Abstract
Targeting MDM2 by disrupting its interaction with p53 or inhibiting its E3 ligase activity is a promising strategy to restore p53 functionality. However, achieving anticancer efficacy while minimizing dose-limiting toxicities remains a significant challenge. Moreover, MDM2 also ubiquitinates various non-p53 targets, complicating its therapeutic targeting. In this study, we demonstrate that MDM2 directly facilitates K48-linked polyubiquitination of MEIS1 at K178, leading to its proteasomal degradation. Notably, MEIS1 forms a non-competitive ternary complex with MDM2 and p53, effectively promoting ubiquitin transfer to itself and preventing p53 ubiquitination. The MEIS1 K178R mutant, which is deficient in ubiquitination, fails to suppress MDM2-mediated p53 ubiquitination, demonstrating a mechanistic link between MEIS1 self-ubiquitination and p53 stabilization. Furthermore, MDM2-mediated MEIS1 ubiquitination is a prerequisite for p53 activation in the DNA damage response. Importantly, a MEIS1-derived peptide, which mimics the MDM2-mediating ubiquitination motif, enhances both MEIS1 and p53 stability, suppresses cell proliferation and tumor growth. Collectively, our findings identify MEIS1 as a molecular decoy that competes for ubiquitin transfer to protect p53 and highlight that MEIS1 ubiquitination could be a novel therapeutic target for reactivating p53-dependent tumor suppression.
Similar content being viewed by others
Introduction
DNA damage is one of the major threats to cellular integrity, as it can lead to cell death or genomic instability, both of which contribute to tumorigenesis [1, 2]. The p53 protein is often referred to as the “guardian of the genome” due to its critical role in detecting DNA damage and responding to various stress signals by initiating processes such as cell cycle arrest, apoptosis, and senescence [3]. Under normal cellular conditions, p53 is maintained at a low level through rapid turnover to prevent the activation of unnecessary apoptotic and senescence pathways [4, 5]. However, in response to DNA damage, p53 becomes stabilized, leading to its accumulation and the transcription of a variety of target genes, such as p21 and BAX, which initiate cell cycle arrest and apoptosis [6]. Disruption of this regulatory pathway is commonly observed in cancers, particularly in colorectal cancer (CRC), where p53 dysfunction contributes to tumor progression and resistance to therapy [7, 8]. Given its vital tumor-suppressive role, restoring p53 function presents a promising therapeutic strategy.
MDM2 is a key E3 ubiquitin ligase that promotes the degradation of p53 via the ubiquitin-proteasome pathway [6]. It inhibits p53 via two primary mechanisms: binding to its transcriptional domain to block its activity and promoting its degradation via the proteasome [9, 10]. Notably, p53 also induces the expression of MDM2, establishing a self-regulating feedback loop [6]. Normally, MDM2 serves to keep p53 levels low [11]. However, in response to DNA damage or cellular stress, p53 is released from MDM2-mediated inhibition, allowing its accumulation and activation of genes that lead to cell cycle arrest and apoptosis. Crucially, before p53 can be activated, MDM2-mediated ubiquitination of p53 must be suppressed. Although several regulatory mechanisms for p53 have been identified, the precise processes that inhibit MDM2-mediated p53 ubiquitination to activate p53 in response to cellular stress remain poorly understood.
A major approach to restoring p53 function involves inhibiting the MDM2-p53 interaction or blocking MDM2’s E3 ligase activity [12]. Despite this, the clinical application of MDM2 inhibitors remains challenging due to potential toxicity and the need to balance tumor-suppressive effects [12]. Additionally, MDM2 participates in the ubiquitination of multiple substrates beyond p53, which complicates therapeutic targeting of itself [13, 14]. These challenges underscore the importance of understanding the mechanisms that regulate p53 stability, particularly the selective targeting of p53 by MDM2 among its numerous substrates.
MEIS1, a homeodomain-containing transcription factor, was initially identified as a regulator of hematopoiesis and development but has since been implicated in various cancers. It promotes the progression of acute myeloid leukemia [15], neuroblastoma [16] and skin cancer [17], while acting as a suppressor in prostate cancer [18] and non-small cell lung cancer [19]. Previous research showed that the lncRNA ELFN1-AS1 interacts with EZH2 and DNMT3a, leading to DNA methylation and H3K27me3 modifications that suppress MEIS1 expression and enhance its downstream target, FEN1 [20]. Although MEIS1 has been linked to tumor progression through its transcriptional activity, the post-translational mechanisms regulating its stability remain poorly understood.
In this study, we investigated the regulatory mechanisms governing MEIS1 protein stability and explored its potential role in modulating the MDM2-p53 axis. We identified MEIS1 as a novel substrate of MDM2 and demonstrated that it functions as a molecular decoy, competing with p53 for MDM2-mediated ubiquitination, thus preventing p53 degradation. Specifically, we show that MEIS1 is ubiquitinated at K178, a modification essential for its own degradation and for stabilizing p53. This mechanism suggests that MEIS1 may act as a protective factor for p53 in CRC, providing new insights into the regulation of p53 stability beyond direct MDM2 inhibition. Furthermore, we examined the role of MEIS1 in the DNA damage response and demonstrated that MDM2-mediated ubiquitination of MEIS1 is essential for p53 activation under genotoxic stress. Based on these findings, we design a MEIS1-derived peptide that suppresses cell proliferation and tumor growth by enhancing the stabilization of both MEIS1 and p53. Together, our findings provide a mechanistic framework for understanding the interplay between MEIS1, MDM2, and p53, highlighting the potential of targeting MEIS1 ubiquitination as a strategy for enhancing p53-dependent tumor suppression in CRC.
Material and methods
Animal experiments
Five-week-old male nude mice, obtained from the Experimental Animal Center of Central South University, were maintained in a pathogen-free environment. Prior to inoculation, the mice were randomly allocated into different experimental groups. Subcutaneous injection of 3 × 106 HCT8 or HCT116 cells, suspended in 100 μl of serum-free medium, was administered to the mice. Subsequently, after a period of 7 days since the inoculation, the designated peptides were administered intraperitoneally to the mice at a dose of 30 mg/kg once daily. Tumor dimensions were monitored three times weekly using an electronic caliper, and tumor volume was calculated employing the formula: total tumor volume (mm3) = 0.52 × length × width2. At the end of 12 days following the initial treatment, the mice were euthanized humanely, and their tumors were excised and weighed in a double-blinded evaluation method. Additionally, selected tumor samples underwent further analysis via Western blot assay and immunohistochemical staining.
Human tissue samples
Colorectal cancer (CRC) tissue specimens were obtained from Xiangya Hospital of Central South University. A total of 16 paired paraffin-embedded CRC tissues and adjacent non-tumorous tissues were included in this study.
Cell lines and cell culture
The human embryonic kidney cell line HEK-293T and several human CRC cell lines, including HCT116, HCT8, RKO and LoVo, were procured from reliable sources. Specifically, HEK-293T and HCT116 were obtained from the American Type Culture Collection, while HCT8, RKO and LoVo were generously provided by Professor Wancai Yang of Jining Medical University. Additionally, HCT116 P53+/+ and HCT116 P53−/− cell lines, which differ in their P53 gene status, were kindly supplied by Professor Mian Wu of the University of Science and Technology of China. All these cell lines were cultured in RPMI-1640 medium (BioInd, Beit Haemek, Israel), supplemented with 10% fetal bovine serum, and maintained in an incubator set at 37 °C with 5% CO2 to ensure optimal conditions for cell growth and proliferation.
