Abstract
Metastasis is the hallmark of lethal renal cell carcinoma (RCC), and the underlying mechanism driving RCC metastasis remains insufficiently understood. Amino acid metabolism remodeling plays a key role in the matastasis of RCC. In this study, we identified SLC6A19, a tryptophan transporter, as a novel suppressor gene of RCC, which is closely correlated with the metastasis and survival of RCC patients. Overexpression of SLC6A19 significantly inhibits RCC cell proliferation, migration and invasion both in vitro and in vivo. Mechanistically, SLC6A19 actively transports tryptophan into cells, facilitating de novo NAD+ biosynthesis, which in turn activates the NAD+-dependent deacetylase SIRT1. This activation drives the deacetylation of histone H3 at lysine 27 (H3K27), thereby restraining the transcription of NF-κB subunit p65, and effectively suppressing the epithelial-mesenchymal transition (EMT) of RCC cells. What’s more, the inactivation of the transcription factor KLF4 is the key factor for the low expression of SLC6A19 in RCC cells. In conclusion, this study uncovers a novel key pathway that drives RCC invasion and metastasis, offering a promising therapeutic target for clinical intervention.
Similar content being viewed by others
Introduction
Renal cell carcinoma (RCC) is a prevalent malignancy of the urinary system, comprising 3–5% of all adult cancers, with clear cell renal cell carcinoma (ccRCC) being the predominant histological subtype [1]. RCC is characterized by high aggressiveness and recurrence rates [2]; Metastasis commonly occurs in advanced stages, and the prognosis is generally poor [3]. Furthermore, identifying key targets involved in the development and metastasis of RCC can enable earlier detection of high-risk patients, allowing for proactive prevention and intervention strategies aimed at prolonging survival and enhancing quality of life. Therefore, investigating the mechanisms underlying the metastasis of RCC is of irreplaceable value in clinical treatment.
Recent studies have shown that the metabolic state of tumor cells changes during cancer progression [4, 5]. So, it is essential to determine how metabolic reprogramming supports tumor progression and how associated reprogramming activities contribute to tumor progression and metastasis. The solute carrier (SLC) superfamily consists of 67 families and represents the largest group of transporters, responsible for the secondary active transport or facilitative shuttling of polar molecules. Solute carrier 6 (SLC6) is a family of transmembrane proteins that transport various small polar compounds [6, 7]. SLC6A19 is a transporter protein involved in the absorption and reabsorption of various neutral amino acids (such as branched-chain amino acids, methionine, phenylalanine, glycine and tryptophan) in the proximal renal tubules and intestinal epithelial cells of normal tissues. Mutations in SLC6A19 can lead to Hartnup disorder, characterized by aminoaciduria and pellagra-like symptoms caused by tryptophan deficiency [8]. However, the role of SLC6A19 in cancer progression and metastasis remains unclear.
Nicotinamide adenine dinucleotide exists in two forms: oxidized (NAD+) and reduced (NADH). The ratio between NAD+ and NADH plays a crucial role in maintaining redox homeostasis. NAD+ can be synthesized de novo from tryptophan or salvaged from its precursors [9]. The de novo biosynthesis of NAD+, mediated by tryptophan, is a key intracellular source. Dysregulation of tryptophan metabolism components can disrupt NAD+ levels, thereby impairing cellular functions, including energy metabolism [10]. Sirtuins (SIRTs) are a family of NAD+-dependent deacetylases with diverse physiological and pathological roles, among which SIRT1 has been the most extensively studied in recent research [11]. Furthermore, SIRT1, an NAD+-dependent deacetylase, regulates the activity of various key proteins through deacetylation, influencing biological processes such as metabolism, aging and stress responses [12]. In RCC, low expression of SIRT1 is associated with tumor malignancy, metastasis and patient prognosis [13, 14].
Spread and metastasis are often associated with EMT, with the reprogramming activities of these cells linked to the regulation of cancer cell metabolism and metabolites, inflammation and immune responses, oxidative stress, matrix remodeling and extracellular matrix interactions [15,16,17]. Furthermore, epigenetic regulation plays an essential role, with changes in histone modification levels directly influencing the expression of EMT-related transcription factors [18,19,20,21]. These modifications, through complex regulatory networks, synergistically affect multiple transcription factors, thereby altering cellular motility and invasiveness.
In our study, we found that the loss of SLC6A19 is associated with poor prognosis and tumor progression in RCC, and further investigated how SLC6A19 was dysregulated in RCC cells and its suppressive effects on RCC metastasis by altering NAD+ de novo biosynthesis. These data provided new insights into the functional significance of SLC6A19 and it mediated NAD+ metabolism in RCC progression.
Results
Low expression of SLC6A19 in RCC
We analyzed the RCC transcriptome data from the TCGA database and found that SLC6A19 expression was the most significantly decreased gene between primary RCC tissues from patients with distant metastasis and those from non-metastasis patients (Fig. 1A). Meanwhile, SLC6A19 expression was lower in tumors with T3–4 stages than T1–2 stages (Fig. 1B). Furthermore, Kaplan-Meier (K-M) analysis showed that higher expression of SLC6A19 was associated with better clinical prognosis in patients with RCC (Fig. 1C).
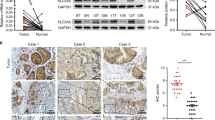
A Volcano plot analysis of differentially expressed genes in the TCGA dataset (M1 vs. M0). B Dot plot of SLC6A19 mRNA levels in the TCGA database (Normal, n = 72, Tumor, n = 524. M0, n = 470, M1, n = 96. T1, n = 299, T2, n = 81, T3, n = 204, T4, n = 13). C Survival analysis of RCC patients with SLC6A19 expression levels in the TCGA cohort (p < 0.0001) (High SLC6A19, n = 300, Low SLC6A19, n = 302). D–F Representative IHC images and scores of SLC6A19 expression in local clinical samples, including cancer and adjacent normal tissues, non-metastatic and metastatic primary lesions, and samples from different clinical stages. G Western blot analysis of SLC6A19 protein levels in 8 pairs of clinical RCC samples. H Western blot analysis of SLC6A19 protein levels in RCC cell lines and the control HK-2 cell line. Scale bar = 100 μm, *p < 0.05, ***p < 0.001, ****p < 0.0001.
