Abstract
Brain organoids have been proposed as suitable human brain model candidates for a variety of applications. However, the lack of appropriate maturation limits the transferability of such functional tools. Here, we present a method to facilitate neuronal maturation by integrating astrocyte-secreted factors into hPSC-derived 2D and 3D neural culture systems. We demonstrate that protein- and nutrient-enriched astrocyte-conditioned medium (ACM) accelerates neuronal differentiation with enlarged neuronal layer and the overproduction of deep-layer cortical neurons. We captured the elevated changes in the functional activity of neuronal networks within ACM-treated organoids using comprehensive electrophysiological recordings. Furthermore, astrocyte-secreted cues can induce lipid droplet accumulation in neural cultures, offering protective effects in neural differentiation to withstand cellular stress. Together, these data indicate the potential of astrocyte secretions to promote neural maturation.
Similar content being viewed by others
Introduction
Organoids modeling the central nervous system (CNS) have emerged as powerful ex vivo models that hold immense potential for advancing understanding of human brain development and neuropsychiatric disorders1,2. Unlike traditional 2D and sub-primate models, brain organoids more faithfully recreate the complex cytoarchitecture, cellular diversity, and multicellular interactions of the environment of the human CNS3,4. Despite offering remarkable advantages over conventional models, human brain organoids (HBOs) generated using contemporary technologies in vitro, still possess remaining challenges, especially in the requirement for long-term culture and promotion of an adequate level of maturity, functionality, and holistic physiological similarities with the human brain5,6,7.
Recent advances in HBOs have been reported by use of novel physical and/or biochemical approaches. Unlike conventional bioreactors and orbital shakers, innovative strategies leveraging mechanobiological regulation, such as mechanical slicing8,9, hyperoxic culture10, and air-liquid interface culture11, have effectively reduced cellular hypoxia and improved cell viability. Furthermore, the integration of advanced biophysical techniques, including microfluidics12, magnetic nanoparticles13, and volumetric compression14,15, has enabled more precise control over the internal microenvironment of organoids, including elevated possession of neural progenitors and functionally differentiated neurons. Complementing these biophysical modifications, the supplementation of growth factors and extracellular matrix components has provided more favorable conditions to support brain cell growth and differentiation. Additionally, the incorporation of diverse cell types including microglia16,17 and endothelial cells18,19, as well as the fusion of patterned organoids, has further enhanced the physiological relevance of HBOs. Moreover, transplanting neural organoids into rodent hosts has shown an unprecedented improved survival of cells and more complex neural connectivity between the organoid and the host brain20,21. Furthermore, whereas great strides have been made in the maturation of brain organoids, the existing approaches employed remain far from satisfactory in their scalability and efficiency. This is largely due to the inherent complexity and time-consuming nature of the current interventions aimed at promoting organoid maturation. Moreover, the limited diversity of the current solutions available further underscores the pressing need for more efficient and innovative strategies to drive this field forward.
Astrocyte-secreted signals are pivotal in the formation, maturation, and plasticity of neural circuits22,23,24. In the presence of astrocytes or astrocyte-conditioned medium (ACM), functional maturation of 2D primary and human pluripotent stem cell-derived neurons is significantly increased25,26. Though these findings suggest that astrocyte-derived factors hold great promise for HBO maturation, the mechanism and effect of promoting HBO maturation using ACM remains relatively unclear. Existing studies have focused primarily on assessing the effects of ACM on specific cell populations within the organoids, without conducting rigorous comparative analyses between organoids cultured with and without ACM27,28. Moreover, these studies have largely lacked comprehensive electrophysiological characterization, limiting the ability to truly validate the functional maturation of these models. It is worth mentioning that the lack of transparency surrounding the composition of commercially available ACM products, making it difficult to assess their precise effects, while raising concerns about the reproducibility and consistency of results.
In this study, to explore the positive effect of astrocyte-secreted cues on promoting cerebral organoid development, an ACM-treated forebrain organoid (MACMO) model with enhanced functional capabilities is presented. The MACMOs obtained the analogous regionalization and formed characteristic neuroepithelial organization with functional neurons as human cortical tissues. Critically, under ACM incubation, the neuronal layer of the organoid was thickened and increased with TBR1+ and CTIP2+ deep-layer cortical neurons, suggesting greater potential to integrate both cortical and extracortical synaptic inputs. Calcium imaging studies evidenced functional recapitulation and augmentation in MACMOs. More importantly, real-time neuronal network activity using MEA was recorded and studied statistically over the course of organoid maturation, revealing that MACMOs gained significantly improved neuronal development, cortical maturation, and function recapitulation. Finally, by omic analysis and lipid staining, the improved neural maturation might be attributed to the accumulation of protective lipid droplets in both neurons and astrocytes of organoids under signaling by astrocyte-secreted cues.
Results
ACM-treated and control forebrain organoids mimics stages of human brain development
A robust method for generating forebrain organoids from hPSCs was developed by comparing and combining previously described differentiation protocols29,30,31 (Supplementary Table 1). Cell aggregates were generated by rapid aggregation in a 96-well ultra-low-attachment plate to obtain homogeneous embryonic bodies29. Dual SMAD and WNT inhibition strategies were employed to enhance cortical identity30,31. Characterizations for various region-specific markers were performed to confirm the regional identity of the organoids (Supplementary Fig. 1). Astrocyte-conditioned medium (ACM) and trophic factors were added to promote neurodifferentiation (Fig. 1a). The ACM was obtained from primary astrocytes of the fetal mouse cortex (MACM) and human astrocytes (HACM) (Supplementary Fig. 2 and Supplementary Fig. 3). The effects of ACM on organoids were evaluated using four PSC lines (H1, H9, DYR, and DXR). Organoid generation and maturation were consistent across all cell lines, demonstrating that the impact of ACM on organoid growth is not limited to specific genetic backgrounds (Supplementary Fig. 4 and Supplementary Fig. 5). The results of S100β+ and GFAP+ immunoreactivity demonstrated the enrichment of astrocytes, while TUJ1-, NEUN- and O4- indicated minimal contamination of neurons and oligodendrocyte progenitor cells (Supplementary Fig. 6). In continuous culture, the dorsal forebrain organoids exhibited a gradual increase in size over a period of 60 days, before reaching a plateau due to the absence of nutrients and oxygen (Supplementary Fig. 4a–d). In contrast, the average size of the ACM-treated organoids (ACMOs) was slightly larger than that of the control organoids, although no significant differences were observed (Supplementary Fig. 4e–h).
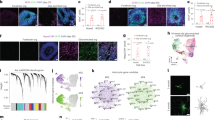
a Schematic diagram of forebrain organoid protocol (Created in BioRender. Yang, H. (2025) https://BioRender.com/r00p936). b Schematic representing the rosette-like cortical region of brain organoids. VZ ventricular zone, MZ marginal zone, CP cortical plate, SVZ subventricular zone, oSVZ outer subventricular zone (created with Biorender.com). c Sample images of immunostaining of MACM conditioned organoids (MACMOs) at day 35 showing proliferating NPCs (NESTIN) and neurons (MAP2). d Confocal image of an immunostained MACMO section at day 45 showing dorsal cortical marker (PAX6). Representative immunostainings showing apical (pVIM) - basal (LAMININ) polarity (e) and oSVZ of day-60 MACMOs. Proliferation marker KI67, NPC marker SOX2, and oRG marker HOPX (f). g Sample images of immunostaining of MACMOs at day 80 showing the rosette structure. IPC marker TBR2, cortical layer 5 neuronal marker CTIP2. h, i Sample images of immunostaining of MACMOs at days 60 and 90, respectively, showing astrogenesis over time. Astrocyte marker, GFAP, and S100β. Magnified views of the boxed region are shown as inset or on the right. Scale bars, (c, d) left 200 μm; (c) inset 100 μm; (d) right 50 μm; (e–i) left, 500 μm; right, 100 μm. The experiments were repeated at least three times independently with similar results. The immunostaining images of forebrain organoids cultured under control conditions are shown in Supplementary Fig. 7.
To characterize the cellular diversity and cytoarchitecture of ACM and control organoids, a series of immunohistochemical staining were performed. At day 35, the organoids consisted mostly of NESTIN+ neural progenitors, which exhibited typical radial glial morphology and neural tube-like structures called neural rosettes32. The MAP2+ neuronal layers were located exclusively away from the surrounding rosettes (Fig. 1c and Supplementary Fig. 7a), indicating the apical-basal polarity of the neuroepithelium. To validate the successful regionalization of organoids, PAX6+ (a frequently used dorsal cortical marker33) and FOXG1+ (a marker for tissues with telencephalon identity34) were stained. These results indicated that the organoids had been correctly regionalized, thereby demonstrating that ACM incubation can reproduce and maintain dorsal forebrain identity (Fig. 1d, Supplementary Fig. 7g, h).
Positive staining for phospho-VIMENTIN (pVIM) during mitosis and for the basement membrane component laminin at day 60 provided further confirmation of the apicobasal organization, which recapitulated the apicobasal orientation of the developing human brain29 (Fig. 1e and Supplementary Fig. 7b). Furthermore, the cell type complexity and cytoarchitecture of the organoids were assessed (Fig. 1b). Cortical regions exhibited evidence of typical progenitor zones, displaying distinct ventricular-like zones (VZ) containing SOX2+ radial glial cells dividing at the apical surface, and TBR2+ intermediate progenitors (IPs) surrounding the VZ (Fig. 1f, g and Supplementary Fig. 7c, d). In addition, the presence of HOPX + SOX2+ cells outside the ventricular zone (VZ) was demonstrated, indicating the presence of outer radial glial cells (oRGs) and the outer subventricular-like zone (OSVZ). Cells that were stained positive for pVIM and the proliferation marker KI67+ indicated the highly proliferative ventricular-like zones in the cortical structure (Fig. 1f and Supplementary Fig. 7c). Further, S100β+ and GFAP+ astrocytes were also observed in the marginal zone of day-60 and day-90 organoids, where neurons settle in (Fig. 1h, i and Supplementary Fig. 7e, f). The astrogliogenesis between MACMOs and control organoids was compared, showing a higher expression of GFAP at days 60–90 with no significance (Supplementary Fig. 8).
