Abstract
An interplay of growth factors and signaling pathways governs the development and maintenance of lymphatic vasculature, ensuring proper fluid homeostasis and immune function. Disruption of these regulatory mechanisms can lead to congenital lymphatic disorders and contribute to various pathological conditions. However, the mechanisms underlying the molecular regulation of these processes remain elusive. Here, we reveal a critical and previously unappreciated role for the signaling scaffold protein Shoc2 in lymphangiogenesis. We demonstrate that loss of Shoc2 results in near-complete loss of lymphatic vasculature in vivo and senescence of lymphatic endothelial cells in vitro. Mechanistically, Shoc2 is required for balancing signaling through the ERK1/2 pathway, and its loss results in increased mTORC1 signaling. This dysregulation impairs mitochondrial respiration and triggers an IRF/IFN-II response, ultimately leading to cellular senescence. Strikingly, expression of the Noonan Syndrome with Loose anagen Hair (NSLH)-causing Shoc2 variant S2G phenocopies the effects of Shoc2 loss. Together, these studies establish the critical role of Shoc2 in lymphangiogenesis and uncover a novel mechanistic link between Shoc2 signaling, mitochondrial function, innate immune response, and lymphatic development, with significant implications for Ras-pathway-related congenital disorders.
Similar content being viewed by others
Introduction
The lymphatic vasculature plays a multifaceted role that extends beyond the continuous collection of excess interstitial fluid and macromolecules and their transport to the bloodstream [1]. A critical component of the immune system, the lymphatic system serves as a conduit for immune cell trafficking and is essential for both adaptive and innate immune responses [2, 3]. The intricate function of the lymphatic system, including its role in immune surveillance and response, relies heavily on the precise regulation of lymphatic endothelial cells (LECs). LECs are typically quiescent, but this minimal activity is actively maintained by complex intracellular signaling. Disrupting this balance has profound consequences [3, 4]. Abnormal activation of pathways like PI3K/AKT and Ras/ERK can force LECs out of quiescence, leading to uncontrolled proliferation and lymphatic malformations (abnormal vessel growth, swelling, or more severe conditions) that impair organ function [5, 6].
The pivotal role of the Ras/ERK signaling pathway in LEC function is underscored by syndromes collectively known as RASopathies, which arise from mutations in genes encoding components of this pathway [7]. Over 20% of RASopathy patients develop lymphatic dysfunction, ranging from lymphedema (a chronic condition characterized by the accumulation of fluid in the interstitial spaces) to more severe conditions, such as chylothorax (fluid in the pleural cavity), hydropericardium (fluid around the heart), or chylous ascites (fluid in the abdomen) [8, 9]. Despite their clinical impact, the causes of these lymphatic abnormalities remain poorly understood, including their delayed onset and triggers.
Noonan Syndrome with loose anagen hair (NSLH) (OMIM #607721) is a RASopathy caused by hereditary variants in the Shoc2 gene [10, 11]. NSLH patients manifest many typical RASopathy presentations, including lymphatic abnormalities [11,12,13,14,15]. Due to NSLH’s rarity, literature describing lymphatic presentation is limited. However, studies show patients harboring the Shoc2 variant c. A4 > G, p. S2G present with severe lymphatic dysfunction, including fetal hydrops, pleural effusions, and generalized edema resulting in death [14, 16, 17].
Shoc2 is a key, evolutionarily conserved Ras/ERK1/2 pathway regulator. It tethers Ras to the catalytic subunit of protein phosphatase 1c (PP1c, also known as PP1CA) to dephosphorylate the inhibitory phospho-S259 of RAF-1 and accelerate ERK1/2 signaling [18]. Shoc2 also interacts with the proteins of the ubiquitin system (HUWE1, VCP, USP7, PSMC5, and FBXW7) [11, 19,20,21,22,23], scaffold proteins SCRIB and Raptor, and signaling enzymes such as the p110α subunit of PI3K[24]. Although the roles of these interactions remain unclear, Shoc2 is vital for embryonic development, morphogenesis, and adult tissue maintenance, as evidenced by vertebrate models [25, 26]. Both Shoc2 null mice and zebrafish CRISPR/Cas9 knockouts show systemic defects and early embryonic lethality. Notably, zebrafish mutants develop unusual edemic swellings, and endothelial-specific Shoc2 knockout in mice leads to hemorrhaging, subcutaneous edema, and fetal lung congestion, suggesting potential vascular deficits [25, 26].
Given the importance of Shoc2 in ERK1/2 signaling, we asked whether Shoc2 regulates lymphatic development and cell maintenance. Using an in vivo approach, we demonstrate that Shoc2 is vital for developmental lymphangiogenesis, as Shoc2-null zebrafish embryos exhibit a near-total lack of lymphatic vessels. This lymphatic vessel deficiency can be rescued by endothelial-specific expression of wild-type Shoc2, demonstrating an endothelial cell-autonomous requirement for Shoc2 in lymphatic development. Our in vitro studies indicate that Shoc2 loss in primary human LECs impairs mTOR pathway activation, thereby disrupting mitochondrial function, activating nucleic acid-sensing machinery, and the IFN/JAK1/STAT1-mediated innate immune response, ultimately leading to cell senescence. Re-expression of wild-type, but not NSLH mutant Shoc2, in Shoc2-deficient LECs restores mitochondrial function and reverses innate immune responses. Furthermore, inhibiting mTOR activity rescues lymphangiogenesis in Shoc2-depleted zebrafish larvae. Hence, this study uncovers the cell-type-specific and essential role of Shoc2 in orchestrating lymphangiogenesis and LEC homeostasis, providing novel insights into the molecular mechanisms that may underlie lymphatic dysfunction in NSLH.
Results
Developmental abnormalities in Shoc2 mutant fish
To understand the role of Shoc2 in lymphatic development, we used a stable zebrafish line (ZDB-ALT-131217-17590, or sa24200) carrying a germline Shoc2 mutation. This c.1546 G > A mutation, which we refer to as shoc2SA, inserts a 31-bp cryptic exon into the mRNA, leading to a premature stop codon (p. Val345*) after exon 5 (Fig. S1A–C). Quantitative PCR and immunoblotting showed reduced Shoc2 mRNA and complete absence of Shoc2 protein in shoc2SA/SA larvae (Fig. S1D, E), indicating nonsense-mediated mRNA decay. The shoc2SA/SA mutants were outcrossed to wild-type fish for at least three generations before further study.
While heterozygous shoc2+/SA carriers had no apparent abnormalities, homozygous shoc2SA/SA larvae displayed severe developmental defects. By 5 dpf, nearly 80% of shoc2SA/SA larvae exhibited widespread edema (heart cavity, trunk, yolk), underinflated swim bladders, and lethargy, mirroring our previously described shoc2Δ22 line (Figs. 1A, B, and S1F) [25]. A complementation test between shoc2Δ22 and shoc2SA/SA confirmed that the edema phenotype is specific to Shoc2 loss (shoc2Δ22/SA). shoc2SA/SA larvae began dying at 6 dpf, with none surviving past 11 dpf (Fig. S1G), demonstrating that the c.1546 G > A substitution severely impairs zebrafish embryonic development.
A Brightfield image of a lateral view of the entire zebrafish larvae of wild-type sibling (top panel) and shoc2SA/SA mutant (bottom panel). The red arrow is pointing to an edematous site. B Brightfield images of a dorsal view of the entire zebrafish larvae of wild-type sibling (top panel) and shoc2SA/SA mutant (bottom panel). Red arrows are pointing to edematous sites. C Schematic of areas of confocal imaging highlighting the trunk and craniofacial vessels. D Confocal image of trunk vessels of wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 5 dpf. Scale bar: 100 μm. TD, thoracic duct; ISLV, intersegmental lymphatic vessel; DLLV, dorsal longitudinal lymphatic vessel. E Confocal image of trunk vessels of a wild-type sibling Tg(mrc1a:eGFP)y251 showing a separated green channel from (D). Scale bar: 100 μm. F Confocal image of trunk vessels of a wild-type sibling Tg(kdrl:mCherry)y171 showing a separated red channel from(D). Scale bar: 100 μm. G Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 5 dpf. Asterisks indicate the absence of lymphatic vessels. Scale bar: 100 μm. H Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251 showing separated green channel from (D). Scale bar: 100 μm. I Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(kdrl:mCherry)y171 showing a separated red channel from (D). Scale bar: 100 μm. J Quantitation of the number of somites with TD segments in wild-type or shoc2SA/SA mutants at 5 dpf (n = 6 larvae in each group). K Quantitation of the number of somites with DLLV segments in wild-type or shoc2SA/SA mutants at 5 dpf (n = 6 larvae in each group). L Quantitation of the number of somites with ISLV in wild-type or shoc2SA/SA mutants at 5 dpf (n = 6 larvae in each group). M Confocal image of the craniofacial lymphatics of a wild-type sibling at 5 dpf. LFL lateral facial lymphatics, MFL medial facial lymphatics, OLV otolithic lymphatic vessel, LAA lymphatic branchial arches. Scale bar: 100 μm. N Confocal image of the craniofacial lymphatics of a shoc2SA/SA mutant at 5 dpf. Asterisks indicate the absence of lymphatic vessels. Scale bar: 100 μm.
Shoc2SA/SA mutants exhibit craniofacial abnormalities, including deformed, hypoplastic Meckel’s cartilage with disorganized chondrocyte stacking (Fig. S2A). Alizarin Red S staining showed significantly reduced craniofacial bone calcification (Fig. S2B). shoc2SA/SA mutants also displayed abnormal pigmentation, characterized by loss of iridophores and irregular melanocyte patterning (Fig. S2C–E). These data confirm that the shoc2 c.1546 G > A variant phenocopies the shoc2Δ22 mutant [25], making it a suitable model for studying Shoc2’s role in zebrafish development.
Defective lymphatic development in Shoc2 mutant fish
Zebrafish are widely used as a model for lymphatic studies due to their optical clarity, external development, conserved vasculature, and availability of lymphatic-specific transgenic reporters [27, 28]. To investigate the role of Shoc2 in lymphatic development, we crossed the shoc2SA/SA mutants with Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 double transgenic zebrafish. This line marks lymphatics, arteries, and veins with green, red, or both fluorescent proteins, respectively [28]. We focused on two major lymphatic vascular beds, the trunk and craniofacial lymphatic networks (Fig. 1C).
At 5 dpf, wild-type larvae exhibited normal lymphatic development, including the thoracic duct (TD), intersegmental lymphatic vessels (ISLV), and dorsal longitudinal lymphatic vessels (DLLV) (Fig. 1D–F). In contrast, shoc2SA/SA mutants showed a marked loss of lymphatic vessels, including the TD, ISLV, and DLLV (Fig. 1G, H), while blood vessel development remained largely unaffected (Fig. 1I). Quantification of TD, ISLV, and DLLV formation in wild-type siblings and shoc2SA/SA mutants revealed a nearly complete loss of all three sets of trunk lymphatic vessels in Shoc2-deficient animals (Fig. 1J–L). Craniofacial lymphatic vessels were also strongly reduced in the mutants (Fig. 1M, N). These findings indicate that lymphatic development is disrupted in Shoc2-deficient zebrafish.
