxidative damage associated with diet-induced fatty liver disease. Gastroenterology 152, 243–256 (2017).
McKenzie, J. et al. Effects of HRT on liver enzyme levels in women with type 2 diabetes: a randomized placebo-controlled trial. Clin. Endocrinol. 65, 40–44 (2006).
Zhang, W. & Wu, F. Linoleic acid induces human ovarian granulosa cell inflammation and apoptosis through the ER-FOXO1-ROS-NFκB pathway. Sci. Rep. 14, 6392 (2024).
Ouyang, Q. et al. Estrogen receptor gene 1 (ESR1) mediates lipid metabolism in goose hierarchical granulosa cells rather than in pre-hierarchical granulosa cells. Biology https://doi.org/10.3390/biology12070962 (2023).
Gregorio, K. C. R., Laurindo, C. P. & Machado, U. F. Estrogen and glycemic homeostasis: the fundamental role of nuclear estrogen receptors ESR1/ESR2 in glucose transporter GLUT4 regulation. Cells https://doi.org/10.3390/cells10010099 (2021).
Huang, B. X. et al. Synaptamide activates the adhesion GPCR GPR110 (ADGRF1) through GAIN domain binding. Commun. Biol. 3, 109 (2020).
Chow, C. W. & Davis, R. J. Integration of calcium and cyclic AMP signaling pathways by 14-3-3. Mol. Cell. Biol. 20, 702–712 (2000).
Rosenkranz, A. C., Rauch, B. H., Freidel, K. & Schrör, K. Regulation of protease-activated receptor-1 by vasodilatory prostaglandins via NFAT. Cardiovasc. Res. 83, 778–784 (2009).
Pérez Riba, M, Aranguren Ibáñez, Á, Matsoukas, M. -T. & Pardo Carrasco, L. Therapeutic uses of non-peptide inhibitors of the calcineurin-NFAT signaling pathway. US patent WO2016207212A1 (2016).
Qu, X. et al. Structural basis of tethered agonism of the adhesion GPCRs ADGRD1 and ADGRF1. Nature 604, 779–785 (2022).
Barros-Álvarez, X. et al. The tethered peptide activation mechanism of adhesion GPCRs. Nature 604, 757–762 (2022).
Hu, B. et al. Therapeutic siRNA: state of the art. Signal Transduct. Target. Ther. 5, 101 (2020).
Tan, D. J. H. et al. Global burden of liver cancer in males and females: changing etiological basis and the growing contribution of NASH. Hepatology 77, 1150–1163 (2023).
Petrick, J. L. et al. International trends in hepatocellular carcinoma incidence, 1978–2012. Int J. Cancer 147, 317–330 (2020).
Wang, Z., Xu, M., Hu, Z. & Shrestha, U. K. Prevalence of nonalcoholic fatty liver disease and its metabolic risk factors in women of different ages and body mass index. Menopause 22, 667–673 (2015).
Wong, V. W. -S. et al. Prevalence of non-alcoholic fatty liver disease and advanced fibrosis in Hong Kong Chinese: a population study using proton-magnetic resonance spectroscopy and transient elastography. Gut 61, 409–415 (2012).
Ganz, M., Csak, T. & Szabo, G. High fat diet feeding results in gender specific steatohepatitis and inflammasome activation. World J. Gastroenterol. 20, 8525–8534 (2014).
Kirsch, R. et al. Rodent nutritional model of non-alcoholic steatohepatitis: species, strain and sex difference studies. J. Gastroenterol. Hepatol. 18, 1272–1282 (2003).
Ohnishi, S., Murakami, T., Moriyama, T., Mitamura, K. & Imawari, M. Androgen and estrogen receptors in hepatocellular carcinoma and in the surrounding noncancerous liver tissue. Hepatology 6, 440–443 (1986).
Cherubini, A., Della Torre, S., Pelusi, S. & Valenti, L. Sexual dimorphism of metabolic dysfunction-associated steatotic liver disease. Trends Mol. Med. https://doi.org/10.1016/j.molmed.2024.05.013 (2024).