The collective data indicated that forebrain organoids treated with astrocyte-conditioned or control medium underwent both neurogenesis and astro-gliogenesis, resulting in the generation of a broad diversity of cells and tissue architecture that was comparable to the developing human brain.
Diverse subtype formation of cortical neurons with functional activity in MACMOs and control organoids
The development of neural circuits involves the wiring and generation of diverse neuron subtypes in the cortical plate, allowing the cortex to integrate and coordinate various information, accomplishing sophisticated information processing. To investigate the neuronal subtypes of the forebrain organoids, immunostaining was performed to examine the detailed expression of six cortical layer markers (Supplementary Fig. 9a). At day 35, neurons expressing the deep-layer cortical neuron markers TBR1 were observed to be predominantly distributed outwardly, as well as neurons expressing the Cajal-Retzius cell marker REELIN (Supplementary Fig. 9b, c). The stereotypical cortical plate architecture was generated in an inside-out order, with early-born neurons populating the deep layers and later-born neurons populating the more superficial layers. To ascertain whether this layer organization was recapitulated in our organoids, layer V cortical neuron marker CTIP2, layer IV cortical neuron marker SATB2, and layer III/II cortical neuron marker BRN2 and CUX1 were stained in day-90 organoids. Observation of CTIP2+ neurons revealed their predominant location in the deeper part of the cortical plate. In contrast, neurons expressing SATB2, CUX1, and BRN2 were observed to be located near the pial surface (Supplementary Fig. 9d–k). These findings suggest that both MACMOs and control organoids had formed the structure of six cortical layers.
Spontaneous and drug-treated neural activity of the organoids was examined utilizing calcium imaging (Fig. 2a, b). Calcein-AM staining and bright-field imaging demonstrated the viability and formation of neuronal networks in the confocal plate (Supplementary Fig. 10a, b). At day 45, spontaneous, asynchronous calcium surges were observed in the MACMOs and control organoids. Subsequently, more frequent and synchronous spikes were observed at the calcium activity of day-90 and day-120 samples, resembling the network activity patterns observed in developing neocortical circuits (Fig. 2c and Supplementary Fig. 10c). The activity of glutamatergic and GABAergic synaptic transmission was evaluated in the MACMOs and control organoids (Supplementary Figs. 11 and 12). To further validate the physiological correctness of the MACMOs and control organoids, the presence of exogenous glutamate resulted in an increase of calcium spikes and synchronized events (Supplementary Fig. 11b). Furthermore, the application of glutamate receptor antagonists, APV and CNQX (NMDA and AMPA receptors, respectively), resulted in a reduction in neural activity (Supplementary Fig. 11c). Conversely, calcium activity was found to be decreased by the administration of GABA (Supplementary Fig. 12a), whereas the activity was significantly upregulated by the addition of saclofen (GABA-B receptor) and SR-95531 (GABA-A receptor) (Supplementary Fig. 12b, c). Quantitative analysis revealed an elevated ratio of VGLUT1+ glutamatergic excitatory neurons in the experimental group compared to the control ones, while GABA+ inhibitory neurons were at comparable levels across both conditions (Supplementary Fig. 13).
a Generation and air-liquid culture of forebrain organoid slices for calcium imaging (Created in BioRender. Yang, H. (2025) https://BioRender.com/r36w725). b Sample images of Fluo-4 AM of a forebrain organoid and acquisition of regions of interests (ROIs) illustrating the identification of active neurons. Activity profiles of all active neurons are shown as normalized ΔF/F values and colorized amplitude plots. Activities of one spiking neuron (blue) and a multi-spiking neuron (red) are identified and visulaized. Scale bar, 100 μm. c Calcium imaging of MACMOs on days 45, 90, and 120. Activity profile is shown as normalized ΔF/F values and colorized amplitude plot. The experiments were repeated at least three times independently with similar results. The calcium imaging of forebrain organoids cultured under control conditions is shown in Supplementary Fig. 6.
These findings indicated that the organoids developed structured and functionally interconnected neural networks over the culture duration, in which glutamatergic and GABAergic receptor activity plays important roles in modulating network activity.
Enlarged neuronal layers and increased deep-layer neurons in MACMOs over 30-90 days
The previous data demonstrated appropriate lineage specification of MACMOs, but did not investigate which specific cell types were affected by ACM. The area and relative thickness of SOX2+ progenitor zones and TUJ1+/MAP2+ neuronal layers were quantified over time (Fig. 3a). The ratio of area and relative thickness between the progenitor zone and neuronal layer in the cortical region decreased along with the culture period for each condition, indicating the process of progenitor cells differentiating into neurons (Fig. 3b, c). At day 60, the MACMOs exhibited a lower progenitor-to-neuron ratio but with no significant differences. However, a significantly lower ratio of thickness between the progenitor zone and neuronal layer was observed in the MACMOs compared to the control. By day 90, the MACMO organoids demonstrated a significantly lower ratio of progenitors-to-neurons, coupled with a reduced relative thickness ratio of the progenitor zone-to-the neuronal layer (Fig. 3b, c). Furthermore, it was observed that the layer thickness of the epithelium and neuronal layer were numerically increased in MACMOs at day 90 (Fig. 3d).
a Sample immunostaining images of organoids against TUJ1, SOX2, and MAP2 at days 60 and 90. Created in BioRender. Yang, H. (2025) https://BioRender.com/b20n239. b SOX2 and TUJ1 area ratio within the cortical region of organoids. each point represents one cortical region from organoids. Day-60: MACM n = 19; CTR n = 23; day-90: MACM n = 36; CTR n = 33. Progenitor-to-neuronal layer thickness ratio (c) the layer thickness of neuronal and total layer (d) within cortical regions of MACMOs and control organoids at days 60 and 90. c, d Each point represents one cortical region from organoids. Day-60: MACM n = 23; CTR n = 16; day-90: MACM n = 31; CTR n = 32. e Sample images of organoids stained against CTIP2, TBR1 at days 30, 60, and 90. f–i Percentage of CTIP2+ and TBR1+ cells in the cortical region or cortical plate of organoids. Each point represents one cortical region or cortical plate from one organoid. f Day-30: MACM n = 23; CTR n = 22; Day-60: MACM n = 16, CTR n = 14; Day-90: MACM: n = 26; CTR n = 30. g Day-30: MACM n = 33; CTR n = 31; Day-60: MACM n = 29; CTR n = 22; Day-90: MACM n = 64; CTR n = 53. h Day-30 MACM: n = 23; CTR n = 22; Day-60: MACM n = 16; CTR n = 12; Day-90: MACM n = 26; CTR n = 30. i Day-30: MACM n = 28; CTR n = 20; Day-60: MACM n = 28; CTR n = 16; Day-90: MACM n = 64; CTR n = 52. j, k Layer thickness of TBR1+/CTIP2+ cells at day-90 organoids. Each point represents one cortical plate column from one organoid. TBR1+: MACM: n = 81; CTR n = 79; CTIP2+: MACM, n = 81; CTR n = 70. The data are represented as mean ± SEM. a and e are created with BioRender.com. Significant difference is determined by two-tailed Mann–Whitney U test: (b–d) day90 total layer, (g) day60 and day90, (j, h) day30, (i) day90, (k); two-tailed unpaired t test: (d) day60, (f) day30 and day60, (g) day30, (h) day60 and day90, i day60; two-tailed unpaired t test with welch correction: (f) day60. *P < 0.05, **P < 0.01, ***P < 0.001; ****P < 0.0001. Source data are provided as a Source Data file.
To identify the expanded neuronal populations, the markers of distinct cortical neurons were analyzed in MACMOs. Cell counts at days 30, 60, and 90 demonstrated that the ratios of TBR1+ and CTIP2+ cells in both the cortical region and cortical plate of the MACMOs were significantly higher than the control group, except for the TBR1+ cells in day-30 control organoids (Fig. 3 f–i). This discrepancy may be attributed to the fact that, at day 30, the early stage of differentiation, a greater number of CTIP2+ layer V cortical neurons were generated in the MACMOs than in the control group. This resulted in a lower ratio of TBR1+ layer VI neurons in the cortical region in MACMOs, indicating that ACM treatment may accelerate the process of neuronal differentiation.
In parallel, organoids were stained for SATB2 and CUX1 at days 60 and 90 (Supplementary Fig. 14a) for the upper cortical layers. Similarly, the thickness of the SATB2+ and CUX1+ upper layer zone exhibited an increase throughout the culture period (Supplementary Fig. 14b, e). Results showed that the length and ratio of CUX1+ at days 60, 90 in the cortical region and cortical plate did not exhibit significant differences between MACMOs and the control group (Supplementary Fig. 14b–d). The day-60 MACMOs exhibited a significantly greater length and a higher ratio of SATB2+ cells in the cortical region compared to the control group, while there was no significant difference in the ratio of SATB2+ cells in the cortical plate (Supplementary Fig. 14e–g). The limited supply of nutrients in the non-vascularized organoids resulted in the discontinuous generation of upper-layer neurons, which may account for the loss of differences between the groups at the late culture stage, e.g. days 60-90.