Loss of lymphatic progenitors in Shoc2 mutant fish
In zebrafish, lymphatic development begins with secondary sprouts emerging from the posterior cardinal vein around 1.5 dpf [29, 30]. By 3 dpf, these sprouts form the parachordal line (PL, or parachordal angioblasts), a transient progenitor structure that gives rise to the trunk lymphatic vessels (Fig. 2A). Time-lapse imaging of Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 transgenic fish (32 h post-fertilization (hpf) to 50 hpf) revealed that wild-type and Shoc2 mutant siblings developed normal secondary sprouts migrating along the intersegmental blood vessels (Fig. 2B–D and Video S1). In contrast, shoc2SA/SA mutants produced only small abortive secondary sprouts (Fig. 2E–G and Video S2). By 3 dpf, wild-type siblings formed PLs (Fig. 2H, I), while shoc2SA/SA mutants completely failed to form PLs (Fig. 2J, K, asterisks), leading to a near-total loss of PL lymphatic progenitors in shoc2SA/SA mutants (Fig. 2L).
A A schematic showing the region of interest where lymphatic vessels originate. Secondary sprouts emerge from a posterior cardinal vein (PCV) and migrate dorsally around 1.5 dpf, giving rise to the parachordal lines (PL) at 3 dpf. These PLs are lymphatic progenitors that migrate and proliferate to form the trunk lymphatic vessels. B Confocal image of trunk vessels of a wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 32 hpf. The image is converted to a greyscale. Scale bar: 100 μm. C Confocal image of trunk vessels of a wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 41 hpf. The image is converted to a greyscale, and secondary sprouts are pseudocolored in green. Scale bar: 100 μm. D Confocal image of trunk vessels of a wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 50 hpf. The image is converted to a greyscale, and secondary sprouts are pseudocolored in green. Scale bar: 100 μm. E Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 32 hpf. The image is converted to a greyscale. Scale bar: 100 μm. F Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 41 hpf. The image is converted to a greyscale, and secondary sprouts are pseudocolored in green. Scale bar: 100 μm. G Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 50 hpf. The image is converted to a greyscale, and secondary sprouts are pseudocolored in green. Scale bar: 100 μm. H Confocal image of trunk vessels of a wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 3 dpf. PL, parachordal line. Scale bar: 100 μm. I Greyscale image of a wild-type sibling Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 in (H). PLs are pseudocolored in green. Scale bar: 100 μm. J Confocal image of trunk vessels of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 zebrafish larvae at 3 dpf. Asterisks indicate the absence of PLs. Scale bar: 100 μm. K Greyscale image of a shoc2SA/SA mutant Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 in (J). Asterisks indicate the absence of PLs. Scale bar: 100 μm. L Quantification of the number of somites containing PL segments in wild-type or shoc2SA/SA mutants at 3 dpf. Seven somites per larvae were counted (n = 7 larvae in each group). M Confocal image of trunk vessels of an uninjected Tg(mrc1a:kalTA4, UAS:eGFP) zebrafish larvae at 3 dpf. The arrows indicate PLs. Scale bar: 100 μm. N Confocal image of trunk vessels of a Tg(mrc1a:kalTA4, UAS:eGFP) shoc2 MO-injected zebrafish larvae at 3 dpf. The arrows indicate PLs. Scale bar: 100 μm. O Confocal image of trunk vessels of a Tg(mrc1a:kalTA4, UAS:eGFP) shoc2 MO and UAS:mCherry co-injected zebrafish larvae at 3 dpf. The arrows indicate PLs. Scale bar: 100 μm. P Confocal image of trunk vessels of a Tg(mrc1a:kalTA4, UAS:eGFP) shoc2 MO and UAS:mCherry-2A-shoc2 co-injected zebrafish larvae at 3 dpf. The arrows indicate PLs. Scale bar: 100 μm. Q Quantification of the number of somites containing PL segments in (M–P). Ten somites per larvae were counted. Data points indicate the number of larvae in each group.
In contrast, dorsal aorta-derived primary sprouts emerged normally to generate trunk intersegmental blood vessels, in both wild-type and shoc2SA/SA mutants (Fig. S3 and Video S3, 4), showing that Shoc2 loss-of-function specifically impairs secondary sprout formation and subsequent lymphatic development.
Endothelial expression of wild-type Shoc2 rescues lymphatic development in Shoc2 mutants
To examine whether endothelial Shoc2 is sufficient to support lymphatic development, we drove mosaic transgenic expression of wild-type Shoc2 in venous and lymphatic endothelial cells of Shoc2-deficient zebrafish. Tol2(uas:mcherry-2A-shoc2) or control Tol2(uas:mcherry) transgenes were injected into single-cell Tg(mrc1a:KalTA4; UAS:eGFP) transgenic animals together with a morpholino targeting the translation start site of endogenous but not transgene-driven Shoc2. The animals were raised and examined for PL formation at 3 dpf. As expected, morpholino injection phenocopied Shoc2 mutants, resulting in a dramatic loss of PL formation compared to uninjected siblings (Fig. 2M, N, Q). Co-injection of the control Tol2(uas:mcherry) transgene did not mitigate the effects of the morpholino on lymphatic development, whereas co-injection of the Shoc2-expressing Tol2(uas:mcherry-2A-shoc2) transgene partially rescued parachordal line formation in Shoc2 morphants (Fig. 2P, Q). These results confirm that Shoc2 function is required autonomously within endothelial cells for lymphatic development.
Shoc2 controls the amplitude of VEGF-mediated ERK1/2 signals
Recognizing the challenge posed by the near-complete loss of lymphatics in zebrafish Shoc2 null larvae for studying LEC homeostasis, we employed human dermal lymphatic endothelial cells (HDLEC) as a tractable in vitro system to investigate Shoc2 function. We found that Shoc2 regulates VEGF-initiated ERK1/2 pathway activation in HDLEC (via VEGF-C) and human umbilical vein endothelial cells (HUVEC) (via VEGF-A). Shoc2 shRNA depletion (KD) significantly decreased ERK1/2 phosphorylation, which was rescued by ectopic expression of Shoc2 tagged with tagRFP (SE), confirming its role in VEGF-mediated ERK1/2 signaling in endothelial cells (Fig. 3A–D). Interestingly, Shoc2-depleted HDLEC cultured for 6 days exhibited morphological changes. We found increased senescence-associated β-galactosidase (SA-β-gal) activity in nearly the entire population of Shoc2-depleted cells (Fig. 3E) and enlarged cell sizes (Fig. 3F). Shoc2-depleted HDLEC also exhibited elevated levels of TP53, phospho-TP53, and Senescence Associated Secretory Phenotype (SASP) markers p21Cip1 (CDKNA1), p16INK4a (CDKN2A), IL-6, and pH2A.X (Fig. 3G), indicative of a senescence phenotype. The minimal impact of Shoc2 loss on HUVEC (Fig. 3G) suggests a stringent requirement for Shoc2 in HDLEC proliferation and growth, potentially involving LEC-specific downstream effectors.
A Western blot of whole cell lysates extracted from human dermal lymphatic endothelial cells (HDLEC) expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), or Shoc2 shRNA together with Shoc2-tagRFP (SE), serum-starved for 16 h and then stimulated or not with VEGF-C (2 ng/ml) for 7 min at 37 °C. Cell lysates were probed using anti-pERK1/2, -ERK1/2, and -Shoc2 antibodies. Shoc2-tR indicates Shoc2-tagRFP with a higher molecular weight. B Levels of pERK1/2 in HDLEC treated with 2 ng/ml VEGFa at 7 min, normalized to the total amount of ERK1/2 and compared to pERK1/2 levels in HDLEC expressing non-targeting (NT) treated with 2 ng/ml VEGFa (set at 1.0). Values are shown in arbitrary units ± S.D. (n = 3) (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns, one-way ANOVA Dunnett’s multiple comparison test). C Western blot of whole cell lysates extracted from human umbilical vein endothelial cells (HUVEC) expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD) or Shoc2 shRNA together with Shoc2-tagRFP (SE), serum-starved for 16 h and then stimulated or not with VEGF-A (2 ng/ml) for 7 min at 37 °C. Cell lysates were probed using anti-pERK1/2, -ERK1/2, and -Shoc2 antibodies. D Levels of pERK1/2 in HUVEC treated with 2 ng/ml VEGFa at 7 min, normalized to the total amount of ERK1/2 and compared to pERK1/2 levels in HUVEC cells expressing non-targeting (NT), treated with 2 ng/ml VEGFa (set at 1.0). Values are shown in arbitrary units ± S.D. (n = 3) (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns, one-way ANOVA Dunnett’s multiple comparison test). E HDLEC expressing lentivirally-supplied non-targeting shRNA (NT) or Shoc2 shRNA (KD) were stained to examine β-Galactosidase activity (left panels). GFP, expressed independently by the lentivirus, served as a reporter to monitor delivery efficiency (right panels). Scale bar–150 μm. F Size distribution of HDLEC expressing lentivirally-supplied non-targeting shRNA (NT) or Shoc2 shRNA (KD), 6 days after transfection, as measured by surface area. G Western blot of whole cell lysates extracted from HDLECs and HUVECs expressing lentivirally supplied non-targeting shRNA (NT) or Shoc2 shRNA (KD) for 6 days. Cell lysates were probed using anti-Shoc2, -TP53, -pTP53, -CDKNA1, -CDKN2A, -IL-6, -pH2A.X and -GAPDH antibodies. The results in each panel are representative of those from three independent experiments.
Transcription changes in Shoc2-depleted HDLEC
To uncover mechanisms underlying Shoc2 depletion-induced senescence, we performed RNA sequencing of control HDLECs (non-targeting siRNA, NT), Shoc2 shRNA-depleted (KD), and HDLECs expressing Shoc2 shRNA together with siRNA-insensitive Shoc2-tagRFP (SE, rescue) (Fig. 4). Three days post-transfection, after confirming a reduction in Shoc2 protein (Fig. S4A), mRNA was isolated from each group for transcriptome profiling. DESeq2 analysis identified 316 differentially expressed genes (DEGs) with a fold change ≥0.5 across pairwise comparisons of Shoc2 KD and SE cells (Venn diagram; Fig. S4B), which were the focus of subsequent studies.
