Abstract
Acute pancreatitis-induced acinar-to-ductal metaplasia involves global chromatin remodeling and contributes to normal tissue regeneration. Oncogenic KRAS hijacks this process to promote PDAC formation. Here we show that regeneration and KRASG12D-driven oncogenesis can be decoupled from tissue regeneration through a chromatin remodeler, SMARCA5. We show that SMARCA5 maintains KRASG12D-dependent chromatin accessibility at regions specifically required for malignancy, without affecting chromatin opening that occurs during normal regeneration. Without SMARCA5, regeneration can be restored in the presence of KRASG12D. Mechanistically, regeneration-related or malignancy-related chromatin remodeling activities occur in a time-sensitive manner. The activity of SMARCA5 is controlled spatiotemporally by transcription factor RUNX1, which only accumulates at sufficient levels with sustained MAPK signals. We further show that inhibition of the SMARCA5-containing NoRC complex specifically inhibits the growth of PDAC organoid but not that of normal tissue derived from patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Proteomics data are available via ProteomeXchange with identifier PXD047272. All ATAC-seq, RNA-seq, ChIP-seq, CUT&RUN and CUT&Tag data are deposited at the Gene Expression Omnibus (GEO) under series GSE248948. Accession codes of the published data in GEO reanalyzed in this study are as follows: single-cell RNA-seq of mouse PDAC, GSE166298; ATAC-seq of PDAC cells, GSE134230; RNA-seq of duct, ADM day 2, ADM day 7 and PDAC cells, GSE134235. All other data supporting the findings of this study are available from the corresponding author on reasonable request. All unique materials generated in this study are available from the corresponding authors upon request. Source data are provided with this paper.
Code availability
The following open-source code and software were used in this study: Trim Galore (v.0.6.4), Bowtie2 (v.2.3.5.1), STAR (v.2.7.9a), SAMtools (v.1.10), featureCounts (v.2.0.1), Genrich (v.0.6), MACS2 (v.2.2.7.1), deepTools (v.3.4.3), Bedtools (v2.29.2), HOMER (v.2.0.1) and GSEA (v.4.2.2). R package DESeq2 (v.1.24.0), R package pheatmap (v.1.0.12), R package ComplexHeatmap (v.2.2.0), R package Seurat (v.4.1.3) and R package AUCell package (v.1.16.0) were also used.
References
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Merrell, A. J. & Stanger, B. Z. Adult cell plasticity in vivo: de-differentiation and transdifferentiation are back in style. Nat. Rev. Mol. Cell Biol. 17, 413–425 (2016).
Storz, P. Acinar cell plasticity and development of pancreatic ductal adenocarcinoma. Nat. Rev. Gastroenterol. Hepatol. 14, 296–304 (2017).
Morris, J. P. T., Cano, D. A., Sekine, S., Wang, S. C. & Hebrok, M. ß-catenin blocks Kras-dependent reprogramming of acini into pancreatic cancer precursor lesions in mice. J. Clin. Invest. 120, 508–520 (2010).
Giroux, V. & Rustgi, A. K. Metaplasia: tissue injury adaptation and a precursor to the dysplasia-cancer sequence. Nat. Rev. Cancer 17, 594–604 (2017).
Strobel, O. et al. In vivo lineage tracing defines the role of acinar-to-ductal transdifferentiation in inflammatory ductal metaplasia. Gastroenterology 133, 1999–2009 (2007).
Rooman, I. & Real, F. X. Pancreatic ductal adenocarcinoma and acinar cells: a matter of differentiation and development?. Gut 61, 449–458 (2012).
Alonso-Curbelo, D. et al. A gene-environment-induced epigenetic program initiates tumorigenesis. Nature 590, 642–648 (2021).
Li, Y. et al. Mutant Kras co-opts a proto-oncogenic enhancer network in inflammation-induced metaplastic progenitor cells to initiate pancreatic cancer. Nat. Cancer 2, 49–65 (2021).
Halbrook, C. J. et al. Mitogen-activated protein kinase kinase activity maintains acinar-to-ductal metaplasia and is required for organ regeneration in pancreatitis. Cell. Mol. Gastroenterol. Hepatol. 3, 99–118 (2017).
