Abstract
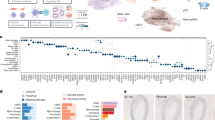
Growing evidence suggests that the induction of cellular senescence in vascular cells is causally linked to the etiology of cardiovascular diseases. To investigate systematically the heterogeneity of senescent vascular cells in atherosclerosis, we used a high-fat diet and PCSK9 overexpression to induce atherosclerosis in a senescence reporter mouse model (p16-tdTomato+/−) and performed single-cell RNA sequencing on whole aortas. Using the SenMayo and CellAge gene sets, we identified four clusters of vascular smooth muscle cells (VSMCs), fibroblasts and T cells enriched in features of senescence, which were reduced upon treatment with the senolytic agent ABT-737. We then derived a global senescence signature of atherosclerosis including Spp1, Ctsb and Tnfrsf11b mRNAs. We validated the enrichment of these mRNAs in senescence by using spatial transcriptomics in a second mouse model of atherosclerosis and senolysis (Ldlr−/−; p16-3MR), as well as by analyzing in vitro models of human VSMC senescence. Our results uncover a vascular-specific transcriptomic signature of senescence that may be exploited for tracking and treating age-related vascular diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-sequencing data are deposited in the Gene Expression Omnibus under accession nos. GSE239591 and GSE239602. Commercially available reagents and their catalog numbers are listed in Supplementary Table 4. Source data are provided with this paper. All other data will be provided upon request.
Code availability
No unique code was used in this study. All code used in this article is open source and listed, and additional information can be provided upon request.
References
Matthews, C. et al. Vascular smooth muscle cells undergo telomere-based senescence in human atherosclerosis: effects of telomerase and oxidative stress. Circ. Res. 99, 156–164 (2006).
Minamino, T. et al. Endothelial cell senescence in human atherosclerosis: role of telomere in endothelial dysfunction. Circulation 105, 1541–1544 (2002).
Kunieda, T. et al. Angiotensin II induces premature senescence of vascular smooth muscle cells and accelerates the development of atherosclerosis via a p21-dependent pathway. Circulation 114, 953–960 (2006).
Hernandez-Segura, A., Nehme, J. & Demaria, M. Hallmarks of cellular senescence. Trends Cell Biol. 28, 436–453 (2018).
Demaria, M. et al. An essential role for senescent cells in optimal wound healing through secretion of PDGF-AA. Dev. Cell 31, 722–733 (2014).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Roos, C. M. et al. Chronic senolytic treatment alleviates established vasomotor dysfunction in aged or atherosclerotic mice. Aging Cell 15, 973–977 (2016).
Liu, J.-Y. et al. Cells exhibiting strong p16INK4a promoter activation in vivo display features of senescence. Proc. Natl Acad. Sci. USA 116, 2603–2611 (2019).
Roche-Molina, M. et al. Induction of sustained hypercholesterolemia by single adeno-associated virus-mediated gene transfer of mutant hPCSK9. Arterioscler. Thromb. Vasc. Biol. 35, 50–59 (2015).
Bjørklund, M. M. et al. Induction of atherosclerosis in mice and hamsters without germline genetic engineering. Circ. Res. 114, 1684–1689 (2014).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Avelar, R. A. et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. 21, 91 (2020).
Yosef, R. et al. Directed elimination of senescent cells by inhibition of BCL-W and BCL-XL. Nat. Commun. 7, 11190 (2016).
Thim, T. et al. Unreliable assessment of necrotic core by virtual histology intravascular ultrasound in porcine coronary artery disease. Circ. Cardiovasc. Imaging 3, 384–391 (2010).
Jana, S., Hu, M., Shen, M. & Kassiri, Z. Extracellular matrix, regional heterogeneity of the aorta, and aortic aneurysm. Exp. Mol. Med. 51, 1–15 (2019).
Garrido, A. M. et al. Efficacy and limitations of senolysis in atherosclerosis. Cardiovasc Res. https://doi.org/10.1093/cvr/cvab208 (2022).
Chi, C. et al. Vascular smooth muscle cell senescence and age-related diseases: state of the art. Biochim. Biophys. Acta Mol. Basis Dis. 1865, 1810–1821 (2019).
Mavrogonatou, E., Pratsinis, H., Papadopoulou, A., Karamanos, N. K. & Kletsas, D. Extracellular matrix alterations in senescent cells and their significance in tissue homeostasis. Matrix Biol. 75–76, 27–42 (2019).
Smit, M. A. & Peeper, D. S. Epithelial–mesenchymal transition and senescence: two cancer-related processes are crossing paths. Aging (Albany NY) 2, 735–741 (2010).
