Abstract
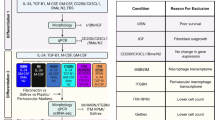
Mucopolysaccharidoses are inherited metabolic disorders caused by the deficiency in lysosomal enzymes required to break down glycosaminoglycans. Accumulation of glycosaminoglycans leads to progressive, systemic degenerative disease. The central nervous system is particularly affected, resulting in developmental delays, neurological regression, and early mortality. Current treatments fail to adequately address neurological defects. Here we explore the potential of human induced pluripotent stem cell (hiPSC)-derived microglia progenitors as a one-time, allogeneic off-the-shelf cell therapy for several mucopolysaccharidoses (MPS). We show that hiPSC-derived microglia progenitors, possessing normal levels of lysosomal enzymes, can deliver functional enzymes into four subtypes of MPS knockout cell lines through mannose-6-phosphate receptor-mediated endocytosis in vitro. Additionally, our findings indicate that a single administration of hiPSC-derived microglia progenitors can reduce toxic glycosaminoglycan accumulation and prevent behavioral deficits in two different animal models of MPS. Durable efficacy is observed for eight months after transplantation. These results suggest a potential avenue for treating MPS with hiPSC-derived microglia progenitors.
Similar content being viewed by others
Introduction
Lysosomal storage disorders (LSDs) are a group of inherited metabolic conditions characterized by deficiency in a lysosomal enzyme and pathological accumulation of undegraded substrates, causing systemic dysfunction and reduced life expectancy. Over 50 different LSDs exist, each one with distinctive disease manifestation. As a group, they have an incidence of 1 in 5000-8000 live births1, with specific LSDs further varying in rarity. Mucopolysaccharidoses are a class of LSDs characterized by the accumulation of glycosaminoglycans (GAGs), and although they share many clinical features, they present varying degrees of severity. While some MPS subtypes include primarily neurological findings with rapid progression, others such as MPS I-S (Scheie syndrome) and MPS I H-S (Hurler-Scheie syndrome) lead to comprehensive systemic clinical findings, including skeletal deformities, respiratory complications, cardiac valve deficits, hepatosplenomegaly, among other defects. Notably, central nervous system (CNS) impairment is characteristic of MPS, such as MPS I-H (Hurler syndrome), MPS II (Hunter syndrome), MPS III (Sanfilippo syndrome), and MPS VII (Sly syndrome)2,3. Neurological regression and developmental delays happen gradually, and an abbreviated life span is typically observed3.
The current standard of care (SOC) for MPS I, MPS II, and MPS VII is enzyme replacement therapy (ERT), whereas MPS III lacks etiology-based therapy and relies on support and ancillary care. In some severe cases, hematopoietic stem cell transplantation (HSCT) has been performed, with equivocal results2. Both ERT and HSCT provide a source of the deficient enzyme in the patient4,5. In the case of ERT, the Blood Brain Barrier (BBB)6 restricts efficacy mostly to the periphery and requires frequent administration, becoming a cumbersome, lifelong, and high-cost treatment. Meanwhile, HSCT typically requires 4 or more weeks of lead time and results in limited CNS engraftment, which can take up to 12 months7 and a suitable donor cannot always be identified8. This delay in CNS treatment is in some cases detrimental as CNS manifestations can be irreversible.
We considered if these shortcomings could potentially be overcome by introducing a CNS-native cell type directly into the brain. In LSDs, donor cells can rescue neighboring, diseased cells through metabolic cross-correction, where the functional enzyme is secreted and transported into the deficient cell via receptor-mediated endocytosis9. Healthy microglia, as tissue-resident macrophages of the CNS, thus could serve the role of an enzyme donor to correct the neurological pathologies seen in LSDs. Microglia originate from the embryonic yolk sac10 as primitive hematopoietic progenitors that give rise to tissue-resident macrophages. Several protocols to derive microglia from human induced pluripotent stem cells (hiPSCs) have been developed in the last decade11,12,13,14. Here we describe the use of hiPSC-derived microglia progenitors (MG01) from healthy donors as a potential one-time treatment of the CNS biology in MPS. We show that in animal models, MG01 can be delivered into the CNS and provide functional enzymes in an immediate and sustained manner, preventing further neurodegeneration. This opens the opportunity to develop MG01 as a therapy for MPSs and, potentially, other LSDs.
Results
MG01 are functional microglia derived from hiPSCs
Microglia progenitors were derived from hiPSCs following the addition of a series of morphogens and growth factors recapitulating primitive myelopoiesis (Fig. 1A). For studies in this manuscript, we used a publicly available cell line (Line 6), and 2 proprietary cell lines. For the differentiation, the mesodermal lineage was induced by high levels of bone morphogenetic protein 4 (BMP4, Day 0 to 4) followed by myeloid lineage specification and expansion (Day 4 onwards). Floating cells, termed MG01, separated from a layer of hemogenic endothelium and were collected from the culture supernatant around day 24–30 of the differentiation and then cryopreserved. This protocol was highly efficient (generating approximately 43 MG01 cells per input hiPSC), and reproducible across 3 hiPSC lines (Fig. 1B). MG01 recovered from cryopreservation with high viability in all 3 hiPSC lines (Fig. 1C) and exhibited canonical microglia markers, including CD45, CD14, CD11B and CX3CR1 (Fig. 1D). Upon plating, MG01 exhibited expected morphology for (primary and iPSCs-derived) microglia14,15,16 (Fig. 1E, Fig. 1F) and exhibited immunoreactivity for prototypical pan-myeloid and microglia markers, including CD45, CD68, TMEM119, IBA1, P2RY12, PU.1 and TREM2 (Fig. 1E). Cytokine secretion in response to external stimuli and phagocytosis are two main characteristics of microglia. When MG01 were challenged with IFNγ and LPS for 48hrs, they became activated (M1-like) (Fig. 1F), secreted pro-inflammatory cytokines (Fig. 1G), and showed reduced phagocytosis of bacterial particles when compared to non-stimulated MG01 (M0-like) after 2 days in vitro (DIV). Extended culture of MG01 maintains phagocytic activity (Fig. 1H, Supplementary Fig. 1A, B) and activation occurs in a dose-dependent manner, with differential phagocytosis and cytokine expression phenotypes according to the dose and type of challenge (Supplementary Fig. 1C), as previously described for iPSC-derived15 and primary human microglia16.
A Schematic of the differentiation process for MG01. B Total MG01 cells produced in a T-75 flask for 3 hiPSC lines. C Viability of MG01 after thaw for each of the 3 hiPSC lines. D Representative flow-cytometry plots for CD45/CD14 and CX3CR1/CD11B in post-thaw MG01 and graphs depicting quantification of 3 biological replicates per line. E Panel of representative images depicting immunoreactivity of MG01 (Line 82) to CD45, CD68, P2RY12, TREM2, IBA1, TMEM119 and PU.1 after two days in culture. F Representative phase contrast images of polarized MG01 (Line 82) 12 hours after the addition of pHrodo-conjugated E. coli particles. Upper left insets depict the detected fluorescence of pHrodo particles after phagocytosis. G Pro-inflammatory cytokine (TNFα, CXCL10 and IL-6) expression levels after polarization of MG01 to M1-like state using LPS and IFNγ. H Quantification of representative phagocytosis assay (2DIV). Graph depicts pHrodo fluorescence after addition to MG01. Images were acquired every 30 m for a span of 24 h. Data points in B, and C represent biological replicates; in D represent a combination of biological and technical replicates, in G–H represent technical replicates from one representative experiment. All cell lines tested behaved similarly and data depicted in E–H is representative of all hiPSC-derived MG01. Graphs denote mean +/− SEM. Scale bars in panel E represent 50 µm. Panel A was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Engrafted MG01 upregulate microglial signature genes
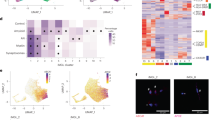
To further characterize MG01, we performed single-cell RNA sequencing (scRNAseq) and compared the data to publicly available dataset from CD45+ primary hematopoietic cells, isolated from human embryos17. Among the 15 clusters of primary hematopoietic cells found in the Bian et al. dataset17 (Fig. 2A), there were 4 macrophage clusters (clusters 1-3 are treated as one to simplify analysis). The Mac 4 cluster shares many genes with Mac 1-3 but also has a unique expression of SALL1 and higher expression of TMEM119, TREM2, MERTK, and P2RY12, suggesting a microglia-specific profile. Using principal-component analysis (PCA), we observed that cryopreserved MG01 groups closely with Mac 1-3, and to a lesser extent, Mac 4, while fetal non-macrophage cells (the remaining 11 clusters) lie farther along the PC axes (Fig. 2C). We scored the identity of MG01 compared to cell signatures of fetal macrophages and found it to be most similar to Mac 1-3 with a fraction of cells similar to Mac 4. In vitro systems are unable to fully recapitulate the in vivo environmental, spatial, and temporal conditions, and thus, as expected in vitro cultured MG01 share the Mac1-3 identity, recapitulating the cells arising earlier in development (CS11-CS15). However, both clusters are stages of sequential development17 of Myb-independent18 yolk sac-derived macrophages into the microglial fate, indicating that MG01 cells are in the developmental trajectory towards microglial fate.
A UMAP plots of the Bian et al., 2020 dataset of fetal hematopoietic (CD45+) cells depicting the origin, Carnegie stage and cell type grouping of all cells. B Same UMAP plot as (A) displaying expression level of representative markers for Mac 1-3 and Mac 4 groups. C PCA of MG01 (Line 82) and Bian et al. dataset (N = 3 MG01 differentiations). PC plots on the right display the identity score for each cell when compared to the Mac 1-3 or Mac 4 signature. Violin plots display the distribution of results for each sample. D Heatmap of differentially expressed markers in each cell group of Bian et al. when compared to all other groups combined. E Schematic depicting the workflow for bulk RNAseq of engrafted MG01. F Dot plot displaying the average expression and percentage of post-thaw MG01 cells expressing Mac 4 and Mac 1-3 genes in each group. G Heatmap of differentially expressed genes between MG01 and engrafted MG01, including genes in the Mac 4 cell group in D, F (N = 1 MG01 differentiation, 3 technical replicates). Transcript data is scaled by gene. Green arrows highlight known microglial markers. Panel E was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
To further address this, we reasoned that in vivo maturation of MG01 may establish microglia identity and start expressing Mac 4 signature genes more robustly. To assess the fate of MG01 in vivo, we transplanted them in the immunocompromised NSG-Quad (NSG-Q) mouse model that expresses the human cytokines CSF1, GM-CSF, SCF, and IL-3, supporting human myeloid survival and engraftment19. We then collected and compared engrafted cells and post-thaw MG01 (non-transplanted) by bulk RNAseq (Fig. 2E) and probed for differential expression of genes in clusters Mac 4 and Mac 1-3 (Fig. 2D, F) again. As predicted, engrafted MG01 now show upregulation of microglia-signature genes when compared to post-thaw MG01, such as SALL1, TMEM119, P2RY12, OLFML3, P2RY13, MERTK, and TREM2 (Fig. 2G). These genes are part of the Mac 4 transcriptomic signature, suggesting that MG01 are yolk sac-derived macrophages that mature into a microglial fate after in vivo engraftment in the brain.
