Abstract
Primary antiphospholipid syndrome (PAPS) is a life-threatening clotting disorder mediated by pathogenic autoantibodies. Here we dissect the origin of self-reactive B cells in human PAPS using peripheral blood and bone marrow of patients with triple-positive PAPS via combined single-cell RNA sequencing, B cell receptors (BCR) repertoire profiling, CITEseq analysis and single cell immortalization. We find that antiphospholipid (aPL)-specific B cells are present in the naive compartment, polyreactive, and derived from the natural repertoire. Furthermore, B cells with aPL specificities are not eliminated in patients with PAPS, persist until the memory and long-lived plasma cell stages, likely after defective germinal center selection, while becoming less polyreactive. Lastly, compared with the non-PAPS cells, PAPS B cells exhibit distinct IFN and APRIL signature as well as dysregulated mTORC1 and MYC pathways. Our findings may thus elucidate the survival mechanisms of these autoreactive B cells and suggest potential therapeutic targets for the treatment of PAPS.
Similar content being viewed by others
Introduction
The ability to bind self-antigens is widespread in the antibody repertoire within the immature and the naive B cell developmental stages. In healthy individuals, self-reactive specificities are counter-selected or regulated before reaching the memory of plasma cell (PC) compartment1,2,3. Failure in B cell tolerance checkpoints can lead to the production of autoantibodies and autoimmune diseases (AD)4. Such autoantibodies can be pathogenic with devastating consequences, as seen in systemic lupus erythematosus (SLE), rheumatoid arthritis, or antiphospholipid syndrome (APS).
Understanding the origins of self-reactivity holds therapeutic significance. While B cell depletion therapies are often beneficial, their inconsistent effectiveness suggests that either non-B cells or treatment-resistant B cell subpopulations may contribute to disease persistence. Upon antigen exposure and in the presence of a second signal from T cell help or Toll-like receptors, naive B cells can rapidly differentiate into plasmablasts via the extrafollicular (EF) pathway5,6. Alternatively, they may migrate to the germinal center (GC) and mature to class switched memory cells and long-lived PC that migrate to bone marrow (BM) to produce antibodies for a lifetime6. The affinity and activation of the B cell receptor (BCR) play critical roles in these developmental decisions7,8. In SLE, increased disease activity and the production of anti-dsDNA antibodies have been linked to an expansion of the EF B cell response9. The EF response has also been implicated in the breakdown of B cell tolerance in conditions such as rheumatoid arthritis and multiple sclerosis. Besides, the role of GC in these diseases is supported by defects in clonal redemption, in which somatic hypermutations (SHM) allow B cells to ‘mutate’ away from self-reactivity10,11,12.
APS is a life-threatening autoimmune clotting disorder characterized by a wide range of clinical symptoms, including venous and arterial thrombosis, or severe pregnancy complications13,14. These thrombotic events are directly linked to pathogenic autoantibodies known as anti-phospholipid antibodies (aPL), which target membrane phospholipids (PL) and/or their associated plasma proteins15,16. Diagnosis depends on the presence of at least one “criteria” aPL autoantibody (i.e. lupus anticoagulant, anticardiolipin, and/or anti-β2 glycoprotein I (β2GP1) antibodies)13,14. Patients with triple-aPL-positivity carry the highest thrombosis risk17. Primary APS (PAPS) occurs independently of other systemic AD, although APS frequently coexists with SLE18. Despite this association, distinct susceptibility genes highlight the differences between these two entities19, suggesting that the autoreactive B cell maturation pathways in SLE may not apply to aPL B cells in APS. Even though the disease is driven by pathogenic autoantibodies, no clear characteristics of B cells in APS have emerged. In a study of 11 PAPS patients an increased frequency of naive and transitional B cells was observed, along with a decrease in switched memory B cells compared to controls or patients who experienced a thrombotic event without aPL20. However, the observed variations were modest and not replicated in a larger cohort of 37 PAPS patients21. BCR repertoire has been extensively studied in several immune-mediated diseases, and profound changes have been noted in SLE, rheumatoid arthritis, or vasculitides22, but in-depth BCR repertoire analysis is still lacking in APS. Furthermore, little is known about aPL-producing B cells in APS. In fact, the diversity of self-antigens targeted by the autoantibodies represents a major challenge in identifying them. Systematic analysis of aPL variable region sequences suggested similarities in heavy chain variable region (VH) gene or family usage compared to control, but this study was performed on 36 sequences only and mostly derived from asymptomatic patients23. We previously demonstrated that unmutated aPL are less pathogenic than mutated aPL24,25,26. This raises the possibility that pathogens or commensals could drive the affinity maturation of these antibodies or their production27, or that additional intrinsic B cell factors favor the expansion of pathogenic clones.
To investigate the B cell characteristics underlying APS, we study a homogeneous cohort of PAPS patients with triple-aPL positivity, representing the most severe form of the disease and serving as a model of profound B cell tolerance breakdown, without confounding influences from other autoimmune disorders (e.g., SLE). Through single B cell repertoire analysis, transcriptomics, and reactivity profiling, we identify that aPL B cells arise from a polyreactive B cell pool with reduced BCR diversity. In PAPS patients, these autoreactive cells evade elimination from the post-antigenic compartment, likely due to defective germinal center selection. This contrasts with SLE, where autoantibodies predominantly emerge from the extrafollicular (EF) response. Moreover, our findings suggest that the persistence of these autoreactive B cells is sustained by elevated levels of interferon (IFN) and A proliferation-inducing ligand (APRIL), alongside dysregulation of the mammalian target of rapamycin complex 1 (mTORC1) and MYC pathways, highlighting potential therapeutic targets for APS treatment.
Results
B cell subpopulations distribution in triple positive PAPS is not altered
We studied 17 non-consanguineous male and female PAPS patients with triple-aPL positivity of European ancestry, aged 29 to 65 years. These patients met both the Sydney and 2023 ACR/EULAR criteria13,14 and had no other AD (Supplementary Data 1). Flow cytometry analysis revealed that numbers and percentages of total B cells were comparable between patients and age/sex-matched healthy donors (HD, 8.0% ± 0.9 of lymphocytes vs 9.9% ± 1.1, P = 0.3) (Fig. 1A, B). Consistent with previous report21, B cell subset distribution was similar between patients and controls. Although a slight, non-significant increase in transitional B cells (CD19+CD27-CD24highCD38high; 4.9% ± 0.7 vs 3.5% ± 0.4, P = 0.2) was observed, memory B cells frequency (CD19+CD27+; 24.4% versus 22.7%, P = 0.81), antibody secreting cells (ASC) frequency, and the distribution of B cells expressing distinct Ig subclasses were comparable to controls (Fig. 1A–D).
A Composite UMAP projection of B cell subsets, generated using 12 parameters analyzed in spectral flow cytometry of PAPS patient and HD samples (n = 30). Composite sample was derived from 5000 representative cells. B Frequencies of B cell subsets from PAPS patients (n = 15) and HD (n = 15). C Composite UMAP projection and frequencies of antigen-experienced B cells. D BCR isotype repartition in patients and HD samples. E Frequencies of aN and rN B cells from patients (n = 15) and HD (n = 15). F Composite UMAP projection and frequencies of DN B cells from patients (n = 15) and HD (n = 15). Significance was determined using a Two-tailed Mann–Whitney test. Data are presented as mean values ±SEM. aN activated naive B cells, DN double negative, HD healthy donor, PAPS primary antiphospholipid syndrome; rN resting naive B cells.
Extrafollicular B cell compartment is not expanded in PAPS
The EF response is characterized by B cells that express CD11c, the transcription factor T-bet and low level of CD2128. Increase of CD11c+ B cells, which include CD27−IgD+ activated naive (aN) B cells and a subpopulation of switched CD27−IgD− cells (double-negative 2, DN2), has been described in different conditions including SARS-Cov2 or autoimmune-prone Down syndrome patients9,28,29. In SLE, aN and DN2 cells are largely expanded and represent a pool of “pre-plasmablasts” enriched for autoreactive specificities5,30. Given the frequent association between APS and SLE, we investigated whether EF activation is a hallmark of PAPS. Neither DN2 nor aN B cells were significantly increased in PAPS patients compared to controls (0.46% ± 0.1 vs. 0.58% ± 0.1, P = 0.15; 0.69% ± 0.1 vs. 0.70% ± 0.1, P = 0.45, respectively) (Fig. 1E, F). We rather found a trend toward an increased DN1/DN2 B cell ratio (Fig. 1F). DN1 B cells are thought to be switched memory B cells precursors5. Frequencies of regulatory T cells and of other T cell subsets, including the circulating T follicular helper cells, frequently correlating with CD11c+ B cell expansion28, were also comparable between patients and controls (Supplementary Fig. 1A). Overall, PAPS patients exhibited mild B cell abnormalities without evidence of EF activation.
BCR diversity is reduced in PAPS
Within the BCR variable regions, distinct VH segments have been associated with autoimmunity22. We analyzed the BCR repertoire by extracting BCR sequences from a multimodal single-cell analysis of sorted B cells, combining CITE-seq analysis (i.e. single cell RNA sequencing (scRNA-Seq) and antibody-derived tags analysis) with single-cell V(D)J sequencing. This analysis was performed on B cells isolated from peripheral blood mononuclear cells (PBMC) of 10 PAPS patients and 7 HD (Fig. 2A).
A Schematic of peripheral B cell isolation from PAPS (n = 10) and HD (n = 7) blood followed by CITE seq including single-cell transcriptomic, antibody repertoire sequencing and ADT. VDJ sequencing was used to obtain paired heavy- and light-chain V(D)J with an average of 5,231 cells recovered per library. B–D IGHV (B), IGKV (C), and IGLV gene (D) usage frequencies in peripheral B cells from PAPS patients (red) and HD (blue). E, F IGHD families (E) and IGHJ gene (F) usage frequencies in peripheral B cells from PAPS patients (red) and HD (blue). G, H CDR3 length (G) and somatic hypermutations per VH segment (H) (nucleotide exchanges compared with the nearest germline gene segment). I, J Chao1 (I) and D50 diversity index (J) according to isotype in PAPS patients (red) and HD (blue). For IGHA diversity, n = 9 in patient group, as one patient did not have IgA+ B cells. Data are presented as mean values ±SEM. Significance was determined using a Two-tailed Mann–Whitney test. ADT antibody derived tag, HD healthy donor, PAPS primary antiphospholipid syndrome.
