Abstract
Behavioral regression occurs in approximately 40% of individuals with SHANK3-associated autism spectrum disorder (ASD). We previously reported that significant behavioral regression in a small cohort of patients with SHANK3 haploinsufficiency, triggered by subclinical infections, responded to immunomodulator treatments. We hypothesize that behavioral regression results from the interplay between SHANK3 deficiency and neuroinflammation. Using Shank3 exon 4–22 deletion heterozygous mutant (Sh3+/−) mouse, which shows no significant behavior impairments, we established a preclinical model – Shank3 haploinsufficiency mouse undergoing a systemic inflammation challenge via intraperitoneal injection of lipopolysaccharides (LPS). We found that, two weeks after LPS challenge, wild-type mice (WT) recovered but Sh3+/− mice exhibited motor impairment, anxiety-like behaviors, and excessive grooming, similar to Shank3 exon 4–22 deletion homozygous mutants. Anti-inflammatory treatment partially reversed LPS-induced behavioral changes. Transcriptomic analysis revealed upregulation of neuroinflammation-related genes and downregulation of synaptic function-related genes in LPS challenged Sh3+/− mice. Especially, pro-inflammatory genes and microglia markers were overly activated that may result from the increased toll-like receptor 4 (TLR4) in Sh3+/− mice. Microglia overactivation elevated synapse engulfment and disrupted synaptic protein may underlie LPS-triggered worsen behavior phenotypes in Sh3+/− mice. Together, our findings indicate that neuroinflammation increases the penetrance of behavioral impairment in Shank3 haploinsufficiency mice and support a potential mechanism for the behavioral regression in human SHANK3 related disorders for future investigations.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a group of neurodevelopmental conditions with typical onset before the age of three [1]. A significant proportion of children with ASD shows developmental arrest at an early developmental age and behavioral regression throughout the lifespan. Behavioral regression is characterized by loss of previously acquired skills, including loss of purposeful hand use, loss of acquired speech, and gait abnormalities [2,3,4]. Catatonic features are frequently associated with behavioral regression observed in ASD [5, 6]. However, the underlying mechanism for the early developmental arrest and behavioral regression remains poorly understood.
The SHANK3 mutation is one of the most common monogenic defects associated with ASD in humans [7,8,9,10]. Genetic mutations in SHANK3 gene can manifest as Phelan-McDermid Syndrome (PMS), characterized by a chromosomal deletion of 22q13.3 or single nucleotide variants, indels, exonic deletions within the SHANK3 gene in individuals with primary ASD [11]. Patients with SHANK3 haploinsufficiency also exhibit severe intellectual disability, motor deficits, and speech delays [12,13,14]. Developmental regression with catatonic features has been reported in ~40% of cases with SHANK3 haploinsufficiency [15,16,17,18,19,20]. We previously described four patients with pathogenic SHANK3 mutations who experienced acute and severe behavioral regression following possible subclinical viral infections [21]. Notably, treatment with intravenous immunoglobulins (IVIG) combined with anti-inflammatory drugs and immunomodulators significantly reversed the behavioral regression in these cases [21]. This clinical report strongly suggests a potential link between inflammation and behavioral regression associated with SHANK3 haploinsufficiency. However, the direct causality could not be investigated and determined from human studies.
SHANK3 protein is primarily characterized as a scaffold protein located at the postsynaptic density of neurons, interacting with other synaptic proteins and affecting synaptic function [22]. However, the role of SHANK3 in function of non-neuronal cells such as microglia has been poorly studied [23,24,25,26]. Microglia activation can sculpt synapses and brain circuits and change animal behavioral phenotypes during normal development as well as in response to neuroinflammation [27, 28]. Aberrant microglia activation has been associated with ASD [29,30,31,32]. Our previous collaborative studies showed that inflammation induces acute behavioral changes [33] and aggravates hypothermia in Shank3 mutant mouse model [34]. These initial findings suggest a potential mechanistic link between SHANK3 deficiency and inflammatory response.
Our lab previously generated a Shank3 mutant mouse line with a deletion of exon 4 to 22 (Shank3Δe4-22) that disrupts most of the Shank3 coding region [35]. Shank3Δe4-22 homozygous but not heterozygous mice showed robust face validity for autism-like behaviors [35]. Therefore, Shank3Δe4-22 heterozygous mice offer a unique opportunity to examine the interaction between genes and environment, specifically whether neuroinflammation may modify the penetrance of behavioral phenotypes associated with SHANK3 deficiency in humans. In this study, we utilized the Shank3Δe4-22 heterozygous mice and investigated the behavioral outcome, transcriptomic profiles, microglia activation, and synapse marker after systemic inflammatory challenge induced by lipopolysaccharides (LPS). These findings provide a general insight on the role of neuroinflammation in ASD susceptibility.
Results
Systemic inflammation via LPS modified the penetrance of motor dysfunction, anxiety-like behavior, and repetitive behavior in Shank3 Δe4-22 heterozygous (Sh3 +/−) mice
To explore whether inflammation plays a role in regression of ASD-associated behaviors, we challenged Sh3+/− mice by intraperitoneal injection of LPS (Sh3+/−+ LPS). Control mice were given the same volume of normal saline (PBS) (Sh3+/− + PBS). Both LPS-treated Sh3+/− and wild-type (WT) mice experienced weight loss at 24 h compared to PBS-treated mice (Fig. 1A). This indicated systemic inflammation induced by LPS, consistent with previous reports [36]. LPS-treated mice regained body weight at 72 h post-injection, suggesting recovery from the LPS challenge (Fig. 1A). Acute behavioral changes, including decreased locomotor activity [37], social avoidance [38], and depressive-like behavior [39], have been reported in LPS-treated mice around 24 h post-injection. However, interpretating these behavioral changes in the context of ASD model may be confounded by immediate sickness of test mice [33, 40, 41]. Therefore, we performed behavioral analysis two weeks after LPS injection to distinguish “LPS and SHANK3 haploinsufficiency interaction effect” vs. “acute LPS effect” (Fig. 1B). The Shank3Δe4-22 homozygous (Sh3−/−) group was included for a comparison [35].
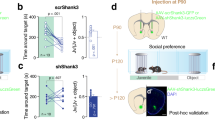
A The body weight loss after LPS intraperitoneal injection. Mouse was weighed before injection, and at 3, 6, 24, 48, and 72 h after LPS (1 mg/kg) or the same volume of PBS injection. Each mouse’s body weight was normalized to the weight before injection. Two-way repeated measures ANOVA was performed (genotype/LPS effect F (3, 50) = 34.32, p < 0.0001; Hour effect F (4, 200) = 84.15, p < 0.0001; genotype/LPS X Hour interaction F (12, 200) = 22.64, p < 0.0001). Post-hoc Tuckey’s test showed no significant difference between WT + PBS (n = 8) and Sh3+/−+PBS (n = 17) or between WT + LPS (n = 8) and Sh3+/−+LPS (n = 21) at 3 h, 6 h, 24 h, 48 h and 72 h after injection. Mouse body weight started to reduce significantly after LPS injection compared to PBS injection at 24 h, 48 h and 72 h. Mouse body weight from LPS groups significantly increased at 72 h compared to 48 h after injection in both WT + LPS and Sh3+/−+LPS groups. See multiple comparison statistics in Supplementary Table 5. B. Schematic drawing for the experimental design. C-D. LPS challenge impaired motor function in Sh3+/− mice. Steady speed rotarod (16 rpm, C) and accelerated speed rotarod (4–40 rpm, D) tests were performed and latency to fall was measured. No significant difference was found between WT + PBS and Sh3+/−+PBS (16 rpm, p = 0.5487, 4–40 rpm, p = 0.3283) or between WT + PBS and WT + LPS (16 rpm, p = 0.9797, 4–40 rpm, p = 0.6711). Sh3+/−+LPS significantly reduced latency to fall compared to Sh3+/−+PBS (16 rpm, p = 0.0126, 4–40 rpm, p = 0.0061) or compared to WT + LPS (16 rpm, p = 0.002, 4–40 rpm, p = 0.001). Sh3−/− mice significantly reduced latency to fall compared to WT + PBS (16 rpm, p < 0.0001, 4–40 rpm, p < 0.0001). E-F. LPS challenge increased anxiety-like behavior in Sh3+/− mice. Mouse duration spent in the center of the open field test and travel distance in light chamber of light/dark box test were measured. No significant difference found between WT + PBS and WT + LPS (OFT, p = 0.5956, light/dark box p = 0.6004). Sh3+/−+LPS showed less center duration than Sh3+/−+PBS (p = 0.0033) despite that Sh3+/−+PBS showed increased center duration than WT + PBS (p = 0.0005), and Sh3+/−+LPS showed less travel distance in light chamber than Sh3+/−+PBS (p = 0.0158) or WT + LPS (p = 0.0107). G. LPS challenge increased repetitive behavior in Sh3+/− mice. Grooming behavior was hand scored and compared. Sh3+/−+LPS showed longer grooming duration than Sh3+/−+PBS (p = 0.0348) or WT + LPS (p = 0.0135). H-I. LPS challenge had no effect on sociability of Sh3+/− mice. Three-chamber tests were conducted and duration spent in mouse chamber (S), empty chamber (E), proximity regions in mouse chamber (SP) and empty chamber (EP) were compared (Two-way repeated measures ANOVA. H. genotype effect F (4, 68) = 1.590, p = 0.1869, S/E effect F (1, 68) = 25.46, p < 0.0001, genotype X S/E interaction F (4, 68) = 0.8135, p = 0.5209; I. genotype effect F (4, 68) = 1.398, p = 0.2441, SP/EP effect F (1, 68) = 34, p < 0.0001, genotype X SP/EP interaction F (4, 68) = 0.4236, p = 0.7911). No significant difference of duration was found between S vs. E (p = 0.5325) or SP vs. EP (p = 0.2529) in Sh3−/− mice. All other groups of mice showed significantly increased duration in S vs. E or SP vs. EP. * p < 0.05, ** p < 0.005, *** p < 0.0005, **** p < 0.0001.
To assess the motor coordination, latency to fall from the rotarod was measured in both steady speed (16 rpm) and accelerated speed (4–40 rpm) conditions. As expected, Sh3−/− mice showed significantly reduced latency to fall compared to WT + PBS group [35]. No significant difference was observed between WT + PBS and WT + LPS groups, but Sh3+/− mice treated with LPS showed significantly shorter latency to fall than the PBS group (Fig. 1C, D). The duration of mouse spent in center of open field test (OFT) showed no significant difference between WT + PBS and WT + LPS groups, but Sh3+/−+LPS spent significantly less time in the center compared to Sh3+/−+PBS (Fig. 1E). In the light/dark box test, distance of mouse moved in light chamber showed no significant difference between WT + PBS and WT + LPS groups while Sh3+/−+LPS significantly moved less in light chamber compared to Sh3+/−+PBS (Fig. 1F). Repetitive behavior was assayed through scoring grooming behavior. Sh3+/−+LPS showed significantly longer grooming duration than Sh3+/−+PBS while WT + PBS and WT + LPS groups were comparable (Fig. 1G). We also conducted a three-chamber social preference test. As expected, Sh3−/− mice showed no preference for social targets compared to WT. However, both LPS- and PBS-treated Sh3+/− and WT mice spent significantly more time in the chamber with the social target (S) or in the proximity region of the social target (SP) (Fig. 1H, I).