MG01 provides functional enzymes in an in vitro cross-correction model
MG01 as a potential therapy for MPS relies on the secretion of enzyme that can be transferred to enzyme-deficient cells, to restore lysosomal function and reduce the accumulation of toxic GAG substrates. MG01 can secrete the missing enzymes of MPS I and MPS VII, as demonstrated by measurable alpha-L-iduronidase (IDUA) and beta-glucuronidase (GUSB) activity respectively, in MG01-conditioned medium (Fig. 3A) across 3 hiPSC lines. We next tested if MG01-secreted enzymes can be transferred to enzyme-deficient cells. For this, we generated four knockout iPSC lines by genetically ablating one lysosomal enzyme gene (namely IDUA, GUSB, SGSH encoding sulfamidase or NAGLU encoding alpha-N-acetylglucosaminidase) in a parental hiPSC line and then differentiated these individual knockout hiPSCs into microglia (MG-KO). We developed a transwell co-culture assay where MG01 were seeded in the lower compartment, and MG-KO lines were seeded on top of the insert (Fig. 3B). After nine or ten days of culture, in the absence of MG01, all four MG-KO lines had accumulated GAGs, as one would expect from patient-derived tissues and cells20,21,22. However, when the same cells were co-cultured with MG01 they showed reduced levels of GAGs (approaching healthy levels), suggesting successful transfer of functional enzyme from MG01 (Fig. 3C–F). Indeed, lysates from enzyme-deficient MG-KO lines co-cultured with MG01 exhibited detectable intra-cellular enzymatic activity, confirming the successful transfer of the functional enzyme that was missing (Supplementary Fig. 2A–D). Notably, even an increase in enzymatic activity, substantially below wild-type levels, was sufficient to achieve complete restoration of GAG levels over 10 days (Fig. 3D, F). This was especially noticeable in the SGSH MG-KO cells (Fig. 3E), where GAG levels were reduced to wild-type levels, despite our inability to detect a considerable amount of SGSH activity, likely because of limitations of the assay system available. As a surrogate method, we demonstrated the presence of measured SGSH protein in MG-KO lysate only after co-culturing with MG01 (Supplementary Fig. 2D).
A Enzymatic activity levels of IDUA and GUSB were found in the supernatant of MG01 cells after 96 h in vitro. B Schematic depicting the transwell-based system. Each graph denotes test cross-correction of enzymes by MG01 to MG-KO cells. Accumulation of GAGs in cell lysates of MG01, MG-KOs, and MG-KOs co-cultured with MG01 for 10 days. Each graph denotes a MG-KO deficiency in a different enzyme: C IDUA, D GUSB, E SGSH, and F NAGLU. Values are depicted as a percentage of the GAG levels found in untreated MG-KOs at the end of the assay. G Accumulation of GAGs in cell lysates of MG-KO co-cultured with MG01 in the presence and absence of a M6P receptor ligand. GAG levels were measured as μg/ml and values are depicted as a percentage of the GAG levels found in untreated MG-KOs at the end of the assay. Graphs denote mean +/− SEM. Data points in A represent the analysis of biological replicates, in C–G represent technical replicates from one representative experiment. Panel B was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
To further provide evidence of the modulation of lysosomal processing due to the uptake of MG01-derived enzymes, we assessed lysosomal changes in IDUA and GUSB MG-KOs with and without co-culture with MG01. LAMP1 and LAMP2 are lysosomal membrane proteins often assayed as markers for lysosomal function and modulation23,24. In MG01 co-cultures, the total LAMP1 protein was unchanged in GUSB−/− MG-KO and in IDUA−/− MG-KO with a variable LAMP1 protein amount in the latter one (Supplementary Fig. 2E). Using an orthogonal method, we quantified intensity of LAMP1 immunoreactivity, and we detected normalization of LAMP1 total signal in MG-KO exposed to MG01-conditioned media (Supplementary Fig. 2F-G). Conversely, total LAMP2 remained unchanged in IDUA−/− MG-KO while co-culture with MG01 restored GUSB−/− MG-KO to WT levels (Supplementary Fig. 2H). Finally, to underscore the potential of MG01 to rescue other cell types, we differentiated GUSB−/− hiPSCs to cortical neurons and co-cultured them with MG01, where we found the expected restoration of GAG accumulation to WT levels and modulation of total LAMP1 and LAMP2 levels (Supplementary Fig. 2I, J).
Lysosomal enzyme uptake is mainly mediated via the mannose-6-phosphate (M6P)25 receptor but there is evidence of M6P receptor-independent pathways. To elucidate the mechanism of MG01 enzyme transfer and GAG reduction, we performed the transwell co-culture assay in the presence of a competitive ligand (e.g. M6P). We show that in the presence of M6P, MG01 partially reduced the accumulation of GAGs, suggesting that M6P pathway is partly responsible for the transfer of the missing enzyme (Fig. 3G), with alternative pathways of lysosomal targeting being responsible for the remainder of the transfer (LIMP-2, sortilin26).
Distribution of MG01 in the CNS of mouse models
We and others have shown that human myeloid engraftment in the mouse can be impacted by limited cross-reactivity of murine cytokines with human cells, especially hCSF127. However, we were encouraged by pilot studies using MG01 in mouse models that were immunocompromised, but lacked the optimal human cytokine environment (Supplementary Fig. 3A, B). Thus, to assess the biodistribution and efficacy of MG01 in disease models we leveraged existing immunodeficient mouse models of MPS I and MPS VII, which phenocopy the human diseases by showing similar peripheral and CNS degeneration while also accepting xenografts. We chose the intracerebroventricular (ICV) route for MG01 delivery as it facilitates CNS-wide engraftment of MG01 for increased biodistribution of donor enzymes, and is a suitable clinical route of administration28.
Initially, we evaluated the engraftment in NOD/scid-MPS I mice 4-months post-administration and found successful engraftment of MG01 derived from a publicly available hiPSC line (Line 6). We then assessed MG01 engraftment 1-, 5- or 8-months after ICV delivery in juvenile (p21) or adult (p60) MPS I or MPS VII animals with MG01 derived from cell lines developed for clinical use (Line 82 or Line 83, Fig. 4A, Supplementary Figs. 4, 5). MG01 derived from all three (6, 82, 83) iPSC lines engrafted in the choroid plexus and meninges, surrounding the brain at all time points examined (Fig.4B, Supplementary Fig. 3B, Supplementary Fig. 4). In some cases, MG01 further integrated into the brain parenchyma and assumed a ramified morphology, reminiscent of resting primary microglia (Fig. 4B, E; Supplementary Fig. 3B, C). As expected from this route of delivery, MG01 was also found engrafted throughout the length of the spinal cord, in the pial surface or perivascular spaces (Supplementary Fig. 5). ICV administration resulted in heterogenous patterns of engraftment between animals, with the majority of MG01 found in the meningeal space, but representation was found consistently across the brain in all cohorts (Supplementary Fig. 4).
A Schematic denoting the experimental design of ICV transplantations in MPS I (IDUA-KO) and MPS VII (GUSB-KO) mouse models. B Representative pictures of engraftment of MG01 in MPS I and MPS VII mouse brain sections detectable by the presence of hNA (human nuclear antigen). MG01 will consistently be immunoreactive to IBA1 (microglia marker) but only positive for CD163 (meningeal macrophages) in certain niches. C PU.1 immunoreactivity in engrafted MG01. D CD45 and ku80 (human nuclei) immunoreactivity on MG01 engrafted in the meninges. E TMEM119 and CD45 immunoreactivity in arborized MG01 that have engrafted in the parenchyma. F STEM121 (human cells) and SALL1 immunoreactivity in arborized MG01 engrafted in the parenchyma. Micrographs are representative of results from N ≥ 3 animals for each condition in all cohorts. Scale bars in B represent 100 µm; in C 50 µm; in D 20 µm; in E–F 50 µm. Panel A was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Further biodistribution data were obtained by droplet digital PCR (ddPCR) detecting hTERT (human telomerase reverse transcriptase; Supplementary Fig. 6) in blood and highly perfused tissues, at 1-month after treatment with MG01, confirmed that engraftment of ICV administered MG01 is restricted to the CNS.
Additionally, to demonstrate engraftment and biodistribution, we counted the presence of human cells in one brain hemisphere of MG01-treated animals, where technically possible. MG01 was found to be distributed across the brain (by sagittal sampling), in a similar pattern for all animals, with higher amounts of MG01 located midway through the hemisphere, although with substantial variability across animals and doses (Supplementary Fig. 7A).
During development, both meningeal macrophages and microglia originate from the same yolk-sac progenitor29 and adopt distinctive phenotypes influenced by their microenvironment. Thus, MG01 engrafted in the meninges were immunoreactive for IBA1, CD163 (Fig. 4B, Supplementary Fig. 3D, yellow arrows), PU.1 (Fig. 4C, Supplementary Fig. 3D) and CD45 (Fig. 4D), while parenchymal MG01 were arborized, maintained IBA1 (Fig. 4B, Supplementary Fig. 3B, green arrows) and CD45 (Fig. 4E) expression, and were positive for TMEM119 (Fig. 4E Supplementary Fig. 3C) and SALL1 (Fig. 4F). We further quantified the proliferative fraction of MG01 across timepoints and found that at 1-month, in both disease models, Ki67+ cells comprised 5-6.7% of total MG01, which was reduced to 1-2.2% after 5-months in vivo (Supplementary Fig. 8). MG01 were detected up to 8-months after transplantation (Supplementary Fig. 4B), indicating long-term engraftment and survival. No tumors were observed by histopathology at any time point evaluated. Later timepoints were not evaluated due to limited life expectancy of mice on NOD.Prkdcscid background (on average 8.5-months30).
As discussed, the presence of hCSF1 and other myeloid-supportive human cytokines (hGM-CSF, hIL-3, hSCF) increases engraftment when compared to NOD/scid models and may differentially impact meningeal versus parenchymal engraftment31,32,33. To better establish engraftment potential of MG01, we assessed engraftment in NSG-Q mice, 1 month after ICV transplantation of MG01 and compared it to MPS I and MPS VII engraftment at the same timepoint (Supplementary Fig. 7B). The animal age at time of grafting also has a large effect in engraftment success. And whilst grafting prior to P21 exhibits the best engraftment (P1/P7)19,27,34, we reasoned it would be challenging to translate to humans at an equivalent developmental period, as P0-P7 in mice is equivalent to pre-term infants35. NOD/scid animals showed similar, modest levels of MG01 engraftment at both P21 and P60-90, modeling juvenile and adult engraftment respectively (Fig. 4A, Supplementary Figs. 3–5, 7B). However, we conclude that, although not optimal, NOD/scid models allow for sufficient engraftment to test the therapeutic effects of MG01 in MPS I and MPS VII models. Further studies in models that optimally support human myeloid engraftment will be needed to fully describe and quantify MG01 engraftment.
MG01 prevents biochemical pathology in the CNS of MPS I mice
GAG accumulation and IDUA deficiency are the biochemical hallmarks of MPS I which we intended to evaluate in MPS I mice that received an ICV injection of MG01 as juvenile or adult animals. We initially tested Line 6-derived MG01 in adult MPS I mice and found a significant reduction of GAGs 4-months post-delivery of MG01 compared to animals transplanted with IDUA−/− MG-KO (Fig. 5A, B, Supplementary Table 1). MG-KO (IDUA−/−) served as an additional, cell only control, over vehicle in these experiments to demonstrate that the activity is in fact IDUA dependent. IDUA enzyme activity also increased in MG01-treated animals, although this effect was not statistically significant (Supplementary Fig. 9A, B).
A Schematic depicting experimental design of the initial experiment in MPS I mice with Line 6-derived MG01. B Accumulation of GAGs in the brain of adult-treated MPS I cohorts at 4-months post-treatment with Line 6-derived MG01. C Schematic depicting the experimental design of adult (MPS I: P60; MPS VII: P90) and juvenile (P21) MG01 transplantation in MPS I and MPS VII mouse models. D Accumulation of GAGs at 1-, 5- and 8-months post-treatment in the brain, spinal cord (SpC), or CSF of the juvenile-treated MPS I cohorts. MG01 was administered at a low (0.4 × 106) or high dose (1.4 × 106). E Correlation between CSF and brain GAG levels at 5 months post-treatment in the juvenile MPS I cohort. F Accumulation of GAGs at 1- and 5-months post-treatment in the brain, spinal cord, or CSF of the adult-treated MPS I cohorts. G Accumulation of GAGs at 1-month post-treatment in the brain and spinal cord of the juvenile-treated MPS VII cohorts. H Accumulation of GAGs at 1- and 5-months post-treatment in the brain and spinal cord of the adult-treated MPS VII cohorts. GAG levels were measured as μg/ml and are depicted as percentage of GAGs in vehicle controls of MPS I or MPS VII, respectively. Significance was tested with a one-way ANOVA using Dunnet’s multiple comparison test. Graphs denote mean +/− SEM. Data points in B, D, and H represent individual animals. (n.s.: not significant, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). Panels A, and C were in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Further experiments were conducted with Line 82- or 83-derived MG01. After administering MG01 in juvenile animals, we found a dose-dependent reduction of GAGs in CNS tissues at 1-, 5-, and 8-months post-transplantation, when compared to vehicle controls (Fig. 5C, D, Supplementary Table 1). At 5 months, ICV delivery of MG01 reduced GAG levels in the CSF, which positively correlated with GAG levels in the brain (Fig. 5E). In the adult-treated cohort, we found similar GAG reduction across timepoints, albeit with delayed CNS-wide onset. In the low dose group GAG reduction was initially (1-month) only detectable in the CSF but was later (5-month) evident in all CNS tissues, suggesting a dose-dependent treatment response when MG01 is delivered in adults compared to juvenile administration (Fig. 5F).