Della Torre, S. Non-alcoholic fatty liver disease as a canonical example of metabolic inflammatory-based liver disease showing a sex-specific prevalence: relevance of estrogen signaling. Front. Endocrinol. https://doi.org/10.3389/fendo.2020.572490 (2020).
Kuiper, G. G. et al. Comparison of the ligand binding specificity and transcript tissue distribution of estrogen receptors alpha and beta. Endocrinology 138, 863–870 (1997).
Gao, X. et al. Estrogen receptor α regulates metabolic-associated fatty liver disease by targeting NLRP3-GSDMD axis-mediated hepatocyte pyroptosis. J. Agric. Food Chem. 69, 14544–14556 (2021).
Della Torre, S. et al. An essential role for liver ERα in coupling hepatic metabolism to the reproductive cycle. Cell Rep. 15, 360–371 (2016).
Meda, C. et al. Hepatic ERα accounts for sex differences in the ability to cope with an excess of dietary lipids. Mol. Metab. 32, 97–108 (2020).
Winn, N. C. et al. Estrogen receptor-α signaling maintains immunometabolic function in males and is obligatory for exercise-induced amelioration of nonalcoholic fatty liver. Am. J. Physiol. Endocrinol. Metab. 316, e156–e167 (2019).
Erkan, G., Yilmaz, G., Konca Degertekin, C., Akyol, G. & Ozenirler, S. Presence and extent of estrogen receptor-alpha expression in patients with simple steatosis and NASH. Pathol. Res Pract. 209, 429–432 (2013).
Cherubini, A. et al. Interaction between estrogen receptor-α and PNPLA3 p.I148M variant drives fatty liver disease susceptibility in women. Nat. Med. 29, 2643–2655 (2023).
Walker, R. W. et al. A common variant in PNPLA3 is associated with age at diagnosis of NAFLD in patients from a multi-ethnic biobank. J. Hepatol. 72, 1070–1081 (2020).
Park, T., Chen, H. & Kim, H. -Y. GPR110 (ADGRF1) mediates anti-inflammatory effects of N-docosahexaenoylethanolamine. J. Neuroinflammation 16, 225 (2019).
Kim, H. Y. & Spector, A. A. N-Docosahexaenoylethanolamine: a neurotrophic and neuroprotective metabolite of docosahexaenoic acid. Mol. Asp. Med. 64, 34–44 (2018).
Kim, H. Y., Spector, A. A. & Xiong, Z. M. A synaptogenic amide N-docosahexaenoylethanolamide promotes hippocampal development. Prostaglandins Other Lipid Mediat. 96, 114–120 (2011).
Hamann, J. et al. International union of basic and clinical pharmacology. XCIV. Adhesion G protein-coupled receptors. Pharm. Rev. 67, 338–367 (2015).
Lin, H. H., Ng, K. F., Chen, T. C. & Tseng, W. Y. Ligands and beyond: mechanosensitive adhesion GPCRs. Pharmaceuticals https://doi.org/10.3390/ph15020219 (2022).
Boyden, S. E. et al. Vibratory urticaria associated with a missense variant in ADGRE2. N. Engl. J. Med. 374, 656–663 (2016).
Kwakkenbos, M. J. et al. Expression of the largest CD97 and EMR2 isoforms on leukocytes facilitates a specific interaction with chondroitin sulfate on B cells. J. Leukoc. Biol. 77, 112–119 (2005).
Wang, T. et al. CD97, an adhesion receptor on inflammatory cells, stimulates angiogenesis through binding integrin counterreceptors on endothelial cells. Blood 105, 2836–2844 (2005).
Luo, R. et al. G protein-coupled receptor 56 and collagen III, a receptor-ligand pair, regulates cortical development and lamination. Proc. Natl Acad. Sci. USA 108, 12925–12930 (2011).
El-Kassas, M. & Alswat, K. Letter to the Editor: a multi-society Delphi consensus statement on new fatty liver disease nomenclature. Hepatology https://doi.org/10.1097/HEP.0000000000000966 (2024).
