Abstract
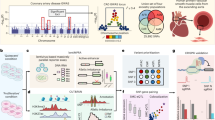
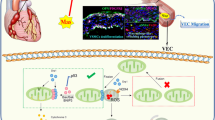
Mitophagy is a major quality control pathway that removes unwanted or dysfunctional mitochondria and plays an essential role in vascular health. Here we show that MCM8 expression is significantly decreased in children with Kawasaki disease (KD) who developed coronary artery aneurysms. Mechanistically, we discovered that nitric oxide signaling promotes TRIM21-mediated MCM8 ubiquitination, which disrupts its interaction with MCM9 and promotes its cytosolic export. In the cytosol, MCM8 relocates to the mitochondria pore-forming proteins and promotes their ubiquitination by TRIM21. In addition, MCM8 directly recruits LC3 via its LC3-interacting region (LIR) motif and initiates mitophagy. This suppresses mitochondrial DNA-mediated activation of type I interferon via cGAS and STING. Mice that are deficient in Mcm8, Trim21 and Nos2 or reconstituted with the East-Asian-specific MCM8-P276 variant develop more severe coronary artery vasculopathy in the Lactobacillus casei extract-induced KD model. Collectively, the data suggest that MCM8 protects vascular health in the KD setting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Protein structures were downloaded from the Protein Data Bank (PDB), including the N-terminal domain of MCM8 (PDB: 7DP3), the N-terminal domain of MCM hexamer from Pyrococcus furiosus (PDB: 4POF) and the N-terminal domain of MCM9 (PDB: 7DPD) were used to reconstruct the MCM8-MCM9 hexamer ring structure. Bulk RNA sequencing data from WT and Mcm8−/− mice are deposited at the National Center for Biotechnology Information (accession number GSE235994), and the count matrix is available in Supplementary Table 1. Source data used for statistical analysis are available as Supplementary Table 3. Uncropped immunoblot files from the same or different exposure time of the same gel are available as supplementary information.
References
Meier, T. et al. Cardiovascular mortality attributable to dietary risk factors in 51 countries in the WHO European Region from 1990 to 2016: a systematic analysis of the Global Burden of Disease Study. Eur. J. Epidemiol. 34, 37–55 (2019).
Ungvari, Z., Tarantini, S., Donato, A. J., Galvan, V. & Csiszar, A. Mechanisms of vascular aging. Circ. Res. 123, 849–867 (2018).
Forstermann, U. Nitric oxide and oxidative stress in vascular disease. Pflugers Arch. 459, 923–939 (2010).
Csiszar, A. et al. Aging-induced phenotypic changes and oxidative stress impair coronary arteriolar function. Circ. Res. 90, 1159–1166 (2002).
Farah, C., Michel, L. Y. M. & Balligand, J. L. Nitric oxide signalling in cardiovascular health and disease. Nat. Rev. Cardiol. 15, 292–316 (2018).
Riley, J. S. & Tait, S. W. Mitochondrial DNA in inflammation and immunity. EMBO Rep. 21, e49799 (2020).
Luo, W. et al. Critical role of cytosolic DNA and its sensing adaptor STING in aortic degeneration, dissection, and rupture. Circulation 141, 42–66 (2020).
Oduro, P. K. et al. The cGAS-STING signaling in cardiovascular and metabolic diseases: future novel target option for pharmacotherapy. Acta Pharm. Sin. B 12, 50–75 (2022).
Diao, R. Y. & Gustafsson, A. B. Mitochondrial quality surveillance: mitophagy in cardiovascular health and disease. Am. J. Physiol. Cell Physiol. 322, C218–C230 (2022).
Eiyama, A. & Okamoto, K. PINK1/Parkin-mediated mitophagy in mammalian cells. Curr. Opin. Cell Biol. 33, 95–101 (2015).
Novak, I. et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 11, 45–51 (2010).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Tong, M., Zablocki, D. & Sadoshima, J. The role of Drp1 in mitophagy and cell death in the heart. J. Mol. Cell. Cardiol. 142, 138–145 (2020).
Shulman, S. T. & Rowley, A. H. Kawasaki disease: insights into pathogenesis and approaches to treatment. Nat. Rev. Rheumatol. 11, 475–482 (2015).
Uehara, R. & Belay, E. D. Epidemiology of Kawasaki disease in Asia, Europe, and the United States. J. Epidemiol. 22, 79–85 (2012).