Consistent with GAG reduction, IDUA enzyme activity was significantly increased compared to vehicle-injected controls at all timepoints in the juvenile (p21) high-dose cohort, although this effect was not statistically significant in the low-dose group at earlier timepoints (Supplementary Fig. 9C, D, Supplementary Table 1). In the adult cohort, considerable IDUA activity was seen in the brain and CSF of animals that received high-dose MG01 but not in the spinal cord, although GAGs were reduced in all CNS tissues. Similarly, in the group that received a low-dose MG01 there was no significant IDUA activity detected (Supplementary Fig. 9E) even though GAG levels were reduced in the 5-month timepoint (Fig. 5F, Supplementary Table 1). Consistent with the in vitro data, GAG levels are indicative of the cumulative effect of enzymatic activity in vivo, while enzyme levels are a mere snapshot in time. We thus primarily focused on GAG readouts to assess the effect of MG01 and used enzyme levels for confirmation, where possible.
To confirm that MG01 enzyme is distributed equally throughout the brain and the whole tissue lysate results reflect correction across the brain, we sampled different brain regions using a small punch biopsy in the 8-month juvenile cohort and tested for GAG levels and IDUA enzyme activity. MG01 treatment had a dose-dependent increase in IDUA activity and a concomitant reduction in GAGs in all brain regions examined in similar proportion to whole brain lysates (Supplementary Fig. 10, Supplementary Table 1). These data further support the suggested mechanism of action of enzyme secretion by MG01 and uptake by diseased cells.
Overall, these data show that MG01 provide functional enzyme in vivo for durable CNS-wide rescue of the abnormal biochemical phenotype in an MPS I mouse model after a single administration of MG01.
MG01 reduces GAG accumulation in the CNS of MPS VII mice
We next sought to test MG01 in a second LSD model. Accordingly, we administered MG01 in an immunodeficient MPS VII model. MG01 administration in juvenile animals achieved similar results to the MPS I model, reducing GAG accumulation in the brain and spinal cord in a dose-dependent manner (Fig. 5G, Supplementary Table 1). In the adult cohort, only high-dose MG01 was tested and although at 1-month post-treatment only a marginal reduction in GAGs was observed, by 5-months MG01-transplanted animals achieved a reduction to levels close to WT animals in both brain and spinal cord (Fig. 5H). The timing in GAG reduction seen in this cohort appears consistent with the delayed enzyme kinetics found in adult delivery in the MPS I model. Overall, GAG reduction in MPS VII mice corroborate long-term capacity of MG01 as a singular treatment to prevent GAG accumulation in the CNS of different MPS models.
ICV administration of MG01 reduces GAG accumulation in the periphery of MPS mouse models
For most peripherally administered LSD therapies, the ability of the enzyme to cross the BBB in sufficient amounts remains a key challenge. Consequently, we explored how enzyme secreted from CNS-delivered MG01 affects the periphery. We first measured urinary GAG levels in MG01-treated animals, a measurement used as a diagnostic tool in the clinic2,36 (Fig. 6A, Supplementary Table 1). MPS I mice that received either low- or high-dose MG01 had reduced GAG levels at 1-month and the reduction was sustained up to 8-months, when compared to vehicle control groups (Fig. 6B). Similarly, in both juvenile and adult animals of the MPS VII model, we observed GAG reduction 2 weeks after transplantation and was sustained up to 5 months (Fig. 6C).
A Schematic depicting the experimental design of adult (MPS I: P60; MPS VII: P90) and juvenile (P21) MG01 transplantation in MPS I and MPS VII mouse models. B Accumulation of GAGs in urine of MPS I mice sampled at regular intervals for the 1-, 5-, or 8-month cohort of the juvenile- or adult-treated MPS I mice. C Accumulation of GAGs in urine of MPS VII mice sampled at regular intervals for the 1- or 5-month cohort of the juvenile- or adult-treated MPS I mice. Urine GAG levels were measured as μg GAGs/ml creatinine and as μg/ml for all other tissues. All GAG measurements are depicted as percentage of GAGs in vehicle controls of MPS I or MPS VII, respectively. Significance was tested using a 2-way ANOVA in a mixed effects analysis followed by Dunnet’s multiple comparison test. Graphs denote mean +/− SEM, N = 3–8 per timepoint in each cohort. (n.s.: not significant, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). Panel A was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Consistent with the urine GAG findings, serum IDUA activity was detectable in MG01-treated animals, and followed the kinetics observed in CNS tissues. In adults, serum IDUA activity increased at 2-weeks in the high-dose regimen but was not detectable in the low-dose group (Supplementary Fig. 11B, Supplementary Table 1). In contrast, the juvenile cohort had sustained increase in IDUA activity in both doses at 2-weeks. Finally, to assess penetrance of the enzyme to peripheral organs, hearts and livers were collected from the 5-month cohort of MPS I mice to evaluate GAG levels. We observed a small reduction compared to vehicle controls in the animals treated with high-dose but not low-dose MG01 in the heart while in the liver we detected a reduction with both doses (Supplementary Fig. 11C). Concordantly, in MPS VII, we detected GAG reduction at 5 months in the liver (Supplementary Fig. 11D).
MG01 prevents ataxia in both MPS mouse models
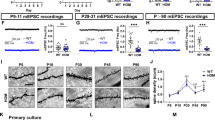
Typical behavioral measurements, especially skeletomuscular, are confounded by the systemic deficiencies found in MPS I and MPS VII models that MG01 is not designed to correct. Our own assessment of novel object recognition and repeated open field could not reproduce the findings from the primary reports37,38 (Supplementary Fig. 12). We detected no difference between wildtype and MPS I, possibly due to mouse background strain differences. Thus, for the behavioral evaluation of the animals, we focused on a readily observable endpoint, the hindlimb clasping assay39, a CNS ataxia measure that is affected in MPS40. Vehicle-treated juvenile MPS I animals had a positive hindlimb clasping reflex upon tail suspension at 5-months and 8-months when compared to WT littermates, indicating detectable ataxia (Fig. 7B). In contrast, juvenile MPS I animals that received either high- or low-dose MG01 had no statistically significant deficiency 5-months post-treatment when compared to wild-type animals and this effect was sustained up to 8 months (Fig. 7B).
A Schematic depicting the experimental design of adult (MPS I: P60; MPS VII: P90) and juvenile (P21) MG01 transplantation in MPS I and MPS VII mouse models. B Hindlimb clasping scores at 5- or 8-months post-treatment in juvenile MPS I mice. C Hindlimb clasping scores at 1-, 2- and 5-months post-treatment in adult MPS VII mice. D Hindlimb clasping scores at 1-month post-treatment in juvenile MPS VII mice. Graphs denote median. Significance was tested using a Kruskal-Wallis test followed by a Conover-Iman post hoc test. Data points in B–D represent individual animals. (n.s.: not significant, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). Panel A was in part created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
In MPS VII, we detected differences in the hindlimb clasping reflex between vehicle and WT groups as early as 1-month post-treatment, but we only saw significant prevention of deficits of the adult-treated animals beginning at 2-months post-treatment (Fig. 7C). We observed durable prevention of this behavioral deficit up to at least 5-months post-treatment in the adult cohort (Fig. 7C). In contrast, in the juvenile MPS VII animals, we observed significant prevention of hindlimb clasping reflex as early as 1-month post-MG01 administration (Fig. 7D). These results are in agreement with the observed reduction in GAGs (Fig. 5G, H) showing an age-dependent effect when treating MPS VII mice and is in accordance with similar observations in clinical trials41,42. These data underscore the importance of early intervention in neuropathic LSDs.
Discussion
ERT as today’s available treatment option for MPS I, MPS II and MPS VII has been transformative for patients, improving their quality of life. However, the burden of treatment with life-long infusions, and the challenges in reversing substrate accumulation in immune-privileged organs such as the brain retain significant unmet medical need5,43. HSCT is a considered therapeutic avenue for patients with severe and rapidly progressive forms of MPS; however, donor availability, the need for early intervention and the extended management of posttransplant life are important considerations. HSCT, comprising of a one-time treatment that provides functional peripheral enzyme production, shows evidence of CNS penetrance, as some cells can infiltrate the brain parenchyma2,7. However, variable CNS penetrance restricts cognitive rescue as it is observed in only a limited number of LSDs, such as MPS I-H6. Plausible factors that could account for this inconsistency are the timing of intervention and the local amount of enzyme required. Even if administered early, HSCT can take up to 12 months for CNS engraftment7,44, therefore critically delaying donor enzyme availability. This is problematic as treatment prior to symptom onset correlates with better prevention of cognitive decline45,46. Our animal data supports this observation, as treatment of juvenile animals with MG01 showed earlier efficacy signals when compared to animals treated as adults. Importantly, MG01 as a ready-to-use product can theoretically be given shortly after diagnosis and can readily provide therapeutic doses of the deficient enzyme throughout the CNS. As rapid, CNS-wide enzyme distribution is critical for the prevention of symptoms, we concluded based on our data that ICV delivery would be the most appropriate route of administration of MG01. This route of administration has been reported as safe in the clinic47,48. Intraparenchymal delivery could be an alternative approach, but it may require multiple injections to reach CNS-wide engraftment.
An unexpected finding in our studies is the small but physiologically relevant presence of donor enzyme and prevention of GAG accumulation in the periphery after delivery of MG01 to the CNS. While systemically delivered enzymes do not readily cross the BBB to gain entrance into the CNS, enzymes in the CSF appear able to cross the CSF-blood barrier and enter peripheral circulation, presumably following the lymphatic drainage of CSF49. Future studies will need to expand on this finding and assess the extent of peripheral rescue in different MPSs under sustained functional enzyme production from the CNS. This will determine if MG01 is sufficient for non-CNS rescue for specific MPSs or if it will require combination therapy with peripherally targeted therapeutics.
MG01 and the co-culturing method developed here allow for fast and reliable initial testing of MG01 as a therapy for its disease correction ability in LSDs and the assay itself can serve as a proxy for patient stratification in aid of the design of the best strategy for future clinical applications.
There are several limitations to this study. We tested MG01’s potential to change CNS disease in MPS I-H, but not against HSCT, the standard treatment for MPS I-H. While a comparison of that nature would be insightful, the aim of this work is to test the universality and immediate availability of MG01. Our data suggests that MG01 aids recovery in neuronopathic MPSs due to its CNS-targeted delivery and rapid engraftment, but future studies will need to test this therapy directly against the SOC of different MPSs, whether it is ERT or HSCT.
Behavioral readouts that rely on animal mobility raised a particular challenge as MPS models have reduced mobility due to musculoskeletal symptoms. The intended purpose of MG01 is to treat neuronopathic MPSs, therefore this mobility phenotype limits the possible behavioral readouts. Additionally, the necessity for an immunocompromised (NOD/scid) background for xenograft studies became a confounding factor, as it is well-known that background strain can affect behavioral readouts50,51,52. In fact, assays reported previously as showing differences in MPS I did not replicate in our model37,38. Future work should attempt additional behavioral readouts after combinatorial therapy or rely on MPS models with less severe peripheral pathology (e.g., MPS III)53.