Li, Y. et al. The emerging role of ISWI chromatin remodeling complexes in cancer. J. Exp. Clin. Cancer Res. 40, 346 (2021).
Clapier, C. R., Iwasa, J., Cairns, B. R. & Peterson, C. L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 18, 407–422 (2017).
Baldi, S. et al. Genome-wide rules of nucleosome phasing in Drosophila. Mol. Cell 72, 661–672 e664 (2018).
Barisic, D., Stadler, M. B., Iurlaro, M. & Schubeler, D. Mammalian ISWI and SWI/SNF selectively mediate binding of distinct transcription factors. Nature 569, 136–140 (2019).
Bomber, M. L. et al. Human SMARCA5 is continuously required to maintain nucleosome spacing. Mol. Cell https://doi.org/10.1016/j.molcel.2022.12.018 (2023).
Racki, L. R. et al. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature 462, 1016–1021 (2009).
Klement, K. et al. Opposing ISWI- and CHD-class chromatin remodeling activities orchestrate heterochromatic DNA repair. J. Cell Biol. 207, 717–733 (2014).
Alvarez-Saavedra, M. et al. Snf2h-mediated chromatin organization and histone H1 dynamics govern cerebellar morphogenesis and neural maturation. Nat. Commun. 5, 4181 (2014).
Zhang, C. et al. The chromatin remodeler Snf2h is essential for oocyte meiotic cell cycle progression. Genes Dev. 34, 166–178 (2020).
Ding, Y., Li, Y., Zhao, Z., Cliff Zhang, Q. & Liu, F. The chromatin-remodeling enzyme Smarca5 regulates erythrocyte aggregation via Keap1-Nrf2 signaling. Elife https://doi.org/10.7554/eLife.72557 (2021).
Iurlaro, M. et al. Systematic assessment of ISWI subunits shows that NURF creates local accessibility for CTCF. Nat. Genet. 56, 1203–1212 (2024).
Liu, X. et al. Aberrant accumulation of Kras-dependent pervasive transcripts during tumor progression renders cancer cells dependent on PAF1 expression. Cell Rep. https://doi.org/10.1016/j.celrep.2023.112979 (2023).
Zhuo, M., Mao, J. & David, C. J. Protocol to package and concentrate adeno-associated virus serotype 8 for use in autochthonous mouse models of pancreatic cancer. STAR Protoc. https://doi.org/10.1016/j.xpro.2023.102108 (2023).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Miyabayashi, K. et al. Intraductal transplantation models of human pancreatic ductal adenocarcinoma reveal progressive transition of molecular subtypes. Cancer Discov. 10, 1566–1589 (2020).
Raghavan, S. et al. Microenvironment drives cell state, plasticity, and drug response in pancreatic cancer. Cell 184, 6119–6137 e6126 (2021).
Moffitt, R. A. et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 47, 1168–1178 (2015).
Chan-Seng-Yue, M. et al. Transcription phenotypes of pancreatic cancer are driven by genomic events during tumor evolution. Nat. Genet. 52, 231–240 (2020).
Del Poggetto, E. et al. Epithelial memory of inflammation limits tissue damage while promoting pancreatic tumorigenesis. Science 373, eabj0486 (2021).
He, P., Yang, J. W., Yang, V. W. & Bialkowska, A. B. Kruppel-like factor 5, increased in pancreatic ductal adenocarcinoma, promotes proliferation, acinar-to-ductal metaplasia, pancreatic intraepithelial neoplasia, and tumor growth in mice. Gastroenterology 154, 1494–1508 e1413 (2018).
Kopp, J. L. et al. Identification of Sox9-dependent acinar-to-ductal reprogramming as the principal mechanism for initiation of pancreatic ductal adenocarcinoma. Cancer Cell 22, 737–750 (2012).
Li, X. et al. COL1A1: a novel oncogenic gene and therapeutic target in malignancies. Pathol. Res. Pract. 236, 154013 (2022).
Zhao, Y. et al. Distinct molecular profiles drive multifaceted characteristics of colorectal cancer metastatic seeds. J. Exp. Med. https://doi.org/10.1084/jem.20231359 (2024).