Wang, Z. et al. Investigating linear and nonlinear associations of LDL cholesterol with incident chronic kidney disease, atherosclerotic cardiovascular disease and all-cause mortality: a prospective and Mendelian randomization study. Atherosclerosis 387, 117394 (2023).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Lee, J. H. et al. Hepatic steatosis index: a simple screening tool reflecting nonalcoholic fatty liver disease. Dig. Liver Dis. 42, 503–508 (2010).
Luengo-Mateos, M. et al. Protocol for ovariectomy and estradiol replacement in mice. STAR Protoc. 5, 102910 (2024).
Wang, Y., Li, X., Huang, R., Chen, X. W. & Wang, X. Apolipoprotein B secretion assay from primary hepatocytes. Bio Protoc. 14, e4982 (2024).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Li, J. et al. LONP1 regulation of mitochondrial protein folding provides insight into beta cell failure in type 2 diabetes. Nat. Metab. 7, 1570–1592 (2025).
Jung, S. et al. Dietary fibre-adapted gut microbiome clears dietary fructose and reverses hepatic steatosis. Nat. Metab. 7, 1801–1818 (2025).
Yang, F. et al. Hepatic GPR110 in sex disparity of MASH. figshare https://doi.org/10.6084/m9.figshare.28532336 (2025).
Acknowledgements
We thank the Core Facility and Technical Service Center for School of Life Sciences and Biotechnology and the Laboratory Animal Center at Shanghai Jiao Tong University. This work was supported by the National Key Research and Development Program of China (2022YFA1004800), National Natural Science Foundation of China (82370865, 82100906), the Natural Science Foundation of Shanghai by Shanghai Municipal Science and Technology Commission (22ZR1430400) to Z.H. and Program of Science and Technology Cooperation with Hong Kong, Macao and Taiwan (22410760500) to S.Z. The schematics in Figs. 2a, 4a, 5a, 7b and 8a–c and Extended Data Figs. 3a,n, 5a, 7d and 9a were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Z.H. conceptualized the study and supervised the project. F.Y. and W.W. carried out the research, analysed and interpreted the results. R.Q. and Q.G. performed the molecular dynamics stimulation study. R.D., G.J. and L.H. carried out the formal analysis of the UK Biobank SNP data and curated the dataset. F.Q., X.Y., S.H., Y.Z., S.J. and L.X. provided additional support for the animal experiments. C.Z. and D.X. provided support for the clinical experiment. D.W. and H.X.H. reviewed and edited the manuscript. C.L. analysed the human transcriptomic data. A.C. and L.V. discussed the human data and provided support for the human data interpretation. P.G., S.Z. and W.J. provided support for the human study and discussed the results. F.Y. and Z.H. wrote and edited the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Robert Schwabe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Revati Dewal, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Expression of GPR110 is reduced in human MASH liver and transcriptionally repressed by rosiglitazone.
a, Hepatic expression of GPR110 in patients with MASH and control individuals by analyzing the following GEO datasets: GSE159676 and GSE66676 (n = 54). b, Schematic diagram showing the locations of four predicted PPARγ response elements (PPREs, sequence indicated in red) within the promoter region of mouse Gpr110 gene as identified by in silico analysis with JASPAR (2024) and PROMO. c, Luciferase reporter assay on Gpr110 promoter in response to rosiglitazone treatment by truncation analysis. Hepa1-6 cells were transfected with luciferase-expressing vector driven by the Gpr110 promoter with different lengths as indicated. The cells were treated with rosiglitazone (20 μM) or DMSO control 4 hours post-transfection for 24 hours. The firefly luciferase activity was measured and normalized to renilla luciferase activity (n = 4/group). Data are presented as mean ± SD; Statistical significance was calculated using two-tailed Welch’s t-test (a) or two-tailed Student’s t-test (c).
Extended Data Fig. 2 AAV-mediated hepatic overexpression of GPR110 does not affect serum lipid levels in mice.