Herman, A. B. et al. DPP4 inhibition impairs senohemostasis to improve plaque stability in atherosclerotic mice. J. Clin. Invest. https://doi.org/10.1172/JCI165933 (2023).
Wiley, C. D. et al. SILAC analysis reveals increased secretion of hemostasis-related factors by senescent cells. Cell Rep. 28, 3329–3337 (2019).
Herman, A. B. et al. Reduction of lamin B receptor levels by miR-340-5p disrupts chromatin, promotes cell senescence and enhances senolysis. Nucleic Acids Res. 49, 7389–7405 (2021).
Wechter, N. et al. Single-cell transcriptomic analysis uncovers diverse and dynamic senescent cell populations. Aging (Albany NY) 15, 2824–2851 (2023).
Tacutu, R. et al. Human Ageing Genomic Resources: new and updated databases. Nucleic Acids Res. 46, D1083–D1090 (2018).
Chen, R., McVey, D. G., Shen, D., Huang, X. & Ye, S. Phenotypic switching of vascular smooth muscle cells in atherosclerosis. J. Am. Heart Assoc. 12, e031121 (2023).
Yap, C., Mieremet, A., de Vries, C. J. M., Micha, D. & de Waard, V. Six shades of vascular smooth muscle cells illuminated by KLF4 (Krüppel-like factor 4). Arterioscler. Thromb. Vasc. Biol. 41, 2693–2707 (2021).
Frendéus, K. H., Wallin, H., Janciauskiene, S. & Abrahamson, M. Macrophage responses to interferon-γ are dependent on cystatin C levels. Int. J. Biochem. Cell Biol. 41, 2262–2269 (2009).
Anerillas, C. et al. The YAP–TEAD complex promotes senescent cell survival by lowering endoplasmic reticulum stress. Nat. Aging 3, 1237–1250 (2023).
Freund, A., Laberge, R.-M., Demaria, M. & Campisi, J. Lamin B1 loss is a senescence-associated biomarker. Mol. Biol. Cell 23, 2066–2075 (2012).
Park, H., Kim, C.-H., Jeong, J.-H., Park, M. & Kim, K. S. GDF15 contributes to radiation-induced senescence through the ROS-mediated p16 pathway in human endothelial cells. Oncotarget 7, 9634–9644 (2016).
Basisty, N. et al. A proteomic atlas of senescence-associated secretomes for aging biomarker development. PLoS Biol. 18, e3000599 (2020).
Coppé, J.-P., Desprez, P.-Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Mosquera, J. V. et al. Integrative single-cell meta-analysis reveals disease-relevant vascular cell states and markers in human atherosclerosis. Cell Rep. 42, 113380 (2023).
Chen, H., Li, Y. & Tollefsbol, T. O. Cell senescence culturing methods. Methods Mol. Biol. 1048, 1–10 (2013).
Krtolica, A., Parrinello, S., Lockett, S., Desprez, P. Y. & Campisi, J. Senescent fibroblasts promote epithelial cell growth and tumorigenesis: a link between cancer and aging. Proc. Natl Acad. Sci. USA 98, 12072–12077 (2001).
Childs, B. G. et al. Senescent intimal foam cells are deleterious at all stages of atherosclerosis. Science 354, 472–477 (2016).
Lear, T. B. & Finkel, T. Senolytic vaccination: a new mandate for cardiovascular health? J. Cardiovasc. Aging 2, 17 (2022).
Childs, B. G., Durik, M., Baker, D. J. & van Deursen, J. M. Cellular senescence in aging and age-related disease: from mechanisms to therapy. Nat. Med. 21, 1424–1435 (2015).
Baker, D. J. & Petersen, R. C. Cellular senescence in brain aging and neurodegenerative diseases: evidence and perspectives. J. Clin. Invest. 128, 1208–1216 (2018).
Niedernhofer, L. J. & Robbins, P. D. Senotherapeutics for healthy ageing. Nat. Rev. Drug Discov. 17, 377 (2018).
Kirkland, J. L., Tchkonia, T., Zhu, Y., Niedernhofer, L. J. & Robbins, P. D. The clinical potential of senolytic drugs. J. Am. Geriatr. Soc. 65, 2297–2301 (2017).
Anerillas, C. et al. A BDNF–TrkB autocrine loop enhances senescent cell viability. Nat. Commun. 13, 6228 (2022).
Suda, M. et al. Senolytic vaccination improves normal and pathological age-related phenotypes and increases lifespan in progeroid mice. Nat. Aging 1, 1117–1126 (2021).