To further confirm the downregulation of SLC6A19 in RCC, we performed immunohistochemistry (IHC) to assess the protein levels of SLC6A19 in 119 RCC patients. We observed that the expression of SLC6A19 was significantly lower in tumor tissues compared to adjacent non-tumor tissues (Fig. 1D). Additionally, SLC6A19 expression levels were lower in high-grade tumors than low-grade tumors, as well as lower in primary tumors from patients with distant metastasis than non-metastasis patients (Fig. 1E, F, Table 1). Correspondingly, Western blot analysis showed a significant reduction in SLC6A19 expression in tumor tissues compared to adjacent non-tumor tissues. All paired samples (Case 1–8) were obtained from patients with clear cell renal cell carcinoma (ccRCC) classified as stage I and Fuhrman grade 1 (Fig. 1G). Furthermore, SLC6A19 expression was significantly downregulated in RCC cell lines compared to the control HK-2 cells (Fig. 1H). These findings suggest that SLC6A19 may play a significant role in the pathogenesis and progression of RCC.
SLC6A19 inhibits RCC progression in vitro and in vivo
To investigate the role of SLC6A19 in RCC progression, stable SLC6A19 overexpression cell lines were constructed in the 786-O and Caki-1 cell lines with lentivirus system. The overexpression levels were verified by Western blot (Fig. 2A). Subsequently, to evaluate the role of SLC6A19 in RCC cell migration and invasion, we performed transwell assays, wound-healing assays in 786-O and Caki-1 cells. We found that stable overexpression of SLC6A19 in 786-O and Caki-1 cells significantly inhibited cell migration and invasion (Fig. 2B–D). Subsequently, we assessed the expression of several EMT-related transcription factors and classical molecular markers by Western blot (Fig. 2E). Cellular morphological changes were observed under microscope and we found that overexpression of SLC6A19 effectively inhibited cell polarity (Fig. 2F).
A Efficiency of SLC6A19 overexpression validated in 786-O and Caki-1 cells. B, C Wound-healing assays of CONTROL and SLC6A19-overexpressing 786-O and Caki-1 cell lines in vitro. D Transwell assays of CONTROL and SLC6A19-overexpressing 786-O and Caki-1 cell lines in vitro. E Protein expression levels of EMT-related cellular markers and transcription factors in 786-O and Caki-1 cell lines. F Representative images of morphologic alteration of cells with SLC6A19 overexpression. Scale bar = 100 μm. G Dissected tumors from subcutaneously injected mice with SLC6A19-overexpression ACHN cells or ACHN cells transduced with control lentivirus vectors; n = 5/group. Tumor volume and weight of subcutaneous tumors. H H&E staining in dissected subcutaneous tumor samples. Scale bar = 100 μm. I IHC staining of N-cadherin, E-cadherin and Ki-67 in dissected subcutaneous tumor samples. Scale bar = 100 μm. J Representative gross images of lung samples showing metastatic nodules (black arrows). K H&E staining of lungs. Number of the pulmonary metastatic lesions. Scale bar = 2.5 mm *p < 0.05, ***p < 0.001, ****p < 0.0001.
Stable SLC6A19 overexpression ACHN cells and ACHN cells transduced with control lentivirus vectors were subcutaneously injected into the right flanks of mice. Five weeks post-injection, we analysis the tumors and found that stable overexpression of SLC6A19 significantly reduced the volume and weight of subcutaneous tumors compared to these in the CONTROL group (Fig. 2G). Hematoxylin and eosin (H&E) and IHC analysis revealed that, compared to the CONTROL group, tumors in SLC6A19 overexpression group exhibited reduced tumor growth, lower Ki-67 expression, decareased N-cadherin levels, and increased E-cadherin expression (Fig. 2H, I).
Additionally, to assess the effect of SLC6A19 on metastasis of RCC, we performed lung metastasis model using stable SLC6A19 overexpression ACHN cells. Lung metastasis ratio was assessed four weeks after tail vein injection of ACHN cells. Consistently, mice in SLC6A19 overexpression group exhibited smaller and fewer lung metastatic nodules (Fig. 2J, K). These results show that SLC6A19 effectively inhibits tumor growth and metastasis in vitro and in vivo.
SLC6A19 activates the de novo NAD+ biosynthesis pathway by transporting tryptophan
To investigate whether metastasis-suppressive effect of SLC6A19 was dependent on its tryptophan transporter function, we removed tryptophan in the culture medium of both SLC6A19 overexpressing and CONTROL RCC cells. Remarkably, we found that SLC6A19 overexpression could obviously inhibit the migration and invasion abilities of RCC cells in the culture medium supplement with tryptophan, however, SLC6A19 overexpression could not alter the migration and invasion abilities of RCC cells when cultured in tryptophan-free medium (Fig. 3A–D). Similarly, Western Blot results showed that overexpression of SLC6A19 could no more regulate the protein levels of multiple EMT modulators (Fig. 3E). These results suggested that the function of SLC6A19 to restrain RCC metastasis depends on tryptophan metabolism.
A, B Transwell assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and removal of tryptophan in vitro. C, D Wound-healing assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and removal of tryptophan in vitro. E Protein expression of EMT-related cellular markers and transcription factors in cell lines. Scale bar = 50 μm ***p < 0.001, ****p < 0.0001.
