Abstract
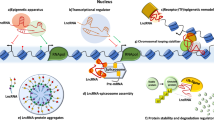
Long noncoding RNAs (lncRNAs) regulate transcriptional and epigenetic programs during aging and senescence. However, no comprehensive studies have systematically integrated multilayered analyses to reveal their diverse regulatory roles. Moreover, lncRNAs with therapeutic potential in age-related diseases remain unexplored. Here we systematically perturbed 32 high-abundance aging- and senescence-associated lncRNAs (PtbAlncs) using a Perturb-seq-based CRISPR–dCas9–KRAB knockdown system coupled with single-nucleus multiomics profiling, enabling simultaneous transcriptomic and chromatin accessibility analysis. This analysis uncovered essential roles for previously uncharacterized lncRNAs in senescence regulation, validated computationally and experimentally. These lncRNAs modulate distinct single-cell RNA-sequencing modules through diverse yet overlapping epigenetic motifs in single-cell ATAC-sequencing modules. Among them, HOTAIRM1, a DNA repair-associated PtbAlnc, stabilizes DNA repair by cooperating with BANF1 and p53 at double-strand break loci within condensates. Its deficiency impairs DNA repair and triggers p53-mediated senescence. In aged mouse lungs, adeno-associated virus-mediated HOTAIRM1 overexpression reduced fibrosis, alleviated tissue damage, and promoted cellular proliferation, underscoring its therapeutic potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Public datasets used for lncRNA selection include GEO: GSE106670, GSE109700, and OMIX: OEP001041. Tissue-wide lncRNA expression data included GTEx and GEO: GSE165192. Additional RBP profiling for HOTAIRM1 used PRIDE: PXD034470; BANF1 ChIP–seq used GEO: GSE227263. Sequencing data generated in this study (Perturb-multiome single cell, CHIRP-seq, and RNA-seq) have been deposited in the Genome Sequence Archive at the China National Center for Bioinformation (CNCB-NGDC) under accession PRJCA057749 (HRA016755 for Perturb-multiome single cell data; HRA016746 for CHIRP-seq and RNA-seq). Mass spectrometry data generated in this study have been deposited in PRIDE under dataset identifier PXD074193.
Code availability
Custom scripts framework used for processing and analysis of the Perturb-multiome datasets are available via Github at https://github.com/Ramzipits/Perturb-multiome and archived on Zenodo at https://doi.org/10.5281/zenodo.18559236 (ref. 97).
References
Melzer, D., Pilling, L. C. & Ferrucci, L. The genetics of human ageing. Nat. Rev. Genet. 21, 88–101 (2020).
Han, J. D. J. The ticking of aging clocks. Trends Endocrinol. Metab. 35, 11–22 (2024).
Wu, Z. M., Zhang, W. Q., Qu, J. & Liu, G. H. Emerging epigenetic insights into aging mechanisms and interventions. Trends Pharmacol. Sci. 45, 157–172 (2024).
Singh, P. P., Demmitt, B. A., Nath, R. D. & Brunet, A. The genetics of aging: a vertebrate perspective. Cell 177, 200–220 (2019).
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. Hallmarks of aging: an expanding universe. Cell 186, 243–278 (2023).
Song, S. L., Lam, E. W. F., Tchkonia, T., Kirkland, J. L. & Sun, Y. Senescent cells: emerging targets for human aging and age-related diseases. Trends Biochem. Sci. 45, 578–592 (2020).
Zhang, W., Sun, H. S., Wang, X., Dumont, A. S. & Liu, Q. Cellular senescence, DNA damage, and neuroinflammation in the aging brain. Trends Neurosci. 47, 461–474 (2024).
Yousefzadeh, M. J. et al. An aged immune system drives senescence and ageing of solid organs. Nature 594, 100–105 (2021).
Herr, L. M., Schaffer, E. D., Fuchs, K. F., Datta, A. & Brosh, R. M. Jr. Replication stress as a driver of cellular senescence and aging. Commun. Biol. 7, 616 (2024).
Kamiya, M. et al. Immune mechanisms in fibrotic interstitial lung disease. Cell 187, 3506–3530 (2024).
O’Reilly, S., Tsou, P. S. & Varga, J. Senescence and tissue fibrosis: opportunities for therapeutic targeting. Trends Mol. Med. 30, 1113–1125 (2024).
Ferrarelli, L. K. Promoting DNA repair to prevent fibrosis. Sci. Signal. 17, eado0928 (2024).
Chaib, S., Tchkonia, T. & Kirkland, J. L. Cellular senescence and senolytics: the path to the clinic. Nat. Med. 28, 1556–1568 (2022).
