Abstract
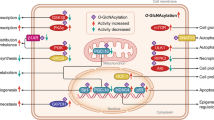
The restoration of the microvasculature is essential to cardiovascular regeneration. Our previous work demonstrated that angiogenic transdifferentiation of fibroblasts into endothelial cells facilitates vascular recovery following limb ischemia and is accompanied by a metabolic shift toward glycolysis. However, a comprehensive characterization of the metabolic alterations that contribute to the transdifferentiation process is still lacking. Here we identify a marked upregulation of uridine diphosphate N-acetylglucosamine (UDP-GlcNAc), the substrate for O-GlcNAcylation, during transdifferentiation. Enhancing this pathway promotes, whereas inhibiting it impairs, the efficiency of transdifferentiation. Mechanistically, we demonstrate that O-GlcNAcylation facilitates chromatin remodeling through modification of HIRA, a histone chaperone responsible for de novo deposition of the noncanonical histone variant H3.3, a process intimately linked to transcriptional activation. These findings are further supported by in vivo lineage tracing and conditional knockout mouse models. Collectively, our study demonstrates that O-GlcNAcylation enhances angiogenic transdifferentiation through a metabolic-and-epigenetic-coupled mechanism, thereby strengthening vascular recovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data, analytical methods and study materials supporting the findings of this study are available within the article and its Supplementary Information. Source data are provided with this paper.
References
Mensah, G. A., Roth, G. A. & Fuster, V. The global burden of cardiovascular diseases and risk factors: 2020 and beyond. J. Am. Coll. Cardiol. 74, 2529–2532 (2019).
Teraa, M., Conte, M. S., Moll, F. L. & Verhaar, M. C. Critical limb ischemia: current trends and future directions. J. Am. Heart Assoc. 5, 26908409 (2016).
Annex, B. H. Therapeutic angiogenesis for critical limb ischaemia. Nat. Rev. Cardiol. 10, 387–396 (2013).
Graf, T. Historical origins of transdifferentiation and reprogramming. Cell Stem Cell 9, 504–516 (2011).
Sayed, N. et al. Transdifferentiation of human fibroblasts to endothelial cells: role of innate immunity. Circulation 131, 300–309 (2015).
Lee, J. et al. Activation of innate immunity is required for efficient nuclear reprogramming. Cell 151, 547–558 (2012).
Zhou, G., Meng, S., Li, Y., Ghebre, Y. T. & Cooke, J. P. Optimal ROS signaling is critical for nuclear reprogramming. Cell Rep. 15, 919–925 (2016).
Meng, S. et al. Transdifferentiation requires iNOS activation: role of RING1A S-nitrosylation. Circ. Res. 119, e129–e138 (2016).
Sayed, N. et al. Retinoic acid inducible gene 1 protein (RIG1)-like receptor pathway is required for efficient nuclear reprogramming. Stem Cells 35, 1197–1207 (2017).
Lai, L., Reineke, E., Hamilton, D. J. & Cooke, J. P. Glycolytic switch is required for transdifferentiation to endothelial lineage. Circulation 139, 119–133 (2019).
Chanda, P. K. et al. Nuclear S-nitrosylation defines an optimal zone for inducing pluripotency. Circulation 140, 1081–1099 (2019).
Meng, S. et al. Reservoir of fibroblasts promotes recovery from limb ischemia. Circulation 142, 1647–1662 (2020).
Dai, Z., Ramesh, V. & Locasale, J. W. The evolving metabolic landscape of chromatin biology and epigenetics. Nat. Rev. Genet. 21, 737–753 (2020).
Etchegaray, J.-P. & Mostoslavsky, R. Interplay between metabolism and epigenetics: a nuclear adaptation to environmental changes. Mol. Cell 62, 695–711 (2016).
Arnold, P. K. et al. A non-canonical tricarboxylic acid cycle underlies cellular identity. Nature 603, 477–481 (2022).
Kaelin, W. G. Jr. & McKnight, S. L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013).
Kimball, T. H. & Vondriska, T. M. Metabolism, epigenetics, and causal inference in heart failure. Trends Endocrinol. Metab. 31, 181–191 (2020).
Gleason, R. J. et al. Developmentally programmed histone H3 expression regulates cellular plasticity at the parental-to-early embryo transition. Sci. Adv. 9, eadh0411 (2023).
Probst, A. V., Desvoyes, B. & Gutierrez, C. Similar yet critically different: the distribution, dynamics and function of histone variants. J. Exp. Bot. 71, 5191–5204 (2020).