Another limitation is that the animal models utilized are not the most suitable for myeloid engraftment and survival, given incomplete myeloid cytokine support for human cells via the mouse cytokines. Despite this, the amount of MG01 engraftment allowed for CNS-wide rescue of GAG accumulation and for enzyme recovery, demonstrating a critical proof-of-concept. We expect that in humans MG01 engraftment will be optimally supported across meningeal macrophages and parenchymal microglia31,32,33, which is in contrast to the NOD/scid animal models where the minority of MG01 became parenchyma-engrafted ramified microglia while the majority remained engrafted to the meninges. Taken together, we expect that MG01 will engraft more readily and survive in higher numbers in humans, providing more efficient restoration of the deficient enzyme. Prior to clinical use, suitable engraftment, modeling, and scaling studies will need to be performed to determine an efficacious human clinical dose.
Beyond engraftment, additional questions remain unanswered, and further research will be required to bring an allogeneic cell therapy of this kind to the clinic. For example, immunogenicity is of particular interest in allogeneic therapies. MG01 is not genetically engineered, however, there is evidence of long-term persistence of allogeneic grafts in the brain without rejection54,55 and several clinical trials for allogeneic neuronal cell therapies are ongoing, demonstrating that cells can survive in the CNS with limited immunosuppression56,57,58,59,60. However, cells of the lymphoid lineage may be more immunogenic than neurons and additional studies could test the genetic modification of MG01 conferring immune evasion61. While there is necessary work to fully understand this therapeutic modality, evidence at present points to a potential clinical path forward for cell therapies designed for indications with insufficient standard of care like many of the LSDs.
In summary, we suggest that allogeneic off-the-shelf microglia delivered behind the BBB directly into the CNS could be a one-time treatment option providing durable CNS disease correction in patients suffering from MPSs. Although the standard of care (ERT) with its limited biodistribution to the CNS, and experimental treatments, such as gene therapies62 and protein-brain-shuttles63 are currently in clinical trials, to our knowledge none of these modalities can be used in several MPSs. Here we have shown the possibility of treating several neuropathic MPSs (MPS I/IIIA/IIIB/VII) with a single therapeutic agent and we believe MG01 has the potential to treat many other LSDs. We believe that microglia, as authentic CNS cells, also have the potential for therapeutic application beyond LSDs, where additional functional engineering may occur to create “mini factories” delivering always-on64, inducible65, or environment-sensing payload66 behind the BBB. This could be an advantage, as many of the current therapeutics, such as antibody therapies67 or many small molecules68, are limited in their potential for brain penetrance.
Methods
Cell line generation
Human iPSC line generation and maintenance culture
The hiPSC lines 82 and 83 were derived by reprogramming peripheral blood mononuclear cells (PBMCs) to induced pluripotent stem cells. Peripheral blood was collected by the National Marrow Donor Program and donor informed consent was obtained (IRB protocol number Pro00025408). Erythroblasts were isolated and cultured from PBMCs according to Perriot et al.69. Erythroblasts were electroporated with a synthetic self-replicating Venezuelan equine encephalitis (VEE) RNA replicon as described by Yoshioka and colleagues70. The VEE replicon is a positive-sense, single-stranded RNA that does not utilize a DNA intermediate, so there is no potential for genomic integration and the absence of viral structural genes renders the RNA incapable of viral packaging and pathology. Human iPSC line 6 was obtained from Lonza71, donor informed consent for this line has been examined by NINDS and found to meet the highest standards.
hiPSCs were maintained using Essential 8 medium (Thermo Fisher Scientific, A26559-01) with recombinant laminin (Biolamina, LN521-05) or Vitronectin (Thermo Fisher Scientific, A14700) as the hiPSC attachment substrate. Cell passaging was performed using EDTA as a non-enzymatic method (Thermo Fisher Scientific, AM9260G) or Accutase (Thermo Fisher Scientific, 00-4555-56). Dissociated cells were plated onto Laminin 521 coated tissue culture vessels in Essential 8 medium supplemented with 10 µM of Y-27632 (Bio-Techne, TB1254-GMP) for the first 24 hr. Parental and clonally-derived gene-edited hiPSC lines were screened for normal karyotypes by Cell Line Genetics (Madison, WI). All hiPSC lines were routinely analyzed for appropriate expression of the pluripotency markers NANOG, SOX2, OCT4, SSEA-3, SSEA-4, and TRA-1-60 (BD, 560873, 561506, 560329, 560236, 563119, and 561153 respectively). Pluripotency and karyotype information is found in Supplementary Fig. 13.
hiPSC knockout cell line generation
hiPSC lines lacking IDUA, GUSB, NAGLU, and SGSH were generated to test the ability of MG01 to in vitro cross-correct this enzyme deficiency. CRISPR/Cas9 single guide RNAs (sgRNAs) targeting a coding exon of each of these four genes were designed and tested. The sequences of these sgRNAs are listed in Table 1. A vial of the parental hiPSC line 82 was thawed and transfected with individual Cas9:sgRNA ribonucleoprotein (RNP) complex targeting each of the four genes to generate functional knockouts. The transfected cells were plated onto Vitronectin-coated tissue culture vessels in Essential 8 medium containing Y-27632 (10 µM). Clones were manually isolated by picking and then screened for bi-allelic gene KO using PCR primers flanking the sgRNA target site followed by Sanger sequencing. PCR primers used for genotyping are listed in Table 2. Multiple clones of mono- and bi-allelic KO genotypes were identified for each target gene, but only bi-allelic KO clones were selected for MG01 differentiation. Final genotypes for each KO line are shown under the “Genotype” column in Table 1.
MG01 differentiation
hiPSCs were plated onto Vitronectin (Thermo Scientific, A14700) at 1.0 × 104 cells/cm2 in Essential 8 medium (Thermo Scientific, A1517001) containing 10 µM of Y-27632 (Tocris, 1254) for 24 hours. hiPSCs were cultured for an additional 2 days in Essential 8 medium with daily medium changes before being induced by Essential 6 (Thermo Scientific, A1516401) medium supplemented with 80 ng/mL of BMP-4 (R&D Systems, 314E-GMP-050). BMP-4 induction continued for 4 days with daily medium changes before the cultures were changed to StemPro-34 SFM medium (Thermo Scientific, 10639011) (containing 1X GlutaMAX Thermo Scientific, 35050061) supplemented with 100 ng/mL of SCF (R&D Systems, 255B-GMP-050), 80 ng/mL of VEGF (R&D Systems, 293-GMP-050) and 25 ng/mL of bFGF (R&D Systems, 233-GMP-025) for 2 days with daily medium changes. On day 6 and day 8, the cells were cultured with StemPro-34 SFM medium containing 50 ng/mL of SCF, 50 ng/mL of IL-3 (R&D Systems, 203-GMP-050), 50 ng/mL of M-CSF (R&D Systems, 216-GMP-500), 50 ng/mL of FLT3L (R&D Systems, 308E-GMP-050) and 5 ng/mL of TPO (R&D Systems, 288-TPE-050). Starting on day 10, cells from the supernatant fraction were pelleted, resuspended in the same fresh medium as days 6 and 8, and placed back into their respective vessel. Beginning on day 14, cells in the supernatant fraction were pelleted, resuspended in StemPro-34 SFM medium containing 50 ng/mL of FLT3L, 50 ng/mL of M-CSF and 25 ng/mL of GM-CSF (R&D Systems, 215-GMP-050) and placed back into their respective vessel. Day 14 medium change was repeated every other day until the cells in the supernatant reached a concentration of >1.0 × 106 live cells per mL. Beginning on day 18 and every other day after, cell counts were performed prior to pelleting the cells in the supernatant for medium exchange to determine the harvest date.
Cryopreservation and thawing of MG01
MG01 were isolated and counted on the NucleoCounter NC-200 (Chemometec) prior to freezing. Cells were resuspended in either BamBanker (Wako Chemicals, 30214681)) or STEM-CELLBANKER GMP grade (amsbio, 11924) freezing medium and transferred to cryogenic vials (Thermo Scientific). Cryogenic vials with cells were frozen with a controlled rate freezer (CBS CRF2101). Cells were transferred to liquid nitrogen (vapor phase) for long-term storage. To thaw MG01, cryogenic vials were transferred to a 37 °C water bath for ~2 minutes until a small ice crystal remained, transferred to a centrifuge tube, and quenched with 1 mL of either RPMI-1640 (Thermo Scientific, 11-875-101) or StemPro-34 SFM medium in a drop-wise fashion. The cells were counted on the NC-200 and quenched with an additional 3 mL of its respective medium. Cells were centrifuged at 250 g for 5 minutes and resuspended in the appropriate assay or culture medium.
Replating and polarization of MG01
MG01 were plated at 1.0 × 105 cells per well of a 96-well tissue culture treated plate in 50 µL of RPMI-1640 (Thermo Scientific, 11-875-101) containing 1X GlutaMAX supplemented with 100 ng/mL of IL-34 (R&D Systems, 5265-IL-010/CF) and 10 ng/mL of GM-CSF (maturation medium). For M0-like microglia, an additional 50 µL of maturation medium was added directly on top of the cells. For M1-like microglia, an additional 50 µL of maturation medium supplemented with 200 ng/mL of LPS (Sigma-Aldrich, L4391) and 200 ng/mL of IFNγ (R&D Systems, 285-GMP) was added directly on top of the cells, unless otherwise stated in the figure legend. For extended microglia culture, MG01 was replated as previously described for 6 days or 11 days with media exchanges at 48 hour intervals. Cultures were polarized as previously described 48 hours prior to E.coli phagocytosis challenge. After addition of pHrodo Red E.coli bioparticles (Thermo Fisher Scientific, P35361), wells were imaged every hour for 24 hours. All cell lines tested behaved similarly and the data depicted is representative of all hiPSC-derived MG01.
Cytokine release assay (FRET)
Spent medium collected from 48 hour polarized microglia were centrifuged at 250 g for 5 minutes and the supernatants were frozen at −80 °C. Supernatants were thawed at room temperature and 16 µL were transferred to small-volume 96 well plates for the addition of donor and acceptor antibodies from either the IL-6, TNFα, or CXCL10 kit (Cisbio, 62HIL06PEG, 62HTNFAPEG, 62HCX10PEG). Following the manufacturer’s instructions, samples were incubated for 2–24 hours at room temperature and processed along with standards, and analyzed on a Clariostar microplate reader (BMG Labtech) using the recommended parameters from Cisbio. Data was interpreted by calculating the ratio of the acceptor and donor emission signals and determining the delta ratio (ratio standard or sample–ratio standard 0). Supplementary Fig. 1C does not include CXCL10 data as it could not be collected due to technical failure.
Flow cytometry analysis
MG01 cells were thawed in 2 mL maturation medium as previously described and a cell count was performed on the NucleoCounter NC-200 (Chemometec). Based on the live cell count per mL, 5.0 × 105 cells were transferred to a 5 mL FACS tube (Falcon). Aliquoted samples of cells were prepared and placed into separate tubes for unstained and viability dye controls. Additional maturation medium was added to the cells at a 1:5 dilution to further dilute the cryopreservation medium. The cells were pelleted by centrifugation at 250 g for 5 minutes, the supernatant was carefully removed, and the cells were resuspended in 100 µL of surface marker staining cocktail shown in Table 3. The unstained sample was resuspended in 100 µL of 1X DPBS (Thermo Scientific, 14190250) and viability dye (Thermo Scientific, NC0476349) control was resuspended in 100 µL of 1X DPBS containing 0.33 µL of the viability dye. The samples were incubated at 4 °C for 30 minutes, washed with 2 mL of 1X DPBS, and pelleted by centrifugation at 200 g for 5 minutes. Samples were resuspended in 200 µL of FACS buffer (Table 3) and then analyzed on the CytoFLEX LX (Beckman Coulter). FCS files were exported and analyzed using FlowJo v10.10 software. The representative gating strategy is included in Supplementary Fig. 14.
