Abstract
Clinical trial-site selection is often inefficient, leading to low enrolment, poor participant diversity and costly delays. We developed DocTr, a cross-modal deep learning framework to optimize this process. DocTr uniquely integrates patient encounter data from medical claims, unstructured trial documents and historical enrolment relationships from OpenPayments data to recommend clinician investigators, specifically optimizing for recommendation accuracy, demographic fairness and operational efficiency. Evaluated on 24,984 clinicians and 5,210 trials, DocTr achieved 58% higher match similarity than leading baselines. A genetic optimization algorithm further refines recommendations, improving fairness scores related to patient race and ethnicity by up to 25% compared with the ground-truth enrolment while minimizing competing trials to near zero. DocTr also provides accurate recruitment cost estimations. By making site selection substantially more efficient, accurate and fair, this model offers a powerful method to accelerate patient access to new therapies.
Similar content being viewed by others
Main
Clinical trials are essential for medical advancement, but their success depends on effective trial site and investigator selection, which significantly impacts recruitment and outcomes. Identifying suitable clinicians remains challenging1. Traditional manual recruitment methods are slow, biased, geographically limited and often overlook optimal sites. This inefficiency leads to poor enrolment (37% of sites under-enrol, 11% enrol none) and costly delays (80% of trials miss timelines, costing $600,000 to $8 million daily)2,3. A study analysing trial recruitment from 2007 to 2010 revealed that more than 60% of planned recruitment sites enrolled fewer than 100 patients, and nearly 15% failed to recruit any participants4.
An artificial intelligence (AI) system matching clinicians to trials using their patient data and trial criteria offers a promising solution. Different from existing patient–trial matching methods that assess individual eligibility5,6, we focus on clinician-level site recommendation based on population characteristics. By aligning clinicians’ patient populations with trial needs, such systems can improve recruitment efficiency, boost enrolment success, reduce screening failures and mitigate bias. However, building this system faces challenges. On the data level, acquiring large-scale, high-quality linked data (clinician history, patient encounters, trial criteria, enrolment information) is hindered by data silos, privacy regulations and integration complexity. On the task level, effectively using diverse real-world data (free-text trial documents, tabular patient and clinician data) requires cross-modal methods. Recommendations must also balance multiple objectives: accuracy, fairness, diversity and minimizing concurrent trial competition.
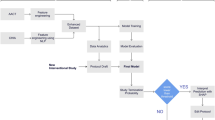
To address these challenges, we introduce DocTr, a cross-modal deep learning model for clinician recommendation (Fig. 1). Our contributions are fourfold. First, we constructed a large-scale linked trial-site dataset by integrating public data sources such as trial documents, patient claims, census data and clinician–trial payment relationships from OpenPayments (https://openpaymentsdata.cms.gov/). Second, DocTr demonstrates superior site recommendation performance. It utilizes a cross-modal graph architecture with genetic optimization for ranking and, when evaluated on 24,984 US clinicians and 5,210 trials, achieved a recommendation similarity score of 0.6, which is 58% higher than leading baselines. This result highlights its strong zero-shot capability and robustness. Third, DocTr produces fair recommendations while minimizing competing trials. A genetic optimization sub-module enhances fairness, increasing race and ethnicity fairness entropy by 9% and 25%, respectively, compared with the ground truth, and reduces the average number of competing trials to near zero. Finally, the model provides practical cost estimations for both trials and specific clinicians, offering valuable financial insights for planning and budgeting.
The pretrained ClinicalBERT processes the unstructured trial summary and criteria text, which are inputted into a text semantic encoder to obtain summary and criteria embeddings, denoted as es and er, respectively. Patient encounter vectors, which include counts of all ICD-10 codes, are processed through a code semantic encoder to produce encounter embeddings, denoted as ec. The heterogeneous graph convolutional network processes the enrolment graph between trials and clinicians to generate structural-enriched trial embeddings gt and clinician embeddings gc. All clinician and trial embeddings are merged to compute the final matching score. Fairness scores are calculated based on sensitive features, including geographic locations, race and ethnicity. These scores are then ranked by a genetic algorithm to generate the final recommendation list for the trial. The final recommendation performance is evaluated from different perspectives including accuracy, fairness, competing trials and cost. GAT, Graph Attention Network; NCTID, National Clinical Trial Identification Number; NPI, National Provider Identifier. Clinical, medical term, USA and fairness icons from Freepik (www.freepik.com); all other icons from Flaticon.com (www.flaticon.com).
Results
Here we report our dataset, task formulation and how DocTr substantially improves the accuracy of clinician recommendations for clinical trials while simultaneously enhancing fairness and operational efficiency.
The linked dataset and evaluation metrics
To build a model capable of effective clinician–trial matching, we constructed a unique linked dataset by integrating data from four primary sources. We sourced trial details, including eligibility criteria and summaries, from ClinicalTrials.gov (https://clinicaltrials.gov/). Patient encounter records were derived from the large, de-identified IQVIA medical claims database (https://www.iqvia.com/solutions/real-world-evidence/real-world-data-and-insights), from which we profiled each clinician using a five-year average (2016–2020) of their patient counts across 1,718 distinct three-digit ICD-10 (The International Classification of Diseases, 10th Revision) codes. We also incorporated regional census data for subsequent fairness evaluations7. The final integrated dataset used for this study encompasses 5,210 clinical trials and 24,984 clinicians across the USA.