AlAsiri, S. et al. Exome sequencing reveals MCM8 mutation underlies ovarian failure and chromosomal instability. J. Clin. Invest. 125, 258–262 (2015).
Golubicki, M. et al. Germline biallelic Mcm8 variants are associated with early-onset Lynch-like syndrome. JCI Insight 5, e140698 (2020).
Lutzmann, M. et al. MCM8- and MCM9-deficient mice reveal gametogenesis defects and genome instability due to impaired homologous recombination. Mol. Cell 47, 523–534 (2012).
Lehman, T. J., Walker, S. M., Mahnovski, V. & McCurdy, D. Coronary arteritis in mice following the systemic injection of group B Lactobacillus casei cell walls in aqueous suspension. Arthritis Rheum. 28, 652–659 (1985).
Ibrahim, N. E. & Januzzi, J. L. Jr. Established and emerging roles of biomarkers in heart failure. Circ. Res. 123, 614–629 (2018).
Griffin, W. C. & Trakselis, M. A. The MCM8/9 complex: a recent recruit to the roster of helicases involved in genome maintenance. DNA Repair (Amst.) 76, 1–10 (2019).
Nelson, I., Hanna, M. G., Wood, N. W. & Harding, A. E. Depletion of mitochondrial DNA by ddC in untransformed human cell lines. Somat. Cell Mol. Genet. 23, 287–290 (1997).
Nicolli, A., Basso, E., Petronilli, V., Wenger, R. M. & Bernardi, P. Interactions of cyclophilin with the mitochondrial inner membrane and regulation of the permeability transition pore, and cyclosporin A-sensitive channel. J. Biol. Chem. 271, 2185–2192 (1996).
Kim, J. et al. VDAC oligomers form mitochondrial pores to release mtDNA fragments and promote lupus-like disease. Science 366, 1531–1536 (2019).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 359, eaao6047 (2018).
White, M. J. et al. Apoptotic caspases suppress mtDNA-induced STING-mediated type I IFN production. Cell 159, 1549–1562 (2014).
Minamikawa, T., Williams, D. A., Bowser, D. N. & Nagley, P. Mitochondrial permeability transition and swelling can occur reversibly without inducing cell death in intact human cells. Exp. Cell. Res. 246, 26–37 (1999).
Soutar, M. P. M. et al. FBS/BSA media concentration determines CCCP’s ability to depolarize mitochondria and activate PINK1-PRKN mitophagy. Autophagy 15, 2002–2011 (2019).
LeBleu, V. S. et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 992–1003 (2014).
Sun, N. et al. A fluorescence-based imaging method to measure in vitro and in vivo mitophagy using mt-Keima. Nat. Protoc. 12, 1576–1587 (2017).
Kalvari, I. et al. iLIR: a web resource for prediction of Atg8-family interacting proteins. Autophagy 10, 913–925 (2014).
Li, J. et al. Structural study of the N-terminal domain of human MCM8/9 complex. Structure 29, 1171–1181 (2021).
Maiorano, D., Cuvier, O., Danis, E. & Mechali, M. MCM8 is an MCM2-7-related protein that functions as a DNA helicase during replication elongation and not initiation. Cell 120, 315–328 (2005).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif—crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Li, T. & Chen, Z. J. The cGAS-cGAMP-STING pathway connects DNA damage to inflammation, senescence, and cancer. J. Exp. Med. 215, 1287–1299 (2018).
Lee, K. Y. et al. MCM8-9 complex promotes resection of double-strand break ends by MRE11-RAD50-NBS1 complex. Nat. Commun. 6, 7744 (2015).
Ragu, S., Matos-Rodrigues, G. & Lopez, B. S. Replication stress, DNA damage, inflammatory cytokines and innate immune response. Genes (Basel) 11, 409 (2020).
Wolf, C. et al. RPA and Rad51 constitute a cell intrinsic mechanism to protect the cytosol from self DNA. Nat. Commun. 7, 11752 (2016).
Coquel, F. et al. SAMHD1 acts at stalled replication forks to prevent interferon induction. Nature 557, 57–61 (2018).
Yang, Y. G., Lindahl, T. & Barnes, D. E. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 131, 873–886 (2007).
Kondo, T. et al. DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking. Proc. Natl Acad. Sci. USA 110, 2969–2974 (2013).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Jiang, H. et al. Chromatin-bound cGAS is an inhibitor of DNA repair and hence accelerates genome destabilization and cell death. EMBO J. 38, e102718 (2019).