To ensure the physiological relevance of our findings, we conducted parallel experiments comparing the effects of HACM and MACM on neural progenitor cells (NPCs) and forebrain neuron precursors (Supplementary Fig. 15). Both HACM and MACM promoted neuronal differentiation, as indicated by increased MAP2 signals, while cell proliferation and apoptosis remained unaffected (Supplementary Fig. 16a–d). In maturing forebrain neuron precursors into neurons, MACM demonstrated a more pronounced effect, showing higher MAP2 + /DAPI+ ratios and reduced apoptosis (Cleaved Caspase 3+/DAPI+) compared to HACM (Supplementary Fig. 16e–h). Both conditioned media increased the proportion of deep-layer neurons (CTIP2+), but the effect was more significant with MACM, resulting in greater neuronal density (Supplementary Fig. 17). Proteomic analysis revealed a high degree of similarity between HACM and MACM, with 2001 shared proteins identified (Venn diagram) (Supplementary Fig. 18). Further analysis of the top 50 most abundant proteins in each sample showed a significant overlap, as illustrated in the protein linkage map (Supplementary Fig. 19). These results suggest the conserved molecular components in both HACM and MACM, supporting their shared biological effects on neuronal maturation while explaining the stronger impact observed with MACM.
These results highlighted the beneficial impact of ACM treatment on neural development within the organoid model. Organoids exposed to ACM exhibited a significant increase in the thickness of neuronal layers, accompanied by a reduction in progenitor-to-neuron ratios and relative thickness ratios. Notably, this was coupled with an increase in the proportion of deep-layer projection neurons, a critically important neuronal subtype for cortical circuit formation and function.
MACMOs showed augmented excitability and more frequent firing in calcium activity
Astrocyte-derived signals have been demonstrated to play a pivotal role in regulating neural circuit development and function in 2D neurons in vitro26,35,36,37,38. Here, we aim to evaluate whether similar effects exist in 3D neural cultures and to determine whether the thickened neuronal layer and increased deep-layer neurons observed in MACMOs result in any functional changes or improvements in neuronal activity and information processing. The data indicated that MACMOs exhibited slightly elevated neural activity, yet no discernible statistical discrepancies were observed in the calcium dynamics of day-45 MACMOs in comparison with control organoids (Supplementary Fig. 20).
In contrast, statistically significant differences were observed between day-90 MACMOs and control organoids (Fig. 4a). Notably, a greater number of spikes were observed in all spiking and multi-spiking neurons in day-90 MACMOs (Fig. 4b, c). The factors that contribute to the enhancement of the calcium transients were therefore investigated. Firstly, the numbers of spiking neurons (i.e., cells spike at the recording period) and multi-spiking neurons (i.e., a subset of spiking neurons, cells that spike at least three times during one minute) in MACMOs were significantly higher than the control (Fig. 4d, e). Furthermore, it was observed that individual spiking neurons in ACM-treated organoids exhibited a higher frequency of firing compared to the control, whereas for individual multi-spiking neurons, no significant differences were observed between MACMOs and the control (Fig. 4f, g). The mean and peak amplitudes of one multi-spiking neuron in MACMOs were found to be higher than those in control organoids, indicating an enhanced dynamic range of Fluo-4 fluorescence in MACMOs (Fig. 4h, i). Taken together, whereas the day 45 MACMOs exhibited a slight increase in electrical activity, the day 90 MACMOs exhibited augmented excitability and a higher frequency of firing.
a Normalized ΔF/F colorized amplitude plot and sample trace of spiking and multi-spiking neurons of day-90 MACMOs and control organoids. b–i Quantification of calcium activities of day 90 organoids of different conditions. Number of spikes (b) in spiking neurons (c) in multi-spiking neurons. d, e Number of cells (d) in spiking neurons (e) in multi-spiking neurons. Number of spikes per cell (f) in spiking neurons (g) in multi-spiking neurons. Normalized mean (h) and peak (i) amplitude of multi-spiking neurons. The data are represented as mean ± SEM. Each point represents 5 mins recording from one organoid. n = 32 MACMOs, and n = 33 control organoids; at least 4 organoids per differentiation; Significant difference is determined by two-tailed Mann–Whitney U test; *P < 0.05, **P < 0.01, ***P < 0.001; ****P < 0.0001. Source data are provided as a Source Data file.
These findings also suggested that astrocyte-secreted factors can influence the functionality of organoids, in a manner analogous to the previously reported enhanced synaptic efficacy observed in 2D neuron cultures35,36. Moreover, HBOs possess a 3D microenvironment and cellular composition that is more akin to the in vivo condition, thereby better preserving and recapitulating the genuine physiological characteristics of neurons compared to 2D cultures. Whereas the effects observed are subtle, their accumulation and synergistic impact over time ultimately lead to observable changes in the overall neuronal functionality within the organoid system.
ACM accelerated the formation of more active and synchronous human neuronal circuits via MEA recording
Although calcium imaging offers a valuable tool for visualizing the spatiotemporal patterns of neuronal activation, this technique relies on indirect measures of electrical activity. Multi-electrode array (MEA) provides a non-invasive, direct, and more physiologically relevant measurement of extracellular field potentials of the forebrain organoids. Comparative experiments using both intact and sliced organoids for MEA recordings indicate that more robust and reliable recording of electrical activity achieved on sliced organoids, which is consistent with the previous report39. On day 45, the organoids were sliced into 500 µm sections and plated per well in six wells of a MEA plate for adherent culture with neurite outgrowth. The system enabled rapid and precise capturing of the neuronal network dynamics across different scales from the overall network level, down to the activity of individual electrodes and even single neurons. Specifically, key electrophysiological parameters were visualized, including the amplitude of spikes (μV), the spiking rate (s), raster plots depicting spike patterns, and the waveforms of the recorded spikes (Fig. 5a). Besides, LFP and drug treatment to day-60 organoids showed response to glutamate and gaba can be assessed in our system (Supplementary Fig. 21).
a Schematic representation of the electrical activity feature plots from the MEA recording software (Created in BioRender. Yang, H. (2025) https://BioRender.com/q32h063). b Example spike raster plots from one well, raw traces, and spike waveform, recorded from single electrodes in connection with MACMOs and control organoids at the early stage (days 45–60) and the late stage (days 61–90). Quantification of electrophysiological activity of the number of (c) spikes, (d) bursts, (e) network bursts, (f) active electrodes, (g) bursting electrodes, and (h) electrodes participating in bursts from forebrain organoids with MEA recording in 5 minutes. The data are represented as mean ± SEM. Each dot represents 5-min recording from one organoid. N = 27 and 99 recordings at early stages and late stages for each condition, respectively; Significant difference is determined by two-tailed Mann–Whitney U test; *P < 0.05, **P < 0.01, ***P < 0.001; ****P < 0.0001. Source data are provided as a Source Data file.
Qualitative analyses revealed that notable spiking activity, though featuring fewer electrode bursts or synchronized network bursts, was observed in both MACMOs and control organoids at an early stage (days 45–60). Notably, an increased spiking frequency and a more synchronously active neuronal network were observed in the late-stage cultures (days 61–90), in line with previously reported findings (Fig. 5b)39. These findings demonstrated the feasibility of monitoring the spontaneous electrophysiological activity in MEAs over an extended period, with increased network activity observed as the organoid increasingly matured.
Subsequently, the quantitative electrophysiological assessments demonstrated the striking functional improvements in the MACMOs. Differences in functionality between MACMOs and control organoids were further distinguished using parameters including neuronal spiking, electrode bursts, and network bursts of organoids at early and late stages. Across multiple developmental stages, the MACMOs exhibited a significantly elevated number of spikes, a greater number of active electrodes, and more pronounced electrode and network bursting events (Fig. 5c–e). Notably, the augmented bursting characteristics in the MACMOs, including increased numbers of bursting electrodes and electrodes engaged in synchronized bursts, substantiated the enhanced connectivity and functional integration of the neural networks within these organoids (Fig. 5f–h).
Together, these findings suggest the advantageous effects of ACM on the functional maturation of organoids, providing corroboration for the differences observed in both immunostaining features and calcium activity between MACMOs and control organoids (Figs. 3 and 4). By promoting more robust neuronal activity patterns and strengthening network-level connectivity, ACM treatment appears to accelerate the development of functional neural circuitry in organoid models.
scRNA-seq reveals the correct organoid cellular diversity and the enhanced maturation of layer neurons of MACMOs
To identify cell-type-specific transcriptomic changes induced by ACM, forebrain organoids were profiled on day 90 through 10× genomics chromium single-cell RNA-Seq (scRNA-Seq). Unbiased clustering of cell populations was conducted using principal component analysis (PCA) with highly variable genes as input, and the results were visualized using Uniform Manifold Approximation and Projection (UMAP) (Fig. 6a). All cells were categorized into seven distinct clusters (Fig. 6b). The combined analysis of the highly differentially expressed genes (DEGs) of each cluster and the expression of known cell-type markers was employed to annotate the cluster identity (Fig. 6c, Supplementary Fig. 22d–f). Gene Ontology (GO) analysis revealed that the differentially expressed genes (DEGs) of the cell clusters were significantly enriched in biological processes, including forebrain development, glial cell differentiation, and neurogenesis (Supplementary Fig. 22c). Pseudotime analyses also revealed a distinct progenitor-to-neuron trajectory, from the dividing progenitor to the excitatory lineage in organoids (Supplementary Fig. 23i, j). Moreover, the density plot of layer neurons illustrated that deep- and upper-layer neurons in MACMOs are more mature than control organoids (Supplementary Fig. 23k). In conclusion, these data indicated that both MACMOs and control organoids underwent a correct trajectory of cell diversification and enhanced maturation in layer neurons in MACM treated group, which is in line with our previous results.
a Schematic illustrating the single-cell transcriptomic analysis of MACMOs and control organoids at day 90. b UMAP plot of the integrated datasets colored by seven cell clusters 0–6: brown as dividing cells; dark blue as radial glial cells; purple as astrocyte; green as intermediate progenitor cells (IPCs); and light blue as deep layer neurons; pink as upper layer neurons; yellow as neurons. Separate UMAP plots of different conditions are shown on the Supplementary Fig. 22. c Dot plot shows the rational of cluster annotation based on cell-type specific markers. d GO term analysis of differentially expressed genes (DEGs) in layer neurons with an adjusted p value cutoff of 0.05 showing neurodevelopment and synaptic related pathway altered between MACMOs and control organoids. DEGs are shown at Supplementary Fig. 23f. e GSEA analysis of GO terms related to the synaptic and axon development pathway in astrocytes between MACMOs and control organoids. a, b, d and e are created in BioRender. Yang, H. (2025) https://BioRender.com/g67p106.
