Abstract
Colorectal cancer (CRC) remains a major global health burden with limited therapeutic options. This study identifies phosphomannomutase 2 (PMM2) as a key oncogenic driver in CRC. PMM2 is significantly upregulated in CRC tissues and cell lines, correlating with advanced tumor stages, lymphatic metastasis, and poor patient survival. Functional assays reveal that PMM2 knockdown inhibits CRC cell proliferation, migration, invasion, and glycolytic activity (reducing glucose uptake, ATP/lactate production, and extracellular acidification rate). Mechanistically, PMM2 interacts with transcriptional regulator TRIM28, promoting TRIM28 nuclear translocation, recruiting transcription factor E2F4, and enhancing KIFC3 transcription by binding to its promoter. KIFC3 mediates PMM2-driven glycolysis, as KIFC3 knockdown partially reverses PMM2-induced metabolic reprogramming and tumor growth in xenograft models. Patient-derived organoid studies further confirm PMM2’s role in promoting CRC progression through the PMM2-KIFC3 axis. Collectively, these findings establish PMM2 as a prognostic biomarker and potential therapeutic target in CRC, highlighting its critical role in metabolic reprogramming and tumorigenesis.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC), one of the most prevalent malignancies of the digestive system, arises from epithelial cells in the colon or rectum due to cumulative genetic and environmental perturbations, progressing from adenomatous polyps to invasive carcinoma [1]. Epidemiologically, CRC ranks as the third most incident cancer globally (10.0% incidence rate) and the second leading cause of cancer-related mortality (8.8%), with developing regions like China experiencing rising incidence rates despite declining trends in developed nations [2]. Current diagnostic modalities, including colonoscopy and histopathological evaluation, remain foundational, while therapeutic strategies—surgical resection, chemotherapy, and radiotherapy—show limited efficacy in advanced or metastatic stages [3]. Targeted therapies (e.g., anti-EGFR/cetuximab, anti-VEGF/bevacizumab) and immunotherapies have emerged as adjuncts, yet their applicability is restricted to <15% of metastatic CRC patients due to primary or acquired resistance [4, 5]. Prognostic biomarkers such as microsatellite instability (MSI) and mutations in KRAS, NRAS, and BRAF offer partial guidance but fail to address the unmet need for novel therapeutic targets and mechanistic insights [6, 7]. Thus, elucidating CRC pathogenesis and its molecular regulatory networks remains critical to overcoming treatment stagnation.
Phosphomannomutase 2 (PMM2), encoded by the PMM2 gene on chromosome 16p13, catalyzes the conversion of mannose-6-phosphate (Man-6-P) to mannose-1-phosphate (Man-1-P), a pivotal step in glycosylation—the enzymatic attachment of oligosaccharides to proteins or lipids to form functional glycoconjugates [8, 9]. Dysregulation of this pathway underlies PMM2-CDG (Congenital Disorder of Glycosylation type Ia), a severe multisystem disorder characterized by early mortality due to neuronal and hepatic dysfunction [9,10,11]. In cancer, PMM2 exhibits importantly roles. The accumulation of studies reveal PMM2 upregulation in breast [12, 13], renal [14], and cervical cancers [15], where it correlates with tumor aggressiveness and therapeutic resistance. In addition, PMM2 overexpression in CRC correlates with poor prognosis and resistance to copper-induced cell death and metastatic potential [16,17,18]. Moreover, PMM2 drives estrogen receptor α (ERα) stability in ERα Y537S-mutant breast cancer via interaction with FOXA1, suggesting its role in metabolic reprogramming and transcriptional regulation [12]. Despite these advances, the molecular mechanisms underlying PMM2’s oncogenic functions in CRC—particularly its crosstalk with glycolytic enzymes and metastasis regulators—remain poorly defined, underscoring the urgency of targeted investigation.
This study elucidates PMM2’s role in CRC progression: (1) promoting proliferation, migration, and metastasis through glycolytic activation and (2) regulating KIFC3 (a gene pivotal for mitotic fidelity and metabolic adaptation) through interaction with TRIM28 to recruit E2F4. Using CRC cell lines, patient-derived xenografts, and multi-omics approaches, we demonstrate that PMM2/TRIM28 complex formation enhances KIFC3-mediated glycolysis, thereby sustaining tumor growth and metastasis. Clinically, PMM2 overexpression in CRC tissues correlates with advanced tumor stages and reduced survival, validating its prognostic utility. By elucidating this axis, our findings not only advance the understanding of CRC pathogenesis but also identify PMM2 as a tractable therapeutic target, particularly for metabolic reprogramming strategies. This work bridges a critical gap in CRC biology and provides actionable insights for developing precision therapies.
Materials and methods
Experimental design
Sample size selection prioritizes ≥80% statistical power (with α = 0.05) to detect a pre-specified effect size—drawn from prior literature or preliminary pilot data. Within-group variability is estimated using pilot results or published values from similar studies, while practical limits (like limited tissue availability or reagent costs) are balanced to avoid over/undersampling. All choices are pre-documented in the study protocol to ensure transparency and methodological rigor.
For animal studies, the sample size was determined based on preliminary pilot experiments and common practices in colorectal cancer xenograft models. We used 6 mice per group, which is a standard sample size in similar studies, to observe consistent tumor growth and metabolic phenotype changes, ensuring that the observed effects are representative and not due to random variation.
Randomization strategies varied across experimental types: cell lines were allocated to groups using a random number generator; mice were assigned via block randomization stratified by body weight; human tissue samples were collected consecutively without randomization. Investigators were blinded to group allocation during data collection and outcome assessment.
Cell lines, cell culture and reagents
Human colon cancer cell lines RKO, HCT-116, HT-29, Caco-2 and HT-16, human normal colon epithelial cell line NCM460, and human embryonic kidney cell line HEK293T were purchased from the American Type Culture Collection (ATCC, USA). These cell lines were tested without mycoplasma contamination. Cells were cultured in DMEM (Invitrogen, USA) supplemented with 10% fetal bovine serum (Gibco, USA) and 1% Penicillin–Streptomycin Solution 100× (Corning, USA) in a humidified atmosphere with 5% CO2 at 37 °C.
Anti-PMM2 (10666-1-AP), anti-E2F4 (10923-1-AP), anti-GAPDH (60004-1-lg), anti-β-actin (66009-1-lg), anti-LDHA (19987-1-AP), anti-PKM2 (15822-1-AP), anti-TRIM28 (15202-1-AP), anti-Histone H3 (17168-1-AP), anti-Flag (66008-3-lg) and anti-HA (51064-2-AP) were purchased from Proteintech; Goat anti-mouse (A0216) and goat anti-rabbit (A0208) were obtained from Beyotime. Anti-KIFC3 (DF12287) was obtained from Affinity Biosciences.
Plasmid construction, transfection and viral infection
To generate PMM2 or TRIM28 overexpression vectors, PCR-amplified fragments were subcloned into the pcDNA3 backbone (Invitrogen). For short hairpin RNA (shRNA) expression, lentiviral vectors were assembled by inserting target shRNA sequences into the pSIH-H1-Puro vector (System Biosciences). Specific shRNA target sequences are detailed in Table S3. KIFC3 promoter luciferase reporter constructs were prepared by cloning PCR-amplified promoter fragments (derived from genomic DNA) into the pGL4-Basic vector (Promega). Site-directed mutagenesis of the KIFC3 promoter was performed using the Mut Express II Fast Mutagenesis Kit V2 (Vazyme, China) following the manufacturer’s protocol.
Plasmid transfection was conducted with Lipofectamine 3000 reagent (Invitrogen) according to the optimized procedures provided in the reagent manual. For lentivirus production, HEK293T cells were cotransfected with recombinant lentiviral vectors and the pPACK Packaging Plasmid Mix (System Biosciences) using Megatran transfection reagent (Origene). The resulting viruses were then used to infect HCT-116 and RKO cells as per the manufacturer’s guidelines. Stably transduced cell lines were established by selecting infected cells with 1 μg/ml puromycin.
Western blot analysis
Cells were lysed on ice for 30 min using RIPA lysis buffer supplemented with protease inhibitors. Equivalent protein samples were separated via 10% SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred onto nitrocellulose membranes. Membranes were blocked with 5% non-fat milk at room temperature for 1 h, followed by overnight incubation with primary antibodies at 4 °C. Protein bands were visualized using the Vazyme enhanced chemiluminescence (ECL) detection system.