An, Y. J. et al. Lactate as a major epigenetic carbon source for histone acetylation via nuclear LDH metabolism. Exp. Mol. Med. 55, 2238–2247 (2023).
Ye, C. & Tu, B. P. Sink into the epigenome: histones as repositories that influence cellular metabolism. Trends Endocrinol. Metab. 29, 626–637 (2018).
Franklin, R., Murn, J. & Cheloufi, S. Cell fate decisions in the wake of histone H3 deposition. Front. Cell Dev. Biol. 9, 654915 (2021).
Ray-Gallet, D. et al. Functional activity of the H3.3 histone chaperone complex HIRA requires trimerization of the HIRA subunit. Nat. Commun. 9, 3103 (2018).
Typas, D. Histone H3 and its chaperones. Nat. Struct. Mol. Biol. 30, 405 (2023).
Armache, A. et al. Histone H3.3 phosphorylation amplifies stimulation-induced transcription. Nature 583, 852–857 (2020).
Esteves de Lima, J. et al. HIRA stabilizes skeletal muscle lineage identity. Nat. Commun. 12, 3450 (2021).
Hart, G. W., Slawson, C., Ramirez-Correa, G. & Lagerlof, O. Cross talk between O-GlcNAcylation and phosphorylation: roles in signaling, transcription, and chronic disease. Annu. Rev. Biochem. 80, 825–858 (2011).
Lu, P. et al. Cryo-EM structure of human O-GlcNAcylation enzyme pair OGT–OGA complex. Nat. Commun. 14, 6952 (2023).
Yang, X. & Qian, K. Protein O-GlcNAcylation: emerging mechanisms and functions. Nat. Rev. Mol. Cell Biol. 18, 452–465 (2017).
Chatham, J. C., Zhang, J. & Wende, A. R. Role of O-linked N-acetylglucosamine protein modification in cellular (patho)physiology. Physiol. Rev. 101, 427–493 (2021).
Tran, D. H. et al. Chronic activation of hexosamine biosynthesis in the heart triggers pathological cardiac remodeling. Nat. Commun. 11, 1771 (2020).
Zuurbier, C. J. et al. Cardiac metabolism as a driver and therapeutic target of myocardial infarction. J. Cell. Mol. Med. 24, 5937–5954 (2020).
Wang, Z. et al. Increasing O-GlcNAcylation is neuroprotective in young and aged brains after ischemic stroke. Exp. Neurol. 339, 113646 (2021).
Wu, Q. P. et al. O-GlcNAc regulates anti-fibrotic genes in lung fibroblasts through EZH2. J. Cell. Mol. Med. 28, e18191 (2024).
Martin, S. E. S. et al. Structure-based evolution of low nanomolar O-GlcNAc transferase inhibitors. J. Am. Chem. Soc. 140, 13542–13545 (2018).
Coffey, G. L. et al. 6-diazo-5-oxo-l-norleucine, a new tumor-inhibitory substance. I. Biologic studies. Antibiot. Chemother. 6, 487–497 (1956).
Yuzwa, S. A. et al. A potent mechanism-inspired O-GlcNAcase inhibitor that blocks phosphorylation of tau in vivo. Nat. Chem. Biol. 4, 483–490 (2008).
Tafessu, A. et al. H3.3 contributes to chromatin accessibility and transcription factor binding at promoter-proximal regulatory elements in embryonic stem cells. Genome Biol. 24, 25 (2023).
Lee, J.-S. & Zhang, Z. O-linked N-acetylglucosamine transferase (OGT) interacts with the histone chaperone HIRA complex and regulates nucleosome assembly and cellular senescence. Proc. Natl Acad. Sci. USA 113, E3213–E3220 (2016).
Torne, J., Orsi, G. A., Ray-Gallet, D. & Almouzni, G. Imaging newly synthesized and old histone variant dynamics dependent on chaperones using the SNAP-tag system. Methods Mol. Biol. 1832, 207–221 (2018).
Annex, B. H. & Cooke, J. P. New directions in therapeutic angiogenesis and arteriogenesis in peripheral arterial disease. Circ. Res. 128, 1944–1957 (2021).
He, L. et al. Preexisting endothelial cells mediate cardiac neovascularization after injury. J. Clin. Invest. 127, 2968–2981 (2017).