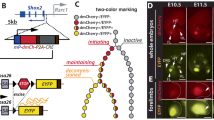
A, B Gene set enrichment analysis (GSEA) is presented as a score plot with the normalized enrichment score (NES) showing upregulation of interferon (IFN) response signature in Shoc2-depleted HDLEC (Shoc2 KD, KD) compared with control (non-targeting, NT) or HDLEC depleted of endogenous Shoc2 and expressing Shoc2-tagRFP (SE). The total height of the curve indicates the extent of enrichment (ES), and the normalized enrichment score (NES), false discovery rate (FDR), and p-value are also provided. Source data are provided as a source data file. Heatmap: the top elected correlated genes for each biological triplicate are presented in the corresponding heat map. Red and blue indicate increased and decreased expression, respectively, across all heatmaps. C The relative GAPDH-normalized expression of selected genes was measured by RT-qPCR. Data represent the mean and standard error of three independent experiments’ mean (SEM) values. P-values were calculated using a t-test (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns). Black bars represent control (NT), and red bars represent Shoc2-depleted cells (KD). D Cell lysates of Shoc2-depleted (KD), control (NT), or Shoc2-tagRFP expressing (SE) HDLEC were analyzed using Western blot 3 days and 6 days post-transfection to determine the expression levels of the indicated proteins. The results in each panel are representative of at least three independent experiments.
Gene Set Enrichment Analysis (GSEA) using the “Hallmark” database revealed negative enrichment in “Cholesterol Homeostasis,” “Fatty acid metabolism,” “Peroxisome,” and “MYC targets” gene sets (normalized enrichment scores (NES) of –2.28, –1.93, –1.87, and –1.80, respectively). Intriguingly, in Shoc2-depleted cells, we observed significant upregulation of the “Interferon-alpha response” (NES = 2.72) and “Interferon-gamma response” (NES = 3.10) genes (Figs. 4A, B, and S4C). “Biological Processes” analysis further showed enrichment in “Negative regulation of viral processes,” “Antigen processing and presentation,” and “Defense response to virus” (normalized p-value = 0.00) (Fig. S5A, B). Detailed analysis confirmed increased expression of key IFN response genes, including IFIT family members (IFIT1, IFIT3, IFI27, IFI6, IFIT30, IFIT35), ISGs (ISG15 and ISG20), and ISGylation regulators (USP18 and HERC6) (Figs. 4A, B, and S5C, S5D) [31, 32]. Many of these upregulated genes contain interferon-sensitive response elements (ISRE) and/or gamma-activated sequences (GAS) cis-regulatory sites (e.g., IFIT3, MX1, ISG15, BST2, IRF7, and OAS1) [33,34,35].
qRT-PCR analysis of mRNA isolated from Shoc2-depleted HDLEC at 3 and 6 days after transfection confirmed the RNA-seq data (Fig. 4C) and showed that loss of Shoc2 also affected the expression of lymphatic vascular markers PROX1 and LYVE1 (Fig. S6A). Gene expression changes became more pronounced at 6 days post-transfection, indicating a progressively stronger IFN response upon Shoc2 depletion, as validated by immunoblotting (Fig. 4D). Shoc2-tagRFP rescued the IFN response, confirming the specificity of the Shoc2-induced effect. Additionally, Shoc2-depleted HDLEC exhibited higher levels of the transcription factors IRF7 and IRF9, which regulate interferon (IFN) production, including IFN-γ [36]. Similarly, there was an increase in the expression of IFN response genes (USP18, IRF7, ISG15, and MDA5) in Shoc2-depleted zebrafish larvae (Fig. S6B). These results suggest that loss of Shoc2 in HDLECs induces transcriptional changes associated with the innate immune response.
Shoc2 loss in HDLEC activates the JAK1/STAT1 intracellular signaling pathway
Type I and II IFNs initiate ISG transcription primarily through the Janus-Activated Kinase (JAK)-Signal Transducer and Activator of Transcription (STAT) pathway (Fig. 5A) [36]. We found that Shoc2 loss in HDLECs significantly increased JAK1 (Y1034/1035) and STAT1 (Y701) phosphorylation, which was reversed by Shoc2-tagRFP (Fig. 5B), confirming that the IFN response in Shoc2-depleted HDLECs is associated with JAK1/STAT1 pathway activation.
A Schematic representation of the major signaling pathway activated by IFNs - the Janus kinase/signal transducer and activator of transcription (JAK1/STAT1) pathway. Activation of the JAK1/STAT1 pathway at the plasma membrane and rapid phosphorylation and nuclear translocation of STAT1 lead to the expression of IFN-stimulated genes (ISGs) containing ISREs (Interferon-Stimulated Response Elements) and GAS (Gamma Interferon-Activated Sites) DNA cis-elements. B Western blot of whole cell lysates extracted from human dermal lymphatic endothelial cells (HDLEC) expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), or Shoc2 shRNA together with Shoc2-tagRFP (SE) for 3 and 6 days. Cell lysates were probed using antibodies to the indicated proteins. C Western blot of whole cell lysates extracted from HDLEC expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), or Shoc2 shRNA together with Shoc2-tagRFP (SE) for 6 days. Cells were treated with 10 µM MEK inhibitor PD50042 for 24 h before Western blot. Cell lysates were probed using antibodies to the indicated proteins. D Western blot of whole cell lysates extracted from HDLEC expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), or Shoc2 shRNA together with Shoc2-tagRFP (SE) for 6 days. Cells were treated with 5 µM JAK1 inhibitor ruxolitinib for 24 h before Western blot. Cell lysates were probed using antibodies to the indicated proteins. The results in each panel are representative of at least three independent experiments.
Earlier studies suggested that ERK1/2 might regulate CCAAT/enhancer-binding protein-β (C/EBP-β), a transcription factor that controls the expression of various ISGs [37, 38]. Therefore, we tested whether blocking ERK1/2 activation would replicate the IFN response seen in Shoc2-depleted HDLECs. Treating Shoc2-depleted, Shoc2-tagRFP-expressing, and control HDLECs with the MEK1/2 inhibitor PD98059 reduced ERK1/2 phosphorylation but did not affect STAT1 phosphorylation or the expression of IFN response proteins (BST2, ISG15, IRF7, and IRF9) (Fig. 5C). Conversely, the JAK1 inhibitor ruxolitinib successfully suppressed the IFN response and JAK1/STAT1 pathway activation in Shoc2-depleted HDLECs (Fig. 5D). The reduction in ERK1/2 phosphorylation observed with ruxolitinib is probably a downstream effect of JAK1/STAT1 pathway inhibition [39]. These findings demonstrate that the IFN response and JAK-STAT pathway activation in Shoc2-depleted HDLECs are independent of ERK1/2 pathway activity.
We then investigated whether Shoc2 loss induces a similar IFN/JAK1/STAT1 response in human umbilical vein endothelial cells (HUVECs) and how the Shoc2 S2G variant impacts this response (Fig. S7). As anticipated, Shoc2 depletion in HDLECs increased levels of ISGs (ISG15, MX1, BST2, RSAD2), the transcription factors IRF7 and IRF9, and IFN-γ and STAT1 phosphorylation. Importantly, in HDLECs depleted of Shoc2 and expressing the Shoc2 S2G variant (S2G), the IFN response and the JAK1/STAT1 pathway activation were comparable to those of the Shoc2-depleted HDLEC (KD) (Fig. S7A). Conversely, the IFN/JAK1/STAT1 response remained at basal levels in HUVECs and primary fibroblasts, regardless of Shoc2 expression or mutation status (Fig. S7B, C). Moreover, 6 days of IFN-γ treatment in HDLEC induced cell senescence (Fig. S8A, B). These data indicate that the aberrant IFN/JAK1/STAT1 pathway activation observed after Shoc2 depletion or S2G variant expression is unique to HDLECs.
Activation of the RIG-I-like receptor signaling pathway in Shoc2-depleted HDLEC
Transcriptome analysis of Shoc2-depleted HDLECs revealed significant enrichment for genes involved in the RIG-I-like receptor (RLR) signaling pathway (KEGG pathway; NES = 1.49). Key RLR pathway genes, including IFIH1 (MDA5), DDX60, DDX58 (RIG-I), TRIM25, and TRIM14, were significantly upregulated in Shoc2-depleted cells (Fig. 6A). MDA5 and RIG-I are cytoplasmic pattern recognition receptors that detect viral double-stranded RNA (dsRNA) and initiate an innate immune response. This response also involves MAVS (mitochondrial antiviral signaling protein) recruitment, which activates transcription factors such as IRF7 and IRF9, driving the production of IFNs and proinflammatory cytokines (Fig. 6B) [40]. qRT-PCR confirmed the increased expression of MDA5 (IFIH1), RIG-I (DDX58), and other RLR response components (E3 ligases TRIM25 and TRIM14, and helicase DDX60) in Shoc2-depleted HDLECs. Notably, MDA5 and RIG-I expression was significantly elevated as early as 3 days post-transfection (Fig. 6C).
A Gene set enrichment analysis (GSEA) is presented as a score plot, with NES indicating the upregulation of selected genes in a RIG-1-like signaling signature in Shoc2-depleted HDLEC (Shoc2 KD, KD) compared with the control (non-targeting, NT). The total height of the curve indicates the extent of enrichment (ES), and the normalized enrichment score (NES), false discovery rate (FDR), and p-value are also provided. Source data are provided as a source data file. Heatmap: the top elected correlated genes for each biological triplicate are presented in the corresponding heat map. Red and blue indicate increased and decreased expression, respectively, for all heatmaps. B Schematic representation of the RIG-I/MDA5 nucleic acid sensing pathway activated by dsRNA. Activation of MDA5 and its interaction with MAVS leads to the phosphorylation of the transcription factors IRF7 and IRF9, promoting their nuclear translocation and driving the expression of IFN genes. C The relative GAPDH-normalized expression of selected genes was measured by RT-qPCR. Data represent the mean and standard error of three independent experiments’ mean (SEM) values. P-values were calculated using a t-test (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns). Black bars represent control (NT), and red bars represent Shoc2-depleted cells (KD). D MDA5 was precipitated from HDLECs depleted of Shoc2 (KD), control (NT), expressing Shoc2-tagRFP (SE), or the Shoc2 S2G variants using anti-MDA5 antibodies. Immunoblots were analyzed with anti-pMDA5 (Ser88), anti-MDA5, and anti-GAPDH antibodies. E MDA5 was precipitated from HUVECs depleted of Shoc2 (KD), control (NT), expressing Shoc2-tagRFP (SE), or the Shoc2 S2G variants using anti-MDA5 antibodies. Immunoblots were analyzed using anti-pMDA5 (Ser88), MDA5, and GAPDH antibodies. F Western blot of whole-cell lysates extracted from HDLECs expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), and transfected with the indicated siRNAs for 3 days. Cell lysates were probed using antibodies to the indicated proteins. G Western blot of whole-cell lysates extracted from HDLECs expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), and transfected with MAVS siRNA for 3 days. Cell lysates were probed using antibodies to the indicated proteins. The results in each panel are representative of at least three independent experiments. H HDLEC expressing lentivirally-supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), or MAVS siRNA were stained to examine β-Galactosidase activity (left panels). Scale bar–150 μm.
MDA5 activation, crucial for the innate immune response, requires dephosphorylation of Ser88 and ISGylation [39,40,41]. We observed that in Shoc2-depleted or S2G-expressing HDLECs, increased MDA5 expression (Fig. 6D, input) correlated with significantly decreased Ser88 phosphorylation of MDA5 precipitated from either Shoc2-depleted or S2G-expressing HDLECs (Fig. 6D, IP), indicating MDA5 activation. Such a response was not observed in HUVECs (Fig. 6E).