Luciferase reporter assay
Luciferase activity was measured using the Vazyme Dual Luciferase Reporter Assay Kit following the manufacturer’s protocol. Briefly, 24 h prior to transfection, cells were seeded into 24-well plates at an appropriate density. For reporter activity analysis, cells were co-transfected with wild-type/mutant KIFC3 promoter reporter plasmids, TRIM28 expression vectors (or negative controls), and Lipofectamine 3000 transfection reagent. Post-transfection, cells were harvested and luciferase activities were quantified. Each experiment was independently repeated three times, with three technical replicates per group.
Revere transcription-quantitative PCR (RT-qPCR)
Total RNA was extracted from the collected cells using Invitrogen Trizol reagent. RNA was reverse-transcribed into cDNA with oligo(dT) primers using the TaKaRa ExScript RT-PCR Kit (Japan). Primers for real-time PCR are detailed in Table S4. Relative expression levels of target genes were calculated using the 2-ΔΔCt method, with normalization to corresponding internal control genes.
Co-immunoprecipitation (co-IP) assay
Harvested cells were lysed with ice-cold lysis buffer on ice for 30 min. Following centrifugation at 12,000 rpm for 15 min at 4 °C, the clarified supernatant was incubated with primary antibodies and rotated overnight at 4 °C. Protein G/A Agarose beads (Santa Cruz Biotechnology) were subsequently added to the antibody-supernatant mixture, and the samples were rotated for an additional 2 h at 4 °C to capture immune complexes. After washing the agarose beads three times with ice-cold lysis buffer, the beads were centrifuged, and the resulting pellets were resuspended in 2× SDS loading buffer. Samples were then boiled at 100 °C for 10 min to dissociate proteins from the beads, and the eluted proteins were analyzed via immunoblotting using the specified primary antibodies.
Chromatin immunoprecipitation (ChIP) assay
ChIP was conducted using the Magna ChIP G Assay Kit (Millipore) following the manufacturer’s protocol. Briefly, cells were cross-linked by treatment with 1% formaldehyde, harvested, and resuspended in ChIP lysis buffer. Chromatin was sheared via sonication, and lysates were centrifuged to remove debris. The resulting supernatants were collected and incubated overnight at 4 °C with target-specific antibodies and Protein G magnetic beads. After immunoprecipitation, cross-links in the precipitated chromatin complexes were reversed, and DNA fragments were purified using the kit-provided reagents. The purified DNA samples were subjected to RT-qPCR analysis using primer sequences detailed in Table S5.
Cell proliferation and colony formation assays
For the cell proliferation assay, treated cells were seeded into 96-well plates at a density of 3000 cells per well. Cell viability was assessed using the CCK-8 reagent (Dojindo) following the manufacturer’s protocol, with measurements taken every 24 h over a 5-day period. For colony formation assay, cells were seeded in 6-well plates (2000 cells per well, three technical replicates) and cultured for 10–14 days to allow colony expansion. Colonies with a diameter exceeding 1.0 mm were counted and recorded as representative growth clusters.
Cell migration and invasion assays
Cell migration was examined by wound healing assays. Cells were seeded into 6-well plates to reach 90% confluence. A uniform scratch wound was created using a 200 μl pipette tip, followed by gentle PBS washing to remove detached cells. Cells were then cultured for an additional 24 h. Wound widths at 0 h (immediately post-scratch) and 24 h were measured, and migration rates were calculated by comparing the reduction in wound area over time. Cell invasion assay was evaluated using Matrigel-coated Transwell chambers (BD Biosciences) according to the manufacturer’s guidelines. Cells were seeded into the upper chamber, and after 24 h of incubation, non-invasive cells on the upper surface were removed. Cells that migrated to the lower surface were fixed with 4% paraformaldehyde, stained with 0.5% crystal violet, and imaged. Invasive cell counts were performed by randomly selecting 5 microscopic fields per well.
Cell apoptosis
Cells were seeded into 6-well plates and cultured overnight to achieve ~70% confluence the following day. Apoptosis was detected using the Annexin V-FITC/Propidium Iodide (PI) Apoptosis Detection Kit (eBioscience). Cells were harvested, stained with Annexin V and PI according to the kit protocol, and apoptotic rates were analyzed via flow cytometry.
Glycolytic phenotype assay
To characterize cellular glycolytic activity, three detection systems were applied: the Lactate Content Assay Kit, ATP Content Assay Kit, and Glucose Uptake Assay Kit (Solarbio), which were used to assess lactate production, ATP generation, and glucose uptake, respectively, following the respective kit instructions. For measurement of lactate production, cells were plated in 96-well plates at 1000 cells per well and cultured under standard growth conditions (DMEM with 10% FBS) for 10 h. After aspirating the culture medium, cells were incubated in FBS-free DMEM for 1 h to minimize serum interference. Supernatants were then collected for lactate quantification. For measurement of ATP production, approximately 1 × 106 cells were lysed in 100 μL of the kit-provided lysis buffer. Lysates were centrifuged to remove debris, and ATP levels in the soluble fraction were measured using the assay reagents. For measurement of glucose uptake, 1 × 106 cells were seeded in 96-well plates and subjected to glucose starvation by incubation in Krebs-Ringer-Phosphate-HEPES (KRPH) buffer containing 2% BSA for 40 min. The glycolytic inhibitor 2-deoxy-D-glucose (2-DG) was added to the medium for 20 min prior to glucose uptake detection using the kit-specific reagents.
Extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) assays
ECAR and OCR were measured using the Seahorse XFe 96 Extracellular Flux Analyzer (Seahorse Bioscience) following the manufacturer’s operational guidelines. The Seahorse XF Glycolysis Stress Test Kit and Seahorse XF Cell Mito Stress Test Kit (Agilent Technologies) were employed for ECAR and OCR measurements, respectively. Cells were plated in Seahorse XF 96-well cell culture microplates at 10,000 cells per well and allowed to adhere for 10 h. For ECAR profiling, glucose, the oxidative phosphorylation inhibitor oligomycin, and the glycolysis inhibitor 2-DG were injected sequentially into each well at predefined time points to perturb metabolic flux. For OCR profiling, oligomycin (ATP synthase inhibitor), FCCP (oxidative phosphorylation uncoupler), and rotenone/antimycin A (complex I/III inhibitors) were injected sequentially to assess mitochondrial respiratory function. Data were acquired using Seahorse XF-96 Wave software and normalized to cell count for inter-sample comparison.
In vivo animal experiments
Animal experiments were performed following the ARRIVE guidelines and approved by the Institutional Animal Care and Use Committee (IACUC) of Chinese PLA General Hospital. All animal ethics were also taken care of in accordance with the relevant guidelines and regulations. For tumor xenograft models, 1 × 107 RKO cells (stably infected with different constructs) were subcutaneously injected into the right dorsal region of NCG mice (purchased from GemPharmatech). Tumor growth was monitored by measuring the longest (L) and shortest (W) tumor diameters at predefined time points using a digital caliper. Tumor volume was calculated using the formula: volume (mm³) = (L × W²)/2. At the experimental endpoint, mice were humanely euthanized according to IACUC guidelines. Tumor tissues were excised, weighed immediately, and snap-frozen in liquid nitrogen for subsequent molecular analysis.
Clinical samples and immunohistochemistry (IHC)
Ten pairs of CRC and adjacent normal tissues used for RNA-seq were collected from the Chinese PLA General Hospital with written informed consent from all patients. This study was approved by the Institutional Ethics Committee of the Chinese PLA General Hospital (approval number: S2022-148-01) and conducted in strict accordance with the Declaration of Helsinki. Colon cancer tissue microarrays (TMA) were obtained from Shanghai Outdo Biotech, encompassing 87 CRC tissue cores and 83 paired adjacent normal tissue cores. IHC staining of formalin-fixed paraffin-embedded (FFPE) sections was conducted following the standard protocol according to the manufacturer’s instructions. Briefly, TMA slides were deparaffinized with xylene and rehydrated through a series of graded ethanol solutions. Heat-mediated antigen retrieval was performed using citrate buffer (pH 6.0) in a pressure cooker for 10 min. Slides were incubated with 3% hydrogen peroxide for 15 min at room temperature to quench endogenous peroxidase activity. Nonspecific binding was blocked with 5% bovine serum albumin (BSA) for 30 min. Slides were then probed with primary antibodies (targeting PMM2) overnight at 4 °C, followed by incubation with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature. Diaminobenzidine (DAB) was used as the chromogenic substrate for visualization of immunoreactivity. Nuclei were counterstained with hematoxylin, and slides were dehydrated, cleared, and mounted with neutral resin. Two pathologists blinded to clinical data independently evaluated PMM2 expression using the H-scoring system (percentage of positive cells × staining intensity). Samples were categorized as “low PMM2” (H-score ≤ 6) or “high PMM2” (H-score > 6) based on inter-observer consensus.