To explore potential sex effect, we compared motor function and sociability in both PBS- and LPS-treated female and male Sh3+/− mice. LPS-treated male Sh3+/− mice exhibited significant reduced latency to fall in rotarod, although no significant difference found between LPS and PBS group in female Sh3+/− mice (Supplementary Fig. 1A, B). Both LPS- and PBS-treated male Sh3+/− mice spent significantly more time in S or SP, but no significant difference found in female Sh3+/−mice (Supplementary Fig. 1C, D). Taken together, our results indicated that, LPS treatment in Sh3+/− mice significantly impaired the motor performance on rotarod, changed exploratory or anxiety-like behaviors in OFT and light/dark box test, and increased grooming duration, but did not affect sociability in three-chamber test. By comparison, LPS treatment in WT mice did not have any significant effect on those behaviors. These findings suggest the interplay of inflammation and Shank3 haploinsufficiency in select domains of behaviors.
To assess whether anti-inflammation treatment can reverse the behavioral changes induced by LPS in Sh3+/− mice, we applied mefenamic acid (MFA), a nonsteroidal anti-inflammatory drug (NSAID). One of the MFA targets is the intracellular cyclooxygenase (COX) signaling, which can be activated by LPS [42,43,44]. Previous study also reported that MFA has neuroprotective effect [45]. Given that both WT + LPS and Sh3+/−+LPS group exhibited impaired motor function at 1 week after the LPS injection (Supplementary Fig. 2A, B) but such behavior deficits of WT + LPS group recovered to baseline at 2-week post-injection (Fig. 1C, D), we chose day 7 post-injection as the endpoint of MFA treatment (Fig. 2A). We administered MFA via intraperitoneal injection once a day for seven days following LPS injection and then performed rotarod test 24 h after the final dose of MFA to evaluate the treatment efficacy (Fig. 2A). In the steady speed (16 rpm) rotarod test, both WT + LPS + MFA and Sh3+/−+LPS + MFA groups showed significantly increased latency to fall compared to vehicle-treated groups (Fig. 2B), suggesting that MFA effectively mitigated the motor function deficits induced by LPS. Sh3+/−+LPS + MFA showed trend of shorter latency to fall than WT + LPS + MFA (Fig. 2B, p = 0.091), suggesting potential less pronounced anti-inflammatory efficacy and the presence of exacerbated inflammatory responses in Sh3+/− mice. In contrast, in the accelerated speed (4–40 rpm) rotarod test, MFA did not restore motor function in either WT or Sh3+/− mice (Fig. 2C). Moreover, when comparing MFA treatment in female and male Sh3+/− mice, MFA significantly increased latency to fall in female Sh3+/− mice but not in male Sh3+/− mice (Fig. 2D, E). Overall, our results showed that LPS-induced motor function impairment in Sh3+/− mice can be partially reversed by anti-inflammatory treatment. Future studies will include additional behavioral assays, as well as time-course and dose-dependent responses to MFA treatment.
A Schematic drawing for experiment design. Anti-inflammatory treatment with intraperitoneal MFA injection was given to mice once every day for 7 days after LPS injection. Rotarod tests were performed 24 h after the last does of MFA injection. B-C. MFA partially improved the rotarod test performance. After 1-week treatment of MFA, steady speed rotarod (16 rpm, B) and accelerated speed rotarod (4–40 rpm, C) tests were performed and latency to fall was compared. (Two-way ANOVA, B. 16 rpm, LPS/MFA effect F (1, 37) = 23.66, p < 0.0001, genotype effect F (1,46) = 1.859, p = 0.1794, LPS/MFA X genotype interaction F (1, 37) = 0.8697, p = 0.3571. C. 4–40 rpm, LPS/MFA effect F (1, 85) = 1.721, p = 0.1932, genotype effect F (1,85) = 4.822, p = 0.0308, LPS/MFA X genotype interaction F (1, 85) = 0.8220, p = 0.3672). Compared to vehicle group, MFA-treated Sh3+/− and WT mice showed increased latency to fall in 16 rpm rotarod test (Sh3+/−, p = 0.0091; WT, p = 0.0002). D-E. MFA treatment was more effective in female Sh3+/− mice than male ones. Two-way ANOVA was performed to compare sex difference in MFA treatment (D. 16 rpm, LPS/MFA effect F (1, 43) = 4.626, p = 0.0372, sex effect F (1,43) = 9.227, p = 0.004, LPS/MFA X sex interaction F (1, 43) = 1.858, p = 0.1799. E. 4–40 rpm, LPS/MFA effect F (1, 16) = 2.065, p = 0.17, sex effect F (1,29) = 2.369, p = 0.1346, LPS/MFA X sex interaction F (1, 16) = 8.046, p = 0.0119). MFA-treated female Sh3+/− showed significantly higher latency to fall compared to vehicle-treated female Sh3+/− (16 rpm, p = 0.0229; 40 rpm, p = 0.0106) and MFA-treated male Sh3+/− (16 rpm, p = 0.0009; 40 rpm, p = 0.0011). * p < 0.05, ** p < 0.005, *** p < 0.0005, **** p < 0.0001.
LPS-induced differentially expressed genes (DEGs) were associated with neuroinflammation and neurotransmission in Sh3 +/− mice
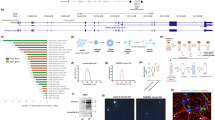
To understand how LPS-induced inflammation modifies the penetrance of motor impairment in Sh3+/− mice, we analyzed the transcriptomic profiles in the forebrains by bulk RNA-sequencing (RNA-seq) (Supplementary Fig. 3A, B, Supplementary Tables 1 and 2). LPS treatment induced hundreds of differentially expressed genes (DEGs) in both WT and Sh3+/− mice compared to PBS-treated groups. The number of upregulated genes (488 DEGs in Sh3+/−+LPS vs. Sh3+/−+PBS and 433 DEGs in WT + LPS vs. WT + PBS) was greater than that of downregulated genes (75 DEGs in Sh3+/−+LPS vs. Sh3+/−+PBS and 108 DEGs in WT + LPS vs. WT + PBS) (Fig. 3A, B). We identified 88 upregulated and 37 downregulated DEGs between Sh3+/−+LPS vs. WT + LPS (Fig. 3C). Although we found 67 upregulated and 33 downregulated genes when comparing transcriptomic profiles between Sh3+/−+PBS and WT + PBS (Fig. 3D), those genes barely overlapped with any LPS-induced DEGs observed in Sh3+/− mice. Next, we compared the overlapping genes among four comparisons (Fig. 3E, F, Supplementary Tables 3 and 4). A total of 303 shared upregulated and 12 shared downregulated DEGs were identified between WT + LPS vs. WT + PBS and Sh3+/−+LPS vs. Sh3+/−+PBS. Those shared DEGs probably represent the LPS-induced neuroinflammatory responses in mouse brains. Eight upregulated genes shared among Sh3+/−+LPS vs. WT + LPS, Sh3+/−+LPS vs. Sh3+/−+PBS and WT + LPS vs. WT + PBS were well-known for their roles in neuroinflammation (Fig. 3E highlighted with white underline, Table 1). Interestingly, 10 upregulated genes and 5 downregulated genes were found only in Sh3+/− mice not in WT mice in response to LPS (Fig. 3E, F highlighted with green underline, Tables 2–3), suggesting that LPS-induced neuroinflammation in Sh3+/− mice is mediated by distinct signaling pathways. Additionally, when we compared the overlapping genes between Sh3+/−+PBS vs. WT + PBS and Sh3+/−+LPS vs. WT + LPS, only 4 genes in total (two are protein-coding genes) were identified (Fig. 3E, F highlighted with red underline). Particularly, Acr is a Shank3 subordinate adjacent gene, whose expression increased when Shank3 is deleted [46]. To further characterize the function of DEGs from RNA-seq, we performed the pathway analysis using the QIAGEN Ingenuity Pathway Analysis (IPA) [47]. We identified more activated inflammation-associated pathways in response to LPS challenges in both Sh3+/− and WT (Supplementary Fig. 4 and 5). In contrast, inhibited neurotransmission-related pathways including NMDA receptor and calcium signaling pathways were only observed in LPS-treated Sh3+/− but not in LPS-treated WT (Supplementary Fig. 4 and 5). To understand the distinct molecular pathways in LPS-treated Sh3+/− mice, we delved into the pathway analysis from Sh3+/−+LPS vs. WT + LPS (Supplementary Fig. 6). The top five activated pathways were all related to inflammatory processes. The top two inhibited pathways, cardiac conduction and CREB signaling in neurons, represent inhibited neuron signal transmission and neuronal functions. The other three inhibited pathways, HIF1α signaling, eNOS signaling and phagosome formation pathways, represent increased neuronal vulnerability to stress and inflammatory stimuli. In addition to the top five pathways with highest z score value, we noticed that most of the upregulated signaling pathways in response to LPS were all inflammation-related, such as cytokine storm pathway and neuroinflammation pathway (Fig. 3G). In contrast, the downregulated pathways in LPS-treated Sh3+/− mice were neuronal and synaptic function-related, such as potassium channel pathway, glutamatergic signaling pathway, acetylcholine signaling pathway, and gap junction signaling pathway (Fig. 3G). Overall, our RNA-seq results suggested that a combination of exacerbated neuroinflammatory responses and impaired neuronal functions is attributed to the LPS-induced behavioral regression in Sh3+/− mice.
A–D. The volcano plots show differentially expressed genes (DEGs) distribution. DEGs were identified in four comparisons: Sh3+/−+LPS vs Sh3+/−+PBS (A), WT + LPS vs. WT + PBS (B), Sh3+/−+LPS vs. WT + LPS (C), Sh3+/−+PBS vs. WT + PBS (D). Upregulated genes are indicated in red color, downregulated genes are indicated in blue color and DEGs below threshold are indicated in grey color. E-F. The venn diagrams show shared upregulated and downregulated genes among four comparisons. Shared upregulated (E) and downregulated genes (F) numbers are shown within the diagrams. Comparison of Sh3+/−+LPS vs Sh3+/−+PBS is red diagram, WT + LPS vs. WT + PBS is blue diagram, Sh3+/−+LPS vs. WT + LPS is green diagram, and Sh3+/−+PBS vs. WT + PBS is yellow diagram. G. The bubble chart shows functions of DEGs of Sh3+/−+LPS vs. WT + LPS comparison. Volcano plot of the negative log of p-value (Y-axis) vs. the z-score (X-axis). p-value was calculated using the right-tailed Fisher’s Exact Test. A negative log of p-value cutoff of 1.3 (p < 0.05) was used to identify significantly changed functional pathways. A z-score range cutoff of ±0.1 was used to identify the activation or inhibition state of a canonical pathway. Some names of pathways are annotated in the chart. Activated pathways are orange bubbles and inhibited pathways are blue bubbles. The size of the bubbles reflects the number of genes overlapped with the pathway database.