To demonstrate the mechanism by which SLC6A19 suppressed RCC metastasis, we further analyzed metabolite profiles of both SLC6A19 overexpression and CONTROL 786-O cells using metabolomics. Metabolomics analysis revealed that, compared to the CONTROL group, 66 secondary metabolites were upregulated, and 41 metabolites were downregulated in 786-O cells overexpressing SLC6A19 (Supplementary Fig. 1A–B). Among these altered metabolites, the level of NAD+ was significantly increased in SLC6A19 overexpression cells (Fig. 4A). By further KEGG and GSEA enrichment analysis, we found that SLC6A19 was closely related to biosynthesis of cofactors (Fig. 4B, Summlemetary Fig. 1C, D). As previously reported, NAD+ could be synthesized de novo from tryptophan [9]. Our results suggested that the effect of SLC6A19 was likely associated with the upregulation of NAD+. Next, we analyzed NAD+/NADH levels in the 786-O and Caki-1 cells, and confirmed that NAD+ levels were significantly elevated after SLC6A19 overexpression. However, under tryptophan-depleted conditions, NAD+ levels showed no significant changes in SLC6A19 overexpression group when compared to the CONTROL group (Fig. 4C), indicating that the cellular upregulation of NAD+ is associated with SLC6A19-transported tryptophan. NMNAT1, a key enzyme in the de novo NAD+ biosynthesis pathway that converts nicotinamide mononucleotide (NAMN) to NAD+ [22, 23]. To prove that SLC6A19 suppressed metastasis through upregulating cellular NAD+ levels, we impeded NAD+ de novo biosynthesis by transfecting siNMNAT1 into RCC cells with CONTROL or overexpressed SLC6A19. Subsequent transwell and wound healing assays indicated that SLC6A19 could not alter the migration and invasion abilities of RCC cells when NAD+ de novo biosynthesis was hindered by siNMNAT1 (Fig. 4D–G). These results suggested that SLC6A19 restrained the metastasis of RCC cells through a function link with the de novo NAD+ biosynthesis pathway by transporting tryptophan.
A Heatmap of metabolite changes in 786-O cells with overexpression of SLC6A19 or CONTROL. B Bubble plot of KEGG enrichment analysis for differential metabolites. C Changes of NAD+ and NADH levels in 786-O and Caki-1 cell lines with overexpression of SLC6A19 and removal of tryptophan in the culture medium. D, E Transwell assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and knockdown of NMNAT1 in vitro. F, G Wound-healing assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and knockdown of NMNAT1 in vitro. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
NAD+-dependent SIRT1 downregulates NF-κB p65 expression by decreasing H3K27 acetylation
Previous studies have shown that SIRT1 is an NAD+-dependent deacetylase that plays a significant role in cancer progression. Therefore, we hypothesized that the activation of SIRT1 could be a key target through which SLC6A19 inhibited RCC metastasis.
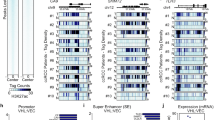
To verify this hypothesis, we first performed Western blot analysis to measure the levels of Acetylated lysine (Ac.k) proteins in 786-O and Caki-1 cell lines. The results demonstrated that SLC6A19 overexpression significantly decreased total protein acetylation, particularly histones. However, this effect is primarily rescued by siSIRT1 rather than by siSIRT2, siSIRT3, or siSIRT6, particularly in histone deacetylation (Fig. 5A, Supplementary Fig. 2A–C). SIRT1 is a crucial NAD⁺-dependent deacetylase that modulates chromatin structure and gene expression by deacetylating various histone lysine residues, including H3K27ac [24]. Previous studies have reported that NAD+ supplementation in humans enhances the removal of H3K27ac from zygotes, an effect mediated by SIRT1 [25]. H3K27ac is a critical epigenetic marker of gene activation, highly enriched in promoter regions, where it facilitates chromatin accessibility and enhances gene transcription [26]. Subsequently, we measured the acetylation level of histone H3 at lysine 27 (H3K27ac) in 786-O and Caki-1 cell lines, and found that overexpression of SLC6A19 could significantly decreased the level of H3K27ac, while the effect was blocked when SIRT1 deficiency (Fig. 5B). The results indicated that SLC6A19 activated the SIRT1-medicated deacetylation of H3K27ac. Therefore, we further investigated the potential downstream mechanism caused by H3K27ac. We analyzed ChIP-seq data using Cistrome DB (http://cistrome.org/db/#/), and found that H3K27ac was greatly enriched in the promoter region of p65, an NF-κB subunit that played a crucial role in driving EMT process, in 786-O cells (Fig. 5C). Furthermore, our ChIP-qPCR assay using H3K27ac specific antibody also confirmed the binding of H3K27ac to the NF-κB p65 promoter region and SLC6A19 overexpression could obviously attenuate the enrichment of H3K27ac in p65 promoter region, while the effect was mostly blocked when SIRT1 deficiency (Fig. 5D).
A Western blot analysis of Acetylated lysine levels in 786-O and Caki-1 cell lines with overexpression of SLC6A19 and knockdown of SIRT1. B Western blot analysis of the effects of SLC6A19 and SIRT1 knockdown on the acetylation levels of H3K27 in 786-O cells. C Analysis of the binding sites of HK327ac to the RELA gene in 786-O cells based on two ChIP-seq datasets from the UCSC genome browser. D ChIP-qPCR results show changes in the binding levels of H3K27ac with the NF-κB p65 promoter with overexpression of SLC6A19 and knockdown of SIRT1. E Western blot analysis of EMT-related cellular markers and transcription factors in cell lines. F, G Transwell assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and knockdown of SIRT1 in vitro. H, I Wound-healing assays of 786-O and Caki-1 cell lines with overexpression of SLC6A19 and knockdown of SIRT1 in vitro. *p < 0.05, ***p < 0.001, ****p < 0.0001.
We further detected the protein levels of p65 and its downstrem regulator, such as Twist and Snail, as well as other EMT maker proteins. The results showed that SLC6A19 could greatly decreased the protein levels of p65, Twist, Snail, and N-cadherin, upregulated E-cadherin protein level. However, overexpression of SLC6A19 no more altered the expression of these proteins when SIRT1 was deficient in cells (Fig. 5E). Consistenly, in cell function assays, SIRT1 depletion could greatly block the suppressive effect of SLC6A19 on the migration and invasion abilities of RCC cells (Fig. 5F–I). These above results proved that SLC6A19 activated the SIRT1-medicated deacetylation of H3K27ac, thus reduced the transcription of p65, impeding EMT process and suppressing RCC metastasis. Our SIRT1 siRNA knockdown experiments showed that silencing SIRT1 abolished the NAD+-induced deacetylation and suppression of EMT, indicating that SIRT1 plays a functionally relevant regulatory role in this process.