Phagocytosis of E. coli pHrodo bioparticles
MG01 were plated at 1.0 × 105 cells per well of a 96-well tissue culture plate and then polarized to either M0- or M1-like microglia. After 2 days, medium change was performed by including pHrodo Red E. coli Bioparticles (Thermo Scientific, P35361, prepared according to manufacturer’s instructions) in both M0- and M1-like microglia maturation medium (see polarization section) at a final concentration of 45.45 µg/mL per well. Cells on the plate were placed into the IncuCyte S3 (Sartorius) within a 37 °C incubator with 5% CO2 and imaged every 2 hours for phagocytic activity. The orange signal is detected from the background, partitioned according to edge sensitivity parameters, filtered out if <100 μm2 in the area, and then the average OCU of the object will be multiplied by the area of the orange object. Total integrated intensity is the total sum of the objects’ fluorescent intensity after background subtraction per image and it is calculated at each timepoint on each well then averaged within the condition. One well with M0 medium including pHrodo Red E. coli Bioparticles only (no cells), and a single well with 10 × 103 MG01 with M0 medium alone (no pHrodo) were also imaged as controls. All cell lines tested behaved similarly and the data depicted is representative of all hiPSC-derived MG01.
Immunofluorescent staining for imaging
Cells were fixed with cold 4% paraformaldehyde (Thermo Scientific, AAJ61899AK) for 10 minutes at room temperature (RT). After fixation, cells were washed with 3 rounds of 1X DPBS (Thermo Scientific, 14190250) for 5 minutes at RT before being permeabilized with three washes of 1X DPBS containing 0.2% Triton X-100 (Sigma-Aldrich, T8787) for 10 minutes each at RT. Next, 10% normal donkey serum (Jackson ImmunoResearch, 017-000-121) in 1X DPBS was added to the cells for 1 hour at RT, and then cells were stained with primary antibody cocktail (Table 4) in a blocking solution at 4 °C overnight. Cells underwent 3 rounds of washes with 1X DPBS containing 0.2% Triton X-100 (DPBS-T) for 10 minutes each at RT the next morning prior to adding the secondary antibody cocktail in 1X DPBS at RT for 1 hour. An additional round of three 10-minute DPBS-T washes preceded a 20-minute incubation with DAPI (Invitrogen, D21490) in 1X DPBS at RT. Lastly, cells were washed with 1X DPBS and stored at 4 °C.
Quantification of LAMP1 staining was performed utilizing QuPath 0.4.372 utilizing the Cell Detection workflow using DAPI for nuclei detection, with a cell expansion radius of 10um, and analyzed with Kruskal-Wallis test followed by Dunn’s multiple comparisons test.
In-vitro metabolic cross-correction
Wild-type and enzyme-deficient (MG-KO) microglia were thawed separately in the maturation medium. Wild-type microglia were plated in 6 well tissue culture-treated plates at 1.0 × 106 in 2 mL of maturation medium. In a separate 6-well tissue culture treated plate holding 0.4 µM porous transparent polyester membrane inserts (Falcon, 353090), MG-KO were plated at 5.0 × 105 in 1 mL of maturation medium (2 mL of maturation medium was added to the well below to prevent the membrane from drying out). After 48 hours, MG-KO on inserts were transferred and placed on top of wild-type wells to begin co-culture. Medium change was performed at the start of co-culture and every other day for 10-14 days. Standalone MG-KOs on inserts were cultured alone in parallel, as a diseased control. To measure metabolic cross-correction wild-type and enzyme-deficient microglia were harvested separately from the insert and wells, lysed in 1% Triton X-100, and evaluated for intracellular enzymatic activity and total GAG content. For the conditioned media experiments, thawed MG01 and MG-KO (IDUA) were plated into separate wells of an 8-well chamber slide at 3.0 × 105 in 400 uL of maturation medium. Beginning on day 0, and every other day onwards, till day 10, the spent medium was collected from MG01 samples and replaced with 400uL of fresh maturation medium. MG01 conditioned medium was prepared by centrifuging the MG01 spent medium at 250 x g to remove any cellular debris. To condition MG-KO (IDUA−/−), MG-KO spent medium was replaced with 400 uL of the MG01 conditioned medium (supernatant) containing 100 ng/mL of IL-34 and 10 ng/mL of GM-CSF during each medium exchange, for a duration of 10 days.
Neuronal differentiation and co-culture
Human iPSCs were differentiated into cortical neuronal progenitors as reported by Ciceri et al.73 On day 20 neural progenitor cells (NPCs) were dissociated to single cells using Accutase and were resuspended in STEM-CELLBANKER (Amsbio, 11924) freezing medium and transferred to cryogenic vials. Cryogenic vials with cells were frozen with a controlled rate freezer (CBS CRF2101). Cells were transferred to liquid nitrogen (vapor phase) for long-term storage. 5.0 × 105 NPCs were plated into separate 6-well tissue culture-treated wells holding 0.4 µM porous transparent polyester membrane inserts (Falcon, 353090). Plated NPCs were cultured in Neurobasal complete media supplemented 20 ng/mL of BDNF (R&D Systems,11166-BD-050), 10 ng/mL of GDNF (R&D Systems, 212-GD-050/CF), 0.5 mM of cAMP (Sigma, D0627), 0.2 mM ascorbic acid (Sigma, A4403) and 10uM DAPT (2634, Tocris) for 10 days with media exchanges every 3 days. Wild-type microglia were plated in 6 well tissue culture-treated plates at 1.0 ×106 in 2 mL of maturation medium. Day 30 NPCs in transwells were transitioned to BrainPhys media (Stem Cell Technologies, 05790) containing N2, BDNF, GDNF, asorbic acid, IL-34, and GM-CSF and placed on top of wild-type microglia wells. Medium change was performed at the start of co-culture and every other day for 10 days. Standalone NPC-KOs on inserts were cultured alone in parallel, as controls.
Dose formulation for animal transplantation
MG01 were thawed and transferred to a centrifuge tube. Before performing a cell count on the NucleoCounter (Chemometec), 1 mL of StemPro-34 SFM medium supplemented with 50 ng/mL of FLT3L, 50 ng/mL of M-CSF, and 25 ng/mL of GM-CSF (SP34-d14) was added directly on top in a drop-wise fashion. An additional 3 mL of SP34-d14 medium was added on top to further dilute the freezing medium. Cells were centrifuged at 250 g for 5 minutes at room temperature and the supernatant was aspirated. The cell pellet volume was determined using a 20–200 µL single channel pipette and transferred to a 1.5 mL microcentrifuge tube. Based on the live cell count per mL, the cell pellet was diluted with additional StemPro-34 SFM-d14 medium for a final formulation of 7 × 104 (high-dose) or 2 × 104 (low-dose) live cells per µL. The prepared cells were placed on ice until ready for transplantation.
Enzymatic activity assay
To determine enzyme activity, cells from the membrane insert were lysed in 1% Triton-X 100 containing protease inhibitors followed by assessment of enzyme activity and GAG levels. The method was adapted from Ou et al., 201474. Enzyme catalytic activity for each lysosomal enzyme was determined by quantifying 4-Methylumbelliferone (4-MU), the fluorescent moiety produced after cleavage of an artificial fluorogenic substrate that is unique to each enzyme (Table 5). The resulting fluorescence of cleaved substrate was read on the CLARIOstar (MARS software v3.31) with an excitation setting at 355 nm and emission at 460 nm. Enzyme levels and activity were interpolated using a 4-MU standard curve. For animal experiments, frozen tissue was homogenized using 1% Triton X-100 in HBSS (Cytiva, SH30588.01) with Halt protease inhibitor (Thermo Scientific, 87786). A BCA protein assay was performed on the resulting lysates to estimate the amount of protein isolated. Up to 50 µg of total protein was used to measure enzyme activity and GAG levels from brain lysates, spinal cord lysates, and serum. For CSF, 1 µl of the sample was used for the assay and results were post-normalized to total protein concentration. Enzyme activity in urine was measured using 5 μL upon thaw. In parallel, creatinine levels were quantified to normalize urine enzymatic activity. Values before normalization to vehicle or wildtype controls are included in Supplementary Table 1.
Enzymatic activity for Line 6 after engraftment studies was assayed with a first generation of the IDUA enzyme activity assay, which needed to be further optimized. Prior to optimization, the strong signal from the fluorogenic compound used to measure IDUA activity showed some background, especially at higher enzyme concentrations. We thus re-optimized the assay to measure the lower enzyme levels expected in our grafted animals. As such, a direct comparison between Line 6 and Lines 82-83 is not possible.
Glycosaminoglycan (GAG) comprised of; dermatan sulfate (DS), heparan sulfate (HS), keratan sulfate (KS), chondroitin sulfate (CS), hyaluronic acid (HA).
GAG quantification
Frozen tissue was homogenized as described above (refer to Enzymatic Activity Assay section). Quantification of GAGs from the various cell lysates was performed using a cationic dye 1,9 dimethylmethelene blue (DMB) (Sigma Aldrich, 341088-1G) which binds to highly charged sulfated GAGs. The resulting absorbance at 525 nm is proportional to the concentration of GAGs in the sample. Values were interpolated using a GAG standard curve. GAG levels were measured using 5 μL of urine upon thaw. In parallel, creatinine levels were quantified to normalize urine GAGs. Values before normalization to vehicle or wildtype controls are included in Supplementary Table 1.
SGSH automated western
Samples were homogenized as described above (refer to Enzymatic Activity Assay section), and a BCA protein assay was performed to estimate the total protein concentration. 5ug of total protein (or a maximum of 4uL) was loaded into each Jess Automated Western reaction. A standard curve was also generated using recombinant human SGSH (Bio-Techne, 8380-SU). A 12-230 kDa separation cartridge was loaded with samples, standards, anti-SGSH antibody (20ug/mL, Bio-Techne MAB83801), and anti-mouse HRP secondary (Bio-Techne 042-205). The instrument was run according to manufacturer instructions, and the resulting peak heights that corresponded to the molecular weight of SGSH (in the electropherogram) were used to generate a standard curve and interpolate results.
LAMP1 and LAMP2 quantification
To study modulation of LAMP1 and LAMP2 levels, MG-KO, MG01, and cortical neurons were homogenized as described above (refer to Enzymatic Activity Assay section), and a BCA protein assay was performed to estimate the total protein concentration. 0.5 µg of total protein was diluted in the sample diluents provided by each kit, and LAMP1 (Abcam, ab302755) and LAMP2 (Abcam, ab302757) ELISAs were performed according to manufacturer protocols.
Animals and husbandry
Experiments were performed with male and female mice (P21 and P60-90 old; n = 6–8/age group/genotype) with comparative groups (NOD.Cg-Prkdcscid Iduatm1Clk/J [Idua −/−] versus their wild-type [Idua+/+] littermates) (Strain #:004083, Jackson Laboratory) and NOD.Cg-Prkdcscid Gusbmps/SndsJ [Gusb−/−] versus their wild-type [Gusb+/+] littermates (Jackson Laboratory). For engraftment studies, male and female mice NSG-Q (NOD.Cg-PrkdcscidIl2rgtm1WjlTg(CMV-IL3,CSF2,KITLG)1Eav/MloySzj; Strain #013062, Jackson Laboratory, crossed with NOD.Cg-PrkdcscidIl2rgtm1WjlTg(CSF1)3Sz/SzJ; Strain #028654, Jackson Laboratory; P7, P14 and P21; n = 1-2/age group/sex) were used. Animals were maintained on a 12 h light-dark cycle (lights on at 6:00 am) at 22–25 °C with ad libitum access to food and water. Mice were housed 2–5 per cage. All experiments were conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee under protocol number 2021-BRT-8.
Stereotaxic surgery
Briefly, 30 minutes prior to surgery, adult animals (P21 and older) were injected subcutaneously with 4 mg/kg Meloxicam SR (Putney, RXMELOXICAM-INJ), and all animals were anaesthetized by inhalation of ~2% isoflurane (Attane, RXISO-250) in oxygen throughout the entire surgical procedure. Anaesthetized animals were positioned in a stereotaxic frame (Kopf instruments) and 10 µL of either MG01 low dose (0.4×106 cells/brain), MG01 high dose (1.4 × 106 cells/brain), or vehicle (StemPro-34 SFM medium containing 50 ng/mL of FLT3L, 50 ng/mL of M-CSF, and 25 ng/mL of GM-CSF; 0 cells/brain) were injected bilaterally with a blunt, 27-gauge needle attached to a 10 µL Model 701 RN Hamilton syringe into the lateral ventricles using the following coordinates from bregma:
-
i.