No significant differences were observed in V(D)J gene usage patterns (Fig. 2B–F) or in complementary determining region 3 (CDR3) amino acid length distribution (Fig. 2G). Similarly, SHM frequencies were similar, apart from a slight increase in immunoglobulin G (Fig. 2H). However, PAPS patients showed reduced clonotype diversity within the IGHV repertoire compared to HD (Fig. 2I, J), as indicated by the Chao1 index (representing the abundance of unique clones in a repertoire) and the D50 index (representing the number of clonotypes that account for 50% of cumulative unique CDR3 sequences). The constrained clonotype diversity within the IGHM compartment suggests impaired generation or selection of the primary repertoire in the BM.
aPL-specific B cells escape negative selection and persist after the naive B cell stage in PAPS
To identify aPL B cells in PAPS patients and examine their BCR profile, we performed single-cell EBV lymphoblastic transformation on isolated CD27- and CD27+ B cells from five PAPS patients with triple aPL positivity31,32,33. In this system, B cells were seeded individually and cultured without competition. Clonality was confirmed by Ig isotype and kappa/lambda staining. Based on Ig staining, the sorted transformed cells were classified as naive (CD27-IgD+), unswitched memory (CD27+IgD+), switched memory (CD27+IgD-), and double negative (CD27-IgD-) B cells.
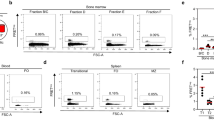
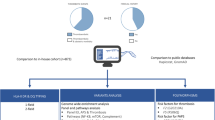
Amongst the criteria aPL, the phospholipid-binding protein β2GPI is the major antigenic target for thrombogenic aPL antibodies15,16,34,35. Patients may also develop non-criteria aPL, such as anti-phosphatidylserine/prothrombin complex, though the pathophysiological significance of this group is still under debate35,36,37,38,39,40. Therefore, we tested the monoclonal antibodies secreted by the B cell clones for reactivity against criteria PL (i.e., cardiolipin and β2GPI) to differentiate PL-reactive from non-PL-reactive clones (Fig. 3A). Additionally, all clones were tested for reactivity against non-criteria PL-related autoantigens using the “Pool PL” ELISA (Fig. 3A) to confirm that the non-aPL group indeed lacked PL reactivity. Only clones exhibiting either (i) criteria PL reactivity or (ii) no reactivity against both criteria and non-criteria PL were included in further analyzed.
A Flowchart of single B cell immortalization and supernatant screening. B Frequency of PL-reactive in each B cell subsets. C IgG subclasses repartition in non-PL and PL-reactive clones. D Kappa/lambda ratio of non-PL and PL-reactive clones in each B cell subsets. aPL antiphospholipid antibody, CL cardiolipin, PL phospholipid.
The frequency of B cell clones specific for PL decreased across maturation stages, with 30.9%, 21.9% and 15.2% of aPL-producing B cells in naive, unswitched memory and switched memory B cells, respectively (Fig. 3B). PL-specific B cell clones were found in the naive compartment in both HD and PAPS patients (39.3% and 30.9%, respectively), suggesting that aPL B cells are part of the natural repertoire. In HD, the frequency of aPL B cell clones dropped within the CD27+ memory B cells (12% vs 19.4% in PAPS patients, P = 0.02) especially in the switched memory population (7.4% vs 15.2%, P = 0.06; Fig. 3B and Supplementary Fig. 2). IgG+ aPL clones were of IgG1 and IgG3 isotype, consistent with a previous findings linking anticardiolipin specificity and IgG3 subtype41, with a significantly higher frequency of IgG3 compared to non-aPL clones (30.8% vs 4.7%, P = < 0.0001) (Fig. 3C). Autoantibody silencing can occur via light chain substitution, particularly through the use of lambda light chains1. However, we observed a predominant use of kappa light chains in aPL clones (Fig. 3D), suggesting that impaired receptor editing may support the maintenance of aPL in the B cell pool.
These data suggest a defect in peripheral tolerance in triple positive PAPS, allowing the persistence of memory aPL B cells in PAPS patients.
The naive pool of aPL B cells is largely polyreactive in PAPS
aPL are present in up to 20% of HD25,26, supporting the hypothesis that aPL are part of the natural repertoire. Natural autoantibodies are typically polyreactive1,42, hence we screened the polyreactive features of aPL and non-aPL antibodies secreted by B cell clones. Their reactivities were tested against HEp-2 cells by immunofluorescence assay and against a panel of defined antigens using ELISA. Naive and memory IgM+ aPL clones displayed significantly higher HEp-2 reactivity as compared to non-aPL clones (80.6% vs 17.6% for naive, and 44.7% vs 9.4% for memory, P < 0.0001, Fig. 4A and Supplementary Fig. 3A). Although IgG aPL progressively lost their HEp-2 reactivity, there was a trend toward higher reactivity compared to their non-aPL counterparts (18.2% vs 10%, P = 0.06). Notably, a recurrent cytoplasmic staining pattern suggestive of reactivity against vimentin, a proposed cofactor of cardiolipin and a potential target of aPL, was observed (Supplementary Fig. 3B)43. A high proportion of naive aPL clones exhibited reactivity to nucleosome, thyroglobulin, myosin, insulin and/or tetanus antigens (Fig. 4B). Antibodies with multiple reactivities (clusters 2,3 and 4) were predominantly from the aPL-positive pool, as indicated by unsupervised clustering approach based on ELISA reactivity (Fig. 4C). Specifically, 46.8%, 19.1%, and 0% of aPL from IgM+CD27-, IgM+CD27+, and IgG+CD27+ clones, respectively, recognized more than three antigens by ELISA (compared to 0% in non-aPL clones). Thus, (i) naive IgM+CD27- aPL clones were largely polyreactive, while (ii) switched clones lost this characteristic (Fig. 4D). Antigen array testing reactivity for 97 different foreign and self-antigens confirmed the high frequency of polyreactivity in IgM aPL clones (Supplementary Fig. 4). It has recently been shown that human IgG anti-β2GP1 cross-react with infectious mimotopes, suggesting that cross-reactivity may drive affinity maturation of natural PL-reactive B cells, which will then exhibit pathogenic features44. However, no reproducible pattern of reactivity was observed among aPL clones against bacterial or viral epitopes (eg. SARS-CoV2 peptides45) (Supplementary Fig. 4).
A HEp-2 reactivity of non-PL and PL-reactive clones, derived from PAPS patients, considering IgM+CD27-, IgM+CD27+ and IgG clones. Pie charts represent the frequencies of reactive (solid) and nonreactive (open) clones, with the number of clones tested (n) indicated in the center (left). B ELISA reactivity of non-PL and PL-reactive clones, derived from PAPS patients, against nucleosome, thyroglobulin, myosin, insulin and tetanus toxin. C K-means clustering plot showing cluster analysis of clones, derived from PAPS patients, reactivity in ELISA (left) and representation of the dimensional repartition of clones according to the PL reactivity (top right) and the number of different antigens recognized among nucleosome, thyroglobulin, myosin, insulin and tetanus toxin (down right). D Pie charts represent the number of reactivity detected among aPL (solid) and non-aPL clones (open), derived from PAPS patients, with the number of clones tested (n) indicated in the center. Significance was determined using a χ2 test. aPL antiphospholipid antibody.
As expected, HD clones exhibited similar characteristics, with 79.8% of naive IgM+ aPL clones testing positive on HEp-2 (vs 79.5% in PAPS patients, P = 0.9) and 48.5% recognizing more than three antigens by ELISA (vs 45.0% in PAPS patients, P = 0.9). A gradual reduction in polyreactivity was also observed as B cells matured (Supplementary Fig. 2B–D). These findings were further confirmed by the auto-antigen array (Supplementary Fig. 5).
In conclusion, aPL B cells in PAPS patients originate from a natural polyreactive B cell pool, present in HD, and lose polyreactivity as they reach the post-antigenic compartment. This suggests that affinity maturation had occurred in aPL-specific B cells in PAPS patients.
aPL antibodies have a constrained repertoire usage and distinct CDR3 features
Given that the variable region of the BCR in PAPS patients show lower diversity compared to controls, we explored the hypothesis that a restricted repertoire encoding the variable regions of heavy and light chain and/or intrinsic BCR features may explain a predetermined reactivity of aPL B cells. Distinct self-reactive BCR characteristics have been described in various autoimmune conditions, including skewed usage of VDJ families, longer CDR3, and impaired Ig light chain recombination1,22. Ig variable regions (heavy and light chain) from 376 clones were sequenced. We found that aPL clones compared to non-aPL had a restricted usage of V genes, with an enrichment of specific IGHV3-11, 3-21, 3-74, 4-34 and 4-39 genes (cumulative frequency: 26.9% in aPL clones vs 7.1% in the non-aPL clones, P = < 0.0001, Fig. 5A). An increased frequency of IGHV4-34 (and its 9G4 idiotype) has been described in AD and both IGHV3-74 and 4-39 have been isolated in patients with primary and secondary APS, respectively4,10,46,47. Considering individual samples, four out of five patients had a bias in BCR gene segments toward IGHV4-39 and three toward IGHV-3-74 and/or IGHV4-34 (Fig. 5A, see boxed figure). Of note, 71% of IGHV4-34 and 60% of IGHV4-39 aPL clones from the IgM+CD27- compartment belonged to cluster 3 and 4, indicating recognition of at least three different antigens (Fig. 4C).
A IGHV gene frequency of aPL B cell clones (red) compared to non-aPL clones (blue) for each patient. The box depicts the relative frequencies of IGHV3-11, 3-21, 3-74, 4-34, and 4-39 usage in aPL clones and non-aPL clones for each PAPS patient. B IGHD family frequency of aPL compared to non-aPL clones. Significance was determined using a χ2 test. C IGHJ gene frequency of aPL compared to non-aPL clones for each patient. D Number of CDR3 positively charged amino acid and arginine residues in CDR3 in aPL (n = 184) compared to non-aPL clones (n = 192). E Number of additional nucleotides in aPL (n = 182) compared to non-aPL clones (n = 188). F Length of CDR3 in aPL (n = 184) compared to non-aPL (n = 192) clones. G Number of SHM per VH segment (nucleotide exchanges compared with the nearest germline gene segment) in naive B cells, unswitched and switched memory B cells in aPL (n = 173) compared to non-aPL (n = 184) clones. H Selected IGHV gene frequency in CD27+ and CD27- population of aPL and non-aPL clones for each patient. I IGHV gene frequency of bone marrow plasma cells from two PAPS patients. Data are presented as mean values ±SEM. Significance was determined using a Two-tailed Mann-Whitney test, unless otherwise stated. aPL antiphospholipid antibody, HD healthy donor, PAPS primary antiphospholipid syndrome, R arginine, SHM somatic hypermutation.