Camell, C. D. et al. Senolytics reduce coronavirus-related mortality in old mice. Science 373, eabe4832 (2021).
Casella, G. et al. Transcriptome signature of cellular senescence. Nucleic Acids Res. 47, 7294–7305 (2019).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660 (2017).
Clayton, Z. S. et al. Cellular senescence contributes to large elastic artery stiffening and endothelial dysfunction with aging: amelioration with senolytic treatment. Hypertension 80, 2072–2087 (2023).
Childs, B. G. et al. Senescent cells suppress innate smooth muscle cell repair functions in atherosclerosis. Nat. Aging 1, 698–714 (2021).
Ahmet, I. et al. Chronic administration of small nonerythropoietic peptide sequence of erythropoietin effectively ameliorates the progression of postmyocardial infarction-dilated cardiomyopathy. J. Pharmacol. Exp. Ther. 345, 446–456 (2013).
Gabunia, K. et al. IL-19 halts progression of atherosclerotic plaque, polarizes, and increases cholesterol uptake and efflux in macrophages. Am. J. Pathol. 186, 1361–1374 (2016).
Ellison, S. et al. Attenuation of experimental atherosclerosis by interleukin-19. Arterioscler. Thromb. Vasc. Biol. 33, 2316–2324 (2013).
Ohayon, J. et al. Necrotic core thickness and positive arterial remodeling index: emergent biomechanical factors for evaluating the risk of plaque rupture. Am. J. Physiol. Heart Circ. Physiol. 295, H717–727 (2008).
Kadoglou, N. P. et al. The anti-inflammatory effects of exercise training promote atherosclerotic plaque stabilization in apolipoprotein E knockout mice with diabetic atherosclerosis. Eur. J. Histochem. 57, e3 (2013).
Stein, S. et al. ApoE−/− PGC-1α−/− mice display reduced IL-18 levels and do not develop enhanced atherosclerosis. PLoS ONE 5, e13539 (2010).
Chen, Y., Yu, Q. & Xu, C.-B. A convenient method for quantifying collagen fibers in atherosclerotic lesions by ImageJ software. Int. J. Clin. Exp. Med. 10, 14904–14910 (2017).
Baidoo, N., Sanger, G. J. & Belai, A. Histochemical and biochemical analysis of collagen content in formalin-fixed, paraffin embedded colonic samples. MethodsX 11, 102416 (2023).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb.) 2, 100141 (2021).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Fabregat, A. et al. Reactome pathway analysis: a high-performance in-memory approach. BMC Bioinformatics 18, 142 (2017).
Griswald, M. et al. GeoMxWorkflows: GeoMx Digital Spatial Profiler (DSP) data analysis workflows. R package version 1.6.0 https://doi.org/10.18129/B9.bioc.GeoMxWorkflows (2023).
Batish, M. & Tyagi, S. Fluorescence in situ imaging of dendritic RNAs at single-molecule resolution. Curr. Protoc. Neurosci. 89, e79 (2019).
Kumar, R. et al. Imaging architecture of granulomas induced by Mycobacterium tuberculosis infection with single-molecule fluorescence in situ hybridization. J. Immunol. 213, 526–537 (2024).
Koppula, A., Abdelgawad, A., Guarnerio, J., Batish, M. & Parashar, V. CircFISH: a novel method for the simultaneous imaging of linear and circular RNAs. Cancers (Basel) 14, 428 (2022).
Acknowledgements
This work was supported in part by the NIA IRP, NIH and National Science Foundation grant number 2244127 (to M.B.), and the Paul F. Glenn Center for Biology of Aging Research at the Mayo Clinic (B.G.C. and D.J.B.). We would like to thank the sequencing and data processing support from the CCR Genomics Core at the National Cancer Institute/NIH.
Author information
Authors and Affiliations
Contributions
A.B.H., D.T. and K.M.-M. conceptualized the study; A.B.H., D.T., K.M.-M., B.G.C., N.K., O.V.F., D.J.B., S.D., M.B. and M.G. designed experiments; A.B.H., K.M.-M., D.T., A.E.C., B.R., C.R.D.S., C.A., J.M.G., C.H.-S., A.N.O., M.M., R.M., J.L.M., Y.P., J.F., M.O.H., N.K., M.L.V. and M.M.F.W. performed and analyzed experiments and analyzed data; A.B.H. and M.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
B.G.C. and D.J.B. have potential financial interests related to this research. They are co-inventors on patents held by the Mayo Clinic and patent applications licensed to or filed by Unity Biotechnology. They are also Unity Biotechnology shareholders. Research in the laboratory of D.J.B. has been reviewed by the Mayo Clinic Conflict of Interest Review Board and is being conducted in compliance with the Mayo Clinic conflict of interest policies. The other authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Minna Kaikkonen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Atherosclerotic plaque pathology analysis and quality control analysis.