Eming, S. A., Wynn, T. A. & Martin, P. Inflammation and metabolism in tissue repair and regeneration. Science 356, 1026–1030 (2017).
Rossi, M. & Gorospe, M. Noncoding RNAs controlling telomere homeostasis in senescence and aging. Trends Mol. Med. 26, 422–433 (2020).
Yang, Z. et al. LncRNA: shedding light on mechanisms and opportunities in fibrosis and aging. Ageing Res. Rev. 52, 17–31 (2019).
Herman, A. B., Tsitsipatis, D. & Gorospe, M. Integrated lncRNA function upon genomic and epigenomic regulation. Mol. Cell 82, 2252–2266 (2022).
Huang, Y. et al. Unraveling aging from transcriptomics. Trends Genet. 41, 218–235 (2025).
Baysoy, A., Bai, Z., Satija, R. & Fan, R. The technological landscape and applications of single-cell multi-omics. Nat. Rev. Mol. Cell. Biol. 24, 695–713 (2023).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 e17 (2016).
Replogle, J. M. et al. Mapping information-rich genotype-phenotype landscapes with genome-scale Perturb-seq. Cell 185, 2559–2575 e28 (2022).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Xia, X. et al. Three-dimensional facial-image analysis to predict heterogeneity of the human ageing rate and the impact of lifestyle. Nat. Metab. 2, 946–957 (2020).
Cheng, H. et al. Repression of human and mouse brain inflammaging transcriptome by broad gene-body histone hyperacetylation. Proc. Natl Acad. Sci. USA 115, 7611–7616 (2018).
Lee, S. et al. Molecular programs of fibrotic change in aging human lung. Nat. Commun. 12, 6309 (2021).
Grammatikakis, I., Panda, A. C., Abdelmohsen, K. & Gorospe, M. Long noncoding RNAs (lncRNAs) and the molecular hallmarks of aging. Aging 6, 992–1009 (2014).
Fang, S. et al. NONCODEV5: a comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 46, D308–D314 (2018).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376 e17 (2019).
Norman, T. M. et al. Exploring genetic interaction manifolds constructed from rich single-cell phenotypes. Science 365, 786–793 (2019).
Papalexi, E. et al. Characterizing the molecular regulation of inhibitory immune checkpoints with multimodal single-cell screens. Nat. Genet. 53, 322–331 (2021).
Jiang, L. D. et al. Systematic reconstruction of molecular pathway signatures using scalable single-cell perturbation screens. Nat. Cell Biol. 27, 505–517 (2025).
Luo, Y. et al. Long non-coding RNA NEAT1 promotes colorectal cancer progression by competitively binding miR-34a with SIRT1 and enhancing the Wnt/beta-catenin signaling pathway. Cancer Lett. 440-441, 11–22 (2019).
Shao, K., Xi, L., Cang, Z., Chen, C. & Huang, S. Knockdown of NEAT1 exerts suppressive effects on diabetic retinopathy progression via inactivating TGF-beta1 and VEGF signaling pathways. J. Cell. Physiol. 235, 9361–9369 (2020).
Liu, F., Tai, Y. & Ma, J. LncRNA NEAT1/let-7a-5p axis regulates the cisplatin resistance in nasopharyngeal carcinoma by targeting Rsf-1 and modulating the Ras-MAPK pathway. Cancer Biol. Ther. 19, 534–542 (2018).
Yu, H., Xu, A., Wu, B., Wang, M. & Chen, Z. Long noncoding RNA NEAT1 promotes progression of glioma as a ceRNA by sponging miR-185-5p to stimulate DNMT1/mTOR signaling. J. Cell. Physiol. 236, 121–130 (2021).
de Magalhaes, J. P. et al. Human Ageing Genomic Resources: updates on key databases in ageing research. Nucleic Acids Res. 52, D900–D908 (2024).
Leisegang, M. S., Warwick, T., Stotzel, J. & Brandes, R. P. RNA-DNA triplexes: molecular mechanisms and functional relevance. Trends Biochem. Sci. 49, 532–544 (2024).
Xiao, Y. et al. Tracking single-cell evolution using clock-like chromatin accessibility loci. Nat. Biotechnol. 43, 784–798 (2025).
Tao, W., Yu, Z. & Han, J. J. Single-cell senescence identification reveals senescence heterogeneity, trajectory, and modulators. Cell Metab. 36, 1126–1143 e5 (2024).
Guney, I., Wu, S. & Sedivy, J. M. Reduced c-Myc signaling triggers telomere-independent senescence by regulating Bmi-1 and p16(INK4a). Proc. Natl Acad. Sci. USA 103, 3645–3650 (2006).