Another distinguishing feature of aPL clones was the increased IGHD-6 family usage (29.4% vs 17.2%, P = 0.04, Fig. 5B), while a similar repartition of IGHJ gene usage was observed between the two groups (Fig. 5C). The CDR3 of aPL clones exhibited a significant higher number of positively charged amino acids (2.0 ± 0.1 vs 1.7 ± 0.1, P = 0.003), a feature associated with self-reactivity1 that is consistent with facilitated binding to negatively charged PL (Fig. 5D). Arginine residues in CDR are important in conferring the ability to bind β2GP148 and their frequency was increased in the CDR3 from aPL clones (Fig. 5D). No significant differences were observed in CDR3 length, number of non-templated (N) nucleotide or SHM (Fig. 5E–G). Light chain sequencing confirmed increased usage of kappa isotype in aPL clones (68% vs 52% in non-aPL clones) and revealed a preferential usage of IGKV3-20 (24% of kappa aPL clones vs 16%). Among lambda aPL clones, IGLV2-14 and IGLV2-23 were the most frequently used genes (mean frequency 21% and 15%, respectively, compared to 14% and 7% in non-aPL clones) (Supplementary Fig. 6A–H). No significant differences were found in the CDR3 regarding positively charged amino acids, SHM, or length. Antibodies encoded by IGHV3-11, 3-21, 3-74, 4-34, and 4-39 were more frequently associated with kappa light chain when they exerted PL reactivity (Supplementary Fig. 6I).
Our data suggests that VDJ usage is restricted in aPL B cells since the naive stage. The similar level of SHM in switched memory aPL clones compared to non-aPL clones supports the GC-dependent post-antigen maturation.
ASC secreting aPL persist in the BM of patients
In healthy individuals, several antibodies arising from the BM are self-reactive, but many of these are removed from the repertoire during B cell development through peripheral checkpoints1,2,3,49. In this view, B cells with IGHV4-34 BCR, which are intrinsically autoreactive, are censored in the GC and prevented from differentiating into ASC, a mechanism that is defective in SLE50. In our cohort IGHV3-21, IGHV3-74, and IGHV4-39 genes were still enriched in the memory compartment of aPL clones with a significantly higher frequency than in memory non-aPL clones (cumulative frequency: 20.9% vs 5.0%, P = 0.003, Fig. 5H), arguing that BCRs with specific aPL features persist in the post-antigenic compartments of PAPS patients. Conversely, IGHV4-34 and IGHV3-11 clones were no longer detected in the memory compartment of aPL clones (Fig. 5H). In HD, IGHV3-11, 3-21, 4-34, and 4-39 usage was also found in naive aPL clones but none of these VH families were detected in aPL memory B cells (Supplementary Fig. 7A, B). Long-lived PC are generated during the GC reaction, then migrate and survive in the BM. We characterized the antibody repertoire of BM PC in PAPS patients. We obtained the IGH and IGL chain genes of 69 single CD138+CD27+CD38+ PC isolated from the BM of two patients of our series. The frequency of IGHV4-39 usage, which was enriched in self-reactive naive B cells and memory cells, was found to be four times higher in PC from PAPS patients when compared with public data from BM PC of HD51 (Fig. 5I), supporting the hypothesis that autoantibodies present in the serum are produced by long-lived PC.
Thus, in PAPS patients PL-reactive B cells are not eliminated from the post antigenic compartment and become long-lived PC.
B cells from PAPS patients display MYC and mTORC1 activation and an increased level of TACI
To gain insight into the immunological mechanisms leading to the persistence and activation of aPL-reactive B cells in PAPS patients, scRNA-Seq libraries were dimensionally reduced and visualized in a single two-dimensional uniform manifold approximation and projection (UMAP). Unsupervised hierarchical clustering grouped the cells into 14 clusters (Fig. 6A, B). Differentially expressed genes (DEGs) were utilized to associate these clusters with previously identified B cell subpopulations (Fig. 6C and Supplementary Fig. 8A). Isotype determination using V(D)J sequences confirmed that naive, transitional and unswitched B cells segregated with IGHM, while switched and DN B cells segregated with IGHG/A (Supplementary Fig. 8B). The distribution of B cell subsets was similar between PAPS and HD, consistent with flow cytometry analysis. Additionally, the kappa/lambda ratio across all B cell subpopulations was also comparable (Supplementary Fig. 8C, D).
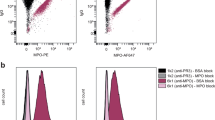
A UMAP projection and clustering of 78,355 B cells analyzed by scRNA-seq. B Bar plot showing cell frequencies from PAPS patients (n = 10) and HD (n = 7) that contributed to cluster c1 to c14. C UMAP projection showing six annotated B cell subtypes. D Plot showing the top 10 GSEA signaling pathway significantly enriched in total B cells from PAPS patients as opposed to HD. E Plots depicting the enrichment of MYC and mTORC1 GSEA signaling pathways in naive and memory B cells from PAPS patients as opposed to HD. F Heatmap listing top genes of mTORC1 signaling pathway enriched in PAPS patients. G Projected enrichment score of indicated pathway for each cell onto UMAP from APS patients. The darker shading represents higher enrichment. H Dot plot for expression of B cell cytokine receptors among naive clusters in all samples. Colors represent minimum-maximum normalized mean expression, and sizes indicate the proportion of cells expressing respective genes. I TACI MFI on naive B cells from two representative APS and two HD (left); TACI and BAFF MFI ratio on naive B cells from APS patients as opposed to HD (right) using FCM. Data are presented as mean values ±SEM. Significance was determined using a Two-tailed Mann–Whitney test. FCM flow cytometry, FDR False Discovery Rate, GSEA Gene Set Enrichment Analysis, HD healthy donor, PAPS primary antiphospholipid syndrome.
Gene set enrichment analysis (GSEA) revealed significantly increased expression of MYC target genes (normalized enrichment score (NES) = 7.0), genes related to oxidative phosphorylation (NES = 7.92), genes involved in IFNα/γ response (NES = 6.70), and genes related to mTORC1 axis (NES = 6.0) in PAPS B cells compared to HD B cells (Fig. 6D). The serine-threonine kinase mTOR is a component of two protein complexes, mTORC1 and mTORC2, which regulate protein synthesis, survival and cell growth52. mTORC1 is involved in B cell proliferation, and is needed for B cells to prepare PC functions and subsequent antibody synthesis53. Together with MYC, mTORC1 is essential in GC initiation, maintenance and positive selection by allowing selected light-zone B cells to proliferate and to re-enter the dark zone54,55. In PAPS patients, genes involved in MYC and mTORC1 signaling pathways were significantly enriched in both naive and memory B cells from patients (Fig. 6E). Individual heatmap on genes from the mTORC1 pathway showed that most of these genes, including mTORC1 positive regulators such as RPTOR or RHEB, were upregulated. (Fig. 6F). CD148 is a receptor phosphatase that is expressed in response to AKT phosphorylation in B cells56. We assessed CD148 expression, as proxy for Akt-mTOR activity, and found increased CD148 expression in naive B cells from PAPS patients (Supplementary Fig. 8E), in line with their mRNA changes.
Projecting scaled enrichment score for individual cells indicated that, among naive B cells, clusters 11 and 14 displayed the strongest enrichment in MYC, mTORC1 and IFNα activation (Fig. 6A, G). These clusters also exhibit the highest levels of IFN receptor genes expression and, together with cluster 7, show higher expression of TNFRSF13B, which encodes the transmembrane activator and CAML interactor (TACI), compared to other naive clusters (Fig. 6G, H). Of note, the frequencies of B cells in cluster 7, 11, or 14 were increased in PAPS patients (Fig. 6B). TACI serves as a receptor for B cell-activating factor (BAFF) and APRIL cytokines and is expressed on mature naive B cells. Engagement of TACI plays a crucial role in mitigating apoptosis, driving class-switch recombination and GC maintenance, as well as promoting antibody production57. In agreement with RNAseq data, a higher surface expression of TACI (but not BAFF receptor) was found on naive B cells using flow cytometry (Fig. 6I and Supplementary Fig. 8F), as well as increased level of APRIL in the sera of patients (Supplementary Fig. 8G).
Overall, our data suggests that B cells from PAPS patients, since the naive stage, experience proliferation and activation pressures involving IFN and APRIL cytokines, as well as the mTORC1 and MYC pathways.
MYC/mTORC1 activation is associated with hallmarks of GC differentiation and defective positive selection in aPL-specific B cells
To investigate the specific transcriptomic and protein expression profiles of aPL B cells in PAPS patients, we identified IGHV3-11, 3-74, 3-21, 4-34, and 4-39 BCR on CITE-Seq data. First, we confirmed a reduced diversity in these clonotypes which were associated to restrained IGLV in patients as compared to HD (Fig. 7A). The proportions of IGHV3-21, 3-74, and 4-39 B cells were comparable between patients and HD within the naive IGHM and IGHG/A populations (Fig. 7B).
A Circos plot depicting the IGHV:IGLV and IGHV:IGKV pairings for a HD and two representative PAPS patients. Pairing between IGHV3-11, 3-21, 3-74, 4-39, or 4-34 (pink segment), and IGKV (blue segment) or IGLV (green segment) gene are shown by the connecting lines inside the circle with thickness corresponding to pairing frequencies. B Frequencies of IGHV3-21, 3-74, 4-39 clonotypes in IGHM and IGHG/A isotypes in PAPS patients (n = 10) and HD (n = 7). C Violin plot representing normalized ADT level for B cell signaling and activation markers in IGHV4-39 clonotypes from APS patients (dark red), HD (dark blue), other clonotypes from PAPS patients (light red) and HD (light blue). D Dot plot for expression of mTORC1 and MYC selected genes among naive and memory cells according to clonotypes in PAPS patients and HD. E Dot plot for expression of pre-GC and plasmablast selected genes in naive cells according to clonotypes in PAPS patients and HD. F Violin plot representing pre-GC signature, SELL and CXCR5 gene expression according to clonotypes in patients and HD. G Frequency of plasmablast (CD38++CD27++) after B cell stimulation with CD40L + IL-21 + IL-4 for 6 days. Two-tailed Mann–Whitney test was used to compare groups. H Representative lineage trees for two clones in APS patients. I Representative amino acid VH alignment for IGHV4-39 clone (clone 1 depicted in H). J Frequency of positively charged amino acid induced by SHM in VH genes (top) of switched IgG/A B cells from HD (n = 8191) and PAPS patients (n = 9406). K Frequency of arginine residues induced by SHM in VH genes CDR of switched IgG/A B cells from HD (n = 7229) and PAPS patients (n = 6101). Data are presented as mean values ±SEM. Significance was determined using a Two-tailed Mann–Whitney test. ADT antibody-derived tag, CDR complementary determining region, HD healthy donor, MFI Mean fluorescence intensity, PAPS primary antiphospholipid syndrome, SHM somatic hypermutation.