Plasmid, siRNA, and transfection
To construct the required expression plasmids, we utilized the vectors pEGFP-N1, pCDNA3.1-Flag, and pCDNA3.1-HA. MEIS1 and P53 were cloned into the pEGFP-N1 and pCDNA3.1-Flag. MDM2 was cloned into the pEGFP-N1 and pCDNA3.1-HA. The mutants of MEIS1 or P53 were cloned into pEGFP-N1 vectors. MDM2-Flag, MDM2 1-199, MDM2 100-299, MDM2 299-391 plasmids were kindly provided by Professor Mian Wu (Jining Medical University). The various ubiquitin mutants (Ub-K6, Ub-K11, Ub-K27, Ub-K29, Ub-K33, Ub-K48, Ub-K63, Ub-K48R) and HA-Ub plasmids were kindly provided by Professor Ceshi Chen from the Kunming Institute of Zoology, Chinese Academy of Sciences. All the siRNAs used in our experiments were synthesized by RiboBio and were administered at a final concentration of 5 nM. For the transfection of plasmids or siRNAs, we employed Lipofectamine 2000 (Invitrogen, California, USA), following the manufacturer’s instructions.
Western blot (WB)
The WB assays were conducted in strict adherence to the previously established protocol [20]. The following antibodies were used: MEIS1 (1:1000, A18273, Abconal), p53 (1:5000, 60283-2-Ig, Proteintech), MDM2 (1:1000, ab16895, Abcam), p21 (1:1000, 10355-1-AP, Proteintech), BAX (1:1000, 50599-2-Ig, Proteintech), p53 p-ser15 (1:1000, ab223868, Abcam), γ-H2AX (1:1000, #7631S, Cell Signaling Technology), Ubiquitin K48 (1:1000, R24785, Zenbio), GFP (1:3000, UM3002, Utibody), Flag (1:3000, UM3009, Utibody), HA (1:3000, UM3004, Utibody), HERC2 (1:500, 27459-1-AP, Proteintech), GAPDH (1:5000, UM4002, Utibody). The secondary antibodies employed in this study were procured from HUABIO. The original WB images have been presented in the Original Data Files.
Quantitative real-time PCR (qPCR)
The qPCR assays were conducted in strict adherence to the previously established protocol [20]. Primer sequences are as followed: MEIS1 (F: CTGTTTGAAAGGGAAAATGCC; R: TGGAAGGGCCTGGGGTT); MDM2 (F: TTGATGAAAGCCTGGCTCTGTGTG; R: CGATGGCGTCCCTGTAGATTCAC); P53 (F: GCCCATCCTCACCATCATCACAC; R: GCACAAACACGCACCTCAAAGC); CDKN1A (F: GCCCGTGAGCGATGGAACTTC; R: CCTGCCTCCTCCCAACTCATCC); BAX (F: GATGCGTCCACCAAGAAGCTGAG; R: CACGGCGGCAATCATCCTCTG); GAPDH (F: CTGGGCTACACTGAGCACC; R: AAGTGGTCGTTGAGGGCAATG).
Immunoprecipitation (IP)
Cells were lysed using IP lysis buffer (Beyotime, China) supplemented with protease inhibitor cocktail (Roche, Swiss) and PMSF (Sigma, USA) on ice for 30 minutes to extract the protein. The resulting supernatants were then incubated with the designated antibodies overnight at 4 °C to enable specific binding interactions. On the following day, Protein A/G PLUS-Agarose beads (Santa Cruz, USA) were gently added to the protein-antibody complex mixture and incubated for another overnight period at 4 °C to enhance immunoprecipitation efficiency. Following incubation, the beads were thoroughly washed with the IP lysis buffer to remove unbound components. To elute the captured protein complexes, the beads were boiled in loading buffer, and the resulting extracts were analyzed via western blot techniques. The antibodies used for IP were as follows: MEIS1 (1:500, ab307544, Abcam), p53 (1:200, 60283-2-Ig, Proteintech), MDM2 (1:100, 66511-1-Ig, Proteintech), GFP (1:200, ab290, Abcam), Flag (1:100, UM3009, Utibody), HA (1:100, UM3004, Utibody).
Two-step co-immunoprecipitations were conducted essentially as described previously [21]. HEK293T cells were transfected with plasmids expressing Flag-MEIS1 (or no-tag MEIS1 as control), p53 and MDM2, and treated with MG132 (10 μM) for 6 h. The steps of the first immunoprecipitation are the same as above. Following centrifugation, the beads were thoroughly washed three times with IP lysis buffer to eliminate unbound components. The Flag-MEIS1 protein complex was then eluted using a 3× Flag peptide. Subsequently, the eluted fractions were subjected to a second round of immunoprecipitation, utilizing either anti-MDM2 antibodies or IgG as a control.
In vitro binding assay
In accordance with the instructions of the manufacturer, the TNT Quick Coupled Transcription/Translation System (Promega) was utilized to express the MEIS1 and MDM2 proteins. Binding assays were conducted by combining MEIS1 and MDM2 proteins, subsequently subjecting the mixture to IP using antibodies specific to either MEIS1 or MDM2, and finally verifying the interaction through WB utilizing antibodies against either MDM2 or MEIS1.
In vitro and in vivo ubiquitination assay
For the in vitro ubiquitination assay, the TNT Quick Coupled Transcription/Translation System (Promega) was employed to express the MEIS1, MEIS1 K178R mutant, MDM2, and p53 proteins, following the manufacturer’s recommended protocols. E1, UbcH5b, E3, substrate, reaction buffer, ubiquitin, ATP, and MgCl2 were mixed. After incubating the reaction mixture at 30 °C for 90 min, the reaction was terminated by adding loading buffer and heating the sample at 100 °C for 10 min.
For the in vivo ubiquitination assay, cells were transfected with the indicated constructs. 24 h after transfection, cells were treated with the proteasome inhibitor MG132 (10 mM) for 6 h. The cells were then collected and lysed using the IP lysis buffer. The ubiquitinated MEIS1 or p53 proteins in the lysates were immunoprecipitated using appropriate antibodies and detected via immunoblotting with anti-HA or anti-K48 antibodies.
Immunohistochemistry (IHC)
The IHC assays and evaluations were conducted in strict adherence to the previously established protocol [20]. The following antibodies were used: MEIS1 (1:200, ab19867, Abcam), p53 (1:1500, 60283-2-Ig, Proteintech).
Statistical analysis
All experiments were independently replicated at least three times, and the results are presented as mean ± SD or SEM. The sample size was determined based on statistical analysis of variance during experimental design. Statistical significance was assessed using GraphPad Prism 8.0. The threshold for statistical significance was set at p < 0.05, and the levels of significance are denoted as follows: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Detailed information about the materials and methods used in the present study is available in the Supplementary Materials and Methods.