Jia, Y. et al. In vivo CRISPR screening identifies BAZ2 chromatin remodelers as druggable regulators of mammalian liver regeneration. Cell Stem Cell 29, 372–385 e378 (2022).
Chen, P. et al. Discovery and characterization of GSK2801, a selective chemical probe for the bromodomains BAZ2A and BAZ2B. J. Med. Chem. 59, 1410–1424 (2016).
Wells, J. M. & Watt, F. M. Diverse mechanisms for endogenous regeneration and repair in mammalian organs. Nature 557, 322–328 (2018).
Ge, Y. & Fuchs, E. Stretching the limits: from homeostasis to stem cell plasticity in wound healing and cancer. Nat. Rev. Genet. 19, 311–325 (2018).
Brahma, S. & Henikoff, S. Epigenome regulation by dynamic nucleosome unwrapping. Trends Biochem. Sci. 45, 13–26 (2020).
Yan, L. & Chen, Z. A unifying mechanism of DNA translocation underlying chromatin remodeling. Trends Biochem. Sci. 45, 217–227 (2020).
Mueller, S. et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature 554, 62–68 (2018).
Fu, Y., Sinha, M., Peterson, C. L. & Weng, Z. The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet. 4, e1000138 (2008).
Reichert, M. et al. Isolation, culture and genetic manipulation of mouse pancreatic ductal cells. Nat. Protoc. 8, 1354–1365 (2013).
Han, J. SMARCA5 paper. protocols.io https://doi.org/10.17504/protocols.io.n92ld623ng5b/v1 (2025).
Liu, X. et al. The PAF1 complex promotes 3′ processing of pervasive transcripts. Cell Rep. https://doi.org/10.1016/j.celrep.2022.110519 (2022).
Li, B., Ren, N., Yang, L., Liu, J. & Huang, Q. A qPCR method for genome editing efficiency determination and single-cell clone screening in human cells. Sci. Rep. https://doi.org/10.1038/s41598-019-55463-6 (2019).
Chen, Q. et al. LYL1 facilitates AETFC assembly and gene activation by recruiting CARM1 in t(8;21) AML. Proc. Natl Acad. Sci. USA 119, e2213718119 (2022).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Zhao, Y. et al. Colorectal cancer patient-derived 2D and 3D models efficiently recapitulate inter- and intratumoral heterogeneity. Adv. Sci. https://doi.org/10.1002/advs.202201539 (2022).
Klöppel, G. & Adsay, N. V. Chronic pancreatitis and the differential diagnosis versus pancreatic cancer. Arch. Pathol. Lab. Med. 133, 382–387 (2009).
Gopinathan, A., Morton, J. P., Jodrell, D. I. & Sansom, O. J. GEMMs as preclinical models for testing pancreatic cancer therapies. Dis. Models Mech. 8, 1185–1200 (2015).
Basturk, O. et al. A revised classification system and recommendations from the Baltimore consensus meeting for neoplastic precursor lesions in the pancreas. Am. J. Surg. Pathol. https://doi.org/10.1097/pas.0000000000000533 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience https://doi.org/10.1093/gigascience/giab008 (2021).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Martin, F. J. et al. Ensembl 2023. Nucleic Acids Res. 51, D933–D941 (2023).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. https://doi.org/10.1186/gb-2008-9-9-r137 (2008).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Chen, Y. et al. Type I collagen deletion in αSMA(+) myofibroblasts augments immune suppression and accelerates progression of pancreatic cancer. Cancer Cell 39, 548–565 e546 (2021).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–W560 (2019).
Chandrashekar, D. S. et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25, 18–27 (2022).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
We thank members of the David and Chen laboratories for discussion and comments during the SMARCA5 study, and the Animal Center, Flow Cytometry Facility and Mass Spectrometry Facility at Tsinghua University for their support. We thank Z. Chen (MOE Key Laboratory of Protein Science, School of Life Sciences, Tsinghua University, Beijing, China) for the SMARCA5-expressing plasmid. This work was funded by Beijing Municipal Natural Science Foundation (grant JQ23024 to M.C.), National Science and Technology Major Project (2025ZD0544700 to M.C.), Beijing Municipal Science & Technology Commission (grant Z231100007223008 to M.C.), National Key R&D Program of China (grant 2021YFA1300100 to M.C.), Natural Science Foundation of China (grant 92068114 to M.C), Tsinghua Dushi Program (to M.C. and C.J.D.), National Key R&D Program of China (grant 20211250193 to C.J.D.) and the Natural Science Foundation of China (grant 20231100054 to C.J.D.).