Neuroinflammation was increased in Sh3 +/− mice
To investigate how LPS leads to aggravated neuroinflammation in Sh3+/− brains, we first measured the RNA expression of the pro-inflammatory cytokines Il1b and chemokines Cxcl10 using RT-qPCR. Il1b and Cxcl10 have been identified as common upregulated genes across all LPS-treated groups in RNA-seq (Table 1). RT-qPCR confirmed that LPS significantly increased Il1b and Cxcl10 expression in both WT and Sh3+/− mice compared to PBS groups. Although no significant difference in Il1b and Cxcl10 expression was found between Sh3+/−+PBS vs. WT + PBS, Sh3+/−+LPS showed significantly increased expression compared to WT + LPS group (Fig. 4A, B). A previous study showed that LPS treatment upregulates microglia marker gene Cx3cr1 while downregulates microglia homeostasis gene P2ry12 expression [48]. Therefore, we examined the expression of microglia markers Cx3cr1 and P2ry12 using RT-qPCR. We found that, as expected, LPS significantly increased Cx3cr1 expression and decreased P2ry12 expression in both WT and Sh3+/− mice compared to PBS groups (Fig. 4C, D). However, Cx3cr1 expression was comparable between Sh3+/−+PBS and WT + PBS but significantly increased in Sh3+/−+LPS compared to WT + LPS group (Fig. 4C). In contrast, P2ry12 expression was significantly higher in Sh3+/−+PBS than in WT + PBS and higher in Sh3+/−+LPS than in WT + LPS (Fig. 4D).
A-D. Differential expression of neuroinflammatory marker genes Il1b, Cxcl10, Cx3cr1 and P2ry12. At 24 h after LPS injection, forebrain RNA was extracted, and RT-qPCR was performed. Target gene expression was normalized to the expression of the housekeeping gene GAPDH. One-way ANOVA was performed to compare gene expression in four groups (Il1b: F = 14.93, p = 0.0004, Cxcl10: F = 16.73, p = 0.0002, Cx3cr1: F = 14.8, p < 0.0001, P2ry12: F = 8.442, p = 0.0002). As expected, compared to PBS treatment, LPS significantly increased expression of Il1b, Cxcl10, and Cx3cr1, and decreased expression of P2ry12 in both WT and Sh3+/− mice (Il1b: WT + PBS (n = 10) vs. WT + LPS (n = 9), p = 0.0005, Cxcl10: WT + PBS (n = 10) vs. WT + LPS (n = 6), p = 0.0105, Cx3cr1: WT + PBS (n = 10) vs. WT + LPS (n = 9), p = 0.0388, P2ry12: WT + PBS (n = 10) vs. WT + LPS (n = 8), p = 0.0241; Il1b: Sh3+/− + PBS (n = 12) vs. Sh3+/− + LPS (n = 11), p = 0.002, Cxcl10: Sh3+/− + PBS (n = 12) vs. Sh3+/− + LPS (n = 11), p = 0.0015, Cx3cr1: Sh3+/− + PBS (n = 12) vs. Sh3+/− + LPS (n = 12), p < 0.0001, P2ry12: Sh3+/− + PBS (n = 11) vs. Sh3+/− + LPS (n = 11), p = 0.0046). Significant increase of Il1b, Cxcl10, Cx3cr1 was observed in Sh3+/−+LPS vs. WT + LPS (Il1b: p = 0.0166, Cxcl10: p = 0.0076, Cx3cr1: p = 0.0004). In contrast, significantly increased expression of P2ry12 was observed in Sh3+/−+PBS vs. WT + PBS (p = 0.0102) and Sh3+/−+LPS vs. WT + LPS (p = 0.036). E-F. Increased immunohistochemistry fluorescence intensity of IBA1 in Sh3+/− + LPS group. Mouse brain slices were collected at 3 h and 24 h after injection, respectively. Microglia activation was labeled with IBA1 antibody (E1-E4, F1-F4). Quantitative fluorescence intensity of IBA1 at cortex regions from WT + LPS, Sh3+/−+PBS, and Sh3+/−+LPS groups were normalized to WT + PBS group and then compared (Kruskal-Wallis test, E5. p = 0.0002; F5. p < 0.0001). LPS increased microglia activation in both WT and Sh3+/− mice at both 3 h and 24 h after injection compared to PBS groups (3 h: WT + PBS (n = 10) vs. WT + LPS (n = 13), p = 0.0067, Sh3+/− + PBS (n = 12) vs. Sh3+/− + LPS (n = 11), p = 0.0004. 24 h: WT + PBS (n = 9) vs. WT + LPS (n = 16), p = 0.0053, Sh3+/− + PBS (n = 15) vs. Sh3+/− + LPS (n = 12), p < 0.0001). Moreover, LPS induced significant overactivation of microglia in Sh3+/− mice at 24 h after injection compared to WT mice, not 3 h (WT + LPS vs. Sh3+/− + LPS, 3 h, p = 0.6204, 24 h, p = 0.0017). G-I. Increased TLR4 and decreased SHANK3 protein expression in Sh3+/− mice. LPS receptor TLR4 and SHANK3 expression were examined in whole-cell lysates using western blot at 24 h after injection. Target protein expression was normalized to β-ACTIN expression (G). Quantitative protein expression from WT + LPS, Sh3+/−+PBS, and Sh3+/−+LPS were normalized to WT + PBS group and one-way ANOVA was performed (TLR4, F = 14.38, p = 0.0001, SHANK3, F = 7.681, p = 0.0033). Significantly elevated TLR4 expression was observed in Sh3+/− mice compared to WT mice (H, WT + PBS vs. Sh3+/− + PBS, p < 0.0001; WT + LPS vs. Sh3+/− + LPS, p = 0.0081). As expected, significantly reduced SHANK3 expression was detected in Sh3+/− mice (I, WT + PBS vs. Sh3+/− + PBS, p = 0.0172; WT + LPS vs. Sh3+/− + LPS, p = 0.002). J-P. Colocalization analysis of TLR4/IBA1 between WT + LPS vs. Sh3+/− + LPS group. Mouse brain slices were collected at 24 h after LPS injection in WT and Sh3+/− mice. IBA1 and TLR4 antibody were used for immunocytochemistry. Colocalization of IBA1 and TLR4 at cortex regions were analyzed and compared (J-K). Unpaired t-test was conducted. No difference in colocalization Pearson’s coefficient between WT + LPS vs. Sh3+/− + LPS was observed (L, two-tailed p = 0.2309). Sh3+/− showed smaller normalized colocalization by TLR4 (M, two-tailed p = 0.0093) but similar normalized colocalization by IBA1 (N, two-tailed p = 0.9141) compared to WT. Sh3+/− showed decreased TLR4 cellular fluorescence intensity (O, two-tailed p = 0.0183) but increased IBA1 cellular fluorescence intensity (P, two-tailed p = 0.0001) than WT. * p < 0.05, ** p < 0.005, *** p < 0.0005, **** p < 0.0001.
We further examined microglia activation by examining the expression of IBA-1, a marker widely used for the assessment of microglia activation state [49]. Compared to PBS treatment, LPS increased IBA-1 staining intensity significantly at 3 h and 24 h after LPS treatment in both WT and Sh3+/− mice. However, Sh3+/−+LPS showed significantly higher microglia activation than WT + LPS group at 24 h but not at 3 h post-injection (Fig. 4E, F). These results indicated that the prolonged activation of microglia contributed to increase of neuroinflammatory responses in Sh3+/− mice.
Next, to understand the mechanism underlying the LPS-induced microglia overactivation in Sh3+/− mice, we analyzed the LPS binding receptor toll-like receptor-4 (TLR4) expression in WT + LPS and Sh3+/−+LPS groups at 24 h post-injection [50]. We found that TLR4 expression was significantly higher in Sh3+/− than WT with either PBS or LPS treatment (Fig. 4G, H). Additionally, we measured the SHANK3 protein expression and found that either PBS- or LPS-treated Sh3+/− groups showed a reduction in SHANK3 protein expression compared to WT groups as expected, but no changes in SHANK3 expression were observed after LPS treatment (Fig. 4G, I). TLR4 is known to associate with LPS-induced microglia activation [51,52,53]. To investigate whether observed TLR4 augmentation directly facilitates microglia overactivation in LPS treated Sh3+/− mice, we analyzed colocalization between TLR4 and IBA1. We found that TLR4 and IBA1 expression were well-colocalized in WT (Fig. 4J, K, arrowhead), but a small fraction of TLR4 signals in Sh3+/− mice were not colocalized with IBA1 (Fig. 4J, K, arrow). Although colocalization Pearson’s coefficient was similar between WT + LPS and Sh3+/−+LPS group, colocalization coefficient normalized by TLR4 was lower in Sh3+/− mice and the colocalization coefficient normalized by IBA1 was similar (Fig. 4L–N). Interestingly, cellular immunofluorescence intensity of TLR4 in Sh3+/− mice is slightly lower than WT (Fig. 4O). In addition, cellular immunofluorescence intensity of IBA1 was significantly higher in Sh3+/− brains (Fig. 4P). Given that TLR4 was expressed in many different cell types including astrocytes, neutrophils, and brain endothelial cells [54, 55], these findings suggested that elevated TLR4 expression in other cell types might contribute to microglia activation in Sh3+/− mice.
Microglia activation affected synaptic structure in Sh3 +/− mice
To uncover how microglia overactivation leads to behavioral changes in Sh3+/− mice, we examined microglia engulfment and synaptic structures. We compared lysosome engulfment (lysosome marker CD68 occupancy) and synapse engulfment (presynaptic marker vGluT1 occupancy) between WT + LPS and Sh3+/−+LPS brains at 24 h post-injection. While CD68 occupancy was comparable between the two groups, vGluT1 occupancy — both within CD68-positive lysosomal puncta in microglia (CD68+IBA1+) and within the microglial cytoplasm (IBA1+) — was significantly higher in Sh3+/− mice compared to WT mice (Fig. 5A–D). These findings suggest that microglial activation is associated with increased phagocytosis of synaptic materials in the Sh3+/−+LPS group.