Next, we focused on IGHV4-39 encoding aPL, as these were found up to the PC stage in PAPS patients. We assessed the expression levels of various cell surface proteins, BCR complex transmembrane proteins, and activation markers according to IGHV4-39 or non IGHV4-39 usage. Within each cohort, B cells expressing IGHV4-39 BCR were similar to those using other V-gene segments (Fig. 7C). However, naive B cells in PAPS patients displayed increased levels of HLA-DR molecules and decreased surface expression of IgM compared to those in HD. This change may enhance the likelihood of T cell help and affect the threshold of BCR engagement, thereby altering early peripheral negative selection. The memory B cell subset was similar between patients and controls (Fig. 7C).
We did not find differences in gene expression between IGHV4-39 and other B cells in patients, but we observed that MYC and mTORC1 pathway signatures were globally enriched compared to HD B cells, in both naive and memory compartments (Fig. 7D). Thus, MYC and mTORC1 activation appears to be a hallmark of all B cells in PAPS, rather than a specific signature of PL-reactive B cells. Additionally, both IGHV4-39 and other clonotypes in patients displayed an increased pre-GC signature58 and increased transcription of follicle homing receptors SELL (coding CD62L) and CXCR5 (Fig. 7E, F). Moreover, naive B cells in PAPS patients did not exhibit a plasmablast signature that would indicate that they were poised to generate short lived plasmablasts (Fig. 7E). Consistently, a trend of increased CXCR4 expression observed on circulating ASC (Supplementary Fig. 1B, C) suggested that they are inclined to migrate to the BM as long-lived PC, a process typically associated with GC maturation. In line with this, the in vitro formation of plasmablasts (CD27highCD38high), upon T help mimicking condition (stimulation with CD40L and IL-21), was enhanced in PAPS patients (Fig. 7G).
We next tested whether dysregulation of the mTORC1 or MYC axis, along with hyperresponsiveness to T cell help, could alter positive selection process in periphery. Given that BCR with IGHV3-21, 74 or 4-39 were only slightly increased in the post antigenic compartment, we hypothesized that patients fail to attenuate their self-reactivity in late processes, such as clonal redemption. Analysis of patients’ intra-clonal VH mutations in patients revealed that SHM more frequently introduced positively charged amino acids, particularly arginine residues in the CDR compared to HD (Fig. 7H–K), consistent with the features of aPL clones (Fig. 5D).
Altogether, these results reinforce the perspective that in patients with PAPS, B cells are preferentially oriented towards GC maturation while failing to eliminate or attenuate the self-reactivity of PL-reactive BCR.
Discussion
APS is a model for auto-antibody mediated disease37, in which the genesis of pathogenic aPL remains incompletely understood due to the complex pattern of aPL reactivities and the challenges of identifying aPL-specific B cells. To address these challenges, we focused on PAPS patients to avoid the confounding influence of SLE, which is frequently associated with APS. Additionally, we studied triple-positive PAPS, a subset characterized by the highest thrombosis risk17. By employing B cell scRNAseq in conjunction with single-cell EBV immortalization, we were able to link the phenotype and function of self-reactive B cell specificities. Our findings indicate that in triple positive PAPS, self-reactive naive B cells preferentially mature via the follicular pathway and become long-lived PC. The altered positive selection in the post antigenic compartment suggests that dysregulation of mTORC1 or MYC axis and hyperresponsiveness to T cell help could enable natural naive PL-reactive B cells to enhance their specificity and become pathogenic.
It is widely accepted that most self-reactive B cells, including those specific to antinuclear antigen, are primarily regulated during early B cell development in the BM through mechanisms such as clonal deletion, receptor editing, or silencing during the transition from early immature to immature stages2. A second tolerance checkpoint further eliminates autoreactive B cells during the transition from new emigrant or transitional B cells and mature naive B cells. Despite these processes, the naive B cell repertoire of HD still retains ~20% of self-reactive B cells1. Given that most naive or IgM+ aPL B cell clones in patients with PAPS exhibit polyreactivity towards self or non-self-antigens, we conclude that aPL B cells originate from a heterogenous natural polyreactive pool of B cells. This finding holds true for both PAPS patients and healthy individuals, as supported by our data. Furthermore, this conclusion is further supported by previous data on two anti-β2GP1 clones generated by phage display, which have highly conserved germline VH and VL genes and a reactivity against cationic and zwitterionic phospholipids, characteristic of human oligoreactive natural autoantibodies59. Our findings are also consistent with the observation that an scFv-Fc (single-chain fragment variable) isolated from a normal human phage library and targeting the D1 domain of β2GPI, exhibits pathogenic behavior similar to that of spontaneous aPL in animal models60.
After antigen encounter, activated B cells either differentiate into short-lived PC or, following cognate interaction with antigen-specific T cells, can form GCs61,62,63. B cells that successfully pass the GC process increase their affinity, through iterative cycles of clonal expansion and SHM in the variable regions of the heavy and light chains. These cells can then become long-lived PC or alternatively memory B cells and long-lived quiescent cells capable of reactivation58. However, class switch recombination (CSR) and SHM can occur both inside and outside of GC64. Indeed, SLE is characterized by B cells lacking IgD and CD27 (DN), which include CXCR5-CD11c+ cells (DN2), actors of EF response, that represent pre-PC5. While we confirm the presence of an IFN transcriptional signature in PAPS B cells, neither our phenotypic or transcriptomic analysis show similar B cells subpopulation repartition and neither of them argue for a disequilibrium of DN2 or EF B cell activation in PAPS. Furthermore, considering the similar occurrence of SHM in both antigen-experienced aPL and non-aPL B cell clones, we conclude that activated aPL B cells in PAPS likely undergo a GC maturation. In some PAPS patients, an expansion of TFH may contribute to breakdown of tolerance in the GC65. We did not find a TFH expansion in our cohort, consistent with the findings of Faliti et al. who demonstrated that the frequency of CCR7lowPD1high CXCR5+ helper T cells is normal in PAPS but increased in SLE patients66. Since APS is often associated with SLE, the specific fate of aPL B cells in SLE remains an open question.
During B cell development in the fetal liver and BM, recombination of immunoglobulin genes generates the naive BCR repertoire. This repertoire is then refined by eliminating or suppressing self-reactive B cells1,4,67, to mitigate the likelihood of AD. Ultimately, this repertoire is shaped through interactions with antigens, including pathogens. As a result, the features of the BCR repertoire correlate with pathogen interactions and, should tolerance mechanisms fail, immune-mediated diseases22. For instance, a comparison of B cell repertoires in SLE and anti-neutrophil cytoplasmic antibody-associated vasculitis (ANCA-AAV) reveals a skewed IGH repertoire in both diseases with increased expression of genes of the IGHV4 family. Notably IGHV4-34 encoded BCR, which can bind both autoantigens and commensal bacterial, has been associated with SLE in both the naive and in the post-antigenic compartments. Extensive BCR repertoire analysis has not yet been conducted in APS. BCR diversity appears significantly reduced in PAPS, as observed in other AD, with a clonal expansion and restricted usage of light and heavy chains. Specific VH encoded BCR, such as IGHV4-39, previously described in a few SLE and APS patients4, are associated to autoreactive positively charged BCRs and can be tracked from the naive stage to the long-lived PC compartment. Although the self-reactive antibodies expressed by polyreactive naive B cells in PAPS may not be directly pathogenic, they could readily be restricted to some IGH or IGL families and undergo class switching upon B cell activation. Furthermore, aPL specificities might be selected or expanded during this maturation process, leading to the loss of polyreactivity, as suggested by the critical role of SHM in driving the thrombotic process in vivo24,25,26. In this context, cross reactivity with exogenous pathogens might contribute to the development of pathogenic B cells44. However, our experiments, which involved testing the reactivity of B cell clones against an antigen array, which included various infectious triggers like SARS-CoV2 peptides, did not provide conclusive evidence on this matter.
We hypothesized that autoreactive B cells possess distinct signaling characteristics, as even modest alteration in B cell signaling within the appropriate environmental context may be sufficient to promote or sustain auto-immunity68. To investigate this further, we conducted a two-step analysis. First, we performed single-cell transcriptomics on sorted B cells. Second, we focused on the previously identified specific IGHV and IGLV rearrangements. Our study demonstrates that in PAPS patients, B cells exhibit a global activation state with a higher prevalence of specific BCR. This activation state appears to be associated with mTORC1 signaling, independently of specific rearrangements. The increased expression of CD148, a proxy for Akt-mTOR activity56, is consistent with the activation of the mTORC1 pathway. Interestingly, one of the exceedingly rare monogenic defects observed in PAPS involves a gain-of-function mutation in mTOR69. The involvement of the mTOR pathway in APS has been demonstrated in patients with APS-related nephropathy at the endothelial level70. However, the role of mTOR signaling in PAPS B cells has not been previously explored. Among the upstream signals of mTORC1, the cytokine BAFF, also known as TNFSF13B, plays a crucial role in B cell survival and homeostasis71,72. Based on our findings, we propose that increased TACI engagement on naive B cells in PAPS contributes to mTOR signaling, which, in turn, likely drives the GC maturation of polyreactive autoreactive B cells towards a pathogenic state. Increase of IFN levels, which is known to alter B cell response to MYD88 signaling73, could potentiate this B cell tolerance checkpoint disruption, as suggested by the IFN transcriptional signature we observed in B cells and previously found in whole blood transcriptome of patients with triple-positive PAPS74,75.
In summary, our findings indicate that PL-reactive B cells in patients with PAPS originate from a pool of naturally polyreactive B cells. While further validation in larger cohorts is needed to generalize our observations, it appears that these B cells exhibit a restricted BCR repertoire and evade peripheral tolerance mechanisms at the naive stage. This escape is sustained by TACI and IFN-R engagement, along with increased MYC/mTOR signaling. At this stage, PAPS B cells are likely exposed to antigenic stimuli, although the specific triggers remain unidentified. Previous evidence suggests possible cross-reactivity with commensal or pathogenic antigens, but the role of local environmental triggers mediated by monocyte or neutrophil activation, along with mTOR signaling, remains under debate. Our results suggest that EF differentiation is not a predominant pathway for PAPS B cells. Instead, GC maturation seems to play a more significant role, promoting the development of memory B cells and ASC with enhanced affinity for PL.
Interestingly, mTOR has been identified as a potential therapeutic target for vascular lesions associated with APS nephropathy. In renal transplant patients with APS-related nephropathy showed improved graft survival when treated with rapamycin (an mTORC1 inhibitor), which also prevented vascular lesion recurrence, such as cell proliferation and intimal hyperplasia70. However, only ten patients were treated, four of whom had only lupus anticoagulant (making titer variation assessment impossible), and only two had anti-β2GPI antibodies. Therefore, how mTOR inhibition affects APS B cells remains an unresolved question. Nevertheless, our data support the potential benefits of rapamycin in mitigating abnormal B cells responses and aPL production in APS.
Methods
Ethics
All patient data and samples were obtained in accordance with the Declaration of Helsinki and the Ethical Board of Strasbourg University Hospital (Comité de Protection des Personnes: CPP-Est IV N° 13/48). All participants provided informed written consent.