Chan, M. et al. Novel insights from a multiomics dissection of the Hayflick limit. eLife 11, e70283 (2022).
Han, J. J. LncRNAs: the missing link to senescence nuclear architecture. Trends Biochem. Sci. 48, 618–628 (2023).
Cai, D. & Han, J. J. Aging-associated lncRNAs are evolutionarily conserved and participate in NFkappaB signaling. Nat. Aging 1, 438–453 (2021).
Zhang, X. et al. KCNQ1OT1 promotes genome-wide transposon repression by guiding RNA-DNA triplexes and HP1 binding. Nat. Cell Biol. 24, 1617–1629 (2022).
Chuang, T. W., Su, C. H., Wu, P. Y., Chang, Y. M. & Tarn, W. Y. LncRNA HOTAIRM1 functions in DNA double-strand break repair via its association with DNA repair and mRNA surveillance factors. Nucleic Acids Res. 51, 3166–3184 (2023).
Burgess, J. T. et al. Barrier-to-autointegration-factor (Banf1) modulates DNA double-strand break repair pathway choice via regulation of DNA-dependent kinase (DNA-PK) activity. Nucleic Acids Res. 49, 3294–3307 (2021).
Park, J. H., Park, E. J., Hur, S. K., Kim, S. & Kwon, J. Mammalian SWI/SNF chromatin remodeling complexes are required to prevent apoptosis after DNA damage. DNA Repair 8, 29–39 (2009).
Yao, D. et al. Scalable genetic screening for regulatory circuits using compressed Perturb-seq. Nat. Biotechnol. 42, 1282–1295 (2024).
Talbott, H. E., Mascharak, S., Griffin, M., Wan, D. C. & Longaker, M. T. Wound healing, fibroblast heterogeneity, and fibrosis. Cell Stem Cell 29, 1161–1180 (2022).
Lu, T. C. et al. Aging Fly Cell Atlas identifies exhaustive aging features at cellular resolution. Science 380, 1145-+ (2023).
Zhu, H. M. et al. Human PBMC scRNA-seq-based aging clocks reveal ribosome to inflammation balance as a single-cell aging hallmark and super longevity. Sci. Adv. 9, eabq7599 (2023).
Teo, G. et al. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J. Proteomics 100, 37–43 (2014).
Jager, S. et al. Global landscape of HIV-human protein complexes. Nature 481, 365–370 (2011).
Jamin, A. & Wiebe, M. S. Barrier to Autointegration Factor (BANF1): interwoven roles in nuclear structure, genome integrity, innate immunity, stress responses and progeria. Curr. Opin. Cell Biol. 34, 61–68 (2015).
Mijit, M., Caracciolo, V., Melillo, A., Amicarelli, F. & Giordano, A. Role of p53 in the regulation of cellular senescence. Biomolecules 10, 420 (2020).
Shao, S. et al. TagBiFC technique allows long-term single-molecule tracking of protein-protein interactions in living cells. Commun. Biol. 4, 378 (2021).
Bauer, N. C. et al. A cross-nearest neighbor/Monte Carlo algorithm for single-molecule localization microscopy defines interactions between p53, Mdm2, and MEG3. J. Biol. Chem. 296, 100540 (2021).
Uroda, T. et al. Conserved pseudoknots in lncRNA MEG3 are essential for stimulation of the p53 pathway. Mol. Cell 75, 982–995 e9 (2019).
Wang, Y. H. et al. Rapid recruitment of p53 to DNA damage sites directs DNA repair choice and integrity. Proc. Natl Acad. Sci. USA 119, e2113233119 (2022).
Menon, V. & Povirk, L. Involvement of p53 in the repair of DNA double strand breaks: multifaceted Roles of p53 in homologous recombination repair (HRR) and non-homologous end joining (NHEJ). Subcell. Biochem. 85, 321–336 (2014).
Hafner, A., Bulyk, M. L., Jambhekar, A. & Lahav, G. The multiple mechanisms that regulate p53 activity and cell fate. Nat. Rev. Mol. Cell Biol. 20, 199–210 (2019).
Bolderson, E. et al. Barrier-to-autointegration factor 1 (Banf1) regulates poly [ADP-ribose] polymerase 1 (PARP1) activity following oxidative DNA damage. Nat. Commun. 10, 5501 (2019).
Heltberg, M. S. et al. Enhanced DNA repair through droplet formation and p53 oscillations. Cell 185, 4394–4408 e10 (2022).
Li, C. et al. Deciphering the molecular mechanism underlying morphology transition in two-component DNA-protein cophase separation. Structure 33, 62–77.e8 (2025).
Schneider, J. L. et al. The aging lung: physiology, disease, and immunity. Cell 184, 1990–2019 (2021).