Human CRC organoid construction and culture
Fresh human colorectal tumor tissues were first washed twice with ice-cold PBS containing 1% Penicillin/Streptomycin to remove blood and necrotic debris, then transferred to a sterile environment and minced into ~1 mm3 fragments using sterile scissors. The minced tissues were digested in a solution containing 2.5 mg/ml collagenase IV (Sigma), 0.1 mg/ml DNase I (Sigma), and 2.5 mg/ml Dispase at 37 °C with gentle shaking for 30–45 min until dispersed. The resulting suspension was filtered through a 100 μm nylon cell strainer to remove undigested tissue clumps, then centrifuged at 2000 rpm for 5 min at 4 °C. The pellet was resuspended and washed twice with ice-cold advanced DMEM/F12 (Invitrogen) to obtain a single-cell suspension. The cells were mixed with ice-cold Matrigel (Corning) at a 1:1 ratio, seeded into 48-well plates (50–100 μl per well), and incubated at 37 °C for 15–20 min to allow Matrigel solidification. Following solidification, CRC organoid medium (composed of advanced DMEM/F12 supplemented with 1% Penicillin/Streptomycin, 4 mM L-glutamine, 10 mM HEPES, 1:50 B-27 supplement, 1:100 N-2 supplement, 1.25 mM N-acetylcysteine, 10 mM nicotinamide, 10 nM (Leu15)-gastrin I human, 50 ng/ml recombinant human EGF, 100 ng/ml recombinant human Noggin, 25 ng/ml recombinant human HGF, 10 μM forskolin, 5 μM A83-01, 500 ng/ml recombinant human R-spondin 1, 300 ng/ml recombinant human Wnt3a, and 10 μM Y-27632) was added to each well. Organoids were cultured at 37 °C with 5% CO2, with medium refreshed every 2 days.
For passaging, when organoids reached 100–200 μm in diameter (typically after 7–10 days of culture, at 70–80% confluency), the medium was aspirated, and the Matrigel layer was gently washed twice with ice-cold PBS. Organoids were then dissociated by adding 200 μl of 0.25% TrypLE or Accutase per well, incubating at 37 °C for 5–10 min, and terminating digestion with an equal volume of medium containing 10% FBS. The suspension was centrifuged at 1500 rpm for 5 min at 4 °C, the pellet was resuspended in advanced DMEM/F12, and the cells were mixed with Matrigel (1:1 ratio) before being seeded into new 48-well plates at a 1:3-1:6 dilution. Fresh medium was added after Matrigel solidification, and culture was continued under standard conditions.
Statistical analysis
Data normality was evaluated using the Shapiro-Wilk test, and variance homogeneity was verified via Levene’s test. Data are presented as mean ± standard deviation (SD) unless specified otherwise. Student’s t test was applied for pairwise comparisons between two groups, while one-way analysis of variance (ANOVA) was employed to assess differences among three or more groups. Survival curves were generated using the Kaplan-Meier method, with inter-group differences evaluated by the log-rank test. Associations between PMM2 expression levels and clinical parameters were analyzed using Spearman’s rank correlation. All statistical computations were executed using SPSS 20.0 and GraphPad Prism 9 software. Statistical significance was defined as P < 0.05. All the experiments in vitro were performed in triplicate and repeated three times.
Ethics statements
Tissue arrays (HColA180Su18) were obtained from Shanghai Outdo Biotech Company. All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of Chinese PLA General Hospital. Ten pairs of CRC and adjacent normal tissues used for RNA-seq were collected from the Chinese PLA General Hospital with written informed consent from all patients. This study was approved by the Institutional Ethics Committee of the Chinese PLA General Hospital (approval number: S2022-148-01) and conducted in strict accordance with the Declaration of Helsinki.
Results
PMM2 knockdown inhibits the proliferation, migration, and invasion of CRC cells
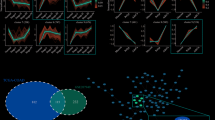
To elucidate molecular drivers of CRC progression, we performed transcriptomic profiling of 10 CRC tissues and matched normal tissues. Using stringent criteria (|log2FC|>1, FDR < 0.05), differential expression analysis revealed 10,829 upregulated and 6594 downregulated genes in CRC. Among these, PMM2 emerged as a top upregulated gene (log2FC = 2.1, FDR = 0.003), alongside known oncogenes such as PBX1, ZEB2, and TWIST1 (Fig. 1A). Notably, ZEB2 and TWIST1 have been implicated in epithelial-mesenchymal transition (EMT) and metastasis in CRC [19,20,21]. To validate PMM2’s functional relevance, we knocked down candidate genes in RKO CRC cells using lentiviral shRNAs. PMM2 knockdown (shPMM2-1/2/3) caused the most pronounced inhibition of proliferation (Fig. 1B), prompting focused investigation of PMM2.
A Volcano plot showing differentially expressed genes in CRC tissues vs. matched normal tissues (|log2FC|>1, FDR < 0.05). PMM2 is highlighted as a top upregulated gene. B Proliferation of RKO cells transfected with shRNAs targeting indicated genes or control shRNA (shCtrl), measured by CCK-8 assay. C RT-qPCR analysis of PMM2 mRNA levels in CRC cell lines (RKO, HT-29, HCT-116, Caco-2, HT-15) and normal rectal epithelial cell line NCM460. D Proliferation of HCT-116 cells with PMM2 knockdown using PMM2 shRNA-1 (shPMM2-1) and shRNA-2 (shPMM2-2), assessed by CCK-8 assay. E Colony formation assay in HCT-116 cells with PMM2 knockdown. F Flow cytometry analysis of apoptosis in HCT-116 cells with PMM2 knockdown. G Wound healing assay showing migration of HCT-116 cells with PMM2 knockdown. H Transwell invasion assay in HCT-116 cells with PMM2 knockdown. Data are mean ± SD (n = 3). *p < 0.05, **p < 0.01 vs. shCtrl.
PMM2 expression was systematically analyzed across CRC cell lines (RKO, HT-29, HCT-116, Caco-2, HT-15) and the normal rectal epithelial cell line NCM460. PMM2 mRNA level was significantly elevated in CRC cells, with RKO and HCT-116 showing the highest expression (Fig. 1C). Subsequently, three shRNA sequences targeting PMM2 were designed, and selected the most efficient knockdown sequence was selected, shPMM2-1, via qPCR and Western blot screening (Fig. S1A, B). Using green fluorescent protein-labeled lentiviral infection efficiency evaluation (Fig. S1C), along with qPCR and Western blot validation (Fig. S1D, E), effective PMM2 knockdown was confirmed in HCT 116 and RKO cells, laying a solid foundation for further functional studies.
Cell proliferation assay demonstrated that PMM2 knockdown significantly inhibited the proliferation capacity of CRC cells (Figs. 1D and S2A). Additionally, colony formation assay further confirmed the inhibitory effect of PMM2 knockdown on the colony-forming ability of CRC cells (Figs. 1E and S2B), underscoring the critical role of PMM2 in sustaining CRC cell proliferation potential. Furthermore, flow cytometry analysis revealed that PMM2 knockdown potentially promoted apoptosis in CRC cells (Figs. 1F and S2C), indicating that PMM2 might be involved in the malignant progression of CRC through the regulation of apoptotic mechanisms. Finally, wound healing and Transwell assays demonstrated that PMM2 knockdown markedly inhibited the migration (Figs. 1G and S2D) and invasion (Figs. 1H and S2E) abilities of CRC cells, further corroborating the pivotal role of PMM2 in the malignant migration and invasion of CRC cells. In summary, these results highlight the crucial role of PMM2 in regulating malignant phenotypes such as proliferation, apoptosis, migration, and invasion in CRC cells.