Study and design
The objective of this study was to analyze the features (i.e. BCR characteristics, phenotypic and transcriptomic profiles) of B cells and aPL-producing B cells in a cohort of PAPS patients with triple positivity. We used high dimensional flow-cytometry, integrated CITE-Seq and BCR-Seq approaches to compare the B cells of a cohort of HD with a cohort of PAPS patients. Single B cells repertoire and aPL reactivity were further characterized in smaller sub-cohorts with an EBV single-cell immortalization method applied on sorted B cells31.
Recruitment of participants and sample collection
The diagnosis of PAPS was confirmed by two senior physicians specialized in Internal Medicine and in Clinical Immunology, according to revised Sydney criteria13. All patients also fulfilled 2023 ACR/EULAR classification criteria14. Triple positivity was considered when persistent positivity (on two or more occasions at least 12 weeks apart) for lupus anticoagulant, anticardiolipin and anti-β2GPI antibodies (IgG or IgM at medium or high levels) was present. Patients were enrolled in the Clinical Immunology Department (National Reference Center for Rare Autoimmune Systemic Diseases, CRMR RESO) of Strasbourg University Hospital. Although some of them presented antinuclear positivity, they were not affected by another AD including SLE, and none of them had significant levels of anti-dsDNA.
Clinical and immunological features of the patients are available in Supplementary Data 1.
Cell staining and phenotyping
Human B cell phenotype was assessed on fresh EDTA peripheral blood samples immediately after collection. BM cells were subjected to density gradient centrifugation to get mononuclear cells suspensions and remove dead cells. Samples were incubated with antibodies for 15 minutes at room temperature. Dead cells were excluded Zombie NIRTM (423105, Biolegend). The cells were analyzed on a spectral flow cytometer (Aurora, Cytek Biosciences). Data analysis was performed using FlowJo software (TreeStar). The following anti-human antibodies were used: PerCP anti-CD3 (300427, clone UCHT1 Biolegend), SB436 anti-CD3 (62-0037-42, clone OKT3, Invitrogen), BV750 anti-CD4 (344643, clone SK3, Biolegend), BV570 anti-CD8a (301037, clone RPA-T8, Biolegend), BV711 anti-CD10 (312225, clone Hl10a, Biolegend), BV421 anti-CD11c (371511, clone S-HCL-3, Biolegend), SB436 anti-CD16 (62-0166-42, clone 3G8, Invitrogen), APC/Cy7 anti-CD19 (302218, clone HIB19, Biolegend), PB anti-CD20 (302320, clone 2H7, Biolegend), FITC anti-CD21 (561372, clone B-ly4, BD Bioscience), AF700 anti-CD21 (354918, clone Bu32, Biolegend), FITC anti-CD24 (555427, clone ML5, BD Bioscience), PE anti-CD24 (555428, clone ML5, BD Bioscience), Kiravia Blue 520 anti-CD25 (356143, clone M-A251, Biolegend), APC anti-CD27 (558664, clone M-T271, BD Biosciences), BV650 anti-CD27 (563229, clone L128, BD Bioscience), BV786 anti-CD27 (563327, clone L128, BD Bioscience), SB436 anti-CD33 (62-0338-42, clone WM-53 (WM53), Invitrogen), PE-Cy7 anti-CD38 (560677, clone HIT2, BD Bioscience), PerCP-eFluor710 anti-CD38 (46-0388-42, clone HB-7, Invitrogen), BV650 anti-CD38 (356620, clone HB-7, Biolegend), APC-Fire750 anti-CD45RA (304151, clone Hl100, Biolegend), BV480 anti-CD69 (747519, clone FN50, BD Bioscience), APC-R700 anti-CD127 (565185, clone HIL-7R-M21, BD Bioscience), APC-Fire750 anti-CD138 (352315, clone DL-101, Biolegend), BV480 anti-CD138 (352301, clone DL-101, Biolegend), BV605 anti-BAFF receptor (743571, clone 11C1, BD Bioscience), PE/Dazzle 594 anti-TACI (311910, clone 1A1, BioLegend), BV421 anti-CCR7 (740052, clone 3D12, BD Bioscience), AF647 anti-CXCR5 (558113, clone RF8B2, BD Bioscience), BV605 anti-IgM (314524, clone MHM-88, Biolegend), PECF594 anti-IgM (562539, clone G20-127, BD Bioscience), AF647 anti-IgM (109-605-129, polyclonal, Jackson ImmunoResearch), PerCP-Cy5.5 anti-IgD (561315, clone IA6-2, BD Bioscience), PerCP-eFluor710 anti-IgD (46-9868-42, clone IA6-2, Invitrogen), PerCP anti-IgG (109-125-098, polyclonal, Jackson ImmunoResearch), PE anti-IgG1 (CYT-IGG1PE, clone SAG1, Cytognos), FITC anti-IgG2 (CYT-IGG2F, clone SAG2 Cytognos), PE anti-IgG2 (CYT-IGG2PE, clone SAG2 Cytognos), FITC anti-IgG3 (CYT-IGG3F, clone SAG3, Cytognos), APC anti-IgG4 (CYT-IGG4AP, clone SAG4, Cytognos), PerCP-Cy5.5 anti-IgA1 (CYT-IGA1C, clone SAA1, Cytognos), APC anti-IgA1 (CYT-IGA1AP, clone SAA1, Cytognos), PerCP-Cy5.5 anti-IgA2 (CYT-IGA2C2, clone SAA2, Cytognos), PE-Cy7 anti-IgE (325509, clone MHE-18, BioLegend), FITC anti-light chain κ (316506, clone MHK-49, Biolegend), PE anti-light chain lambda (316607, clone MHL-38, Biolegend) and BV480 TCRgd (746498, clone 11F2, BD Bioscience).
Details concerning the phenotypic characterization and the gating strategies used for B cells are available in Supplementary Data 2 and Supplementary Fig. 9, respectively.
B cell sorting and immortalization
B cells were FACS sorted in naive (CD3-CD19+CD27-) and memory (CD3-CD19+CD27+) subsets. Sorted cells were then incubated in complete medium containing 50% EBV supernatant (of B95-8 cells) and 2.5 mg/mL CpG ODN 2006 (tlrl-2006, Invivogen) for 4 hours at 37 °C. Cells were then seeded on 10 000 irradiated allogeneic mononuclear cells at 3 cells per well in 96 U-bottom plates31.
Assessment of clonality of single EBV B cell clones in the final population was performed staining for the kappa or lambda light chain by flow cytometry and complemented with BCR sequencing data and ELISA. Clonality was defined by: a) presence of cells with only one type of light chain (kappa or lambda) using flow cytometry, and b) presence in supernatant of only one isotype of immunoglobulin heavy chain. Cells were then categorized, based on their isotype and the initial sort, in naive (CD3-CD19+CD27-IgG/A-), unswitched memory (CD3–CD19+CD27+IgG/A-), switched memory (CD3-CD19+CD27+IgG/A+) or DN (CD3–CD19+27-IgG/A+) subsets.
ELISA
Immunoglobulin concentration
Immunoglobulin concentration in the supernatant of the EBV clones was analyzed by ELISA. 96 wells (442404, Nunc Maxisorp, Dutscher) were coated with anti-human immunoglobulin, either anti-IgM (109-006-129, polyclonal, Jackson ImmunoResearch), anti-IgG (109-006-098, polyclonal, Jackson ImmunoResearch) or anti-IgA (109-006-011, polyclonal, Jackson ImmunoResearch) in bicarbonate buffer and serial dilution of clone supernatants were added.
Polyreactivity assays
96 wells (442404, Nunc Maxisorp, Dutscher) were coated with 100 µL of either insulin (I0908, Sigma-Aldrich; 2 µg/mL), thyroglobulin (609312, Sigma-Aldrich, Germany; 2 µg/mL), myosin (M1636, Sigma-Aldrich; 2 µg/mL) or nucleosome solution (DNA final concentration: 1 µg/mL) in bicarbonate buffer at 37 °C overnight. Plates were saturated with PBS/0.05% (v/v) Tween/1% (w/v) bovine serum albumine (BSA) for 1 hour at 37 °C. 100 µL of EBV clones’ supernatants ([IgM, IgG or IgA] = 2 µg/mL) were incubated overnight at +4 °C. Antibodies were considered polyreactive when they recognized at least three analyzed antigens. For nucleosome obtention HEK293 cells were cultured, and 1 × 109 cells were centrifuged at 400 g for 5 minutes. Before use, 100 µL of protease inhibitor cocktail (P8340, Sigma-Aldrich) was added to buffers A through D. The cells were washed twice (centrifugation: 5 min, 400 g) using 10 mL of buffer A (15 mM Tris-HCl, pH 7.4, 15 mM NaCl, 60 mM KCl, 5 mM MgCl2). The cells were then washed with 10 mL of buffer B (10 mM Tris-HCl, pH 7.4, 1 mM KCl, 1.5 mM MgCl2) and resuspended in 10 mL of buffer B. The cells were homogenized on ice for 5 minutes. Next, 10 mL of buffer A was added, followed by centrifugation at 1000 g for 10 minutes. The resulting pellet was washed with 10 mL of buffer A and centrifuged again (10 min, 1000 g). The pellet was then resuspended in 10 mL of buffer C (15 mM Tris-HCl, pH 7.4, 15 mM NaCl, 60 mM KCl, 5 mM MgCl2, 1 mM CaCl2), and the absorbance was measured at 260 nm using a 1:100 dilution in 0.1% (v/v) sodium dodecyl sulfate (SDS) solution. Micrococcal nuclease (N3755, Sigma-Aldrich) was added at a concentration of 0.1 units per absorbance unit to digest free DNA. The samples were incubated at 37 °C for 20 minutes, after which 500 µL of 200 mM ethylenediaminetetraacetic acid (EDTA) was added to stop the reaction. After centrifugation at 1500 g for 10 minutes at 4 °C, the pellet was resuspended in 10 mL of buffer D (1 mM Tris-HCl, pH 7.4, 1 mM EDTA, pH 7.4). The solution was incubated on ice for 30 minutes with regular shaking. Following centrifugation (10 min, 1500 g, 4 °C), the supernatant containing nucleosomes was collected and dialyzed (20 kDa molecular weight cut-off) against 500 mL of buffer D in two steps at 4 °C: first for 2 hours, then overnight with fresh buffer. DNA concentration was measured using a 1:100 dilution in 0.1% (v/v) SDS solution. The samples were stored at −20 °C until use76.
Revelation step
Bound Ig were detected with peroxidaxe conjugated anti-human IgM (109-036-129, polyclonal, Jackson ImmunoResearch), IgG (109-036-088, polyclonal, Jackson ImmunoResearch) or IgA (109-036-011, polyclonal, Jackson ImmunoResearch), and developed with TMB (3,3’,5,5’-tetramethylbenzidine). After 20 minutes, the reaction was stopped with 1 M HCl, and the absorbance was measured at 450 nm using a MultiScan FC (Thermo Fisher Scientific).