The crucial link between these disparate datasets is the OpenPayments database (https://openpaymentsdata.cms.gov/), administered by the Centers for Medicare & Medicaid Services (CMS). This public resource tracks payments from industry sponsors to healthcare providers. By matching trial National Clinical Trial Identification Number from ClinicalTrials.gov and clinician National Provider Identifiers from claims data with the payment records in OpenPayments, we established a historical connection between specific clinicians and the trials they participated in. These documented payment relationships serve as the ground truth for our model, representing a real-world proxy for a clinician’s enrolment in a given trial. This integrated dataset provides a comprehensive view of clinician expertise, their access to relevant patient populations and their historical trial engagement. The data statistics about trials and clinicians used in this work are shown in Supplementary Table 2.
Using this dataset, we formulate the site selection problem as a clinician ranking task. The objective is to develop a model that takes a clinical trial and a clinician as input and computes a matching score. For any given trial, the model ranks a pool of candidate clinicians based on their matching scores, producing an ordered list of recommendations.
To benchmark our model, we compared it against a diverse set of baselines, including naive methods (random selection, popularity-based selection), general-purpose recommendation models (for example, LightGCN8, NCL9 and ConvNCF10), and specialized clinical trial models (Doc2Vec11, FRAMM12). Furthermore, to isolate the contributions of our model’s key components, we conducted an ablation study by evaluating two variants: DocTr-semantic, which relies only on semantic information from trial documents and patient data; and DocTr-structural, which uses only structural information from the historical enrolment graph.
Our primary evaluation metric is composite similarity (CS), which measures the similarity between the profiles of recommended clinicians and those of the ground truth clinicians who were actually enrolled in a trial. A higher CS score indicates a better match. We report this metric for fixed-size recommendation lists with the top K highest matching scores, denoted as CS@K (for K = 5, 10, 20), and for a list size matching the actual number of enrolled clinicians in each trial, denoted as CS@GT (Ground Truth). To assess fairness, we use entropy to measure the diversity of recommendations across sensitive attributes like race and ethnicity, where higher entropy signifies greater diversity. Finally, for cost and enrolment predictions, we evaluate accuracy using the concordance correlation coefficient (CCC), where a value of 1 indicates perfect agreement. A detailed description of all inputs, models and metrics can be found in the Methods. The experiments were designed to assess DocTr’s recommendation accuracy, its ability to promote fairness while minimizing competing trials, and its utility in practical financial planning.
The clinician recommendation performance for new unseen clinical trials
This approach mimics the real-world scenario in which trial-site managers are required to identify suitable clinicians from scratch for upcoming trials. To assess DocTr’s generalization and adaptability, we evaluate it under a challenging temporal split setting, dividing data 8:1:1 based on trial start times. This ensures test set trials (starting 5 October 2020 to 31 December 2026) are entirely unseen during training (1 October 1989 to 3 October 2019) and validation (4 October 2019 to 5 October 2020), potentially including unseen clinicians. Of the 3,767 unique clinicians in the test set, 1,027 (27.3%) were entirely new to the model.
Performance results (Table 1) show DocTr significantly outperforms all baselines. Compared with the best baseline, NCL, DocTr achieves a 58%–62% higher CS score (0.60 versus 0.37 and 0.38). This score also exceeds the ground truth CS, which measures the similarity among clinicians actually enrolled in the trials. Both ablated versions also outperform baselines but show a significant gap compared with the full DocTr, indicating each model component is important. As test trials are new and absent from the training relationship graph, structural information provides less predictive power here compared with the semantic model.
We also evaluated DocTr under regular settings, using a typical random 8:1:1 data split. Although the same clinicians and trials might appear across sets, each unique clinician–trial enrolment relationship is exclusive to one set. Results (Supplementary Table 1) show DocTr significantly surpasses all baselines, achieving up to 19% higher CS than the best baseline FRAMM. When comparing performance between regular and temporal settings (Supplementary Fig. 2), DocTr’s performance declined only 12%, markedly less than the average baseline drop of 43%. This smaller performance gap highlights DocTr’s stability, stemming from its effective use of both semantic and structural information when encountering unseen data. Furthermore, DocTr consistently outperformed FRAMM across all clinical trial phases and major disease categories even in the challenging temporal split evaluations (Supplementary Fig. 1).
Fairness and competing score of the clinician recommendations
This section evaluates how our genetic algorithm enhances fairness and reduces competing trials in recommendations. The algorithm assigns equal weight to sensitive attributes (race, ethnicity, geography). Fairness is measured using entropy12, where higher values indicate greater diversity. As the genetic algorithm alters the recommendation results, we also report the altered CS score. All results are from the temporal split setting (Table 2).