Results
MEIS1 protein stability is regulated by E3 ubiquitin ligase MDM2
By analyzing MEIS1 mRNA and protein levels in tumor and paired non-tumor tissues from CRC patients, we found that although MEIS1 mRNA expression was elevated or unchanged in some cases (Fig. 1A), its protein levels were significantly downregulated (Fig. 1B). This discrepancy suggests that MEIS1 expression in CRC is regulated not only at the transcriptional level but also through post-translational mechanisms, potentially involving protein degradation pathways. To investigate the regulation of MEIS1 protein level in CRC, we analyzed the stability of MEIS1 protein following treatment with the protein synthesis inhibitor cycloheximide (CHX). Treatment of RKO and LoVo cells with CHX resulted in a time-dependent decline in MEIS1 protein levels (Fig. S1A), indicating that MEIS1 is an unstable protein in CRC cells.
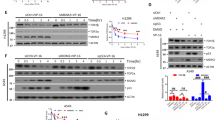
A qRT-PCR assay was performed to measure the mRNA expression level of MEIS1 in paired samples of tumor and paired non-tumor tissues from CRC patients. B Western blot assay was conducted to determine the protein expression level of MEIS1 in paired samples of tumor and paired non-tumor tissues from CRC patients. C Venn diagram depicting the common E3 ligases predicted to mediate MEIS1 degradation, based on IP/MS analysis and data from Ubibrowser. D Western blot analysis of MEIS1 protein expression in cells transfected with MDM2 or HERC2. E Cells transfected with MDM2 were treated with MG132 to inhibit proteasomal degradation, and MEIS1 protein level was subsequently measured by western blot to assess the impact of MG132 on MEIS1 stability. F Cell lysates were subjected to IP/IB using specific antibodies against MEIS1 or MDM2, along with IgG as a control, to detect the endogenous interaction between MEIS1 and MDM2. G HEK293T cells were co-transfected with HA-Ub, Flag-MEIS1, MDM2 WT or MDM2 C464A, followed by MG132 treatment. Flag-tagged proteins were immunoprecipitated, and the ubiquitination status of MEIS1 was analyzed.
Protein degradation primarily occurs via the ubiquitin-proteasome system or the autophagy-lysosome system [22]. To determine which pathway is responsible for MEIS1 degradation, we treated CRC cells with the proteasome inhibitor MG132 and the autophagy inhibitors MRT, CQ, and 3-MA. Western blot analysis revealed that inhibition of the proteasome pathway significantly increased MEIS1 protein levels (Fig. S1B). Furthermore, MG132 treatment markedly extended the half-life of MEIS1 (Fig. S1C). In addition, ubiquitination assays confirmed that MEIS1 undergoes substantial ubiquitination in CRC cells (Fig. S1D). These data indicated that MEIS1 degradation in CRC is primarily mediated via the ubiquitin-proteasome system.
Given MEIS1 is degraded through ubiquitin-proteasome system, we sought to identify its potential E3 ubiquitin ligases. Immunoprecipitation and mass spectrometry (IP/MS) analyses identified several E3 ligases interacting with MEIS1, including MDM2, RNF2, TRIM41, ARIH1, RFWD3, HERC2, and TRIM71 (Fig. S2A and Supplementary Table 1). Additionally, bioinformatics predictions using the Ubibrowser database suggested several candidate E3 ligases for MEIS1 (Fig. S2B). A Venn diagram comparing the experimentally identified and predicted E3 ligases revealed two common candidates: MDM2 and HERC (Fig. 1C).
To determine which of these E3 ligases regulates MEIS1 stability, we overexpressed MDM2 or HERC2 in CRC cells and analyzed their effects on MEIS1 protein levels. Notably, only MDM2 overexpression led to a significant reduction in MEIS1 protein expression (Fig. 1D). Conversely, MDM2 knockdown resulted in a marked increase in MEIS1 protein level (Fig. S2C). To exclude the possibility that MDM2 affects MEIS1 at the transcriptional level, we performed qPCR analysis and confirmed that MDM2 knockdown did not alter MEIS1 mRNA level (Fig. S2D). These findings suggest that MDM2 regulates MEIS1 expression by modulating its protein stability rather than its transcription.
To further confirm that MDM2-mediated MEIS1 degradation occurs via the ubiquitin-proteasome system, we treated MDM2-overexpressing cells with MG132. Western blot analyses revealed that MDM2 downregulated MEIS1 protein level under normal conditions, whereas MG132 treatment effectively rescued MEIS1 expression (Fig. 1E), reinforcing the role of MDM2 in ubiquitin-proteasome system-mediated MEIS1 degradation.
Next, we assessed the impact of MDM2 on MEIS1 protein stability by measuring its half-life. CRC cells transfected with either MDM2 overexpression (oe-MDM2) or MDM2 knockdown (si-MDM2) constructs were treated with CHX. The results revealed that MDM2 overexpression significantly reduced the half-life of MEIS1, whereas MDM2 knockdown prolonged it (Fig. S3A, B).
To elucidate the molecular mechanism underlying MDM2-mediated MEIS1 degradation, we performed co-immunoprecipitation (Co-IP) assays, which confirmed that MDM2 physically interacts with MEIS1 at both endogenous (Fig. 1F) and exogenous (Fig. S3C) levels. To further investigate the role of MDM2’s E3 ubiquitin ligase activity in MEIS1 ubiquitination, we examined the effect of the MDM2 C464A mutant, which lacks E3 ligase activity. Notably, this mutant failed to induce MEIS1 ubiquitination, indicating that its E3 ligase activity is required for this process. Moreover, although the MDM2 C464A mutant retained its binding affinity for MEIS1, it was unable to promote MEIS1 degradation (Fig. 1G), further confirming that MDM2-mediated MEIS1 ubiquitination is crucial for its proteasomal degradation.
MDM2 promotes K48-linked polyubiquitination of MEIS1
Ubiquitination of substrate proteins occurs through seven lysine residues on ubiquitin (K6, K11, K27, K29, K33, K48, and K63), each of which contributes to formation distinct polyubiquitin chain with specific functional consequences [23]. To determine the specific type of polyubiquitination mediated by MDM2 on MEIS1, we co-transfected HEK293T cells with MEIS1, MDM2, and various ubiquitin mutants, each containing only a single intact lysine residue. Western blot analysis revealed that MDM2 promoted MEIS1 ubiquitination in the presence of HA-tagged wild-type ubiquitin and K48-only ubiquitin (Fig. S4A). Conversely, MEIS1 ubiquitination was significantly reduced in the presence of K48R ubiquitin (Fig. 2A), indicating that MDM2-mediated MEIS1 ubiquitination is predominantly K48-linked.
A HEK293T cells expressing MEIS1-GFP were co-transfected with MDM2-Flag and Ub/K48/K48R-HA. Cellular extracts were immunoprecipitated with anti-GFP antibody and the ubiquitination level of MEIS1 was detected by HA antibody. Cells expressing MEIS1-Flag were co-transfected with MDM2-HA (B) or siMDM2 (C), followed by treatment with MG132. The ubiquitination level of MEIS1 was measured. D MEIS1 and MDM2 proteins were obtained through in vitro transcription and translation system. An in vitro ubiquitination assay was conducted, both with and without the presence of MDM2. Ubiquitination status of MEIS1 was detected via western blot using a specific MEIS1 antibody. The interaction between MEIS1 and MDM2 was assayed by IP with MEIS1 or MDM2 antibody, followed by western blot with MDM2 or MEIS1 antibody.