9-week-old male and female C57BL/6 J mice placed on STC were administered with AAV encoding Gpr110 (AAV-GPR110) or Gfp (AAV-GFP) driven by thyroxine-binding globulin (Tbg) promoter via tail-vein injection (2×1011 copies of AAV/mouse). After 1 week, the mice were switched to CDAHF60 diet for 12 weeks (n = 6/group). a,c, AAV-mediated selective overexpression of Gpr110 in the liver, but not in subcutaneous white adipose tissue (sWAT), epididymal white adipose tissue (eWAT), gastrocnemius muscle (Gastroc) or hypothalamus (Hypo) in male (a) and female (c) mice was confirmed by quantitative real-time PCR analysis. b,d, Hepatic protein expression of GPR110 induced by AAV-GPR110 in male (b) and female (d) mice was validated by western blot. The bottom panels are the densitometric analyses for the relative abundance of GPR110 normalized with GAPDH. e-h, Serum levels of triglyceride (TG) and total cholesterol (TC) in male (e,f) and female (g,h) mice. Data are presented as mean ± SD; Statistical significance was calculated using two-tailed Student’s t-test (a-h).
Extended Data Fig. 3 AAV-mediated hepatic overexpression of GPR110 exacerbates MASH development and progression in female mice, but not male mice.
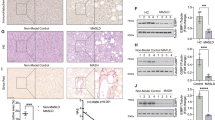
9-week-old male and female C57BL/6 J mice placed on STC were administered with AAV encoding Gpr110 (AAV-GPR110) or Gfp (AAV-GFP) driven by thyroxine-binding globulin (Tbg) promoter via tail-vein injection (2×1011 copies of AAV/mouse). One week later, mice were switched to CDAHF60 diet for 12 weeks (n = 6/group). a,n, Experimental design for male mice (a) and female mice (n). b,o, Body weight of male (b) and female (o) mice. c,p, Liver/body weight ratio of male (c) and female (p) mice. d,e,q,r, Serum ALT and AST levels in male (d,e) and female (q,r) mice. f,s, Representative images of H&E staining (top), immunohistochemistry for F4/80 (middle) and Sirius Red staining (bottom) of liver sections from male (f) and female (s) mice; Scale bars, 100 μm. g-k, NAFLD activity score (g), hepatic steatosis score (h), inflammatory foci per field (i), percentage of F4/80-positive area (j) and percentage of fibrosis area (k) in the liver of male mice. t-x, NAFLD activity score (t), hepatic steatosis score (u), inflammatory foci per field (v), percentage of F4/80-positive area (w) and percentage of fibrosis area (x) in the liver of female mice. l,y, Hepatic triglyceride (TG) content in male (l) and female (y) mice. m,z, Hepatic hydroxyproline content in male (m) and female (z) mice. Data are presented as mean ± SD; Statistical significance was calculated using two-tailed Student’s t-test (b-e,g-m,o-r,t-z). Schematic in a and n created with BioRender.com.
Extended Data Fig. 4 Generation and characterization of Gpr110 LKO mice.
a, Schematic diagram for the generation of hepatocyte-specific Gpr110 knockout (LKO) mice. loxP sites were inserted upstream and downstream of exon 4 of Gpr110. Locations of the primers for PCR analysis of the wild-type (WT) and Gpr110-floxed alleles are indicated. The primers F1 and R1 detect the presence of the loxP region and amplify a band of 320 bp in Gpr110-floxed mice and 286 bp in WT mice. A set of Cre-recombinase (Cre) coding sequence-specific primers amplifies a band of 390 bp in Cre+ mice. b, Genotyping of LKO mice by genomic PCR analysis. c, Selective deletion of Gpr110 in the liver, but not in eWAT, colon or Hypo in LKO mice was confirmed by RT-qPCR analysis (n = 4 mice/group). d, Gpr110 mRNA expression in hepatocytes and non-parenchymal cells (NPCs) from LKO and WT mice (n = 4/group). e, Reduction of GPR110 in the liver of LKO mice as compared to the WT mice was validated at the protein level (n = 3/group). f-k, 10-week-old male and female LKO mice and their Gpr110fl/fl (WT) littermates were fed STC or CDAHF60 for 12 weeks (n = 5/group for male and female WT and LKO mice on STC, n = 9/group for male WT mice on CDAHF60, n = 7/group for male LKO mice on CDAHF60, n = 11/group for female WT mice on CDAHF60, n = 13/group for female LKO mice on CDAHF60). f,i, Body weight of male (f) and female (i) mice. g,h,j,k, Serum levels of TG and TC in male (g,h) and female (j,k) mice. l,m, Livers from male WT or LKO mice fed STC or CDAHF60 were harvested for transcriptomic analysis. Differentially expressed genes (DEGs) were identified (Adjusted p value < 0.05 and fold change > 1.5) (n = 3/group). GO enrichment analysis of DEGs induced by GPR110 ablation in male (upper) and female (lower) mice (l). Venn diagram identifies the genes induced by MASH and reversed in LKO or suppressed by MASH and reversed in LKO mice (m). Data are presented as mean ± SD; Statistical significance was calculated using two-tailed Student’s t-test (c-e) or one-way ANOVA followed by Tukey’s multiple comparisons test (f-k), one-sided p values by hypergeometric test and performed adjustments for multiple testing (l).