A-D. Increased vGluT1 engulfment in Sh3+/− mice at 24 h after LPS injection. A. Representative images of maximum projection of z-stack imaging of IBA1/CD68/vGluT1 co-staining (left), 3D rendering masked by IBA1 and CD68 (center), and enlarged images of 3D rendering (right). Total volumes of masked vGluT1 and CD68 were calculated and unpaired t-test was conducted. No significant difference of IBA1 masked CD68 occupancy was detected (B. two-tailed p = 0.7149). IBA1 and CD68 double masked vGluT1 occupancy and IBA1 masked vGluT1 occupancy was higher in Sh3+/− mice than WT mice (C. two-tailed p < 0.0001, D. two-tailed p = 0.0003). E-H. Reduced synapses in Sh3+/− mice at 24 h after LPS injection. E. Representative images of super-resolution imaging of vGluT1/PSD95 co-staining at 24 h post-injection. Colocalized puncta of vGluT1and PSD95 were indicated by arrowheads and were counted as synapse puncta. Non-colocalized puncta of PSD95 were indicated by arrows. Colocalization of vGluT1and PSD95 was analyzed, and number of colocalized puncta was counted as synapse density. Unpaired t-test was conducted. Sh3+/− mice showed a lower colocalization Pearson’s coefficient (F, two-tailed p = 0.0002), a reduced normalized colocalization coefficient by PSD95 (G, two-tailed p = 0.0056) and decreased synapse density (H, two-tailed p = 0.0107) compared to WT mice. I-M. Reduced synapses in Sh3+/− mice at 2 weeks after LPS injection. I-J. Representative images of super-resolution imaging of vGluT1/PSD95 co-staining at 2 weeks post-injection among four groups, WT + PBS, WT + LPS, Sh3+/−+PBS, and Sh3+/−+LPS. Mask was generated based on the colocalized puncta of vGluT1and PSD95. Non-colocalized puncta of PSD95 were indicated by arrows in J. One-way ANOVA was conducted for comparing colocalization Pearson’s coefficient (K. F = 4.254, p = 0.0088), colocalization coefficient by PSD95 (L. F = 26.51, p < 0.0001), and synapse density (M. F = 10.94, p < 0.0001). Sh3+/−+LPS group showed a lower colocalization Pearson’s coefficient than WT + LPS (p = 0.0067) or Sh3+/−+PBS groups (p = 0.0022), a reduced normalized colocalization coefficient by PSD95 than WT + LPS (p < 0.0001) or Sh3+/−+PBS groups (p < 0.0001), and decreased synapse density than WT + LPS (p < 0.0001) or Sh3+/−+PBS groups (p < 0.0001). N-S. Reduced PSD95, HOMER 1b/c, NR2A and GluA1 expression in Sh3+/− mice at 2 weeks after LPS injection. Postsynaptic scaffold proteins PSD95 and HOMER 1b/c (N), and excitatory receptors NMDA subunit NR2A and AMPA subunit GluA1 expression (Q) were examined in forebrain PSD fractions using western blot at 2 weeks after injection. Target protein expression was normalized to β-ACTIN expression. Quantitative protein expression from WT + LPS, Sh3+/−+PBS, and Sh3+/−+LPS were normalized to WT + PBS group and one-way ANOVA was conducted (PSD95, F = 4.763, p = 0.006, HOMER 1b/c, F = 12.28, p < 0.0001, NR2A, F = 5.06, p = 0.0091, GluA1, F = 4.951, p = 0.0313). O. Significantly decreased PSD95 expression was detected in Sh3+/− mice but not WT mice after LPS treatment (Sh3+/−+LPS (n = 11) vs. WT + LPS (n = 12), p = 0.0009; Sh3+/−+LPS vs. Sh3+/−+PBS (n = 11), p = 0.0354). No significant change in PSD95 was detected between WT + PBS (n = 12) vs. WT + LPS (p = 0.5153) or WT + PBS vs. Sh3+/−+PBS (p = 0.4845). P. Significantly reduced HOMER 1b/c expression was observed in Sh3+/− mice compared to WT mice (WT + PBS (n = 6) vs. Sh3+/− + PBS (n = 6), p < 0.0001; WT + LPS (n = 6) vs. Sh3+/−+LPS (n = 6), p = 0.0088). No significant change of HOMER 1b/c was detected after LPS treatment (WT + PBS vs. WT + LPS (p = 0.1781) or Sh3+/−+PBS vs. Sh3+/−+LPS (p = 0.318)). R. Sh3+/−+LPS group (n = 6) showed significant decreased NR2A expression compared to both the Sh3+/−+PBS group (n = 6) (p = 0.0053) and the WT + LPS group (n = 6) (p = 0.002). No significant change of NR2A was observed between WT + PBS (n = 6) vs. WT + LPS (p = 0.1404) or WT + PBS vs. Sh3+/−+PBS (p = 0.2827). S. Sh3+/−+LPS group (n = 3) showed significantly reduced GluA1 expression compared to both the Sh3+/− + PBS group (n = 3) (p = 0.0378) and the WT + LPS group (n = 3) (p = 0.0169). No significant change in GluA1 was found between WT + PBS (n = 3) vs. WT + LPS (p = 0.5898) or WT + PBS vs. Sh3+/−+PBS (p = 0.3102). * p < 0.05, ** p < 0.005, *** p < 0.0005, **** p < 0.0001.
To determine whether increased synaptic engulfment leads to synapse dismantling in the Sh3+/− brain, we examined the colocalization of the presynaptic marker vGluT1 with the postsynaptic density marker PSD95 at 24 h following LPS injection. In WT mice, vGluT1 and PSD95 puncta were well colocalized (Fig. 5E, arrowhead). In contrast, Sh3+/− mice displayed discrete PSD95 clusters that did not colocalize with vGluT1 puncta (Fig. 5E, arrow). Quantitative analysis revealed that the Sh3+/−+LPS group showed a significantly lower Pearson’s correlation coefficient for colocalization, a reduced colocalization coefficient normalized by PSD95, and decreased overall synaptic puncta density compared to the WT + LPS group (Fig. 5F–H).
Next, we compared the colocalization of vGluT1 and PSD95 among four groups — WT + PBS, WT + LPS, Sh3+/−+PBS, and Sh3+/−+LPS — at two weeks after injection. In both WT + PBS and WT + LPS groups, vGluT1 and PSD95 puncta remained closely aligned (Fig. 5I). The Sh3+/−+PBS group also exhibited substantial colocalization of vGluT1 and PSD95; however, some displaced PSD95 fragments were observed, which became more pronounced in the Sh3+/−+LPS group (Fig. 5J, arrow). Consistent with the findings at 24 h post-injection, the Sh3+/−+LPS group showed a significantly lower Pearson’s correlation coefficient for colocalization, a reduced colocalization coefficient normalized by PSD95, and decreased synapse density compared to the other groups (Fig. 5K–M). These results support the conclusion that microglial overactivation enhances synapse engulfment and contributes to synaptic structure disorganization in Sh3+/− mice in response to LPS-induced inflammation.
To investigate whether disrupted synaptic structure caused PSD protein reduction and synaptic dysfunction in Sh3+/− mice, we explored excitatory postsynaptic scaffold proteins PSD95 and HOMER 1b/c expression (Fig. 5N). No significant change of PSD95 was observed between Sh3+/−+PBS vs. WT + PBS. However, PSD95 was significantly decreased in the Sh3+/−+LPS compared to both the WT + LPS group (~28% reduction) and the Sh3+/−+PBS group (~17% reduction), suggesting that LPS treatment reduced the PSD95 protein level in Sh3+/− mice not in WT mice (Fig. 5O). In contrast, HOMER 1b/c expression was significantly decreased in Sh3+/−+PBS vs. WT + PBS (~41% reduction) and in Sh3+/−+LPS vs. WT + LPS (~23% reduction) but showed no significant change between Sh3+/−+LPS vs. Sh3+/−+PBS. It indicated that LPS treatment did not further reduce HOMER expression (Fig. 5P). PSD95 is critical for excitatory synapse function through its interactions with NMDA and AMPA receptors [56]. We examined whether the change of PSD95 may lead to the change of the expression of major subunits of NMDA receptor subunit NR2A and the AMPA receptor subunit GluA1 associated with LPS treatment (Fig. 5Q). We found that NR2A expression was reduced in the Sh3+/−+LPS group compared to both the WT + LPS group and the Sh3+/−+PBS group (Fig. 5R). The GluA1 expression was significantly reduced in Sh3+/−+LPS mice compared to both the WT + LPS group and Sh3+/−+PBS group (Fig. 5S). Taken together, these results suggested a mechanistic convergence between LPS-induced neuroimmune responses, and the synaptic disruptions known to arise from Shank3 deficiency.
Discussion
In this study, we successfully established a preclinical model and experimental protocol to assess the role of neuroinflammation induced by LPS in Sh3+/− mice. We demonstrated that LPS challenge modified the penetrance of motor function deficits, anxiety-like behavior, and repetitive behavior but did not alter social preference in Sh3+/− mice. More interestingly, the motor and emotion specific behavioral changes are reminiscent of catatonia features that are often observed in humans with SHANK3 deficiency and idiopathic ASD [57,58,59]. Additionally, our extensive comparative transcriptome analysis among the different groups validated the experimental protocol of LPS induction and revealed distinct underlying molecular mechanisms. The significantly upregulated genes ubiquitously associated with inflammatory pathways in LPS treated groups indicated the effectiveness of LPS induction. The downregulated genes in multiple pathways associated with neuronal and synaptic functions strongly support a potential causal link between LPS treatment and increased penetrance of abnormal behaviors in Sh3+/− mice. Moreover, increased microglia activation and reduced synaptic function from LPS treatment were further validated by molecular experiments. Finally, the partial reversal of motor impairment in LPS treated Sh3+/− mice through the anti-inflammation treatment with MFA further supports this causality. Notably, MFA treatment did not restore LPS-treated WT and Sh3+/− mice performance in the accelerating speed rotarod. These findings suggest that the mechanisms underlying inflammation-induced motor impairment are different between steady speed and accelerating speed rotarod tests [60]. Additionally, one-week MFA treatment was more effective in female Sh3+/− mice than male mice (Fig. 2D, E) and LPS-treated male Sh3+/− mice performance was worse than female ones (Supplementary Fig. 1A, B), indicating potential sex difference in motor function [61] and neuroinflammation process [62].
The behavior impairments induced by LPS in Sh3+/− mice could attribute to the dysfunction of neuronal cells resulted from neuroinflammation. In the previous study that our lab contributed, LPS challenge increased the excitability in D1 but not D2 medium spiny neurons using whole-cell patch clamp recordings and decreased neuron calcium transients in nucleus accumbens in response to social stimuli using in vivo Ca2+ imaging [33]. Our data showed that increased neuroinflammation downregulated genes associated with neuronal and synaptic functions in Sh3+/− mice compared to WT mice and reduced synaptic protein PSD95 in Sh3+/− mice.