a, Quantification of plaque size presented by sex and condition (ND, HFD, and HFD + ABT-737). For female mice, ND (n = 4), HFD (n = 3), and HFD + ABT-737 (n = 2) and HFD vs. ND, p = 0.0006 and HFD + ABT-737 vs. HFD, p = 0.576, whereas for male mice, ND (n = 4), HFD (n = 2), and HFD + ABT-737 (n = 3) and HFD vs. ND, p = 0.0004 and HFD + ABT-737 vs. HFD, p = 0.8876. b, Quantification of necrotic core area presented by sex and condition (ND, HFD, and HFD + ABT-737). For female mice, HFD (n = 3) and HFD + ABT-737 (n = 2) HFD + ABT-737 vs. HFD, p = 0.1929, whereas for male mice, HFD (n = 2) and HFD + ABT-737 (n = 2) and HFD + ABT-737 vs. HFD, p = 0.4985. c, Quantification by plaque fibrous cap thickness presented by sex and condition (ND, HFD, and HFD + ABT-737). For female mice, HFD (n = 3) and HFD + ABT-737 (n = 2) and HFD + ABT-737 vs. HFD, p = 0.2033, whereas for male mice, HFD (n = 2) and HFD + ABT-737 (n = 2) and HFD + ABT-737 vs. HFD, p = 0.4552. d, Representative H&E staining of aortic roots from ND, HFD, and HFD + ABT-737. The scale bar is 200 µm. e, Quantification of collagen fibers (Aniline blue) area / area of aortic root presented by sex and condition (ND, HFD, and HFD + ABT-737). For female mice, HFD (n = 2) and HFD + ABT-737 (n = 2) HFD + ABT-737 vs. HFD, p = 0.2467, whereas for male mice, HFD (n = 2) and HFD + ABT-737 (n = 2) and HFD + ABT-737 vs. HFD, p = 0.7036. f, Representative senescence-associated beta galactosidase (SA-β-gal) activity staining of aortic roots from ND, HFD, and HFD + ABT-737. The scale bar is 200 µm. g, Quantification of SA-β-gal (ND, HFD, and HFD + ABT-737). ND (n = 5), HFD (n = 5) and HFD + ABT-737 (n = 5) and for significance, HFD vs. ND, p ≤ 0.0001 and HFD + ABT-737 vs. HFD, p = 0.069. h, Pulse wave velocity (PWV) measurements presented by sex and condition (ND, HFD, and HFD + ABT-737). For female mice, ND (n = 3), HFD (n = 2), and HFD + ABT-737 (n = 4) and HFD vs. ND, p = 0.748 and HFD + ABT-737 vs. HFD, p = 0.9597, whereas for male mice, ND (n = 3), HFD (n = 2), and HFD + ABT-737 (n = 2) and HFD vs. ND, p = 0.0294 and HFD + ABT-737 vs. HFD, p = 0.1001. i, Quality control table of the scRNA-seq samples including estimated number of cells, fraction reads in cells, mean reads per cell, median UMI counts per cell, median genes per cell, and total genes detected. j, Heatmap of normalized expression levels of classical senescence associated mRNAs either enriched in HFD and reduced by ABT-737 treatment, or conversely, reduced by HFD and increased by ABT-737 treatment in mice (Lmnb1 and Lbr mRNAs). Significance was established using Two-Way ANOVA (for data in graphs a-c, e, and h) and One-Way ANOVA (for data in graph g) with multiple comparisons. *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001.
Extended Data Fig. 2 scRNA-seq analysis of senescence gene expression.
a, UMAPs of p16-tdTomato mRNA expression in ND, HFD, and HFD + ABT-737 treated mice. b, UMAP plot of Cdkn2a mRNA expression across all clusters and conditions (left). Violin plot of Cdkn2a mRNA expression across all clusters (right). c, UMAP plot of Cdkn1a mRNA expression across all clusters and conditions (left). Violin plot of Cdkn1a mRNA expression across all clusters (right). d, GSEA SenMayo plots comparing HFD vs ND and HFD + ABT-737 vs HFD for Clusters 15, 20, 21, 3, 16, and 27. e, GSEA CellAge plots comparing HFD vs ND and HFD + ABT-737 vs HFD for Clusters 6, 17, 7, and 11.