Subsequently, gene set enrichment analysis of DEGs between neurons in MACMOs and the control group identified biological processes linked to neurodevelopment pathways, including forebrain development, neuron migration, and regulation of neurogenesis (Fig. 6d). Similarly, gene set enrichment analysis (GSEA) of GO terms in astrocytes identified significant enrichment in pathways related to synapse-related terms in MACMOs, like axon development, postsynaptic membrane and neuron to neuron synapse (Fig. 6e). The pathway analysis provided further supports for the enhanced functional capabilities observed in the MACMOs (Figs. 4 and 5).
Lipid droplet accumulation in neurons and astrocytes of MACMOs
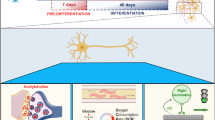
It is noteworthy that further analysis of scRNA-seq data revealed that treatment with ACM altered the transcriptomic signature of astroglia and intermediate progenitor cells, with significant enrichment in biological pathways related to lipid metabolic processes (Fig. 7a, b). In particular, the enriched pathways included phospholipid and glycerolipid metabolic process, and cholesterol transport. This further indicated that ACM treatment influenced the metabolic state and signaling pathways of astrocytes, which may be closely related to their regulatory functions in neural development.
GSEA of GO terms related to the lipid metabolism in astrocytes (a) and radial glial cells (b) between control and ACMOs. c Sample confocal images of day-60 ACMOs and control organoids, for immunostaining of LipidSpot488, GFAP, and TUJ1 at day 60. Scale bar, 20 μm. d, g Quantification of the number and average size of lipid droplets per field in organoids. e, f, h, i Quantification of the number and average size of lipid droplets per field colocalized with TUJ1+ or GFAP+ cells in organoids. The data are represented as mean ± SEM. Each point represents one region of interest of one day 60 organoid section. N = 9 replicates for each condition; Significant difference is determined by two-tailed unpaired t test for (e, g–i); two-tailed unpaired t test with welch correction for (d); two-tailed Mann–Whitney U test for (f). *P < 0.05, **P < 0.01, ***P < 0.001. a–c are created in BioRender. Yang, H. (2025) https://BioRender.com/p79r180. Source data are provided as a Source Data file.
Transcriptional changes in the cholesterol metabolic process pathways were also observed among neurons of MACMOs and control organoids (Fig. 6e). Given the critical role of lipids in supporting neuronal function and energy metabolism, we carried out further investigations to closely examine the lipid profiles of the organoids to better elucidate the metabolic alterations induced by ACM. In particular, our analysis focused on the lipid droplets (LDs), which have been identified as a key indicator of metabolic alterations. LDs play a vital role in the regulation of brain energy homeostasis and are specialized for energy storage40. The co-localization of LipidSpot488, GFAP, and TUJ1 staining demonstrated the presence of LDs in both neurons and astrocytes of MACMOs (Fig. 7c). These results provide further evidence for the presence of LDs in neurons, which challenges the prevailing view that LDs are absent in neurons41. The analysis of the colocalization of LDs with neurons and astrocytes revealed that astrocytes exhibited a greater accumulation of LDs compared to neurons (Fig. 7e, f), in accordance with previous reports indicating that neurons have a lower storage capacity for neutral lipids than astrocytes42.
It is noteworthy that a significant increase in the number and size of LDs was observed in both neurons and astrocytes of MACMOs (Fig. 7h–i). Generally, the accumulation of LD plays an antioxidant role during development or hallmarks of cellular stress and degeneration. Organoids inherently experience a degree of cellular stress, and our morphological analyses revealed that ACMO organoids are slightly larger than control organoids (Supplementary Fig. 4), suggesting that ACMO organoids may develop more extensive zones of oxygen and nutrient deficiency. Additionally, ACMOs exhibited an enhanced neuronal activity (Figs. 4 and 5), which—according to the mechanism proposed by researchers—could potentially trigger higher lipotoxicity43,44. Surprisingly, our scRNA-seq analysis revealed that the ACM-treated group exhibits a lower proportion of stressed cells compared to the control group (Supplementary Fig. 23a–d). Further scRNA-seq analysis focusing on markers associated with hypoxia, glycolysis, and ER stress pathways demonstrated that ACM-treated organoids displayed lower expression of stress markers, indicating reduced cellular stress (Supplementary Fig. 23e, l, m). Moreover, wild-type organoids younger than 150 days are less likely to recapitulate disease phenotypes. In our study, both astrocyte cultures and the PSC-derived organoids for analysis were maintained under normal, non-pathological conditions and were younger than 150 days, suggesting a low possibility of expressing neurological diseaserisk genes. These data suggested that the LD accumulation observed in ACMOs likely represents a neuroprotective and developmental adaptation rather than a manifestation of disease.
To further investigate the role of ACM in LD regulation, we analyzed LD distribution in NPCs and forebrain neuron precursors cultured with MACM-, HACM-supplemented or control differentiation medium. Lipid staining combined with immunofluorescence markers revealed significant increases in LD accumulation in ACM-treated group compared to controls (Fig. 8). Both MACM and HACM treatments significantly increased the total number of lipid droplets in NPC compared to control groups (Fig. 8b, c). Quantification of LDs colocalized with auto-differentiated GFAP+ cells showed a significant increase in MACM-treated groups, but not in HACM-treated cells, suggesting that murine-derived factors may enhance lipid metabolism in astrocytes (Fig. 8d). While both MACM and HACM promoted LD accumulation, MACM showed a slightly stronger effect in differentiated forebrain neurons(Fig. 8e, f). Similarly, the number of LDs colocalized with TUJ1+ neurons and GFAP+ astrocytes were significantly elevated in ACM-treated forebrain neurons compared to controls (Fig. 8g, h), indicating the involvement of LDs in neuronal function and maturation.
a Schematic diagram illustrating the treatment of NPCs and forebrain neurons with different culture media, followed by immunocytochemistry. b, e Sample confocal images of NPCs or forebrain neurons, for immunostaining of lipid droplets marker, LipidSpot; astrocyte marker, GFAP; neuron marker, TUJ1. Scale bar, 20 μm. c, d, f, g, h Quantitative analysis of lipid droplets number and the number of lipid droplets colocalized with GFAP+ or TUJ1+ cells. The data are represented as mean ± SEM. N = 9 replicates for each condition; The significant difference is determined by one-way ANOVA, followed by Tukey’s test. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. a, b and e are created in BioRender. Yang, H. (2025) https://BioRender.com/j55r024. Source data are provided as a Source Data file.
To further investigate the function of lipid droplets, immunostaining for apoptosis and oxidative stress markers showed that ACM treatment reduced apoptosis (indicated by lower cleaved-caspase3 levels) and increased TRX levels in forebrain neuron differentiation with no significance in SOD1 levels, suggesting a neuroprotective effect (Supplementary Fig. 17g and Supplementary Fig. 24a–c). No significant changes were observed in NPCs (Supplementary Fig. 24d–g). These findings are consistent with Ramosaj et al.42, which demonstrated that LD availability supports neural differentiation of neural stem/progenitor cells.
Collectively, it is indicated that the protective rather than cellular stress or pathological nature of LDs in ACM-treated organoids and 2D neural cultures.
Discussion
The findings herein underscore the influence of astrocyte-secreted factors on neuronal wiring and functional maturation of brain organoids, as evidenced by the calcium imaging and MEA recording. The formation of interconnected neural circuits is a remarkable feature of neural development. Abnormalities like circuit formation, functionality, and refinement are known to be involved in many neuropsychiatric diseases3,45,46. Therefore, reliably assessing the electrical functions and large-scale network dynamics of these intricate neural circuits becomes especially critical. Nonetheless, probing network activity is absent in current attempts to apply ACM to mature brain organoids. We highlight that MACMOs displayed an enhancement of electrophysiological network activity. The multifaceted analysis, combination of calcium imaging and MEA recording, provides ways to investigate the physiology of neurons and circuits during the process of formation, maturation, and wiring.