We then confirmed the essential roles of MAVS and MDA5 in driving the IFN response in Shoc2-depleted HDLECs. Depleting either MDA5 or MAVS eliminated the IFN response (i.e., ISGs expression), the activation of the JAK1/STAT1 pathway (i.e., JAK1/ STAT1 phosphorylation) (Fig. 6F), and the cell senescence caused by Shoc2 loss (i.e., SASP response) (Fig. 6G, H). These results demonstrate that the activation of the IFN-JAK1/STAT1 pathway in Shoc2-depleted HDLECs depends on the MDA5-MAVS cytoplasmic pattern recognition response.
Mitochondrial dysfunction of Shoc2-depleted HDLEC
The mitochondrial outer-membrane protein MAVS influences mitochondrial homeostasis and dynamics, including fusion and fission [42, 43]. Therefore, we investigated how Shoc2 loss impacts mitochondrial function. Compared to controls, Shoc2-depleted HDLECs and those expressing the Shoc2 S2G variant showed significantly higher phosphorylation of Drp1 (Ser616), a mitochondrial fission marker (Fig. 7A). While total DRP1 protein levels remained stable, levels of other mitochondrial fusion proteins (MTFP1, MFF, and OPA1) were altered. Transcriptome analysis also revealed changes in oxidative phosphorylation (OXPHOS) gene expression, including those encoding ATP synthase, cytochrome c oxidase, and other electron transport chain components (Fig. S9).
A Western blot of whole-cell lysates extracted from HDLECs expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), Shoc2 shRNA together with Shoc2-tagRFP (SE), or Shoc2 shRNA together with Shoc2(S2G)-tagRFP (S2G) for 3 and 6 days. Cell lysates were probed using antibodies relevant to mitochondrial fission and fusion. GAPDH was used as a loading control. B Relative oxygen consumption rate (OCR), measured by the Seahorse XFe96 analyzer, is normalized to cell number. Data are presented as mean ± SEM of three independent experiments. Statistical significance was determined by one-way ANOVA (Brown-Forysthe test), (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns). FCCP, carbonyl cyanide-p trifluoromethoxyphenylhydrazone; Rot rotenone, AA antimycin A. C–F Basal and maximal OCR analysis, Mitochondrial ATP production, and proton leak of HDLEC depleted of Shoc2 (KD) or expressing either Shoc2-tagRFP (SE) or Shoc2-tagRFP S2G (S2G). Statistical significance was determined by one-way ANOVA (Brown-Forysthe test), (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns). G Immunostaining of dsRNA (J2) (green) in control and Shoc2-depleted HDLECs. Mitochondria are labeled with Mitotracker (red). Scale bar–10 μm. H RT–qPCR analysis of mtRNA heavy (H) and light (L) strands in cytosolic fractions of HDLEC depleted of Shoc2 for 6 days. The gene names indicate the location of the RNA segment amplified. Data shown as fold change with respect to cells expressing non-targeting siRNA. Each reaction was performed in triplicate. I Western blotting of membrane/nuclear (mem/N), mitochondrial (MT), and cytosolic (cyto) fractions of control (NT) and Shoc2-depleted (KD) cells, analyzed using Tom20 and GAPDH antibodies.
Furthermore, Seahorse metabolic flux analysis showed that Shoc2 loss significantly decreased basal mitochondrial oxygen consumption rates (OCR), but not maximal rates (Fig. 7B–D). Mitochondrial ATP production and proton leak were also considerably reduced in cells depleted of Shoc2 or expressing the Shoc2 S2G variant (Fig. 7E, F). Together, these data indicate that mitochondrial function is impaired in HDLEC depleted of Shoc2 or expressing the S2G variant.
The release of mt-dsRNA into the cytosol is a common feature of senescent cells and a key driver of the SASP phenotype [44]. HDLECs transfected with poly (I: C) (a synthetic analog of dsRNA) enter senescence and activate the immune response (Fig. S8C). Immunofluorescence analysis assessing the abundance and subcellular localization of dsRNA revealed higher levels of cytosolic dsRNA in Shoc2-depleted HDLECs than in control cells (Figs. 7G and S9C). Furthermore, both heavy and light mt-dsRNA transcripts were significantly increased in RNA extracted from the mitochondria and nucleus-free cytosol of Shoc2-depleted HDLEC (Fig. 7H). The purity of the cytosolic fraction was confirmed by probing the mitochondria marker Tom20, ensuring that the sample was free of mitochondrial or nuclear contamination (Fig. 7I). Together, these data indicate that cytosolic accumulation of mt-dsRNA is a likely trigger of cellular senescence in Shoc2-depleted HDLECs.
Regulation of the mTOR pathway by Shoc2
Given the ERK1/2-independent IFN-JAK1/STAT1 activation (Fig. 5) and impaired mitochondrial function in Shoc2-depleted HDLECs (Fig. 7), we explored other signaling pathways. Shoc2 was previously shown to negatively regulate mTORC1 signaling by competing with Raptor, a key component of the mTORC1 complex, for binding to mTOR (Fig. 8A) [45]. Therefore, the effect of Shoc2 loss on mTOR pathway activation was examined (Fig. 8B). Increased mTORC1 activity, evidenced by higher phosphorylation of mTOR and its downstream targets (i.e., eukaryotic translation initiation factor 4E-binding protein 1 (4E-BP1) and ribosomal protein S6 kinase beta-1 (S6K1), was observed in Shoc2-depleted or S2G-expressing HDLECs. Immunoprecipitation confirmed significantly increased Raptor-mTOR binding in Shoc2-depleted or S2G-expressing HDLECs, reversed by Shoc2-tagRFP (Fig. 8C). mTOR-Rictor binding remained unaffected by the Shoc2 loss (Fig. 8C). We did not observe mTOR pathway hyperactivation in Shoc2-depleted HUVECs (Fig. S10A, B). Although Raptor-mTOR binding was slightly increased in Shoc2-depleted or S2G-expressing HUVECs, the lack of mTOR signaling upregulation might be due to the substantially lower densitometric Raptor-mTOR ratio (1:0.2 vs. 1:1 in HDLEC) (Fig. S10B).
A Schematic representation of the Shoc2-mTOR axis in which Raptor binds to mTOR, stimulates mTORC1 complex activity, indicated by S6K and 4EBP phosphorylation. mTORC1 activation is followed by activation of the IFN/Jak/STAT response. B Western blot of whole cell lysates extracted from HDLEC expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), Shoc2 shRNA together with Shoc2-tagRFP (SE), or Shoc2 shRNA together with Shoc2(S2G)-tagRFP (S2G) for 3 days. Cell lysates were probed using antibodies to proteins relevant to mTORC1 signaling. GAPDH was used as a loading control. C Cell lysates were prepared from HDLEC cells expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), Shoc2 shRNA together with Shoc2-tagRFP (SE), or Shoc2 shRNA together with Shoc2(S2G)-tagRFP (S2G) for 3 days. mTOR was immunoprecipitated with an anti-mTOR antibody. Immunoprecipitated proteins were subjected to SDS-PAGE and western blot analysis. Blots were probed with antibodies against mTOR, Raptor, and Rictor. D, E Western blot of whole cell lysates extracted from HDLEC expressing lentivirally supplied non-targeting shRNA (NT), Shoc2 shRNA (KD), Shoc2 shRNA together with Shoc2-tagRFP (SE), or Shoc2 shRNA together with Shoc2(S2G)-tagRFP (S2G) for 3 days and treated with Rapamycin. Cell lysates were probed using the indicated antibodies. GAPDH was used as a loading control. F HDLEC expressing lentivirally-supplied Shoc2 shRNA (KD) were treated with 10 nM Rapamycin 24 h after transfection for 6 days and then stained to examine β-Galactosidase activity (left panels). GFP, expressed independently by the lentivirus, served as a reporter to monitor delivery efficiency (right panels). Scale bar–150μm. G As shown by the schematic illustration, one-cell stage Tg(mrc1a:eGFP)y251; Tg(kdrl:mCherry)y171 embryos were injected with shoc2 MO, and then treated with 25 μM of Rapamycin for 40 h starting at 28 hpf. Confocal images show the trunk vessels of 3 dpf zebrafish larvae injected with shoc2 MO (upper image) and shoc2 MO-injected larvae treated with 25 μM of Rapamycin (lower image). The asterisks or arrows indicate the absence or presence of PLs, respectively. Scale bar: 100 μm. Quantification of the number of somites containing PL segments (Fig. S10). Ten somites per larva were counted. Data points indicate the number of larvae in each group.
To confirm the mechanistic link between mTORC1 activation and the IFN response in Shoc2-depleted HDLEC, cells were treated with an mTOR inhibitor, rapamycin. Rapamycin effectively blunted mTOR signaling, blocked the IFN/JAK/STAT response in Shoc2-depleted and S2G-expressing HDLECs (Fig. 8D), normalized mitochondrial fusion protein activation and expression (Fig. 8E), and prevented the cell senescence in Shoc2-depleted HDLECs (Fig. 8F). Additionally, rapamycin treatment partially restored lymphangiogenesis in zebrafish larvae depleted of Shoc2 (Figs. 8G and S10C–G).
Our data shows that mTORC1, but not mTORC2, mediates Shoc2’s effects in HDLECs. Disrupting the Shoc2-Raptor-mTOR equilibrium by Shoc2 loss or the S2G mutant increases Raptor-mTOR binding and mTORC1 signaling, which affects mitochondrial function and leads to the accumulation of mt-dsRNA in the cytosol, triggering an innate interferon response and ultimately causing cell senescence (Fig. S11).
Discussion
Lymphatic vessel development and homeostasis are governed by intricate signaling [1, 5, 46]. Our study uncovers a critical new role for the signaling scaffold Shoc2 in lymphatic regulation, revealing its multifaceted impact on endothelial signaling and homeostasis. We demonstrate LEC-specific Shoc2 functions: Shoc2 loss activates mTOR and IFN/JAK/STAT signaling pathways, disrupts mitochondrial respiration and function, and induces cellular senescence, ultimately leading to lymphangiogenic defects. Notably, the Shoc2 (S2G) NSLH variant mirrors these dysfunctions, providing relevant in vitro evidence for understanding the molecular mechanisms underlying NSLH-related lymphatic dysfunction, particularly in the absence of accessible patient-derived cell models. Together, these findings establish a novel mechanistic link between the Shoc2 scaffold, mitochondrial function, mTOR signaling, and innate immune response in LECs. Importantly, these effects are cell-type specific, as Shoc2 depletion in primary fibroblasts and venous endothelial cells failed to induce comparable mTOR activation or immune responses.