To further validate this finding, we utilized an antibody that specifically recognizes K48-linked polyubiquitin chains. The results demonstrated that MDM2 overexpression markedly increased K48-linked ubiquitination of MEIS1 (Fig. 2B). In contrast, MDM2 knockdown resulted in a substantial reduction in MEIS1 ubiquitination compared to control cells (Fig. 2C), further supporting the role of MDM2 in facilitating K48-linked polyubiquitination.
Moreover, in vitro binding and ubiquitination assays confirmed that MDM2 directly interacts with MEIS1 and efficiently promotes its ubiquitination (Fig. 2D). Collectively, these findings establish MDM2 as a key E3 ubiquitin ligase for MEIS1, facilitating its K48-linked polyubiquitination and subsequent degradation via the proteasome pathway.
The Ring domain of MDM2 binds to the Ser/Thr-rich domain of MEIS1 and promotes ubiquitination at K178
To elucidate the structural basis of the MDM2–MEIS1 interaction, we performed Co-IP assays using a series of MDM2 deletion constructs to assess their binding affinity to MEIS1 (Fig. S5A). Our results demonstrated that MDM2 interacts with MEIS1 primarily through its Ring domain (Fig. 3A). Notably, among the truncated MDM2 constructs, only the Ring domain was sufficient to promote K48-linked polyubiquitination of MEIS1, akin to full-length MDM2 (Fig. S5B), indicating that the Ring domain is essential for MDM2-mediated MEIS1 ubiquitination.
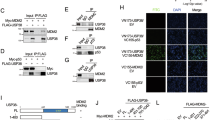
A MDM2 and its truncates were co-transfected with MEIS1 into HEK293T cells, followed by detection of the binding between MEIS1 and the MDM2 domain. MEIS1 and its truncates were co-transfected with MDM2 into HEK293T cells, followed by detection of MDM2 binding to the MEIS1 domain (B) or the ubiquitination level of MEIS1 (C). D MEIS1-WT, K72R, K75R, K93R, K126R, K132R, K162R, K178R or K180R were co-transfected with Flag-MDM2 into HEK293T cells and treated with MG132. The ubiquitination level of MEIS1 was detected. E Schematic representation of MEIS1 and MDM2 binding domains.
To further determine the MEIS1 domain responsible for binding MDM2, we generated six MEIS1 deletion mutants based on its structural domains (Fig. S6A). Co-IP assays demonstrated that MEIS1 interacts with MDM2 through its Ser/Thr-rich domain (Fig. 3B). Notably, ubiquitination assays showed that deletion of this domain completely abolished MEIS1 ubiquitination (Fig. 3C). Surprisingly, although deletion of the PBX interaction domain did not disrupt the interaction between MEIS1 and MDM2, it resulted in a marked reduction in overall ubiquitination level of MEIS1 (Fig. 3C), suggesting that this domain may contain a key ubiquitination site.
Subsequently, to identify the specific lysine residue required for MDM2-mediated ubiquitination, we generated a series of lysine-to-arginine (K-to-R) mutants within the PBX interaction domain (Fig. S6B). Among these mutants, only the K178R substitution completely abolished MDM2-induced K48-linked polyubiquitination of MEIS1 (Fig. 3D), indicating that K178 is the primary ubiquitination site. Importantly, the K178 mutation did not affect the MEIS1-MDM2 interaction (Fig. S6C), supporting the hypothesis that the PBX region is not involved in the interaction between these two proteins but rather contains a key ubiquitination site necessary for MDM2-mediated MEIS1 degradation.
In summary, these findings demonstrate that MDM2 interacts with MEIS1 via its Ring domain, binding to the Ser/Thr-rich domain of MEIS1, thereby promoting K48-linked polyubiquitination at K178 within the PBX interaction domain, ultimately leading to MEIS1 degradation (Fig. 3E).
MEIS1 stabilizes p53 protein by suppressing its ubiquitination
As a pivotal guardian of the genome, p53 plays a central role in regulating key cellular processes, including cell cycle regulation, DNA damage response, and apoptosis [24]. MDM2, an E3 ubiquitin ligase, promotes the ubiquitination of p53, leading to its proteasomal degradation and thereby modulating its tumor suppressor function. Upon reviewing our IP/MS analysis results, we identified p53 as a potential MEIS1-associated protein (Supplementary Table 1). Given our previous findings that MEIS1 acts as a tumor suppressor in CRC—regulating cell proliferation, cell cycle progression, and drug resistance [20]—and considering that these functions align with those of p53, we hypothesized a functional link between MEIS1 and p53. Supporting this hypothesis, Gene Set Enrichment Analysis (GSEA) revealed a positive correlation between high MEIS1 expression and activation of the p53 signaling pathway in CRC (Fig. S7A).
To explore the regulatory effect of MEIS1 on p53, we manipulated MEIS1 levels in CRC cells and examined both the mRNA and protein levels of p53. qRT-PCR analysis revealed no significant changes in TP53 mRNA level upon MEIS1 overexpression or knockdown, compared to controls (Fig. 4A). However, the mRNA levels of CDKN1A and BAX, two canonical p53 downstream targets, were markedly decreased upon MEIS1 knockdown and significantly elevated upon MEIS1 overexpression (Fig. 4A). These findings suggest that MEIS1 does not regulate p53 at the transcriptional level but may instead influence its protein stability.
A The mRNA expressions of P53, CDKN1A and BAX were analyzed by qPCR in cells with MEIS1 overexpression or knockdown. B The protein levels of p53, p21 and BAX were detected by western blot in cells with MEIS1 overexpression or knockdown. The ubiquitination level of p53 was measured by IP/IB assays in cells with MEIS1 overexpression (C) or knockdown (D).
To further investigate this, we assessed p53 protein level following MEIS1 modulation. Western blot analysis revealed that MEIS1 overexpression significantly increased the protein levels of p53, p21, and BAX, whereas MEIS1 depletion led to a marked reduction in their expression (Fig. 4B). Since p53 protein level is tightly regulated by proteasomal degradation, we hypothesized that MEIS1 may modulate p53 stability at the post-translational level.
To test this hypothesis, we examined whether MEIS1 affects the ubiquitination of endogenous p53. Overexpression of MEIS1 in CRC cells led to a substantial reduction in p53 ubiquitination, supporting the idea that MEIS1 protects p53 from proteasomal degradation (Figs. 4C and S7B). Conversely, MEIS1 knockdown significantly enhanced the ubiquitination of p53 (Figs.4D and S7C), further reinforcing the notion that MEIS1 inhibits p53 degradation by suppressing its ubiquitination. Together, these findings demonstrate that MEIS1 enhances p53 protein stability by inhibiting its ubiquitination, thereby preventing its proteasomal degradation.
Non-competitive complex formation of MEIS1, MDM2 and p53
Given the potential role of MEIS1 in binding to either p53 or MDM2, we further explored the interactions among these three proteins. Specifically, we investigated the interaction between endogenous MDM2 and p53 in the context of MEIS1 overexpression. Our findings revealed that the interaction between MDM2 and p53 was independent of MEIS1 expression level (Fig. 5A). Furthermore, overexpression of p53 did not alter the interaction between MEIS1 and MDM2, nor did overexpression of MDM2 affect the interaction between MEIS1 and p53 (Fig. 5B, C). These findings suggest that MEIS1 interacts with both p53 and MDM2 in the endogenous context, potentially forming a ternary complex. To confirm this hypothesis, we performed two-step Co-IP assays, which demonstrated that MEIS1 binds simultaneously to both p53 and MDM2 (Fig. 5D), supporting the existence of a ternary complex comprising MEIS1, p53, and MDM2.
