Abstract
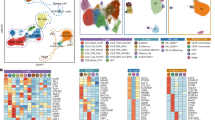
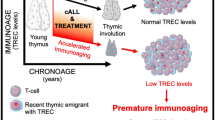
Age-related thymic involution increases vulnerability to cancers and infection in older adults, yet the driving mechanisms and its impact on peripheral T cells remain unclear. Using single-cell sequencing, we here analyzed 387,762 cells from human thymus and peripheral blood of young and aged individuals. Within thymus, we found aging reduced T-lineage potential in early thymic progenitors but increased innate lymphocyte lineage potential. Aged thymus were enriched in mature T cells with low SOX4 expression and inflammatory profiles but depleted of thymic epithelial cells and expression of tissue-restricted antigens. In the periphery, we identified transcriptional features of T cell aging and established a naive T cell-based model for immune age prediction. Furthermore, we identified CD38 as a marker of recent thymic emigrants. Finally, single-cell T cell receptor (TCR) repertoire sequencing identified shifts in TCR repertoire diversity within memory/effector T cells and expanded virus-specific T cells during aging. Collectively, our data offer insights into human thymic involution and peripheral T cell aging and could inform strategies to restore compromised T cell immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
scRNA-seq, scTCR-seq and CITE-seq datasets generated in this study have been deposited in NCBI GEO with the accession code GSE231906. Source data are provided within this paper. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Computer code used in this study has been deposited to the Github repository (https://github.com/dengyj-2019/thymus_and_periphery_aging) for public access.
References
Lobach, D. F. & Haynes, B. F. Ontogeny of the human thymus during fetal development. J. Clin. Immunol. 7, 81–97 (1987).
Steinmann, G. G. Changes in the human thymus during aging. Curr. Top. Pathol. 75, 43–88 (1986).
Rezzani, R., Nardo, L., Favero, G., Peroni, M. & Rodella, L. F. Thymus and aging: morphological, radiological, and functional overview. Age 36, 313–351 (2014).
Mackall, C. L., Punt, J. A., Morgan, P., Farr, A. G. & Gress, R. E. Thymic function in young/old chimeras: substantial thymic T cell regenerative capacity despite irreversible age-associated thymic involution. Eur. J. Immunol. 28, 1886–1893 (1998).
Jamieson, B. D. et al. Generation of functional thymocytes in the human adult. Immunity 10, 569–575 (1999).
Thome, J. J. et al. Longterm maintenance of human naive T cells through in situ homeostasis in lymphoid tissue sites. Sci. Immunol. 1, eaah6506 (2016).
Kooshesh, K. A., Foy, B. H., Sykes, D. B., Gustafsson, K. & Scadden, D. T. Health consequences of thymus removal in adults. N. Engl. J. Med. 389, 406–417 (2023).
Valkenburg, S. A. et al. Early priming minimizes the age-related immune compromise of CD8⁺ T cell diversity and function. PLoS Pathog. 8, e1002544 (2012).
Qi, Q. et al. Diversity and clonal selection in the human T-cell repertoire. Proc. Natl Acad. Sci. USA 111, 13139–13144 (2014).
Egorov, E. S. et al. The changing landscape of naive T cell receptor repertoire with human aging. Front Immunol. 9, 1618 (2018).
Goronzy, J. J. & Weyand, C. M. T cell development and receptor diversity during aging. Curr. Opin. Immunol. 17, 468–475 (2005).
Mittelbrunn, M. & Kroemer, G. Hallmarks of T cell aging. Nat. Immunol. 22, 687–698 (2021).
Hashimoto, K. et al. Single-cell transcriptomics reveals expansion of cytotoxic CD4 T cells in supercentenarians. Proc. Natl Acad. Sci. USA 116, 24242–24251 (2019).
Sayed, N. et al. An inflammatory aging clock (iAge) based on deep learning tracks multimorbidity, immunosenescence, frailty and cardiovascular aging. Nat. Aging 1, 598–615 (2021).
Kousa, A. I. et al. Age-related epithelial defects limit thymic function and regeneration. Nat. Immunol. 25, 1593–1606 (2024).