Author information
Authors and Affiliations
Contributions
M.C. and C.J.D. conceived the project. W.X., C.J.D. and M.C. supervised the project. J.H., M.C. and C.J.D. designed the experiments. J.H. performed most of the experiments and bioinformatics analysis with the help from X. Lu and S.W. M.Z. provided single-cell RNA-seq analyses of mouse PDAC. Y.L. generated data of ATAC-seq and small-scale RNA-seq of mouse acinar cells and PanIN cells and provided PanIN tissue samples for H&E and IF. X. Liu provided ATAC-seq data in PDAC cell lines upon CHD4 KO and performed Chd4 KO tumor formation assay. J.M. performed Ino80 KO tumor formation assay. Q.C. performed co-IP in Extended Data Fig. 8b. M.G. performed some of the IF experiments and quantification. D.Z. provided mouse tissue samples for Extended Data Fig. 1l,m. J.W. and X.W. performed the initial CUT&RUN experiments and shared the protein A-MNase for CUT&RUN. J.P. and R.H. provided human PDAC tissue sections for IF. M.G. and X.W. supervised and performed the drug treatment of human patient organoid experiments. H.J., M.C. and C.J.D. prepared most of the figures and wrote the manuscript with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Cigall Kadoch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SMARCA5 is a specific regulator of PDAC initiation.
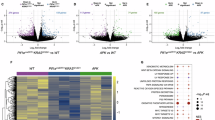
a, Results from a CRISPR KO screening22 showing the relative representation of sgRNAs targeting different chromatin remodelers in a PDAC cell line. b, FPKM of marker genes in mCherryneg-acinar cells (see Method for details). n = 2 mice with 3 technical replicates each. One dot represents the average of the three technical replicates. Short horizontal lines indicate the mean. c, Western blotting indicating SMARCA5 and SMARCA4 depletion after sgRNA treatment. d, RT-qPCR showing depletion efficiencies of sgRNA-treated PDAC cells. e, ATAC-seq signals at chromatin regions more accessible in tumor than acinar (n = 59735) in the mouse PDAC cell line treated with another set of sgRNAs, in relation to Fig. 1a. f, ChIP-seq reads for SMARCA4 and SMARCA5 on chromatin regions that are shown in Fig. 1a. g, ATAC-seq signals at chromatin regions more accessible in tumor than acinar which are bound by SMARCA5 (right) or SMARCA4 (left). h, The numbers of overlapping ATAC-seq peaks regulated by either SMARCA4 or SMARCA5 in PDAC cell line. i, ATAC-seq signal at chromatin regions shared by tumor and acinar cells (n = 39586). j, SMARCA5-regulated peaks in “806” cells, with Smarca5 KO rescued with either WT SMARCA5 (SMARCA5WT) or enzymatically dead SMARCA5 (SMARCA5K211R). k, ATAC-seq signals at indicated chromatin regions in the mouse PDAC cell line “Scf” treated with indicated sgRNAs. l, Representative IF images of SMARCA5, amylase and KRT19. m, Representative IHC staining of SMARCA5. l and m, Scale bar, 20 μm. n and o, Box plots showing quantification of SMARCA5 signals in pancreatic cells. The box in box plot shows the interquartile range, spanning from the first quartile to the third quartile. The line inside the box represents the median, while the whiskers extend from the box to the minimum and maximum values of the dataset. For n, Acinar: n = 91 cells; ADM: n = 503 cells; PanIN: n = 224 cells; Tumor: n = 333 cells. For o, Acinar: n = 45 cells; ADM: n = 101 cells; PanIN: n = 121 cells; Tumor: n = 207 cells.
Extended Data Fig. 2 SMARCA5 is a specific regulator of PDAC initiation without affecting normal pancreas.