Volkman, H. E., Cambier, S., Gray, E. E. & Stetson, D. B. Tight nuclear tethering of cGAS is essential for preventing autoreactivity. eLife 8, e47491 (2019).
Chen, H. et al. cGAS suppresses genomic instability as a decelerator of replication forks. Sci. Adv. 6, eabb8941 (2020).
Michalski, S. et al. Structural basis for sequestration and autoinhibition of cGAS by chromatin. Nature 587, 678–682 (2020).
Kujirai, T. et al. Structural basis for the inhibition of cGAS by nucleosomes. Science 370, 455–458 (2020).
Paull, T. T. Mechanisms of ATM activation. Annu. Rev. Biochem. 84, 711–738 (2015).
Kudo, N. et al. Leptomycin B inactivates CRM1/exportin 1 by covalent modification at a cysteine residue in the central conserved region. Proc. Natl Acad. Sci. USA 96, 9112–9117 (1999).
Hirayama, S. et al. Nuclear export of ubiquitinated proteins via the UBIN-POST system. Proc. Natl Acad. Sci. USA 115, E4199–E4208 (2018).
Hyer, M. L. et al. A small-molecule inhibitor of the ubiquitin activating enzyme for cancer treatment. Nat. Med. 24, 186–193 (2018).
Hendriks, I. A. et al. Site-specific mapping of the human SUMO proteome reveals co-modification with phosphorylation. Nat. Struct. Mol. Biol. 24, 325–336 (2017).
Espinosa, A. et al. The autoantigen Ro52 is an E3 ligase resident in the cytoplasm but enters the nucleus upon cellular exposure to nitric oxide. Exp. Cell. Res. 314, 3605–3613 (2008).
Lundberg, J. O. & Weitzberg, E. Nitric oxide signaling in health and disease. Cell 185, 2853–2878 (2022).
Nishimura, K. et al. Mcm8 and Mcm9 form a complex that functions in homologous recombination repair induced by DNA interstrand crosslinks. Mol. Cell 47, 511–522 (2012).
McCrindle, B. W. et al. Diagnosis, treatment, and long-term management of Kawasaki disease: a scientific statement for health professionals from the American Heart Association. Circulation 135, e927–e999 (2017).
Eriksson, U. et al. Lethal autoimmune myocarditis in interferon-γ receptor-deficient mice: enhanced disease severity by impaired inducible nitric oxide synthase induction. Circulation 103, 18–21 (2001).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
This study is supported by the National Natural Science Foundation of China (82125015, 92042303, 81901665 and 32100735), the Guangdong Province Key Project (2018B030335001), the Guangzhou City Key Project (20200703002 and 202206010002), the Guangzhou Women and Children’s Medical Center Fund and the Chongqing International Institute for Immunology (2021YJC02).
Author information
Authors and Affiliations
Contributions
Yuxia Zhang, M. Lin, X.G. and Yuzhang Wu conceived the ideas. M. Lin conducted all experiments, with assistance from H.X., S.W., M. Liu, W.L., Q.T., Y.L., W.H., C.G., E.I., R.F., M.A., J.C. and Z.H. Z.H., X.Z. and Yan Zhang performed bioinformatic analysis. D.C. and X.G. organized patient samples. Yuxia Zhang supervised the project and wrote the manuscript, with input from A.M.L., J.Z., Y.T., L.L., K.Z., F.B., Yan Wu, S.J., J.C. and Yuzhang Wu. All authors discussed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Yuxia Zhang, M. Lin, H.X., Z.H., F.B., J.C. and X.G. declare application of patents: use of MCM8-cGAS-STING-I-type interferon signal pathway as disease target (CN202110959959.8, authorized; CN202310692614.x, submitted; PCT/CN2021/113811, submitted; US18022505, submitted; EP21857767.4, submitted), by the Guangzhou Women and Children’s Medical Center. Patent contents include use of MCM8 or its fragments to enhance MCM8-mediated mitophagy for therapeutic purposes in diseases.