McDonough, J. E. et al. Small-airway obstruction and emphysema in chronic obstructive pulmonary disease. N. Engl. J. Med. 365, 1567–1575 (2011).
Biffo, S., Ruggero, D. & Santoro, M. M. The crosstalk between metabolism and translation. Cell Metab. 36, 1945–1962 (2024).
Zhang, X. et al. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 113, 2526–2534 (2009).
Rea, J. et al. HOTAIRM1 regulates neuronal differentiation by modulating NEUROGENIN 2 and the downstream neurogenic cascade. Cell Death Dis. 11, 527 (2020).
Fu, L. et al. LncRNA HOTAIRM1 promotes osteogenesis by controlling JNK/AP-1 signalling-mediated RUNX2 expression. J. Cell. Mol. Med. 23, 7517–7524 (2019).
Liu, M. et al. The regulations of telomerase reverse transcriptase (TERT) in cancer. Cell Death Dis. 15, 90 (2024).
Bi, Y. et al. Exosomal miR-302b rejuvenates aging mice by reversing the proliferative arrest of senescent cells. Cell Metab. 37, 527–541 e6 (2025).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733 e12 (2016).
Chen, Z. H. et al. The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ. 24, 212–224 (2017).
Jing, Y. et al. Mutant NPM1-regulated lncRNA HOTAIRM1 promotes leukemia cell autophagy and proliferation by targeting EGR1 and ULK3. J. Exp. Clin. Cancer Res. 40, 312 (2021).
Liang, L. et al. The long noncoding RNA HOTAIRM1 controlled by AML1 enhances glucocorticoid resistance by activating RHOA/ROCK1 pathway through suppressing ARHGAP18. Cell Death Dis. 12, 702 (2021).
Khanduja, J. S., Calvo, I. A., Joh, R. I., Hill, I. T. & Motamedi, M. Nuclear noncoding RNAs and genome stability. Mol. Cell 63, 7–20 (2016).
Goni, E. et al. Uncovering functional lncRNAs by scRNA-seq with ELATUS. Nat. Commun. 15, 9709 (2024).
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011).
Yin, J. et al. Imaging of mRNA-protein interactions in live cells using novel mCherry trimolecular fluorescence complementation systems. PLoS ONE 8, e80851 (2013).
Miller, K. E., Kim, Y., Huh, W. K. & Park, H. O. Bimolecular fluorescence complementation (BiFC) analysis: advances and recent applications for genome-wide interaction studies. J. Mol. Biol. 427, 2039–2055 (2015).
Bennardo, N., Cheng, A., Huang, N. & Stark, J. M. Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair. PLoS Genet. 4, e1000110 (2008).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Buske, F. A., Bauer, D. C., Mattick, J. S. & Bailey, T. L. Triplexator: detecting nucleic acid triple helices in genomic and transcriptomic data. Genome Res. 22, 1372–1381 (2012).
Millstein, J., Chen, G. K. & Breton, C. V. cit: hypothesis testing software for mediation analysis in genomic applications. Bioinformatics 32, 2364–2365 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Giurgiu, M. et al. CORUM: the comprehensive resource of mammalian protein complexes-2019. Nucleic Acids Res. 47, D559–D563 (2019).
Oughtred, R. et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
& Cerezo, M. et al. The NHGRI-EBI GWAS Catalog: standards for reusability, sustainability and diversity. Nucleic Acids Res. 53, D998–D1005 (2025).
Zhu, S.X. & Han, J.-D. J. Ramzipits/Perturb-multiome: Perturb-multiome v1.0.0. Zenodo https://doi.org/10.5281/zenodo.18559236 (2026).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (32330017, 32088101, 92374207, 82361148130) and Beijing Natural Science Foundation (IS23077) to J.-D.J.H. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.-D.J.H. conceived and supervised the project. S.Z. conducted and helped conceive all computational analyses and performed all experiments of multiomic perturbation, RNA profiling, cell-based assay, and molecular assays. S.Y. helped conceive and execute the mechanism and mouse assays and conducted tissue staining and pathology quantification. S.Y. also assisted with cell-based assays. D.C. helped conceive the multiomic perturbation framework and contributed to bleomycin treatment transcriptomic data. Y.R. assisted with a small subset of vector construction. L.W. provided support with vector extraction and related assays. Z.Q. provided the truncated recombinant p53 protein. S.Z. and J.-D.J.H wrote the manuscript. V.G. suggested experiments and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.-D.J.H. and S.Z. are listed as inventors on a patent application related to the use of HOTAIRM1 for anti-aging therapy (Chinese patent application number 2026100645625). All other authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Mikel Hernaez, Xin Jin, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Selection of aging- and senescence-associated lncRNAs for single-cell CRISPR screening.