LPS-induced neuroinflammation could also alter the function of non-neuronal cells in Sh3+/− mice. Our findings of augmented TLR4 expression and microglia activation associated with LPS treatment suggested that microglia are primarily involved in more significant neuroinflammatory responses in Sh3+/− mice. Consistent with the findings in many reports in literature supporting that TLR4 mediates microglial activation in brains [51,52,53], our findings also support that the upregulation of TLR4 contributes to the observed microglial activation following LPS administration. The direct causality between upregulation of TLR4 and microglia activation due to LPS induction has been demonstrated in a recent study [63]. Our observations are consistent with this finding and support the augmented TLR4 as the cause of microglia activation in SHANK3 deficiency mice. Further analysis of TLR4 and IBA1 cellular colocalization indicated that additional cell types might also be involved in LPS-TLR4 (ligand-receptor) mediated neuroinflammation. Cellular fluorescence intensity of TLR4 in fixed brain cortex regions decreased while total protein level of TLR4 in freshly prepared forebrain tissue increased in Sh3+/− mice than WT. This inconsistence implied that changes of TLR4 in Sh3+/− might be brain region specific and cell type specific. In most of the literature, SHANK3 is characterized as a PSD protein. Thus, it is an interesting and important question how a PSD protein could regulate the expression of other genes or proteins, such as the TLR4 changes we observed. Here we hypothesizes several potential mechanisms (Fig. 6A): 1. Isoform-specific nuclear functions: As shown in our previous reports, SHANK3 has numerous mRNA and protein isoforms, and not all isoforms localize to the PSD [46, 64]. We have demonstrated that some isoforms localize to the nucleus, although their functions remain unknown. It is possible that these nuclear isoforms act as transcriptional regulators. The deficiency of nuclear SHANK3 isoforms may therefore contribute to the altered expression of TLR4. 2. Non-neuronal isoform expression: Not all SHANK3 isoforms are expressed in synapses or neurons. For example, certain isoforms are highly expressed in the thymus [65] and in endothelial cells [66]. This supports the idea that SHANK3 may have broader biological functions. The loss of SHANK3 isoforms in non-neuronal cells may contribute to changes in TLR4 expression. 3. Altered PSD interactome: As a scaffolding protein in the PSD, loss of SHANK3 will inevitably affect other neuronal proteins that normally interact with SHANK3. Disruption of these interacting proteins may secondarily alter TLR4 expression. Thus, it warranted a future mechanistic study on the TLR4 signaling pathways in Shank3 haploinsufficiency mouse model.
A. SHANK3 haploinsufficiency may regulate TLR4 expression through several distinct cellular mechanisms. 1) SHANK3 at PSD can modulate PSD interactome and downstream signaling pathways which regulate TLR4 expression. 2) Nuclear SHANK3 isoforms may function as a transcriptional regulator affecting Tlr4 gene expression. 3) In non-neuronal cells, SHANK3 may participate in cell-specific signaling pathways altering TLR4 expression. B. The schematic drawing proposes a model wherein LPS treatment exacerbates neuroinflammation and drives altered behavioral phenotypes in Shank3 haploinsufficiency mouse. LPS evokes neuroinflammation via three interconnected pathways. 1) LPS binding to TLR4 expressed on peripheral neutrophils promotes blood-brain barrier infiltration and microglial activation. 2) LPS binding to TLR4 expressed on cerebral endothelial cells compromises blood-brain barrier integrity, increasing permeability for infiltrating immune components. 3) LPS that penetrates blood-brain barrier directly activates neuronal and glial TLR4, triggering proinflammatory cytokine release. Notably, Sh3+/− mice exhibited exacerbated neuroinflammation compared to WT, as evidenced by elevated proinflammatory gene expression, increased proinflammatory cytokine levels, microglial overactivation at 24 h after LPS injection. Enhanced TLR4 expression observed in Sh3+/− mice may result from three distinct cellular signaling pathways in A. These findings suggest that SHANK3 haploinsufficiency increased brain susceptibility to LPS-induced neuroinflammation. Microglia activation can sculpt synapses and brain circuits and change animal behavioral phenotypes long-term. In Sh3+/− brains, observed elevated microglia engulfment, reduced synapses, and decreased synaptic transmission-related gene expression at 24 h after LPS injection suggests increased synapse elimination in Sh3+/− mice. This early synaptic vulnerability persisted through 2 weeks post-injection, when reduced synapse density, decreased PSD95 and excitatory receptor expression were observed. It indicates that neuroinflammation-induced synaptic dysfunction was associated with behavioral deterioration found in Sh3+/− mice. Together, it supports a mechanistic link between behavioral regression and interplay between SHANK3 deficiency and neuroinflammation.
Moreover, we uncovered a mechanistic convergence between LPS-induced neuroimmune responses, and the synaptic disruptions known to arise from Shank3 deficiency. We discovered increased microglia engulfment and disrupted synaptic structure in Sh3+/− mice. Such dysfunction sustained until 2 weeks later when we identified LPS-triggered behavioral changes only in Sh3+/− mice instead of WT mice. Moreover, in a previous study, we reported that the scaffold protein PSD95 expression did not change significantly while HOMER1b/c was significantly reduced in Sh3+/− mice [35]. In the current study, we replicated the finding that HOMER1b/c expression is significantly decreased in Sh3+/− mice. However, LPS treatment did not further reduce HOMER levels. We also replicated the finding that PSD95 is not changed in Sh3+/− mice. However, LPS treatment significantly reduced PSD95 expression in Sh3+/− mice not WT mice. This pattern of PSD95 changes in response to LPS, influenced by SHANK3 haploinsufficiency, differs from that observed for HOMER1b/c. Although we previously reported no change of NR2A or GluA1 in Sh3+/− mice [35], we found that LPS treatment reduced NR2A and GluA1 expression in Sh3+/− mice instead of WT mice.
Our findings represent a first step toward understanding how SHANK3 haploinsufficiency is linked to neuroimmune signaling. Our previous study showed that SHANK3 deficiency in periphery nerve system contributes to the exaggerated inflammatory responses [67]. Our recently published collaborative study has shown that conditional knock-out of SHANK3 major isoform in brain endothelial cells recapitulated social behavior impairment and other synaptic phenotypes [66]. Our findings are aligned with previous studies showing that Shank3 deficiency exacerbates the inflammatory responses in peripheral neurons either induced by LPS [34, 67] or by maternal immune activation during the pregnancy [68]. Overall, our findings showed that LPS exacerbated neuroinflammation in Sh3+/− mouse brains through unique molecular mechanisms as summarized in Fig. 6 for further investigations in future.
In humans, haploinsufficiency of SHANK3 is highly penetrant for neurodevelopmental impairments [13]. Approximately ~70% of PMS and SHANK3 deficiency patients met the diagnostic criteria for ASD. The comorbidity of intellectual disability, anxiety, and epilepsy are highly penetrant or common [58]. There is an increasing recognition of behavioral regression and catatonia among young teens with SHANK3 deficiency [57]. However, the triggers and mechanisms underlying the behavioral regression are largely unknown. Although few clinical case reports implicated neuroinflammation as a trigger [21, 69], our findings may offer a mechanistic explanation for the behavioral regressions observed in SHANK3 patients after experiencing a subclinical infection responded to the immune modulator treatment. Our study may support a link between neuroinflammation and autism susceptibility in general.
Our results establish an important proof-of-concept for dissecting the link between neuroinflammation and autism susceptibility. The apparent limitations of this study include that we have not provided definitive causality for the cell types and specific molecular mechanism directly implicated in the increased neuroinflammatory responses indued by LPS in SHANK3 haploinsufficiency mice. Future studies are warranted to examine dosage- or treatment duration-dependent effects of LPS on behavioral outcomes at different neurodevelopmental stages and in different sexes of Shank3 deficiency mice. Despite these limitations, our results suggest that Shank3 haploinsufficiency increases susceptibility to neuroinflammation-induced behavior impairment through distinct molecular pathways. These findings provide new insights into the role of neuroinflammation in ASD and support the further investigation of anti-inflammatory treatments as both preventive and therapeutic strategies for behavior regression in ASD.
Materials and methods
Animals
Animals were housed under standard 12 h/12 h light/dark cycles (7 AM-7 PM) with food, water ad libitum, controlled temperature and humidity in the Yale University animal facility. Shank3Δe4-22 mouse line (Jax Strain #: 039524) was generated in-house and had been crossed with C57BL/6 J mouse line (Jax Strain #: 000664) over 20 generations. Wild-type (WT), Shank3Δe4-22 homozygous (Sh3−/−) and Shank3Δe4-22 heterozygous mice (Sh3+/−) were obtained through heterozygotes breeding as previously described [35]. All experiment procedures were performed within a time frame of 9 AM-5 PM. Animals of the same sex were group housed (2–5 mice per cage). Both male and female mice were used. One cage of mice with same genotype and same sex were assigned to PBS or LPS group randomly. Experimental order of different group was randomized. Mice from multiple groups for same experiments were carried out at same time. Before experiments, each mouse was cataloged with identification number in an excel sheet which included detailed information such as genotype and drug treatment. During experiments, experimenter handled mice based on its identification number and attempted to be semi-blinded. Post-processing, data handling and statistical analysis were semi-automated performed using software stated below. Mice with health problems such as runt, fight wounds, malocclusion, and eye problems, were excluded from experiments. For RNA and protein extraction, mice were deeply anesthetized with 5% isoflurane, then mouse brains were quickly collected, flash frozen in liquid nitrogen and stored in −80 °C freezer until further use. The number of animals used in each experiment was determined based on our lab prior experience and a prior power analysis was calculated using G*power statistics software [70] (Supplementary table 5). Total number of mice used for each experiment was as follows: 86 mice for MFA treatment rotarod test, 52–95 mice for open field, rotarod and three-chamber behavior test, 49–66 mice for light/dark box and grooming behavior test, 15 mice for RNA sequencing, 40 mice for RT-qPCR, 24 mice for wide-field IBA1 staining, 6 mice for TLR4/IBA1 co-staining, 6 mice for microglia engulfment and vGluT1/PSD95 staining at 24 h post-injection, 12 mice for vGluT1/PSD95 staining at 24 h post-injection, 18 mice for whole-cell protein analysis and 24 mice for synaptic protein analysis. All experiments were replicated at least once, except that RNA sequencing was done one time.
Drug
Three to five-week-old female and male mice were intraperitoneally injected with LPS (Sigma, L6511) in saline at 1 mg/kg or same volume of saline (PBS). To test the mefenamic acid (MFA) anti-inflammatory effect on behavior improvement, MFA (Thermofisher, AAJ6270522) dissolved in DMSO at 5 mg/kg or the same volume of DMSO (vehicle) was injected intraperitoneally to the mice with LPS injection once a day for 7 days. Before injection, freshly made stock solution of MFA in DMSO (MFA treatment) or DMSO (vehicle treatment) was diluted with 1:100 ratio in PBS to minimize DMSO toxicity.
Behavior assay
For behavioral experiments, mice were handled for 3 days and habituated in the procedure room for 1 h before behavioral assays. All behavioral tests were conducted once for each mouse.