Peer review
Peer review information
Nature Cardiovascular Research thanks Michael Trakselis, A. Phillip West and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 MCM8 deficiency promotes IFN-I signaling and cardio vasculopathy.
a, CRISPR/Cas9 strategy used to generate Mcm8−/− mouse line. b, Gel electrophoresis showing representative genotyping data for WT and Mcm8−/− mice. c, Bar plots showing expressions of Nppa and Nppb from heart tissues of WT (PBS, n = 12 animals; LCWE, n = 12 animals; LCWE+anti-IFNAR, n = 8 animals), Mcm8+/− (PBS, n = 10 animals; LCWE, n = 8 animals; LCWE+anti-IFNAR, n = 6 animals) or Mcm8−/− (PBS, n = 6 animals; LCWE, n = 8 animals; LCWE+IFNAR, n = 8 animals) mice under indicated conditions. d, Bar plots comparing infiltration of neutrophils and macrophages from heart tissues of WT (PBS, n = 4 animals; LCWE, n = 5 animals; LCWE+anti-IFNAR, n = 4 animals), Mcm8+/− (PBS, n = 4 animals; LCWE, n = 5 animals; LCWE+anti-IFNAR, n = 4 animals) or Mcm8−/− (PBS, n = 3 animals; LCWE, n = 5 animals; LCWE+anti-IFNAR, n = 4 animals) mice under indicated conditions. e, CRISPR/Cas9 strategy used to generate Mcm8f/f mouse line. f, Representative H&E sections of heart and coronary artery tissues from Mcm8f/f, Mcm8f/f Cdh5Cre or Mcm8f/f Lyz2Cre mice under PBS or LCWE-treatment (n = 5 animals). For violin plots, middle line indicates median. g, Bar plots comparing infiltration of neutrophils and macrophages from heart tissues of Mcm8f/f (PBS, n = 8 animals; LCWE, n = 10 animals), Mcm8f/fCdh5Cre (PBS, n = 5 animals; LCWE, n = 8 animals) or Mcm8f/f Lyz2Cre (PBS, n = 8 animals; LCWE, n = 10 animals) mice under indicated conditions. h, Bar plots showing plasma cytokine concentrations at day 5 from Mcm8f/f (PBS, n = 8 animals; LCWE, n = 10 animals), Mcm8f/f Cdh5Cre (PBS, n = 5 animals; LCWE, n = 8 animals) or Mcm8f/f Lyz2Cre (PBS, n = 8 animals; LCWE, n = 10 animals) mice under PBS or LCWE treatment. i, Representative staining for Ifna4 and CD31 on heart tissues from Mcm8f/f (PBS, n = 8 animals; LCWE, n = 10 animals), Mcm8f/fCdh5Cre (PBS, n = 5 animals; LCWE, n = 8 animals) or Mcm8f/fLyz2Cre (PBS, n = 8 animals; LCWE, n = 10 animals) mice under PBS or LCWE treated conditions. Data are mean ± s.d. For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. P values were calculated by two-tailed Student’s t test for all experiments.
Extended Data Fig. 2 MCM8 deficiency promotes mtDNA accumulation and type I IFN production.
a, Bar plots showing MCM8 expression and IFN-β concentration, and immunoblot showing MCM8 knockdown efficiency in THP-1 cells. MCM8-1-siRNA was used in following experiments as MCM8-siRNA (n = 3 independent samples). b, Dot plots showing the expression of IFNB from ctrl- or MCM8-siRNA transfected THP-1 cells under indicated conditions (n = 3 independent samples). c, Immunoblot showing phosphorylation of IRF3 (pIRF3) in ctrl- or MCM8-siRNA transfected THP-1 cells under SeV stimulation at indicated time. d, Immunoblot showing depletion efficiency of cGAS, STING, IFI16 or DDX58 protein from WT-, CGAS-, STING-, IFI16- or DDX58-knockout THP-1 cells. e, Dot plots comparing the expressions of ISG15 in ctrl- and MCM8-siRNA transfected WT-, CGAS-, STING-, IFI16- or DDX58-knockout THP-1 cells, under indicated conditions. Fold change is relative to ctrl-siRNA transfected THP-1 cells under unstimulated conditions (n = 3 independent samples). f, Dot plots comparing relative abundance of DLOOP and TERT in the cytosol of ctrl- or MCM8-siRNA transfected HUVEC cells (n = 3 independent samples). g, Dot plots showing the expression of IFNB from ctrl- or MCM8-siRNA transfected HUVEC cells with or without ddC treatment under indicated conditions (n = 3 independent samples). h, Dot plots showing the expression of Ifnb from WT or Mcm8−/− MEF cells with or without ddC treatment under indicated conditions (n = 3 independent samples). i, Dot plots showing the expression of ISG15, ISG54 or ISG56 from ctrl- or MCM8-siRNA transfected THP-1 cells with or without ddC treatment under indicated conditions (n = 3 independent samples). j, k, Dot plots showing the expression of IFNB from ctrl- or MCM8-siRNA transfected THP-1 cells with CsA (j) or VBIT4 (k) treatment under indicated conditions (n = 3 independent samples). Data are mean ± s.d. P values were calculated by two-tailed Student’s t test for all experiments.