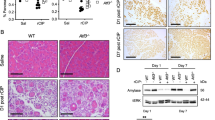
a, Representative IHC staining of SMARCA4 and SMARCA5 in PDAC patient samples. Red asterisks indicate tumor cells, and blue arrowheads indicate stromal cells. Data were obtained from The Human Protein Atlas (https://www.proteinatlas.org). Scale bar, 50μm. b, Quantification of SMARCA4 and SMARCA5 intensities in tumor versus stromal cells. Each dot represents a cell (SMARCA5 quantification: n = 29 for tumor cells, n = 25 for stroma cells; SMARCA4 quantification: n = 25 for tumor cells, n = 28 for stroma cells). c, Representative IF staining for SMARCA4 in mouse normal pancreas. White arrows indicate the nuclei of acinar cells. Scale bar, 5μm. d, Representative IF co-staining for GFP and SMARCA5 in Rosa and Smarca5 knockout samples obtained from tumor formation assays described in Fig. 1d. Scale bars, left 200μm, right 20μm. e, Representative IF staining for SMARCA4 in mouse ADM Day 2 samples. Scale bar, 50μm. f, Representative H&E staining in the presence of sgRNAs targeting Trp53, Cdkn2a and the indicated experimental sgRNAs. Scale bar, 100 μm. g, Representative IF co-staining for GFP and SMARCA5 in sgRNA treated samples obtained from survival assay described in Figs. 1f, g at the time of endpoint (mice sacrificed with tumor burden). Scale bar, 50μm. h, Quantification of SMARCA5 intensity in GFP-positive cells from survival assay. n = 3414 cells for sgRosa; n = 5707 cells for sgSmarca5. i, Representative flow cytometric data displaying the percentage of RFP and BFP in Rosa and Smarca5 knockout samples described Fig. 1i at the indicated time points. j, Representative IF co-staining for GFP, RFP and amylase, corresponding to Fig. 1i. Scale bar, 50μm. k, Quantification of amylase intensity in GFP+&RFP+ cells. Significance was determined by one-way ANOVA. n = 3 biological independent mice. Data are present as mean ± SEM, with each dot representing one mouse.
Extended Data Fig. 3 Smarca5 ablation does not affect ADM formation and regeneration in KRASwt context and SMARCA5 is activated by KRAS signaling.
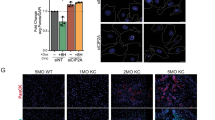
a, Representative H&E staining and IHC staining for GFP of the pancreas from the Ca mice treated with indicated AAV-Cre-sgRNA viruses in KRASwt context described in Fig. 2a. Scale bar, 100 μm. b, Quantification of histological appearance of GFP-positive cells at ADM Day 2 in KRASwt context (sgRosa, n = 2; sgSmarca5, n = 3 biologically independent animals). Short horizontal lines indicate the mean. c, Representative IF staining for GFP and SMARCA5 at ADM Day 2. Scale bar, 50μm. d, Representative IF co-staining for GFP and SMARCA5 in sgRosa and sgFosl1&Junb samples (as shown in Fig. 2a). Scale bar, 20μm. e, Quantification of SMARCA5 signals in GFP-positive cells in d. Significance was determined by unpaired two-tailed Student’s t-test. n = 3 biological independent mice. Data are present as mean ± SEM, with each dot representing one mouse. f, Western blot showing SMARCA5 levels after KRAS depletion in mouse PDAC cell line “3-4” (see method for details). g, Representative H&E staining and IHC staining for GFP of the pancreas from sgRosa and sgSmarca4 mice on ADM Day 7 in KRASG12D context. Scale bar, 100 μm.