Because Shoc2 knockout mice are embryonically lethal at E8.5 (26), zebrafish are currently the only in vivo vertebrate model for examining global Shoc2 loss. Our previous work showed severe neural crest defects in shoc2 mutant zebrafish [25, 47]. This study uncovers a critical, lymphatic endothelium-autonomous role of Shoc2 (Figs. 1 and 2). Shoc2 mutant zebrafish display a striking loss of facial and trunk lymphatic vessels, with no apparent effect on other endothelial cells. Given the high homology between zebrafish and human Shoc2, these findings are highly relevant to human biology. Our endothelial-specific transgenic Shoc2 “rescue” experiments confirm its autonomous requirement and sufficiency for normal lymphatic development. It is tempting to hypothesize that the tissue microenvironment (e.g., stiffness, immune cell composition) or a specific need for mTOR metabolic signaling in different endothelial cells makes lymphatic cells particularly sensitive to Shoc2 function. Future research will clarify why the loss of this ubiquitously expressed signaling scaffold causes such diverse outcomes across cell types.
Shoc2 function in lymphatic endothelial cells: mechanisms
Beyond its role in developmental lymphangiogenesis, this study elucidates how Shoc2 maintains LEC homeostasis. Shoc2’s function is best explored in the ERK1/2 pathway [48]. in its impact on the proliferation and motility of immortalized/cancer cells [49, 50], and in its role in mTORC1 signaling in various cancers [20, 51]. Yet the function of Shoc2 in primary cells remains largely unexplored. We provide compelling evidence that, in primary LECs, Shoc2 regulates VEGF-ERK1/2 activity and balances mTORC1 signals (Figs. 3, 5, and 8). While pharmacological modulation with rapamycin provided preliminary insight into a potential relationship between Shoc2-associated signaling and mTORC1 activity, its systemic effects, particularly in vivo, preclude definitive mechanistic conclusions, positioning this observation as hypothesis-generating and warranting further investigation with more targeted approaches. Consistent with mTORC1’s role in regulating mitochondrial function and cellular senescence [52,53,54,55,56], upregulated mTORC1 in Shoc2-depleted cells leads to mitochondrial dysfunction. Given mTORC1’s pleiotropic roles in cellular processes such as nutrient and energy sensing, protein synthesis, and metabolism, further work is critical to fully understand Shoc2’s effects on cellular homeostasis, especially its role in adaptive autophagy and nutrient sensing.
Mitochondria are key immune signaling hubs [43, 57]. This study reveals that Shoc2 dysfunction impairs mitochondrial-regulated innate immune responses (Fig. 7), leading to mt-dsRNA release, activation of nucleic acid sensing, and the production of proinflammatory cytokines and interferons, ultimately resulting in cellular stress and senescence (Fig. S11). Importantly, Shoc2’s role in MDA5 activation is independent of the Shoc2-PP1c-MRas holoenzyme, as PP1c was absent from mTOR-Shoc2 immunoprecipitates and MDA5 Ser338 was dephosphorylated in Shoc2-depleted cells. Future work will investigate whether the lymphatic endothelium’s sensitivity to Shoc2 dysfunction is linked to mTORC1 signaling-mediated regulation of cell quiescence.
The inhibitory effects of IFNγ on lymphatic vessels are well-documented [1, 58,59,60]. Thus, increased IFNγ signaling in Shoc2-depleted LECs likely contributes to their senescence and underlies the lymphatic vessel defects in Shoc2-lacking larvae. Future studies employing selective inhibition of the interferon pathway will be essential for defining the molecular circuitry by which endogenous interferon signaling regulates LEC proliferation and senescence, and for delineating the role of Shoc2 within this pathway. Investigating whether lymphatic cells in NSLH patients modulate adaptive immunity or antigen presentation, and the physiological consequences of the Shoc2 S2G-associated IFN response on lymphatic vessel development, including the potential contribution of chronic IFN activation to lymphatic dysfunction, is crucial. These studies align with recent findings on Shoc2’s role in invertebrate antibacterial and eukaryotic antiviral immune responses [61, 62].
Implications for NSLH pathology
One of the most interesting findings is that the Shoc2 NSLH (S2G) variant fails to effectively compete with mTOR for Raptor binding in LEC. Similarly to Shoc2 loss, this Shoc2 variant disrupts mitochondrial function and triggers the IFN response. Given the Shoc2 S2G variant’s aberrant N-myristoylation and PM localizaLon [10b], this suggests the formaLon of spaLally defined Shoc2-Raptor-mTOR complexes. However, why this variant increases mTOR activity, despite binding to mTOR, remains to be elucidated. Notably, mitochondrial dysfunction, decreased adipogenesis, and metabolic changes are found in other RASopathies [63,64,65]. For example, PTPN11 mutations (associated with Noonan syndrome) reduce mitochondrial membrane potential and ATP levels, thereby increasing ROS levels, whereas B-RAF and NF1 mutations affect respiration [66,67,68]. While specific mechanisms remain unclear, our results underscore the need to elucidate the mechanistic link between mitochondrial dysfunction and specific RASopathy gene mutations.
RASopathy patients, including those with the NSLH Shoc2 S2G variant, face an increased risk of autoimmune disorders [69,70,71,72,73,74,75], though the triggers for clinically overt disease remain poorly understood. Our study presents a novel mechanism: Shoc2-deficient lymphatic cells exhibit mitochondrial dysfunction and an impaired IFN response, which may underlie NSLH autoimmune presentations.
In conclusion, we demonstrate that the progressive IFN response observed in both Shoc2-deficient and S2G variant-expressing cells is mTOR-dependent and reversible with rapamycin. Our findings underscore Shoc2’s critical role as a molecular gatekeeper that balances ERK1/2 and mTOR signaling and highlight its cell-type-specific functions, providing novel insights into the roles of ERK1/2 and mTOR during developmental lymphangiogenesis. These results highlight the importance of a more comprehensive understanding of Shoc2’s complex role in both healthy and diseased organisms. Given the intricate interactions among Shoc2, the IFN response, and mTORC1, targeting these pathways offers significant therapeutic potential for other RASopathies associated with lymphatic dysfunction or abnormal immune activation. Specifically, JAK or mTOR inhibitors may provide therapeutic benefits for patients with Shoc2 mutations, possibly reducing lymphatic and immune-related pathologies.
Materials and methods
Antibodies and other reagents
Table S1 lists the specific primary and secondary antibodies used for protein detection. Ruxolitinib and PD98059 were purchased from LC Labs, and hexadimethrine bromide (polybrene) was obtained from Sigma-Aldrich.
Constructs
tagRFP-tagged Shoc2 (Shoc2-tRFP), the pLVTHM constructs expressing Shoc2 short hairpin RNA (shRNA), and non-targeting shRNA were described previously [76]. The pLVTHM constructs expressing Shoc2 and non-targeting shRNA also express GFP to visualize transfected cells [76].
Cell culture and lentivirus transduction
Human dermal lymphatic endothelial cells (HDLEC) (PromoCell) and normal human umbilical vein endothelial cells (HUVEC) (Lifeline Cell Technology) were grown in VascuLife basal media (Lifeline Cell Technology) containing 5 ng/mL rh FGF basic, 50 µg/mL ascorbic acid, 1 µg/mL hydrocortisone hemisuccinate, 2% FBS, 10 mM L-glutamine, 15 ng/mL rh IGF-1, 5 ng/mL rh EGF, 5 ng/mL rh VEGF, 0.75 U/mL heparin sulfate, and supplemented with 30 µg/mL gentamicin and 15 ng/mL amphotericin B.
For lentivirus transduction, cells were plated in 12-well dishes at 50–60% confluence. The ratio of viral particles to target cells was determined for each viral prep. The lentiviral stock was diluted in a fresh culture medium containing polybrene. Shoc2–tagRFP (SE or S2G) and shShoc2/GFP (KD), or non-targeting-GFP (NT) fluorescence, were detected 36–48 h after lentivirus transduction. The efficiency of shRNA knockdown and cDNA expression was validated using Shoc2-specific antibodies 3 and 6 days after lentivirus transduction. To ensure well-controlled experiments and avoid bias from clonal variation, individual cell clones were not selected for analysis.
Separation of cytoplasmic and mitochondrial fractions
Cytoplasmic and mitochondrial fractions were prepared using the Mitochondria Isolation Kit (89874, Thermo Fisher Scientific). The purity of the fractions was assessed by Western blotting according to standard protocols. RNA was extracted using the RNeasy Plus isolation kit with gDNA eliminator columns (Qiagen).
Real-time quantitative PCR (qPCR): RNA extraction, cDNA synthesis, and PCR
Total RNA was isolated from cells using the PureZOL/Aurum total RNA isolation kit (Bio-Rad) according to the manufacturer’s instructions. Aliquots containing equal amounts of RNA were used for reverse transcription-PCR (RT-PCR) analysis. Quantitative RT-PCR was performed using the SoAdvanced SYBR Green Supermix and a Bio-Rad CFX detection system (Bio-Rad). Sequence-specific primer sets are listed in Table S2. The relative RNA levels were calculated using the comparative threshold cycle method. Sample values were normalized to GAPDH mRNA, and results are shown as fold changes in mRNA levels relative to cells transfected with non-targeting siRNA. The data represent the mean ± standard deviation (S.D.) from three independent experiments. For RNA isolated from the cytosolic fractions of HDLEC, RevertAid reverse transcriptase (ThermoFisher Scientific) was used to synthesize cDNA. RT–qPCR was performed using strand-specific primers containing CMV promoter sequences to target different genes. Reactions were performed in triplicate, and all qRT-PCR data were normalized to the housekeeping genes B-actin and GAPDH. The primers used are listed in Tables S2, S3, S5, and S6.
siRNA transfections
To silence protein expression by RNA interference, cells were seeded in 12-well plates (at 50–60% confluence) with 1 ml of VascuLife media per well at least 20 h before transfection. Small interfering RNA (siRNA) transfections were performed according to the manufacturer’s recommendations using Dharmafect 2 (Dharmacon). The siRNA sequences targeting MDA5 and MAVS are listed in Table S4. The siRNA knockdown efficiency was validated by Western blotting.
Immunoprecipitation
Cells were placed on ice and washed with Ca2+, Mg2+-free phosphate buffered saline (PBS), and the proteins were solubilized in 50 mM Tris (pH 7.5) containing 150 mM NaCl, 1% Triton X-100, 1 mM Na3VO4, 10 mM NaF, 0.5 mM phenylmethylsulfonyl fluoride (PMSF, Sigma, St. Louis, MO, USA), 10 μg/ml of leupeptin, and 10 μg/ml of aprotinin (Roche, Basel, Switzerland) for 15 min at 4 °C. Lysates were centrifuged at 14,000 rpm for 15 min to remove insoluble material. Lysates were incubated with appropriate antibodies for 2 h, and the immunocomplexes were precipitated using protein A- or G-Sepharose (GE Healthcare Life Sciences, Chicago, IL, USA). Immunoprecipitates and aliquots of cell lysates were denatured in a sample buffer at 95 °C, resolved by electrophoresis by SDS-PAGE, transferred to a nitrocellulose membrane, and probed by western blotting with various antibodies, followed by chemiluminescence detection.
mTOR immunoprecipitation was done following the protocol of Kim et al. [77]. Briefly, cells were rinsed once with PBS, lysed in 600 µl of ice-cold lysis buffer (40 mM HEPES [pH 7.5], 120 mM NaCl, 1 mM EDTA, 50 mM NaF, 1.5 mM Na3VO4, 0.5 mM phenylmethylsulfonyl fluoride, 10 μg/ml of leupeptin, and 10 μg/ml of aprotinin and 0.3% CHAPS) and incubated for 30 min on ice. After clearing, 40 µl of a 50% slurry of protein G-Sepharose and 4 µg of the anti-mTOR antibody were added to the supernatant, which was then rotated for 3 h at 4 °C.