Li, I. M. H., Horwell, A. L., Chu, G., de Crombrugghe, B. & Bou-Gharios, G. Characterization of mesenchymal–fibroblast cells using the Col1a2 promoter/enhancer. In Fibrosis. Methods in Molecular Biology Vol. 1627 (ed. Rittié, L.) (Humana, 2017).
Olivier-Van Stichelen, S., Abramowitz, L. K. & Hanover, J. A. X marks the spot: does it matter that O-GlcNAc transferase is an X-linked gene? Biochem. Biophys. Res. Commun. 453, 201–207 (2014).
Xiong, J. et al. A metabolic basis for endothelial-to-mesenchymal transition. Mol. Cell 69, 689–698 (2018).
Dupas, T., Lauzier, B. & Mcgraw, S. O-GlcNAcylation: the sweet side of epigenetics. Epigenet. Chromatin 16, 49 (2023).
Fang, H. T. et al. Global H3.3 dynamic deposition defines its bimodal role in cell fate transition. Nat. Commun. 9, 1537 (2018).
Alhusban, S. et al. Glucosamine-mediated hexosamine biosynthesis pathway activation uses ATF4 to promote ‘exercise-like’ angiogenesis and perfusion recovery in PAD. Circulation 150, 1702–1719 (2024).
Kronlage, M. et al. O-GlcNAcylation of histone deacetylase 4 protects the diabetic heart from failure. Circulation 140, 580–594 (2019).
Ong, Q., Han, W. & Yang, X. O-GlcNAc as an integrator of signaling pathways. Front. Endocrinol. 9, 599 (2018).
Thompson, J. W., Griffin, M. E. & Hsieh-Wilson, L. C. Methods for the detection, study, and dynamic profiling of O-GlcNAc glycosylation. Methods Enzymol. 598, 101–135 (2018).
Li, S. et al. A relay velocity model infers cell-dependent RNA velocity. Nat. Biotechnol. 42, 99–108 (2024).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Lau, K. S. et al. Complex N-glycan number and degree of branching cooperate to regulate cell proliferation and differentiation. Cell 129, 123–134 (2007).
Fehl, C. & Hanover, J. A. Tools, tactics and objectives to interrogate cellular roles of O-GlcNAc in disease. Nat. Chem. Biol. 18, 8–17 (2022).
Zhu, Y. & Hart, G. W. Dual-specificity RNA aptamers enable manipulation of target-specific O-GlcNAcylation and unveil functions of O-GlcNAc on β-catenin. Cell 186, 428–445 (2023).
Swonger, J. M., Liu, J. S., Ivey, M. J. & Tallquist, M. D. Genetic tools for identifying and manipulating fibroblasts in the mouse. Differentiation 92, 66–83 (2016).
Keembiyehetty, C. et al. Conditional knock-out reveals a requirement for O-linked N-acetylglucosaminase (O-GlcNAcase) in metabolic homeostasis. J. Biol. Chem. 290, 7097–7113 (2015).
Xia, J. & Wishart, D. S. Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat. Protoc. 6, 743–760 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank the MD Anderson Metabolomics Core Facility (L. Tan and P. Lorenzi) for their guidance with LC–MS metabolomic experiments. The Metabolomics Core Facility is supported by National Institutes of Health grants S10OD012304-01 and P30CA016672. We also thank the HMRI RNA Core for their assistance with plasmid generation. Additionally, we thank the Houston Methodist Research Institute Flow Cytometry Core (D. Haviland and E. Jardinella) for their guidance with flow cytometry. Finally, we thank J. Hanover from the NIDDK of the National Institutes of Health for his generous sharing of Ogafl/fl mice. This work was supported by grants from National Institutes of Health R01s, HL169204-01A1 (L.L.), HL133254 (J.P.C.), HL1577990 (J.P.C.), HL148338 (J.P.C.) and GM151280 (J.L.), F30 HL167547 (A.J.L.) and AHA 25POST1378365 (S.L.) and the National Institute of General Medical Sciences of the National Institutes of Health (1R35GM150460 to G.W.) as well as the Cancer Prevention and Research Institute of Texas CPRIT RP150611 (J.P.C.) and CPRIT RP250213 (J.L.).