Extended Data Fig. 3 VSMC phenotype analysis.
a, Dot plot representation of the gene expression profile for VSMC phenotype mRNA markers for cluster 0 subclustered VSMCs. b, Dot plot representation of the gene expression profile for VSMC phenotype mRNA markers for cluster 12 subclustered VSMCs.
Extended Data Fig. 4 Analysis of UMI counts, mitochondrial and ribosomal gene expression in VSMC, fibroblast, and T cell clusters with senescence features.
a-c, UMAPs of scRNA-seq UMI counts (a), mitochondrial gene expression (b) and ribosomal gene expression (c) across cluster 0 subclustered VSMCs in ND, HFD, and HFD + ABT-737 treated mice. d-f, UMAPs of scRNA-seq UMI counts (d), mitochondrial gene expression (e) and ribosomal gene expression (f) across cluster 12 subclustered VSMCs in ND, HFD, and HFD + ABT-737 treated mice. g-i, UMAPs of scRNA-seq UMI counts (g), mitochondrial gene expression (h) and ribosomal gene expression (i) across cluster 2 subclustered fibroblasts in ND, HFD, and HFD + ABT-737 treated mice. j-l, UMAPs of scRNA-seq UMI counts (j), mitochondrial gene expression (k) and ribosomal gene expression (l) across Cluster 9 subclustered T cells in ND, HFD, and HFD + ABT-737 treated mice.
Extended Data Fig. 5 Vascular senescence score on individual cell clusters.
a, UMAPs of vascular-senescence scoring across cluster 0 VSMCs in ND, HFD, and HFD + ABT-737. b, UMAPs of vascular-senescence scoring across cluster 12 VSMCs in ND, HFD, and HFD + ABT-737. c, UMAPs of vascular-senescence scoring across cluster 2 fibroblasts in ND, HFD, and HFD + ABT-737. d, UMAPs of vascular-senescence scoring across cluster 9 T cells in ND, HFD, and HFD + ABT-737.
Extended Data Fig. 6 Gene score of the most prominent transcripts expressed in senescent vascular cells.
UMAPs visualization of the expression of the 17 vascular senescent transcripts individually across all clusters. The arrows indicate the senescent cell clusters for VSMCs (V), fibroblasts (F), and T cells (T) as well as non-senescent macrophages (M) in which the transcripts were expressed.
Extended Data Fig. 7 Doxorubicin-induced cellular senescence in wild type mice.
a, RT-qPCR analysis of the levels of Cdkn2a and Lmnb1 mRNAs in aortic roots of mice treated with vehicle, doxorubicin, and doxorubicin followed by ABT-737. b, RT-qPCR analysis of the levels of Ltbp2, Sparc, Mgp, Lum, and Cd9 mRNAs in aortic root tissue as described (a). c, RT-qPCR analysis of the levels of Apoe, Lgals3, Cst3, Tnfrsf11b, and Prg4 mRNAs in aortic root tissue as described (a). d, RT-qPCR analysis of the levels of Mmp3, Ctss, Lcp1, and Comp mRNAs in aortic root tissue as described (a). In a-d, data represent the means ± SD for DMSO, n = 4; Doxo, n = 4; Doxo + ABT-737, n = 3 (females), and DMSO, n = 4; Doxo, n = 4; Doxo + ABT-737, n = 4 (males). n represents biological replicates. In panel (a), Cdkn2a, Doxo vs. DMSO, p = 0.0162 (females) and p = 0.002 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0226 (females) and p = 0.0034 (males); Lmnb1, Doxo vs. DMSO, p = 0.5863 (females) and p = 0.1807 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.1013 (females) and p = 0. 3449 (males). In panel (b), Ltbp2, Doxo vs. DMSO, p = 0.0004 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0007 (females) and p ≤ 0.0001 (males); Sparc, Doxo vs. DMSO, p ≤ 0.0001 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p ≤ 0.0001 (females) and p = 0.0002 (males); Mgp, Doxo vs. DMSO, p = 0.9928 (females) and p = 0.9928 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.7181 (females) and p > 0.9999 (males); Lum, Doxo vs. DMSO, p = 0.0018 (females) and p = 0. 0002 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0018 (females) and p = 0.001 (in males); Cd9, Doxo vs. DMSO, p = 0.0014 (females) and p = 0. 0011 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0031 (females) and p = 0.0019 (males). In panel (c), Apoe, Doxo vs. DMSO, p = 0.0194 (females) and p = 0. 3607 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.5746 (females) and p = 0.4692 (males); Lgals3, Doxo vs. DMSO, p ≤ 0.0001 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0002 (females) and p ≤ 0.0001 (males); Cst3, Doxo vs. DMSO, p = 0.0008 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0021 (females) and p ≤ 0.0001 (males); Tnfrsf11b, Doxo vs. DMSO, p = 0.3051 (females) and p = 0.9339 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.1425 (females) and p = 0.9833 (males); Prg4, Doxo vs. DMSO, p ≤ 0.0001 (females) and p = 0.0177 (males), whereas Doxo + ABT-737 vs. Doxo, p ≤ 0.0001 (females) and p = 0.0283 (males). In panel (d), Mmp3, Doxo vs. DMSO, p ≤ 0.0001 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p ≤ 0.0001 (females) and p ≤ 0.0001 (males); Ctss, Doxo vs. DMSO, p = 0.0007 (females) and p = 0.0025 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0017 (females) and p = 0.005 (males); Lcp1, Doxo vs. DMSO, p ≤ 0.0001 (females) and p ≤ 0.0001 (males), whereas Doxo + ABT-737 vs. Doxo, p ≤ 0.0001 (females) and p ≤ 0.0001 (males); Comp, Doxo vs. DMSO, p = 0.0005 (females) and p = 0.0029 (males), whereas Doxo + ABT-737 vs. Doxo, p = 0.0027 (females) and p = 0.004 (males). Significance was established using One-Way Anova for each sex, *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001.