Extended Data Fig. 4 SMARCA5 regulates chromatin opening of peaks near malignant genes on Day 7.
a-b, Metaplots (a) and heatmap (b) showing ATAC-seq signals gained at indicated time points after CAE treatment (samples were collected as in Fig. 2a). In b, each column represents a biologically independent mouse. c, The numbers of overlapping “shared peaks” (n = 18693) and K-lockedopen peaks9 (n = 10134). d, The percentages of “late peaks” classified by different genomic regions. e, Bar plots indicating the percentages of indicated peaks in the current study in the A1, A2, N1, N2, R and S groups identified in the previous study8. f, Diagram showing mouse model and procedure for PanIN sample collection (left). H&E staining showing a representative PanIN sample obtained from a Pdx1-Cre; LSL-KrasG12D; Klf5-CreERT2-mCherry; Rosa26LSL-YFP mouse 6 weeks after treated with caerulein. Scale bar, 100 μm. g, FPKM of acinar, PanIN, fibroblast and immune cells marker genes in mCherrypos cells sorted from PanIN mouse models. n = 1 mice with 3 technical replicates. Short horizontal lines indicate the mean. h, Heatmap demonstrating PanIN-specific and tumor-specific peaks. ATAC-seq data are obtained from the previous study9. “PanIN-specific and tumor-specific peaks” are defined in Method. i, Venn diagram illustrates the intersection of genes near “early”, “shared” and “late” peaks. Gene near a peak refers to the gene whose transcription start sites (TSS) is closest to that peak. j, GO analysis of genes annotated in i. GO analysis was performed by Enrichr using the term of “Biological process”. Significance was determined by hypergeometric test. k, ATAC-seq tracks showing representative “early”, “shared” and “late” peaks near Klf5, Mapk1 and Fn1 gene loci. 0 h, 8 h Day 2 and Day 7 tracks are from mice with KRASG12D. “Early peaks” (green), “shared peaks” (blue), and “late peaks” (red).
Extended Data Fig. 5 SMARCA5 regulates chromatin opening of peaks near malignant genes on Day 7.
a-b, Metaplot analyses of ATAC-seq signals at all open chromatin regions in sgRosa and sgSmarca5 samples at indicated time in either KRASG12D (a) or KRASwt (b). “All open peaks” refer to all peaks called by the Genrich software in the sgRosa samples at the corresponding time points. c, CTCF signals on “CTCF binding sites” in Rosa and Smarca5 knockout samples from ADM assay on Day 2. “CTCF binding sites” were determined by MACS2 using CTCF CUT&RUN data from Day 2 samples, with a threshold set at p < 0.01. d, Venn diagram showing the intersection of BRG1 CUT&RUN peaks and SMARCA5-bound®ulated peaks on Day 7. “BRG1” CUT&RUN peaks were determined by MACS2 using BRG1 CUT&RUN data from Day 7 samples, with a threshold set at p < 0.01. e, ATAC-seq, CUT&RUN (SMARCA5 and BRG1) and CUT&Tag (H3K27ac) tracks showing peaks regulated by SMARCA5 on Day 7 near Col6a1 gene locus. “Shared peaks” are labeled in blue and “late peaks” are labeled in red. f, Heatmap showing the ATAC-seq signals of the peaks activated by SMARCA5 on ADM Day 7 and in tumor samples.
Extended Data Fig. 6 SMARCA5 activates malignant genes and MAPK genes.
a, GO analysis of SMARCA5-activated genes in tumor assay (n = 342). GO analysis was performed with Metascape66. b, GO analysis of SMARCA5-activated genes in tumor (n = 342). GO analysis was performed by Enrichr using the term “MSigDB Hallmark”. Significance was determined by Enrichr using hypergeometric test. c, GO analysis of SMARCA5-repressed genes in tumor (n = 294). GO analysis was performed by Metascape. For a and c, significance was determined by Metascape using hypergeometric test and Benjamini-Hochberg p-value correction algorithm. d, GSEA comparing the expression of Basal-like genes and Classical genes (Moffit et al.27), Squamous genes and Progenitor genes (Bailey et al.24), Basal-A & Basal-B genes and Classical-A & Classical-B genes (Chan-Seng Yue et al.28) in Smarca5-knockout tumor assay. The acquisition for these gene lists is described in the Methods section. e, Representative IF co-staining for GFP and ITGA3 in Rosa and Smarca5 knockout tumor samples. Quantifications are shown on the right indicating ITGA3 signals in GFP-positive cells in Rosa and Smarca5 knockout tumor samples. Significance was determined by unpaired two-tailed Student’s t-test. Scale bars, 50μm. n = 3 biological independently mice. Data are present as mean ± SEM, with each dot representing one mouse.