Gray, D. H. et al. Developmental kinetics, turnover, and stimulatory capacity of thymic epithelial cells. Blood 108, 3777–3785 (2006).
Bertho, J. M. et al. Phenotypic and immunohistological analyses of the human adult thymus: evidence for an active thymus during adult life. Cell Immunol. 179, 30–40 (1997).
Park, J. E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Lavaert, M. et al. Integrated scRNA-seq identifies human postnatal thymus seeding progenitors and regulatory dynamics of differentiating immature thymocytes. Immunity 52, 1088–1104 (2020).
Le, J. et al. Single-cell RNA-seq mapping of human thymopoiesis reveals lineage specification trajectories and a commitment spectrum in T cell development. Immunity 52, 1105–1118 (2020).
Ma, S. et al. Spatial transcriptomic landscape unveils immunoglobin-associated senescence as a hallmark of aging. Cell 187, 7025–7044 (2024).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Chopp, L. B. et al. An integrated epigenomic and transcriptomic map of mouse and human αβ T cell development. Immunity 53, 1182–1201 (2020).
Nobrega, C. et al. T cells home to the thymus and control infection. J. Immunol. 190, 1646–1658 (2013).
Bautista, J. L. et al. Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla. Nat. Commun. 12, 1096 (2021).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Romi, F., Aarli, J. A. & Gilhus, N. E. Myasthenia gravis patients with ryanodine receptor antibodies have distinctive clinical features. Eur. J. Neurol. 14, 617–620 (2007).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Mogilenko, D. A. et al. Comprehensive profiling of an aging immune system reveals clonal GZMK+ CD8+ T cells as conserved hallmark of inflammaging. Immunity 54, 99–115 (2021).
Satoh, Y. et al. The Satb1 protein directs hematopoietic stem cell differentiation toward lymphoid lineages. Immunity 38, 1105–1115 (2013).
Wang, Y. et al. Integrating single-cell RNA and T cell/B cell receptor sequencing with mass cytometry reveals dynamic trajectories of human peripheral immune cells from birth to old age. Nat. Immunol. 26, 308–322 (2025).
Kock, K. H. et al. Asian diversity in human immune cells. Cell 188, 2288–2306 (2025).
Luo, O. J. et al. Multidimensional single-cell analysis of human peripheral blood reveals characteristic features of the immune system landscape in aging and frailty. Nat. Aging 2, 348–364 (2022).
Zhu, H. et al. Human PBMC scRNA-seq-based aging clocks reveal ribosome to inflammation balance as a single-cell aging hallmark and super longevity. Sci. Adv. 9, eabq7599 (2023).
Liu, C. et al. Time-resolved systems immunology reveals a late juncture linked to fatal COVID-19. Cell 184, 1836–1857 (2021).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913 (2021).
Thompson, J. S. et al. The immune status of healthy centenarians. J. Am. Geriatr. Soc. 32, 274–281 (1984).
Karagiannis, T. T. et al. Multi-modal profiling of peripheral blood cells across the human lifespan reveals distinct immune cell signatures of aging and longevity. EBioMedicine 90, 104514 (2023).
Wang, M. et al. HIV-1-infected T cell clones are shared across cerebrospinal fluid and blood during ART. JCI Insight 9, e176208 (2024).
Chen, C., et al. The core cellular network modulates immune phenotype switching in hepatitis B. Sci. Bull. 70, 2717–2720.
Nehar-Belaid, D. et al. Mapping systemic lupus erythematosus heterogeneity at the single-cell level. Nat. Immunol. 21, 1094–1106 (2020).
Okuzono, Y. et al. B-cell immune dysregulation with low soluble CD22 levels in refractory seronegative myasthenia gravis. Front Immunol. 15, 1382320 (2024).
Tanaskovic, S., Fernandez, S., Price, P., Lee, S. & French, M. A. CD31 (PECAM-1) is a marker of recent thymic emigrants among CD4+ T-cells, but not CD8+ T-cells or gammadelta T-cells, in HIV patients responding to ART. Immunol. Cell Biol. 88, 321–327 (2010).