Extended Data Fig. 5 Loss of hepatic Gpr110 ameliorates MASH and its related hepatic fibrosis induced by an CDHF45 diet in female mice, but not in male mice.
10-week-old male and female LKO mice and their WT littermates were fed with STC or choline-deficient high-fat (CDHF45) diet for 32 weeks (n = 5/group for male and female WT and LKO mice on STC, n = 6/group for male and female WT and LKO mice on CDHF45 for b-y). a, Experimental design. b,d, Body weight in male (b) and female (d) mice. c,e, Liver/body weight ratio of male (c) and female (e) mice. f-i, Serum ALT and AST levels in male (f,g) and female (h,i) mice. j,r, Representative images of H&E staining (top), immunohistochemistry for F4/80 (middle) and Sirius Red staining (bottom) of liver sections from male (j) and female (r) mice; Scale bars, 100 μm. k-o, NAFLD activity score (k), hepatic steatosis score (l), inflammatory foci per field (m), percentage of F4/80-positive area (n) and percentage of fibrosis area (o) in the liver of male mice. s-w, NAFLD activity score (s), hepatic steatosis score (t), inflammatory foci per field (u), percentage of F4/80-positive area (v) and percentage of fibrosis area (w) in the liver of female mice. p,x, Hepatic TG content in male (p) and female (x) mice. q,y, Hepatic hydroxyproline content in male (q) and female (y) mice. z,aa, Western blot analysis for α-SMA protein expression in the liver of male (a) and female (aa) mice. The right panels are the densitometric analyses for the relative abundance of α-SMA normalized with GAPDH (n = 3/group). Data are presented as mean ± SD; Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test (b-i,k-q,s-aa). Schematic in a created with BioRender.com.
Extended Data Fig. 6 Loss of hepatic Gpr110 has no obvious effects on serum lipid and glucose levels or glucose metabolism in male or female mice.
10-week-old male and female LKO mice and their WT littermates were fed with STC or choline-deficient high-fat (CDHF45) diet for 32 weeks. a,b,d,e, Serum levels of TG and FFA in male (a,b) and female (d,e) mice. c,f, Blood glucose levels at fed and fasting states in male (c) and female (f) mice. g-l, Serum levels of TC, HDL and LDL in male (g,h,i) and female (j,k,l) mice. m,n, Glucose tolerance test (GTT) performed at the 30th week of study and the area under curve (AUC) during GTT in male (m) and female (n) mice. o,p, Insulin tolerance test (ITT) performed at the last week of study and AUC during ITT in male (o) and female (p) mice. n = 5/group for male and female WT and LKO mice on STC, n = 6/group for male and female WT and LKO mice on CDHF45. Data are presented as mean ± SD; Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test (a-l,m-p right panels) or two-way ANOVA followed by Tukey’s multiple comparisons test (m-p left panels).