Western blot analysis
Cells were placed on ice and washed with Ca2+, Mg2+-free phosphate buffered saline (PBS), and the proteins were solubilized in 50 mM Tris (pH 7.5) containing 150 mM NaCl, 1% Triton X-100, 1 mM Na3VO4, 10 mM NaF, 0.5 mM phenylmethylsulfonyl fluoride (PMSF, Sigma, St. Louis, MO, USA), 10 μg/ml of leupeptin, and 10 μg/ml of aprotinin (Roche, Basel, Switzerland) for 15 min at 4 °C. Lysates were centrifuged at 14,000 rpm for 15 min to remove insoluble material. Aliquots of cell lysates were denatured in a loading buffer at 95 °C, resolved by electrophoresis, and probed by western blotting with various antibodies, followed by chemiluminescence detection. Proteins transferred from SDS-polyacrylamide gels to nitrocellulose membranes were visualized using a ChemiDoc analysis system (Bio-Rad, Hercules, CA, USA). Several exposures were analyzed to determine the linear range of the chemiluminescence signals. Quantification was performed using the densitometry analysis mode of Image Lab software (Bio-Rad, Hercules, CA, USA). To visualize proteins with similar molecular weights, in some instances, several identical SDS-PAGE gels were used to resolve protein lysates.
Immunofluorescence of dsRNA with J2 antibody
Cells were grown on a coverslip for 24–48 h before treatment. Cells were incubated with MitoTracker™ Red CMXRos (7512, Thermo Fisher) at 200 nM for 30 min at 37 °C before fixing in 4% PFA for 15 min at room temperature. Cells were washed three times with PBS, permeabilized with 0.25% Triton X-100 in PBS, washed with 0.05% Tween 20 in PBS, and incubated with 3% BSA in TBS for 60 min at room temperature. Anti-dsRNA (J2) antibodies were diluted 1:200 in TBST containing 1% BSA and incubated for 60 min at room temperature. The coverslips were washed three times with 0.05% Tween 20-PBS, incubated with secondary goat anti-mouse IgG (H + L) conjugated to Alexa Fluor 488 at a dilution of 1:300, and then washed three times with 0.05% Tween 20-PBS and PBS. Finally, they were mounted with Mowiol (Calbiochem) medium. Images were acquired with a spinning-disk Marianas system based on a Zeiss Axio Observer Z1 inverted fluorescence microscope equipped with 100x Plan Apo PH NA 1.6 oil immersion objective, piezo stage controller, spherical aberration correction module, temperature- and CO2-controlled chamber, all controlled by Slidebook6 software (Intelligent Imaging Innovation, Denver, CO). For immunofluorescence imaging, a Hamamatsu ORCA-Fusion BT CMOS camera was used to obtain a z-stack of 10 x-y confocal images acquired at 250–300 nm z-steps. Images were analyzed using Slidebook software.
Seahorse extracellular flux analysis
The Seahorse XF96 Extracellular Flux Analyzer (Agilent) was used to measure cellular respiration activity in HDLEC. Cells were seeded at a density of 2 × 104 cells per well in an XF96 plate ~12 h before the measurement. The following day, cells were washed twice with assay medium (unbuffered DMEM supplemented with 25 mM glucose, 2 mM L-glutamine, and 1 mM pyruvate) and incubated at 37 °C in a non-CO2 incubator for 1 h. OCR was measured at baseline and following sequential injections of oligomycin (1 µM), carbonyl cyanide-4-(trifluoromethoxy)phenylhydrazone (FCCP) (1.5 µM), and rotenone/antimycin A (0.5 µM). Data were normalized to total protein content determined by a BCA assay.
The mitochondrial stress tests were performed per the manufacturer’s protocol. The relative levels of non-mitochondrial, basal, maximal, and ATP production-related respiration were calculated using the Mito stress tests using the Seahorse Wave software (Agilent) for XF analyzers.
Cell senescence assay
HDLEC were stained for senescence-associated β-galactosidase activity using the β-Galactosidase Staining Kit (Cell Signaling) according to the manufacturer’s recommendation. Briefly, equal numbers of cells 2 × 104 were washed once with 1X PBS and then fixed. Cells were incubated with the β-Galactosidase Staining Solution at 37 °C for at least overnight. Cells were overlaid with 70% glycerol for long-term storage and stored at 4 °C. To quantify cells stained for senescence-associated β-galactosidase activity, we acquired GFP fluorescence to visualize the transfected cells. The percentage of senescent cells was divided by the total number of cells counted using immunofluorescence (n = 3 dishes per genotype).
Zebrafish strains and maintenance
This study was conducted in AAALAC-accredited facilities under active research projects approved by the University of Kentucky Animal Care and Use Committee (#2019-3208), the University of Illinois at Chicago Animal Care and Use Committee (#23-112), and the Eunice Kennedy Shriver National Institute of Child Health and Human Development (Animal Study Proposal #21-015, 24-015). All zebrafish strains were bred, raised, and maintained according to established animal care protocols for zebrafish husbandry. Embryos were staged as previously described [78]. The following transgenic and mutant zebrafish lines were used in this study: Tg(mrc1a:eGFP)y251 (28), Tg(kdrl:mcherry)y205 [79]. and Tg(fli1:eGFP)y1 [80]. The Tg(mrc1a: KalTA4) line was generated from this study. The Tg(UAS:eGFP) line was generously provided by the Kawakami lab. The shoc2Δ22 has been previously described in [25]. and the ZDB-ALT-131217-17590; sa24200 (shoc2SA/SA) line was obtained from the Sanger Institute Zebrafish Mutation Project.
Generation of transgenic zebrafish lines
The mrc1a: KalTA4 transgene construct was assembled using an LR recombination reaction, using the destination vector pDestTol2pA/pA2 [81] and three entry clones: p5E-mrc1a [28], pME-KalTA4 (PCR-amplified from plasmid #10080, Addgene), and p3E-polyA from the Tol2kit [81]. The pME-KalTA4 construct was amplified using primers containing attB1/attB2 sites, and then cloned into the pDONR211 vector using BP Clonase (Cat# 11789-020, Thermo Fisher Scientific). The transgenic line Tg(mrc1a: KalTA4) was generated by microinjecting the mrc1a: KalTA4 transgene construct (55 pg) along with Tol2 transposase (50 pg) into one-cell stage zebrafish embryos from the Tg(UAS:eGFP) line. Embryos were subsequently screened for GFP expression in the lymphatic vasculature, leading to the establishment of the germline-transmitting line Tg(mrc1a: KalTA4; UAS:eGFP) y721.
Imaging methods
Embryos were anesthetized using 1x tricaine, mounted in 0.8-1.5% low-melting-point agarose dissolved in egg water, and mounted on a depression slide. Confocal images were acquired using a Nikon Ti2 inverted microscope with a Yokogawa CSU-W1 spinning disk equipped with a 10× Air 0.45 NA objective or a Zeiss LSM880 microscope. Brightfield images were obtained using a Leica M205 microscope. The images were analyzed using ImageJ, Adobe Photoshop (Adobe), and NIS-Elements (Nikon) software.
Morpholino injection
All MOs were obtained from Gene Tools, LLC (Philomath, OR), and injected into 1-2-cell-stage zebrafish embryos. The following MOs were used in this study: standard control MO: 5 5’-CCTCTTACCTCAGTTACAATTTATA-3’; shoc2 MO1: 5 5’-TACTGCTCATGGCGAAAGCCCCGCA-3’. Embryos were injected with 5.2 ng of Shoc2 MOs.
Shoc2 expression construct
The 6x-UAS:mCherry-2A-shoc2 construct was generated through a three-part LR recombination reaction using Gateway Technology. This process involved creating three entry clones and a destination vector. The 5’ entry clone (p5E 6xUAS) was constructed via traditional cloning methods. A linearized p5E-MCS vector, digested with XhoI and SpeI, served as the backbone, into which a 6xUAS fragment was inserted. This fragment was PCR-amplified from the 6x-UAS: EGFP plasmid, kindly provided by Marnie Halpern. The middle entry clone (pME mCherry-2A-shoc2) was generated using SLiCE (Seamless Ligation Cloning Extract, where 3 fragments were assembled by short stretches of homologous DNA recombination with the PPY SLiCE strain. The first fragment was a linearized pME-MCS vector, digested with EcoRI and SpeI. The second fragment was a PCR-amplified mCherry-2A fragment from a plasmid encoding mCherry-2A, and the third fragment was a full coding sequence of zebrafish shoc2 amplified using PCR. The 3’ entry clone, p3E polyA, containing the polyA sequence, was obtained from the Tol2kit. The final 6xUAS mCherry-2A-shoc2 construct was assembled by performing an LR recombination reaction using the destination vector, pDestTol2pA/pA2 [81], resulting in the insertion of the three entry clones into the destination vector.
Shoc2 rescue in zebrafish
Shoc2 rescue experiments were conducted in shoc2 morphants with the Tg(mrc1a:kalTA4; UAS:eGFP) transgenic background for endothelial rescue. The shoc2 MO (5’-TACTGCTCATGGCGAAAGCCCCGCA-3’; Gene Tools, LLC) was used at a concentration of 4.6 ng per embryo. For Shoc2 expression rescue, embryos were injected with either 6x-UAS:mCherry-2A-shoc2 or 6x-UAS:mCherry (control group) at a concentration of 40–45 ng per embryo, with or without shoc2 MO. DNA constructs were injected at the one-cell stage, followed by MO injection into the yolk at the one-to-four-cell stage.
Analysis of RNA-seq data
mRNAs were isolated from HDLECs expressing Shoc2 shRNA. For library preparation, mRNA was first isolated from total RNA using oligo(dT) magnetic beads and then sheared into approximately 200-base fragments. The cDNA library was sequenced using the Illumina NextSeq 500 sequencer. Quality control (QC) of the raw sequence data was performed using FastQC (version 0.11.7). The concatenated sequences were directly aligned to the Homo sapiens reference genome assembly (GRCh38) using STAR (version 2.6), producing BAM alignment files. Sequencing reads were trimmed and filtered using Trimmomatic (V0.39) to remove adapters and low-quality bases. Trimmed reads were mapped to the human reference genome assembly GRCh38 transcript annotation using RSEM. RSEM results normalization and differential expression analysis were performed using the R package EdgeR. Significantly up- or downregulated genes between groups were determined as having a fold change >= 2 and a q-value < 0.05. Transcript per million values were used to generate heatmaps, with represented values ranging from +1.5 (red, high expression) to −1.5 (blue, low expression). The RNA-seq data are publicly available as GEO series GSE288882. The data is MIAME compliant (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE288882).