Haines, C. J. et al. Human CD4+ T cell recent thymic emigrants are identified by protein tyrosine kinase 7 and have reduced immune function. J. Exp. Med. 206, 275–285 (2009).
McFarland, R. D., Douek, D. C., Koup, R. A. & Picker, L. J. Identification of a human recent thymic emigrant phenotype. Proc. Natl Acad. Sci. USA 97, 4215–4220 (2000).
Bohacova, P. et al. Multidimensional profiling of human T cells reveals high CD38 expression, marking recent thymic emigrants and age-related naive T cell remodeling. Immunity 57, 2362–2379 (2024).
Lanfermeijer, J., Borghans, J. A. M. & van Baarle, D. How age and infection history shape the antigen-specific CD8+ T-cell repertoire: Implications for vaccination strategies in older adults. Aging Cell 19, e13262 (2020).
Smithey, M. J. et al. Lifelong CMV infection improves immune defense in old mice by broadening the mobilized TCR repertoire against third-party infection. Proc. Natl Acad. Sci. USA 115, E6817–E6825 (2018).
Reinhardt, R. L., Khoruts, A., Merica, R., Zell, T. & Jenkins, M. K. Visualizing the generation of memory CD4 T cells in the whole body. Nature 410, 101–105 (2001).
Agus, D. B., Surh, C. D. & Sprent, J. Reentry of T cells to the adult thymus is restricted to activated T cells. J. Exp. Med. 173, 1039–1046 (1991).
Jang, G. et al. Stem cell decoupling underlies impaired lymphoid development during aging. Proc. Natl Acad. Sci. USA 120, e2302019120 (2023).
Elsaid, R. et al. A wave of bipotent T/ILC-restricted progenitors shapes the embryonic thymus microenvironment in a time-dependent manner. Blood 137, 1024–1036 (2021).
Kadouri, N., Nevo, S., Goldfarb, Y. & Abramson, J. Thymic epithelial cell heterogeneity: TEC by TEC. Nat. Rev. Immunol. 20, 239–253 (2020).
Lehar, S. M., Dooley, J., Farr, A. G. & Bevan, M. J. Notch ligands Delta 1 and Jagged1 transmit distinct signals to T-cell precursors. Blood 105, 1440–1447 (2005).
Lancaster, J. N. et al. Central tolerance is impaired in the middle-aged thymic environment. Aging Cell 21, e13624 (2022).
Cowan, J. E., Takahama, Y., Bhandoola, A. & Ohigashi, I. Postnatal involution and counter-involution of the thymus. Front. Immunol. 11, 897 (2020).
Cheng, K. et al. Spatial interactions of immune cells as potential predictors to efficacy of toripalimab plus chemotherapy in locally advanced or metastatic pancreatic ductal adenocarcinoma: a phase Ib/II trial. Signal Transduct. Target Ther. 9, 321 (2024).
Gayoso., A., Shor., J., Carr., A.J., Sharma., R. & Pe’er., D. GitHub: DoubletDetection. Zenodo https://doi.org/10.5281/zenodo.2678041 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J. P. GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23, 3251–3253 (2007).
Clarke, J. et al. Single-cell transcriptomic analysis of tissue-resident memory T cells in human lung cancer. J. Exp. Med. 216, 2128–2149 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Hou, R., Denisenko, E., Ong, H. T., Ramilowski, J. A. & Forrest, A. R. R. Predicting cell-to-cell communication networks using NATMI. Nat. Commun. 11, 5011 (2020).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Schnell, A. et al. Stem-like intestinal Th17 cells give rise to pathogenic effector T cells during autoimmunity. Cell 184, 6281–6298 (2021).
Acknowledgements
We thank members of the flow cytometry core laboratory and histological core laboratory at West China Hospital, Sichuan University for technical help. This study was supported by grants from the Chongqing International Institute for Immunology (2020YJC01), National Natural Science Foundation of China (82025002, 32230036, 82394411, 82394410, 82170113 and 81670327), the 1.3.5 Project of disciplines of excellence (ZYGD22005, RHM24203 and ZYGD20010) and National Clinical Research Center for Geriatrics (Z20201001 and Z2021JC009), West China Hospital, Sichuan University.