We compare DocTr-genetic (optimized recommendations) with DocTr-regular (based solely on matching scores). The genetic algorithm significantly improved fairness without compromising performance (composite similarity). Compared with the best baseline model, DocTr-genetic increased race and ethnicity fairness entropy by 7% and 17%, respectively. Geographical entropy showed no significant change, potentially because of the initially wide USA-based candidate distribution; improvements might be more evident in geographically focused searches. The ground-truth clinician fairness scores were higher than most general recommenders, suggesting fairness considerations in trial design. Figure 2a,b compares race and ethnicity distributions for DocTr-genetic, the baseline FRAMM and the ground truth. Our study focused on race and ethnicity as primary optimization targets because of the profound and persistent disparities in these areas. Although gender was also considered in our model, the gender distribution is already diverse in the dataset. Therefore, baseline disparities and subsequent improvements were less pronounced in our dataset (entropy = 0.999).
a, Race distributions of the clinician list are recommended by DocTr, FRAMM and Ground Truth. Numbers are absolute increase ratios or decrease ratios compared with the ground truth distribution. b, Ethnicity distribution of the clinician list is recommended by DocTr, FRAMM and Ground Truth. c, Average number of ongoing competing trials in the recommended clinicians. DocTr-genetic improves the recommendation results to near-zero competing trials. Bars represent the mean and error bars indicate ± s.d. The sample size, n = 5 independent model evaluations, was performed using different random seeds.
Figure 2c shows that DocTr-genetic reduced the average number of competing trials per recommended clinician to near zero (0.03), a marked improvement over DocTr-regular (2.1 competing trials). This reduction enhances operational efficiency. Traditional methods might favour frequently chosen clinicians, increasing competition. By contrast, the genetic algorithm explores diverse clinician combinations, considering expertise, patient data, history and current trial load, thereby improving diversity and reducing competition without sacrificing relevance. This balanced approach is crucial for ethical and practical aspects of clinical trial recruitment.
Estimating recruitment costs and enrolments for trials and clinicians
Estimating trial recruitment costs and enrolments is crucial for optimizing financial planning and operational efficiency. Accurate predictions enable better resource allocation and minimize financial waste. DocTr utilizes its cross-modal embeddings to achieve precise estimations by identifying similarities with existing trials and clinicians. We evaluate accuracy at both trial and clinician levels (Fig. 3). In the trial-level estimation, for a test trial, we identify the top three closest trial embeddings via cosine distance. The average costs and enrolments from these similar trials (sourced from payment data and trial documents) provide the estimate. In the clinician-level estimation, for a target clinician, we find the top three closest clinician embeddings. Their average recruitment costs (from payment data) estimate the target clinician’s cost. As ground truth enrolment per clinician is unavailable, we use payment amounts as a proxy, assuming enrolment is proportional to payment.
a, Trial cost and enrolment estimation. b, Estimating cost per new clinician. Clinical and money icons from Freepik (www.freepik.com); doctor and budget icons from Flaticon.com (www.flaticon.com).
As shown in Table 3, the model demonstrated strong predictive accuracy across multiple financial and enrolment metrics. The estimation of total cost per trial achieved a CCC of 0.83, indicating a high degree of agreement between predicted and actual costs. Similarly, at the clinician level, DocTr accurately estimated the total cost per clinician (CCC = 0.75) and the average cost per enrolment per clinician (CCC = 0.84). Although the prediction for total enrolment per trial was more moderate (CCC = 0.57), the strong performance on cost-related metrics reflects the precision of DocTr’s learned embeddings. These results show the efficacy of the joint embeddings in capturing latent financial and operational patterns. This foresight can facilitate strategic decision-making, such as selecting cost-effective recruitment strategies and prioritizing trials based on financial feasibility, thereby helping to balance cost management with the advancement of medical research.
Discussion
In this study, we introduced DocTr, a cross-modal deep learning recommendation model designed to effectively match clinicians with clinical trials. DocTr integrates diverse data sources—clinical trial documents, patient encounter claims and clinician–trial payment relationships—creating comprehensive embeddings enriched with semantic and structural insights from historical enrolment data. Tested rigorously on a large-scale dataset (24,984 clinicians, 5,210 trials) using regular and temporal splits, DocTr demonstrated superior performance. It achieved 58% higher similarity in clinician recommendations compared with the best baseline when evaluated on unseen, newly proposed trials. DocTr also showed robustness and versatility across all trial phases and disease categories.
Beyond matching accuracy, a key advancement of our work is the integration of a genetic optimization algorithm to enhance recommendation fairness and operational efficiency. This component significantly enhanced recommendation fairness regarding ethnicity, race and geography. DocTr also effectively minimized competing trials among recommended clinicians to near zero, a common recruitment challenge. Furthermore, the model provided accurate recruitment cost estimates, aiding budget management and planning. Race and ethnicity are not used in any part of the primary matching score calculation. Instead, demographic data are used exclusively as a post-processing optimization factor within the genetic algorithm. The categories for race and ethnicity are drawn directly from the US Census Bureau to align with national standards. Although DocTr cannot address inequities embedded within the protocol’s text13, it is designed to address inequities in the site selection process. By identifying a broad pool of qualified clinicians beyond the usual networks and then using a genetic algorithm to explicitly optimize for geographic and demographic diversity among the recommended sites, DocTr works to mitigate the systemic biases that occur during recruitment.