Gene set enrichment analysis was performed using GSEA software and gene sets from the Molecular Signature Database (MSigDB). As recommended for GSEA from RNA sequencing experiments, genes with no counts in any sample were removed, along with low-expressing genes (mean or geometric mean <10 reads across all samples). As recommended in the GSEA user guide (https://www.gsea-msigdb.org/gsea/doc/GSEAUserGuideFrame.html), a false discovery rate (FDR) cut of 25% was used for Hallmark, KEGG, and Biological pathway gene set analysis. These data were the basis for the Venn diagram, heatmaps, and KEGG pathway analysis.
Statistical analyses
A sample size calculation was performed to ensure sufficient statistical power (usually 80% or higher) to detect the pre-determined effect size with a low risk of a Type I error (α, typically 0.05). This calculation identified the minimum number of samples needed to be reasonably confident that a true difference or relationship, as defined by the effect size, if present, would be detected in the study results. In animal studies, the number of animals per group was determined using a power analysis to ensure adequate statistical power to detect the pre-specified effect size between experimental groups, while maintaining a low probability of a false-positive result (Type I error, α, typically 0.05). This calculation aligns with the ethical principle of Replacement, Reduction, and Refinement (the 3Rs) by ensuring the fewest possible animals are used to achieve statistically meaningful and robust results. Results are expressed as means ± S.D. The statistical significance of the differences between groups was determined using a t-test or a one-way or two-way ANOVA (followed by Dunnett’s or Brown-Forsythe test, as indicated in the figure legends). p = 0.01 to 0.05 was considered statistically significant. (*p = 0.01 to 0.05, **p = 0.001 to 0.01, ***p = 0.0001 to 0.001, ****p < 0.0001, ≥ 0.05- non-significant, ns). GraphPad Prism 8.0 (GraphPad Prism Software Inc., Chicago, IL, USA) was used to perform all statistical analyses.
Data availability
This published article and its supporting information files include all data generated or analyzed during this study. The uncropped blots from this study are included as Supplementary Data. Other raw data from this study are available from the corresponding authors upon request.
References
Hu Z, Zhao X, Wu Z, Qu B, Yuan M, Xing Y, et al. Lymphatic vessel: origin, heterogeneity, biological functions and therapeutic targets. Signal Transduct Target Ther. 2024;9:9.
Petrova TV, Koh GY. Biological functions of lymphatic vessels. Science. 2020;369:eaax4063.
Stritt S, Koltowska K, Makinen T. Homeostatic maintenance of the lymphatic vasculature. Trends Mol Med. 2021;27:955–70.
Ricard N, Bailly S, Guignabert C, Simons M. The quiescent endothelium: signalling pathways regulating organ-specific endothelial normalcy. Nat Rev Cardiol. 2021;18:565–80.
Bui K, Hong YK. Ras pathways on prox1 and lymphangiogenesis: insights for therapeutics. Front Cardiovasc Med. 2020;7:597374.
Deng Y, Atri D, Eichmann A, Simons M. Endothelial ERK signaling controls lymphatic fate specification. J Clin Investig. 2013;123:1202–15.
Zenker M. Clinical overview on RASopathies. Am J Med Genet C Semin Med Genet. 2022;190:414–24.
Chen CP. Syndromic and single gene disorders associated with fetal pleural effusion (I): Noonan syndrome, RASopathy and congenital lymphatic anomalies. Taiwan J Obstet Gynecol. 2024;63:174–7.
Kleimeier LER, van Schaik C, Leenders E, Itkin M, Klein WM, Draaisma JMT. Lymphatic phenotype of Noonan syndrome: innovative diagnosis and possible implications for therapy. J Clin Med. 2022;11:3128.
Cordeddu V, Di Schiavi E, Pennacchio LA, Ma’ayan A, Sarkozy A, Fodale V, et al. Mutation of SHOC2 promotes aberrant protein N-myristoylation and causes Noonan-like syndrome with loose anagen hair. Nat Genet. 2009;41:1022–6.
Hannig V, Jeoung M, Jang ER, Phillips JA 3rd, Galperin E. A novel SHOC2 variant in rasopathy. Hum Mutat. 2014;35:1290–4.
Baldassarre G, Mussa A, Banaudi E, Rossi C, Tartaglia M, Silengo M, et al. Phenotypic variability associated with the invariant SHOC2 c.4A>G (p.Ser2Gly) missense mutation. Am J Med Genet A. 2014;164A:3120–5.
Garavelli L, Cordeddu V, Errico S, Bertolini P, Street ME, Rosato S, et al. Noonan syndrome-like disorder with loose anagen hair: a second case with neuroblastoma. Am J Med Genet A. 2015;167A:1902–7.
Gargano G, Guidotti I, Balestri E, Vagnarelli F, Rosato S, Comitini G, et al. Hydrops fetalis in a preterm newborn heterozygous for the c.4A>G SHOC2 mutation. Am J Med Genet A. 2014;164A:1015–20.
Motta M, Solman M, Bonnard AA, Kuechler A, Pantaleoni F, Priolo M, et al. Expanding the molecular spectrum of pathogenic SHOC2 variants underlying Mazzanti syndrome. Hum Mol Genet. 2022;31:2766–78.
Digilio MC, Lepri F, Baban A, Dentici ML, Versacci P, Capolino R, et al. RASopathies: clinical diagnosis in the first year of life. Mol Syndromol. 2011;1:282–9.
Zmolikova M, Puchmajerova A, Hecht P, Lebl J, Trkova M, Krepelova A. Coarctation of the aorta in Noonan-like syndrome with loose anagen hair. Am J Med Genet A. 2014;164A:1218–21.
Rodriguez-Viciana P, Oses-Prieto J, Burlingame A, Fried M, McCormick F. A phosphatase holoenzyme comprised of Shoc2/Sur8 and the catalytic subunit of PP1 functions as an M-Ras effector to modulate Raf activity. Mol Cell. 2006;22:217–30.
Wilson P, Abdelmoti L, Norcross R, Jang ER, Palayam M, Galperin E. The role of USP7 in the Shoc2-ERK1/2 signaling axis and Noonan-like syndrome with loose anagen hair. J Cell Sci. 2021;134:jcs258922.
Xie CM, Tan M, Lin XT, Wu D, Jiang Y, Tan Y, et al. The FBXW7-SHOC2-raptor axis controls the cross-talks between the RAS-ERK and mTORC1 signaling pathways. Cell Rep. 2019;26:3037–50.e4.
Jang ER, Jang H, Shi P, Popa G, Jeoung M, Galperin E. Spatial control of Shoc2-scaffold-mediated ERK1/2 signaling requires remodeling activity of the ATPase PSMC5. J Cell Sci. 2015;128:4428–41.
Jang H, Jang ER, Wilson PG, Anderson D, Galperin E. VCP/p97 controls signals of the ERK1/2 pathway transmitted via the Shoc2 scaffolding complex: novel insights into IBMPFD pathology. Mol Biol Cell. 2019;30:1655–63.
Jang ER, Galperin E. The function of Shoc2: a scaffold and beyond. Commun Integr Biol. 2016;9:e1188241.
Kaduwal S, Jeong WJ, Park JC, Lee KH, Lee YM, Jeon SH, et al. Sur8/Shoc2 promotes cell motility and metastasis through activation of Ras-PI3K signaling. Oncotarget. 2015;6:33091–105.
Jang H, Oakley E, Forbes-Osborne M, Kesler MV, Norcross R, Morris AC, et al. Hematopoietic and neural crest defects in zebrafish shoc2 mutants: a novel vertebrate model for Noonan-like syndrome. Hum Mol Genet. 2019;28:501–14.
Yi J, Chen M, Wu X, Yang X, Xu T, Zhuang Y, et al. Endothelial SUR-8 acts in an ERK-independent pathway during atrioventricular cushion development. Dev Dyn. 2010;239:2005–13.
Yaniv K, Isogai S, Castranova D, Dye L, Hitomi J, Weinstein BM. Imaging the developing lymphatic system using the zebrafish. Novartis Found Symp. 2007;283:139–48.
Jung HM, Castranova D, Swift MR, Pham VN, Venero Galanternik M, Isogai S, et al. Development of the larval lymphatic system in zebrafish. Development. 2017;144:2070–81.
Yaniv K, Isogai S, Castranova D, Dye L, Hitomi J, Weinstein BM. Live imaging of lymphatic development in the zebrafish. Nat Med. 2006;12:711–6.
Cha YR, Fujita M, Butler M, Isogai S, Kochhan E, Siekmann AF, et al. Chemokine signaling directs trunk lymphatic network formation along the preexisting blood vasculature. Dev Cell. 2012;22:824–36.
Diamond MS, Farzan M. The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nat Rev Immunol. 2013;13:46–57.
Kang JA, Kim YJ, Jeon YJ. The diverse repertoire of ISG15: more intricate than initially thought. Exp Mol Med. 2022;54:1779–92.
Bluyssen HA, Vlietstra RJ, van der Made A, Trapman J. The interferon-stimulated gene 54 K promoter contains two adjacent functional interferon-stimulated response elements of different strength, which act synergistically for maximal interferon-alpha inducibility. Eur J Biochem. 1994;220:395–402.
Ronni T, Matikainen S, Lehtonen A, Palvimo J, Dellis J, Van Eylen F, et al. The proximal interferon-stimulated response elements are essential for interferon responsiveness: a promoter analysis of the antiviral MxA gene. J Interferon Cytokine Res. 1998;18:773–81.
Ohtomo T, Sugamata Y, Ozaki Y, Ono K, Yoshimura Y, Kawai S, et al. Molecular cloning and characterization of a surface antigen preferentially overexpressed on multiple myeloma cells. Biochem Biophys Res Commun. 1999;258:583–91.
Platanias LC. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat Rev Immunol. 2005;5:375–86.
Ko CY, Chang WC, Wang JM. Biological roles of CCAAT/Enhancer-binding protein delta during inflammation. J Biomed Sci. 2015;22:6.
Hu J, Roy SK, Shapiro PS, Rodig SR, Reddy SP, Platanias LC, et al. ERK1 and ERK2 activate CCAAAT/enhancer-binding protein-beta-dependent gene transcription in response to interferon-gamma. J Biol Chem. 2001;276:287–97.
Wies E, Wang MK, Maharaj NP, Chen K, Zhou S, Finberg RW, et al. Dephosphorylation of the RNA sensors RIG-I and MDA5 by the phosphatase PP1 is essential for innate immune signaling. Immunity. 2013;38:437–49.
Reikine S, Nguyen JB, Modis Y. Pattern recognition and signaling mechanisms of RIG-I and MDA5. Front Immunol. 2014;5:342.