PMM2 enhances glycolytic activity to promote CRC progression
To elucidate the metabolic reprogramming mediated by PMM2 in CRC, we evaluated its role in glucose metabolism. PMM2 knockdown in HCT-116 and RKO cells significantly repressed aerobic glycolytic activity, evidenced by reduction in glucose uptake (Figs. 2A and S3A), decrease in ATP production (Figs. 2B and S3B), and inhibition of lactate (Figs. 2C and S3C). This metabolic shift was further confirmed by reduced extracellular acidification rate (ECAR, Figs. 2D and S3D) and elevated oxygen consumption rate (OCR, Figs. 2E and S3E). Furthermore, knockdown of PMM2 decreased the expression levels of PKM2 and LDHA (Figs. 2F and S3F), two key enzymes in glycolysis, indicating a metabolic shift toward glycolysis.
A Glucose uptake in HCT-116 cells with PMM2 knockdown (shPMM2-1). B ATP production in HCT-116 cells with PMM2 knockdown. C Lactate production in HCT-116 cells with PMM2 knockdown. D Extracellular acidification rate (ECAR) measured by Seahorse XFe analyzer in HCT-116 cells with PMM2 knockdown. E Oxygen consumption rate (OCR) in HCT-116 cells with PMM2 knockdown. F Western blot analysis of glycolytic enzymes (PKM2, LDHA) in HCT-116 cells with PMM2 knockdown. Effects of 2-deoxy-D-glucose (2-DG, 5 mM) on glucose uptake (G), ATP production (H), ECAR (I), and OCR (J) in PMM2-overexpressing HCT-116 cells. K Western blot analysis of PKM2 and LDHA in PMM2-overexpressing HCT-116 cells treated with 2-DG. CCK-8 (L) and colony formation (M) assays in PMM2-overexpressing HCT-116 cells with or without 2-DG. Data are mean ± SD (n = 3). **p < 0.01 vs. corresponding control.
Notably, the glycolytic inhibitor 2-deoxy-D-glucose (2-DG) significantly inhibited PMM2-driven glycolysis: glucose uptake and ATP levels were decreased, respectively (Figs. 2G, H and S3G, H), while ECAR (Figs. 2I and S3I) and OCR (Figs. 2J and S3J) normalized to baseline levels. Additionally, the increase in PKM2 and LDHA protein levels driven by PMM2 was also obviously restrained by 2-DG (Figs. 2K and S3K). Functional assays revealed that 2-DG co-treatment notably attenuated PMM2-mediated cell proliferation and colony formation in HCT-116 and RKO cells, respectively (Figs. 2L, M and S3L, M).
Since PMM2 is a critical enzyme that catalyzes the reversible isomerization of Man-6-P to Man-1-P, we investigated whether the catalytic activity of PMM2 is associated with its regulation of tumor glycolytic activity. First, a catalytically inactive PMM2 mutant (T171R/I174R) was constructed [22]. The results showed that both PMM2 wild-type and the mutant could promote glycolytic activity in HCT-116 and RKO cells (Fig. S4A–C) and increase the expression level of LDHA (Fig. S4D), indicating that PMM2-driven tumor glycolysis is independent of its catalytic activity. These findings establish PMM2 as a critical regulator of CRC glycolytic reprogramming, linking it to metabolic adaptation and tumorigenic potential.
PMM2-TRIM28 interaction is indispensable for metabolic adaptation and tumorigenic potential in CRC
Bioinformatics analysis using the STRING database predicted a robust interaction between PMM2 and tripartite motif-containing protein 28 (TRIM28), a multifunctional E3 ubiquitin ligase and transcriptional regulator implicated in tumorigenesis [23,24,25]. To validate this interaction, co-immunoprecipitation (Co-IP) assays were performed in CRC cells. As shown in Fig. 3A, B PMM2 or TRIM28-specific antibodies effectively immunoprecipitated another endogenous protein, confirming their physical association. Structural analysis further revealed three functional domains in PMM2: domain1 (188-246 aa), domain2 (86-187 aa), and domain3 (1-85 aa). Domain deletion mutants (Δ1, Δ2, Δ3) were generated and transiently transfected into HEK293T cells (Fig. S5A). Exogenous Co-IP demonstrated that only the PMM2-Δ1 mutant lost interaction with TRIM28, identifying domain 1 as the critical binding interface (Fig. 3C).
Co-immunoprecipitation (Co-IP) assays in HCT-116 and RKO cells showing interaction between endogenous PMM2 and TRIM28 (A: IP with anti-PMM2; B: IP with anti-TRIM28). C Co-IP of HA-tagged TRIM28 with FLAG-tagged full-length (FL) or domain-deleted PMM2 (Δ1, Δ2, Δ3) in HEK293T cells. Glucose uptake (D), Lactate production (E), and ECAR (F) in HCT-116 shCtrl or shPMM2 cells transfected with NC, PMM2-FL or PMM2-Δ1. G Western blot analysis of PKM2, LDHA, and PMM2 in HCT-116 cells grouped as in (D). CCK-8 (H) and wound healing (I) assays in HCT-116 cells grouped as in (D). Data are mean ± SD (n = 3). **p < 0.01 vs. corresponding control.
Functional rescue experiments were performed by transfecting PMM2-FL or PMM2-Δ1 plasmids into PMM2-knockdown cells. The results showed that PMM2 knockdown in HCT-116 and RKO cells significantly decreased aerobic glycolytic activity, while transfection of PMM2-FL into these knockdown cells restored glycolytic activity, whereas transfection of PMM2-Δ1 failed to do so (Figs. 3D–F and S5B–D). Furthermore, transfection of PMM2-FL, but not PMM2-Δ1, into PMM2-knockdown cells rescued the expression levels of PKM2 and LDHA (Figs. 3G and S5E). Notably, PMM2-Δ1 also lost its pro-proliferative and pro-migratory activities (Figs. 3H, I and S5F, G). These findings indicate that domain 1, a key domain of PMM2, is crucial for the interaction between PMM2 and TRIM28 and for promoting CRC progression.
PMM2 drives TRIM28 nuclear translocation and KIFC3 transcription
Bioinformatics analysis using the GTRD database predicted that TRIM28, an E3 ubiquitin ligase and transcriptional regulator, may bind to the promoter region of KIFC3—a gene overexpression and poor prognosis in CRC (Fig. 4A, B). Further using the hTFtarget prediction tool, three potential binding sites were obtained (site 1, site 2, and site 3, Fig. S6A). To validate this interaction, dual-luciferase reporter assays were performed using KIFC3 promoter wild-type and mutant constructs. TRIM28 overexpression significantly enhanced luciferase activity in HCT-116 and RKO cells transfected with the wild-type KIFC3 promoter (Figs. 4C and S6B), but not with a mutant promoter lacking the TRIM28-binding site (site 1). Site-directed mutagenesis revealed that site 1 (positions –780 to –765) was critical for TRIM28-mediated transcriptional activation, as mutations here abolished luciferase activity (Figs. 4C and S6B). Chromatin immunoprecipitation (ChIP) assay further demonstrated that TRIM28 specifically enriched at KIFC3 promoter site 1 in CRC cells (Figs. 4D and S6C), confirming its direct regulatory role.
A KIFC3 mRNA levels in CRC tissues vs. normal tissues (TCGA dataset). B Kaplan–Meier survival analysis of CRC patients with high vs. low KIFC3 expression (TCGA dataset). C Dual-luciferase reporter assay in HCT-116 cells transfected with wild-type (WT) or mutant (Mut1, Mut2, Mut3) KIFC3 promoter constructs, with or without TRIM28 overexpression. D Chromatin immunoprecipitation (ChIP) assay showing TRIM28 enrichment at the KIFC3 promoter (site 1, site 2, or site 3) in HCT-116 cells. E Western blot of TRIM28 in nuclear/cytoplasmic fractions of HCT-116 cells with PMM2 knockdown (Histone H3 and GAPDH as nuclear/cytoplasmic markers). F Immunofluorescence staining of TRIM28 in HCT-116 cells with PMM2 overexpression. Images of TRIM28 (green) and nuclei (blue) were merged. Scale bar, 20 μm. RT-qPCR (G) and Western blot (H) analysis of KIFC3 expression in HCT-116 cells with PMM2 knockdown. I ChIP assay showing PMM2 overexpression enhances TRIM28 binding to the KIFC3 promoter (site 1). J Dual-luciferase assay in HCT-116 cells co-transfected with PMM2 shRNA, TRIM28, and KIFC3 promoter constructs. Western blot analysis indicates TRIM28 and PMM2 expression. Data are mean ± SD (n = 3). **p < 0.01 vs. corresponding control. ns, not significant.