Although our model focuses on data-driven factors, we recognize that real-world site selection is also influenced by strategic considerations, particularly the involvement of key opinion leaders (KOLs). Pharmaceutical sponsors may prioritize engaging KOLs to enhance a trial’s credibility, facilitate adoption of the new therapy upon approval and guide the trial’s scientific direction. DocTr, by design, focuses on optimizing for recruitment potential by matching trial criteria to clinician practice patterns and enrolment history. It does not explicitly model a clinician’s influence or reputation, because these qualitative attributes are not captured in claims or payment data. Therefore, we see DocTr not as a replacement for strategic planning, but as a powerful complementary tool. It can be used to generate a robust, data-validated list of high-potential sites, which trial managers can then refine based on strategic priorities like KOL inclusion. It is worth noting, however, that the OpenPayments data used to build our enrolment graph may be correlated with KOL status, because influential clinicians often receive higher payments for trial leadership and consulting roles. This suggests an implicit signal of influence may already be captured in the model’s structural component. Future research could explore incorporating proxies for KOL status, such as publication records or grant awards, to create an even more comprehensive recommendation system.
A key distinction of our work is its focus on clinician-level site recommendation, which differs fundamentally from existing patient–trial matching approaches5,6,14 focused on individual eligibility (a classification task requiring granular electronic health records). By operating upstream to identify optimal sites before individual recruitment, DocTr complements patient-matching systems, enhancing outreach efficiency and maximizing population-level recruitment probability.
Despite these strengths, This research has certain limitations. Cost estimation accuracy may be affected by biases or ambiguities in OpenPayments data (for example, unclear payment purposes, underrepresentation of non-industry trials, shared payments). Using ICD-10 codes as input might introduce bias, because they are primarily billing codes potentially influenced by reimbursement factors. This is particularly relevant in highly subspecialized fields like oncology and cardiology, in which our model showed slightly lower performance. An analysis of our data confirmed that clinicians in these fields have significantly sparser patient encounter vectors (average L0 norm of 0.4 versus 0.6 for other domains). When a clinician’s practice is highly focused on a narrow set of conditions (for example, a specific cancer type), the sparsity in clinician vectors will have a larger effect on the cosine similarity, which leads to worse performance. Using more comprehensive clinician information beyond ICD codes could mitigate this issue in future work. However, our results show they provide sufficient signal for population-level matching. Similarly, our approach to evaluating fairness relies on using the clinician’s practice zip code and census data as a proxy for their patients’ demographic distribution. This is a notable simplification, because the actual demographic makeup of a clinician’s patient panel can differ substantially from that of the surrounding area, especially for specialists who draw patients from a wider region. This approximation was necessary because of the absence of patient-level demographic data in our claims dataset, but it should be considered a key constraint when interpreting our fairness entropy scores. Moreover, the concept of diversity in clinical trials is multi-faceted, encompassing not only race, ethnicity, and geography, but also other critical factors like age, gender, socio-economic status and disability, as expertly detailed in recent literature15,16,17. The exclusion of these fairness metrics is a decision necessitated by the absence of reliable patient-level data in our source claims dataset and the inadequacy of geographical census data as a proxy for these variables. Another limitation is the model’s geographical scope, which is currently limited to US clinicians because of data constraints.
Furthermore, the accuracy of the cost estimation results is constrained by assumptions. The OpenPayments data do not specify whether payments are directly for recruitment versus other activities (for example, consulting), and they represent only a fraction of total trial costs, most of which are paid to institutions rather than individuals. Our assumption that payments are proportional to enrolment is a necessary proxy, not a reflection of complex real-world financial arrangements. Therefore, the cost estimations should be interpreted as directional financial insights rather than comprehensive budgetary predictions. The challenge of disambiguating payments linked to multiple clinicians also persists. Lastly, our use of ClinicalBERT with mean pooling to generate sentence embeddings, although effective, could potentially be improved by using dedicated sentence transformer models18, which are specifically optimized for semantic retrieval and may offer enhanced performance. Future work could address these limitations by incorporating richer datasets (institutional information, detailed enrolment history, structured electronic health records, clinical notes, recruitment status and indications for competing trials calculation), extending or testing DocTr globally, and conducting public engagement, especially in diverse and hard-to-reach populations.
Methods
This section details our methodology for recommending clinical trial sites, integrating NLP, graph-based learning and fairness-aware optimization. It comprises three key components: (1) generating semantic-enriched embeddings from trial and patient data; (2) creating structural-enriched embeddings from enrolment history; and (3) a matching algorithm balancing accuracy, fairness and practical constraints. All patient data in this study are statistically de-identified and certified as compliant with the Health Insurance Portability and Accountability Act. Because this study utilized fully anonymized, pre-existing data, it was considered non-human subjects research, and thus a waiver of consent and specific institutional review board approval for this secondary analysis were not required.