A The interaction of endogenous p53 and MDM2 was detected by IP/IB assays in cells with MEIS1 overexpression. B The interaction of endogenous MEIS1 and MDM2 was detected by IP/IB assays in cells with p53 overexpression. C The interaction of endogenous p53 and MEIS1 was detected by IP/IB assays in cells with p53 overexpression. D HEK293T cells were co-transfected with Flag-MEIS1 (or no-tag MEIS1 as control), p53 and MDM2 then treated with MG132. Two-step co-immunoprecipitation identifies a MEIS1-p53-MDM2 complex. The procedure is outlined in the box at the top. E MEIS1 and its truncates were co-transfected with p53 into HEK293T cells, followed by detection of the binding between p53 and the MEIS1 domain. F p53 and its truncates were co-transfected with MEIS1 into HEK293T cells, followed by detection of the binding between MEIS1 and the p53 domain. G Schematic representation of MEIS1, p53 and MDM2 binding domains.
Next, to identify the specific regions involved in the interaction between MEIS1 and p53, we utilized a series of MEIS1 mutants. Upon transfection of these mutants into cells overexpressing p53 and subjected to Co-IP, we found that, in addition to the full-length MEIS1, the deletion of the Asp/Glu-rich domain alone disrupted its interaction with p53 (Fig. 5E). This emphasizes the critical role of the Asp/Glu-rich domain in mediating the binding of MEIS1 to p53.
To further characterize the regions of p53 involved in its interaction with MEIS1, we generated a series of p53 deletion mutants (Fig. S8A). Co-IP results revealed that the Regulatory domain of p53 is essential for the interaction with MEIS1 (Fig. 5F). In summary, our results indicate that MEIS1 uses its Asp/Glu-rich domain to bind to the Regulatory domain of p53, while its Ser/Thr-rich domain mediates the interaction with the Ring domain of MDM2 (Fig. 5G).
MEIS1 mutants lacking ubiquitination capability fail to shield p53 from MDM2-mediated ubiquitination
To investigate the mechanism by which MEIS1 stabilizes p53, we examined whether p53, which is also a substrate of MDM2, could influence MEIS1 stability. Co-IP assays confirmed that, consistent with our previous findings, MEIS1 significantly inhibited MDM2-mediated p53 ubiquitination (Fig. S9A, B). In contrast, p53 had no effect on the ubiquitination of MEIS1 (Fig. S9C, D).
We next assessed the effect of the MEIS1 K178R mutant, which lacks ubiquitination capability on MDM2-mediated p53 ubiquitination. Our results indicated that MDM2-mediated ubiquitination of MEIS1-WT suppressed p53 ubiquitination. However, despite retaining its ability to bind p53, the K178R mutant failed to inhibit MDM2-mediated p53 ubiquitination (Figs. 6A and S9E). Given the role of ubiquitination in regulating p53 stability, it was not surprising that the K178R mutant was unable to elevate p53 protein level (Fig. 6B). Moreover, whereas MEIS1-WT effectively prevented MDM2-induced p53 downregulation, mutants lacking ubiquitination capability lost this protective effect (Figs. 6C and S9F). Additionally, MEIS1 WT also extended the half-life of p53, while the K178R mutant had no effect on p53 stability (Fig. S9G). These findings suggest that MDM2-mediated ubiquitination of MEIS1, particularly at K178, is critical for stabilizing p53.
A MDM2-HA, MEIS1 WT/K178R were co-transfected with p53 into HEK293T cells and treated with MG132. The ubiquitination levels of p53 and MEIS1 were detected. B Cells overexpressing MEIS1 WT or the K178R mutant variant were analyzed by western blot to determine the protein level of p53. C HCT116 p53–/– cells were co-transfected with MDM2-HA, MEIS1 WT or K178R, and p53. The protein levels of p53 were analyzed. D MDM2, p53 and MEIS1 WT or MEIS1 ΔSer/Thr-rich or MEIS1 ΔAsp/Glu-rich were co-transfected. After MG132 treatment for 6 hours, the ubiquitination level of p53 was detected by IP/IB assay. E Schematic diagram of the hypothesis that MEIS1 protects p53 from MDM2-mediated degradation. F To acquire the proteins of MEIS1 WT/K178R, MDM2, and p53, an in vitro transcription and translation approach was utilized. An in vitro ubiquitination assay was conducted, incorporating Ub, E1, UbcH5b, E3(MDM2), p53, and the MEIS1 WT/K178R. The ubiquitination status of both MEIS1 and p53 was then analyzed by western blot.
To assess whether the protective effect of MEIS1 on p53 requires direct interaction with p53, we co-transfected MDM2 and p53 in HEK 293 T cells, along with MEIS1-WT, MEIS1 Ser/Thr deletion mutants (which lose the ability to bind MDM2), or MEIS1 Asp/Glu deletion mutants (which lose the ability to bind p53) (Figs. 6D and S9H). Compared with the control, co-expression of MEIS1-WT with MDM2 and p53 enhanced MEIS1 ubiquitination while significantly reducing p53 ubiquitination (Fig. 6D, lane3). However, when the Ser/Thr deletion mutants were expressed, MEIS1 ubiquitination decreased, its protective effect on p53 was lost, and p53 ubiquitination increased significantly (Fig. 6D, lane4). In contrast, the Asp/Glu deletion mutants, despite their inability to bind p53, still protected it from MDM2-mediated ubiquitination (Fig. 6D, lane5) These results suggest that the protective effect of MEIS1 on p53 does not require direct interaction between MEIS1 and p53, but instead relies on MEIS1’s own ubiquitination-dependent mechanism.
The Ring domain of MDM2 directly interacts with the E2-conjugated enzyme, enabling the transfer of ubiquitin to its target substrate [25]. Given that MEIS1 binds to this domain and undergoes ubiquitination at residues near the MDM2 binding site, we hypothesized that MEIS1 may act as a dominant substrate for MDM2. This proximity likely biases ubiquitin transfer toward MEIS1, thereby reducing p53 ubiquitination (Fig. 6E). To test this hypothesis, we performed in vitro ubiquitination assays, which confirmed that both MEIS1 and p53 are ubiquitinated by MDM2 (Fig. 6F, lane3 and lane6). Notably, MEIS1-WT effectively inhibited MDM2-mediated p53 ubiquitination (Fig. 6F, lane4), whereas the K178R mutant of MEIS1 lost its ability to block p53 ubiquitination (Fig. 6F, lane5).
Taken together, these results suggest that the K178 residue in MEIS1 functions as a switch for ubiquitin transfer, directing MDM2-mediated ubiquitination and thereby regulating the stability of p53.
The crucial role of MDM2-mediated MEIS1 ubiquitination in activating p53 during DNA damage response
The activation of p53 is a central event in the DNA damage response and plays a critical role in determining cell fate [26]. Because p53 activation is tightly controlled by both post-translational modifications and MDM2-dependent degradation, we next investigated how MEIS1 contributes to p53 stabilization under genotoxic stress.