Author information
Authors and Affiliations
Contributions
Y.D. performed the majority of data analysis; Z.P., K.M., X.Q., B.Y. and Y.L. processed the samples and performed the biological experiments, with help from L.L., H.Y., H.C., X.Z., K.W. and P.G.; P.Y., H.W., Y.Z., Y.X., K.Z., Q.H. and W.G. collected and validated human samples; B.D., X.C., X.W. and J.H. supervised human sample collection; Y.D., Z.P., K.M., X.Q., B.Y. and Y.L. prepared the figures and tables; and H.Z., Y.W. and H.H. conceived of the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The dynamic changes of thymocytes with aging.
(a-b) Uniform and manifold approximation and Projection (UMAP) visualization of thymocyte clusters (a), based on the differentially expressed genes (DEGs) visualized in violin plots (b). Colors represent the annotation of cell identity. (c-d) Representative flow cytometry data of CD45+ and CD45− cells in thymus across age groups (c). Statistics of the percentages of CD45+ and CD45− cells in live cells, along with the total cell number in thymi across Groups I to III (d) (n = 9 for Group III; n = 7 for Groups I and II). P values (from left to right): ns, P = 0.8451; **P = 0.0025; **P = 0.0094; ns, P = 0.8428; **P = 0.0025; **P = 0.0096; ****P < 0.0001; ****P < 0.0001; ns, P = 0.5519; **P = 0.0020; ***P = 0.0003; ns, P = 0.8186. (e) Representative flow cytometry data of T, CD19+ B and CD33+ myeloid cells in thymus across age groups. All cells were firstly gated on CD45+ cells. T cells are divided into double negative (DN), double positive (DP), CD4 single positive (SP) and CD8 SP cells, gated on CD45+CD19−CD33− cells. (f) Statistics of cell numbers of DN, DP, CD4 SP, CD8 SP, CD19+ B cells and CD33+ myeloid cells in CD45+ cells in thymi across Groups I to III (n = 9 for Group III; n = 7 for Groups I and II). P values (row-wise: from left to right): ***P = 0.0002; ****P < 0.0001; ns, P = 0.6188; ****P < 0.0001; ****P < 0.0001; ns, P = 0.5985; ****P < 0.0001; ****P < 0.0001; ns, P = 0.4760; ***P = 0.0003; ****P < 0.0001; ns, P = 0.6310; **P = 0.0015; ***P = 0.0005; ns, P = 0.9676; *P = 0.0183; **P = 0.0081; ns, P = 0.9814. (g-h) Flow cytometry gating strategies (g) and representative data (h) of NK cells, ILC in thymi across Groups I to III. All cells are gated on Lin−CD45+ cells. NK, CD56+; ILC1, CD56−CD161+ CD117−CRTH2−; ILC2, CD56−CD161+ CRTH2+; ILC3, CD56−CD161+ CD117+CRTH2−. (i) Statistics of the percentages of ILCs and NK in Lin−CD45+ cells in thymus with aging (n = 7 for Group II; n = 9 for Groups I and III). P values (from left to right): ns, P = 0.5107; ***P = 0.0006; *P = 0.0178; ns, P = 0.9917; *P = 0.0302; ns, P = 0.0570; ns, P = 0.5592; ***P = 0.0003; **P = 0.0084; ns, P = 0.3243; *P = 0.0143; ns, P = 0.3498; ns, P = 0.5336; ***P = 0.0008; *P = 0.0203. All data represent biological replicates. Data are presented as mean ± standard error of mean (s.e.m.). Statistical analysis was performed using Bonferroni-adjusted two-tailed Wilcoxon rank sum test (b) or One-way ANOVA with Tukey’s multiple comparisons test (d, f, i).
Extended Data Fig. 2 Reduced T cell differentiation lineage potentials in aged human thymus.