P60 & P90 mice: AP −0.3, ML+/− 1.4, DV 2.34 at a 10° angle
-
ii.
P14 & P21 mice: AP + 0.1, ML+/− 1.4, DV −2.34 at a 10° angle
-
iii.
P7 mice: AP + 0.1, ML+/− 1.4, DV −2 at a 10° angle
Hindlimb clasping Test
Behavioral testing was performed at 1, 5, and/or 8 months post-surgery, depending on the cohort. Briefly, animals were removed from cage and suspended by their tail for ~10-15 seconds and their abilities of hindlimb clasping were monitored via video recording. Hindlimb clasping was scored on a scale from 0-3 based on the following:
-
i.
Score 0: Both hindlimbs were splayed outward and away from the abdomen
-
ii.
Score 1: One hindlimb is retracted toward the abdomen for more than 50% of the time
-
iii.
Score 2: Both hindlimbs are partially retracted toward the abdomen for more than 50% of the time
-
iv.
Score 3: Both hindlimbs are entirely retracted and touching the abdomen for more than 50% of the time
Tissue collection and analysis
For in life analysis, blood was collected via submandibular bleed (50 ul) one day prior to surgery. Post surgery, blood (collected via submandibular bleed; 50 ul) and urine (collected via rubbing abdomen above bladder; 20 ul) samples were collected 2 weeks and then every month before termination. Terminal tissue collection was performed 1-, 5-, or 8-months post MG01 transplantation, depending on the cohort. Briefly, animals were anaesthetized by inhalation of ~2% isoflurane in oxygen and a small incision was made along the back of the neck to expose the muscle tissue. Next, muscle tissue was carefully removed to expose the cisterna magna and a 20 µm glass pulled pipette was inserted into the cisterna magna to collect ~3–8 µl of cerebrospinal fluid (CSF). Following CSF collection, blood was collected via intracardial puncture, urine was extracted from the bladder and then animals were perfused transcardially with cold phosphate buffered saline (PBS). Brains were dissected into two hemispheres and the left hemisphere was placed in cold 4% paraformaldehyde (PFA) for 48 h for histological analysis and the right hemisphere was flash frozen for biochemical analysis. One spinal column was collected from each group and placed in 4% PFA for 48 h for histological analysis and the remaining spinal cords were extruded via PBS flush and flash frozen for biochemical analysis. For serum extraction, blood was allowed to coagulate at RT for 1 hr and then centrifuged at 1300 g for 15 min at 4 °C. Clear supernatant, containing serum, was carefully removed to a new tube and flash frozen for biochemical analysis. For brain region specificity of enzymatic activity and substrate accumulation, the right hemisphere was placed into a brain block and cut at 1 mm thick sections coronally. Each region was removed with a 2 mm tissue puncher, and 2 punches per region were pooled for biochemical analysis.
Histology and immunofluorescence
Harvested tissues for histological evaluation were fixed in 4% PFA for 48 h. Left brain hemispheres were transferred into 20% sucrose (MP Biomedicals, 0219474705) in PBS for 24 h and then into 30% sucrose in PBS for 24 h. Brains were then embedded in OCT (Sakura, 4583)/30% sucrose solution (1:1) and snap-frozen in an ethanol bath, over dry ice. 20 µm thick sections were then obtained in a sagittal orientation using a Cryostat (LEICA; CM3050 S) and mounted onto positively charged microscope slides (Electron Microscopy Sciences, 71873-02). The slides were allowed to air dry for ~1 h and then stored at −80 °C. Spinal columns were washed in PBS and then the spinal cord was dissected from the vertebral column. Extracted spinal cords followed the same sucrose gradient submersion as described above for brain hemispheres. Following the last sucrose solution, spinal cords were dissected into cervical, thoracic, and lumbar regions, embedded in OCT solution, and stored at −80 °C. Spinal cords were sectioned into serial 20 µm slices on the Cryostat, mounted onto positively charged slides, and stored at −80 °C.
For immunofluorescence staining, slides were incubated at RT for 30 m, permeabilized with 0.2% Triton-X in PBS (PBS-T), blocked for 1 h with 10% donkey serum in PBS-T and incubated with primary antibodies (Table 4) in blocking solution overnight at 4 oC. The next day, slides were washed with PBS-T 3 times and incubated with secondary antibodies (Table 4) for 1 h at RT. Slides were then washed 3 times with PBS-T, incubated with 300 nM DAPI in PBS for 30 m, and mounted with Fluorsave™ Reagent (MilliporeSigma, 345789) for imaging. Slides were imaged in an Axioscan 7 microscope (Zeiss).
For DAB, sections were processed the same way as above with the addition of heat-induced antigen (epitope) retrieval (HIER) step for 30 minutes at 95 °C in 1x Citrate Buffer (Abcam, ab64214), permeabilization using 0.8% PBS-T, and utilizing a biotinylated secondary followed by PBS-T washing steps and streptavidin-DAB incubation for 30 m. After, DAB was developed utilizing a DAB peroxidase substrate kit (Fisher Scientific, SK-4100). Slides were imaged in an Axioscan 7 microscope (Zeiss).
hNA+ cell count statistical analysis
Quantification of MG01 was performed by cryosectioning of formalin fixed brain hemispheres followed by immunohistochemistry for the human marker hNA or ku80. Sagittal cryosections 200-400um apart spanning the whole brain of every animal were selected to check for hNA immunoreactivity after manual assessment of image and section quality. Sections were imaged using a Zeiss Axioscan 7 Slide Scanner and hNA counts were performed using an automated workflow based on pixel classification using the Intellesis platform in Zen Blue 3.3 (Zeiss).
Ki67+ colocalization analysis
Quantification of ki67+/ku80+ cells was done manually using the Cell Counter plugin in ImageJ or Fiji75. Three comparable sections per animal were selected to check for ki67/ku80 immunoreactivity after manual assessment of image and section quality. All ku80 cells detected were counted and the proportion of ki67-positive cells in that population was calculated. All data points are the average of the percentage per section per animal.
Statistical analysis
All statistical analyses were performed using Prism 9.4.1 (GraphPad Software LLC). In all graphs: *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. For all IDUA levels, GAG quantification, and behavior, data points represent biological replicates.
For enzymatic activity and GAG readouts, data was tested for normality using the Shapiro-Wilk normality test. If normally distributed, it was analyzed using Prism with a one-way ANOVA using Dunnet’s multiple comparison test, comparing all groups to the vehicle group. If not normally distributed, it was analyzed with Kruskal-Wallis (KW) test followed by Dunn’s multiple comparisons test. Data is represented as mean and SEM.
For Fig. 5, there was a significant effect of treatment on GAG levels in the 4-month MPS I adult cohort treated with Line 6 (F2,8 = 96.76, p < 0.0001). Similarly, a treatment effect on GAG levels for the juvenile MPS I cohort brain lysates at 1 M (F (3, 21) = 1107, P < 0.0001), 5 M (F (3, 16) = 270.3, p < 0.0001) and 8 M (KW: H = 16.02, p < 0.001); in spinal cords at 1 M (F (3, 13) = 306.5, P < 0.0001), 5 M (F (3, 12) = 151.8, p < 0.0001), 8 M (F(3, 11) = 26.63, p < 0.0001); and CSF at 1 M (F (3, 21) = 179.5, P < 0.0001) and 5 M (F (3, 16) = 304.7, p < 0.0001). There was a treatment effect on GAG levels for the Adult MPS I cohort brain lysates at 1 M (F (3, 20) = 182.6, p < 0.0001) and 5 M (F (3, 18) = 140.3, p < 0.0001); in spinal cord at 1 M (F (3, 20) = 182.6, p < 0.0001) and 5 M (F (3, 14) = 72.50, p < 0.0001); and CSF at 1 M (F (3, 20) = 41.89, p < 0.0001). In the MPS VII model, there was a significant effect on adult cohort brain lysates at 1 M (KW: H = 13.66, p < 0.0001) and 5 M (F (2, 23) = 252.4, p < 0.0001), spinal cord at 1 M (F (2, 15) = 204.2, p < 0.0001) and 5 M (F (3, 20) = 76.32, p < 0.0001). In the juvenile cohort, there was a significant effect at 1 M on brain lysates (F (3, 21) = 76.21, p < 0.0001) and spinal cords (F (3, 17) = 31.38, p < 0.0001). p values from post-hoc tests when compared to the vehicle group are displayed on the corresponding figure.
For Supplementary Fig. 9, there was a significant effect of treatment on IDUA levels in the juvenile MPS I cohort brain lysates at 1 M (F (3, 21) = 559.7, p < 0.0001), 5 M (KW: H = 17.86, p = 0.0005)) and 8 M(F (3, 15) = 764.9, p < 0.0001); spinal cords at 1 M (F (3, 21) = 559.7, p < 0.0001), 5 M (F (3, 12) = 3898, p < 0.0001) and 8 M (F (3, 11) = 1569, p < 0.0001); and CSF at 1 M (F (3, 21) = 179.5, p < 0.0001), 5 M (F (3, 16) = 575.9, p < 0.0001) and 8 M (F (3, 14) = 116.4, p < 0.0001). In the adult MPS I cohort, there was a significant effect in brain lysates at 1 M (F (3, 20) = 7921, p < 0.0001) and 5 M (F (3, 18) = 2256, p < 0.0001), and in CSF at 1 M (KW: H = 20.42, p = 0.0001) and 5 M (F (3, 16) = 304.7, p < 0.0001). p values from post-hoc tests when compared to the vehicle group are displayed on the corresponding figure.
For hindlimb clasping, a minimum of 5 animals per group were used across all studies. As the scoring from hindlimb clasping is not normally distributed, data were analyzed using R and the nonparametric Kruskal-Wallis test and Conover-Iman post hoc tests were used if significant main effects were observed via Kruskal-Wallis. All data are represented as median.
For Fig. 7, there was a significant effect of treatment in hindlimb clasping score in MPS I animals at 5 M (chi-squared = 9.7119, df = 3, p = 0.02) and 8 M (chi-squared = 8.8107, df = 3, p = 0.03) post-treatment. In MPS VII animals, there was a significant effect on treatment in hindlimb clasping score in the adult cohort at 1 M (chi-squared = 13.6137, df = 2, p = 0), 2 M (chi-squared = 22.6845, df=2, p = 0), and 5 M (chi-squared = 16.733, df = 2, p = 0) post-treatment; and in the juvenile cohort at 1 M (chi-squared=8.0653, df = 2, p = 0.02) post-treatment. p values from post-hoc tests are displayed on the corresponding figure.
For urine and serum samples, data was analyzed using Prism using a 2-way ANOVA in a mixed-effects analysis followed by Dunnet’s multiple comparison test, comparing all groups to the vehicle group. Data is represented as mean and SEM.
For Fig. 6, a statistically significant interaction between the effects on time and treatment on GAG levels in urine of the adult MPS I 1 M (F (3, 20) = 396.5, p < 0.0001) and 5 M (F (12, 75) = 274.4, p < 0.0001) cohorts; in the MPS I juvenile 1 M (F (3, 20) = 85.57, p < 0.0001), 5 M (F (12, 47) = 29.64, p < 0.0001) and 8 M (F (18, 71) = 15.89, p < 0.0001) cohort; in the MPS VII adult 1 M (F (4, 36) = 21.98, p < 0.0001) and 5 M (F (12, 107) = 6.824, p < 0.0001) cohort. In the MPS VII juvenile 1 M cohort there was a significant effect of treatment (F (3, 32) = 176.1, p < 0.0001) on GAG levels in urine. p values from post-hoc tests are displayed on the corresponding figure.
For Supplementary Fig. 11, there was a significant effect of treatment on IDUA levels of the MPS I adult cohort at 1 M (F (3, 20) = 932.3, p < 0.0001). There a statistically significant interaction between the effects on time and treatment on IDUA levels in serum of the adult MPS I 5 M (F (12, 86) = 10.84, p < 0.0001) cohort; and in the juvenile 1 M (F (6, 42) = 16.09, p < 0.0001), 5 M (F (18, 88) = 49.82, p < 0.0001) and 8 M (F (21, 105) = 38.64, p < 0.0001) cohorts. There was a significant effect of treatment on GAG levels in the heart (F (3, 16) = 964.9, p < 0.0001) and liver (F (3, 16) = 1104, p < 0.0001) of the 5 M MPS I cohort and in the liver of the 5 M MPS VII cohort liver (F (2, 16) = 24.18, p < 0.0001). p values from post-hoc tests are displayed on the corresponding figure.