Antiphospholipid ELISA
Reactivity against PL was determined using modified protocols from Lieby et al.24,25. All supernatants were systematically tested in two types of ELISA: the cardiolipin/β2GPI ELISA and the “pool PL” ELISA. aPL with cardiolipin/β2GPI were selected for further analysis. “Pool PL” ELISA was essential to confirm that our non-aPL group genuinely lacked PL reactivity.
For the cardiolipin/β2GPI ELISA, polystyrene microtiter plates (456529, Dutscher) were coated with 50 μL/well of cardiolipin (C1649, Sigma-Aldrich) at 50 μg/mL in absolute alcohol and evaporated at room temperature for 4 hours. Plates were saturated with 1 µg/mL of purified human cofactor β2GP1 (RP43146, Invitrogen) for 2 hours at room temperature. No FCS was used.
For the “pool PL” ELISA, polystyrene microtiter plates (456529, Dutscher) were coated with 50 μL/well of cardiolipin (C1649, Sigma-Aldrich), phosphatidylserine (P7769, Sigma-Aldrich), phosphatidic acid (840101 C, Sigma-Aldrich), phosphatidylcholine (P3556, Sigma-Aldrich), phosphatidylethanolamine (P7943, Sigma-Aldrich), phosphatidylinositol (P2517, Sigma-Aldrich) and phosphatidylglycerol (841138 C, Avanti Polar Lipids,) at 25 μg/mL in absolute alcohol and evaporated at room temperature for 4 hours. Plates were saturated with PBS 10% FCS with 5 µg/mL of prothrombin (539515, Sigma-Aldrich) for 2 hours at room temperature.
40 µL of supernatant were then incubated overnight at +4 °C. Bound Ig were detected with peroxidaxe conjugated anti-human IgM (109-036-129, Jackson ImmunoResearch), IgG (109-036-088, Jackson ImmunoResearch) or IgA (109-036-011, Jackson ImmunoResearch), and developed with TMB (3,3’,5,5’-tetramethylbenzidine). After 20 minutes, the reaction was stopped with 1 M HCl, and the absorbance was measured at 450 nm using a MultiScan FC (Thermo Fisher Scientific).
Patients with known titer of aPL antibodies were systematically used as positive controls. Additionally, serums from the same 5 HD were systematically added to establish a positivity threshold based on a z-score; z-score = (X-M)/SD in which X is the optical density (OD) of the considered sample (EBV supernatant), M and SD are the mean and standard deviation of the OD for the HD group. Sample was labeled as containing (i) aPL if the z-score was >10, (ii) and non-aPL antibodies if the z-score was <10 in pool PL ELISA and in cardiolipin/β2GPI ELISA. Samples reactive in cardiolipin/β2GPI ELISA with non-reactivity in pool PL ELISA were labeled as doubtful and definitively discarded. Samples reactive in pool PL ELISA with non-reactivity in cardiolipin/β2GPI ELISA were labeled as non-criteria aPL and discarded. Only clones exhibiting either (i) criteria PL reactivity or (ii) classified as non-aPL with a z-score below 2 in both assays were further analyzed. Of note, the blocking buffer in the “pool PL” ELISA may have facilitated the dissociation of the PS-PT complex, which could have allowed rare anti-PS/PT antibodies to be included in the non-aPL group.
HEp-2 reactivity
HEp-2 reactivity was assessed on slides coated with HEp-2 cells (FA2400, ZEUS scientific). cell-coated slides were incubated in a moist chamber at room temperature with 30 µL of EBV clones’ supernatants and detected with AlexaFluor488 goat anti-human IgG (A48276, polyclonal, Invitrogen; dilution 1/200) or AlexaFluor488 goat anti-human IgM (A-21215, polyclonal, Invitrogen; dilution 1/200). ZEISS Axioskop microscope was used to read patterns. Pictures were acquired with Zeiss LSM800 (20x/0.8).
Autoantigens array
Autoantigen microarrays were manufactured in the Microarray & Immune Phenotyping core Facility of University of Texas Southwestern Medical Center (Dallas, TX, USA). Human EBV clones’ supernatants were first treated with DNAse I (AM2222, Invitrogen) to remove free-DNA and then applied onto autoantigen arrays with 1:2 dilution. The autoantibodies binding was detected with laser wavelengths 532 nm (cy3 labeled anti-IgG, 109-166-098, polyclonal, Jackson ImmunoResearch) and 635 nm (cy5 labeled anti-IgM, 109-606-129, polyclonal, Jackson ImmunoResearch).
Genepix Pro 7.0 software was used to analyze the images and generate the Genepix report (GPR) files (Molecular Devices, Sunnyvale, California, USA). The net fluorescent intensity (NFI) of each antigen was generated by subtracting the local background and negative control (Phosphate buffered saline) signal. The signal-to-noise ratio (SNR) was also generated for each antigen. SNR is used as a quantitative measure of the ability to resolve true signal from background noise. A higher SNR indicates higher signal over background noise. NFI was normalized by robust linear model (R MASS package) using positive controls with different dilutions. To avoid outliers in either NFI or SNR, autoantibody score (Ab-score) is calculated by log 2((NFI*SNR) + 1). Values are plotted as fold change versus the mean of the Ab-Score of the non-aPL group for the specified autoantigen within the considered subset (i.e. IgM (CD27-), IgM (CD27+) or IgG).
Details concerning the proteins on the microarray are available in Supplementary Data 3.
B cell clone repertoire analysis
RNA of B-EBV cell clones was isolated using NucleoSpin RNA XS (740990, Macherey Nagel), and then reverse transcribed on a T100™ Thermal cycler (BioRad) using Maxima H Minus Reverse Transcriptase (EP0752, Thermo Fisher Scientific) following the manufacturer’s protocol. To amplify IGH/IGL transcripts, 3.5 μl of cDNA products was amplified by a PCR in 20 μl reactions containing 20pM primers (available in Supplementary Data 4) and 10 µL of HotStarTaq Plus Master Mix Kit (203645, Qiagen, 1U of HotStarTaq Plus DNA Polymerase per reaction). PCR was performed for 45 cycles at 94 °C for 30 sec,57 °C for 30 sec, 72 °C for 55 sec. All PCR products were purified from agarose gel using NucleoSpin Gel and PCR Clean-up (740609, Macherey Nagel) and then sequenced by Sanger sequencing (Eurofins Genomic). Sequences were analyzed using IMGT/HighV-QUEST (http://www.imgt.org/IMGT_vquest/input); and the NCBI’s IgBLAST server (http://www.ncbi.nlm.nih.gov/igblast/). Rare non-germline sequences that contained high numbers of mutations that could not be attributed to sequence polymorphisms were excluded from further analysis.
Bone marrow plasma cell repertoire analysis
Single-cell sorting of PC (CD3−CD19+CD27highCD38highIgD-CD24-CD138+) populations was performed on a FACSAria II flow Cytometer (BD Biosciences) into 96-well PCR plates. RNA from these single cells was reverse-transcribed in the original 96-well plate in 40 μl reactions using SuperScript IV CellsDirect cDNA Synthesis Kit (11750350, Thermo Fisher Scientific) following the manufacturer’s protocol. 4 µl of cDNA product were used to amplify IGH transcripts by two rounds of PCR in 20 μl reactions containing 20 pM primers and 10 µL of HotStarTaq Plus Master Mix Kit (203645, Qiagen). PCR was performed for 45 cycles at 94 °C for 30 sec, 57 °C for 30 sec, 72 °C for 55 sec (1st-PCR) or 45 sec (2nd PCR)1. PCR efficiencies ranged from 30-40%. All PCR products were purified from agarose gel using NucleoSpin Gel and PCR Clean-up (740609, Macherey Nagel), sequenced by Sanger sequencing (Eurofins Genomic) and analyzed for IGHV, D and J gene usage on IMGT/HighV-QUEST and NCBI’s IgBLAST server.
Plasma APRIL level
The APRIL level in the plasma of PAPS patients was assessed using the LEGENDplex™ Human B cell Panel (740527, Biolegend), following the manufacturer’s instructions. An additional cohort PAPS cohort with triple-aPL-positivy (n = 10) was used for this assay.
In vitro plasmablast differentiation assay
PBMC were isolated from the blood of PAPS patients and HD. Total B cells were isolated with the EasySep™ Human B cell Isolation Kit (17954, Stemcell Technologies) following manufacturers’ instructions. A total of 50,000 B cells/well were plated into U96 wells and stimulated with CD40L (rcyec-hcd40l, Invivogen), IL-21 (200-21, Preprotech) and IL-4 (200-04, Preprotech) in Roswell Park Memorial Institute medium (RPMI; Life Technologies) enriched with gentamicine (50 µg/mL), supplemented with 10% FCS. After 6 days in culture, the phenotype of human B cells was determined using Attune NxT Flow Cytometer (Invitrogen).
CITE-SEQ and VDJ sequencing
PBMC was isolated from the blood of ten PAPS patients and seven HD. Total B cells were isolated with the EasySep™ Human B cell Isolation Kit (17954, Stemcell Technologies). B cells were resuspended at 10 × 106/mL and incubated with Human TruStain FcX Fc Blocking reagent (422301, Biolegend) for 10 min at 4 °C. Cells were then incubated with the following anti-human antibodies conjugated to an oligomer for 30 min at 4 °C: TotalSeq™-C0050 anti-CD19 (302265, clone HIB19, BioLegend) TotalSeq™-C0154 anti-CD27 (302853, clone O323, Biolegend), TotalSeq™-C0180 anti-CD24 (311143, clone ML5, Biolegend), TotalSeq™-C0389 anti-CD38 (303543, clone HIT2 Biolegend), TotalSeq™-C0138 anti-CD5 (300637, clone UCHT2 Biolegend), TotalSeq™-C0159 anti-HLA-DR (307663, clone L243, Biolegend), TotalSeq™-C0006 anti-CD86 (305447, clone IT2.2, Biolegend), TotalSeq™-C0136 anti-IgM (314547, clone MHM-88, Biolegend), TotalSeq™-C0384 anti-IgD (348245, clone IA6-2, Biolegend), TotalSeq™-C0181 anti-CD21 (354923, clone Bu32, Biolegend) and TotalSeq™-C0831 anti-CD138 (352327, clone DL-101, Biolegend). The scRNA-seq libraries were generated using Chromium Next GEM Single Cell 5’ V(D)J Reagent Kit v.1.1 with Feature Barcoding (10X Genomics) according to the manufacturer’s protocol. Gene expression (mRNA), antibody-derived tags and VDJ BCR libraries were constructed. Briefly, cells were counted and up to 10 000 cells were loaded in the 10X Genomics Chromium Controller to generate single-cell gel-beads in emulsion. After reverse transcription, gel-beads in emulsion were disrupted. Barcoded complementary DNA was isolated and amplified by PCR. Following fragmentation, end repair and A-tailing, sample indexes were added during index PCR. The purified libraries were sequenced on a Novaseq S2 flowcell (Illumina).