Furthermore, the exact effect of ACM on HBOs has remained elusive. Previous studies introduced ACM for maturing organoids, focusing only on neurons or astrocytes, showing the potential of ACM to mature neurons of hypothalamic arcuate organoids or to accelerate glial differentiation in cortical organoids27,28. Yet, the exact impact of ACM on neurodifferentiation and the structure of forebrain organoids has not been investigated, nor have side-by-side validations been included. Our study provides detailed characterizations of the long-term effects of ACM on forebrain organoid maturation encompassing structure and cellular traits. Surprisingly, ACM exposure promotes the thickening of the neuronal layer and lowers the progenitor-to-neuron ratio, leading to an overproduction of deep-layer neurons. This implies that ACM treatment could enhance the developmental quality of brain organoid tissue. Although brain organoids contain endogenous astrocytes, their number and functionality are far less developed compared to fetal brain conditions, underscoring the critical role of ACM in promoting neuronal maturation and enhancing network formation.
It is well-known that interior hypoxia and nutrient deficiency due to insufficient surface diffusion impede the exploitation of HBOs in translational applications, as well as developmental and disease studies8,47. Studies have confirmed that the lack of oxygen triggers widespread metabolic changes and cellular stress in brain organoids, including hypoxia, oxidative stress, anaerobic glycolysis, and endoplasmic reticulum (ER) stress48. Under hypoxia and metabolic stress conditions, LDs in the brain commonly form and serve a neuroprotective role to alleviate cellular stresses and maintain energy homeostasis49. Interestingly, our results show ACM incubation elevated the level of LDs in both neurons and astrocytes. Studies indicate the over-activation of glycolysis in brain organoids not only impairs certain cell-type specifications but also disturbs energy production in organoids47,48,50,51. Neural stem cells primarily rely on aerobic glycolysis to meet their energy needs, whereas mature neurons predominantly depend on oxidative phosphorylation52. The disrupted balance between glycolysis and oxidative phosphorylation affects the process of neuronal differentiation53. Our results align with findings from Ramosaj et al.42, which demonstrated that increased lipid droplet availability positively affects neural stem/progenitor cell metabolism and neuronal differentiation. The neuroprotective effects mediated by ACM and LD formation were explored through scRNA-seq and proteomic analysis. Key genes involved in lipid metabolism and cholesterol homeostasis pathway were analyzed, such as APOE, LDLR, ABCA1, SOAT1, and HMGCR. Results suggested that astrocyte-secreted cues might regulate LD dynamics and cholesterol metabolism via cholesterol esterification, efflux, and biosynthesis suppression54. A proposed mechanism involves 25-hydroxycholesterol (25HC), which enhances cholesterol efflux and increases extracellular APOE levels in astrocytes. While microglia-mediated activation may not occur in our organoids, ACM likely mimics this pathway, highlighting the neuroprotective role of lipid droplets in cholesterol sequestration and reuptake. The accumulation of LDs in MACMOs suggests a supportive mechanism of ACM to provide a better energy supply for maintaining the energy balance during development. Moreover, during periods of enhanced activity, LDs form in astrocytes to protect neurons from oxidative stress and lipotoxicity44,55.
Unexpectedly, our findings suggest that MACM may offer certain advantages over HACM in accelerating neuronal maturation and network development. For instance, murine astrocytes have been reported to exhibit earlier glutamate sensitivity56 and support dendritic development and stability, potentially through species-specific NMDA receptor interactions. Additionally, MACM production tends to be faster and more scalable than HACM, which may make it a practical option for certain applications, such as disease modeling. In addition, we compared commercially available ACM with our custom-made ACM. SDS-PAGE analysis revealed notable differences between the protein profiles of the commercial ACM and our isolated ACM. Specifically, the commercial ACM exhibited a higher enrichment of proteins within certain molecular weight ranges, whereas the protein composition in our isolated ACM was more evenly distributed. We found that the enriched protein bands are not proteins secreted by human astrocytes, but rather components added to the astrocytes, specifically AGS (Astrocyte Growth Supplement). Besides the pure basal medium of astrocyte medium also contains unknown proteins (Supplementary Fig. 25). Taken together, the presence of AGS and proteins in basal medium of astrocyte medium introduce new variables, making it more difficult to identify the effective astrocyte-secreted factors. The use of commercial ACM becomes impractical, as it is challenging to discern whether the observed effects are due to astrocyte-secreted factors or the added factors themselves.
Moreover, it is important to acknowledge certain limitations that may impact the interpretation and generalizability of our findings. Firstly, due to constraints in the availability of human astrocytes, we utilized MACM in our experiments. This interspecies approach introduces potential discrepancies arising from species-specific differences in astrocyte-secreted factors. While previous studies have demonstrated the efficacy of rodent ACM in influencing human neural cells, the translational relevance of our findings could be affected by these interspecies variations. Future studies employing hPSC-derived astrocyte-conditioned media or mix-species co-culture studies would be essential to further validate and extend our observations. Secondly, the inclusion of serum in our culture media presents another limitation, which introduces additional variables for future applications. The presence of serum may obscure the specific contributions of astrocyte-secreted factors to neural maturation and activity. However, serum-free conditions significantly impaired astrocyte viability, making it challenging to produce sufficient ACM for the experiments (Supplementary Figs. 26 and 27). Given the substantial amount of ACM required to sustain long-term organoid cultures, serum-free conditions were not feasible for our large-scale experiments. Further investigation utilizing serum-free, chemically defined media would allow for more precise delineation of the mechanisms and discovery of the exact molecular agents, such as secreted proteins and exosomes etc., driving the observed effects of astrocyte-secreted factors.
In summary, our study contributes to the understanding of astrocyte-neuron interactions and provides a perspective for optimizing organoids to achieve advanced neural-network maturation. Addressing the aforementioned limitations in future research will be vital for strengthening the utility of HBOs as powerful platforms for studying neurodisorders and advancing the translational potential of these findings.
Methods
Ethical regulations
Our research complies with all relevant ethical regulations. All the acquisitions of human cells were approved by the Ethics Committee at Tsinghua Shenzhen International Graduate School, Tsinghua University [Project 2022/123]. Approved ethics for cell lines are provided by the respective cell banks providing the cells, Wa01 (NIHhESC-10-0043) from WiCell, Wa09 (NIHhESC-10-0062) from WiCell, DXR0109B cell line from ATCC and DYR0100 cell line from the National Collection of Authenticated Cell Cultures, and the primary human astrocytes from ScienCell Research Laboratories. All the animal procedures were approved by the Ethics Committee at Tsinghua Shenzhen International Graduate School, Tsinghua University [Project 2022/123, Project 2024/F021].
Human pluripotent stem cell culture and generation of cerebral organoids
Human H9 ES (WA09) cells, Human H1 ES(WA01) cells (WiCell Agreement No. 22-W0390), DXR0109B hiPSC cells (ATCC ACS-1023TM 70023390), DYR0100 hiPSC cells (National Collection of Authenticated Cell Cultures, Catalog# SCSP-1301) were maintained in MTESRTM Plus (STEMCELL Technologies, catalog # 100-0276) on Matrigel (Corning, catalog number 354277) coated plates and passaged once a week using Relesr (STEMCELL Technologies, catalog # 100-0483). Material transfer agreements (MTAs) for PSCs lines have been duly executed. Dorsal forebrain organoids were generated step by step57. In brief, 70%- 90% confluent cells were dissociated using accutase (Sigma-Aldrich, catalog number A6964) and plated into ultra-low-attachment 96 U-bottomed-well plates (Corning, catalog number 7007) in MTESRTM Plus medium supplemented with 50um Y27632 (MCE, catalogue number HY-10583) at density of 1500 cells per well for 2 days to generate embryoid body-like aggregates. From day 2–6, half of the medium was replaced with EB patterning medium containing 1 μM A83-01 (TGFβ inhibitor, Stem Cell Technologies, Cat# 72022), 2 μM Dorsomorphin (BMP signaling inhibitor, Stem Cell Technologies, Cat# 72102) and 1.25 μM XAV-939 (Wnt inhibitor, Stem Cell Technologies, Cat# 72674). On days 6–9, half of the medium was replaced with a neural induction medium containing 1 μM CHIR99021 (GSK3 inhibitor, reinforces Wnt signaling during early organoid formation, Stem Cell Technologies, Cat# 72052), 1 μM SB431542 (TGFβ inhibitor, Stem Cell Technologies, Cat# 72232) and 2 μM Wnt3a (MCE, Cat# HY-P70453). On day 9, organoids were embedded in the induction medium and Matrigel (Corning, catalog number 354230) with 3:2 dilution for another 5 days. On day 14, embedded organoids were mechanically dissociated from Matrigel and transferred to an orbital shaker in 6 well plates supplementing with differentiation medium containing 20 ng/mL BDNF (Stem Cell Technologies, Cat# 78005) and 20 ng/mL GDNF (Stem Cell Technologies, Cat# 78058). From Day 35 to Day 70, Matrigel (Corning, catalog number 354230) was added into the differentiation medium at 1% (v/v). At Day 70, the differentiation medium was replaced with a maturation medium. For astrocyte-conditioned medium incubation, simply replacing the neurobasal medium with medium conditioned by mouse (CD1) cortex astrocytes in differentiation and maturation medium.
Neural progenitor cells (NPCs) and forebrain neuron differentiation
To induce neural differentiation, human H9 ES cells were cultured in STEMdiff™ SMADi Neural Induction Kit (Stem Cell Technologies, Catalog #08581) following the manufacturer’s instructions. Briefly, H9 cells were seeded as a monolayer and cultured in the neural induction medium for 18–21 days, during which they transitioned into NPCs and can be further passaged and cryopreserved. The generated NPCs were then further differentiated into forebrain neuron precursors using the STEMdiff™ Forebrain Neuron Differentiation Kit (Stem Cell Technologies, Catalog #08600) for one week (Supplementary Fig. 15). NPCs and forebrain neuron precursors were dissociated into single cells and seeded into confocal dishes at a density of 3 × 104 cells/cm2. NPCs were used before passage 4. NPCs and forebrain neuron precursors were fed with differentiation medium prepared using MACM, HACM and Neurobasal as basal medium for 4 weeks. The protein concentration of ACM used in the experiments was 5 ug/ml.