Digital droplet polymerase chain reaction (ddPCR)
ddPCR assay and analysis was conducted at a contract research organization (Charles River, SF). Briefly, the ddPCR assay was a singleplex assay designed to detect hTERT DNA, consisting of oligonucleotide primers and probe mix containing a TaqMan™ hTERT Control Assay (Thermo Fisher) designed to amplify the hTERT sequence in human gDNA. 18 mice (9 males and 9 females) were dosed via bilateral intracerebroventricular (ICV) injection with MG01 or formulation vehicle. The samples were collected 1 month after dosing from mouse whole blood, plasma, brain, spinal cord, liver, spleen, lungs, and kidney fluids and tissues and DNA was isolated. Prepared dosed samples from shedding matrices and biodistribution tissues were analyzed via ddPCR in duplicate. The fluid and tissue samples were analyzed in a BioRad QX ONE ddPCR at a final concentration of 50 ng/rxn of gDNA. All ddPCR analysis contained 2 sets of controls, a no template control, and a negative control.
Bulk RNA sequencing
RNA was isolated from micro-dissected fixed histology sections using the RNeasy FFPE kit (Qiagen) and analyzed on an Agilent 2100 Bioanalyzer using Agilent 6000 RNA Nano Kit (Agilent Technologies). Samples included in analysis are post-thaw MG01 (Line 6) and isolated tissue from an NSG-Q mouse 5.5 months after intracranial transplantation with Line 6-derived MG01 (Coordinates: 7° angle- ML + 2, AP −2.2, DV −2 > −1.8, −0.95). After RNA extraction, all samples were processed in the same batch for library preparation, sequencing, and analysis to control for batch effects. For each sample, 3 cDNA libraries for sequencing were generated per sample using the Collibri 3’ mRNA Library Prep Kit for Illumina (ThermoFisher). Libraries were quantified using the Collibri Library Quantification Kit (ThermoFisher) and evaluated on the Agilent 2100 Bioanalyzer using the Agilent High Sensitivity DNA Kit (Agilent Technologies). Libraries were sequenced on the Illumina NextSeq 500 using the NextSeq 500/550 High Output Kit (75 Cycles). Reads were filtered and trimmed using Cutadapt v3.476 and aligned with STAR v2.7.8a77 to both hg38 (human reference genome) and mm39 (mouse reference genome). Aligned reads were sorted and indexed using SAMtools v1.1278 and filtered to detect only human-specific reads using XenofilteR79. Reads aligning to expressed genes were counted using HTSeq v0.13.580. Counts were CPM normalized and scaled per gene, prior to plotting. Heatmaps were generated using ggplot2v3.5.0.
Single-cell RNA sequencing and analysis
Fetal hematopoietic (CD45+) single-cell data from Bian et al.17 was downloaded from NCBI Gene Expression Omnibus, accession number GSE133345. For downstream analysis, primitive macrophage subpopulations (Mac 1, Mac 2, and Mac 3) were grouped together. UMAP plots were generated and colored by origin, Carnegie stage, and cell type grouping of all cells using Scanpy81.
Single-cell RNA sequencing was performed on MG01 cells (Line 6) (n = 3 independent biological replicates) and hiPSCs using 10X Genomics. The raw data for each dataset was the “Filtered Feature Barcode Matrix” output by Cell Ranger82 alignment and deconvolution pipeline. Each dataset was imported and mitochondrial content, number of genes by counts, and total counts were annotated for each cell. A knee locator-based method83 was used to remove cells that have an overrepresentation of mitochondrial counts. The data was then filtered to include only cells that have a minimum of 1000 non-zero transcripts. Double detection and removal were performed and the dataset subset was to only include 5000 cells (so that each dataset has the same number of cells in the analysis). Each dataset was iteratively joined to the other datasets included in the analysis. Joining is performed on Ensembl IDs on an inner-join basis (only genes common to both datasets are included in the resulting data object). MG01 and hiPSC data were then joined to Bien et al. data on Ensembl IDs. Finally, the total counts per cell was normalized to 1,000,000 (CPM normalization), and log2 transformed.
PCA was run on the concatenated dataset using highly variable genes. Differential gene expression was run using Scanpy’s rank_genes_groups function and filtered based on a minimum log fold change = 4 and a threshold fraction of cells expressing the gene within and outside of the group = 0.5. Genes enriched in the Mac 1-3 groups and Mac 4 group were used for sparsity-based scoring. To score, the normalized expression matrix was binarized and the sparsity of the matrix was calculated. The matrix was subsetted to include the genes in the group of interest (Mac 1-3 or Mac 4). The gene set score for each cell was calculated as the sum of the binarized expression data for the gene group multiplied by the sparsity, normalized by the maximum score so that the value ranged from 0 to 1.
Differential gene expression was run on the Bian et al. dataset17 (with the same settings described previously) based on the cell type groupings of all cells to find unique markers for each group. Expression data from this dataset was standardized between 0 and 1 and shown as a heatmap. Unique genes found in the Mac 4 group and Mac 1-3 groups were displayed as dotplots using the single-cell RNA sequencing datasets and as a heatmap using engrafted MG01 bulk RNAseq expression data.
Software information
Images for brain sections acquired with Axioscan utilizing Zen 3.1. GAG levels, enzymatic activity, and LAMP1/2 assays were measured in the ClarioSTAR platform using MARS software v3.31. FACS data obtained in CytoFLEX LX using CytExpert v2.5 - only flow cytometry analysis performed (no cell sorting occured). Phagocytosis data was acquired and analyzed using the Incucyte platform v2023.1.1. Bulk RNAseq reads were filtered and trimmed using Cutadapt v3.4 and aligned with STAR v2.7.8a to both hg38 and mm39. Aligned reads were sorted and indexed using SAMtools v1.12 and filtered to detect only human-specific reads using XenofilteR (v1.6). Reads aligning to expressed genes were counted using HTSeq v0.13.5. scRNAseq raw data for each dataset was the “Filtered Feature Barcode Matrix” output by Cell Ranger v3.0.2 alignment and deconvolution pipeline UMAP plots were generated colored by origin, Carnegie stage, and cell type grouping of all cells using Scanpy 1.8.1. Biochemical and behavior graphs and statistical tests were performed in Prism v10.1.0. Pixel classification and cell counts performed with Zen 3.3 with Intellesis module. LAMP1 intensity per cell was measured using QuPath 0.4.3. Cell viability was measured with the Nucleocounter platform using Nucleoview v1.4.3
Miscellaneous software
All figures were created and assembled in Affinity Designer 1.10. Graphs and statistical tests performed in Prism v10.1.0.
Material availability
Cell lines #82 and #83 are not available for sharing because they are proprietary and reserved for potential clinical use. All critical experiments have been conducted with cell line #6, which was obtained via NHCDR (https://stemcells.nindsgenetics.org/). We generated additional IDUA, GUSB, NAGLU, and SGSH knockout iPSC cell lines based on parental cell line #6 which are available, subject to prior written authorization from the NHCDR and a material transfer agreement.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Source data are provided with this paper. Bulk and single cell RNAseq data are available through the GEO SuperSeries accession number GSE224139, including accession series GSE224136 and GSE222969. Source data are provided with this paper.
References
Rajkumar, V. & Dumpa, V. Lysosomal Storage Disease. in StatPearls (StatPearls Publishing, Treasure Island (FL), 2022).
Shapiro, E. G. & Eisengart, J. B. The natural history of neurocognition in MPS disorders: A review. Mol. Genet Metab. 133, 8–34 (2021).
Valle, D. J., Antonarakis, S., Ballabio, A., Beaudet, A. L. & Mitchell, G. A. The Online Metabolic and Molecular Bases of Inherited Disease | OMMBID | McGraw Hill Medical. https://ommbid.mhmedical.com/book.aspx?bookID=2709#225069487.
Jb, E. et al. Long-term cognitive and somatic outcomes of enzyme replacement therapy in untransplanted Hurler syndrome. Molecular genetics and metabolism reports 13, 64–68 (2017).
Clarke, L. A. Mucopolysaccharidosis Type I. in GeneReviews® (eds. Adam, M. P. et al.) (University of Washington, Seattle, Seattle (WA), 1993).
Edelmann, M. J. & Maegawa, G. H. B. CNS-targeting therapies for lysosomal storage diseases: current advances and challenges. Front Mol. Biosci. 7, 559804 (2020).
Boelens, J. J. & van Hasselt, P. M. Neurodevelopmental outcome after hematopoietic cell transplantation in inborn errors of metabolism: current considerations and future perspectives. Neuropediatrics 47, 285–292 (2016).
Gragert, L. et al. HLA match likelihoods for hematopoietic stem-cell grafts in the U.S. Registry. N. Engl. J. Med. 371, 339–348 (2014).
Fratantoni, J. C., Hall, C. W. & Neufeld, E. F. Hurler and Hunter syndromes: mutual correction of the defect in cultured fibroblasts. Science 162, 570–572 (1968).
Ginhoux, F., Lim, S., Hoeffel, G., Low, D. & Huber, T. Origin and differentiation of microglia. Front. Cell. Neurosci. 7, 1662–5102 (2013).
Muffat, J. et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nat. Med. 22, 1358–1367 (2016).
McQuade, A. et al. Development and validation of a simplified method to generate human microglia from pluripotent stem cells. Mol. Neurodegen. 13, 67 (2018).
Pandya, H. et al. Differentiation of human and murine induced pluripotent stem cells to microglia-like cells. Nat. Neurosci. 20, 753–759 (2017).
Douvaras, P. et al. Directed differentiation of human pluripotent stem cells to microglia. Stem Cell Rep. 8, 1516–1524 (2017).
Abud, E. M. et al. iPSC-derived human microglia-like cells to study neurological diseases. Neuron 94, 278–293.e9 (2017).
Rustenhoven, J. et al. Isolation of highly enriched primary human microglia for functional studies. Sci. Rep. 6, 19371 (2016).
Bian, Z. et al. Deciphering human macrophage development at single-cell resolution. Nature 582, 571–576 (2020).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Svoboda, D. S. et al. Human iPSC-derived microglia assume a primary microglia-like state after transplantation into the neonatal mouse brain. Proc. Natl Acad. Sci. USA 201913541 https://doi.org/10.1073/pnas.1913541116 (2019).
Matalon, R. & Dorfman, A. Sanfilippo A Syndrome sulfamidase deficiency in cultured skin fibroblasts and liver. J. Clin. Invest. 54, 907–912 (1974).
Brot, F. E., Glaser, J. H., Roozen, K. J., Sly, W. S. & Stahl, P. D. In vitro correction of deficient human fibroblasts by beta-glucuronidase from different human sources. Biochem. Biophys. Res. Commun. 57, 1–8 (1974).
Di Natale, P. et al. In vitro gene therapy of mucopolysaccharidosis type I by lentiviral vectors. Eur. J. Biochem. 269, 2764–2771 (2002).
Eskelinen, E.-L. Roles of LAMP-1 and LAMP-2 in lysosome biogenesis and autophagy. Mol. Asp. Med. 27, 495–502 (2006).
Cao, M., Luo, X., Wu, K. & He, X. Targeting lysosomes in human disease: from basic research to clinical applications. Sig Transduct. Target Ther. 6, 1–28 (2021).
Coutinho, M. F., Prata, M. J. & Alves, S. Mannose-6-phosphate pathway: a review on its role in lysosomal function and dysfunction. Mol. Genet Metab. 105, 542–550 (2012).
Coutinho, M. F., Prata, M. J. & Alves, S. A shortcut to the lysosome: The mannose-6-phosphate-independent pathway. Mol. Genet. Metab. 107, 257–266 (2012).
Hasselmann, J. et al. Development of a chimeric model to study and manipulate human microglia in vivo. Neuron S0896627319306002 https://doi.org/10.1016/j.neuron.2019.07.002 (2019).