ScRNA-Seq data processing
The sequencing reads were demultiplexed and aligned to the human reference transcriptome (GRCh38-2020-A), using the CellRanger Pipeline (v5.0.1). The unfiltered raw UMI counts from cellranger were loaded into Seurat v4.0.4 for quality control, data integration and downstream analyses77. Empty sequencing beads, apoptotic cells and potential doublets were removed by filtering out cells with fewer than 500 features or a mitochondrial content higher than 20%. BCR reads were aligned to the GRCh38 reference genome (vdj_GRCh38_alts_ensembl-5.0.0) and consensus BCR annotation was performed using the cellranger vdj program (v5.0.1). The clonotypes information (all_contig_annotations.csv files) were loaded into the seurat object and only productive contigs of BCR were kept for the analysis. Data from each sample were normalized and scaled using the sctransform method, and batch effect between samples was corrected using Seurat’s FindIntegratedAnchors. On this integrated dataset, we computed the principal component analysis on the 3000 most variable genes. UMAP was carried out using the 30 most significant PCs, and community detection was performed using the graph-based modularity-optimization Louvain algorithm from Seurat’s FindClusters function. Cell type labels were assigned to resulting clusters based on a manually curated list of marker genes. All clusters were annotated, and 78,355 cells were kept for further analysis. Two small clusters with T cell (0.2%) and monocyte (0.1%) signatures were removed from further analysis as well as rare clear doublets. To evaluate the expression of the 10 surface proteins, the antibody-derived tags count for the cells retained after filtering were loaded in the integrated seurat object and normalized using the center log ratio normalization method. Differentially expressed genes between different clusters were identified using the FindAllMarkers function from Seurat using default settings (Wilcoxon test and Bonferroni P value correction). For GSEA analysis, differentially expressed genes were calculated using the Wilcoxon rank sum test via the wilcoxauc function of the presto package using default parameters (including Benjamini-Hochberg false discovery rate correction) and filtered on logFC > 0.5 and padj < 0.01. GSEA was run on pre-ranked genes using the fgsea package implemented in R78.
scBCR-seq data processing
Immune profiling V(D)J data was added to the dataset by matching cell barcodes in the scRNAseq libraries. Cell barcodes that had a BCR heavy or light chain that mapped to “NA” was excluded from downstream analysis, as well as cases of 10X Genomics barcodes with two or more consensus heavy chain sequences. On the BCR data, repertoire abundance and overlap, gene usage, and diversity estimation were performed using the R package immunarch (v0.9.0)79. Mutation quantification and lineage reconstruction was performed on the immcantation framework80 as described by Mathew et al.81. In brief, clones clustering were made with the Change-O toolkit and lineages were reconstructed using the dnapars function of the Phylip package via buildPhylipLineage function82. Sequences that had the same V-gene, same J-gene, and same CDR3 length with maximal length normalized nucleotide hamming distance of 0.2 were considered as potentially belonging to the same clonal group. Mutation frequencies in V genes were then calculated using the calcObservedMutations() function from Immcantation/SHazaM v1.0.2 R package. Circlize package was used for heavy and light chain pairing83.
Statistics
Statistical analyses were performed using Prism v9 (GraphPad). For nonparametric variables, the Mann-Whitney test was used. The χ2 test was used for categorical values. Multiple comparisons were performed using Wilcoxon rank sum test and Bonferroni correction. Numerical values are given by mean ± SEM. A P value of ≤0.05, or <0.01 for multiple comparisons, was considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Single-cell RNA-Seq data were deposited at Gene Expression Omnibus, with the primary accession code: GSE262240. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Wardemann, H. et al. Predominant autoantibody production by early human B cell precursors. Science 301, 1374–1377 (2003).
Nemazee, D. Mechanisms of central tolerance for B cells. Nat. Rev. Immunol. 17, 281–294 (2017).
Meffre, E. & O’Connor, K. C. Impaired B cell tolerance checkpoints promote the development of autoimmune diseases and pathogenic autoantibodies. Immunol. Rev. 292, 90–101 (2019).
Yurasov, S. et al. Defective B cell tolerance checkpoints in systemic lupus erythematosus. J. Exp. Med. 201, 703–711 (2005).
Jenks, S. A. et al. Distinct effector B cells induced by unregulated toll-like receptor 7 contribute to pathogenic responses in systemic lupus erythematosus. Immunity 49, 725–739.e6 (2018).
Elsner, R. A. & Shlomchik, M. J. Germinal center and extrafollicular B cell responses in vaccination, immunity, and autoimmunity. Immunity 53, 1136–1150 (2020).
Turner, J. S., Ke, F. & Grigorova, I. L. B cell receptor crosslinking augments germinal center B cell selection when T cell help is limiting. Cell Rep. 25, 1395–1403.e4 (2018).
Chen, S. T., Oliveira, T. Y., Gazumyan, A., Cipolla, M. & Nussenzweig, M. C. B cell receptor signaling in germinal centers prolongs survival and primes B cells for selection. Immunity 56, 547–561.e7 (2023).
Woodruff, M. C. et al. Extrafollicular B cell responses correlate with neutralizing antibodies and morbidity in COVID-19. Nat. Immunol. 21, 1506–1516 (2020).
Reed, J. H., Jackson, J., Christ, D. & Goodnow, C. C. Clonal redemption of autoantibodies by somatic hypermutation away from self-reactivity during human immunization. J. Exp. Med. 213, 1255–1265 (2016).
Burnett, D. L., Reed, J. H., Christ, D. & Goodnow, C. C. Clonal redemption and clonal anergy as mechanisms to balance B cell tolerance and immunity. Immunol. Rev. 292, 61–75 (2019).
Sabouri, Z. et al. Redemption of autoantibodies on anergic B cells by variable-region glycosylation and mutation away from self-reactivity. Proc. Natl Acad. Sci. USA 111, E2567–E2575 (2014).
Miyakis, S. et al. International consensus statement on an update of the classification criteria for definite antiphospholipid syndrome (APS). J. Thromb. Haemost. 4, 295–306 (2006).
Barbhaiya, M. et al. The 2023 ACR/EULAR antiphospholipid syndrome classification criteria. Arthritis Rheumatol. Hoboken NJ 75, 1687–1702 (2023).
Arad, A., Proulle, V., Furie, R. A., Furie, B. C. & Furie, B. β2-glycoprotein-1 autoantibodies from patients with antiphospholipid syndrome are sufficient to potentiate arterial thrombus formation in a mouse model. Blood 117, 3453–3459 (2011).
Fischetti, F. et al. Thrombus formation induced by antibodies to beta2-glycoprotein I is complement dependent and requires a priming factor. Blood 106, 2340–2346 (2005).
Pengo, V. et al. Incidence of a first thromboembolic event in asymptomatic carriers of high-risk antiphospholipid antibody profile: a multicenter prospective study. Blood 118, 4714–4718 (2011).
Dabit, J. Y., Valenzuela-Almada, M. O., Vallejo-Ramos, S. & Duarte-García, A. Epidemiology of antiphospholipid syndrome in the general population. Curr. Rheumatol. Rep. 23, 85 (2021).
Yin, H. et al. Association of STAT4 and BLK, but not BANK1 or IRF5, with primary antiphospholipid syndrome. Arthritis Rheum. 60, 2468–2471 (2009).
Simonin, L. et al. Lymphocyte disturbances in primary antiphospholipid syndrome and application to venous thromboembolism follow-up. Clin. Rev. Allergy Immunol. 53, 14–27 (2017).
Alvarez-Rodriguez, L., Riancho-Zarrabeitia, L., Calvo-Alén, J., López-Hoyos, M. & Martínez-Taboada, V. Peripheral B cell subset distribution in primary antiphospholipid syndrome. Int. J. Mol. Sci. 19, 589 (2018).
Bashford-Rogers, R. J. M. et al. Analysis of the B cell receptor repertoire in six immune-mediated diseases. Nature 574, 122–126 (2019).
Giles, I. P. et al. A systematic analysis of sequences of human antiphospholipid and anti–β2-glycoprotein I antibodies: The importance of somatic mutations and certain sequence motifs. Semin. Arthritis Rheum. 32, 246–265 (2003).
Lieby, P. et al. The clonal analysis of anticardiolipin antibodies in a single patient with primary antiphospholipid syndrome reveals an extreme antibody heterogeneity. Blood 97, 3820–3828 (2001).
Lieby, P. et al. Memory B cells producing somatically mutated antiphospholipid antibodies are present in healthy individuals. Blood 102, 2459–2465 (2003).
Lieby, P. et al. Pathogenic antiphospholipid antibody: an antigen-selected needle in a haystack. Blood 104, 1711–1715 (2004).
Manfredo Vieira, S. et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 359, 1156–1161 (2018).
Keller, B. et al. The expansion of human T-bethighCD21low B cells is T cell dependent. Sci. Immunol. 6, eabh0891 (2021).
Malle, L. et al. Autoimmunity in Down’s syndrome via cytokines, CD4 T cells and CD11c+ B cells. Nature 615, 305–314 (2023).
Wang, S. et al. IL-21 drives expansion and plasma cell differentiation of autoreactive CD11c hi T-bet + B cells in SLE. Nat. Commun. 9, 1758 (2018).
Janda, A. et al. Disturbed B-lymphocyte selection in autoimmune lymphoproliferative syndrome. Blood 127, 2193–2202 (2016).
Casali, P., Inghirami, G., Nakamura, M., Davies, T. F. & Notkins, A. L. Human monoclonals from antigen-specific selection of B lymphocytes and transformation by EBV. Science 234, 476–479 (1986).
Traggiai, E. et al. An efficient method to make human monoclonal antibodies from memory B cells: potent neutralization of SARS coronavirus. Nat. Med. 10, 871–875 (2004).
McNeil, H. P., Simpson, R. J., Chesterman, C. N. & Krilis, S. A. Anti-phospholipid antibodies are directed against a complex antigen that includes a lipid-binding inhibitor of coagulation: beta 2-glycoprotein I (apolipoprotein H). Proc. Natl Acad. Sci. USA 87, 4120–4124 (1990).
Dieudonné, Y. et al. B cells in primary antiphospholipid syndrome: Review and remaining challenges. Autoimmun. Rev. 20, 102798 (2021).
Vega-Ostertag, M., Liu, X., Kwan-Ki, H., Chen, P. & Pierangeli, S. A human monoclonal antiprothrombin antibody is thrombogenic in vivo and upregulates expression of tissue factor and E-selectin on endothelial cells. Br. J. Haematol. 135, 214–219 (2006).