Extended Data Fig. 3 Apoptotic Caspases suppress IRF3-IFN activation in MCM8 deficient cells.
a, Flow cytometry showing annexin V and PI staining of ctrl- or MCM8-siRNA transfected THP-1 cells. b, Immunoblot showing Caspase-3 activation in ctrl- or MCM8-siRNA transfected THP-1 cells under indicated conditions. c, d, ELISA showing IFN-β concentrations (n = 3 independent samples) (c) and immunoblot showing IRF3 phosphorylation (d) from ctrl- or MCM8-siRNA transfected THP-1 cells under indicated conditions. e, Flow cytometry (top) and quantitative analysis (bottom) comparing CCCP-depolarized mitochondria in ctrl- or MCM8-siRNA transfected THP-1 cells (n = 3 independent samples). Data are mean ± s.d. P values were calculated by two-tailed Student’s t test for all experiments.
Extended Data Fig. 4 MCM8 mediates a mitophagy pathway that is independent of PINK1, PRKN, NIX, FUNDC1 and DRP1.
a, b, Immunoblot showing LC3B on isolated mitochondria (mitoLC3), as well as MCM8 and other mitochondrial proteins including TOM20, PINK1 and COXIV from the whole cell lysate (WCL) of empty vector (EV) or Flag-MCM8 transfected HeLa cells (a) or ctrl- or MCM8-siRNA transfected THP-1 cells (b). c, Flow cytometry showing mitophagic flux in ctrl- or MCM8- siRNA transfected mt-mKeima-HeLa cells. d, e, Dot plots showing the expression of IFNA (left) and immunoblot showing pIRF3 (right) in ctrl- or MCM8-siRNA transfected PINK1-deficient (d) or PRKN-deficient (e) THP-1 cells (n = 3 independent samples). f, g, h, Immunoblot showing IRF3 phosphorylation in ctrl- or MCM8-siRNA transfected NIX-deficient (f), FUNDC1-deficient (g) or DNM1L (DRP1)-deficient (h) THP-1 cells. i, Flow cytometry showing mitophagic flux in EV or MCM8 transfected PINK1-deficient, NIX-deficient, FUNDC1-deficient or DNM1L-deficient mt-mKeima-HeLa cells under indicated conditions. Data are mean ± s.d. P values were calculated by two-tailed Student’s t test for all experiments.
Extended Data Fig. 5 MCM8 interacts with LC3 and functions as a mitophagy receptor.
a, Significantly enriched gene ontology terms of MCM8 co-purified proteins identified by mass spectrometry analysis. b, Immunoblot showing endogenous interaction of MCM8 and LC3B under indicated conditions in THP-1 cells. c, Immunofluorescence showing localization of MCM8, LC3B and MitoTracker Deep Red under indicated conditions in HeLa cells (n = 7 independent images). For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. d, The putative LIR motif from MCM8 and comparison with other MCM family members. e, A structural remodeling showing binding of MCM8 LIR motif with LC3B. F274 and L277 from MCM8-LIR are respectively inserted into the hydrophobic pocket 1 (formed by I23, F108 and L53) and hydrophobic pocket 2 (formed by I35, V54, L63, I66 and I67) of LC3B. Electrostatic interactions are formed between R272 of MCM8-LIR and D19 of LC3. f, Immunoblot showing mitoLC3, as well as γH2AX from the whole cell lysate (WCL) of EV, MCM8 (FL) or MCM8 (LIR) transfected Mcm8−/− MEF cells. g, Dot plots showing relative abundance of DLOOP in the cytosol of EV, MCM8 (FL), MCM8 (LIR) or MCM8 (dLIR) transfected Mcm8−/− MEF cells (n = 3 independent samples). h, i, Immunoprecipitation and immunoblot showing the interaction of MCM8 and LC3 in ZnCl2 (g) or EDTA (h) treated THP-1 cells under indicated condition. Data are mean ± s.d. P values were calculated by two-tailed Student’s t test.