Given PMM2’s interaction with TRIM28, we investigated whether PMM2 regulates TRIM28’s subcellular localization. Western blot analysis of nuclear and cytoplasmic fractions revealed that PMM2 knockdown reduced nuclear TRIM28 levels (Figs. 4E and S6D). Conversely, immunofluorescence staining assay showed that PMM2 overexpression increased nuclear TRIM28 levels (Figs. 4F and S6E), suggesting PMM2 facilitates the nuclear translocation of TRIM28. Furthermore, PMM2 depletion suppressed KIFC3 mRNA and protein expression, respectively (Figs. 4G, H and S6F, G). Moreover, ChIP assay demonstrated that PMM2 overexpression significantly enhanced the association of TRIM28 with the KIFC3 promoter (Figs. 4I and S6H). Dual-luciferase reporter assay showed that TRIM28 increased KIFC3 reporter activity. PMM2 knockdown decreased KIFC3 reporter activity and inhibited TRIM28-mediated promotion of KIFC3 promoter activity. These effects were abolished when binding site 1 was mutated, indicating that PMM2 overexpression promotes the specific binding of TRIM28 to the KIFC3 promoter (Figs. 4J and S6I). In conclusion, PMM2 promotes the nuclear translocation of TRIM28 and binds to site 1 on the KIFC3 promoter in CRC cells, thereby regulating the transcriptional expression of KIFC3.
E2F4 mediates PMM2/TRIM28-regulated KIFC3 transcription
Since TRIM28 is generally recognized as a transcriptional co-regulator that modulates downstream gene expression by recruiting transcription factors, we further sought to identify the key transcription factors involved in TRIM28-mediated KIFC3 regulation. First, we performed immunoprecipitation combined with mass spectrometry (IP-MS) to initially screen proteins interacting with TRIM28. Further selection based on large fold changes between experimental and control groups and high protein abundance yielded two candidate transcription factors: E2F4 and SMARCC1 (Fig. 5A). Using the hTFtarget prediction tool, we found that E2F4 could bind to site 1 of the KIFC3 promoter, prompting us to focus subsequent studies on E2F4. Co-IP assays validated the interaction between TRIM28 and E2F4, as specific antibodies against either protein effectively immunoprecipitated the endogenous counterpart (Figs. 5B, C and S7A, B). RT-qPCR and Western blot results showed that E2F4 significantly upregulated KIFC3 mRNA and protein levels in HCT-116 and RKO cells (Figs. 5D, E and S7C, D). Moreover, dual-luciferase reporter assays revealed that E2F4 enhanced the activity of the wild-type KIFC3 promoter in 293T cells but had no effect on the promoter with site 1 mutation (Fig. 5F). ChIP assays further confirmed that E2F4 was specifically recruited to site 1 of the KIFC3 promoter, verifying its direct regulatory role (Figs. 5G and S7E).
A Immunoprecipitation combined with mass spectrometry (IP-MS) analysis of TRIM28-interacting proteins in HCT-116 cells. Co-immunoprecipitation (Co-IP) assays in HCT-116 cells validating the interaction between endogenous TRIM28 and E2F4 (B: IP with anti-TRIM28; C: IP with anti-E2F4). RT-qPCR (D) and Western blot (E) analysis of KIFC3 mRNA and protein levels in HCT-116 cells with E2F4 overexpression. F Dual-luciferase reporter assay in 293 T cells transfected with WT or Mut1 KIFC3 promoter constructs, with or without E2F4 overexpression. G ChIP assay showing E2F4 enrichment at the KIFC3 promoter site 1 in HCT-116 cells. H Dual-luciferase reporter assay in 293 T cells co-transfected with PMM2, E2F4, and KIFC3 promoter constructs. I ChIP assay showing PMM2 overexpression enhances E2F4 binding to the KIFC3 promoter site 1 in HCT-116 cells. RT-qPCR (J) and Western blot (K) analysis of KIFC3 expression in HCT-116 cells with PMM2 overexpression and/or TRIM28 knockdown. L Dual-luciferase reporter assay in 293T cells co-transfected with PMM2, TRIM28 shRNA, E2F4 shRNA, and KIFC3 promoter constructs. M ChIP assay showing TRIM28 knockdown partially inhibits PMM2-induced E2F4 binding to the KIFC3 promoter site 1 in HCT-116 cells. Data are mean ± SD (n = 3). *p < 0.05, **p < 0.01 vs. corresponding control. ns, not significant.
Next, we investigated whether PMM2/TRIM28 modulates E2F4-mediated KIFC3 regulation. Dual-luciferase reporter assays showed that both PMM2 and E2F4 increased KIFC3 promoter activity in 293T cells, with an additive effect (Fig. 5H). ChIP assays demonstrated that PMM2 further promoted E2F4 recruitment to the KIFC3 promoter (Figs. 5I and S7F). Additionally, TRIM28 knockdown inhibited KIFC3 mRNA and protein expression and partially abrogated PMM2’s promoting effect on KIFC3 (Figs. 5J, K and S7G, H). In 293T cells, TRIM28 knockdown suppressed KIFC3 promoter activity and partially reduced PMM2’s enhancement of promoter activity; these effects were abolished when E2F4 was knocked down (Fig. 5L). ChIP assays yielded similar results: TRIM28 knockdown partially abrogated PMM2-induced E2F4 binding to the KIFC3 promoter (Figs. 5M and S7I). These findings indicate that E2F4 acts as a key transcription factor mediating TRIM28-dependent KIFC3 transcription, and PMM2 enhances this regulatory cascade by promoting TRIM28-E2F4 interaction and E2F4 recruitment to the KIFC3 promoter, thereby linking PMM2 to downstream glycolytic and tumorigenic pathways.
PMM2 accelerates CRC progression by enhancing glycolytic activity through KIFC3
Since PMM2 promotes KIFC3 expression and regulates CRC progression by modulating glycolytic activity, we investigated whether the PMM2/KIFC3 axis also regulates glycolysis, thereby affecting CRC progression. In HCT-116 and RKO cells, we developed models of KIFC3 knockdown and KIFC3 knockdown following PMM2 overexpression, with validations performed via qPCR and Western blot (Fig. S8A, B). Glycolysis-related experimental results revealed that KIFC3 knockdown significantly suppressed glycolytic activity in CRC cells, with decreased glucose uptake, ATP level, and lactic acid content (Figs. 6A–C and S8C–E), reduced ECAR (Figs. 6D and S8F), and increased OCR (Figs. 6E and S8G). In addition, knockdown of KIFC3 reduced the protein levels of PKM2 and LDHA in CRC cells (Figs. 6F and S8H). Notably, KIFC3 knockdown partially rescued the promoting effects of PMM2 overexpression on glycolysis activity (Figs. 6A–F and S8C–H). Moreover, the PMM2/KIFC3 exerted similar roles in cell proliferation and migration (Figs. 6G, H and S8I, J). These findings suggest that the function of PMM2 in CRC is partially dependent on KIFC3 expression, highlighting their synergistic relationship. Thus, these results reveal a novel mechanism by which PMM2 enhances glycolytic activity through KIFC3, thereby promoting the progression of CRC cells.
Glucose uptake (A), ATP production (B), and lactate production (C) in HCT-116 cells with PMM2 overexpression and/or KIFC3 knockdown. ECAR (D) and OCR (E) in HCT-116 cells with PMM2 overexpression and/or KIFC3 knockdown. F Western blot of PKM2, LDHA, PMM2, and KIFC3 in HCT-116 cells with PMM2 overexpression and/or KIFC3 knockdown. CCK-8 (G) and wound healing (H) assays in HCT-116 cells with PMM2 overexpression and/or KIFC3 knockdown. Data are mean ± SD (n = 3). *p < 0.05, **p < 0.01 vs. corresponding control.
In vivo mouse models and clinical significance of the PMM2/KIFC3 axis in regulating CRC progression
To validate the functional role of the PMM2/KIFC3 axis in CRC progression, we established subcutaneous xenograft models using RKO cells with PMM2 overexpression or KIFC3 knockdown. As expected, the PMM2 overexpression group exhibited significantly increased tumor volume compared to controls, while the KIFC3 knockdown group showed a marked reduction in tumor volume. Importantly, KIFC3 knockdown partially inhibited the tumor-promoting effect of PMM2 (Fig. 7A). Additionally, after the mice were sacrificed and the tumors were removed from the mice, the weight of the tumors showed the same trend (Fig. 7B). These results indicate that PMM2 promotes tumor growth through KIFC3. Immunohistochemical (IHC) staining of xenografts revealed elevated PMM2, KIFC3, Ki67, and glycolytic markers (LDHA, PKM2) in PMM2-overexpressing tumors, with KIFC3 knockdown reversing these trends (Fig. 7C). Western blot analysis further demonstrated that PMM2 overexpression upregulated KIFC3 and glycolytic enzymes (LDHA, PKM2), while KIFC3 depletion abolished these effects (Fig. 7D). These data suggest that PMM2 enhances tumor growth via KIFC3-mediated glycolysis.