Extended Data Fig. 7 SMARCA5 activates malignant genes and MAPK genes.
a, GSEA comparing the expression of genes that are activated or repressed by SMARCA5 in tumor assay and ADM Day 7 assays. Gene lists were obtained from tumor assay. b, RNA expression of Basal-like and Classical genes in Rosa and Smarca5 knockout ADM Day 7 samples. Basal and Classical gene lists are same as Extended Data Fig. 6e. c, Representative IF co-staining for GFP and ITGA3, LAMC2, TFF2 in ADM Day 7 samples. Scale bars, 50μm. d, Quantifications showing ITGA3 or LAMC2 signals in GFP-positive cells in ADM Day 7 samples. Significance was determined by unpaired two-tailed Student’s t-test. n = 3 biological independently mice. Data are present as mean ± SEM, with each dot representing one mouse. e, Heatmap showing RNA expression of genes whose FPKM > 1 in sgRosa samples and that belong to MAPK signaling pathway in Rosa and Smarca5 knockout tumor samples. MAPK gene list is obtained from the Kyoto Encyclopedia of Genes and Genomes (KEGG). In b and e, each column represents a biologically independent mouse. f, ATAC-seq signals of “shared peaks” and “late peaks” in tumor samples (data from a previous study9). g, ATAC-seq signal of MAPK genes nearby peaks in tumor samples. MAPK gene list is the same as Extended Data Fig. 7e. “Nearby peaks” are peaks within the range from the transcription start sites (TSSs) of the gene upstream of each MAPK gene to the TSSs downstream of it. h and i, Tracks showing ATAC-seq peaks regulated by SMARCA5 in tumor near Kras, Jun and Fosl1 loci.
Extended Data Fig. 8 RUNX1 associates with SMARCA5.
a, Co-IP experiments using an antibody against endogenous SMARCA5 in “806” cells. Antibodies used for western blotting are shown on the left. b, IP of HA-tagged RUNX1, ATF1 and FOSL1 from 293 T nuclear extract, followed by western blotting. 293 T cells were transfected with the indicated HA-tagged transcription factors. Endogenous SMARCA5 was blotted by western blotting. Antibodies are indicated on the left. Quantification of SMARCA5 signals in IP samples is shown below. c, IHC for RUNX1 at indicated time points (samples were collected as in Fig. 2a). Scale bar, 20μm. d, Dot plots showing quantification of RUNX1 signals in c. n = 22, 32, 33, 30, 30, 43, 24, 45 cells from left to right. e, Heatmap demonstrating RNA expression of transcription factors at the indicated samples. Each column represents a biologically independent mouse. Each condition used replicate mice (bottom) and each mouse was represented by three technical replicates. f, Representative IF co-staining for amylase, KRT19 and JUNB or FOSL1 at indicated time points (samples were collected as in Fig. 2a). Scale bar, 20 μm. g, RNA expression of Runx1, Runx2 and Runx3 in mouse PDAC samples. n = 2 mice with 3 technical replicates each. One dot represents the average of the three technical replicates. Short horizontal lines indicate the mean. h, Box plot showing protein levels of RUNX1 in human normal pancreas samples and PDAC samples. Protein expression data were obtained from UALCAN65. i, Bar plot showing RNA expression levels of Runx1 in human normal pancreas samples and PDAC samples. RNA expression data were obtained from GEPIA264. j, Representative IF co-staining for ITGA3, TFF2 and RUNX1 in the pancreatic tissue dissected from PDAC patients. Scale bar, 20 μm. k, Quantification of ITGA3 and RUNX1 signals in TFF2-positive and TFF2-negative cells in samples from PDAC patients. n = 1033, 364, 1033, 364 cells from left to right.