Inactivation of transcription factor KLF4 mediates low expression of SLC6A19 in RCC
To further investigate the cause of low SLC6A19 expression in RCC cells, we used the online tools to predict potential upstream transcription factors of SLC6A19. By integrating gene expression correlation analyses from TCGA, potential transcription factors predicted from GTRD (Gene Transcription Regulation Database)(https://gtrd.biouml.org/) and JASPAR (The Open-Access Database of Transcription Factor Binding Profiles)(https://jaspar.elixir.no/) databases, we screened out two candidate transcription factors, KLF4 and HNF4A, for further analysis (Fig. 6A). Then we constructed overexpressing plasmids of KLF4 and HNF4A, respectively (Supplementary Fig. 3A–B), and found that overexpression of KLF4 could obviously upregulate the mRNA level of SLC6A19 in RCC cell lines, while HNF4A overexpression did not significantly increase the mRNA expression of SLC6A19 (Fig. 6B). Consistently, KLF4 overexpression could also greatly boost SLC6A19 protein level in both 786-O and Caki-1 cell lines (Fig. 6C).
A Venn diagram showing the predicted upstream transcription factors of SLC6A19 based on JASPAR, GTRD, and TCGA databases. B qRT-PCR analysis of SLC6A19 mRNA levels after performing transient overexpression of KLF4 and HNF4A in 786-O and Caki-1 cells. C Western blot analysis of SLC6A19 protein levels. D JASPAR indicated that KLF4 shared with the binding site with the promoter region of SLC6A19. E ChIP-qPCR using KLF4 antibody in 786-O cells to detect enrichment of potential binding sequences in the promoter region of SLC6A19. F Dual-luciferase reporter assays show that KLF4 binds to the wild-type (WT) sequence, but not the mutant (MUT). G, H IHC shows the expression levels of KLF4 and SLC6A19 in RCC patients. I Analysis of the correlation between KLF4 and SLC6A19 mRNA expression levels in the TCGA database (n = 367). J Representative IHC images and scores of KLF4 expression in local clinical samples, including cancer and adjacent normal tissues. K Survival analysis of RCC patients with SLC6A19 expression levels in the TCGA cohort (p = 0.0038) (High KLF4, n = 302, Low KLF4, n = 303). L The mechanism of SLC6A19-Mediated Inhibition of RCC Metastasis. Scale bar = 100 μm *p < 0.05, **p < 0.01, ****p < 0.0001, ns not significant.
Next, to identify the specific binding sites of KLF4 in SLC6A19 promoter region, we performed ChIP-qPCR and Dual-luciferase reporter assays in 786-O cell line. Figure 6D showed the predicted binding sites by JASPAR. The result of ChIP-qPCR assay indicated that KLF4 could bind to this specific site in SLC6A19 promoter region (Fig. 6E). To confirm the result, we consructed luciferase reporter plasmids containing wild (WT) or the mutation (MUT) binding sequences in the promoter region of luciferase. These results of luciferase reporter assays showed that KLF4 overexpression could significantly increase the luciferase activity in WT group, but not in MUT group (Fig. 6F). These results proved that KLF4 could bind to the specific site in SLC6A19 promoter region and boost the transcription of SLC6A19. Besides, we analyzed the protein levels of SLC6A19 and KLF4 in RCC tissues, found that the levels of SLC6A19 were higher in patients with higher KLF4 expression, and the protein levels of SLC6A19 and KLF4 in RCC tissues were significantly positively correlated (Fig. 6G, H). What’s more, we further confirmed the positive correlation between the mRNA levels of KLF4 and SLC6A19 in RCC tissues using TCGA database (Fig. 6I). Furthermore, local clinical samples showed a significant reduction in KLF4 expression in tumor tissues compared to adjacent non-tumor tissues (Fig. 6J) and Kaplan–Meier (K-M) analysis using TCGA data showed that higher expression of KLF4 was associated with better clinical prognosis in patients with RCC (Fig. 6K). These results indicated that the inactivation of transcription factor KLF4 might be the crucial reason of low SLC6A19 expression in RCC.
Discussion
Cancer metastasis remains an unresolved issue, and the specific mechanisms involved in the metastatic process are still not fully understood. In this study, We identified SLC6A19 is a powerful tumor suppressor gene in RCC, which is significantly down-regulated in RCC cells due to the inactivation of the transcription factor KLF4. In mechanism, SLC6A19 could transport tryptophan into cells, facilitates de novo NAD+ biosynthesis, and activates SIRT1-mediated deacetylation of H3K27ac, thereby restraining the transcription of NF-κB subunit p65 and effectively suppressing the EMT process of RCC cells.
As a neutral amino acid transporter, SLC6A19 has been reported to play a key role in certain metabolic disorders. For example, in mouse models, the absence of SLC6A19 normalizes elevated plasma amino acids in rare metabolic disorders such as phenylketonuria and urea cycle disorders [27]. However, in tumorigenesis and metastasis, the function and specific mechanism are still unclear. By analyzing Both the TCGA database and samples from our center confirmed the low expression of SLC6A19 in tumors, which correlates with metastasis and poor prognosis, suggesting its potential tumor-suppressive role in RCC progression. Further in vitro and in vivo experiments demonstrated that SLC6A19 was indeed a novel critical tumor suppressor gene in RCC.