Author information
Authors and Affiliations
Contributions
L.L. and J.P.C. conceptualized the project and designed the experiments. L.L., S.L. and A.J.L. wrote the manuscript and prepared the figures. L.L. performed metabolomic experiments. Y.L., K.N.C. and G.W. performed the metabolomics analysis. L.L., E.S.N., S.L., A.J.L. and K.N.C. performed in vitro transdifferentiation and western experiments. L.L., S.L., A.J.L., K.N.C. and C.L.K. conducted the in vivo experiments. S.L., A.J.L. and L.L. performed flow cytometry experiments and data analysis. L.L., S.L., A.J.L., K.N.C. and E.M. performed confocal microscopy image and data analysis. A.J.L., M.G., S.L. and E.S.N. performed and analyzed quench–chase–pulse experiments. J.L. helped with plasmid generation. L.L. supervised all aspects of this work. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.L. and J.P.C. are inventors on a patent assigned to Houston Methodist Hospital on the use of modulators of O-GlcNAcylation for the treatment of vascular diseases (inventors J.P.C. (Houston, TX), L.L. (Houston, TX)/Houston Methodist Hospital (Houston, TX); publication number 20250114337; status, application pending; specific aspect of this study covered in the patent application: the invention is a therapeutic approach for modulating cellular transdifferentiation and potentially angiogenesis by modulating O-GlcNAcylation (GlcNAc) of proteins to treat various conditions. The inventors used existing small molecules to prove the concept in vitro and in vivo. The projected use includes (1) wound or fracture healing and treating vascularization (ischemic tissues, coronary, cerebral and peripheral arterial disease) by enhancing the O-GlcNAcylation pathway, potentially enhancing angiogenesis and (2) treating tumor neovascularization or excess vascularization (age-related macular degeneration, diabetic retinopathy, telangiectasias, AV malformations) by inhibiting the GlcNAcylation pathway, potentially inhibiting angiogenesis). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Hossein Ardehali, Brian Annex, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 O-GlcNAcylation regulates angiogenic transdifferentiation by H3.3 deposition.
a. Western blots confirm the overexpression of SNAP-tagging H3.3 in Hela cells. The fusion protein SNAP-H3.3 is detected by the SNAP antibody at 37 kD and SNAP itself at 25 kD. Western blot for H3.3 shows endogenous H3.3 at 15 kD and SNAP-H3.3 at 37 kD. b. Immunofluorescence imaging of positive (P = pulse), negative (QP = quench-pulse), and experimental (QCP = quench-chase-pulse) controls for the quench-chase-pulse experiments (Scale bar, 15 µm). c. Quantification of immunofluorescence from quench-chase-pulse controls. P: n = 126 cells; QP: n = 162 cells; QCP: n = 119 cells. P values are from Kruskal-Wallis test. d. Total protein visualization for indicated groups on gel using Stain-Free Technology. e. Confirmation of HIRA knockdown efficiency in BJ fibroblasts by western blot. f. Putative site-specific O-GlcNAcylation site mutations on HIRA, confirmed by Sanger sequencing. g. Verification of HIRA WT and HIRA mutant overexpression in BJ fibroblasts through Western blot using a Flag-tag antibody. In c, data are mean ± SD. In a, e, g, results are representative of two independent experiments. In d, results are representative of three independent experiments.
Extended Data Fig. 2 O-GlcNAcylation regulates angiogenic transdifferentiation.
a. Schematic figure illustrates lineage tracing strategy in mice for Fsp1-driven fibroblasts labeled with EYFP. b. Negative staining control of secondary antibody staining performed without primary antibody. c. Immunofluorescence staining of YFP (green), O-GlcNAc (gray), and DAPI (blue) in ischemic hindlimb tissue from Fsp1-Cre: R26R-EYFP mice at day 3, with or without (non-operated) femoral artery ligation. (scale bar: 50 µm, z-stack, [1.15 µm/slide]). d. Single-channel immunofluorescence staining shows DAPI (blue), YFP (green), and O-GlcNAc (gray) in the ischemic hindlimb on day 3 post-surgery and in limb from non-operated mice. In b-d, results are representative of two independent experiments.
Extended Data Fig. 3 O-GlcNAcylation plays a role in vascular recovery from ischemia.
a. Schematic figure showing the experimental timeline of hindlimb ischemia surgery, drug injection, laser Doppler perfusion imaging, and tissue processing. Created with BioRender.com. b. Representative plots and densitometry quantification of western blot for O-GlcNAc in hindlimb tissue of wild-type mice with indicated treatments (n = 3 mice per group). P values are from one-way ANOVA. c-d. Weights of treated and untreated mice at days 0, 3, 7, 14, and 21 post-surgery (n = 6 mice per group). e, g. Immunofluorescence staining of CD31 (green) and DAPI (blue) in ischemic hindlimb tissue of vehicle and OSMI-4 (e) or vehicle and TMG (g) treated mice (Scale bar, 50 µm). f, h. Bar graphs show quantification of relative CD31+ cells in the indicated treatment groups (compared to Vehicle) through immunofluorescence imaging (n = 6 mice per group). P values are from two-tailed unpaired Student’s t-test. In b–h, results represent two independent experiments. In b, c, d, f, h, data are mean ± SD.