Extended Data Fig. 6 MCM9 is not required for MCM8-mediated mitophagy.
a, Immunofluorescence showing the localization of MCM9 and MCM8 under indicated conditions in HeLa cells (n = 7 independent images). For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. b, Immunoblot showing nuclear and cytosolic MCM9 in THP-1 cells under indicated conditions. Data are mean ± s.d. P values were calculated by two-tailed Student’s t test.
Extended Data Fig. 7 TRIM21 regulates MCM8 ubiquitination and cytosolic export.
a, Immunoblot showing nuclear and cytosolic MCM8 when treated with Exportin1 inhibitor Leptomycin B (10 nM) under indicated conditions. b, Immunoblot showing mitoLC3B, as well as MCM8 and other mitochondrial proteins (TOM20, MFN2, PINK1) in THP-1 cells treated with leptomycin B under indicated conditions. c, Immunoblot showing nuclear and cytosolic MCM8 when treated with the ubiquitin activating enzyme inhibitor TAK-243 (1 μM) under indicated conditions. d, Immunoprecipitation and immunoblot showing interaction of MCM8 with LC3 in Flag-MCM8 transfected HeLa cells when treated with TAK-243 under indicated conditions. e, f, Immunoprecipitation and immunoblot showing interaction of HA-MCM8 with Flag-TRIM21 (e) or endogenous interaction of MCM8 with TRIM21 (f) in HeLa cells under indicated conditions. g, Immunoblot showing ubiquitination of MCM8 in ctrl- or TRIM21-siRNA transfected HeLa cells over-expressing Flag-MCM8 and HA-Ubiquitin under treatment with poly(dA:dT). h, i, Immunoblot showing endogenous ubiquitination of MCM8 in ctrl- or TRIM21-siRNA transfected HeLa (h) or THP-1 (i) cells. j, Bar plots showing the relative abundance of Dloop in the plasma from WT or Mcm8−/− mice under PBS (WT, n = 12 animals; Mcm8−/−, n = 9 animals) or LCWE (WT, n = 18 animals; treatment (WT, n = 18 animals; Mcm8−/−, n = 15 animals). Fold change is relative to PBS treated WT mice. k, Representative H&E images and coronary artery thickness of WT (PBS, n = 4 animals; LCWE, n = 6 animals) or Trim21−/− (PBS, n = 4 animals; LCWE, n = 6 animals) mouse heart sections under PBS or LCWE treatment. For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. l, Representative staining for Ifna4 and CD31 on heart tissues of WT (PBS, n = 4 animals; LCWE, n = 6 animals) and Trim21−/− (PBS, n = 4 animals; LCWE, n = 6 animals) mice under PBS or LCWE treated conditions. Data are mean ± s.d. P values were calculated by two-tailed Student’s t test.
Extended Data Fig. 8 TRIM21 mediates ubiquitination of MCM8 and the mitochondria pore proteins.
a, Immunofluorescence and statistical analysis showing distribution of TRIM21 and TOM20 under indicated conditions in HeLa cells (n = 6 independent images). For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. b, Immunoblot showing nuclear or cytosolic MCM8 in ctrl- or TRIM21- siRNA transfected THP-1 cells under indicated conditions. c. Immunoprecipitation and immunoblot showing interaction of MCM8 with LC3 in WT or TRIM21−/− THP-1 cells under indicated conditions. d, e, Immunoblot showing ubiquitination of VDAC1 (d) or BAK I in ctrl- or TRIM21-siRNA transfected THP-1 cells under indicated conditions. f, g, Immunoblot showing ubiquitination of VDAC1 (f) or BAK (g) in ctrl- or MCM8-siRNA transfected THP-1 cells under indicated conditions. h, Immunoblot showing ubiquitination of MCM8 (FL) or MCM8 (LIR) in EV or Myc-TRIM21 transfected HeLa cellI. i, Structural remodelling showing Lysine 208 and 232 at the interface of the MCM8-MCM9 hexamer ring structure. j, Dot plot comparing relative abundance of DLOOP and TERT in the cytosol of MCM8-WT, K208R, K232R, or K263R transfected HeLa cells under indicated conditions (n = 3 independent samples). Data are mean ± s.d. P values were calculated by two-tailed Student’s t test.