(a) Dot plots of DEGs among DN cells clusters described in Fig. 2a. Colors represent the scaled expression and dot sizes encode the proportion of gene-expressing cells. (b-c) UMAP visualizations of Monocle3 derived differentiation trajectory of DN cells. Colors indicate cell identity (b) or pseudotime (c). (d) Dot plot shows the transcription levels of CD3, CD4 and CD8 among DN cell subsets and DP cells. Dot colors represent the scaled expression and dot sizes encode the proportion of gene-expressing cells. (e) Heatmaps indicate the age-associated genes (adjusted P values < 0.05, fold change > 1.25) in DN-Q1 (top) and DN-Q2 (bottom) cells. (f) Violin plots of DEGs in each lineage engaged in STEMNET analysis. (g) Statistics of the proportion of T cells differentiated from thymic progenitor cells including Thy1 (CD45+CD34+CD7−CD1a−) and Thy2a (CD45+CD34+CD7+CD1a−CD44+CD2−) sorted from thymi across different age groups in artificial thymic organoid (ATO) (n = 5 for Group I; n = 3 for Group II; n = 4 for Group III). P values (row-wise: from left to right): P = 0.0726; *P = 0.0333; ns, P = 0.9686; P = 0.1394; P = 0.0983; ns, P = 0.9992; ns, P = 0.2285; ns, P = 0.1830; ns, P > 0.9999; ns, P = 0.1562; *P = 0.0259;ns, P = 0.6752; P = 0.0603; **P = 0.0039; ns, P = 0.3886; ns, P = 0.3410; ns, P = 0.1512; ns, P = 0.9180. Data are presented as mean ± s.e.m. (h) Representative flow cytometry data of differentiation results of CD45+CD34+ thymic progenitor cells including Thy1 and Thy2a sorted from thymi across age groups under T cell differentiation conditions, which are derived from 4 independent experiments. All data represent biological replicates. Statistical analysis was performed using Bonferroni-adjusted two-tailed Wilcoxon rank sum test (a, d, e, f) or One-way ANOVA with Tukey’s multiple comparisons test (g).
Extended Data Fig. 3 Aged-related genes and pathways in DP to SP cells.
(a) Violin plots show the DEGs of intrathymic conventional CD8 T cells, CD8ααT and nature killer T cells (NKTs) described in Fig. 3c. (b) The proportions of thymocytes from DP to SP stages change with aging. (c) Heatmaps display the age-related genes (adjusted P values < 0.05, fold change > 1.1) of DP-P (left) and DP-Q (right) cells in different age groups. (d) Dot plots show the age-related pathways (adjusted P values < 0.05) identified by Single Sample Gene Set Enrichment Analysis (ssGSEA), in DP-P (top) and DP-Q (bottom) cells across age groups. Dot colors represent the scaled enrichment score while dot sizes encode -log10 adjusted P values. Annotation colors represent the age group information. (e) Heatmaps display the age-related genes (adjusted P values < 0.05, fold change > 1.25) of mature CD4 (left) and CD8 (right) T cells in different age groups. All data represent biological replicates. Statistical analysis was performed using Bonferroni-adjusted two-tailed Wilcoxon rank sum test (a, c, d, e).
Extended Data Fig. 4 The functional decay of thymic stromal cell with age.
(a) Dot plot shows the representative DEGs of all thymic stromal populations described in Fig. 4a. Dot colors represent the scaled expression and dot sizes encode the proportion of gene-expressing cells. Bonferroni-adjusted two-tailed Wilcoxon rank sum test. (b) The proportions of thymic stromal cell populations excluding sorted EpCAM+ cells in each age group. (c) Bar plot filled by donor information, indicates that the cells of each thymic epithelial cell (TEC) subset in each age group are derived from multiple donors. (d) The distribution of TEC subsets in each age group. (e) Heatmap shows the systematic changes of age-associated genes (Bonferroni-adjusted two-tailed Wilcoxon rank sum test, adjusted P values < 0.05, fold change > 1.5) in TEC subsets, which are clustered into 5 gene modules via hierarchical clustering method. Heatmap tile colors represent fold change. Annotation tile colors represent the cell identity or the age group information. Significant genes (left) and related pathways (Benjamini-Hochberg adjusted hypergeometric test, adjusted P < 0.05) enriched by over representative analysis (ORA) (right) are colored according to the related gene module. (f) Dot plot shows the expression of tissue-restricted antigens (TRAs) from different tissues in different TEC clusters. Dot colors represent the scaled TRA scores, and dot sizes encode the proportion of TRA-expressing cells. (g) Bar plot shows the proportions of skeletal muscle-restricted genes regulated by transcription factors (TFs) highly expressed in TEC-myo (adjusted P values < 0.05, fold change > 1.5). (h) The scatter plot illustrates the Pearson correlation between the expression levels of the top 5 TFs, as shown in Extended Data Fig. 4g, and the expression of skeletal muscle-restricted antigens in TEC-myo. P values were calculated via two-tailed t-test. (i) Violin plots visualize the expression of TFs positively correlated with the expression of skeletal muscle-restricted antigens, as indicated in Extended Data Fig. 4h, in TEC-myo during aging (83 cells for Group I; 138 cells for Group II; 57 cells for Group III). Benjamini-Hochberg adjusted two-tailed Wilcoxon rank sum test. Data are presented as mean ± s.e.m. P values (from left to right): ****P < 0.0001; ****P < 0.0001; ****P < 0.0001; ns, P = 0.311; ****P < 0.0001; ****P < 0.0001; **P = 0.005; ***P = 0.0001; P = 0.0768; ns, P = 0.892; ***P = 0.0003; **P = 0.0014. All data represent biological replicates.