Anxiety-like behaviors were tested by open field test (OFT) and light/dark box test followed previous studies [35, 71]. In OFT, mouse spontaneous activity in Noldus Open Field (4 chambers, 40 (L) x 40 (W) x 30 (H) cm per chamber) over 10 min was recorded with EthoVision XT video tracking software (Noldus). The Noldus two-chambered light/dark box is equipped with the lid of dark chamber made by infrared translucent material. The dimensions of the compartment are one third for the dark chamber (20 (L) x 20 (W) x 20 (H) cm) and two thirds for the light chamber (20 (L) x 40 (W) x 20 (H) cm). Mice were firstly placed into the open-light chamber ( ~ 700 lux) and given 5 min to freely explore both light and dark (0 lux) chambers. The video was recorded with EthoVision XT video tracking software. Ethovision software was used to automatically score the travel distance and duration in arena.
Repetitive behavior was tested by evaluating mouse grooming behaviors. Mouse was placed in a Noldus phenotyper (30 (L) x 30 (W) x 45 (H) cm) and recorded with EthoVision XT video tracking software. Mouse freely explored for 10 min and last 5 min of grooming behavior was hand scored.
Motor coordination was tested by rotarod (Med Associate) performance as described [35, 72]. Mice were tested in two 5 min trials with an inter-trial interval of 30 min. In accelerating session, revolutions per minute (rpm) of the turning barrel is accelerating from 4 to 40. In steady speed session, the rpm of the turning barrel is fixed at 16. Latency to fall, or to rotate off the top of the turning barrel, was measured by the timer.
Social preference was tested by a three-chamber assay as described [72]. An unfamiliar age- and sex-matched mouse was used as a social stimuli mouse in the three-chamber test and habituated under a wired enclosure for 3 days before testing. A customized chamber composed of a rectangular Plexiglas arena (42 (L) x 56.5(W) x 35.8 (H) cm) divided into three chambers (each chamber 18.5 ×42 cm), with 2 doors connecting chambers and two enclosures (wired pencil cups with metal vertical bars) located in the two outer chambers. Briefly, the test mouse was habituated in the center chamber for 5 min. Then the social stimuli mouse was placed under one of the pencil cups and 2 doors were lifted allowing the experimental mouse to freely explore the whole arena. The location of the empty and mouse enclosures, and the social stimuli mouse were changed and counterbalanced for each trial. The EthoVision XT video tracking software was used to score the duration of mouse spending in the empty chamber, the mouse chamber, proximity regions (5 cm in distance) close to the empty enclosure, and proximity regions close to the mouse enclosure.
Bulk RNA sequencing
Total RNA was isolated from the forebrain hemisphere using Trizol (Invitrogen, 15596018) followed manufacturer protocol and then treated with DNase I (Invitrogen, AM1907) to remove contaminating DNA. RNA samples were sent to Yale Center for Genome Analysis (YCGA) for bulk RNA sequencing (RNA-seq). See details about sequencing process in Supplementary Information. After obtaining the data, low quality reads were trimmed, and adaptor contamination were removed using Trim Galore (v0.5.0). Trimmed reads were mapped to the mouse reference genome (mm10) using HISAT2 (v2.1.0) [73]. Gene expression levels were quantified using StringTie (v1.3.3b) [74]. Differentially expressed genes were identified using DESeq2 (v 1.22.1) [75]. Unpaired t-test p value was used for filtering DEGs with significant differences between the two groups. Log2fold cutoff threshold at 1 and p < 0.05 was used to filter DEGs from two groups comparison (WT + LPS vs. WT + PBS, Sh3+/−+LPS vs. Sh3+/−+PBS, Sh3+/+LPS vs. WT + LPS and Sh3+/−+PBS vs. WT + PBS) for genes comparison and analysis. Pathway analysis: QIAGEN Ingenuity Pathway Analysis (IPA) library of canonical pathways was used to identify the pathways most significant to the dataset. Molecules from the dataset meeting the criteria in which log2fold cutoff threshold at 0.25 and p < 0.05 were associated with a canonical pathway in the QIAGEN Knowledge Base were considered for the analysis. Because using log2fold cutoff threshold at 1 only revealed three activated pathways in Sh3+/+LPS vs. WT + LPS comparison which were all inflammation-related, so we expanded the log2fold cutoff threshold to 0.25. The significance of the association between the data set and the canonical pathway was measured based on: 1) a ratio of the number of molecules from the data set that map to the pathway divided by the total number of molecules that map to the canonical pathway was displayed, 2) a right-tailed Fisher’s Exact Test was used to calculate a p-value determining the probability that the association between the genes in the dataset and the canonical pathway was explained by chance alone, 3) A z-score was calculated to indicate the likelihood of activation or inhibition of that pathway. In this paper, pathways with right-tailed Fisher’s Exact Test p < 0.05, z-score > 0.1 was considered as activated pathways and z-score < −0.1 was considered as inhibited pathways. The volcano plot was created using VolcaNoseR [76]. Veen diagram was plotted using InteractiVenn [77].
Real-Time quantitative reverse transcription PCR (RT-qPCR)
Total RNA was isolated using same protocol as RNA-seq. RNA was transcribed into cDNA using iScript cDNA Synthesis Kit (Bio-Rad, 1708890) and qPCR was performed in triplicate using iQ SYBR Green Supermix (Bio-Rad, 1708880). Gene expressions were normalized to the expression of GAPDH. Primers used to amplify cDNAs are: Il1b, forward, TGGACCTTCCAGGATGAGGACA, reverse, GTTCATCTCGGAGCCTGTAGTG; Cxcl10, forward, ATCATCCCTGCGAGCCTATCCT, reverse, GACCTTTTTTGGCTAAA CGCTTTC; Cx3cr1, forward, GAGCATCACTGACATCTACCTCC, reverse, AGAAGGCAGTCGTGAGCTTGCA; P2ry12 forward, CATTGACCGCTACCTGAAGACC, reverse, GCCTCCTGTTGGTGAGAATCATG; Gapdh forward, CAAAATGGTGAAGGTCGGTG, reverse, AATGAAGGGGTCGTTGATGG.
Immunohistochemistry
Transcardiac perfusion was conducted at both 3 h and 24 h after LPS or PBS injection with 20 mL PBS then 20 mL 4% paraformaldehyde (PFA, Santa Cruz, sc-281692). Mouse brains were collected, fixed in 4% PFA for 24 h, then changed to 30% (w/v) sucrose before sectioning. 50 μM brain sections between Bregma 1 mm to −1 mm were obtained using vibratome (Leica, VT1200S) or using cryostat (Leica, CM1860). Primary antibody including anti-rabbit-IBA1 (1:200, FUJIFILM Wako Chemicals, 019-19741), anti-mouse-TLR4 (1:100, Abcam, ab22048), anti-rat-CD68 (1:100, Abcam, ab53444), anti-mouse-vGluT1 (1:100, Synaptic Systems, 135511), anti-guinea pig-vGluT1 (1:200, Synaptic Systems, 135318), and anti-mouse-PSD95 (1:100, Invitrogen, MA1-045) were used. Secondary antibody including goat anti-mouse Alexa Fluor 488 (Abcam, ab150113), donkey anti-mouse Alexa Fluor 488 (Abcam, ab150105), donkey anti-rabbit Alexa Fluor 594 (Invitrogen, A-21207), goat anti-rat Alexa Fluor 594 (Invitrogen, A-11007), goat anti-guinea pig Alexa Fluor 647 (Abcam, ab150187), goat anti-rabbit Alexa Fluor 647 (Jackson ImmunoResearch, 111-605-144) were used at 1:500 dilution. Zeiss LSM 980 confocal microscopy with a 20X objective lens was used for wide-field IBA1 imaging, a 63X oil objective lens with Z stack (0.2 µm thickness/slice, total 30-40 slices/image) was used for engulfment imaging, and Airyscan 2 with a 63X oil objective lens was used for super-resolution imaging of vGluT1 and PSD95 colocalization. Zeiss LSM 900 confocal microscopy with a 63X oil objective lens was used for TLR4/IBA1 co-staining imaging. In total, 49 images at 3 h and 52 images at 24 h post-injection were analyzed for IBA1 fluorescence intensity, 20 images were analyzed for IBA1/TLR4 colocalization, 47 images were analyzed for microglia engulfment, and 26 images at 24 h and 61 images at 2 weeks post-injection were analyzed for vGluT1/PSD95 colocalization. Images were randomly selected from cortex regions of brain slices. Immunostaining and imaging were performed under same conditions and same settings at same time across different groups within same cohort. Cohort 1 with 8 groups was for IBA1 staining. Cohort 2 with 2 groups was for TLR4/IBA1 co-staining. Cohort 3 with 2 groups was for IBA1/CD68/vGluT1 co-staining. Cohort 4 with 2 groups was for 24 h post-injection vGluT1/PSD95 co-staining. Cohort 5 with 4 groups was for 2 weeks post-injection vGluT1/PSD95 co-staining. Biological triplicates of brain slices from one mouse brain were employed. Results from at least three mouse brains per group were analyzed and compared. ImageJ software was used for IBA1 fluorescence intensity quantification. Imaris software (Oxford Instrument) was used for microglia engulfment 3D rendering analysis. ZEN Blue software was used for colocalization analysis. See details on 3D rendering and colocalization analysis protocols in the Supplementary Information.
Western blot
Western blot was performed as previously described [78]. Whole cell lysates from mouse forebrain hemisphere were prepared using 1x lysis buffer (Cell Signaling Technology, 9803S) containing 1x protease/phosphatase inhibitor (Cell Signaling Technology, 5872S). Crude postsynaptic density (PSD) fraction from mouse forebrain hemisphere were prepared following previously described protocol with some modifications [35, 79]. See details in Supplementary Information. Samples were boiled in 4x Laemlli buffer (Bio-Rad, 161-0747) at 98 °C for 5 min before gel electrophoresis. Home-made anti-rabbit SHANK3 C-terminus monoclonal antibody (1:1000), anti-rabbit TLR4 monoclonal antibody (1:1000, Santa Cruz, sc-293072), anti-mouse PSD95 monoclonal antibody (1:1000, Invitrogen, MA1-046), HOMER 1b/c monoclonal HRP-conjugated antibody (1:1000, Santa Cruz, sc-17842-HRP), anti-rabbit GluA1 (AMPA receptor) polyclonal antibody (1:1000, Abcam, ab31232), anti-NR2A (NMDA receptor) polyclonal antibody (1:1000, Millipore, ab31232), and β-ACTIN monoclonal HRP-conjugated antibody (1:1000, Invitrogen, MA5-15739-HRP) were used.