Extended Data Fig. 4 Angiogenic transdifferentiation in vivo.
a. Immunofluorescence images show YFP + CD31+ double-positive cells in hindlimb muscle tissue of Fsp1-Cre: R26R-EYFP mice at day 3 post-surgery. (Red: CD31; Green: YFP; Blue: DAPI; Scale bar = 50 µm, 20 z-stacks, [0.65 µm/slice]). b. Single channel immunofluorescence images of DAPI, YFP, and CD31 from ischemic hindlimb on day 3 post-surgery at 40x magnification. c. Merged immunofluorescence images depict the co-localization of YFP with CD144 in ischemic hindlimb tissue of Fsp1-Cre: R26R-EYFP mice at days 3 post-surgery. The images were captured at 40x and 100x magnification (40x has 15 z-stacks, 1.02 µm/slice, 100x has 15 slices, 1.03 µm/slice). Scale bar = 50 µm. d. Single channel immunofluorescence images of DAPI, YFP, and CD144 in ischemic hindlimb on day 3 post-surgery at 40x magnification. In a–d, results represent two independent experiments.
Extended Data Fig. 5 Angiogenic transdifferentiation in vivo.
a. Merged immunofluorescence images show the co-localization of YFP (green) with CD31 (red) in ischemic hindlimb tissue of Fsp1-Cre: R26R-EYFP mice at day 7 post-surgery. Images were captured at 40x (21 z-stacks, 0.5 µm/slice) and 100x magnification (17 slices, 0.5 µm/slice). Scale bar = 50 µm. b. Single-channel immunofluorescence images of DAPI (blue), YFP (green), and CD31 (red) in ischemic hindlimb tissue at day 7 post-surgery, captured at 40x magnification. c. Immunofluorescence images illustrate YFP + CD144+ (double-positive) cells in hindlimb muscle tissue of Fsp1-Cre: R26R-EYFP mice at day 7 post-surgery. Images were captured at 40x (15 z-stacks, 0.45 µm/slice) and 100x magnification (15 slices, 0.39 µm/slice). Scale bar = 50 µm. d. Single-channel immunofluorescence images of DAPI (blue), YFP (green), and CD144 (red) in ischemic hindlimb tissue at day 7 post-surgery, captured at 40x magnification. In a–d, results represent two independent experiments.
Extended Data Fig. 6 Angiogenic transdifferentiation in vivo.
a. Merged immunofluorescence images show co-localization YFP (green) with ERG1 (red) in ischemic hindlimb tissue of Fsp1-Cre: R26R-EYFP mice at day 3 post-surgery. Images were captured at 40x (15 z-stacks, 1.07 µm/slice) and 100x magnification (15 slices, 1.02 µm/slice). Scale bar: 50 µm. b. Single-channel immunofluorescence images of DAPI (blue), YFP (green), and ERG1 (red) in ischemic hindlimb tissue at day 3 post-surgery, captured at 40x magnification. c. Representative plots and bar graphs of flow cytometry analysis display % of YFP+ cells in contralateral and ligation limbs on day 3 (n = 4 mice per group). and day 7 (n = 3 mice per group). Gating was performed on DAPI-CD11b- cells. Data are presented as mean ± SD. P values are from multiple unpaired Student’s t-test. d. The gating strategy for detecting the YFP+ cells among CD31+ cells from digested hindlimb tissue at day 7 post-surgery. e. Flow cytometry analysis demonstrates the presence of YFP + CD31+ cells predominantly in the ligated limb compared to the contralateral limb in Fsp1-Cre: R26R-EYFP mice at day 7 post-surgery. In a-e, results represent two independent experiments.