Tryptophan metabolism is the source of de novo NAD+ biosynthesis and is closed correlated with cellular NAD+ homeostasis, which is crucial in tumor progression. The underlying mechanism by which NAD+ homeostasis affected cell function was intricate. A recent study reported that prolonged incubation with NAD+ leads to pyrimidine depletion, purine accumulation, replication stress, and cell cycle arrest [28]. The accumulation of lactic acid in tumor cells increases NADH levels, disrupts the NAD+/NADH metabolic balance, lowers intracellular pH, and promotes tumor progression [29]. However, the precise relationship between NAD+ homeostasis and EMT remains unclear. NAD+ homeostasis may regulate EMT through multiple mechanisms, such as activating SIRT1 to modulate transcription factors, activating AMPK to suppress the mTOR pathway, and influencing the NAD+/NADH ratio to regulate oxidative stress [30]. Our study demonstrates that SLC6A19 restrains RCC metastasis through facilitating de novo NAD+ biosynthesis via transporting tryptophan into cells, offering new insights into the relationship between NAD+ homeostasis and RCC metastasis.
Sirtuins are a class of post-translational modification enzymes that rely on NAD+ as a co-substrate to catalyze the deacylation of lysine residues [31]. Increased NAD+ can act as a cofactor to activate various NAD+-dependent enzymes, thus, fluctuations in NAD+ availability directly influence SIRT1-mediated deacetylation activity and its downstream biological effects, including transcriptional regulation. NAD+-dependent deacetylase SIRT1 has garnered significant attention. For instance, SIRT1 deficiency induces acetylation and activation of the cell cycle checkpoint kinase WEE1, thereby inhibiting WEE1 and promoting drug resistance [32]. In gastric cancer, SIRT1 suppresses the expression of ARHGAP5 by physically binding to the transcription factor c-JUN, deacetylating it, and inhibiting its transcriptional activity [33]. Other NAD+-dependent deacetylases, such as SIRT2, primarily regulate the cell cycle and play roles in the nervous system [34]. SIRT3 is located in mitochondria and is involved in regulation of mitochondrial functions, while SIRT6 is found in the nucleus and participates in DNA repair [35]. The role of SIRT1 in RCC has gained increasing attention. RCC cells typically exhibit a high metabolic rate, and alterations in SIRT1 activity may influence tumor growth and metastasis through the regulation of other metabolic pathways [36]. Moreover, SIRT1 modulates various biological processes, including transcription, genome stability, signal transduction, and metabolism, primarily through histone deacetylation [37]. Studies have shown that elevated H3K27ac acetylation levels are associated with cholesterol synthesis and can be reversed by a SIRT1 agonist [38]. H3K27ac is widely recognized as a key epigenetic marker that promotes transcription [39]. Concurrent inhibition of p300/CBP and FLT3 enhances the deacetylation of H3K27, and further reduces the phosphorylation of STAT5, AKT, and ERK, as well as c-Myc expression, which contributes to synergistic antileukemic effects [40]. In this study, we demonstrated that SLC6A19 could activate SIRT1-mediated deacetylation of H3K27ac, further restrained the transcription of NF-κB subunit p65, and eventually suppressed the EMT process of RCC cells.
These provide a metabolic-epigenetic link that integrates nutrient transport with transcriptional control of malignancy-related pathways. While our data collectively support this model—through NAD+ measurement, functional rescue experiments, and downstream molecular readouts—several mechanistic layers warrant further clarification. However, the mechanistic link between SLC6A19, NAD+, SIRT1, and p65 remains incompletely defined. Although we demonstrated that NAD+ accumulation mediated by tryptophan uptake, we did not directly quantify key intermediate metabolites along the de novo NAD+ biosynthesis pathway, such as kynurenine, quinolinic acid, or NMN. Given that NAD⁺ can be synthesized via de novo, salvage, and Preiss–Handler pathways, future studies employing targeted metabolomics, metabolic flux analysis or isotope tracing will be essential to clarify the specific contribution of the tryptophan–kynurenine–NAD+ axis [41, 42]. In addition, since SIRT1 is known to regulate multiple cellular processes, mapping its downstream regulatory network in RCC represents an important direction for future work.
In conclusion, we identified SLC6A19 as a novel metastasis-suppressor gene in RCC, whose low expression is mediated by KLF4 inactivation. Functionally, SLC6A19 promotes de novo NAD⁺ biosynthesis by transporting tryptophan, thereby activating SIRT1 and subsequently reducing NF-κB p65 expression via H3K27 deacetylation. This finding aids in further understanding the metastatic mechanisms of RCC and offers new therapeutic targets and strategies for personalized treatment.
Materials and methods
Patients and clinical samples
A total of 116 paraffin-embedded tissue specimens were obtained from patients with RCC who underwent surgical resection at Nanjing Drum Tower Hospital between 2014 and 2022 for immunohistochemistry (IHC) analysis and 8 paired samples from patients with clear cell renal cell carcinoma (ccRCC) classified as stage I and Fuhrman grade 1 were used for Western blot analysis. For patients with metastasis, systemic targeted therapy with axitinib was administered prior to surgery. Written informed consent was provided by all participating patients, and the study protocol was approved by the Ethics Committee of Nanjing Drum Tower Hospital.
Cell lines and culture
All cell lines were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). All lines underwent short tandem repeat (STR) profiling to confirm their identity and were verified to be free of mycoplasma contamination. The 786-O and 769-P cells were cultured in RPMI 1640 medium (WISENT, Nanjing, China), Caki-1 and ACHN cells were cultured in McCoy’s 5A medium (WISENT, Nanjing, China) and DMEM medium (WISENT, Nanjing, China), each supplemented with 10% FBS (ABW, Shanghai, China) and 1% penicillin-streptomycin, in a humidified incubator at 37 °C with 5% CO2.
Bioinformatics analysis
RCC data were obtained from The Cancer Genome Atlas (TCGA). Samples were grouped into M0 or M1 based on the absence or presence of distant metastasis, respectively. Gene expression levels were estimated using the RSEM method, and RNA-Seq normalized counts underwent logarithmic transformation.
RNA extraction and quantitative real-time PCR (qRT-PCR)
Total RNA was extracted using TRIzol reagent (Vazyme, Nanjing, China) according to the manufacturer’s instructions. cDNA synthesis was performed using the HiScript III RT SuperMix (Vazyme, Nanjing, China). For qRT-PCR, we used SYBR Green-based detection with 2× Taq Master Mix (Vazyme, Nanjing, China) and data were collected using the StepOne Real-Time PCR System (Applied Biosystems, CA, USA). Relative mRNA levels were assessed by normalizing to ACTB expression through the 2−ΔΔCT method. Primer amplification efficiency was assessd by generating a standard curve using serial dilutions of cDNA, and only primers with amplification efficiencies between 90% and 110% were used. The primer sequences used are provided in Supplementary Table 1. All experiments were conducted in biological triplicates.