Extended Data Fig. 8 Human VSMC senescence validation.
a, Phase contrast micrographs of SA-βgal activity (blue) staining in proliferating (P), replicative senescence (RS), doxo-treated, IR-treated, oxLDL-treated, and CoCl2-treated human VSMCs for 7–10 days. b, RT-qPCR analysis of the levels of CDKN2A and IL8 mRNAs in human VSMCs treated as described in (a). c, RT-qPCR analysis of the levels of LTBP2, SPARC, and MGP mRNAs in human VSMCs treated as described in (a). d, RT-qPCR analysis of the levels of LUM, CD9, APOE, LGALS3, CST3 mRNAs in human VSMCs treated as described in (a). e, RT-qPCR analysis of the levels of TNFRSF11b, PRG4, MMP3, CTSS, and LCP1 mRNAs in human VSMCs treated as described in (a). f, Table of human scRNA-seq metadata analysis from atherosclerotic tissue. Each column represents a VSMC phenotype identified in the metadata analysis, and each column includes the vascular senescent scoring genes that were expressed in the indicated VSMC phenotype. In b-e, data represent the means ± SD for P, n = 3; IR, n = 3; oxLDL, n = 3; CoCl2, n = 3; Doxo, n = 3; RS, n = 3; n represents biological replicates. In panel (b), CDKN2A, IR (p = 0.1334), oxLDL (p = 0.0191), CoCl2 (p = 0.001), Doxo (p = 0.4446), RS (p ≤ 0.0001) vs. P; IL8, IR (p = 0.0007), oxLDL (p = 0.0014), CoCl2 (p = 0.0003), Doxo (p = 0.0008), RS (p = 0.0027) vs. P. In panel (c), LTBP2, IR (p = 0.002), oxLDL (p = 0.0021), CoCl2 (p ≤ 0.0001), Doxo (p ≤ 0.0001), RS (p = 0.4192) vs. P; SPARC, IR (p = 0.2171), oxLDL (p ≤ 0.0001), CoCl2 (p = 0.0002), Doxo (p = 0.0106), RS (p = 0.0019) vs. P; MGP, IR (p = 0.0201), oxLDL (p = 0.001), CoCl2 (p = 0.0134), Doxo (p = 0.0015), RS (p = 0.0043) vs. P. In panel (d), LUM, IR (p ≤ 0.0001), oxLDL (p = 0.0002), CoCl2 (p ≤ 0.0001), Doxo (p = 0.0003), RS (p ≤ 0.0001) vs. P; CD9, IR (p = 0.0002), oxLDL (p = 0.0296), CoCl2 (p = 0.0151), Doxo (p = 0.0291), RS (p = 0.0002) vs. P; APOE, IR (p = 0.0171), oxLDL (p = 0.0004), CoCl2 (p = 0.016), Doxo (p = 0.0486), RS (p = 0.1703) vs. P; LGALS3, IR (p = 0.2702), oxLDL (p = 0.0014), CoCl2 (p = 0.007), Doxo (p = 0.0284), RS (p = 0.0014) vs. P; CST3, IR (p = 0.9809), oxLDL (p ≤ 0.0001), CoCl2 (p = 0.1202), Doxo (p = 0.087), RS (p = 0.0209) vs. P. In panel (e), TNFRSF11b, IR (p = 0.0019), oxLDL (p ≤ 0.0001), CoCl2 (p = 0.001), Doxo (p = 0.0003), RS (p ≤ 0.0001) vs. P; PRG4, IR (p = 0.0595), oxLDL (p = 0.0005), CoCl2 (p = 0.0008), Doxo (p = 0.0011), RS (p = 0.0021) vs. P; MMP3, IR (p = 0.0054), oxLDL (p = 0.0007), CoCl2 (p = 0.0003), Doxo (p = 0.0024), RS (p = 0.0098) vs. P; CTSS, IR (p = 0.0085), oxLDL (p ≤ 0.0001), CoCl2 (p = 0.0922), Doxo (p = 0.0019), RS (p = 0.0003) vs. P; LCP1, IR (p = 0.0029), oxLDL (p = 0.0033), CoCl2 (p = 0.0003), Doxo (p = 0. 0008), RS (p = 0.0006) vs. P. Significance was established using Shapiro-Wilk test first, followed by Welch’s t-test. *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001.