Task formulation
Formally, in this paper, NT represents the number of trials and NC and denotes the number of clinicians. The model utilizes the distribution of patients across various disease codes as a proxy for the encounter capabilities of clinicians. The distribution of patients across disease codes in a clinician’s practice provides a quantifiable representation of their clinical focus and expertise domains. This distribution reflects both the clinician’s expertise and their access to relevant patient populations—two critical factors for successful trial recruitment.
To model trials, we incorporate trial document information such as inclusion criteria, research summaries, disease categories and trial phases. The model is trained to generate ranked lists of clinicians for each trial. We outline the model inputs and the matching task as detailed below.
Input 1 (clinician data)
The patient encounter vector for a clinician \({\bf{\text{c}}}\in {{\mathbb{R}}}^{{N}_{D}}\) is the patient counts categorized under ND different ICD-10 codes associated with the clinician, aggregated to three digits. This vector is derived from patient-level claims records by clinicians’ ID (National Provider Identifier) using IQVIA’s real-world claims dataset (https://www.iqvia.com/solutions/real-world-evidence/real-world-data-and-insights). We compute the annual total of patient counts and then calculate the five-year average from 2016 to 2020 to derive c. To capture both the scale of a clinician’s practice and their area of specialization, this input vector is constructed by concatenating two components: (1) the raw five-year average patient counts for each ICD-10 code, and (2) a normalized vector representing the percentage distribution of these counts.
Input 2 (trial data)
Each trial t includes a trial summary s, inclusion criteria r and trial condition category codes d. This information is extracted from ClinicalTrials.gov (https://clinicaltrials.gov/). Both s and r consist of free text. We exclusively focus on the inclusion criteria because exclusion criteria do not directly assist in selecting clinicians or trial sites and it is extremely difficult to reliably model the absence of a condition from claims data. The condition category code d corresponds to ICD-10 and shares the same dimensions as the patient encounter vector c.
Input 3 (clinician–trial relationships)
We use the payment records from clinical trials to clinicians as a proxy for historical trial enrolment patterns. The data are sourced from the OpenPayments dataset (https://openpaymentsdata.cms.gov/). The clinician–trial relationships consist of a list of pairs (c, t, m). Each pair represents a payment record to the clinician c associated with the trial t identified by the trial’s National Clinical Trial Identification Number. The value m indicates the payment amount in USD and is utilized in our cost evaluation.
Task (clinician–trial recommendation)
The clinician–trial recommendation task is framed as a ranking task. Given a clinical trial t and a clinician c, the model calculates the matching score y = ϕ(t, c). For a given clinical trial t and a set of NC clinicians (c1, c2,..., \({\bf{c}}_{N_{C}}\)), the final matching result is a list of clinicians ordered by their matching scores (y1, y2,..., \({\bf{y}}_{N_{C}}\)).
Learning semantic-enriched embeddings from trial summary, eligibility criteria and patient encounter information
We generate embeddings from diverse data sources (disease codes, trial summaries, eligibility criteria) into a unified latent space using different modules.
Learning semantic embeddings using trial documents and patient encounters
Trial summaries and inclusion criteria (extracted via the trial parser19) from ClinicalTrials.gov are embedded using ClinicalBERT20. Formally, using ClinicalBERT, we obtain embeddings for all summary and criteria sentences \([{{\bf{\text{s}}}}_{1},{{\bf{\text{s}}}}_{2},\ldots ,{{\bf{\text{s}}}}_{n}]\) and \([{{\bf{\text{r}}}}_{1},{{\bf{\text{r}}}}_{2},\ldots ,{{\bf{\text{r}}}}_{n}]\). We then compute the trial summary embedding es and the eligibility criteria embedding er by applying mean pooling to all sentences, as follows:
In addition, we derive embeddings for each clinician. Directly using disease code vectors (that is, high-dimensional binary vectors) as embeddings are impractical because of the sparse and high-dimensional nature of the ICD codes. Hence, we use another encoder, parameterized by ϕc, compressing the high-dimensional disease codes into lower-dimensional latent vectors ec. We use different encoders to generate semantic embeddings for the clinicians, criteria and sentences as:
These capture medical concept relationships, improving similarity calculations between clinicians and trials.
Enriching semantic embeddings using enrolment history
We also leverage enrolment history of individual clinicians to enhance the clinician’s semantic embeddings. To achieve this, we generate the enrolment history embeddings for each clinician by consolidating the criteria r and summary s embeddings from the trials they have historically enrolled in. The aggregation for the ith clinician is defined as follows:
here \({{\bf{\text{h}}}}_{r}^{(i)}\) and \({{\bf{\text{h}}}}_{s}^{(i)}\) are the aggregated criteria and trial summary embeddings of historical trials enrolled by the ith clinician, respectively. r(k), s(k) indicates the criteria and summary embeddings of the kth trial where the ith clinician has enrolled.
Structural-enriched embeddings for clinical trials and clinicians
Semantic similarity alone is limited, struggling with new trials and ignoring latent preferences. We introduce structural embeddings, incorporating enrolment history via a graph neural network.