Extended Data Fig. 5 Quantification of multiple immunofluorescence staining validates the age-related changes of interaction between stromal cells and T cell progenitors.
(a) Quantitative ImageJ-based immunofluorescence analysis of age-related changes in the proportions of DLL4+CD205+ cortical TECs (cTECs), JAG1+PDGFRα+ fibroblasts (FBs) and NOTCH1+CD34+CD45+ T-progenitor cells (b) (n = 3 for all Groups). P values (from left to right): P = 0.6425; P = 0.3535; P = 0.8385; P = 0.8466; P = 0.1221; P = 0.2472; P = 0.8851; P = 0.0791; *P = 0.0438. (b-c) Spatial immunofluorescence quantitative analysis of the proportion of cTEC-T- progenitor cell (b) or FB-T-progenitor cell (c) interaction pairs across age groups, which is defined as nuclei centroids ≤15 μm apart, n = 3. (d-e) Representative flow cytometry plots (top) and quantitative analysis (bottom) of CD3+TCRβ+ T cell (d) or CD3−CD161+ ILC (e) populations derived from Thy1 progenitor cells of Groups I and III, following differentiation in the JAG1-ATO system (n = 3 for all Groups). P values (from left to right): P = 0.2802; **P = 0.0083. All data represent biological replicates. Data are presented as mean ± s.e.m. Statistical analysis was performed using One-way ANOVA with Tukey’s multiple comparisons test (a) and two-tailed t-test (d, e).
Extended Data Fig. 6 The age-related changed in the peripheral T cells.
(a) UMAP visualization of peripheral blood mononuclear cell (PBMC) clusters. Colors indicate the annotation of cell identity. (b) Heatmap showing the DEGs of PBMC clusters derived from Extended Data Fig. 6a. (c) The proportions of cell clusters of PBMCs changed with aging. (d) The DEGs of peripheral T cell clusters derived from Fig. 5a. Dot colors represent the scaled expression and dot sizes encode the proportion of gene-expressing cells. (e) Pearson correlation between the chronological and predicted age (based on peripheral pan-T cells) in the training cohorts (left, n = 394 samples) or external healthy validation cohorts (right, n = 81 samples). All data represent biological replicates. Statistical analysis was performed using Bonferroni-adjusted two-tailed Wilcoxon rank sum test (b, d) or two-tailed t-test (e).
Extended Data Fig. 7 Identification of human Recent Thymic Emigrants (RTEs).
(a-b) Partition-based graph abstraction (PAGA) results show the differentiation/development trajectory of peripheral CD4 (a) and CD8 (b) T cells, with edge width reflects the strength of connectivity between cell clusters, and node colors denote the cell identity annotation. Dot colors represent cell identity. (c-d) Spearman correlation among CD4 (c) and CD8 (d) intrathymic mature SOX4hi/lo SP, peripheral RTEs and naive T cells. (e-f) DEGs among CD4 (e) and CD8 (f) intrathymic mature SOX4hi/lo SP, peripheral RTEs and naive T cells. Bonferroni-adjusted two-tailed Wilcoxon rank sum test. All data represent biological replicates.