To assess the impact of DNA damage on MEIS1 and p53 expression, we treated HCT8 and HCT116 cells with increasing concentrations of doxorubicin (DOX), a chemotherapeutic agent known to induce DNA damage and enhance p53 expression [27]. Both MEIS1 and p53 mRNA and protein levels exhibited a dose-dependent increase following DOX treatment (Fig. S10A, B). MEIS1 was similarly induced in HCT116 p53–/– cells (Fig. S10C), demonstrating that its upregulation is p53-independent. Consistently, Co-IP analysis showed that DNA damage did not weaken the MEIS1-MDM2 interaction (Fig. S10D), indicating that increased MEIS1 protein levels do not arise from dissociation from MDM2 but more likely reflect transcriptional induction triggered by DNA damage. Functionally, MEIS1 knockdown markedly reduced DOX-induced p53 accumulation (Figs. 7A and S11A). Importantly, MEIS1 depletion only partially attenuated DOX-mediated p53 induction, suggesting that MEIS1 does not promote p53 activation by increasing DNA damage, but rather acts through a direct post-translational regulatory mechanism.
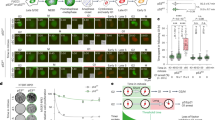
A RKO cells transfected with siMEIS1 were subjected to 1 μM DOX, and the protein levels of p53, p21, and BAX were analyzed by western blot. B HCT116 and HCT8 cells transfected with siMEIS1 were treated with 1 μM DOX, and the total and phosphorylated p53 levels were examined by western blot. C HCT116 and HCT8 cells with reduced MEIS1 expression were treated with ATM/ATR inhibitors and 1 μM DOX, followed by western blot analysis of total and phosphorylated p53. D HCT116 cells expressing MEIS1-WT or the K178R mutant were treated with DOX at the indicated concentrations, and p53, p21, and BAX protein levels were measured by western blot. E RKO cells co-transfected with MEIS1-WT or K178R and siMEIS1 were treated with 1 μM DOX, and p53, p21 and BAX levels were assessed by western blot. F RKO cells transfected with siMEIS1 were treated with DOX (1 μM), etoposide (20 μM), or cisplatin (20 μM), and the protein levels of p53, p21, and BAX were evaluated by western blot.
A well-established mechanism of p53 activation in response to DNA damage involves phosphorylation at Ser15, Thr18, or Ser20, which reduces MDM2 binding and enhances p53 stability [6]. Given the essential role of post-translational modifications in p53 activation, we evaluated total and phosphorylated p53 in MEIS1-deficient cells. DOX robustly induced both p53 phosphorylation and total protein accumulation; however, MEIS1 knockdown significantly reduced total p53 while leaving phosphorylation largely unchanged (Fig. 7B). These data indicate that MEIS1 does not influence upstream ATM/ATR signaling but acts downstream of phosphorylation to stabilize p53.
To further dissect this mechanism, we suppressed p53 phosphorylation using ATM/ATR inhibitors. Although the inhibitors effectively reduced DOX-induced p53 phosphorylation, total p53 levels remained elevated in control cells, likely due to increased MEIS1 expression. In contrast, MEIS1 depletion led to a marked reduction in total p53 under these conditions (Fig. 7C), confirming that MEIS1 acts downstream of canonical DDR signaling and stabilizes p53 through an MDM2-dependent mechanism rather than by modulating its phosphorylation.
We then assessed the requirement of MEIS1 ubiquitination for p53 activation. Consistent with its ability to inhibit p53 ubiquitination, MEIS1-WT maintained strong DOX-induced p53 activity, as reflected by elevated p53, p21, and BAX expression. By contrast, the ubiquitination-deficient MEIS1 K178R mutant failed to support robust p53 activation (Figs. 7D and S11B), highlighting the critical role of MEIS1 ubiquitination in p53 activation. Similarly, MEIS1 knockdown markedly reduced DOX-induced p53 expression. Overexpression of MEIS1-WT rescued p53 protein level, while overexpression of the K178R mutant failed to restore p53 expression in RKO and LoVo cells under DOX treatment (Figs. 7E and S11C). These findings establish MEIS1 ubiquitination as a prerequisite for its ability to protect p53 from MDM2-mediated degradation during DNA damage.
To address whether MEIS1 regulation of p53 is stimulus-specific, we examined p53 activation under three DNA-damaging conditions (DOX, etoposide, cisplatin). All stimuli induced MEIS1, p53, p21, and BAX, and MEIS1 knockdown partially attenuated these inductions (Figs. 7F and S11D). These results establish MEIS1 as a general modulator of the p53 stabilization during genotoxic stress.
Taken together, these findings support a model in which MDM2-mediated ubiquitination of MEIS1 serves as a key regulatory mechanism that facilitates and prolongs p53 stabilization in response to DNA damage.
MEIS1-derived peptide suppresses CRC growth by stabilizing MEIS1 and p53
As previously mentioned, both MEIS1 and the E2 enzyme bind to the Ring domain of MDM2, and their close proximity likely favors ubiquitin transfer to MEIS1, thus reducing p53 ubiquitination. Since the K178 site of MEIS1 is essential for preventing p53 degradation, we designed and synthesized a peptide (NFCHRYISCLK) based on this site, along with three control peptides, all labeled with fluorescein isothiocyanate (FITC). Fluorescence microscopy confirmed that MEIS1-derived peptides were efficiently internalized by cells compared with the control peptides (Fig. S12A). Western blot analysis further revealed that treatment with MEIS1-derived peptides led to a dose-dependent upregulation of MEIS1 and p53 protein levels. (Figs. 8A and S12B, C).
A Cells were treated with MEIS1 peptide at the indicated concentrations, and then the protein level of MEIS1 and p53 were detected. B Cells were treated with MEIS1 peptide at the indicated concentrations, followed by treatment with CHX for various durations. The half-lives of MEIS1 and p53 protein were measured. C Subcutaneous xenografts of cells were harvested from each mouse intraperitoneally injected with PBS or MEIS1 peptide. Scatterplot and curve show the growth and weight of xenografts for each group. D A schematic illustration outlines the pivotal molecular mechanisms that underlie the intricate regulation of the MDM2-p53 axis by MEIS1 in CRC.
To determine whether MEIS1-derived peptides influence protein stability, CHX chase assays were performed, revealing a significant extension of MEIS1 and p53 half-life following peptide treatment (Fig. 8B). Additionally, IP assays showed that a marked reduction in the endogenous ubiquitination levels of MEIS1 and p53 upon the peptide treatment (Fig. S12D), further supporting the notion that MEIS1-derived peptides stabilize these proteins by preventing their degradation.
Next, we examined the functional impact of MEIS1-derived peptides on CRC cell proliferation. CCK8 assays demonstrated a significant inhibition of cell growth in vitro (Fig. S13A). Furthermore, to evaluate the anti-tumor efficacy in vivo, the peptide was administered via intraperitoneal injection in nude mice bearing CRC xenografts. Notably, MEIS1 peptide treatment led to a significant suppression of tumor growth in vivo (Fig. 8C). Consistently, IHC (Fig. S13B) and WB (Fig. S13C) analyses revealed increased MEIS1 and p53 protein levels in xenograft tumors from MEIS1 peptide-treated mice. These findings indicate that MEIS1-derived peptides exert their anti-tumor effects by enhancing MEIS1 and p53 expression, thereby inhibiting CRC growth.