Constructing the heterogeneous trial–clinician graph
To construct the heterogeneous trial–clinician graph G(V, A), we start by defining trial–clinician relationships pairs (c, t, m), where c is a clinician, t is a trial and m indicates the payment amount for clinician c running trial t. The undirected graph comprises a node set V that includes two distinct types: clinicians c and trials t:
where vc,i represents the ith clinician node, and vt, j represents the jth trial node. We use the clinician data c as node embeddings for clinician nodes and trial criteria embeddings t for trial nodes. The adjacency matrix A denotes the graph edges, where each entry Aij = 1 indicates an edge between clinician node vc,i and trial node vt, j whenever there is a record (ci, tj, m) in the trial–clinician pairs data.
Learning structural-enriched embeddings using heterogeneous LightGCN
To derive structurally enriched embeddings from the trial and clinician data, we use the a heterogeneous graph neural network framework. In particular, we apply LightGCN8. The graph convolution operation in LightGCN is defined as:
where Vk is the node embedding matrix at the kth layer, \(\hat{A}\) is the symmetrically normalized adjacency matrix, and D is the diagonal degree matrix with \({D}_{{ii}}={\sum }_{j}{A}_{ij}\).
Given the graph’s heterogeneity—where each trial node connects directly to clinician nodes and vice versa—it is essential to align the trial and clinician node embeddings within the same latent space. This alignment is integrated into the node embedding aggregation process:
where \({\pi }_{t}\) and \({\pi }_{c}\) are two mapping functions that adjust trial or clinician node embeddings to the same space, respectively.
The final node embeddings are obtained by averaging the embeddings across all layers:
where K is the total number of layers. The structural-enriched embeddings for a single trial and clinician are denoted as gt and gc, respectively.
Calculating matching scores and optimizing the fairness of recommendations
We combine embeddings to calculate matching scores and use a genetic algorithm to generate a final recommendation list optimized for accuracy, fairness and minimal competition.
Calculating matching scores and model loss
The final matching score between a trial and a clinician consists of two components: the semantic matching score and the structural matching score.
Given the semantic-enriched trial summary embeddings es, criteria embeddings er, clinician embeddings ec and historical trial embeddings hr and hs, the semantic matching score is calculated using the dot product between the trial and clinician embeddings:
where [·|·] denotes vector concatenation, and Wt and Wc are weight matrices that transform the concatenated vectors to the same dimension.
Similarly, the structural matching score is calculated as:
where gt and gc are structural-enriched summary embeddings and criteria embeddings, respectively.
The overall final matching score is then calculated by combining these two scores:
where α is a learnable parameter adjusted adaptatively to balance the importance of semantic and structural information in the matching process. Sorting the matching scores from highest to lowest, we can identify the top K clinicians as the final recommendations.
To enhance the ranking results of our matching scores, we utilize the Bayesian Personalized Ranking (BPR) loss21. The BPR loss is designed to optimize pairwise ranking, making it particularly suitable for situations in which we aim to maximize the relative score between positive and negative pairs. Specifically, for each trial–clinician pair (denoted as i) we randomly select another clinician from the candidate set (denoted as j) to act as a negative example. The objective is to ensure that the model assigns a higher score to the positive trial–clinician pair \({\hat{y}}_{i}\) than to the randomly selected negative pair \({\hat{y}}_{j}\). Mathematically, the BPR loss is formulated as:
where σ is the sigmoid function, and \(\lambda {||\Theta ||}^{2}\) is a regularization term that helps to prevent overfitting by penalizing large weights in the model. The model is optimized by minimizing this loss function, which effectively maximizes the margin between the positive and negative pairs’ scores. This allows the model to focus on ranking the correct clinician–trial pairs higher in the matching process.
Improving the fairness of recommendations and reducing competing trials through a genetic algorithm
Real-world recruitment requires balancing accuracy with fairness (racial, ethnic, geographic diversity) and minimizing competing trials. We use a genetic algorithm for this multi-objective optimization to select the top K clinicians.
In our methodology, each potential solution is represented as a clinical trial associated with a list of K clinicians, where each clinician is associated with their matching score, number of competing trials, demographics (race and ethnicity) and geographical information of their patients. Note that the claim data do not include patient demographics, so we use the OpenPayments data to get the location of the clinician, and then use the zip code-level census data as a proxy of patient demographic distributions. We define competing trials as trials that start before the current trial and have not finished. The fitness score, which evaluates the quality of each solution, considers both the total matching score and the fairness scores. The fairness scores are calculated using entropy measures for gender, race, ethnicity and geographic diversity. Concretely, given a group of K clinicians, the fitness score f is calculated as:
where m denotes the mth sensitive attribute, \({H}_{m}\in [0,1]\) denotes the normalized entropy of the attribute, n is the number of possible values for the attribute and pi is the probability (ratio) of the ith value in the selected group of K clinicians. Higher entropy values indicate more diversity and fairness in the selected group of clinicians. Ci denotes the number of competing trials of the ith clinician. We use hyper-parameters β to control the contribution of each attribute to the final fitness score. The trial recruiter can easily change the weight to outline specific attributes.