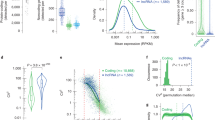
(a-b), Scatter plots showing highly abundant lncRNAs with significant correlations to aging in human blood and brain tissues. Spearman’s rank correlation (two-sided) was used to calculate correlation coefficients and P values. (c-d), Scatter plots showing highly abundant lncRNAs significantly correlated with early and late stages of human fibroblast senescence. e, Scatter plots showing highly abundant lncRNAs significantly associated with the DNA damage response in human fibroblasts. f, Venn diagram summarizing the overlap of candidate lncRNAs identified across dataset. P values in c-e were calculated using a Wilcoxon rank-sum test (two-sided). All P values were adjusted by Benjamini–Hochberg FDR.
Extended Data Fig. 2 Performance evaluation of Perturb-Multiome in PtbAlnc perturbation across chromatin and transcriptomic landscapes.
a, UMAP visualization of expression features from unfiltered Perturb-Multiome on K562 cells with PtbAlnc perturbation, colored by cell cycle states. b, UMAP visualization of chromatin accessibility features from unfiltered Perturb-Multiome on K562 cells with PtbAlnc perturbation, colored by cell cycle states. c, Heatmap showing the genotype distribution of K562 cells post cell identity assignment and guide barcode processing in Perturb-Multiome. d, Silhouette coefficients of K562 single cells for each perturbation group (NT, n = 512; PURPL, n = 45; LINC0511, n = 51; XIST, n = 18; FAM66C, n = 110; LINC01578, n = 72; LINC01638, n = 44; LINC00578, n = 93; MIR155HG, n = 22; MIR222HG, n = 65; NEAT1, n = 17; GAS5, n = 71; SNHG1, n = 60; HOTAIRM1, n = 15; LINC02693, n = 27; MEG3, n = 58) across RNA + ATAC integrated, RNA, and ATAC embeddings. The boxplot centerline represents the median, boxes indicate interquartile ranges; whiskers indicate minima and maxima. Higher silhouette values indicate greater within-cluster consistency. e, Pairwise Euclidean distances between cluster centroids across perturbation groups in the same embeddings as in d, shown as heatmaps. Larger distances indicate stronger separation between clusters. f, Dot plot summarizing significant knockdown effects in promoter chromatin accessibility and expression of target lncRNAs in K562 cells across PtbAlnc-targeted groups versus non-targeting (NT) controls (two-tailed Student’s t-test). g, RT-qPCR validation of CRISPRi-mediated lncRNA knockdown efficiency for target lncRNAs, comparing stable and transient sgRNA transduction. Error bars indicate the mean±s.e.m. from n = 3 independent experiments. h, Mixscape modeling of cell perturbation state scores, distinguishing NEAT1 knockdown (KD) and non-perturbed (NP) groups. i, Position Weight Matrix showing upregulated and downregulated DNA motifs following NEAT1 KD.
Extended Data Fig. 3 PtbAlnc triplex targeting landscape and enrichment of aging- and senescence-associated pathways.
a, Dot plot showing the chromosomal distribution of RNA-DNA triplex targeting sites (TTS) predicted for PtbAlncs, highlighting the proportion of TTS located within repetitive genomic regions. b, Bar plot depicting the enrichment of PtbAlnc RNA-DNA TTS in transposable elements. (c-h), Bar plots summarizing gene set enrichment analysis for 300 up- and down-regulated genes in KD cells of additional PtbAlncs. Enriched pathways related to aging and senescence were identified using the Enrichr package and the human WikiPathways database, with key pathways highlighted by red arrows. P values were calculated using two-sided Fisher’s exact test.
Extended Data Fig. 4 Additional analyses of Perturb-Multiome profiling in BJ cells.
a, Silhouette coefficients of BJ single cells for each perturbation group (NT, n = 698; GAS5, n = 86; LINC01011, n = 60; LINC00511, n = 81; LINC00578, n = 106; LINC01578, n = 80; LINC01638, n = 104; SNHG29, n = 31; MIR155HG, n = 50; MIR222HG, n = 92; PURPL, n = 120; SNHG1, n = 96; SNHG3, n = 67; FENDRR, n = 31; NEAT1, n = 89) across RNA + ATAC integrated, RNA, and ATAC embeddings. The boxplot centerline represents the median, boxes indicate interquartile ranges; whiskers indicate minima and maxima. b, Heatmaps of pairwise Euclidean distances between cluster centroids in BJ cells across RNA + ATAC integrated, RNA, and ATAC embeddings. c, Dot plot summarizing significant knockdown effects in promoter chromatin accessibility and expression of target lncRNAs in BJ cells across PtbAlnc-targeted groups versus non-targeting (NT) controls (two-tailed Student’s t-test).