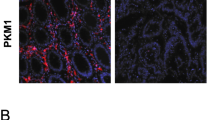
A Images and tumor growth curves of subcutaneous xenografts in NCG mice injected with RKO cells (NC + shCtrl, PMM2 + shCtrl, NC + shKIFC3, PMM2 + shKIFC3). B Tumor weights at endpoint (day 48). C Representative IHC staining of PMM2, KIFC3, Ki67, LDHA, and PKM2 in xenograft tumors. Scale bar, 100 μm. D Western blot of PMM2, KIFC3, LDHA, and PKM2 in xenograft tumors. The lower panel shows the relative protein expression level. E IHC staining of PMM2 in normal vs. CRC tissues (tissue microarray). Scale bar, 50 μm. F Kaplan–Meier survival analysis of CRC patients with high vs. low PMM2 expression (log-rank test). Data are mean ± SD (n = 6 mice/group). *p < 0.05, **p < 0.01 vs. corresponding control.
To establish the clinical significance of PMM2, tissue microarray combined with immunohistochemistry was assessed to evaluate PMM2 expression in normal and cancerous tissues from clinical CRC patients. IHC assay demonstrated that PMM2 expression was significantly elevated in CRC tissues (Fig. 7E). Moreover, patients were divided into high and low expression groups based on the median PMM2 expression in tumor samples. Kaplan-Meier survival analysis revealed that high PMM2 expression correlated with poorer OS (HR = 0.19, 95% CI: 0.09–0.37, p < 0.0001, Fig. 7F), highlighting PMM2 as a potential prognostic biomarker in CRC.
Next, we analyzed the expression difference of PMM2 between cancerous and adjacent normal tissues, as well as its correlation with KIFC3 expression, using public databases. The results showed that PMM2 expression was significantly higher in cancer tissues than in adjacent normal tissues in the TCGA-COAD dataset (Fig. S9A). However, PMM2 exhibited an extremely weak negative correlation with KIFC3 (Fig. S9B). Furthermore, in multiple GEO public datasets, PMM2 expression showed no correlation with KIFC3 (Fig. S9C), which was inconsistent with our expectations. This could be attributed to the fact that mRNA expression does not always align with protein expression, and the sample size available for analysis in some datasets is insufficient, among other reasons.
Additionally, in the tissue microarray, Mann-Whitney U tests identified a significant positive correlation between PMM2 expression and pathological grade (p = 0.003), lymphatic metastasis (p = 0.002), and clinical stage (p = 0.006) (Table S1). Consistently, PMM2 expression also positively correlated with advanced tumor grade (Spearman’s ρ = 0.320, p = 0.002), lymphatic metastasis (Spearman’s ρ = 0.336, p = 0.001) and clinical stage (Spearman’s ρ = 0.296, p = 0.005) (Table S2). These clinical correlations align with our mechanistic findings, positioning PMM2 as a biomarker for CRC prognosis and therapeutic targeting.
PMM2 regulates glycolytic activity and tumorigenicity in CRC patient-derived organoids
To elucidate the physiological relevance of PMM2 in the regulation of CRC progression, patient-derived organoid (PDO) models were established using tumor tissues obtained from six CRC patients (PDO-1 to PDO-6). The mRNA and protein expression levels of PMM2 and KIFC3 were subsequently examined in these six intestinal tumor organoids. qPCR analysis revealed that PMM2 mRNA expression was lowest in PDO-2 and relatively higher in PDO-5 and PDO-6. Similarly, KIFC3 mRNA expression was found to be lowest in PDO-1 and PDO-2, while elevated levels were observed in PDO-4 (Fig. S10A, B). Western blotting results indicated that PMM2 protein levels were lowest in PDO-2 and PDO-3 and highest in PDO-6, whereas KIFC3 protein expression was lowest in PDO-1 and PDO-2 and relatively higher in PDO-4 (Fig. 8A). Cell proliferation, assessed by CCK8 assay, demonstrated the slowest proliferation rate in PDO-2, while higher proliferation rates were observed in PDO-4, PDO-5, and PDO-6 (Fig. 8B). Based on these findings, PDO-2 and PDO-6 were selected for further investigation. Measurements of ATP and lactate content, as well as colony formation assays, showed that PDO-6 exhibited higher ATP and lactate levels and greater colony formation capacity compared to PDO-2 (Fig. S10C–E).
A Western blot of PMM2 and KIFC3 in PDOs (PDO-1 to PDO-6). B CCK-8 proliferation assay in PDOs. RT-qPCR (C) and Western blot (D) of PMM2 and KIFC3 in PDO-2 with PMM2 overexpression. ATP (E) and lactate (F) production in PDO-2 with PMM2 overexpression. CCK-8 (G) and colony formation (H) assays in PDO-2 with PMM2 overexpression. Scale bar, 200 μm. RT-qPCR (I) and Western blot (J) of PMM2 and KIFC3 in PDO-6 with PMM2 knockdown (shPMM2). ATP (K) and lactate (L) production in PDO-6 with shPMM2. CCK-8 (M) and colony formation (N) assays in PDO-6 with shPMM2. Scale bar, 200 μm. Data are mean ± SD (n = 3). *p < 0.05, **p < 0.01 vs. corresponding control.
To further explore the role of PMM2, lentiviral overexpression of PMM2 was performed in PDO-2. Following infection with PMM2-expressing lentivirus, both mRNA and protein levels of PMM2 were significantly upregulated, confirming successful construction of the PMM2-overexpressing model. Concurrently, the expression of KIFC3 at both mRNA and protein levels was also upregulated, consistent with findings from in vitro cell models (Fig. 8C, D). Enhanced production of ATP and lactate was observed in the PMM2-overexpressing organoids (Fig. 8E, F). In vitro CCK8 and colony formation assays demonstrated that PMM2 overexpression significantly promoted proliferation and clonogenic capacity of the organoids (Fig. 8G, H). Conversely, PMM2 knockdown was conducted in PDO-6 using shRNA-expressing lentivirus. Following infection, a significant downregulation of PMM2 at both mRNA and protein levels was confirmed, indicating successful establishment of the PMM2 knockdown model. Simultaneously, a marked reduction in KIFC3 expression was observed at both the transcript and protein levels (Fig. 8I, J). ATP and lactate assays revealed that PMM2 knockdown led to a significant decrease in ATP and lactate production (Fig. 8K, L). In vitro analyses using CCK8 and colony formation assays further showed that suppression of PMM2 markedly inhibited proliferation and colony formation in the organoids (Fig. 8M, N). These findings indicate that PMM2 similarly regulates tumor glycolytic activity and contributes to the progression of CRC in intestinal tumor organoids.
Discussion
Colorectal cancer (CRC), a prevalent malignancy of the digestive tract, continues to exhibit persistently high global incidence and mortality rates. Current therapeutic strategies—including surgical resection, chemotherapy, and targeted therapies—have shown partial clinical efficacy but face substantial challenges due to interindividual genetic heterogeneity, which leads to marked variability in treatment outcomes. Despite progress, our understanding of CRC pathogenesis and its molecular regulatory networks remains incomplete. In this study, we identified significant overexpression of PMM2 in CRC tissues, which strongly correlated with advanced tumor stage, elevated metastatic potential, and poor patient prognosis. Functional analyses revealed that PMM2 drives glycolytic metabolic reprogramming, directly potentiating malignant phenotypes in CRC cells, including enhanced proliferation, migration, and metastatic capacity in both in vitro and in vivo models. Mechanistically, PMM2 interacts with the transcriptional co-regulator TRIM28 to form a functional complex that recruits E2F4 to activate KIFC3 expression. This activation reinforces tumor glycolytic flux, meeting metabolic adaptive demands and driving tumor progression. To our knowledge, this work provides the first evidence that PMM2-mediated metabolic remodeling governs CRC progression, expanding the molecular mechanistic framework of CRC pathogenesis. The PMM2-TRIM28 interaction interface and downstream KIFC3 signaling axis not only establish PMM2 as a potential prognostic biomarker but also reveal novel therapeutic targets for precision interventions against metabolic reprogramming. Therapeutic strategies targeting this molecular axis may hold promise for suppressing CRC progression and overcoming chemoresistance, with significant translational implications.