Immunohistochemistry (IHC)
Clinical RCC and non-cancerous samples, along with subcutaneous tumor tissues from mice, were subjected to immunohistochemistry (IHC). The evaluation of IHC staining involved two parameters: staining intensity and the proportion of stained area. Intensity was graded on a scale of 0 (negative), 1 (low), 2 (moderate), and 3 (high). The extent of staining was scored as follows: 0 for 0% stained, 1 for 1%–25% stained, 2 for 26%–50% stained, 3 for 51%–75% stained, and 4 for over 75% stained. The final IHC score was calculated by multiplying the staining intensity score by the extent score. A final score of 4 or less was considered indicative of low expression, whereas a score greater than 4 signified high expression.
Western blot
Tissue and cell samples were homogenized and lysed in RIPA buffer (Beyotime Biotech, Shanghai, China) supplemented with both proteinase and phosphatase inhibitors (Beyotime Biotech, Shanghai, China). Protein expression levels were subsequently quantified using an electrochemiluminescence (ECL) system (Tanon, Shanghai, China). For a complete list of the primary antibodies used, please refer to Supplementary Table 2.
Transwell
24-well plates with inserts containing membrane filters (Jet Bio-Filtration, Guangzhou, China) were used. Approximately 1 ×105 cells were seeded into the upper chamber of a Transwell insert in serum-free medium, while the lower chamber was filled with medium containing 10% fetal bovine serum (FBS). After 10 h of incubation at 37 °C, non-migrated cells on the upper surface were gently removed, and the migrated cells on the underside were fixed, stained, and counted under a microscope. Diluted Matrigel matrix (ABW, Shanghai, China) was carefully added to the bottom of each upper chamber, followed by the same steps as described above for the Transwell invasion assay. All experiments were conducted in biological triplicates.
Wound-healing
Cells were seeded in 6-well plates and allowed to reach ~90% confluence the next day. A straight-line scratch was made using a sterile pipette tip, and the medium was replaced with medium containing 1% FBS to prevent proliferation. Wound images were taken immediately (0 h) and at 12 h after a scratch was made using a sterile 200 µL pipette. The measurements were then analyzed using Image J software. All experiments were conducted in biological triplicates.
Animal experiments
All animal care and handling followed the NIH Guide for the Care and Use of Laboratory Animals and received approval from the Institutional Review Board of Nanjing Drum Tower Hospital (Nanjing, China). For the xenograft models, male BALB/c nude mice (5 weeks old) were purchased and randomly divided into two groups (n = 5 per group) for subcutaneous injection of 3 × 106 ACHN cells each mouse. The mice were subsequently sacrificed 5 weeks later. The harvested organs were immediately excised and weighed. For the metastatic model, mice were randomly assigned to two groups (n = 5 per group). Approximately 1 × 106 ACHN cells were administered to each mouse via tail vein injections. After 4 weeks of tail vein injections, the mice were sacrificed for histological examination of lung metastases. The number of mice used was consistentent with previous studies. Animals were randomly assigned to experimental groups using simple randomization. The investigator was blinded to group allocation during both the execution of the animal experiments and the outcome assessment.
ChIP-qPCR and ChIP-seq
The ChIP assay for 786-O cells was carried out using the Simple ChIP Plus Sonication Chromatin IP Kit (Cell Signaling Technology, USA) following the manufacturer’s instructions. Interactions with the predicted promoter regions were subsequently examined by qRT-PCR using the specific primers provided in Supplementary Table 1. ChIP-seq data were obtained from the Cistrome DB database. The dataset used in this study corresponds to GEO accession number GSM2067535. All experiments were conducted in biological triplicates.
NAD+/NADH measurement
The NAD+/NADH measurement was performed using NAD+/NADH Assay Kit with WST-8 (Beyotime Biotech, Shanghai, China) according to the manufacturer’s protocol. The amount was measured at 450 nm by a microplate reader. Each measurement was performed in triplicate.
Metabolomics
Samples were extracted using 80% precooled methanol, followed by protein precipitation at −20 °C and centrifugation at 20,000 × g. The supernatants were vacuum dreid and reconstituted in 50% methanol before LC-MS analysis. UPLC separation was performed using an ACQUITY UPLC T3 column (100 mm × 2.1 mm, 1.8 µm) with a flow rate of 0.3 mL/min and a gradient of water (5 mM ammonium acetate + 5 mM acetic acid) and acetonitrile. The Q-Exactive mass spectrometer (Thermo Fisher Scientific) was operated in both postive and negative ion modes with high resolution. Quality control (QC) samples were prepared by pooling equal aliquots from each sample and run periodically to ensure system stability. Raw data were converted to mzXML format and processed using XCMS, CAMERA, and metaX software packages in R. Peak picking, alignment, retention time correction, and metabolite annotation were conducted using these tools. Metabolites were annotated by matching m/z and fragmentation patterns with HMDB, KEGG and in-house spectrla libraries, using a tolerance of 10 ppm. Differential metabolites were identified using statistical filters (VIP > 1, p < 0.05, FDR-adjusted) and multivariate analysis (PLS-DA). Missing values were imputed with k-nearest neighbor, and signal drift was corrected using QC-based robust LOESS.
Lentivirus and plasmids
Human SLC6A19 cDNA was cloned into the lentiviral vector GV492 (pGC-3FLAG-gcGFP-puromycin)(Genechem, Shanghai,China) using Agel and Nhel. Lentiviral particles were generated by co-transfecting GV492-SLC6A19, psPAX2, and pMD2.G into 293T cells with Lipofectamine 2000 (Invitrogen, USA). Virus was collected 72 h later, filtered (0.45 μm), and stored at −80 °C. Lentivirus were transfected into RCC cells according to the manufacturer’s instructions. Cells infected with lentivirus carrying the empty GV492 vector were used as the negative control group.