Liu G, Lee JH, Parker ZM, Acharya D, Chiang JJ, van Gent M, et al. ISG15-dependent activation of the sensor MDA5 is antagonized by the SARS-CoV-2 papain-like protease to evade host innate immunity. Nat Microbiol. 2021;6:467–78.
Sun X, Sun L, Zhao Y, Li Y, Lin W, Chen D, et al. MAVS maintains mitochondrial homeostasis via autophagy. Cell Discov. 2016;2:16024.
Marques E, Kramer R, Ryan DG. Multifaceted mitochondria in innate immunity. NPJ Metab Health Dis. 2024;2:6.
Lopez-Polo V, Maus M, Zacharioudakis E, Lafarga M, Attolini CS, Marques FDM, et al. Release of mitochondrial dsRNA into the cytosol is a key driver of the inflammatory phenotype of senescent cells. Nat Commun. 2024;15:7378.
Xie CM, Sun Y. The MTORC1-mediated autophagy is regulated by the FBXW7-SHOC2-RPTOR axis. Autophagy. 2019;15:1470–72.
Lapinski PE, Lubeck BA, Chen D, Doosti A, Zawieja SD, Davis MJ, et al. RASA1 regulates the function of lymphatic vessel valves in mice. J Clin Investig. 2017;127:2569–85.
Norcross RG, Abdelmoti L, Rouchka EC, Andreeva K, Tussey O, Landestoy D, et al. Shoc2 controls ERK1/2-driven neural crest development by balancing components of the extracellular matrix. Dev Biol. 2022;492:156–71.
Bonsor DA, Simanshu DK. RAS and SHOC2 roles in RAF activation and therapeutic considerations. Annu Rev Cancer Biol. 2024;8:97–113.
Terai H, Hamamoto J, Emoto K, Masuda T, Manabe T, Kuronuma S, et al. SHOC2 is a critical modulator of sensitivity to EGFR-TKIs in non-small cell lung cancer cells. Mol Cancer Res. 2021;19:317–28.
Sulahian R, Kwon JJ, Walsh KH, Pailler E, Bosse TL, Thaker M, et al. Synthetic lethal interaction of SHOC2 depletion with MEK inhibition in RAS-driven cancers. Cell Rep. 2019;29:118–34.e8.
You X, Dou L, Tan M, Xiong X, Sun Y. SHOC2 plays an oncogenic or tumor-suppressive role by differentially targeting the MAPK and mTORC1 signals in liver cancer. Life Med. 2024;3:lnae23.
Yu B, Ma J, Li J, Wang D, Wang Z, Wang S. Mitochondrial phosphatase PGAM5 modulates cellular senescence by regulating mitochondrial dynamics. Nat Commun. 2020;11:2549.
Chen W, Zhao H, Li Y. Mitochondrial dynamics in health and disease: mechanisms and potential targets. Signal Transduct Target Ther. 2023;8:333.
Carroll B, Nelson G, Rabanal-Ruiz Y, Kucheryavenko O, Dunhill-Turner NA, Chesterman CC, et al. Persistent mTORC1 signaling in cell senescence results from defects in amino acid and growth factor sensing. J Cell Biol. 2017;216:1949–57.
de la Cruz López KG, Toledo Guzmán ME, Sánchez EO, García Carrancá A. mTORC1 as a regulator of mitochondrial functions and a therapeutic target in cancer. Front Oncol. 2019;9:1373.
Morita M, Prudent J, Basu K, Goyon V, Katsumura S, Hulea L, et al. mTOR controls mitochondrial dynamics and cell survival via MTFP1. Mol Cell. 2017;67:922–35.e5.
Yoon J, Kim S, Lee M, Kim Y. Mitochondrial nucleic acids in innate immunity and beyond. Exp Mol Med. 2023;55:2508–18.
Shao X, Liu C. Influence of IFN- alpha and IFN- gamma on lymphangiogenesis. J Interferon Cytokine Res. 2006;26:568–74.
Kemp SS, Penn MR, Koller GM, Griffin CT, Davis GE. Proinflammatory mediators, TNFalpha, IFNgamma, and thrombin, directly induce lymphatic capillary tube regression. Front Cell Dev Biol. 2022;10:937982.
Lucas ED, Tamburini BAJ. Lymph node lymphatic endothelial cell expansion and contraction, and the programming of the immune response. Front Immunol. 2019;10:36.
Embry A, Baggett NS, Heisler DB, White A, de Jong MF, Kocsis BL, et al. Exploiting bacterial effector proteins to uncover evolutionarily conserved antiviral host machinery. PLoS Pathog. 2024;20:e1012010.
Zhao BR, Wang XX, Wang XW. Shoc2 recognizes bacterial flagellin and mediates antibacterial Erk/Stat signaling in an invertebrate. PLoS Pathog. 2022;18:e1010253.
Dard L, Blanchard W, Hubert C, Lacombe D, Rossignol R. Mitochondrial functions and rare diseases. Mol Asp Med. 2020;71:100842.
Dard L, Hubert C, Esteves P, Blanchard W, Bou About G, Baldasseroni L, et al. HRAS germline mutations impair LKB1/AMPK signaling and mitochondrial homeostasis in Costello syndrome models. J Clin Investig. 2022;132:e131053.
Bajia D, Bottani E, Derwich K. Effects of Noonan syndrome-germline mutations on mitochondria and energy metabolism. Cells. 2022;11:3099.
Lee I, Pecinova A, Pecina P, Neel BG, Araki T, Kucherlapati R, et al. A suggested role for mitochondria in Noonan syndrome. Biochim Biophys Acta. 2010;1802:275–83.
Zhang EE, Chapeau E, Hagihara K, Feng GS. Neuronal Shp2 tyrosine phosphatase controls energy balance and metabolism. Proc Natl Acad Sci USA. 2004;101:16064–9.
Kleefstra T, Wortmann SB, Rodenburg RJ, Bongers EM, Hadzsiev K, Noordam C, et al. Mitochondrial dysfunction and organic aciduria in five patients carrying mutations in the Ras-MAPK pathway. Eur J Hum Genet. 2011;19:138–44.
Rigante D, Leoni C, Onesimo R, Giorgio V, Trevisan V, Zampino G. Aberrant N-myristoylation as a prelude to autoimmune manifestations in patients with SHOC2 mutations. Autoimmun Rev. 2023;22:103462.
Quaio CR, Carvalho JF, da Silva CA, Bueno C, Brasil AS, Pereira AC, et al. Autoimmune disease and multiple autoantibodies in 42 patients with RASopathies. Am J Med Genet A. 2012;158A:1077–82.
Siano MA, Marchetti V, Pagano S, Di Candia F, Alessio M, De Brasi D, et al. Risk of autoimmune diseases in patients with RASopathies: systematic study of humoral and cellular immunity. Orphanet J Rare Dis. 2021;16:410.
Bader-Meunier B, Cave H, Jeremiah N, Magerus A, Lanzarotti N, Rieux-Laucat F, et al. Are RASopathies new monogenic predisposing conditions to the development of systemic lupus erythematosus? Case report and systematic review of the literature. Semin Arthritis Rheum. 2013;43:217–9.
Wang XO, Liu ZQ, Shangguan SF, Lai JM, Wang PC, Song FY, et al. [Clinical characteristics analysis of children with Noonan-like syndrome with loose anagen hair]. Zhonghua Er Ke Za Zhi. 2025;63:405–10.
Menentoğlu B, Konur Akbaş EN, Kavrul Kayaalp G, Akgün Ö, Aslanger AD, Uyguner ZO, et al. Complex crosstalk: early-onset lupus and antiphospholipid syndrome in a child with SHOC2-related RASopathy. Rheumatology. 2025;64:4852–3.
Okazaki T, Saito Y, Sugita K, Nosaka K, Ohno K, Hiraoka Y, et al. Recurrent erythema nodosum in a child with a SHOC2 gene mutation. Yonago acta Med. 2019;62:159–62.
Jeoung M, Galperin E. Visualizing of signaling proteins on endosomes utilizing a knockdown and reconstitution approach. Methods Enzym. 2014;534:47–63.
Kim DH, Sarbassov DD, Ali SM, King JE, Latek RR, Erdjument-Bromage H, et al. mTOR interacts with raptor to form a nutrient-sensitive complex that signals to the cell growth machinery. Cell. 2002;110:163–75.
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Dev Dyn. 1995;203:253–310.
Fujita M, Cha YR, Pham VN, Sakurai A, Roman BL, Gutkind JS, et al. Assembly and patterning of the vascular network of the vertebrate hindbrain. Development. 2011;138:1705–15.
Lawson ND, Weinstein BM. In vivo imaging of embryonic vascular development using transgenic zebrafish. Dev Biol. 2002;248:307–18.
Kwan KM, Fujimoto E, Grabher C, Mangum BD, Hardy ME, Campbell DS, et al. The Tol2kit: a multisite gateway-based construction kit for Tol2 transposon transgenesis constructs. Dev Dyn. 2007;236:3088–99.
Acknowledgements
The authors thank Dr. Tianyan Gao for critically reading the manuscript and providing valuable reagents, as well as Dr. Saurabh Chattopadhyay and Dr. Michaela Guck for their valuable reagents. The authors also thank Ms. Lina Abdelmoti for the excellent care of the zebrafish colony.
Funding
This project was supported by grants from the National Institute of General Medical Sciences (R35GM136295 and 1S10OD025033-01 to EG) and the Eunice Kennedy Shriver National Institute of Child Health and Human Development (R01HD118715 to EG). BMW was supported by the intramural program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development (ZIA-HD001011 and ZIA-HD008808). This research was supported by the Biostatistics and Bioinformatics and Redox Metabolism Shared Resources of the University of Kentucky Markey Cancer Center (P30CA177558). Its contents are solely the authors’ responsibility and do not necessarily represent the official views of the National Institute of Health. HMJ was supported by the UIC College of Medicine start-up fund.
Author information
Authors and Affiliations
Contributions
Conceptualization: EG, BMW, and HMJ. Investigation: PW, VV, RN, KK, VNP, BMW, HMJ, and EG; Methodology: PW, VV, RN, KK, VNP, and HMJ. Resources: EG, BMW, and HMJ. Supervision: EG, BMW, and HMJ. Funding acquisition: EG, BMW, and HMJ. Writing: EG, BMW, and HMJ co-wrote the paper with input from authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All methods detailed in this study were performed in accordance with the relevant guidelines and regulations. This study only involved live vertebrates, and no human participants were involved. For the component of this study involving live vertebrates the ethics approval was obtained from the University of Kentucky Animal Care and Use Committee (#2019-3208), the University of Illinois at Chicago Animal Care and Use Committee (#23-112), and the Eunice Kennedy Shriver National Institute of Child Health and Human Development Committee (Animal Study Proposal #21-015, 24-015).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wilson, P., Vishwakarma, V., Norcross, R. et al. Signaling scaffold Shoc2 regulates lymphangiogenesis by suppressing mTORC1-mediated IFN responses. Cell Death Differ (2026). https://doi.org/10.1038/s41418-026-01730-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41418-026-01730-9