Discussion
In this study, we identify MEIS1 as a previously unrecognized substrate of MDM2 and reveal a regulatory mechanism in which MEIS1 competes with p53 for MDM2-mediated ubiquitination. By diverting MDM2 catalytic activity toward itself, MEIS1 prevents p53 ubiquitination and degradation, thereby enhancing p53 stability and transcriptional output (Fig. 8D). This mechanism provides an additional layer of control over p53 beyond conventional MDM2-p53 disruption strategies and highlights MEIS1 ubiquitination as a previously overlooked determinant of p53 function in CRC.
The role of MEIS1 in cancer has been recognized as context-dependent, exhibiting both oncogenic [28,29,30] and tumor-suppressive [31, 32] properties. This dual role highlights the complexity of MEIS1’s function in cellular processes and underscores the need for a deeper understanding of its expression and regulatory mechanisms. Our previous work has shown that MEIS1 suppresses tumor growth and enhances sensitivity to oxaliplatin in CRC by regulating FEN1. Additionally, MEIS1 expression is epigenetically silenced in CRC via the lncRNA ELFN1-AS1, which recruits the EZH2-DNMT3a complex to its promoter, leading to DNA methylation and histone modification [20]. Despite these insights, the post-translational regulation of MEIS1 has remained largely unexplored. Our study addresses this gap by identifying the ubiquitin-proteasome system as a key pathway controlling MEIS1 stability in CRC. Specifically, we demonstrate that MDM2 functions as a novel E3 ligase for MEIS1, targeting it for ubiquitination at K178, triggering its degradation. Importantly, this modification is essential not only for MEIS1 degradation but also for preventing MDM2-mediated p53 ubiquitination, establishing MEIS1 as a key stabilizer of p53.
In addition to revealing MEIS1 as a substrate of MDM2, our study provides important mechanistic insights into why MDM2 preferentially ubiquitinates MEIS1. The Ring domain of MDM2 plays a pivotal role in its ubiquitin ligase activity [33, 34], as it recruits the E2 ubiquitin-conjugating enzyme in its thioesterified state, facilitating the transfer of ubiquitin to the target substrate [35]. Our findings show that MEIS1 binds to the Ring domain of MDM2 and undergoes ubiquitination at residues near the MDM2 binding site, suggesting that the proximity between these regions enhances the efficiency of ubiquitination. This interaction not only drives MEIS1 degradation but also provides a mechanism by which MDM2 selectively regulates p53 stability through the modulation of MEIS1 availability as a competitive substrate. Furthermore, this raises the intriguing possibility that MEIS1 may influence the ubiquitination of other MDM2 substrates, potentially modulating additional oncogenic or tumor-suppressive pathways.
Beyond its role in p53 stabilization, our study also highlights the involvement of MEIS1 in the DNA damage response. We demonstrate that MEIS1 ubiquitination by MDM2 is essential for p53 activation under genotoxic stress, suggesting that MEIS1 acts as a molecular sensor that modulates MDM2 activity in response to cellular stress. This regulatory interplay between MEIS1, MDM2, and p53 may represent a broader paradigm governing p53 activation under both physiological and pathological conditions. Future studies should explore whether MEIS1-mediated p53 stabilization extends to other stress responses and whether this pathway is conserved across different cancer types.
While targeting the MDM2-p53 axis has long been a therapeutic strategy in cancer, small-molecule MDM2 inhibitors often induce feedback mechanisms, leading to upregulation of MDM2 and limiting the efficacy of these drugs, while also causing dose-limiting toxicities [36, 37]. Furthermore, because MDM2 ubiquitinates multiple substrates, selective inhibition remains challenging [13, 14]. Importantly, our clinical analyses reveal that a higher proportion of patients with p53-WT was observed in the MEIS1-low group compared with the MEIS1-high group. (Fig. S14A and Supplementary Table 2). This pattern underscores a specific dependency on the MEIS1-MDM2-p53 regulatory axis in p53-WT tumors, providing a rationale for therapeutic strategies aimed at restoring MEIS1 function. Unlike traditional MDM2 inhibitors, which often induce compensatory MDM2 upregulation, stabilizing MEIS1 provides an alternative approach by competitively inhibiting MDM2-mediated p53 ubiquitination without disrupting MDM2-p53 interaction directly. Notably, our designed MEIS1-derived peptide significantly inhibits CRC growth by stabilizing MEIS1 and p53. These results underscore the potential of MEIS1-based therapies, particularly for p53 wild-type tumors. Further studies are required to optimize the pharmacokinetics, specificity, and delivery strategies of these therapeutics for clinical application.
Despite these promising insights, several questions remain. First, although MEIS1 acts as a dominant substrate for MDM2 in CRC, it will be important to determine how MEIS1 stabilization influences the ubiquitination of other MDM2 substrates and whether this contributes to tumor suppression. Second, given the dual role of MEIS1 in different cancer types, it is crucial to determine the factors that dictate its oncogenic versus tumor-suppressive functions in specific tumor contexts. A deeper understanding of these regulatory mechanisms will be crucial for optimizing therapeutic strategies targeting MEIS1. Finally, although our data establish MEIS1 as a critical regulator of p53 in CRC, the extent to which this mechanism is conserved in other p53-WT tumors remains an important question for future exploration.
In conclusion, our study identifies MDM2-mediated ubiquitination of MEIS1 as a key regulatory switch controlling p53 stability and activity. By acting as a dominant substrate for MDM2, MEIS1 protects p53 from degradation and reinforces its tumor-suppressive function in CRC. These findings expand the current understanding of p53 regulation and highlight the potential of targeting MEIS1 ubiquitination as a therapeutic strategy for p53-dependent cancers.
Data availability
All data required for the comprehensive evaluation of the conclusions presented in this paper are included either within the paper itself or in the accompanying Supplementary Materials. Should any additional data be required, the authors are readily available to furnish it upon request.
References
Lagunas-Rangel FA. Role of circular RNAs in DNA repair. RNA Biol. 2024;21:149–61.
Jackson SP, Bartek J. The DNA-damage response in human biology and disease. Nature. 2009;461:1071–8.
Gungordu S, Aptullahoglu E. Targeting MDM2-mediated suppression of p53 with idasanutlin: a promising therapeutic approach for acute lymphoblastic leukemia. Investig N Drugs. 2024;42:603–11.
Kastenhuber ER, Lowe SW. Putting p53 in context. Cell. 2017;170:1062–78.
Zhang H, Xu J, Long Y, Maimaitijiang A, Su Z, Li W et al. Unraveling the guardian: p53’s multifaceted role in the DNA damage response and tumor treatment strategies. Int J Mol Sci. 2024;25:12928.
Liu Y, Su Z, Tavana O, Gu W. Understanding the complexity of p53 in a new era of tumor suppression. Cancer Cell. 2024;42:946–67.
Wang M, Attardi LD. A balancing act: p53 activity from tumor suppression to pathology and therapeutic implications. Annu Rev Pathol. 2022;17:205–26.
Laos M, Sulg M, Herranen A, Anttonen T, Pirvola U. Indispensable role of Mdm2/p53 interaction during the embryonic and postnatal inner ear development. Sci Rep. 2017;7:42216.
Manfredi JJ. Mdm2 and MdmX: partners in p53 destruction. Cancer Res. 2021;81:1633–4.