Extended Data Fig. 9 Human WI-38 fibroblast senescence validation.
a, Phase contrast micrographs of SA-βgal activity (blue) staining in human WI-38 fibroblasts that were either proliferating (P), rendered senescent by replicative senescence (RS), or by treatment with Doxo, IR and additional culture for 7–10 days. b, RT-qPCR analysis of the levels of CDKN2A, CDKN1A, GDF15, LMNB1 and IL8 mRNAs in WI-38 fibroblasts treated as described in (a). c-f, RT-qPCR analysis of the levels of SPP1, SERPINE2, CTSB, TNFRSF11B, PRG4, LTPBP2, FTH1, MGP, LUM, CD9, APOE, LGALS3, CST3, SPARC, THBS1, MMP3, CTSS, and LCP1 mRNAs in human WI-38 fibroblasts treated as described in (a). In b-f, data represent the means ± SD for P, n = 3; IR, n = 3; n = 3; Doxo, n = 3; RS, n = 3; n represents biological replicates. In panel (b), CDKN2A, IR (p = 0.0099), Doxo (p ≤ 0.0001), RS (p = 0.0005) vs. P; CDKN1A, IR (p ≤ 0.0001), Doxo (p ≤ 0.0001), RS (p ≤ 0.0001) vs. P; GDF15, IR (p = 0.001), Doxo (p ≤ 0.0001), RS (p ≤ 0.0021) vs. P; LMNB1, IR (p = 0.001), Doxo (p = 0.0002), RS (p = 0.0009) vs. P; IL8, IR (p ≤ 0.0001), Doxo (p = 0.0005), RS (p = 0.0003) vs. P. In panel (c), SPP1, IR (p = 0.0085), Doxo (p = 0.0035), RS (p = 0.0071) vs. P; SERPINE2, IR (p = 0.0014), Doxo (p = 0.0025), RS (p = 0.0026) vs. P; CTSB, IR (p = 0.0003), Doxo (p = 0.0002), RS (p = 0.0007) vs. P; TNFRSF11B, IR (p = 0.0013), Doxo (p = 0.0003), RS (p ≤ 0.0001) vs. P; PRG4, IR (p = 0.0135), Doxo (p = 0.0043), RS (p = 0.0039) vs. P. In panel (d), LTPBP2, IR (p = 0.0003), Doxo (p = 0.0012), RS (p = 0.0006) vs. P; FTH1, IR (p = 0.0025), Doxo (p = 0.0038), RS (p = 0.0016) vs. P; MGP, IR (p = 0.0007), Doxo (p ≤ 0.0001), RS (p ≤ 0.0001) vs. P; LUM, IR (p = 0.0005), Doxo (p = 0.0059), RS (p = 0.0014) vs. P; CD9, IR (p = 0.0051), Doxo (p = 0.7688), RS (p = 0.0008) vs. P. In panel (e), APOE, IR (p = 0.0008), Doxo (p = 0.007), RS (p ≤ 0.0001) vs. P; LGALS3, IR (p = 0.0108), Doxo (p = 0.0229), RS (p ≤ 0.0001) vs. P; CST3, IR (p = 0.0026), Doxo (p ≤ 0.0001), RS (p ≤ 0.0001) vs. P; SPARC, IR (p = 0.0021), Doxo (p = 0.002), RS (p = 0.0004) vs. P; THBS1, IR (p = 0.0003), Doxo (p = 0.0114), RS (p ≤ 0.0001) vs. P. In panel (f), MMP3, IR (p = 0.0015), Doxo (p = 0.0007), RS (p = 0.0003) vs. P; CTSS, IR (p = 0.0002), Doxo (p = 0.0023), RS (p = 0.0019) vs. P; LCP1, IR (p = 0.0003), Doxo (p = 0.9169), RS (p ≤ 0.0001) vs. P.Significance was established using Shapiro-Wilk test first, followed by Welch’s t-test. *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001.