Extended Data Fig. 5 Validation of lncRNA knockdown and additional support for senescence phenotype screening.
a, qPCR validation of lncRNA knockdown efficiency following siRNA transduction targeting specific genes. Data are from independent experiments (n = 3 for HOTAIRM1, LINC01638, MIR155HG, MEG3, LINC01578, FAM66C and LINC00578; n = 2 for NEAT1, GAS5, XIST, LINC02693, SNHG1, PURPL and MIR222HG). Data are presented as the mean±s.e.m. b, Violin plots quantifying cell counts per high-power field (HPF; 0.96 mm × 0.96 mm) in control siRNA and HOTAIRM1 siRNA groups (n = 50 HPFs per condition). For violin plot, the center line indicates the median; dotted lines indicate the 25th and 75th percentiles; the violin is truncated to the data range (min–max). P values were calculated by two-tailed Student’s t-test. c, SA-β-gal staining in WI-38 cells after siRNA-mediated knockdown of representative PtbAlncs that did not significantly affect senescence. Senescence ratios were quantified per high-power field (HPF; 0.96 mm × 0.96 mm) (control, n = 23; SNHG1, n = 10; MIR222HG, n = 10; LINC00578, n = 13; PURPL, n = 10) from 3 independent experiments. For boxplots, center lines indicate medians; boxes indicate the 25th–75th percentiles; whiskers indicate minima and maxima. P values were calculated by two-tailed Student’s t-test. Scale bar, 100 μm. d, Violin plots showing expression levels of senescence markers in BJ fibroblasts single cells from Perturb-multiome after knockdown of candidate lncRNAs (NT, n = 698; LINC01638, n = 104; MIR155HG, n = 50; NEAT1, n = 89; GAS5, n = 86; MEG3, n = 21; LINC01578, n = 80; XIST, n = 72; SNHG1, n = 96; MIR222HG, n = 92; LINC00578, n = 106; PURPL, n = 120). P values were calculated by two-tailed Student’s t-test. e, Violin plots of senescence markers expression in K562 single cells from Perturb-multiome (NT, n = 258; HOTAIRM1 KD, n = 39). P values were calculated by two-tailed Student’s t-test. For violin plots in d-e, the violin shape represents the data distribution and is truncated to the data range (min–max).
Extended Data Fig. 6 Correlation network analysis of genomic features affected by PtbAlnc perturbation.
a, Heatmap of Pearson correlation matrices for differentially expressed genes (DEGs) following PtbAlnc perturbation in non-targeting cells. Seven highly correlated RNA modules were identified through Ward’s hierarchical clustering. b, Heatmap of Pearson correlation matrices for differentially accessible (DA) motifs following PtbAlnc perturbation in non-targeting cells. Six highly correlated ATAC modules were identified through Ward’s hierarchical clustering, with key features annotated for each module. c, Heatmap showing the average chromVAR scores for RNA modules across different PtbAlnc perturbation groups, reflecting the expression abundance within each module. d, Heatmap displaying the average chromVAR scores for ATAC modules across different PtbAlnc perturbation groups, reflecting the chromatin accessibility within each module. e, Dot plot showing fold change in RNA module scores across different PtbAlnc perturbation groups compared to NT cells. f, Dot plot showing fold change in ATAC module scores across different PtbAlnc perturbation groups compared to NT cells.
Extended Data Fig. 7 HOTAIRM1 links aging-associated transcriptional programs to the DNA damage response and senescence progression.
a, Average expression of cytoplasmic ribosomal protein genes (Z-score normalized module score) was computed at the single-cell level using Seurat’s AddModuleScore() function, based on 17 genes from the ‘cytoplasmic ribosomal proteins’ pathway term significantly downregulated upon HOTAIRM1 knockdown (see Fig. 2e). Module scores were plotted against Epitrace-predicted age scores in K562 cells from our Perturb-Multiome dataset. Correlation coefficients and P values were calculated using Pearson’s correlation (two-sided). b, Bar plot showing increased HOTAIRM1 expression following bleomycin-induced DNA damage in IMR90 cells (<10 passages) RNA-seq data. Data are presented as mean±s.e.m; n = 2. P values were calculated by two-tailed Student’s t-test. c, Bar plot showing upregulation of XRCC5 and XRCC6 expression after bleomycin-induced DNA damage in IMR90 cells RNA-seq data. Data are presented as mean±s.e.m.; n = 2. P values were calculated by two-tailed Student’s t-test. d, RT–qPCR analysis of HOTAIRM1 relative abundance in BJ cells collected at the indicated time points after bleomycin treatment. Values are normalized to the corresponding 0 h sample. e, RT–qPCR analysis of XRCC6 and XRCC5 relative abundance in HEK293 (top) and BJ (bottom) cells collected at the indicated time points after bleomycin treatment. Values are normalized to BJ 0 h (set to 1). f, RT–qPCR analysis of CDKN1A relative abundance in BJ (left) cells and IL6 relative abundance in HEK293 (right top) and BJ (right bottom) cells collected at the indicated time points after bleomycin treatment. Values are normalized to BJ 0 h (set to 1). Data in d-f are presented as the mean±s.e.m. from n = 3 independent experiments.