Extended Data Fig. 7 Global deletion of Gpr110 ameliorates CDAHF60-induced MASH in female mice.
a, Schematic diagram for the generation of global Gpr110 knockout (GKO) mice. A pair of sgRNAs targeting exon 4 of Gpr110 gene was designed. The CRISPR/Cas9 system was utilized to produce GKO mice. Locations of the primers for PCR analysis of the wild-type (WT) and GKO alleles are indicated. The primers F1 and R1 detect the presence of exon 4 and amplify a band of 447 bp in WT mice. The primers F2 and R2 spanning the exon 4 amplify a band of 280 bp in GKO mice. b, Genotyping of GKO mice by genomic PCR analysis. c, Deletion of GPR110 in the liver of GKO mice at the protein level was validated (n = 3/group). d-ab, 10-week-old male and female GKO mice and their WT littermates were fed STC or CDAHF60 for 12 weeks (n = 5/group for male and female WT and GKO mice on STC, n = 7/group for male WT mice on CDAHF60, n = 5/group for male GKO mice on CDAHF60, n = 7/group for female WT mice on CDAHF60, n = 6/group for female GKO mice on CDAHF60 for e-x). d, Experimental design. e,h, Liver/body weight ratio of male (e) and female (h) mice. f,g,i,j, Serum ALT and AST levels in male (f,g) and female (i,j) mice. k,s, Representative images of H&E staining (top), immunohistochemistry for F4/80 (middle) and Sirius Red staining (bottom) of liver sections from male (k) and female (s) mice; Scale bars, 100 μm. l-p, NAFLD activity score (l), hepatic steatosis score (m), inflammatory foci per field (n), percentage of F4/80-positive area (o) and percentage of fibrosis area (p) in the liver of male mice. t-x, NAFLD activity score (t), hepatic steatosis score (u), inflammatory foci per field (v), percentage of F4/80-positive area (w) and percentage of fibrosis area (x) in the liver of female mice. q,y, Hepatic TG content in male (q) and female (y) mice. r,z, Hepatic hydroxyproline content in male (r) and female (z) mice. aa,ab, Western blot analysis for α-SMA protein expression in the liver of male (aa) and female (ab) mice. The right panels are the densitometric analyses for the relative abundance of α-SMA normalized with GAPDH (n = 3/group). Data are presented as mean ± SD; Statistical significance was calculated using two-tailed Student’s t-test (c) or one-way ANOVA followed by Tukey’s multiple comparisons test (e-j,l-r,t-ab). Schematic in d and n created with BioRender.com.
Extended Data Fig. 8 GPR110 expression in male and female mice.
a,b, The expression of GPR110 at mRNA (a) and protein (b) levels in the liver of WT male and female mice fed with STC or CDAHF60 diet for 12 weeks; The expression levels in all other groups were compared to the expression level in male-STC group (n = 3/group). c,d, The mRNA expression levels of Esr2 (c) and Gpr30 (d) in the liver of WT or LKO male (left panel) and female (right panel) mice fed with either STC or CDAHF60 diet; The expression levels in all other groups were compared to the expression level in WT-STC male group (n = 5/group for male and female WT and LKO mice on STC, n = 9/group for male WT mice on CDAHF60, n = 7/group for male LKO mice on CDAHF60, n = 11/group for female WT mice on CDAHF60, n = 13/group for female LKO mice on CDAHF60). Data are presented as mean ± SD; Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test (a-d).
Extended Data Fig. 9 Gender differences in the influence of Gpr110 deletion on liver phenotypes are imposed by the oestrogen signaling.
8-week-old female LKO and WT mice underwent ovariectomy (OVX) or sham operation. After 2-week recovery, all mice were placed on CDAHF60 diet (n = 6/group). a, Experimental design. b, Liver/body weight ratio of these mice. c,d, Serum levels of ALT (c) and AST (d). e, Representative images of H&E staining (top), immunohistochemistry for F4/80 (middle) and Sirius Red staining (bottom) of liver sections; Scale bars, 100 μm. f-j, NAFLD activity score (f), hepatic steatosis score (g), inflammatory foci per field (h), percentage of F4/80-positive area (i) and percentage of fibrosis area (j) in the liver. k,l, Hepatic contents of TG (k) and hydroxyproline (l). Data are presented as mean ± SD; Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test (b-d,f-l). Schematic in a created with BioRender.com.