Statistical analysis
Graphpad Prism 10.0 (GraphPad Software) was used for the statistical analysis and data plot. Differences were analyzed by student’s t-test if comparing two variables or using ANOVA if comparing more than two variables followed by multiple comparisons either using statistical hypothesis testing with correction or planned comparisons without correction. Significance was determined by two-tailed t-test p value or multiplicity adjusted p value. For data, which was not sampled from Gaussian distribution, a nonparametric test was performed. Since unequal variance observed in LPS-evoked cytokines and chemokines, Brown-Forsythe and Welch ANOVA tests were performed (Fig. 4A, B). See detailed statistical analysis for each figure in Supplementary Table 5. n represents the total number of data points per group, usually meaning the mice number with an exception that n represents number of images in immunohistochemistry results. Both sexes of mice were used and balanced across genotype and treatment, with an exception that WT + PBS and WT + LPS groups were all male mice in RNA-seq experiments. Behavioral difference between male and female mice were explored in Supplementary Fig. 1. In box plot, individual data points were plotted as: whiskers go down to the smallest value and up to the largest value, box extends from the 25th to 75th percentiles and line in the middle of the box is plotted at the median. In bar graph, individual data points were plotted and presented as average ± SEM. p < 0.05 was considered statistically significant.
References
Regier DA, Kuhl EA, Kupfer DJ. The DSM-5: Classification and criteria changes. World Psychiatry. 2013;12:92–98.
Stefanatos GA. Regression in autistic spectrum disorders. Neuropsychol Rev. 2008;18:305–19.
Manelis L, Meiri G, Ilan M, Flusser H, Michaelovski A, Faroy M, et al. Language regression is associated with faster early motor development in children with autism spectrum disorder. Autism Res. 2020;13:145–56.
Boterberg S, Van Coster R, Roeyers H. Characteristics, early development and outcome of parent-reported regression in autism spectrum disorder. J Autism Dev Disord. 2019;49:4603–25.
Kohlenberg TM, Trelles MP, McLarney B, Betancur C, Thurm A, Kolevzon A. Psychiatric illness and regression in individuals with Phelan-McDermid syndrome. J Neurodev Disord. 2020;12:7.
Ghaziuddin M. Catatonia: A common cause of late regression in autism. Front Psychiatry. 2021;12:674009.
Betancur C, Buxbaum JD. SHANK3 haploinsufficiency: A “common” but underdiagnosed highly penetrant monogenic cause of autism spectrum disorders. Mol Autism. 2013;4:17.
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, et al. Large-Scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–584 e523.
Zhou X, Feliciano P, Shu C, Wang T, Astrovskaya I, Hall JB, et al. Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes. Nat Genet. 2022;54:1305–19.
Fu JM, Satterstrom FK, Peng M, Brand H, Collins RL, Dong S, et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat Genet. 2022;54:1320–31.
Nevado J, Garcia-Minaur S, Palomares-Bralo M, Vallespin E, Guillen-Navarro E, Rosell J, et al. Variability in Phelan-McDermid Syndrome in a Cohort of 210 Individuals. Front Genet. 2022;13:652454.
Soorya L, Kolevzon A, Zweifach J, Lim T, Dobry Y, Schwartz L, et al. Prospective investigation of autism and genotype-phenotype correlations in 22q13 deletion syndrome and SHANK3 deficiency. Mol Autism. 2013;4:18.
Leblond CS, Nava C, Polge A, Gauthier J, Huguet G, Lumbroso S, et al. Meta-analysis of SHANK Mutations in Autism Spectrum Disorders: A gradient of severity in cognitive impairments. PLoS Genet. 2014;10:e1004580.
Serrada-Tejeda S, Martínez-Piédrola RM, Máximo-Bocanegra N, Sánchez-Herrera-Baeza P, Pérez-de-Heredia-Torres M. Descriptive analysis of adaptive behavior in Phelan-McDermid syndrome and autism spectrum disorder. Front Neurosci. 2022;16:893003.
Reierson G, Bernstein J, Froehlich-Santino W, Urban A, Purmann C, Berquist S, et al. Characterizing regression in Phelan McDermid Syndrome (22q13 deletion syndrome). J Psychiatr Res. 2017;91:139–44.
Denayer A, Van Esch H, de Ravel T, Frijns JP, Van Buggenhout G, Vogels A, et al. Neuropsychopathology in 7 patients with the 22q13 deletion syndrome: presence of bipolar disorder and progressive loss of skills. Mol Syndromol. 2012;3:14–20.
Zwanenburg RJ, Ruiter SAJ, van den Heuvel ER, Flapper BCT, Van Ravenswaaij-Arts CMA. Developmental phenotype in Phelan-McDermid (22q13.3 deletion) syndrome: A systematic and prospective study in 34 children. J Neurodev Disord. 2016;8:16.
Serret S, Thümmler S, Dor E, Vesperini S, Santos A, Askenazy F. Lithium as a rescue therapy for regression and catatonia features in two SHANK3 patients with autism spectrum disorder: Case reports. BMC Psychiatry. 2015;15:107.
Phelan K, McDermid HE. The 22q13.3 Deletion Syndrome (Phelan-McDermid Syndrome). Mol Syndromol. 2012;2:186–201.
Phelan K, Boccuto L, Powell CM, Boeckers TM, van Ravenswaaij-Arts C, Rogers RC, et al. Phelan-McDermid syndrome: a classification system after 30 years of experience. Orphanet J Rare Dis. 2022;17:27.
Bey AL, Gorman MP, Gallentine W, Kohlenberg TM, Frankovich J, Jiang Y-h, et al. Subacute neuropsychiatric syndrome in girls with SHANK3 mutations responds to immunomodulation. Pediatrics. 2020;145:e20191490.
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 2007;39:25–27.
He G, He Y, Ni H, Wang K, Zhu Y, Bao Y. Dexmedetomidine attenuates neuroinflammation and microglia activation in LPS-stimulated BV2 microglia cells through targeting circ-Shank3/miR-140-3p/TLR4 axis. Eur J Histochem. 2023;67:3766.
Fischer I, Shohat S, Levy G, Bar E, Trangle SS, Efrati S, et al. Hyperbaric oxygen therapy alleviates social behavior dysfunction and neuroinflammation in a mouse model for autism spectrum disorders. Int J Mol Sci. 2022;23:11077.
Cope EC, Briones BA, Brockett AT, Martinez S, Vigneron PA, Opendak M, et al. Immature neurons and radial glia, but not astrocytes or microglia, are altered in adult Cntnap2 and Shank3 mice, models of autism. eNeuro. 2016;3:0196-16.2016
Urrutia-Ruiz C, Rombach D, Cursano S, Gerlach-Arbeiter S, Schoen M, Bockmann J, et al. Deletion of the autism-associated protein SHANK3 abolishes structural synaptic plasticity after brain trauma. Int J Mol Sci. 2022;23:6081.
Eyo U, Molofsky AV. Defining microglial-synapse interactions. Science. 2023;381:1155–6.
Pereira-Iglesias M, Maldonado-Teixido J, Melero A, Piriz J, Galea E, Ransohoff RM, et al. Microglia as hunters or gatherers of brain synapses. Nat Neurosci. 2025;28:15–23.
Suzuki K, Sugihara G, Ouchi Y, Nakamura K, Futatsubashi M, Takebayashi K, et al. Microglial activation in young adults with autism spectrum disorder. JAMA Psychiatry. 2013;70:49–58.
Kumar A, Williams MT, Chugani HT. Evaluation of basal ganglia and thalamic inflammation in children with pediatric autoimmune neuropsychiatric disorders associated with streptococcal infection and tourette syndrome: a positron emission tomographic (PET) study using 11C-[R]-PK11195. J Child Neurol. 2015;30:749–56.
Han VX, Patel S, Jones HF, Dale RC. Maternal immune activation and neuroinflammation in human neurodevelopmental disorders. Nat Rev Neurol. 2021;17:564–79.
Michels S, Mali A, Jäntti H, Rezaie M, Malm T. Microglial involvement in autism spectrum disorder: insights from human data and iPSC models. Brain Behav Immun. 2025;130:106071.
Tzanoulinou S, Musardo S, Contestabile A, Bariselli S, Casarotto G, Magrinelli E, et al. Inhibition of Trpv4 rescues circuit and social deficits unmasked by acute inflammatory response in a Shank3 mouse model of Autism. Mol Psychiatry. 2022;27:2080–94.
Zhang L, Bang S, He Q, Matsuda M, Luo X, Jiang YH, et al. SHANK3 in vagal sensory neurons regulates body temperature, systemic inflammation, and sepsis. Front Immunol. 2023;14:1124356.
Wang X, Bey AL, Katz BM, Badea A, Kim N, David LK, et al. Altered mGluR5-Homer scaffolds and corticostriatal connectivity in a Shank3 complete knockout model of autism. Nat Commun. 2016;7:11459.
Piirsalu M, Taalberg E, Lilleväli K, Tian L, Zilmer M, Vasar E. Treatment with lipopolysaccharide induces distinct changes in metabolite profile and body weight in 129Sv and Bl6 mouse strains. Front Pharmacol. 2020;11:371.
Jung H, Lee D, You H, Lee M, Kim H, Cheong E, et al. LPS induces microglial activation and GABAergic synaptic deficits in the hippocampus accompanied by prolonged cognitive impairment. Sci Rep. 2023;13:6547.
Decker Ramirez EB, Arnold ME, McConnell KT, Solomon MG, Amico KN, Schank JR. The effects of lipopolysaccharide exposure on social interaction, cytokine expression, and alcohol consumption in male and female mice. Physiol Behav. 2023;265:114159.
O’Connor JC, Lawson MA, André C, Moreau M, Lestage J, Castanon N, et al. Lipopolysaccharide-induced depressive-like behavior is mediated by indoleamine 2,3-dioxygenase activation in mice. Mol Psychiatry. 2009;14:511–22.
Parrott JM, Oster T, Lee HY. Altered inflammatory response in FMRP-deficient microglia. iScience. 2021;24:103293.
Hodges SL, Nolan SO, Tomac LA, Muhammad IDA, Binder MS, Taube JH, et al. Lipopolysaccharide-induced inflammation leads to acute elevations in pro-inflammatory cytokine expression in a mouse model of Fragile X syndrome. Physiol Behav. 2020;215:112776.
Eliopoulos AG, Dumitru CD, Wang CC, Cho J, Tsichlis PN. Induction of COX-2 by LPS in macrophages is regulated by Tpl2-dependent CREB activation signals. EMBO J. 2002;21:4831–40.
Blais V, Turrin NP, Rivest S. Cyclooxygenase 2 (COX-2) inhibition increases the inflammatory response in the brain during systemic immune stimuli. J Neurochem. 2005;95:1563–74.
Cimolai N. The potential and promise of mefenamic acid. Expert Rev Clin Pharmacol. 2013;6:289–305.
Khansari PS, Halliwell RF. Evidence for neuroprotection by the fenamate NSAID, mefenamic acid. Neurochem Int. 2009;55:683–8.
Lu X, Ni P, Suarez-Meade P, Ma Y, Forrest EN, Wang G, et al. Transcriptional determinism and stochasticity contribute to the complexity of autism-associated SHANK family genes. Cell Rep. 2024;43:114376.
Krämer A, Green J, Pollard J Jr., Tugendreich S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics. 2014;30:523–30.