Extended Data Fig. 8 PtbAlnc–protein interaction network and functional interplay between HOTAIRM1 and BANF1 in DNA repair.
(a-c), High-confidence interaction network between PtbAlnc (red diamonds) and associated proteins (circles), inferred from RNA-binding protein profiling. Protein complexes and biological processes are shaded in yellow and blue. The edge color reflects the MiST score, while edge thickness corresponds to spectral counts. Dashed circles indicate Gene Ontology (GO) annotations. d, Bar plot demonstrating the specific and high-confidence binding of BANF1 to HOTAIRM1. Each bait lncRNA was designed with two distinct groups of specific probes to validate interactions. e, Density plot showing the overlap of HOTAIRM1 and BANF1 binding peaks in DNA repair genes’ TSS (n = 32). f, Venn diagram showing overlap between DNA repair genes with CHIRP peaks and genes with predicted triplex sites in promoters (two-sided Fisher’s exact test, p = 0.035; background = 62,703 genes). g, Bar plot illustrating the pathway annotation of BANF1 ChIP peaks in promoter regions (KEGG2021, Enrichr; two-sided Fisher’s exact test). h, Venn diagram showing the overlap between HOTAIRM1 CHIRP peaks and BANF1 ChIP peaks. Statistical significance was assessed using two-sided Fisher’s exact test, with a background of 100,000 genome-wide peaks. i, Pie chart depicting the genomic distribution of overlapping peaks between HOTAIRM1 and BANF1. j, Bar plot displaying pathway enrichment for promoters bound by both HOTAIRM1 and BANF1, focusing on DNA repair pathways (KEGG2021, Enrichr; two-sided Fisher’s exact test).
Extended Data Fig. 9 Additional insights into the role of HOTAIRM1 in DNA repair, senescence, and condensate formation.
a, Relative abundance of TP53 in Perturb-RNA-seq in all K562 single cells (control, n = 258; HOTAIRM1 KD, n = 39). For violin plots, the violin shape represents the data distribution and is truncated to the data range (min–max). b, Bar plot showing qPCR results corresponding to Fig. 5a. c, Bar plot quantifying relative TP53 mRNA expression levels under each condition, measured by qPCR, corresponding to Fig. 5b. d, Quantification of γH2AX foci per nucleus in control (n = 39) and HOTAIRM1 siRNA–treated cells (n = 58) after bleomycin exposure. e, Representative immunofluorescence images of HEK293 cells transfected with empty vector (Control) or a HOTAIRM1 overexpression plasmid (HOTAIRM1 OE), stained for γH2AX (green) and nuclei (DAPI, blue); violin plot shows γH2AX foci per cell (control, n = 125; HOTAIRM1 OE, n = 129) from 3 independent experiments. Center lines indicate median; dashed lines indicate quartiles. Scale bar, 20 μm. f, RT–qPCR quantification of HOTAIRM1 abundance in RPE-1 (left) and HEK293 (right) cells transfected with empty vector (Control) or HOTAIRM1 overexpression plasmid (OE). g, Schematic representation of the I-SceI-based DSB repair reporter system, showing EJ5-GFP for NHEJ, where successful repair reconstructs the GFP coding sequence, restoring GFP expression, and DR-GFP for HR, where homologous recombination results in GFP+ cells. h, Bar plot showing NHEJ repair efficiency following HOTAIRM1, p53 and BANF1 KD. i. Bar plot quantifying HR repair efficiency following HOTAIRM1, p53 and BANF1 KD. j, Graph depicting intrinsic disorder scores for BANF1 and p53 (PONDR VSL2). k, Representative images of in vitro droplet formation observed with recombinant BANF1-ECFP and Cy5-labeled truncated p53. Scale bar: 5 μm. l, Representative images of in vitro HOTAIRM1 condensates and a violin plot of droplet size at different salt concentration (150 mM, n = 42 droplets; 300 mM, n = 42 droplets) from 3 independent experiments. Scale bar: 2 μm. m, FRAP analysis of BANF1-ECFP droplets. Recovery curves indicate the dynamic behavior of droplets (n = 10). Scale bar: 0.5 μm. n, Bar plot showing FCS F-cross values, reflecting molecular interactions within fusion droplets. o, Predicted secondary structure of HOTAIRM1 (centroid model) using RNA-fold. Mountain plot representation showing cumulative base-pairing count across the transcript (blue line). Red arrows mark approximate boundaries (nt ~120 and ~400) corresponding to structural transitions. p, Densitometric quantification of pulldown signals (integrated band density, a.u.). For violin plots in e and l, the center line indicates the median; dotted lines indicate the 25th and 75th percentiles; the violin is truncated to the data range (min–max). Representative confocal imaging in e, k, l and m are from three independent experiments with similar results. P values in a, d, e-f, h-i and l were calculated by two-tailed Student’s t-test. Data in b–d, f, h-i and n are presented as the mean±s.e.m. from n = 3 independent experiments.