The KLF4 and HNF4A coding sequences were cloned into the GV219 (pcDNA3.1(+)) expression vector using Agel and Nhel. All constructs were verified by sequencing. Plasmids were transfected into RCC cells according to the manufacturer’s instructions. Empty vector was used as control.
Statistical analysis
All cell experiments were independently conducted at least three times, and the data are expressed as the mean ± standard error of the mean (SEM). Inter-group differences were analyzed using Student’s t test (two-tailed) and one-way analysis of variance (ANOVA).
Data availability
The datasets supporting the conclusions of this article have been deposited in the OMlX, China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences, and are available at https://ngdc.cncb.ac.cn/omix under accession number OMIX011101.
References
Rinaldi L, Senatore E, Feliciello S, Chiuso F, Insabato L, Feliciello A. Kidney cancer: from tumor biology to innovative therapeutics. Biochim Biophys Acta Rev Cancer. 2024;1880:189240.
El Zarif T, Semaan K, Xie W, Eid M, Zarba M, Issa W, et al. First-line systemic therapy following adjuvant immunotherapy in renal cell carcinoma: an international multicenter study. Eur Urol. 2024;86:503–12.
Benjamin DJ, Shrestha A, Fellman D, Rezazadeh A. Association between time to treatment initiation and survival in metastatic renal cell cancer: a population-based study of 5,193 cases from the California Cancer Registry (2010-2019). JCO. 2023;41:e16530–e16530.
Wettersten HI, Aboud OA, Lara PN, Weiss RH. Metabolic reprogramming in clear cell renal cell carcinoma. Nat Rev Nephrol. 2017;13:410–9.
Hakimi AA, Reznik E, Lee CH, Creighton CJ, Brannon AR, Luna A, et al. An integrated metabolic atlas of clear cell renal cell carcinoma. Cancer cell. 2016;29:104–16.
Schlessinger A, Zatorski N, Hutchinson K, Colas C. Targeting SLC transporters: small molecules as modulators and therapeutic opportunities. Trends Biochem Sci. 2023;48:801–14.
Kukułowicz J, Pietrzak-Lichwa K, Klimończyk K, Idlin N, Bajda M. The SLC6A15-SLC6A20 neutral amino acid transporter subfamily: functions, diseases, and their therapeutic relevance. Pharmacol Rev 2023;76:142–93.
Kleta R, Romeo E, Ristic Z, Ohura T, Stuart C, Arcos-Burgos M, et al. Mutations in SLC6A19, encoding B0AT1, cause Hartnup disorder. Nat Genet. 2004;36:999–1002.
Zapata-Pérez R, Wanders RJA, van Karnebeek CDM, Houtkooper RH. NAD+ homeostasis in human health and disease. EMBO Mol Med. 2021;13:e13943.
Wettersten HI, Hakimi AA, Morin D, Bianchi C, Johnstone ME, Donohoe DR, et al. Grade-dependent metabolic reprogramming in kidney cancer revealed by combined proteomics and metabolomics analysis. Cancer Res. 2015;75:2541–52.
Ungurianu A, Zanfirescu A, Margină D. Sirtuins, resveratrol and the intertwining cellular pathways connecting them. Ageing Res Rev. 2023;88:101936.
Alves-Fernandes DK, Jasiulionis MG. The role of SIRT1 on DNA damage response and epigenetic alterations in cancer. Int J Mol Sci. 2019;20:3153.
Chen Y, Zhu Y, Sheng Y, Xiao J, Xiao Y, Cheng N, et al. SIRT1 downregulated FGB expression to inhibit RCC tumorigenesis by destabilizing STAT3. Exp Cell Res. 2019;382:111466.
Wang X, Lu Y, Tuo Z, Zhou H, Zhang Y, Cao Z, et al. Role of SIRT1/AMPK signaling in the proliferation, migration and invasion of renal cell carcinoma cells. Oncol Rep. 2021;45:109.
Siddiqui A, Ceppi P. A non-proliferative role of pyrimidine metabolism in cancer. Mol Metab. 2020;35:100962.
Ramesh V, Gollavilli PN, Pinna L, Siddiqui MA, Turtos AM, Napoli F, et al. Propionate reinforces epithelial identity and reduces aggressiveness of lung carcinoma. EMBO Mol Med. 2023;15:e17836.
Li C-W, Xia W, Huo L, Lim S-O, Wu Y, Hsu JL, et al. Epithelial-mesenchymal transition induced by TNF-α requires NF-κB-mediated transcriptional upregulation of Twist1. Cancer Res. 2012;72:1290–1300.
Liu J, Wu Z, Han D, Wei C, Liang Y, Jiang T, et al. Mesencephalic astrocyte-derived neurotrophic factor inhibits liver cancer through small ubiquitin-related modifier (SUMO)ylation-related suppression of NF-κB/snail signaling pathway and epithelial-mesenchymal transition. Hepatology. 2020;71:1262–78.
Hou J, Wang T, Xie Q, Deng W, Yang JY, Zhang SQ, et al. N-Myc-interacting protein (NMI) negatively regulates epithelial-mesenchymal transition by inhibiting the acetylation of NF-κB/p65. Cancer Lett. 2016;376:22–33.
Lou J, Geng S, He W, Liu S-B, Shi X, Chang Y, et al. Zyxin inhibits the epithelial-mesenchymal transition process in gastric cancer by upregulating SIRT1. MedComm. 2023;4:e357.
Yang X, Sun F, Gao Y, Li M, Liu M, Wei Y, et al. Histone acetyltransferase CSRP2BP promotes the epithelial-mesenchymal transition and metastasis of cervical cancer cells by activating N-cadherin. J Exp Clin Cancer Res. 2023;42:268.
Cao Y, Tang L, Du K, Paraiso K, Sun Q, Liu Z, et al. Anterograde regulation of mitochondrial genes and FGF21 signaling by hepatic LSD1. JCI Insight. 2021;6:e147692.