Extended Data Fig. 7 Angiogenic transdifferentiation in vivo.
a. Schematic illustration of the lineage tracing strategy in mice for Col1a2-driven fibroblasts labeled with tdTomato. b. Immunofluorescence images show co-localization of CD31(green) with Tdtomato (red) in ischemic hindlimb tissue of Col1a2-tdTomato mice at day 7 post-surgery. Images were captured at 40x (15 z-stacks, 0.68 µm/slice) and 100x magnification (15 slices, 0.64 µm/slice). Scale bar: 50um. c. Immunofluorescence images depict co-localization of CD144 (green) with Tdtomato (red) in ischemic hindlimb tissue of Col1a2-tdTomato mice at day 7 post-surgery. Images were captured at 40x (15 z-stacks, 0.72 µm/slice) and 100x magnification (15 slices, 0.78 µm/slice). Scale bar: 50 µm. d. Bar graphs of % tdTomato+ cells from CD31+cells in ischemic limbs at day 7 post-surgery (n = 3 mice per group), normalized to non-operated mice. All flow cytometry analysis is gated on live and non-immune cells (DAPI-CD11b-). Data are mean ± SD, P values are from two-tailed unpaired Student’s t-test. In b–d, results represent two independent experiments.
Extended Data Fig. 8 Angiogenic transdifferentiation in vivo.
a. Negative control for secondary antibody staining performed without primary antibody (Blue: DAPI, Red: lectin). b. Whole mount staining images show lectin-perfused vessels in the hindlimb muscle tissue from non-operated mice. (Red: lectin; Blue: DAPI; Scale bar = 50 µm, 10 z-stack, 0.8 um/slide). c. Single-channel and merged images of whole-mount staining of DAPI, YFP, CD31, and lectin from the ischemic hindlimb at day 3 post-surgery, captured at 20x magnification. In a–c, results represent two independent experiments.
Extended Data Fig. 9 Angiogenic transdifferentiation in vivo.
a. Negative control for secondary antibody staining performed without primary antibody (Blue: DAPI, Red: lectin). b. Whole mount staining images show YFP + CD31+ double-positive cells incorporated into lectin-perfused vessels in the hindlimb muscle tissue at day 7 post-surgery. (Red: lectin; Green: YFP; Gray: CD31; Blue: DAPI; Scale bar = 50 µm, 14 z-stacks, [1 µm/slice]). c. Single-channel and merged immunofluorescence images of whole mount staining of DAPI, YFP, CD31, and lectin in ischemic hindlimb at day 7 post-surgery, captured at 20x magnification. d. Representative flow cytometry plots show a reduction in YFP + CD31+ cell population in the ischemic limb of OSMI-4-treated groups compared to vehicle-treated groups at day 7 post-surgery (n = 3 mice per group, results are representative of two independent experiments). In a-c, results represent two independent experiments.
Extended Data Fig. 10 Manipulation of O-GlcNAcylation in fibroblasts orchestrates vascular recovery.
a, b. Western blots illustrate the knockdown efficacy of OGT (a) or OGA (b) in tail-tip fibroblasts obtained from fibroblast-specific knockdown mice and control mice. c, d. Area under the curve analysis of recovery from ischemia in Oga-flox and Oga-cKO mice, separated by sex. Male: n = 7 mice per group; female: n = 8 mice per group. P values are from two-way ANOVA. e–g. Weights of Ogt-flox and Ogt-cKO, Oga-flox and Oga-cKO mice during recovery from ischemia, Oga-flox and Oga-cKO mice separated by sex. h, i. Quantification of CD31+ cells in ischemic recovered limbs of Oga-flox and Oga-cKO mice at day 21, separated by sex. Male: n = 7 mice per group; female: n = 8 mice per group. j–k. Quantification of CD144+ cells in ischemic recovered limbs of Ogt-flox and Ogt-cKO (j) or Oga-flox and Oga-cKO (k) mice on day 21 post-surgery. n = 6 mice per group. In c–k, data are mean ± SD, results represent two independent experiments. In h–k, P values are from two-tailed unpaired Student’s t-test.
Supplementary information
Supplementary Tables 1–3 (download XLSX )
Supplementary Table 1. Identified metabolites from metabolomic analysis. A list of upregulated metabolites identified from the LC–MS-based metabolomic analysis, displayed in rank order of fold change at day 3 versus day 0. Supplementary Table 2. Cells and reagents. Supplementary Table 3. Antibody list.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots and gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, S., Lu, A.J., Nagueh, E.S. et al. O-GlcNAcylation promotes angiogenic transdifferentiation to reverse vascular ischemia. Nat Cardiovasc Res 4, 904–920 (2025). https://doi.org/10.1038/s44161-025-00673-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-025-00673-7
This article is cited by
-
Mechanisms, precision therapies, and technological frontiers in coronary atherosclerosis: a comprehensive review
Acta Pharmacologica Sinica (2026)