Liu Y, Tavana O, Gu W. p53 modifications: exquisite decorations of the powerful guardian. J Mol Cell Biol. 2019;11:564–77.
Liu J, Guan D, Dong M, Yang J, Wei H, Liang Q, et al. UFMylation maintains tumour suppressor p53 stability by antagonizing its ubiquitination. Nat Cell Biol. 2020;22:1056–63.
Yao Y, Zhang Q, Li Z, Zhang H. MDM2: current research status and prospects of tumor treatment. Cancer Cell Int. 2024;24:170.
Venkatesh D, O’Brien NA, Zandkarimi F, Tong DR, Stokes ME, Dunn DE, et al. MDM2 and MDMX promote ferroptosis by PPARα-mediated lipid remodeling. Genes Dev. 2020;34:526–43.
Zafar A, Khan MJ, Naeem A. MDM2- an indispensable player in tumorigenesis. Mol Biol Rep. 2023;50:6871–83.
Meriç N, Kocabaş F. The historical relationship between Meis1 and leukemia. Adv Exp Med Biol. 2022;1387:127–44.
Geerts D, Schilderink N, Jorritsma G, Versteeg R. The role of the MEIS homeobox genes in neuroblastoma. Cancer Lett. 2003;197:87–92.
Okumura K, Saito M, Isogai E, Aoto Y, Hachiya T, Sakakibara Y, et al. Meis1 regulates epidermal stem cells and is required for skin tumorigenesis. PLoS ONE. 2014;9:e102111.
VanOpstall, C, Perike, S, Brechka, H, Gillard, M, Lamperis, S, Zhu, B et al. MEIS-mediated suppression of human prostate cancer growth and metastasis through HOXB13-dependent regulation of proteoglycans. Elife. 2020;9:e53600.
Li W, Huang K, Guo H, Cui G. Meis1 regulates proliferation of non-small-cell lung cancer cells. J Thorac Dis. 2014;6:850–5.
Li Y, Gan Y, Liu J, Li J, Zhou Z, Tian R, et al. Downregulation of MEIS1 mediated by ELFN1-AS1/EZH2/DNMT3a axis promotes tumorigenesis and oxaliplatin resistance in colorectal cancer. Signal Transduct Target Ther. 2022;7:87.
Wang SP, Wang WL, Chang YL, Wu CT, Chao YC, Kao SH, et al. p53 controls cancer cell invasion by inducing the MDM2-mediated degradation of Slug. Nat Cell Biol. 2009;11:694–704.
Nedelsky NB, Todd PK, Taylor JP. Autophagy and the ubiquitin-proteasome system: collaborators in neuroprotection. Biochim Biophys Acta. 2008;1782:691–9.
Han S, Wang R, Zhang Y, Li X, Gan Y, Gao F, et al. The role of ubiquitination and deubiquitination in tumor invasion and metastasis. Int J Biol Sci. 2022;18:2292–303.
Levine AJ, Oren M. The first 30 years of p53: growing ever more complex. Nat Rev Cancer. 2009;9:749–58.
Spano D, Catara G. Targeting the ubiquitin-proteasome system and recent advances in cancer therapy. Cells. 2023;13:29.
Helton ES, Chen X. p53 modulation of the DNA damage response. J Cell Biochem. 2007;100:883–96.
Shamseddine AA, Clarke CJ, Carroll B, Airola MV, Mohammed S, Rella A, et al. P53-dependent upregulation of neutral sphingomyelinase-2: role in doxorubicin-induced growth arrest. Cell Death Dis. 2015;6:e1947.
Argiropoulos B, Yung E, Xiang P, Lo CY, Kuchenbauer F, Palmqvist L, et al. Linkage of the potent leukemogenic activity of Meis1 to cell-cycle entry and transcriptional regulation of cyclin D3. Blood. 2010;115:4071–82.
Crijns AP, de Graeff P, Geerts D, Ten Hoor KA, Hollema H, van der Sluis T, et al. MEIS and PBX homeobox proteins in ovarian cancer. Eur J Cancer. 2007;43:2495–505.
Mahmoudian RA, Bahadori B, Rad A, Abbaszadegan MR, Forghanifard MM. MEIS1 knockdown may promote differentiation of esophageal squamous carcinoma cell line KYSE-30. Mol Genet Genom Med. 2019;7:e00746.
Whitlock NC, Trostel SY, Wilkinson S, Terrigino NT, Hennigan ST, Lake R, et al. MEIS1 down-regulation by MYC mediates prostate cancer development through elevated HOXB13 expression and AR activity. Oncogene. 2020;39:5663–74.
Zhu J, Cui L, Xu A, Yin X, Li F, Gao J. MEIS1 inhibits clear cell renal cell carcinoma cells proliferation and in vitro invasion or migration. BMC Cancer. 2017;17:176.
Nomura K, Klejnot M, Kowalczyk D, Hock AK, Sibbet GJ, Vousden KH, et al. Structural analysis of MDM2 RING separates degradation from regulation of p53 transcription activity. Nat Struct Mol Biol. 2017;24:578–87.
Linke K, Mace PD, Smith CA, Vaux DL, Silke J, Day CL. Structure of the MDM2/MDMX RING domain heterodimer reveals dimerization is required for their ubiquitylation in trans. Cell Death Differ. 2008;15:841–8.
Plechanovová A, Jaffray EG, Tatham MH, Naismith JH, Hay RT. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature. 2012;489:115–20.
Wang W, Qin JJ, Rajaei M, Li X, Yu X, Hunt C, et al. Targeting MDM2 for novel molecular therapy: beyond oncology. Med Res Rev. 2020;40:856–80.
Rinaldo C, Prodosmo A, Siepi F, Moncada A, Sacchi A, Selivanova G, et al. HIPK2 regulation by MDM2 determines tumor cell response to the p53-reactivating drugs nutlin-3 and RITA. Cancer Res. 2009;69:6241–8.
Acknowledgements
We thank Professor Mian Wu for HCT116 p53–/– cell lines and plasmids of MDM2 and its mutants. We appreciate Professor Ceshi Chen for HA-Ub and all ubiquitin mutant plasmids. We acknowledge Professor Wancai Yang for the CRC cell line. We express our sincere gratitude to Professor Wen Liu for designing the peptides derived from MEIS1.
Funding
This project was supported by the National Natural Science Foundation of China (No. 82173376 and No. 82372711), the Key Project of Hunan Province (No. 2022WK2012), and the Student Innovation Project of Central South University (No. 1053320215042 and 1053320220319).
Author information
Authors and Affiliations
Contributions
JXL and GY conceived of and designed the research studies. JXL performed the main experiments and generated figures. YXD, QX, SMD, ALL, DW, JQW, and CL performed some experiments. HXY, MNW and GS provided reagents. All authors participated in the analysis and interpretation of data. JXL wrote the manuscript, and all authors edited the manuscript. GY supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The study was approved by the Protection of Human Subjects Committee of Xiangya Hospital, and written informed consent was obtained from all participants prior to sample collection (2021-KT27). All protocols pertaining to animal care and euthanasia have been duly approved by the Institutional Animal Care and Use Committee at Central South University (CSU-2022-0001-0234).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, J., Duan, Y., Xiao, Q. et al. Ubiquitination of MEIS1 by MDM2 serves as a switch for p53 stabilization and DNA damage response activation. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01714-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01714-9