Extended Data Fig. 9 Nitric oxide induces MCM8 mediated mitophagy.
a, Bar plot showing nitic oxide concentrations in THP-1 cells with or without NO scavenger 1400W treatment under indicated conditions (n = 3 independent samples). b, Immunoprecipitation and immunoblot showing ubiquitination of Flag-MCM8 in EV or myc-TRIM21 transfected HeLa cells with or without 1400W treatment. c, Immunoblot showing nuclear and cytosolic MCM8 in THP-1 cells with or without 1400W treatment under indicated conditions. d, Immunofluorescence and statistical analysis showing MCM8 distribution in HeLa cells with or without 1400W treatment under indicated conditions (n = 7 independent images). For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. e, Representative H&E sections of WT or Nos2−/− mice heart and coronary artery tissues under PBS or LCWE treatment (n = 6 animals). f, Bar plots showing inflammatory cytokine concentrations in the plasma of WT or Nos2−/− mice at day 5 under PBS (WT, n = 10 animals; Nos2−/−, n = 6 animals) or LCWE (WT, n = 7 animals; Nos2−/−, n = 7 animals) treatment. Data are mean ± s.d. P values were calculated by two-tailed Student’s t test for all experiments.
Extended Data Fig. 10 MCM8-P276 promotes type I IFN production and cardio vasculopathy in mice.
a, Schematic graph displaying generation of MCM8-A276 and MCM8-P276 chimeric mice. b, Schematic graph illustrating the generation of KD model using MCM8-A276 and MCM8-P276 chimeric mice. c, Dot plots showing cytokine concentrations at day 14 of MCM8-A276 (PBS, n = 2 animals; LCWE, n = 5 animals) and MCM8-P276 chimeric (PBS, n = 2 animals; LCWE, n = 5 animals) mice, P values were calculated by one-tailed Student’s t test. d, Bar plots showing the relative abundance of Dloop and Tert in the plasma from MCM8-A276 or MCM8-P276 chimeric mice under PBS or LCWE treatment. Fold change is relative to PBS treated MCM8-A276 mice (n = 8 animals), P values were calculated by two-tailed Student’s t test. e, Flow cytometry (left) and quantitative analysis (right) showing neutrophil infiltration in heart tissues of MCM8-A276 (PBS, n = 2 animals; LCWE, n = 5 animals) and MCM8-P276 (PBS, n = 2 animals; LCWE, n = 5 animals) chimeric mice under indicated conditions, P values were calculated by one-tailed Student’s t test. f, Representative H&E sections showing cardiovascular pathology and inflammatory cell infiltration in MCM8-A276 and MCM8-P276 mice at day 14 after LCWE administration. g, Representative staining for Ifna4 on heart tissues of MCM8-A276 or MCM8-P276 mice under PBS or LCWE treatment (n = 7 independent images). For box plots, the box extends from the Q1 to Q3 quartile values of the data, with a line at the median. The whiskers extend from minimum to maximum value of data. P values were calculated by two-tailed Student’s t test.
Supplementary information
Supplementary Table 1 (download TXT )
Combined count matrix for WT and Mcm8−/− mice
Supplementary Table 2 (download XLSX )
MCM8 binding proteins identified by LC–MS
Supplementary Table 3 (download XLSX )
Source data for statistical analysis
Source data
Source Data Fig. 1 (download PDF )
Uncropped scans of blots representative findings in Fig. 2
Source Data Fig. 2 (download PDF )
Uncropped scans of blots representative findings in Fig. 3
Source Data Fig. 3 (download PDF )
Uncropped scans of blots representative findings in Fig. 4
Source Data Fig. 4 (download PDF )
Uncropped scans of blots representative findings in Fig. 5
Source Data Fig. 5 (download PDF )
Uncropped scans of blots representative findings in Fig. 6
Source Data Extended Data Fig. 1 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 2
Source Data Extended Data Fig. 2 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 3
Source Data Extended Data Fig. 3 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 4
Source Data Extended Data Fig. 4 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 5
Source Data Extended Data Fig. 5 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 6
Source Data Extended Data Fig. 6 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 7
Source Data Extended Data Fig. 7 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 8
Source Data Extended Data Fig. 8 (download PDF )
Uncropped scans of blots representative findings in Extended Data Fig. 9
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, M., Xian, H., Chen, Z. et al. MCM8-mediated mitophagy protects vascular health in response to nitric oxide signaling in a mouse model of Kawasaki disease. Nat Cardiovasc Res 2, 778–792 (2023). https://doi.org/10.1038/s44161-023-00314-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-023-00314-x