Meroni, P. L., Borghi, M. O., Raschi, E. & Tedesco, F. Pathogenesis of antiphospholipid syndrome: understanding the antibodies. Nat. Rev. Rheumatol. 7, 330–339 (2011).
Manukyan, D. et al. Cofactor-independent human antiphospholipid antibodies induce venous thrombosis in mice. J. Thromb. Haemost. 14, 1011–1020 (2016).
Li, S. et al. Prevalence and diagnostic value of non-criteria antiphospholipid antibodies for antiphospholipid syndrome in Chinese patients. Front. Immunol. 14, 1107510 (2023).
Pontara, E. et al. Close link between antiphosphatidylserine/prothrombin antibodies, lupus anticoagulant, and activated protein C resistance in tetra antiphospholipid antibody–positive subjects. J. Thromb. Haemost. 21, 3138–3144 (2023).
Tsutsumi, A. et al. IgG subclass distribution of anticardiolipin antibody in patients with systemic lupus erythematosus. J. Rheumatol. 15, 1764–1767 (1988).
Berneman, A., Guilbert, B., Eschrich, S. & Avrameas, S. IgG auto- and polyreactivities of normal human sera. Mol. Immunol. 30, 1499–1510 (1993).
Ortona, E. et al. Vimentin/cardiolipin complex as a new antigenic target of the antiphospholipid syndrome. Blood 116, 2960–2967 (2010).
Ruff, W. E. et al. Pathogenic autoreactive T and B cells cross-react with mimotopes expressed by a common human gut commensal to trigger autoimmunity. Cell Host Microbe 26, 100–113 (2019).
Zuo, Y. et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci. Transl. Med. 12, eabd3876 (2020).
Müller-Calleja, N. et al. Lipid presentation by the protein C receptor links coagulation with autoimmunity. Science 371, eabc0956 (2021).
Tipton, C. M. et al. Diversity, cellular origin and autoreactivity of antibody-secreting cell population expansions in acute systemic lupus erythematosus. Nat. Immunol. 16, 755–765 (2015).
Giles, I. et al. Arginine residues are important in determining the binding of human monoclonal antiphospholipid antibodies to clinically relevant antigens. J. Immunol. 177, 1729–1736 (2006).
Tsuiji, M. et al. A checkpoint for autoreactivity in human IgM+ memory B cell development. J. Exp. Med. 203, 393–400 (2006).
Richardson, C. et al. Molecular basis of 9G4 B cell autoreactivity in human systemic lupus erythematosus. J. Immunol. Baltim. Md 1950 191, 4926–4939 (2013).
Scheid, J. F. et al. Differential regulation of self-reactivity discriminates between IgG+ human circulating memory B cells and bone marrow plasma cells. Proc. Natl Acad. Sci. USA 108, 18044–18048 (2011).
Manning, B. D. & Cantley, L. C. AKT/PKB signaling: navigating downstream. Cell 129, 1261–1274 (2007).
Gaudette, B. T., Jones, D. D., Bortnick, A., Argon, Y. & Allman, D. mTORC1 coordinates an immediate unfolded protein response-related transcriptome in activated B cells preceding antibody secretion. Nat. Commun. 11, 723 (2020).
Luo, W., Weisel, F. & Shlomchik, M. J. B cell receptor and CD40 signaling are rewired for synergistic induction of the c-Myc transcription factor in germinal center B cells. Immunity 48, 313–326.e5 (2018).
Ersching, J. et al. Germinal center selection and affinity maturation require dynamic regulation of mTORC1 kinase. Immunity 46, 1045–1058.e6 (2017).
Cox, E.-M. et al. AKT activity orchestrates marginal zone B cell development in mice and humans. Cell Rep. 42, 112378 (2023).
Schneider, P. The role of APRIL and BAFF in lymphocyte activation. Curr. Opin. Immunol. 17, 282–289 (2005).
King, H. W. et al. Single-cell analysis of human B cell maturation predicts how antibody class switching shapes selection dynamics. Sci. Immunol. 6, eabe6291 (2021).
Languren, M. et al. Characterization of monoclonal anti-beta2-glycoprotein-I and anti-prothrombin antibody fragments generated by phage display from a patient with primary antiphospholipid syndrome. J. Autoimmun. 26, 57–65 (2006).
Agostinis, C. et al. A non-complement-fixing antibody to β2 glycoprotein I as a novel therapy for antiphospholipid syndrome. Blood 123, 3478–3487 (2014).
Taylor, J. J., Pape, K. A. & Jenkins, M. K. A germinal center-independent pathway generates unswitched memory B cells early in the primary response. J. Exp. Med. 209, 597–606 (2012).
Weisel, F. J., Zuccarino-Catania, G. V., Chikina, M. & Shlomchik, M. J. A temporal switch in the germinal center determines differential output of memory B and plasma cells. Immunity 44, 116–130 (2016).
Deenick, E. K. et al. Naive and memory human B cells have distinct requirements for STAT3 activation to differentiate into antibody-secreting plasma cells. J. Exp. Med. 210, 2739–2753 (2013).
Roco, J. A. et al. Class-switch recombination occurs infrequently in germinal centers. Immunity 51, 337–350.e7 (2019).
Long, Y. et al. Follicular helper and follicular regulatory T cell subset imbalance is associated with higher activated B cells and abnormal autoantibody production in primary anti-phospholipid syndrome patients. Clin. Exp. Immunol. 206, 141–152 (2021).
Faliti, C. E. et al. P2X7 receptor restrains pathogenic Tfh cell generation in systemic lupus erythematosus. J. Exp. Med. 216, 317–336 (2019).
Chen et al. Autoreactivity in naïve human fetal B cells is associated with commensal bacteria recognition. Science 369, 320–325 (2020).
Rawlings, D. J., Metzler, G., Wray-Dutra, M. & Jackson, S. W. Altered B cell signalling in autoimmunity. Nat. Rev. Immunol. 17, 421–436 (2017).
Rodríguez-García, M. E. et al. A novel de novo MTOR gain-of-function variant in a patient with Smith-Kingsmore syndrome and Antiphospholipid syndrome. Eur. J. Hum. Genet. 27, 1369–1378 (2019).
Canaud, G. et al. Inhibition of the mTORC pathway in the antiphospholipid syndrome. N. Engl. J. Med. 371, 303–312 (2014).
Jackson, S. W. & Davidson, A. BAFF inhibition in SLE—Is tolerance restored? Immunol. Rev. 292, 102–119 (2019).
Sintes, J. et al. mTOR intersects antibody-inducing signals from TACI in marginal zone B cells. Nat. Commun. 8, 1462 (2017).
Ferri, D. M. et al. Elevated levels of interferon-α act directly on B cells to breach multiple tolerance mechanisms promoting autoantibody production. Arthritis Rheumatol. 75, 1542–1555 (2023).
Verrou, K.-M., Sfikakis, P. P. & Tektonidou, M. G. Whole blood transcriptome identifies interferon-regulated genes as key drivers in thrombotic primary antiphospholipid syndrome. J. Autoimmun. 134, 102978 (2023).
Ugolini-Lopes, M. R. et al. Enhanced type I interferon gene signature in primary antiphospholipid syndrome: association with earlier disease onset and preeclampsia. Autoimmun. Rev. 18, 393–398 (2019).
Gies, V. et al. Identification of autoreactive B cells with labeled nucleosomes. Sci. Rep. 7, 602 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at https://doi.org/10.1101/060012 (2021).
Samokhina, M. et al. immunomind/immunarch: Immunarch 0.9.0. Zenodo https://doi.org/10.5281/zenodo.7446955 (2022).
Vander Heiden, J. A. et al. pRESTO: a toolkit for processing high-throughput sequencing raw reads of lymphocyte receptor repertoires. Bioinforma. Oxf. Engl. 30, 1930–1932 (2014).
Mathew, N. R. et al. Single-cell BCR and transcriptome analysis after influenza infection reveals spatiotemporal dynamics of antigen-specific B cells. Cell Rep. 35, 109286 (2021).
Gupta, N. T. et al. Change-O: a toolkit for analyzing large-scale B cell immunoglobulin repertoire sequencing data. Bioinforma. Oxf. Engl. 31, 3356–3358 (2015).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in. R. Bioinform. 30, 2811–2812 (2014).
Acknowledgements
This study was supported by the PERSONALIS project co-financed by the European regional development fund (ERDF) (to A.S.K., V.G., P.S.S., T.M., R.V., and M.R.), the University of Strasbourg, the Strasbourg University Hospital, the University of Freiburg, the Seed Money program of Eucor – The European Campus (to V.G. and M.R.), the “Fonds de Dotation CSL Behring Pour La Recherche” (to V.G.), the Fondation pour la Recherche Médicale (FRM, grant number FDM201906008804 to Y.D.) and by the German Research foundation (DFG; Project number 468499998 to M.R.). We are grateful to all patients and volunteers who participated in this study. Our appreciation extends to the team at the Microarray Core at UT Southwestern Medical Center in Dallas (TX, USA) for their expert assistance with custom arrays, autoantigen processing, and data analysis. Additionally, we thank the Direction de la Recherche Clinique et de l’Innovation (DRCI) of the Strasbourg University Hospital for their support. Our thanks to BioRender.com to generate under CC-BY license Fig. 2A (Created in BioRender. D, Y. (2024) [www.BioRender.com/k12i608]) and 3A (Created in BioRender. D, Y. (2024) [www.BioRender.com/h87r225]).
Author information
Authors and Affiliations
Contributions
Y.D., P.S., M.R., A.S.K., and V.G. conceived the project and designed the experiments. Y.D., R.L., J.R., I.J., Q.F., L.J., O.V., and V.G. performed and analyzed the experiments. F.C., Z.C., M.L., and M.M. analyzed bioinformatics data. S.D., N.W., S.G., B.N., A.H., R.E.V., A.G., V.P., and T.M. help to perform the experiments. Y.D., M.R., A.S.K., and VG. wrote the manuscript and all authors revised it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Burcu Ayoglu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dieudonné, Y., Lorenzetti, R., Rottura, J. et al. Defective germinal center selection results in persistence of self-reactive B cells from the primary to the secondary repertoire in Primary Antiphospholipid Syndrome. Nat Commun 15, 9921 (2024). https://doi.org/10.1038/s41467-024-54228-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-54228-8
This article is cited by
-
Altered B cell activation contributes to the immunopathogenesis of childhood arthritis-associated uveitis
Nature Communications (2026)
-
B Cell Tolerance and Obstetric Antiphospholipid Syndrome
Clinical Reviews in Allergy & Immunology (2025)
-
Effects of telitacicept in SLE patients with antiphospholipid antibody positivity: a retrospective self-controlled case series
Clinical Rheumatology (2025)