Extended Data Fig. 10 Association of GPR110 rs937057 T > C variant with MASLD and its functional impact.
a, Representative allelic discrimination plot showing genotypes of samples for the GPR110 variant rs937057 T > C (p.I571M); Homozygous: T/T, C/C; Heterozygous: T/C. b, Allele frequencies of rs937057 T > C in MASLD and control groups in the case control cohort (n = 396). c,d, Molecular dynamic simulation on I571M mutation on the autoactivation of GPR110567-910. c, Root mean square deviation (RMSD) values of Cα atoms for wild-type I571stalk GPR110567-910 (green) and stalk mutation I571stalkM GPR110567-910 (cyan). d, The molecular dynamics (MD) stimulated structures of wild-type I571stalk GPR110567-910 and stalk mutation I571stalkM GPR110567-910 were superimposed, showing a deviation of 2.213 Å. Two different side views were shown in the left and middle panels, and the top view was shown in the right panel. e, Human female hepatoma Mahlavu cells were transfected with plasmids overexpressing WT GPR110 (I571), mutant GPR110 (M571) or GFP as the control for 24 hours for the detection of GPR110 mRNA expression (n = 6/group). Data are presented as mean ± SD; Statistical analyses were performed using two-sided χ2 test (b); Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test (e).
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–6, Extended Data Fig. 1–10 legends and Supplementary Figs. 1–10.
Supplementary Data 2 (download XLSX )
Supplementary Fig. 2: Hepatic expression of Scd1 in different interventions. a,b, Hepatic expression of Scd1 in the livers of male (a) and female (b) LKO and WT mice fed with 12-week CDAHF60 (n = 9 per group for male WT mice, n = 7 per group for male LKO mice, n = 11 per group for female WT mice, n = 13 per group for female LKO mice). c,d, Hepatic expression of Scd1 in the livers of male (c) and female (d) mice with AAV-mediated overexpression of GPR110 or GFP control fed with 12-week CDAHF60 (n = 6 per group for male mice receiving AAV, n = 8 per group for female mice receiving AAV). Data are presented as the mean ± s.d. Statistical significance was calculated using a two-tailed Student’s t-test (a–d).
Supplementary Data 3 (download XLSX )
Supplementary Fig. 3: Loss of Gpr110 in hepatocytes ameliorates hepatic steatosis in female, but not male mice. Ten-week-old male and female LKO mice and their WT littermates were fed with CDAHF60 diet for 2 weeks (n = 6 per group for male WT and LKO mice, n = 7 per group for female WT mice, n = 6 per group for female LKO mice). a,e, Liver/body weight ratio of male (a) and female (e) mice. b,c,f,g, Serum ALT and AST levels in male (b and c) and female (f and g) mice. d,h, Serum TG level of male (d) and female (h) mice. i,l, Representative images of H&E staining (top) and Oil-Red O staining (bottom) of liver sections from male (i) and female (l) mice; Scale bars, 100 μm. j,m, hepatic steatosis score in the liver of male (j) and female (m) mice. k,n, Hepatic TG content in male (k) and female (n) mice. o,p, Real-time PCR analysis of liver tissues for genes involved in the regulation of lipid metabolism in male (o) and female (p) mice. Data are presented as the mean ± s.d. Statistical significance was calculated using a two-tailed Student’s t-test (a–h, j, k and m–p).