The genetic algorithm evolves an initial random population of solutions (sets of K clinicians) over generations. It uses fitness-proportional selection to choose parents, crossover to swap segments of clinician sets, and mutation (low-probability replacement of a clinician) to maintain diversity and avoid local optima. The process repeats for a set number of generations, returning the solution with the highest fitness score.
This methodology ensures the recommended clinicians are accurate (high matching scores), diverse (representing different races, ethnicities, geographic locations) and have minimal competing trials. The genetic algorithm is computationally efficient (heuristic) and flexible—additional attributes like cost can be easily incorporated by modifying the fitness score. The algorithm is detailed in Supplementary Table 3, and the inference pipeline is shown in Supplementary Table 4. The DocTr model can be extended to other geographical settings by adding relevant clinician and enrolment data.
Experiment settings
To evaluate the effectiveness of the DocTr model comprehensively, we compare it against a diverse set of baseline models. These baselines encompass a range of methodologies from simple approaches to advanced models tailored for clinical trial recommendations. This selection aims to provide a thorough comparative analysis across different dimensions of recommendation quality. Detailed information about these baseline models and hyper-parameters can be found in Supplementary Section C.
-
Naive recommendation models: random clinicians, popular clinicians.
Note that both clinical trial models, Doc2Vec and FRAMM, were not initially designed for large-scale trial-site recommendations, presenting limitations when adapted to the current dataset. We do not include individual-level patient–trial matching models because their tasks and model inputs and outputs are fundamentally different from this work. We also compare DocTr’s performance against its ablated version.
DocTr-structural
This version of the DocTr model excludes semantic embeddings based on trial summary and eligibility criteria and patient diagnoses, relying solely on structural information derived from the graph neural network based on clinician–trial relationships.
DocTr-semantic
By contrast, this variant removes structural embeddings and does not utilize the graph neural network, basing predictions purely on semantic information.
For evaluating DocTr and comparators, we use CS, a metric tailored to clinical trial recruitment, rather than conventional metrics like hit rate or recall. Standard metrics are inadequate because multiple clinicians can be equally suitable (for example, those with similar patient populations and expertise, reflected in similar ICD-10 distributions); requiring exact ID matches penalizes functionally equivalent recommendations. CS better reflects the real-world goal of finding clinicians with appropriate profiles by quantifying the similarity between the embeddings of recommended clinicians and those actually enrolled in the trial.
The formula for calculating CS for the top K recommended clinicians is defined as follows:
where \({\hat{{\bf{\text{c}}}}}_{k}\) is the embedding of the kth recommended clinician and cj is the embedding of the jth clinician who participated in the trial, ||·||2 denotes the Euclidean norm. CS values range from −1 to 1, with higher values indicating greater similarity between recommended and enrolled groups, and 0 suggesting no relationship. In the evaluation, K represents the number of recommended clinicians, which set to 5, 10 and 20. We also assess CS@GT, which K sets to the actual number of clinicians who participated in each trial, thereby varying K across different trials. We also report the CS score of the ground truth clinicians enrolled in trials, denoted as Ground Truth.
We evaluate cost estimation accuracy using CCC a robust measure of agreement between actual recruitment costs (Y) and predicted costs (\(\hat{Y}\)):
where \({\rm{cov}}(Y,\hat{Y})\) is the covariance between the ground truth Y and the predicted \(\hat{Y}\), σ and μ denote the variance and means respectively. CCC ranges from −1 (perfect disagreement) to 1 (perfect agreement), where 1 indicates perfect agreement, 0 indicates no agreement and −1 indicates perfect disagreement.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The OpenPayments data used in this study are publicly available from the Centers for Medicare & Medicaid Services (CMS) at https://openpaymentsdata.cms.gov/. Clinical trial data analysed in this study are publicly available from ClinicalTrials.gov (https://clinicaltrials.gov/). Census data are available from the United States Census Bureau (https://www.census.gov/)7. Processed data generated during this study are available via Zenodo at https://doi.org/10.5281/zenodo.18207089 (ref. 22).
Code availability
The custom computer code used to implement DocTr and process the data is publicly available via GitHub at https://github.com/v1xerunt/DocTr and available via Zenodo at https://doi.org/10.5281/zenodo.18207089 (ref. 22).
References
Briel, M. et al. Exploring reasons for recruitment failure in clinical trials: a qualitative study with clinical trial stakeholders in Switzerland, Germany, and Canada. Trials 22, 844 (2021).
Tufts Center for the Study of Drug Development 89% of trials meet enrollment, but timelines slip, half of sites under-enroll. Tufts CSDD Impact Rep. 15 (2013).
Johnson, O. An evidence-based approach to conducting clinical trial feasibility assessments. Clin. Invest. 5, 491–499 (2015).
Luo, J., Chen, W., Wu, M. & Weng, C. Systematic data ingratiation of clinical trial recruitment locations for geographic-based query and visualization. Int. J. Med. Inform. 108, 85–91 (2017).
Gao, J., Xiao, C., Glass, L. M. & Sun, J. COMPOSE: cross-modal pseudo-Siamese network for patient trial matching. In Proc. 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining (eds Gupta, R. et al.) 803–812 (Association for Computing Machinery, 2020).