Emerging evidence has demonstrated the multifaceted oncogenic role of PMM2 across malignancies. Notably, PMM2 is also the causal gene for phosphomannomutase 2-congenital disorders of glycosylation (PMM2-CDG), a rare metabolic disease caused by loss-of-function mutations in PMM2’s catalytic domain that impair N-glycosylation [9, 10]. Intriguingly, our study reveals that PMM2’s oncogenic function in CRC is independent of its catalytic activity (as both wild-type and catalytically inactive PMM2 mutants promote glycolysis), indicating a dual functional paradigm: PMM2 drives CDG via catalytic deficiency but exerts oncogenic effects through non-catalytic protein–protein interactions (e.g., with TRIM28), uncoupling its metabolic glycosylation role from its tumor-promoting activity. In breast cancer, PMM2 is overexpressed in tumor tissues compared to normal counterparts, positively correlates with ERα status, and is a direct target of the tumor-suppressive miR-451a, positioning it as a promising therapeutic target [12, 13]. In renal cell carcinoma, PMM2 overexpression correlates with advanced disease progression and unfavorable clinical outcomes, highlighting its therapeutic potential [14]. In cervical squamous cell carcinoma, PMM2 expression patterns exhibit stage-dependent alterations and functional involvement in carcinogenesis [15]. Within CRC biology, prior studies have reported PMM2 upregulation in tumor compared to normal tissues, its participation in copper-dependent cell death pathways, and its prognostic significance (e.g., elevated PMM2 levels associate with high-risk patient stratification and dismal survival) [16,17,18]. Our findings align with these observations, confirming PMM2 overexpression in CRC specimens and its strong association with advanced tumor staging and poor prognosis. Critically, we extend these insights by demonstrating that PMM2 promotes tumor growth and metastatic dissemination through metabolic reprogramming, solidifying its role as a key oncogenic driver in CRC pathogenesis. Mechanistically, we identified a PMM2-TRIM28 protein complex that requires the transcription factor E2F4 to directly bind the KIFC3 promoter. E2F4 is a core member of the E2F transcription factor family, which primarily regulates cell cycle progression, proliferation, and differentiation by controlling the expression of genes involved in DNA synthesis and cell division [26,27,28]. The PMM2-TRIM28 complex recruits E2F4 to the KIFC3 promoter, forming a ternary regulatory module that amplifies KIFC3 transcription. This finding clarifies the missing link in the PMM2-TRIM28-KIFC3 axis, where E2F4 acts as the key intermediary transcription factor bridging TRIM28’s co-regulatory function to KIFC3’s transcriptional activation, revealing a novel regulatory axis in metabolic adaptation.
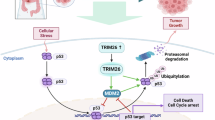
The dual transcriptional regulatory capacity of TRIM28 is well-documented. Agarwal et al. demonstrated that TRIM28 phosphorylation enables hTERT activation by competitively displacing TRIM24 from chromatin [25], whereas Li et al. reported its essential co-repressor function in ZNF331-mediated tumor suppression in head and neck squamous cell carcinoma [29]. Additionally, TRIM28 can act as an E3 ubiquitin ligase, regulating substrate protein ubiquitination and downstream functions. The ubiquitin-proteasome system (UPS) is a pivotal regulator of oncoprotein turnover and tumor suppressor stability, and UPS dysfunction drives malignant progression and presents therapeutic vulnerabilities. For example, Wang et al. identified MAGI3 as a novel E3 ligase that targets c-Myc for proteasomal degradation in CRC [30], and Liu et al. demonstrated hypoxia-dependent regulation of HIF-1α stability via NEDD4L-mediated ubiquitination in CRC models [31]. Notably, TRIM28 itself exhibits E3 ligase activity, as shown by its interaction with BCL2A1 to modulate anti-apoptotic signaling through ubiquitination in melanoma [32]. While our study clarifies the PMM2-TRIM28 interaction in metabolic regulation, the potential E3 ligase activity of TRIM28 toward PMM2 remains unexplored. This intriguing possibility that TRIM28 might post-translationally modify PMM2 through ubiquitination warrants dedicated investigation to fully delineate the complex regulatory interplay within this oncogenic axis.
KIFC3, a member of the C-terminal kinesin-14 subfamily (Kinesin superfamily proteins), is encoded by the KIFC3 gene (chromosome 16q21). As a minus-end microtubule-dependent motor protein, KIFC3 has recently been implicated in tumor progression via activation of the PI3K/AKT pathway, exhibiting oncogenic properties in CRC [33], hepatocellular carcinoma [34], and non-small cell lung cancer [35]. Among the three mammalian kinesin-14 members (KIFC1, KIFC2, KIFC3), KIFC1 drives aerobic glycolysis in endometrial and breast cancers [36, 37], suggesting KIFC3 may similarly regulate tumor glycolytic reprogramming. In tumors, aerobic glycolysis-mediated metabolic reprogramming promotes proliferation, immune escape, and metastasis via lactate production and microenvironmental acidification, acting as a key driver of malignant progression. Our mechanistic studies reveal that the PMM2-TRIM28 complex amplifies KIFC3-mediated aerobic glycolysis, manifesting as enhanced glucose uptake, elevated ATP levels, increased lactic acid production, heightened extracellular acidification rate (ECAR), and reduced oxygen consumption rate (OCR). The PMM2-KIFC3 axis thus emerges as a pivotal regulator of glycolytic flux in CRC, providing the first evidence of KIFC3’s role in governing aerobic glycolysis within CRC biology. These findings expand the molecular network underlying CRC progression, demonstrating that the PMM2-KIFC3 axis critically drives tumor growth and metastasis through metabolic reprogramming. This discovery not only enriches our understanding of KIFC3’s oncogenic mechanisms but also highlights metabolic interventions targeting this axis as promising therapeutic avenues for CRC management.
Collectively, our findings establish PMM2 overexpression as a critical prognostic indicator in CRC, strongly associated with advanced tumor staging and reduced survival rates. Mechanistically, PMM2 forms a functional complex with TRIM28 to amplify KIFC3 transcription and expression. The PMM2-KIFC3 axis drives tumor progression by enhancing aerobic glycolytic activity, thereby modulating CRC growth and metastatic dissemination. These insights position PMM2 as a promising therapeutic target, with interventions targeting PMM2-KIFC3-mediated metabolic reprogramming representing a novel strategy for CRC treatment. While direct PMM2 inhibitors are currently limited [11, 38], our identification of PMM2’s domain 1 (188-246 aa) as the critical TRIM28-binding interface opens alternative avenues: peptide inhibitors blocking this domain or PROTAC molecules degrading PMM2 could disrupt the PMM2-TRIM28-E2F4 complex, inhibiting downstream KIFC3-mediated glycolysis. Additionally, targeting E2F4’s DNA-binding activity or KIFC3’s glycolytic regulatory function may serve as complementary strategies to suppress the oncogenic axis. However, this study has several limitations. First, the limited cohort size necessitates validation in expanded clinical populations to strengthen the robustness of our conclusions. Second, while we characterized the PMM2-TRIM28 interaction, the potential E3 ubiquitin ligase activity of TRIM28 toward PMM2 remains unexplored. Given TRIM28’s documented role in ubiquitination-dependent regulation of other substrates, further investigation is required to determine whether TRIM28-mediated post-translational modifications modulate PMM2 stability or activity. Addressing these questions will refine our understanding of this oncogenic pathway and inform the development of targeted therapeutic strategies.
Data availability
The datasets in the current study are available from the corresponding author on reasonable request.
References
Eng C, Jácome AA, Agarwal R, Hayat MH, Byndloss MX, Holowatyj AN, et al. A comprehensive framework for early-onset colorectal cancer research. Lancet Oncol. 2022;23:e116–e128.
Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229–63.
Cai W, Zhong Y, Zhao J, Mao J, Hua L, He X, et al. TRIM24 drives colorectal cancer progression via CAVIN2 degradation and ERK/RhoA pathway activation. Cell Investig. 2025;1:100033.
Shin AE, Giancotti FG, Rustgi AK. Metastatic colorectal cancer: mechanisms and emerging therapeutics. Trends Pharmacol Sci. 2023;44:222–36.
Biller LH, Schrag D. Diagnosis and treatment of metastatic colorectal cancer: a review. JAMA. 2021;325:669–85.