Extended Data Fig. 10 Spatial transcriptomic profiling of arterial segments.
a, H&E staining (top) and Masson’s trichrome staining (bottom) of brachiocephalic arterial sections from Ldlr−/− + GCV, Ldlr−/−;3MR + Vehicle, and Ldlr−/−;3MR + GCV treated mice. The scale bar is 100 µm. b, Plaque cross sectional area, c, plaque fibrous cap thickness measurements and d, Masson’s trichrome analysis of Aniline blue-positive area over total plaque area in brachiocephalic arterial sections from Ldlr−/− + GCV (n = 4), Ldlr−/−;3MR + Vehicle (n = 4), and Ldlr−/−;3MR + GCV (n = 4) treated mice. In panel (b), Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.8299) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.7933). In panel (c), Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.5444) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.3969). In panel (b), Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.7628) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.9994). e, H&E staining (top) and Masson’s trichrome staining (bottom) of the descending aortic sections from Ldlr−/− + GCV, Ldlr−/−;3MR + Vehicle, and Ldlr−/−;3MR + GCV treated mice. The scale bar is 50 µm. f-h, Plaque cross sectional area (f), plaque fibrous cap thickness measurements (cap is underlined with yellow dashed line) (g), and Masson’s trichrome analysis of Aniline blue-positive area over total plaque area in sections of the descending aorta (h) from Ldlr−/− + GCV, Ldlr−/−;3MR + Vehicle, and Ldlr−/−;3MR + GCV treated mice. In panel (f) and (g), Ldlr−/− + GCV (n = 5), Ldlr−/−;3MR + Vehicle (n = 4), and Ldlr−/−;3MR + GCV (n = 6); in panel (f), Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.9706) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.08), whereas in (g), Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.379) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.0021). In panel (h), Ldlr−/− + GCV (n = 4), Ldlr−/−;3MR + Vehicle (n = 4), and Ldlr−/−;3MR + GCV (n = 5), and significance Ldlr−/−;3MR + Vehicle vs. Ldlr−/− + GCV (p = 0.9867) and Ldlr−/−;3MR + GCV vs. Ldlr−/−;3MR + Vehicle (p = 0.0263) i, Reactome pathway analysis of genes upregulated in cap compared to normal wall across all conditions (left) and of genes upregulated in the normal wall compared to the cap (right). j, Reactome pathway analysis of genes upregulated in the core compared to the normal wall across all conditions. In b-d and f-h, data represent the means ± SD and n represents biological replicates. Significance was established using One-Way Anova with multiple comparisons (b-d and f-h) or Fisher-Irwin test (i and j). *, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001.
Supplementary information
Supplementary Information (download PDF )
Supplementary tables containing information on GSEA results (Supplementary Table 1), set of primers (Supplementary Table 2), smFISH probes (Supplementary Table 3) and reagents used in the study (Supplementary Table 4).
Source data
Source Data Fig. 1 (download XLSX )
Raw measurements for the assays indicated in the figure legend.
Source Data Fig. 5 (download XLSX )
RT–qPCR results.
Source Data Fig. 7 (download XLSX )
Raw measurements for smFISH.
Source Data Fig. 8 (download XLSX )
Raw measurements for smFISH.
Source Data Extended Data Fig. 1 (download XLSX )
Raw measurements for the assays indicated in the figure legend.
Source Data Extended Data Fig. 7 (download XLSX )
RT–qPCR results.
Source Data Extended Data Fig. 8 (download XLSX )
RT–qPCR results.
Source Data Extended Data Fig. 9 (download XLSX )
RT–qPCR results.
Source Data Extended Data Fig. 10 (download XLSX )
Raw measurements for the assays indicated in the figure legend.
Rights and permissions
About this article
Cite this article
Mazan-Mamczarz, K., Tsitsipatis, D., Childs, B.G. et al. Single-cell and spatial transcriptomics map senescent vascular cells in arterial remodeling during atherosclerosis in mice. Nat Aging 5, 1528–1547 (2025). https://doi.org/10.1038/s43587-025-00889-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-00889-z
This article is cited by
-
Immune-mediated interactions between sarcopenia and atherosclerosis in aging
Immunity & Ageing (2025)
-
Vascular smooth muscle cell metabolic reprogramming and phenotypic remodeling in atherosclerosis
Cell Death Discovery (2025)