Extended Data Fig. 9 RUNX1 is a malignancy-associated TF and dictates SMARCA5 specificity.
a, Western blotting indicating RUNX1 depletion efficiency after sgRNA treatment. b, Quantification of histological appearance of GFP-positive cells infected with the indicated AAV-Cre-sgRNA viruses (n = 3 biologically independent animals for sgRosa and sgRunx1 on Day 2; n = 10 for sgRosa and n = 12 for sgRunx1 on Day 7), represented as mean ± SD. Quantification used whole pancreata on Day 7 and representative regions on Day 2.c, Quantification of histological appearance of GFP-positive cells in the presence of the indicated AAV-Cre-sgRNA viruses (n = 8 biologically independent animals for sgRosa; n = 7 for sgRunx1), represented as mean ± SD. d, Representative IF co-staining for GFP and RUNX1 on Day 7 and in tumor of Rosa and Runx1 knockout samples in the context of KRASG12D. Scale bar, 20μm. White dotted circles indicate ADM or tumor regions. e, Quantification of histological appearance of GFP-positive cells in the presence of the indicated AAV-Cre-sgRNA viruses (n = 3 biologically independent animals), represented as mean ± SD. f, Venn diagram showing the numbers of ATAC-seq peaks activated by SMARCA5 and RUNX1 on ADM Day 7 in the context of KRASG12D. g, Heatmap showing the ATAC-seq signals of the peaks not-regulated by SMARCA5 (n = 107302) in Rosa, Smarca5 and Runx1 knockout samples on ADM Day 7. h, ATAC-seq tracks and CUT&RUN (RUNX1) tracks showing peaks regulated by SMARCA5 on in Smarca5 and Runx1 knockout ADM Day 7 samples near Col6a1 and Fosl1 gene loci. In b, c and e, significance was determined by unpaired two-tailed Student’s t-test.
Extended Data Fig. 10 RUNX1 is a malignancy-associated TF and dictates SMARCA5 specificity.
a, The numbers of genes activated or repressed by SMARCA5 or RUNX1 on ADM Day 7. Significance was determined by hypergeometric distribution. b, GSEA comparing the expression of SMARCA5-repressed genes in ADM Day 7 samples with Runx1 knockout. c, GSEA comparing the expression of Basal-like and Classical genes in ADM Day 7 samples with Runx1 knockout. Basal-like/ Classical gene list are same as Extended Data Fig. 6e. d, Heatmap showing RNA expression in ADM Day 7 samples. Gene list are identical to Fig. 4c. e, Representative IF co-staining in tumor and ADM Day 7 samples. Scale bar, 50μm. Significance was determined by unpaired two-tailed Student’s t-test. n = 3 biological independently mice. Data are present as mean ± SEM, with each dot representing one mouse. f, RNA expression of genes in the MAPK signaling pathway in ADM Day 7 samples. MAPK gene list identical to Extended Data Fig. 7e. g, Representative H&E staining for samples obtained before Dox treatment in Fig. 7a. Scale bar, 50μm. h, Representative IF co-staining for samples obtained 3 weeks after Dox treatment in Fig. 7a. Scale bar, 50μm. i, GSEA comparing expression of Basal-like genes in SMARCA5 and RUNX1 knockdown samples in PDCC system. The gene list was selected from ref. 25. j, GO analysis of GSK2801-repressed genes (n = 226). Significance was determined by Metascape using hypergeometric test and Benjamini-Hochberg p-value correction algorithm. k, GSEA comparing the expression of SMARCA5-activated genes in GSK2801 treated PDCCs. Gene lists were same as Extended Data Fig. 6a. Those genes symbols are transferred from mouse to human for GSEA analysis.
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Tables 1–14.
Source data
Source Data Fig. 5 and Source Data Extended Data Figs. 1, 3, 8, 9 (download PDF )
This file contains uncropped scans of blots for Fig. 5d and Extended Data Figs. 1c, 3f, 8a,b and 9a.
Source Data Figs. 1, 2, 4, 5, 7 and Source Data Extended Data Figs. 1–4 and 6–10 (download XLSX )
This file contains statistical source data for Figs. 1e,g,i, 2c,e,g, 4e, 5a,c,d,g,k and 7c,g,h,k,i and Extended Data Figs. 1d,n,o, 2b,h,k, 3b,e, 4e, 6e, 7d, 8d,k, 9b,c,e and 10e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, J., Lu, X., Guo, M. et al. Spatiotemporal control of SMARCA5 by a MAPK–RUNX1 axis distinguishes mutant KRAS-driven pancreatic malignancy from tissue regeneration. Nat Cancer 7, 43–59 (2026). https://doi.org/10.1038/s43018-025-01065-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01065-3