Extended Data Fig. 8 Potential connection of intrathymic and peripheral T cells.
(a-c) Dot plots illustrate the age-related changes in TCR V/J gene segment proportions during T cell development. Proportions significantly changed (Benjamini-Hochberg adjusted tow-tailed Wilcoxon rank sum test, P < 0.05) in at least one of the T cell development stages are displayed, with significant differences highlighted in red boxes. (d) The Pielou’s evenness index of TCR repertoire of intrathymic (top, n = 18) and peripheral T cell populations (bottom, n = 22). Kruskal-Wallis test. Colors indicate cell populations. The box in the boxplot represents the interquartile range (IQR), with its lower and upper edges indicating the 25th and 75th percentiles, respectively. The medium value (50th percentile) is shown within the box, and the whiskers extend to the minimum and maximum values within 1.5 times the IQR of the quartiles. (e) TCR similarity score (TRSS) reflected the similarity in clonal distribution across two sets of cells. Colors and sizes represent TRSS. The numbers on the x-axis represent ages of individual samples. (f) GSEA comparison of intrathymic memory-like CD8 T cells and peripheral CD8 TCM (left) or CD8 TEM-K (GZMK+ CD8 effector memory, right). Permutation test. Absolute normalized enrichment score (NES) > 1, P values < 0.05 and adjusted P values < 0.25. (g) Dot plot shows TRSS of peripheral CD4 subsets and intrathymic memory-like CD4 T cells, colors and sizes represent TRSS. (h) Spearman correlation of peripheral CD4 subsets and intrathymic memory-like CD4 T cells. (i) Dot plots show TRSS of peripheral CD4 TCM (central memory), CD4 TEFF (effector), CD4 TEM (effector memory), CD4 CTL (cytotoxic T lymphocyte) and intrathymic memory-like CD4 T cells with age. Colors and sizes represent TRSS. (j) Spearman correlation of peripheral CD4 TCM, CD4 TEFF, CD4 TEM, CD4 CTL and intrathymic memory-like CD4 T cells with age. All data represent biological replicates.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1. Gating strategies used for flow cytometry.
Supplementary Table 1 (download XLSX )
Volunteer information for different experiments, including sex, age, tissues, cell type and etc.
Supplementary Table 2 (download XLSX )
Reagent and resource include flow antibody, chemicals, peptides, recombinant proteins,software and algorithms.
Supplementary Table 3 (download XLSX )
Cell number of major cell type across different age groups.
Supplementary Table 4 (download XLSX )
Top 30 maker genes of major cell cluster test by Wilcoxon rank-sum test with P value adjusted for multiple testing by Bonferroni method.
Supplementary Table 5 (download XLSX )
DEGs within a specific cell type across different age groups were test by Wilcoxon rank-sum test with P value adjusted for multiple testing by Bonferroni method.
Supplementary Table 6 (download XLSX )
Dynamic pathways (pathways matrices were calculated by ssGSEA algorithm) within a specific cell type across different age groups were test by Wilcoxon rank-sum test with P value adjusted for multiple testing by Bonferroni method.
Supplementary Table 7 (download XLSX )
Over representation analysis in age-related gene module. P values were adjusted for multiple testing by Benjamini-Hochberg method.
Supplementary Table 8 (download XLSX )
Crosstalk between TEC and ETP-TP cells, as well as FB and ETP-TP cells during aging, related to Fig. 4.
Supplementary Table 9 (download XLSX )
Pathways enriched by genes positively or negatively correlated with AgeDiff (predicted age - actual chronological age), calculated using GSEA with P values detected by permutation test.
Supplementary Table 10 (download XLSX )
List of nucleotide sequences of the plasmid MigR1 and the gene JAG1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deng, Y., Peng, Z., Ming, K. et al. Single-cell analysis of human thymus and peripheral blood unveils the dynamics of T cell development and aging. Nat Aging 5, 2494–2513 (2025). https://doi.org/10.1038/s43587-025-00990-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-00990-3