Primary mouse and human astrocyte culture and conditioned medium preparation
All the animal experiments were performed under the ethical regulation of Shenzhen International Graduate School (SIGS) of Tsinghua University. All the mice were housed in a pathogen-free environment with the temperature maintained at 23 ± 2 °C and relative humidity at 50–65% under a 12 h/12 h light/dark cycle with free access to food and water, in accordance with the National Institutes of Health guidelines. Primary mouse astrocytes isolated from postnatal day 1–3 CD-1 mouse cortex were plated on a 10 mm dish in MEM medium containing 10% fetal bovine serum (FBS) (Thermo Fisher Scientific) and 1% Penicillin/Streptomycin (Gibco, Cat# 15140122)58,59,60. Briefly, the pups were euthanized, and their brains were dissected, removing the olfactory bulbs and hindbrain. The cortical plate was isolated, and the meninges were peeled away. Each cortical hemisphere was cut into small pieces (approximately 4–8 cuts) and subjected to enzymatic digestion to obtain a single-cell suspension. At culture day 8–10, confluent astrocytes were shaken at 1.61 g overnight to remove neurons, oligodendrocyte precursor cells and microglial. After shaking, the remaining confluent astrocyte layer was washed twice with PBS and passaged at a density of 2 × 106 cells. Then, these astrocytes reached over 90% confluence, cells were washed three times with PBS to remove the remaining FBS and a fresh neurobasal medium (Thermo Fisher) was added. Astrocyte-conditioned medium was collected and replaced with a fresh medium for 5 days (Supplementary Figs. 2 and 3).
Primary human astrocytes, isolated from the human brain cerebral cortex (catalogue no. 1800, Sciencell), were cultured in Matrigel (Corning, catalog number 354277) coated dishes supplementing the astrocyte medium (AM) (catalogue no. 1801, Sciencell). The culture medium was changed every other day. Cells were dissociated with TrypLE (Thermo Fisher Scientific) and neutralized with 10% FBS containing medium. Human astrocytes were used before passaged 10. For medium collection, same density and treatment procedure were adopted compared to mouse astrocytes. Both human and mouse conditioned medium was centrifuged at 1000 × g for 10 min to remove cell debris and sterile-filtered via a vacuum filter. The conditioned medium was stored at −80 °C until use. The ACM was concentrated using ultrafiltration tube (Thermo Fisher Scientific, 3000 MWCO) and analyzed by BCA assays and SDS-PAGE to monitor protein content and composition (Supplementary Figs. 28 and 29). The final concentration of both MACM and HACM used in the experiments were standardized as 5 µg/ml.
Protein Western blot
Cells were seeded in 10 cm culture dishes at a density of 2 × 106 cells per dish and cultured for 24 hours to allow cell attachment. The entire cell lysis process was performed on ice. First, cells were washed three times with PBS, followed by enzymatic digestion. After digestion, the cell suspension was centrifuged at 400 × g for 5 min at 4 °C to remove the digestion solution. The pellet was then washed twice with PBS and resuspended in an Eppendorf tube. Cells were lysed using RIPA Lysis Buffer (MedChemExpress, HY-K1001), and the supernatant was collected. The total concentration of protein in the supernatant was measured by using a BCA Protein Assay Kit (Thermo Scientific, A53225). Proteins were separated by electrophoresis using a 10% SDS-PAGE gel (Epizyme, PG212) according to their molecular weight. Then, proteins were transferred onto PVDF membranes (BIO-RED, 1620177) and blocked with 5% skim milk/PBS blocking solution at room temperature for 1 h. The membranes were then cut into appropriate sections based on the target band size and incubated overnight at 4 °C with primary antibody solution. Following incubation, the membranes were washed three times with TBST, the corresponding species-specific HRP-conjugated secondary antibody was applied and incubated at room temperature for 1 h. Finally, proteins were visualized by using the ECL detection system (4A BIOTECH, 4AW011-200) and imaged with the ChemiBoc XRS + imaging system (BIO-RED, USA).
Calcium imaging and data analysis
Dorsal forebrain organoids were vibrating sectioned by Leica VT 1200S and cultured using the microwell air-liquid interface attachment method9. Briefly, the slices were plated on 12 well confocal plates attaching polydimethylsiloxane (PDMS) (DOW, catlog# SYLGARD™ 184 Silicone Elastomer kit) microwell each well, ensuring minimal movement during imaging, and cultured at an air-liquid interface for 3 days before imaging. The plate was coated with 100 mg/mL poly-L-ornithine (Sigma-Aldrich, catlog# P4957-50ML) and 1% Matrigel. For calcium dye loading, the slices were incubated with 5 µM Fluo4-AM solution (Invitrogen, Catalog # F14201) for 2 h at 37 °C. Excess dye was removed by washing twice with BrainPhys™ Neuronal Medium (Stem Cell Technologies, Catlog# 05790). Imaging was carried out at 37 °C and 5% CO2 using a confocal microscope (NikonA1R) for 5 min with no delay between frames, and the frame rate was [7.67]. For drug treatment, drugs were loaded in a warm medium and incubated at 37 °C for 1 min. See a list of drugs and their detailed information in Supplementary Table 2 and Supplementary Fig. 11A. Calcium imaging data were processed in MATLAB (MathWorks) using the CNMF-E methodology to identify spikes and obtain fluorescence transients (ΔF/F0)61,62. Quantitative parameters of calcium activity were extracted based on inferred spiking activity neuron.S. The colorized calcium trace was plotted using CNMFeClustering63.
MEA recording and analysis
Brain organoids were sectioned into 500 μm slices at day 40 and seeded on six-well MEA plates (Axion Biosystem). Each well contains 64 0.04 MΩ platinum electrodes with 30 µm microelectrode diameter and spaced by 200 µm. The plate was coated with 100 mg/mL poly-L-ornithine and 1% Matrigel. Medium change was performed twice a week with prewarmed BrainPhys™ Neuronal Medium one day before recording. After recording, ACM was supplementing via medium change. 5 min of spontaneous activity data was recorded 3 rounds after the plate was stable in the Maestro device with the AxIS Navigator using the manufacturer’s Spontaneous Neural Configuration. Data analysis was performed using the manufacturer’s standalone tools Neural Metric Tool (Axion Biosystems). The spike-detecting threshold was set to 5.5 standard deviations, and the electrodes that detected at least 5 spikes/min were considered active electrodes. Bursts of each electrode were identified using an inter-spike interval (ISI) threshold requiring a minimum number of five spikes with a maximum ISI of 100 ms. As for network bursts in the well, a minimum of 10 spikes under the same ISI with a minimum of 25% active electrodes were required.
Single-cell RNA-seq library preparation and sequencing
5 organoids of each condition at day 90 were pooled together and washed three times with HBSS without Ca2+ and Mg2+ (STEMCELL Technologies, 37250). Single-cell suspensions were acquired via incubation with a papain-based dissociation (Miltenyi Biotec, 130-092-628). Prewarmed papain containing mix 1 (2 ml) was added to the organoids and incubated for 15–20 min at 37 °C with gentle agitation every 5 min. After triturating 5–10 times with 1000 µl wide-bore, enzyme mix 2 was added and incubated for 10 min at 37 °C twice with P1000 and P200 pipette tips trituration steps. Cells were washed twice with PBS + 0.2% BSA and filtered through a 40 µm cell strainer. Viability and cell count were assessed using a Trypan Blue assay on the automated cell counter Countess (Thermo Fisher Scientific). Cells were centrifuged at 300 × g at 4 °C for 5 min and resuspended in ice-cold PBS + 0.2% BSA at a cell density of 1000 cells/μL. Single-cell encapsulation and library preparation were performed according to the manufacturer’s protocol. Approximately 12,000 cells in each channel were loaded onto a Chromium Single Cell 3′ Chip (10× Genomics, PN-120236) and processed through a Chromium controller to generate single-cell gel beads in emulsions (GEMs) according to the manufacturer’s instructions. Captured libraries were prepared with the Chromium Single Cell 3′ Library & Gel Bead Kit v.2 (10x Genomics, PN-120237) and sequenced on an Illumina Novaseq with a paired-end 150 bp (PE150) reading strategy.
Single-cell RNA-seq analysis
Preprocessing and quality control
ScRNA-seq libraries were integrated by Seurat (v5.1.0) under the R (v4.4.1) environment. The reads were aligned to the GRCh38 reference genome in Cellranger-7.0.0. We filtered high-quality cells based on the number of genes detected (min. 200, max 6000), gene-level unique molecular identifier (UMI) counts (min. 500), log 10GenesPerUMI > 0.8 and removing cells with high mitochondrial (<20%) and ribosomal (<30%) mRNA content. Subsequently, the feature matrices were multiplied by 10,000 scaling factor and natural-log transformed. The top 2000 highly variable genes were then selected by Variance Stabilizing Transformation (VST) for downstream analysis. Principal component analysis (PCA) was performed on the expression matrices, and the top 20th PCs were used for downstream analysis. Cells were clustered in PCA space using Seurat’s FindNeighbors, followed by FindClusters with resolution = 1.2. Additionally, stressed cells were filtered out using the Gruffi R package47 before the removal of clusters expressing markers associated with unfolded protein response(e.g. DDIT3) with manual inspection64.