Zhu Y, Liu J, Park J, Rai P, Zhai RG. Subcellular compartmentalization of NAD+ and its role in cancer: a sereNADe of metabolic melodies. Pharmacol Ther. 2019;200:27–41.
Wu Q-J, Zhang T-N, Chen H-H, Yu X-F, Lv J-L, Liu Y-Y, et al. The sirtuin family in health and disease. Sig Transduct Target Ther. 2022;7:402.
Li J, Zhang J, Hou W, Yang X, Liu X, Zhang Y, et al. Metabolic control of histone acetylation for precise and timely regulation of minor ZGA in early mammalian embryos. Cell Discov. 2022;8:96.
English DM, Lee SN, Sabat KA, Baker IM, Pham TK, Collins MO, et al. Rapid degradation of histone deacetylase 1 (HDAC1) reveals essential roles in both gene repression and active transcription. Nucleic Acids Res 2025;52:gkae1223.
Xu J, Hu Z, Dai L, Yadav A, Jiang Y, Bröer A, et al. Molecular basis of inhibition of the amino acid transporter B0AT1 (SLC6A19). Nat Commun. 2024;15:7224.
Munk SHN, Merchut-Maya JM, Adelantado Rubio A, Hall A, Pappas G, Milletti G, et al. NAD+ regulates nucleotide metabolism and genomic DNA replication. Nat Cell Biol. 2023;25:1774–86.
Amjad S, Nisar S, Bhat AA, Shah AR, Frenneaux MP, Fakhro K, et al. Role of NAD+ in regulating cellular and metabolic signaling pathways. Mol Metab. 2021;49:101195.
Li H-R, Liu Q, Zhu C-L, Sun X-Y, Sun C-Y, Yu C-M, et al. β-Nicotinamide mononucleotide activates NAD+/SIRT1 pathway and attenuates inflammatory and oxidative responses in the hippocampus regions of septic mice. Redox Biol. 2023;63:102745.
Pande S, Raisuddin S. Molecular and cellular regulatory roles of sirtuin protein. Crit Rev Food Sci Nutr. 2023;63:9895–913.
Zhu X, Su Q, Xie H, Song L, Yang F, Zhang D, et al. SIRT1 deacetylates WEE1 and sensitizes cancer cells to WEE1 inhibition. Nat Chem Biol. 2023;19:585–95.
Dong G, Wang B, An Y, Li J, Wang X, Jia J, et al. SIRT1 suppresses the migration and invasion of gastric cancer by regulating ARHGAP5 expression. Cell Death Dis. 2018;9:977.
Wang Y, Yang J, Hong T, Chen X, Cui L. SIRT2: controversy and multiple roles in disease and physiology. Ageing Res Rev. 2019;55:100961.
Morigi M, Perico L, Benigni A. Sirtuins in renal health and disease. J Am Soc Nephrol. 2018;29:1799–809.
Tozzi R, Cipriani F, Masi D, Basciani S, Watanabe M, Lubrano C, et al. Ketone bodies and SIRT1, synergic epigenetic regulators for metabolic health: a narrative review. Nutrients. 2022;14:3145.
Shen H, Qi X, Hu Y, Wang Y, Zhang J, Liu Z, et al. Targeting sirtuins for cancer therapy: epigenetics modifications and beyond. Theranostics. 2024;14:6726–67.
Hu S, Liu K, Luo H, Xu D, Chen L, Zhang L, et al. Caffeine programs hepatic SIRT1-related cholesterol synthesis and hypercholesterolemia via A2AR/cAMP/PKA pathway in adult male offspring rats. Toxicology. 2019;418:11–21.
Zhu Y, Shang L, Tang Y, Li Q, Ding L, Wang Y, et al. Genome-wide profiling of H3K27ac identifies TDO2 as a pivotal therapeutic target in metabolic associated steatohepatitis liver disease. Adv Sci. 2024;11:e2404224.
Chen Y-J, Zhao Y, Yao M-Y, Wang Y-F, Ma M, Yu C-C, et al. Concurrent inhibition of p300/CBP and FLT3 enhances cytotoxicity and overcomes resistance in acute myeloid leukemia. Acta Pharmacol Sin. 2025;46:1390–403.
Lu D, Grant M, Lim BL. NAD(H) and NADP(H) in plants and mammals. Mol Plant. 2025;18:938–59.
Bi Q, Zhao J, Nie J, Huang F. Metabolic pathway analysis of tumors using stable isotopes. Semin Cancer Biol. 2025;113:9–24.
Funding
This work was supported by grants from National Natural Science Foundation of China (82172777 to CJ, 82473415 to HG), Nanjing Medical Science and technology development Foundation (ZKX22024 to MD), Jiangsu Province Capability Improvement Project through Science, Technology and Education, Jiangsu Provincial Medical Key Discipline (Laboratory) Cultivation Unit (JSDW202221 to HG).
Author information
Authors and Affiliations
Contributions
ZC, CJ, MD and HG designed and coordinated the project, ZC, RD and ZC did the majority of the experiments with the help of XT, WY, YC, YC, HY and WC. ZC and MD performed bioinformatics analyses, ZC, CJ, MD and HG wrote the manuscript. All authors revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All methods conducted in this study were performed in accordance with the Ethics Committee of Nanjing Drum Tower Hospital. For experiments involving human participants, all participants provided explicit and informed consent prior to sample collection and analysis. All patients had been informed of the purpose of the study and agreed to participate. For animals experiments, all procedures were carried out in compliance with the ARRIVE guidelines and relevant national regulations on animals welfare. The animal study protocol was approved by the Animal Ethics Committee of Nanjing Drum Tower Hospital.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Z., Duan, R., Chen, Z. et al. SLC6A19-mediated tryptophan uptake suppresses renal cell carcinoma metastasis via activating NAD+-dependent deacetylase SIRT1. Oncogenesis 14, 38 (2025). https://doi.org/10.1038/s41389-025-00580-3
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41389-025-00580-3