Jung G, Hernández-Illán E, Moreira L, Balaguer F, Goel A. Epigenetics of colorectal cancer: biomarker and therapeutic potential. Nat Rev Gastroenterol Hepatol. 2020;17:111–30.
Zhang N, Ng AS, Cai S, Li Q, Yang L, Kerr D. Novel therapeutic strategies: targeting epithelial-mesenchymal transition in colorectal cancer. Lancet Oncol. 2021;22:e358–e368.
Radenkovic S, Budhraja R, Klein-Gunnewiek T, King AT, Bhatia TN, Ligezka AN, et al. Neural and metabolic dysregulation in PMM2-deficient human in vitro neural models. Cell Rep. 2024;43:113883.
Ligezka AN, Radenkovic S, Saraswat M, Garapati K, Ranatunga W, Krzysciak W, et al. Sorbitol is a severity biomarker for pmm2-cdg with therapeutic implications. Ann Neurol. 2021;90:887–900.
Gallego D, Serrano M, Córdoba-Caballero J, Gámez A, Seoane P, Perkins JR, et al. Transcriptomic analysis identifies dysregulated pathways and therapeutic targets in PMM2-CDG. Biochim Biophys Acta Mol Basis Dis. 2024;1870:167163.
Gamez A, Serrano M, Gallego D, Vilas A, Perez B. New and potential strategies for the treatment of PMM2-CDG. Biochim Biophys Acta Gen Subj. 2020;1864:129686.
Cipolletti M, Acconcia F. PMM2 controls ERα levels and cell proliferation in ESR1 Y537S variant expressing breast cancer cells. Mol Cell Endocrinol. 2024;584:112160.
Hina SS, E. Down-regulation of miRNA 451a promotes many oncogenic signaling pathways in breast cancer patients. Indian J Biochem Biophys. 2023;60:467–77.
Yamada Y, Arai T, Sugawara S, Okato A, Kato M, Kojima S, et al. Impact of novel oncogenic pathways regulated by antitumor miR-451a in renal cell carcinoma. Cancer Sci. 2018;109:1239–53.
Yang C, Ren J, Li B, Zhang D, Ma C, Cheng C, et al. Identification of clinical tumor stages related mRNAs and miRNAs in cervical squamous cell carcinoma. Pathol Res Pract. 2018;214:1638–47.
Cui Z, Sun G, Bhandari R, Lu J, Zhang M, Bhandari R, et al. Comprehensive analysis of glycolysis-related genes for prognosis, immune features, and candidate drug development in colon cancer. Front Cell Dev Biol. 2021;9:684322.
Yang G, Wang H, Sun B. Construction of cuproptosis‑associated prognostic signature in colon adenocarcinoma based on bioinformatics and RT‑qPCR analysis. Oncol Lett. 2023;25:91.
Ye C, Huang Y, Gao Y, Zhu S, Yuan J. Exploring the glycolytic cross-talk genes between inflammatory bowel disease and colorectal cancer. Funct Integr Genom. 2023;23:230.
Slowicka K, Petta I, Blancke G, Hoste E, Dumas E, Sze M, et al. Zeb2 drives invasive and microbiota-dependent colon carcinoma. Nat cancer. 2020;1:620–34.
Sreekumar R, Harris S, Moutasim K, DeMateos R, Patel A, Emo K, et al. Assessment of nuclear ZEB2 as a biomarker for colorectal cancer outcome and TNM risk stratification. JAMA Netw Open. 2018;1:e183115–e183115.
Shen X, Hu X, Mao J, Wu Y, Liu H, Shen J, et al. The long noncoding RNA TUG1 is required for TGF-β/TWIST1/EMT-mediated metastasis in colorectal cancer cells. Cell Death Dis. 2020;11:65.
Ji T, Zhang C, Zheng L, Dunaway-Mariano D, Allen KN. Structural basis of the molecular switch between phosphatase and mutase functions of human phosphomannomutase 1 under ischemic conditions. Biochemistry. 2018;57:3480–92.
Liang M, Sun Z, Chen X, Wang L, Wang H, Qin L, et al. E3 ligase TRIM28 promotes anti-PD-1 resistance in non-small cell lung cancer by enhancing the recruitment of myeloid-derived suppressor cells. J Exp Clin Cancer Res. 2023;42:275.
Wang R, Zhuang A, Wu Y, Shen J, Zhang Y, Liu P, et al. S-palmitoylation: a novel player in cancer and its emerging therapeutic implications. Cell Investig. 2025;1:100032.
Agarwal N, Rinaldetti S, Cheikh BB, Zhou Q, Hass EP, Jones RT, et al. TRIM28 is a transcriptional activator of the mutant TERT promoter in human bladder cancer. Proc Natl Acad Sci USA. 2021;118:e2102423118.
Kent LN, Leone G. The broken cycle: E2F dysfunction in cancer. Nat Rev Cancer. 2019;19:326–38.
Baral S, Yu Y, Sun Q, Jiang M, Li R, Cheng Y, et al. Transcription factor E2F4 promote proliferation, migration, and invasion of gastric cancer cells by transcriptionally activating DSCC1. Int J Biol Sci. 2024;20:4978–98.
Allmann S, Mayer L, Olma J, Kaina B, Hofmann TG, Tomicic MT, et al. Benzo[a]pyrene represses DNA repair through altered E2F1/E2F4 function marking an early event in DNA damage-induced cellular senescence. Nucleic Acids Res. 2020;48:12085–101.
Li J, Cheng H, Zhao Y, Wang Y, Gong C, Gong R, et al. ZNF331 represses the proliferation of head and neck squamous cell carcinoma via co-repressor TRIM28. Oral Dis. 2024;31:1178–88.
Wang H, Yang W, Qin Q, Yang X, Yang Y, Liu H, et al. E3 ubiquitin ligase MAGI3 degrades c-Myc and acts as a predictor for chemotherapy response in colorectal cancer. Mol Cancer. 2022;21:151.
Liu S, Guo R, Xu H, Yang J, Luo H, Yeung S-CJ, et al. 14-3-3σ-NEDD4L axis promotes ubiquitination and degradation of HIF-1α in colorectal cancer. Cell Rep. 2023;42:112870.
Lionnard L, Duc P, Brennan MS, Kueh AJ, Pal M, Guardia F, et al. TRIM17 and TRIM28 antagonistically regulate the ubiquitination and anti-apoptotic activity of BCL2A1. Cell Death Differ. 2018;26:902–17.
Liao H, Zhang L, Lu S, Li W, Dong W. KIFC3 promotes proliferation, migration, and invasion in colorectal cancer via PI3K/AKT/mTOR signaling pathway. Front Genet. 2022;13:848926.
Lu S, Liu Y, Tian S, He Y, Dong W. KIFC3 regulates progression of hepatocellular carcinoma via EMT and the AKT/mTOR pathway. Exp Cell Res. 2023;426:113564.
Ma Y, Zhang Y, Jiang X, Guan J, Wang H, Zhang J, et al. KIFC3 promotes the proliferation, migration and invasion of non-small cell lung cancer through the PI3K/AKT signaling pathway. Sci Rep. 2024;14:20471.
Zhou K, Lin J, Dai M, He Y, Xu J, Lin Q. KIFC1 promotes aerobic glycolysis in endometrial cancer cells by regulating the c-myc pathway. J Bioenerg Biomembr. 2021;53:703–13.
Gao H, Wang J, Liu J, Wang H, Wang T, Li S, et al. FOXD1 activates KIFC1 to modulate aerobic glycolysis and reinforce cisplatin resistance of breast cancer. Reprod Biol. 2025;25:100969.
Vilas A, Yuste-Checa P, Gallego D, Desviat LR, Ugarte M, Pérez-Cerda C, et al. Proteostasis regulators as potential rescuers of PMM2 activity. Biochim Biophys Acta Mol Basis Dis. 2020;1866:165777.
Author information
Authors and Affiliations
Contributions
ZP: manuscript editing, data curation and investigation and formal analysis; BM and ZS: data curation and investigation and formal analysis; YSZ and YY: data curation and investigation; YL: manuscript editing, data curation and investigation; CL and YZ: project administration, formal analysis and manuscript writing and editing. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peng, Z., Ma, B., Song, Z. et al. PMM2 interacts with TRIM28 to recruit E2F4 and promote KIFC3-mediated tumor glycolysis and colorectal cancer progression. Oncogene 45, 1145–1160 (2026). https://doi.org/10.1038/s41388-026-03707-x
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-026-03707-x