Clustering and annotation
Following quality control, cells were clustered (FindClusters function; resolution = 0.6) and visualized with Uniform Manifold Approximation and Projection (UMAP). Cluster identities were assigned based on differentially expressed gene markers, top biological process annotations and literature48,50,65. Differentially expressed genes (DEGs) for each cluster were identified via the FindAllMarkers function (log fold change threshold = 0.35). GO terms enrichment of each cluster was performed using the enrichGO function from the ClusterProfiler package (v.4.14.4). Enrichment was performed on biological process GO terms.
GO analysis of neurons and astrocytes between conditions
We ran a differential gene expression analysis comparing all neurons in MACMOs and control organoids using the FindMarkers function using the two-sided MAST test. Genes were annotated as significantly differentially expressed if they had a p.adjust less than 0.05. Significantly DEGs (including both upregulated and downregulated genes) between conditions were all submitted to pathway analysis by enrichGO function. Gene set enrichment analysis (GSEA) was performed using the gseGO function on all genes ranked by log fold change between astrocytes of the experimental and control groups. Enrichment was performed on biological process GO terms while the q value Cutoff was set at 0.05. False discovery rate (FDR) for GO and GSEA analysis was estimated by Benjamini-Hochberg method with p.adjust function.
Differentiation trajectory estimation
Pseudo-time trajectory estimation was carried out with R package monocle (v2.34.0). The kinetic trend of the expression level of significant genes along pseudotime was plotted utilizing the plot_cell_trajectory function.
Mass spectrometry
Protein extraction
Lysis buffer which contains Tris-HCl (pH 7.6), 8 M urea, 1 mM NaF and 2% (v/v) proteinase inhibitor was added in each sample. Samples were sonicated and centrifuged with 15,000 × g for 10 min. The concentration of protein was quantified using BCA kit according to the instruction manual.
Digestion and desalt
Add 10 mM DTT to the protein extraction solution and incubate at 56 °C for 1 h. Restore to room temperature, then add 55 mM IAA to the solution and incubate at room temperature in dark for 45 min. Subsequently, proteins were digested using FASP strategy at 37 °C overnight. Ultrafiltration tubes were centrifuged at room temperature of 15,000 × g for 10 min. Peptides mixture was transferred to a new tube and dried in vacuum. Peptides were then desalted using C18 column.
LC-MS/MS detection
The peptides were separated on a 25 cm C18 column (75 μm inner diameter, 1.9 μm particle size) using a nanoElute coupled to a timsTOF HT mass spectrometer via a CaptiveSpray ionization source using a 60-min acetonitrile (ACN) gradient: starting with solvent B (0.1% formic acid dissolved in 80% acetonitrile) ranging from 6 to 30% for 0–42 min, 30–42% solvent B for 42–51 min, 42–95% solvent B for 51–56 min and maintain 95% solvent B for 4 min. For the dia-PASEF acquisition, an average window size of 25.5 Da (minimum: 12.27 Da, maximum: 122.81 Da) and consisted of 12 frames with 3 mass windows per frame (75 ms) resulting in a cycle time of 0.975 seconds (including one MS1 frame).
Protein identification and quantification
Spectronaut (Biognosys, Schlieren, Switzerland) was used to analyze the DIA data with library-free method. MS/MS data were searched against protein sequences which were downloaded from Uniprot database with the following settings: enzyme: Trypsin/P; maximum missed cleavages: 2; fixed modification: carbamidomethyl (C); variable modifications: oxidation (M) and acetyl (protein N-term); precursor mass tolerance: 20 ppm; fragment mass tolerance: 0.05 Da. The results were filtered by 1% FDR, and only those protein groups which passed this filter criteria were used in downstream analysis.
Analysis of human and mouse protein sample
Based on protein name, the proteomics of mouse and human sample were mapped to identify shared and unique proteins. Functional enrichment analysis of co-detected proteins in both species, were performed via g:Profiler66. The interaction network analysis of selected proteins was performed using Genemania web server67.
Immunohistochemical analysis
Cells were fixed with 4% paraformaldehyde for 10 min and then washed three times in PBS. For organoids, samples were fixed for 20 min and washed similarly. After incubating overnight in 30% sucrose, samples were embedded in O.C.T. (Sakura). Fixed cells or sectioned organoids were similarly permeabilized with 0.1% Triton X-100, and blocked with 5% normal donkey serum in PBS for 1 h at room temperature, and then incubated with primary antibodies at 4 °C overnight. Alexa Fluor 488-, 568-, 594-, and 647-labeled secondary antibodies (Invitrogen) were used for detection. The nuclei were stained using a DAPI solution (Sigma-Aldrich) or Hochest (Yeasen). Following staining, the cells were imaged in PBS and the organoid sections were mounted on glycerol (Sigma-Aldrich)57. See list of antibodies and their information in Supplementary Table 3. For live cell staining in organoid slices seeded on a confocal plate, Calcein-AM (Yeasen, catlog# 40747-A) was added as manufacturer instructions. For lipid droplet staining, sections were stained with LipidSpot 488 (Biotum, catalog# 70065-T) (1:1000 in PBS) for 20 min at room temperature after the secondary antibody incubation and captured using a 40× water objective lens. Images were captured by a confocal microscope (NikonA1R). Primary antibodies used with corresponding dilutions were: SOX2 (Mouse, Santa Cruz, SC-365823, 1:100), TBR2 (Rabbit, Abcam, ab23345, 1:100), SATB2 (Rabbit, Abcam, ab34735, 1:100), REELIN (Mouse, Millipore, MAB5366, 1:100), FOXG1 (Rabbit, Abcam, ab18259, 1:100), KI67 (Mouse, BD, 550609, 1:100), PAX6 (Rabbit, Covance, PRB-278P, 1:300), NESTIN (Mouse, STEMCELL, 60091, 1:500), PVIM (Mouse, MBL, D076-3, 1:300), TUJ1 (Rabbit, Abcam, ab18207, 1:500), HOPX (Rabbit, Sigma-Aldrich, HPA030180, 1:500), BRN2 (Rabbit, Millipore, MABD51, 1:300), LAMININ (Rabbit, Abcam, ab11575, 1:100), S100 β (Rabbit, Abcam, ab52642, 1:500), TBR1 (Rabbit, Abcam, ab31940, 1:500), CUX1 (Mouse, Abcam, ab54583, 1:100), GFAP (Chicken, Abcam, ab4674, 1:2000), MAP2 (Chicken, Abcam, ab92434, 1:2000), SOX2 (Goat, R&D, AF2018, 1:200), CTIP2 (Rat, abcam, ab18465, 1:100), NEUN (Mouse, Millipore, MAB377, 1:300), O4 (Mouse, R&D, MAB1326, 1:200), vGLUT1 (Mouse, Millipore, MAB5502, 1:500), GABA (Rabbit, Sigma, A2052, 1:500), FZD9 (Rabbit, OriGene, TA314730, 1:300), PROX1 (Mouse, Millipore, MAB5654, 1:300), KIRREL2 (Sheep, R&D, AF2564, 1:500), MATH1 (Mouse, Santa Cruz, Sc136173, 1:100), MPHOX2B (Goat, R&D, AF4940, 1:500), OTX2 (Rabbit, Abcam, Ab21990, 1:300), SOD1 (Rabbit, Abcam, Ab51254, 1:300), TRX (Mouse, PTG, 66475-1-1g, 1:300), Cleaved Caspase-3 (Rabbit, Abcam, Ab32042, 1:500), OCT3/4 (Mouse, STEMCELL, 60093, 1:500). Secondary antibodies used with corresponding dilutions were: Anti-Mouse IgG H&L (Alexa Fluor® 594, Abcam, ab150116, 1:500), Anti-Mouse IgG H&L (Alexa Fluor® 488, Abcam, ab150113, 1:500), Anti-Rabbit IgG H&L (Alexa Fluor® 488, Abcam, ab150077, 1:500), Anti-Chicken IgY H&L (Alexa Fluor® 647, Abcam, ab150171, 1:1000), Anti-Rabbit IgG H&L (Alexa Fluor® 568, Abcam, ab175470, 1:500), Anti-Sheep IgG H&L (Alexa Fluor® 647, Abcam, ab150179, 1:1000), Anti-Rabbit IgG H&L (Alexa Fluor® 647, Abcam, ab150075, 1:1000), Anti-Goat IgG H&L (Alexa Fluor® 488, Abcam, ab150129, 1:500), Anti-Goat IgG H&L (Alexa Fluor® 647, Abcam, ab150131, 1:1000), Anti-Rat IgG H&L (Alexa Fluor 488, Abcam, ab150153, 1:500), Anti-Rat IgG H&L (Alexa Fluor® 647, Abcam, ab150155, 1:1000)
Analyses of layer thickness and percentage of positive marker cells
Cell counts and measurements were performed using ImageJ (NIH). Briefly, z-stacked images underwent maximal intensity projection. The expression of markers was quantified over DAPI+ cells in individual organoid sections using an automatic image threshold. The thickness of the progenitor zone (defined as the length of the SOX2+ region and the neural-tube morphology), neuronal zone (defined as the length from the boundary of the progenitor zone to the pial surface with TUJ1+ or MAP2+ cells), deep layer (defined as CTIP2+ or TBR1+ regions), and upper layer (defined as SATB2+ or CUX1+ regions) were measured. Relative layer thickness is the ratio of thickness to total thickness from the ventricular surface to the pial surface. ROIs of cortical regions were defined by identifiable ventricle-like structures and locally enhanced density of nuclei through checking cross-sections of organoids stained with DAPI. 2-3 region of interests (ROIs) of 100 μm width and 200–500 μm length radial columns per cortical region with positive marker cells were cropped to stand for the cortical plates.