Extended Data Fig. 10 HOTAIRM1 expression, cellular proliferation, and lung pathology following PtbAlnc perturbation.
a, Representative Ki67 IHC staining images showing cellular proliferation ratios following LINC01638 and MIR155HG KD (LINC01638 KD, MIR155HG KD: n = 6; control: n = 12). Ki67-positive cells were quantified across 20 HPFs per sample (HPF = 338 μm × 338 μm). Statistical significance was determined using a two-tailed Student’s t-test. Scale bar: 50 μm. b, Representative H&E staining images illustrating lung structural integrity and pathological changes upon LINC01638 and MIR155HG KD. Violin plots quantify inflammation region counts and lung damage scores in mice based on H&E staining (LINC01638 KD, MIR155HG KD: n = 6; control: n = 12). For inflammation region analysis, one region (200 μm × 200 μm) was selected per sample. Lung damage scoring followed the American Thoracic Society’s Lung Injury Score (2010, Method), with 10 fields selected per sample. Statistical significance was determined using a two-tailed Student’s t-test. Scale bar: 500 μm. c, Representative Sirius Red staining images showing lung fibrosis following LINC01638 and MIR155HG KD. Violin plots quantify fibrosis ratio within vascular areas (LINC01638 KD, MIR155HG KD: n = 6, control: n = 12). Fibrosis analysis was based on 15 fields per sample (HPF = 498 μm × 498 μm). Statistical significance was determined using a two-tailed Student’s t-test. Scale bar: 500 μm. d, Bar plot quantifying endogenous HOTAIRM1 expression levels in the cohort of experimental mice (n = 3). Data are presented as the mean±s.e.m. **P < 0.01, by two-tailed Student’s t-test. Representative imaging in a-c is from three independent experiments for OE and six independent experiments for control with similar results. For violin plots in a-c, the center line indicates the median; dotted lines indicate the 25th and 75th percentiles; the violin is truncated to the data range (min–max).
Supplementary information
Supplementary Table 1 (download XLSX )
This table lists DNA repair-related genes whose TSSs show overlap between HOTAIRM1 CHIRP-seq peaks and BANF1 ChIP-seq binding. DNA repair annotation was based on KEGG pathways: Base excision repair (hsa03410), Nucleotide excision repair (hsa03420), Mismatch repair (hsa03430), Homologous recombination (hsa03440), and Non-homologous end-joining (hsa03450).
Supplementary Table 2 (download XLSX )
This table provides the sequences of the sgRNAs used for CRISPRi knockdown in this study. For each target lncRNA, three independent sgRNAs (sg1–sg3) were designed and cloned into the CRISPRi vector backbone used for pooled screening/perturbation experiments. Two non-targeting sgRNAs (NT1 and NT2) are included as negative controls.
Supplementary Table 3 (download XLSX )
This table lists, for each PtbAlnc perturbation vector, the corresponding target identity (gene/ID) and its associated GBC sequence used for sample/perturbation demultiplexing. The table also provides the primer sequences used for the two-round dial-out PCR amplification during GBC library construction.
Supplementary Table 4 (download XLSX )
This table provides the sequences of all qPCR primers used in this study, including gene-specific primer pairs (sense and antisense) for expression validation and related quantitative assays.
Supplementary Table 5 (download XLSX )
This table lists the sequences of all siRNAs used in this study, including multiple independent siRNAs targeting each indicated gene, as well as the corresponding sense and antisense strands for knockdown experiments.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots and gel.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, S., Ying, S., Cai, D. et al. Multiomic single-cell perturbation screens reveal critical lncRNA regulators of senescence. Nat Aging (2026). https://doi.org/10.1038/s43587-026-01100-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43587-026-01100-7