Jin, Q. et al. Matching patients to clinical trials with large language models. Nat. Commun. 15, 9074 (2024).
Census Regions and Divisions of the United States. Secondary Census Regions and Divisions of the United States (US Census Bureau, 2022); https://www2.census.gov/geo/pdfs/maps-data/maps/reference/
He, X. et al. LightGCN: simplifying and powering graph convolution network for recommendation. In Proc. 43rd International ACM SIGIR Conference on Research and Development in Information Retrieval (eds Huang, J. et al.) 639–648 (Association for Computing Machinery, 2020).
Lin, Z., Tian, C., Hou, Y. & Zhao, W. X. Improving graph collaborative filtering with neighborhood-enriched contrastive learning. In Proc. ACM Web Conference 2022 (WWW '22) (eds Laforest, F. et al.) 2320–2329 (Association for Computing Machinery, 2022).
He, X. et al. Outer product-based neural collaborative filtering. In Proc. 27th International Joint Conference on Artificial Intelligence (IJCAI'18) (ed. Lang, J.) 2227–2233 (AAAI Press, 2018).
Biswal, S., Xiao, C., Glass, L. M., Milkovits, E. & Sun, J. Doctor2Vec: dynamic doctor representation learning for clinical trial recruitment. In Proc. AAAI Conference on Artificial Intelligence (eds Conitzer, V. & Sha, F.) 557–564 (AAAI Press, 2020).
Theodorou, B., Glass, L., Xiao, C. & Sun, J. FRAMM: Fair ranking with missing modalities for clinical trial site selection. Patterns 5, 100944 (2024).
Sliwinski, K., Miller, J. E. & Lynch, H. F. Examining equity-related eligibility criteria in clinical trials supporting 2022 US drug approvals. PLoS ONE 20, e0324807 (2025).
Zhang, X., Xiao, C., Glass, L. M. & Sun, J. DeepEnroll: patient-trial matching with deep embedding and entailment prediction. In Proc. Web Conference 2020 (WWW '20) (eds Huang, Y. et al.) 1029–1037 (Association for Computing Machinery, 2020).
Varma, T., Mello, M., Ross, J. S., Gross, C. & Miller, J. Metrics, baseline scores, and a tool to improve sponsor performance on clinical trial diversity: retrospective cross sectional study. BMJ Med. 2, e000395 (2023).
Varma, T., Gross, C. P. & Miller, J. E. Clinical trial diversity—will we know it when we see it? JAMA Oncol. 9, 765–767 (2023).
Varma, T., Jones, C. P., Oladele, C. & Miller, J. Diversity in clinical research: public health and social justice imperatives. J. Med. Ethics 49, 200–203 (2023).
Reimers, N. & Gurevych, I. Sentence-BERT: sentence embeddings using Siamese BERT-networks. In Proc. 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP) (eds Inui, K. et al.) 3982–3992 (Association for Computational Linguistics, 2019).
META Research Clinical Trial Parser. GitHub https://github.com/facebookresearch/Clinical-Trial-Parser (2020).
Huang K., Altosaar J. & Ranganath R. ClinicalBERT: Modeling clinical notes and predicting hospital readmission. Preprint at https://arxiv.org/abs/1904.05342 (2019).
Rendle, S., Freudenthaler, C., Gantner, Z. & Schmidt-Thieme, L. BPR: Bayesian personalized ranking from implicit feedback. In Proc. 25th Conference on Uncertainty in Artificial Intelligence (UAI '09) (ed. McAllester, D.) 452–461 (AUAI Press, 2009).
Gao J., Xiao C., Glass L. M., Harrison E. M. & Sun J. v1xerunt/DocTr: v1.0. Zenodo https://doi.org/10.5281/zenodo.18207089 (2026)
Acknowledgements
J.G. acknowledges the receipt of studentship awards from the Health Data Research UK-The Alan Turing Institute Wellcome PhD Programme in Health Data Science (grant no. 218529/Z/19/Z) and Baidu Scholarship. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.G. and J.S. conceived and designed the study. J.G. developed the methodology, and performed the data collection and statistical analysis. E.M.H. and L.M.G. provided clinical interpretation and guidance on the healthcare context. J.G. and C.X. wrote the original draft of the paper. J.S. and E.M.H. supervised the project. All authors contributed to the critical revision of the paper and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
C.X. is an employee of GE Healthcare. L.M.G. is an employee of IQVIA. J.S. is a co-founder and equity holder of Keiji AI and holds an academic appointment at the University of Illinois at Urbana-Champaign. Keiji AI develops artificial intelligence technologies related to clinical research and trial analytics. The other authors declare no competing interests. All analyses and conclusions presented in this manuscript are those of the authors and do not necessarily represent the views of GE Healthcare, IQVIA, Keiji AI or their affiliates.
Peer review
Peer review information
Nature Health thanks Jennifer Miller and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Lorenzo Righetto, with the collaboration of the Nature Health team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, J., Xiao, C., Glass, L.M. et al. Matching clinicians with clinical trials using AI. Nat. Health 1, 290–299 (2026). https://doi.org/10.1038/s44360-026-00073-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44360-026-00073-6