Sousa C, Golebiewska A, Poovathingal SK, Kaoma T, Pires-Afonso Y, Martina S, et al. Single-cell transcriptomics reveals distinct inflammation-induced microglia signatures. EMBO Rep. 2018;19:e46171.
Norden DM, Trojanowski PJ, Villanueva E, Navarro E, Godbout JP. Sequential activation of microglia and astrocyte cytokine expression precedes increased Iba-1 or GFAP immunoreactivity following systemic immune challenge. Glia. 2016;64:300–16.
Park BS, Song DH, Kim HM, Choi BS, Lee H, Lee JO. The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature. 2009;458:1191–5.
Lee JW, Nam H, Kim LE, Jeon Y, Min H, Ha S, et al. TLR4 (toll-like receptor 4) activation suppresses autophagy through inhibition of FOXO3 and impairs phagocytic capacity of microglia. Autophagy. 2019;15:753–70.
Yao L, Kan EM, Lu J, Hao A, Dheen ST, Kaur C, et al. Toll-like receptor 4 mediates microglial activation and production of inflammatory mediators in neonatal rat brain following hypoxia: role of TLR4 in hypoxic microglia. J Neuroinflammation. 2013;10:785.
Zusso M, Lunardi V, Franceschini D, Pagetta A, Lo R, Stifani S, et al. Ciprofloxacin and levofloxacin attenuate microglia inflammatory response via TLR4/NF-kB pathway. J Neuroinflammation. 2019;16:148.
TLR4 expression in the Human Protein Atlas. https://www.proteinatlas.org/ENSG00000136869-TLR4/single+cell, Accessed Date Accessed.
Sjöstedt E, Zhong W, Fagerberg L, Karlsson M, Mitsios N, Adori C, et al. An atlas of the protein-coding genes in the human, pig, and mouse brain. Science. 2020;367:eaay5947.
Chen X, Levy JM, Hou A, Winters C, Azzam R, Sousa AA, et al. PSD-95 family MAGUKs are essential for anchoring AMPA and NMDA receptor complexes at the postsynaptic density. Proc Natl Acad Sci USA. 2015;112:E6983–6992.
Dhossche D, de Billy C, Laurent-Levinson C, Le Normand MT, Recasens C, Robel L, et al. Early-onset catatonia associated with SHANK3 mutations: looking at the autism spectrum through the prism of psychomotor phenomena. Front Psychiatry. 2023;14:1186555.
Srivastava S, Sahin M, Buxbaum JD, Berry-Kravis E, Soorya LV, Thurm A, et al. Updated consensus guidelines on the management of Phelan-McDermid syndrome. Am J Med Genet A. 2023;191:2015–44.
Yurumez E, Temelturk RD, Kaymak B, Karabıyıkoğlu Y, Zengin ND. Catatonia in autism spectrum disorders: Prevalence, functional impact, and diagnostic insights. Psychiatry Res. 2024;342:116250.
Monville C, Torres EM, Dunnett SB. Comparison of incremental and accelerating protocols of the rotarod test for the assessment of motor deficits in the 6-OHDA model. J Neurosci Methods. 2006;158:219–23.
Kovács AD, Pearce DA. Location- and sex-specific differences in weight and motor coordination in two commonly used mouse strains. Sci Rep. 2013;3:2116.
Han J, Fan Y, Zhou K, Blomgren K, Harris RA. Uncovering sex differences of rodent microglia. J Neuroinflammation. 2021;18:74.
Hines DJ, Choi HB, Hines RM, Phillips AG, MacVicar BA. Prevention of LPS-induced microglia activation, cytokine production and sickness behavior with TLR4 receptor interfering peptides. PLoS ONE. 2013;8:e60388.
Wang X, Xu Q, Bey AL, Lee Y, Jiang Y-H. Transcriptional and functional complexity of Shank3 provides a molecular framework to understand the phenotypic heterogeneity of SHANK3 causing autism and Shank3 mutant mice. Mol Autism. 2014;5:30.
Redecker P, Bockmann J, Böckers TM. Expression of postsynaptic density proteins of the ProSAP/Shank family in the thymus. Histochem Cell Biol. 2006;126:679–85.
Kim Y-E, Kim M, Kim S, Lee R, Ujihara Y, Marquez-Wilkins EM, et al. Endothelial SHANK3 regulates tight junctions in the neonatal mouse blood-brain barrier through β-Catenin signaling. Nat Commun. 2025;16:1407.
Han Q, Kim YH, Wang X, Liu D, Zhang ZJ, Bey AL, et al. SHANK3 deficiency impairs heat hyperalgesia and TRPV1 signaling in primary sensory neurons. Neuron. 2016;92:1279–93.
Atanasova E, Arévalo AP, Graf I, Zhang R, Bockmann J, Lutz AK, et al. Immune activation during pregnancy exacerbates ASD-related alterations in Shank3-deficient mice. Mol Autism. 2023;14:1.
Jesse S, Delling JP, Schon M, Boeckers TM, Ludolph A, Senel M. Phelan McDermid syndrome: Multiple sclerosis as a rare but treatable cause for Regression-A case report. Int J Mol Sci. 2021;22:2311.
Faul F, Erdfelder E, Lang AG, Buchner A. G*Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav Res Methods. 2007;39:175–91.
Bourin M, Hascoët M. The mouse light/dark box test. Eur J Pharmacol. 2003;463:55–65.
Moy SS, Nadler JJ, Young NB, Perez A, Holloway LP, Barbaro RP, et al. Mouse behavioral tasks relevant to autism: Phenotypes of 10 inbred strains. Behav Brain Res. 2007;176:4–20.
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37:907–15.
Pertea M, Pertea GM, Antonescu CM, Chang T-C, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33:290–5.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Goedhart J, Luijsterburg MS. VolcaNoseR is a web app for creating, exploring, labeling and sharing volcano plots. Sci Rep. 2020;10:20560.
Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics. 2015;16:169.
Wang SE, Ko SY, Jo S, Choi M, Lee SH, Jo HR, et al. TRPV1 Regulates Stress Responses through HDAC2. Cell Rep. 2017;19:401–12.
Carlin RK, Grab DJ, Cohen RS, Siekevitz P. Isolation and characterization of postsynaptic densities from various brain regions: enrichment of different types of postsynaptic densities. J Cell Biol. 1980;86:831–45.
Seaton G, Smith H, Brancale A, Westwell AD, Clarkson R. Multifaceted roles for BCL3 in cancer: a proto-oncogene comes of age. Mol Cancer. 2024;23:7.
Baba T, Mukaida N. Role of macrophage inflammatory protein (MIP)-1α/CCL3 in leukemogenesis. Mol Cell Oncol. 2014;1:e29899.
Bufi AA, Di Stefano J, Papait A, Silini AR, Parolini O, Ponsaerts P. The central role of CXCL10-CXCR3 signaling in neuroinflammation and neuropathology. Cytokine Growth Factor Rev. 2025;84:20–34.
Shenoy AR, Wellington DA, Kumar P, Kassa H, Booth CJ, Cresswell P, et al. GBP5 promotes NLRP3 inflammasome assembly and immunity in mammals. Science. 2012;336:481–5.
Han JH, Lee S, Park YS, Park JS, Kim KY, Lim JS, et al. IFITM6 expression is increased in macrophages of tumor-bearing mice. Oncol Rep. 2011;25:531–6.
Henry CJ, Huang Y, Wynne AM, Godbout JP. Peripheral lipopolysaccharide (LPS) challenge promotes microglial hyperactivity in aged mice that is associated with exaggerated induction of both pro-inflammatory IL-1beta and anti-inflammatory IL-10 cytokines. Brain Behav Immun. 2009;23:309–17.
Zhao J, Liao Y, Miller-Little W, Xiao J, Liu C, Li X, et al. STEAP4 expression in CNS resident cells promotes Th17 cell-induced autoimmune encephalomyelitis. J Neuroinflammation. 2021;18:98.
Lester SN, Li K. Toll-like receptors in antiviral innate immunity. J Mol Biol. 2014;426:1246–64.
Li Y, Fan Q, Li F, Pang R, Chen C, Li P, et al. The multifaceted roles of activating transcription factor 3 (ATF3) in inflammatory responses - Potential target to regulate neuroinflammation in acute brain injury. J Cereb Blood Flow Metab. 2023;43:8–17.
Deshmane SL, Kremlev S, Amini S, Sawaya BE. Monocyte chemoattractant protein-1 (MCP-1): an overview. J Interferon Cytokine Res. 2009;29:313–26.
Krzyzowska M, Kowalczyk A, Skulska K, Thörn K, Eriksson K. Fas/FasL contributes to HSV-1 brain infection and neuroinflammation. Front Immunol. 2021;12:714821.
Wu S, Pei Q, Ni W, Fu X, Zhang W, Song C, et al. HSPA1A protects cells from thermal stress by impeding ESCRT-0–Mediated autophagic flux in epidermal thermoresistance. J Invest Dermatol. 2021;141:48–58.e43.
Dukay B, Csoboz B, Tóth ME. Heat-Shock proteins in neuroinflammation. Front Pharmacol. 2019;10:920.
Frankiw L, Mann M, Li G, Joglekar A, Baltimore D. Alternative splicing coupled with transcript degradation modulates OAS1g antiviral activity. RNA. 2020;26:126–36.
Bao Z, Liu Y, Chen B, Miao Z, Tu Y, Li C, et al. Prokineticin-2 prevents neuronal cell deaths in a model of traumatic brain injury. Nat Commun. 2021;12:4220.
Lei J, Deng Y, Ma S. Downregulation of TGIF2 is possibly correlated with neuronal apoptosis and autism-like symptoms in mice. Brain Behav. 2022;12:e2610.
Acknowledgements
This work was supported by the National Institutes of Health (NIH) grant HD088007 MH104316, MH098114, MH117289, HD087795 (Y-H.J), the Autism Science Foundation (ASF) fellowship (S-N.Q), and the National Research Foundation of Korea (NRF) grant RS-2024-00401971, 2021R1A6A1A03039503 (S.J), Brain Pool program RS-2024-00409006 (S.E.W). BioRender was used to create schematic diagrams (Qiao, S. (2026) https://BioRender.com/lrex0mf).
Author information
Authors and Affiliations
Contributions
S-N.Q. and Y-H.J. designed the experiments. S-N.Q. collected and analyzed the data. S.W. manually scored grooming behavior. S.W., K-Y.K. and S.J. performed western blot. S-N.Q. and Y-H.J. wrote and revised the manuscripts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Ethics approval and consent to participate
Experiments procedures conducted were following the Institutional Animal Care and Use Committee (IACUC) guidelines at Yale University. The animal protocol (No. 2025-20271) was approved by Yale IACUC.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qiao, SN., Wang, S.E., Kim, KY. et al. Inflammation increases the penetrance of behavioral impairment in Shank3 haploinsufficiency mice — can it explain the behavioral regression in Autism?. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03534-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03534-2