Supplementary Data 4 (download XLSX )
Supplementary Fig. 4: AAV-mediated delivery of shRNA specifically knocks down Esr1 expression in the liver. a,b, Hepa1-6 cells were transfected with plasmids encoding shRNA against various sequences in Esr1 (shEsr1-1, shEsr1-2 and shEsr1-3) or scramble sequence (Scramble). The expression of ERα at mRNA (a) and protein (b) levels in the cells were determined by real-time qPCR and western blot analyses (n = 3 per group). shEsr1-1 was selected for subsequent AAV packaging as it demonstrated highest Esr1 knockdown efficiency. c,d, 9-week-old female WT mice were infected with AAV encoding shRNA against Esr1 (AAV-shEsr1) or with scramble sequence (AAV-Scramble) via tail-vein injection (2 × 1011 genome copies of AAV per mouse; n = 4 per group). Three weeks after infection, the mRNA expression of Esr1 in the liver, gastrocnemius and ovaries (c) and the protein expression of ERα in the liver (d) was measured by real-time qPCR and western blot, respectively. e–g, 9-week-old female LKO and WT mice fed with STC were administered with AAV-shEsr1 or AAV-Scramble via tail-vein injection. One week after injection, the mice were switched to a CDAHF60 diet for 12 weeks (n = 8 per group for WT mice receiving Scramble, n = 7 per group for LKO mice receiving Scramble and WT/LKO mice receiving shEsr1). e, Body weight of the mice. f,g, Serum levels of TG (f) and TC (g) in the mice. Data are presented as the mean ± s.d. Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple-comparisons test (a, b and e–g) or two-tailed Student’s t-test (c and d).
Supplementary Data 5 (download XLSX )
Supplementary Fig. 5: The NFAT2–ERα signalling is specifically coupled to GPR110, but not general PKA activation in hepatocytes. Hepa1-6 cells transfected with GFP or GPR110 were treated with P12 (100 μM) or forskolin (FSK, 5 μM) as indicated for 2 h. a–c, The cells were subjected to the detection of PKA activity (n = 4 per group; a), NFAT2 phosphorylation (n = 3 per group; b) and Esr1 mRNA expression (n = 6 per group; c). Data are presented as the mean ± s.d. Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple-comparisons test (a–c).
Supplementary Data 6 (download XLSX )
Supplementary Fig. 6: Effects of P12 on hepatic PKA–NFAT2–ERα signalling in vivo. Ten-week-old female C57BL/6J mice were injected with P12 (low dose: 2 mg per kg, high dose: 5 mg per kg) or PBS control via tail vein. a–c, Liver tissues were collected 4 h after injection for the detection of PKA activity (n = 4 per group; a), NFAT2 phosphorylation (n = 3 per group; b) and Esr1 mRNA expression (n = 4 per group; c). Data are presented as the mean ± s.d. Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple-comparisons test (a–c).
Supplementary Data 8 (download XLSX )
Supplementary Fig. 8: Knockdown efficiency of siRNAs against Gpr110 in vitro. a,b, Hepa1-6 (a) and AML12 (b) cells were transfected with siRNA against various sequences in mouse Gpr110 (siRNA-1, siRNA-2 and siRNA-3) or scramble sequence (Scramble). The mRNA expression of Gpr110 in the cells were determined by real-time qPCR (n = 3 per group). siRNA-3 was selected for subsequent chemical modifications and GalNAc conjugation as it demonstrated highest Gpr110 knockdown efficiency. Data are presented as the mean ± s.d.; Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple-comparisons test (a and b).
Supplementary Data 9 (download XLSX )
Supplementary Fig. 9: GalNAc-conjugated siRNA against Gpr110 suppresses hepatic expression of genes related to inflammation and fibrosis in female mice. Male and female mice fed with CDAHF60 for 2 weeks were subjected to siGpr110 or Scramble via subcutaneous injection (5 mg per kg body weight biweekly, four doses). These mice were maintained on CDAHF60 for another 8 weeks (n = 6 per group for male mice receiving Scramble or siGpr110, n = 8 per group for female mice receiving Scramble or siGpr110 for a–d). Hepatic mRNA expression of genes related to inflammation (a and c) and fibrosis (b and d) in male (a and b) and female (c and d) mice. Data are presented as the mean ± s.d. Statistical significance was calculated using two-tailed Student’s t-test (a–d).
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, F., Wang, W., Qiu, F. et al. Hepatic GPR110 contributes to sex disparity in the development of MASH through oestrogen receptor α-dependent signalling. Nat Metab 8, 116–138 (2026). https://doi.org/10.1038/s42255-025-01436-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01436-1
This article is cited by
-
A sex-specific brake on liver oestrogen signalling
Nature Metabolism (2026)
-
GPR110 as a regulator of MASH
Nature Reviews Endocrinology (2026)