Cohen-Pfeffer, J. L. et al. Intracerebroventricular delivery as a safe, long-term route of drug administration. Pediatr. Neurol. 67, 23–35 (2017).
Utz, S. G. et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 181, 557–573.e18 (2020).
Shultz, L. D. et al. Multiple defects in innate and adaptive immunologic function in NOD/LtSz-scid mice. J. Immunol. 154, 180–191 (1995).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395.e6 (2018).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Brioschi, S., Zhou, Y. & Colonna, M. Brain Parenchymal and Extraparenchymal macrophages in development, homeostasis, and disease. J. Immunol. 204, 294–305 (2020).
Xu, R. et al. Human iPSC-derived mature microglia retain their identity and functionally integrate in the chimeric mouse brain. Nat. Commun. 11, 1577 (2020).
Semple, B. D., Blomgren, K., Gimlin, K., Ferriero, D. M. & Noble-Haeusslein, L. J. Brain development in rodents and humans: Identifying benchmarks of maturation and vulnerability to injury across species. Prog. Neurobiol. 106–107, 1–16 (2013).
Cadaoas, J. et al. Vestronidase alfa: Recombinant human β-glucuronidase as an enzyme replacement therapy for MPS VII. Mol. Genet Metab. 130, 65–76 (2020).
Pan, D., Sciascia, A., Vorhees, C. V. & Williams, M. T. Progression of multiple behavioral deficits with various age of onset in a murine model of Hurler Syndrome. Brain Res. 1188, 241–253 (2008).
El-Amouri, S. S., Dai, M., Han, J.-F., Brady, R. O. & Pan, D. Normalization and improvement of CNS deficits in mice with hurler syndrome after long-term peripheral delivery of BBB-targeted Iduronidase. Mol. Ther. 22, 2028–2037 (2014).
Guyenet, S. J. et al. A simple composite phenotype scoring system for evaluating mouse models of cerebellar Ataxia. J. Vis. Exp. 1787 https://doi.org/10.3791/1787 (2010).
Ebrahimi‐Fakhari, D. et al. The spectrum of movement disorders in childhood‐onset lysosomal storage diseases. Mov. Disord. Clin. Pr. 5, 149–155 (2018).
Aldenhoven, M. et al. Long-term outcome of Hurler syndrome patients after hematopoietic cell transplantation: an international multicenter study. Blood 125, 2164–2172 (2015).
Muenzer, J. Early initiation of enzyme replacement therapy for the mucopolysaccharidoses. Mol. Genet. Metab. 111, 63–72 (2014).
Eisengart, J. B. et al. Long-term cognitive and somatic outcomes of enzyme replacement therapy in untransplanted Hurler syndrome. Mol. Genet Metab. Rep. 13, 64–68 (2017).
de Vasconcelos, P. & Lacerda, J. F. Hematopoietic stem cell transplantation for neurological disorders: a focus on inborn errors of metabolism. Front. Cell. Neurosci. 16, 1662–5102 (2022).
Poe, M. D., Chagnon, S. L. & Escolar, M. L. Early treatment is associated with improved cognition in Hurler syndrome. Ann. Neurol. 76, 747–753 (2014).
Sessa, M. et al. Lentiviral haemopoietic stem-cell gene therapy in early-onset metachromatic leukodystrophy: an ad-hoc analysis of a non-randomised, open-label, phase 1/2 trial. Lancet 388, 476–487 (2016).
Leone, M. A. et al. Foetal Allogeneic Intracerebroventricular Neural Stem Cell Transplantation in People with Secondary Progressive Multiple Sclerosis: A phase I dose-escalation clinical trial. 2022.11.14.22282124 Preprint at https://doi.org/10.1101/2022.11.14.22282124 (2022).
Duma, C. et al. Human intracerebroventricular (ICV) injection of autologous, non-engineered, adipose-derived stromal vascular fraction (ADSVF) for neurodegenerative disorders: results of a 3-year phase 1 study of 113 injections in 31 patients. Mol. Biol. Rep. 46, 5257–5272 (2019).
Brinker, T., Stopa, E., Morrison, J. & Klinge, P. A new look at cerebrospinal fluid circulation. Fluids Barriers CNS 11, 10 (2014).
Bothe, G. W. M., Bolivar, V. J., Vedder, M. J. & Geistfeld, J. G. Behavioral differences among fourteen inbred mouse strains commonly used as disease models. Comp. Med. 55, 326–334 (2005).
Bryant, C. D. et al. Behavioral differences among C57BL/6 substrains: implications for transgenic and knockout studies. J. Neurogenet. 22, 315–331 (2008).
Bothe, G. W. M., Bolivar, V. J., Vedder, M. J. & Geistfeld, J. G. Genetic and behavioral differences among five inbred mouse strains commonly used in the production of transgenic and knockout mice. Genes Brain Behav. 3, 149–157 (2004).
Wagner, V. F. et al. University of Washington, Seattle, Seattle (WA), 1993).
Freed, C. R. et al. Survival of implanted fetal dopamine cells and neurologic improvement 12 to 46 months after transplantation for Parkinson’s disease. N. Engl. J. Med. 327, 1549–1555 (1992).
Freed, C. R. et al. Transplantation of embryonic dopamine neurons for severe Parkinson’s disease. N. Engl. J. Med. 344, 710–719 (2001).
Piao, J. et al. Preclinical efficacy and safety of a human embryonic stem cell-derived midbrain dopamine progenitor product, MSK-DA01. Cell Stem Cell 28, 217–229.e7 (2021).
Therapeutics, BlueRock Phase 1 Study To Assess the Safety and Tolerability of Human Embryonic Stem Cell-Derived Midbrain Dopamine Neuron Cell Therapy (MSK-DA01) For Advanced Parkinson’s Disease https://clinicaltrials.gov/study/NCT04802733 (2022).
Kirkeby, A. et al. Preclinical quality, safety, and efficacy of a human embryonic stem cell-derived product for the treatment of Parkinson’s disease, STEM-PD. Cell Stem Cell 30, 1299–1314.e9 (2023).
The STEM-PD Clinical Trial. The STEM-PD Clinical Trial https://stem-pd.org/.
Announcement of physician-initiated clinical trials for Parkinson’s disease|News & Events. CiRA | Center for iPS Cell Research and Application, Kyoto University http://www.cira.kyoto-u.ac.jp/e.
Petrus-Reurer, S. et al. Immunological considerations and challenges for regenerative cellular therapies. Commun. Biol. 4, 1–16 (2021).
Gene therapies for mucopolysaccharidoses - Rossi − 2024 - Journal of Inherited Metabolic Disease - Wiley Online Library. https://doi.org/10.1002/jimd.12626.
Giugliani, R. et al. Neurocognitive and somatic stabilization in pediatric patients with severe Mucopolysaccharidosis Type I after 52 weeks of intravenous brain-penetrating insulin receptor antibody-iduronidase fusion protein (valanafusp alpha): an open label phase 1-2 trial. Orphanet J. Rare Dis. 13, 110 (2018).
Irion, S. et al. Identification and targeting of the ROSA26 locus in human embryonic stem cells. Nat. Biotechnol. 25, 1477–1482 (2007).
Das, A. T., Tenenbaum, L. & Berkhout, B. Tet-on systems for Doxycycline-inducible gene expression. Curr. Gene Ther. 16, 156–167 (2016).
Khalil, A. S. & Collins, J. J. Synthetic biology: applications come of age. Nat. Rev. Genet 11, 367–379 (2010).
Stanimirovic, D. & Kemmerich, K. Conquering the barriers: are antibody therapeutics feasible for CNS indications? Future Neurol. 10, 67–70 (2015).
Mikitsh, J. L. & Chacko, A.-M. Pathways for small molecule delivery to the central nervous system across the blood-brain barrier. Perspect. Med. Chem. 6, 11–24 (2014).
Perriot, S., Canales, M., Mathias, A. & Du Pasquier, R. Generation of transgene-free human induced pluripotent stem cells from erythroblasts in feeder-free conditions. STAR Protoc. 3, 101620 (2022).
Yoshioka, N. et al. Efficient generation of human iPSCs by a synthetic self-replicative RNA. Cell Stem Cell 13, 246–254 (2013).
Baghbaderani, B. A. et al. cGMP-manufactured human induced pluripotent stem cells are available for pre-clinical and clinical applications. Stem Cell Rep. 5, 647–659 (2015).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Ciceri, G. et al. An epigenetic barrier sets the timing of human neuronal maturation. Nature 626, 881–890 (2024).
Ou, L., Herzog, T. L., Wilmot, C. M. & Whitley, C. B. Standardization of α-L-iduronidase enzyme assay with Michaelis-Menten kinetics. Mol. Genet Metab. 111, 113–115 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kluin, R. J. C. et al. XenofilteR: computational deconvolution of mouse and human reads in tumor xenograft sequence data. BMC Bioinforma. 19, 366 (2018).
Putri, G. H., Anders, S., Pyl, P. T., Pimanda, J. E. & Zanini, F. Analysing high-throughput sequencing data in Python with HTSeq 2.0. Bioinformatics btac166 https://doi.org/10.1093/bioinformatics/btac166 (2022).
SCANPY: large-scale single-cell gene expression data analysis | Genome Biology | Full Text. https://genomebiology.biomedcentral.com/articles/10.1186/s13059-017-1382-0.
Massively parallel digital transcriptional profiling of single cells | Nature Communications. https://www.nature.com/articles/ncomms14049.
Satopaa, V., Albrecht, J., Irwin, D. & Raghavan, B. Finding a ‘Kneedle’ in a Haystack: Detecting knee points in system behavior. 2011 31st International Conference on Distributed Computing Systems Workshops, Minneapolis, USA, 2011, pp. 166–171 (2010).
Acknowledgements
BlueRock would like to thank members of Mispro Biotech Services for assistance with animal husbandry and care, the NYU Genome Technology Center for sequencing and other support functions within BlueRock for assistance with work related to this study. Authors acknowledge the NHCDR and the NINDS Material, line LiPSC-GR1.1, referred to Line 6 in this manuscript. Generation of the GMP line LiPSC-GR1.1 was supported by the NIH Common Fund Regenerative Medicine Program, and reported in Stem Cell Reports. The NIH Common Fund and the National Center for Advancing Translational Sciences (NCATS) are joint stewards of the LiPSC-GR1.1 resource. NHCDR is housed at RUCDR, now known as Sampled.
Author information
Authors and Affiliations
Contributions
P.D., D.B., B.S., A.L., C.Pat., S.I. conceived and planned the experiments. In vivo work was designed and performed by A.L., D.L, J.W., N.S. and C.Pal.. Cell line engineering was designed and executed by C.S., C.M., A.F. and B.K. Bioinformatics analysis was performed by D.W., B.B., E.P., D. D.. Myeloid progenitor generation and in vitro phenotype analysis was performed by B.S., G.L., E.B. and S.F. hiPSC derivation and quality control by P.T. Assay development and implementation by E.S., M.E., N.A. and M.S. Project consultation and interpretation of results by C.Pal., M.T and M.S.. Conceptualization and supervision of the project by P.D., C.Pat. and S.I. Writing of the manuscript by D.B., P.D., C.Pat. and S.I.
Corresponding authors
Ethics declarations
Competing interests
All authors are current or past employees of BlueRock Therapeutics and have financial incentives to develop cellular medicines to reverse devastating diseases. All research was performed using BlueRock’s internal funds. BlueRock is a wholly-owned subsidiary of Bayer AG. There are no more competing interests.
Peer review
Peer review information
Nature Communications thanks Alessandro Aiuti and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Douvaras, P., Buenaventura, D.F., Sun, B. et al. Ready-to-use iPSC-derived microglia progenitors for the treatment of CNS disease in mouse models of neuropathic mucopolysaccharidoses. Nat Commun 15, 8132 (2024). https://doi.org/10.1038/s41467-024-52400-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52400-8
This article is cited by
-
Exploring retinal microglia: development, degeneration, and iPSC-derived model systems
Brain Structure and Function (2026)
-
From intention to implementation: automating biomedical research via LLMs
Science China Information Sciences (2025)