Quantification of lipid droplets
For lipid droplet analysis, maximum z projections were created on each channel using auto-threshold with different methods (‘OSTU’ for TUJ1 and DAPI, ‘MaxEntropy’ for GFAP and LipidSpotTM488) to generate binary images. Overlapping mask of lipid droplets channel and GFAP+ or TUJ1+ were generated via image calculator. The masks were used to quantify astrocytic and neuronal lipid droplets. The ‘Analyze Particles’ function in ImageJ was then used to quantify the number of lipid droplets and average lipid droplet size. For each lipid droplet, the ROI was recorded. Among the ROIs, if the area of the TUJ1+ or GFAP+ mask that overlapped with the LipidSpot+ signal was greater than zero, then the counts of astrocytic and neuronal lipid droplets were incremented accordingly. Detailed macro codes can be found in the provided Github link: https://github.com/Zhengh-honghui/Codes_ACM_orgnaoids.
Statistics and reproducibility
All statistical tests and the exact number of experimental replicates are included in the figure legends. All data are presented as mean ± SEM as stated in Figure Legends. Statistical analysis was performed using GraphPad Prism (GraphPad Software Inc). If the data passed the normality tests, then the data were considered as normally distributed. Normally distributed data with equal standard deviations were analyzed using an unpaired two-tailed Student’s t test or ANOVA followed by post hoc Tukey’s multiple-comparison test. Normally distributed data with unequal standard deviations were analyzed using an unpaired t-test with Welch’s correction. Non-normally distributed data were analyzed using the unpaired Mann–Whitney U test or Kruskal–Wallis test followed by Dunn’s multiple-comparison test. *P < 0.05, **P < 0.01, and ***P < 0.001. All representative fluorescence images, live-cell staining, westen blotting, and bright field images shown in the main text and supplementary information were repeated at least three times independently with similar results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are included in the supporting material with the paper. The scRNA-seq data generated in this study have been deposited in GEO under a accession code: GSE273941. The mass spectrometry proteomics data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier, under an accession code: PXD060495. Any additional requests for information can be directed to, and will be fulfilled by, the corresponding author. Source data are provided with this paper.
Code availability
The code for scRNA analysis, lipid droplet quantification and local field potential of MEA recordings are available at GitHub (https://doi.org/10.5281/zenodo.14968830).
References
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell–derived neocortex. Proc. Natl Acad. Sci. 110, 20284–20289 (2013).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Di Lullo, E. & Kriegstein, A. R. The use of brain organoids to investigate neural development and disease. Nat. Rev. Neurosci. 18, 573–584 (2017).
Kelava, I. & Lancaster, M. A. Stem cell models of human brain development. Cell Stem Cell 18, 736–748 (2016).
Del Dosso, A., Urenda, J.-P., Nguyen, T. & Quadrato, G. Upgrading the physiological relevance of human brain organoids. Neuron 107, 1014–1028 (2020).
Pașca, S. P. The rise of three-dimensional human brain cultures. Nature 553, 437–445 (2018).
Kelley, K. W. & Pașca, S. P. Human brain organogenesis: Toward a cellular understanding of development and disease. Cell 185, 42–61 (2022).
Qian, X. et al. Sliced human cortical organoids for modeling distinct cortical layer formation. Cell Stem Cell 26, 766–781.e9 (2020).
Tang, J., Zheng, H., Feng, Y., Zeng, J. & Ma, S. Attachment culture of cortical organoids at the microwell air-liquid interface. STAR Protoc. 4, 102502 (2023).
Watanabe, M. et al. Self-organized cerebral organoids with human-specific features predict effective drugs to combat Zika virus infection. Cell Rep. 21, 517–532 (2017).
Giandomenico, S. L. et al. Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat. Neurosci. 22, 669–679 (2019).
Wang, Y., Wang, L., Guo, Y., Zhu, Y. & Qin, J. Engineering stem cell-derived 3D brain organoids in a perfusable organ-on-a-chip system. RSC Adv. 8, 1677–1685 (2018).
Kim, H., Lee, Y.-J., Kwon, Y. & Kim, J. Efficient generation of brain organoids using magnetized gold nanoparticles. Sci. Rep. 13, 21240 (2023).
Tang, X. et al. Volumetric Compression by Heterogeneous Scaffold Embedding Promotes Cerebral Organoid Growth and Maturation. Available at SSRN 4321960.
Li, Y. et al. Volumetric compression induces intracellular crowding to control intestinal organoid growth via Wnt/β-catenin signaling. Cell Stem Cell 28, 63–78.e7 (2021).
Schafer, S. T. et al. An in vivo neuroimmune organoid model to study human microglia phenotypes. Cell 186, 2111–2126.e20 (2023).
Popova, G. et al. Human microglia states are conserved across experimental models and regulate neural stem cell responses in chimeric organoids. Cell Stem Cell 28, 2153–2166.e6 (2021).
Shi, Y. et al. Vascularized human cortical organoids (vOrganoids) model cortical development in vivo. PLoS Biol. 18, e3000705 (2020).
Wörsdörfer, P. et al. Generation of complex human organoid models including vascular networks by incorporation of mesodermal progenitor cells. Sci. Rep. 9, 15663 (2019).
Revah, O. et al. Maturation and circuit integration of transplanted human cortical organoids. Nature 610, 319–326 (2022).
Mansour, A. A. et al. An in vivo model of functional and vascularized human brain organoids. Nat. Biotechnol. 36, 432–441 (2018).
Clarke, L. E. & Barres, B. A. Emerging roles of astrocytes in neural circuit development. Nat. Rev. Neurosci. 14, 311–321 (2013).
Allen, N. J. & Eroglu, C. Cell biology of astrocyte-synapse interactions. Neuron 96, 697–708 (2017).
Baldwin, K. T. & Eroglu, C. Astrocytes “chordinate” synapse maturation and plasticity. Neuron 100, 1010–1012 (2018).
Johnson, M. A., Weick, J. P., Pearce, R. A. & Zhang, S.-C. Functional neural development from human embryonic stem cells: accelerated synaptic activity via astrocyte coculture. J. Neurosci. 27, 3069–3077 (2007).
Diniz, L. P. et al. Astrocyte-induced synaptogenesis is mediated by transforming growth factor β signaling through modulation of D-serine levels in cerebral cortex neurons. J. Biol. Chem. 287, 41432–41445 (2012).
Huang, W.-K. et al. Generation of hypothalamic arcuate organoids from human induced pluripotent stem cells. Cell Stem Cell 28, 1657–1670.e10 (2021).
Wang, M. et al. Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain. Nat. Biotechnol. 43, 52–62 (2025).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Qian, X. et al. Generation of human brain region–specific organoids using a miniaturized spinning bioreactor. Nat. Protoc. 13, 565–580 (2018).
SM, C. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Elkabetz, Y. et al. Human ES cell-derived neural rosettes reveal a functionally distinct early neural stem cell stage. Genes Dev. 22, 152–165 (2008).
Georgala, P. A., Carr, C. B. & Price, D. J. The role of Pax6 in forebrain development. Dev. Neurobiol. 71, 690–709 (2011).
Renner, M. et al. Self‐organized developmental patterning and differentiation in cerebral organoids. EMBO J. 36, 1316–1329 (2017).
Pfrieger, F. W. & Barres, B. A. Synaptic efficacy enhanced by glial cells in vitro. Science 277, 1684–1687 (1997).
Ullian, E. M., Sapperstein, S. K., Christopherson, K. S. & Barres, B. A. Control of synapse number by glia. Science 291, 657–661 (2001).
Ullian, E., Harris, B., Wu, A., Chan, J. & Barres, B. Schwann cells and astrocytes induce synapse formation by spinal motor neurons in culture. Mol. Cell. Neurosci. 25, 241–251 (2004).
Krencik, R., Weick, J. P., Liu, Y., Zhang, Z.-J. & Zhang, S.-C. Specification of transplantable astroglial subtypes from human pluripotent stem cells. Nat. Biotechnol. 29, 528–534 (2011).
Zafeiriou, M.-P. et al. Developmental GABA polarity switch and neuronal plasticity in Bioengineered Neuronal Organoids. Nat. Commun. 11, 3791 (2020).
Welte, M. A. Expanding roles for lipid droplets. Curr. Biol. 25, R470–R481 (2015).
Kumar, M., Knapp, J. A., Gupta, K. & Ryan, T. A. Isolation and lipidomic profiling of neuronal lipid droplets: unveiling the lipid Landscape for insights into neurodegenerative disorders. bioRxiv, https://www.biorxiv.org/content/10.1101/2023.12.13.571527v1 (2023).
Ramosaj, M. et al. Lipid droplet availability affects neural stem/progenitor cell metabolism and proliferation. Nat. Commun. 12, 7362 (2021).
Ralhan, I., Chang, C.-L., Lippincott-Schwartz, J. & Ioannou, M. S. Lipid droplets in the nervous system. J. Cell Biol. 220, e202102136 (2021).
Ioannou, M. S. et al. Neuron-astrocyte metabolic coupling protects against activity-induced fatty acid toxicity. Cell 177, 1522–1535.e14 (2019).
Palop, J. J. & Mucke, L. Network abnormalities and interneuron dysfunction in Alzheimer disease. Nat. Rev. Neurosci. 17, 777–792 (2016).
Quadrato, G., Brown, J. & Arlotta, P. The promises and challenges of human brain organoids as models of neuropsychiatric disease. Nat. Med. 22, 1220–1228 (2016).
Vértesy, Á. et al. Gruffi: an algorithm for computational removal of stressed cells from brain organoid transcriptomic datasets. EMBO J. 41, e111118 (2022).
Bhaduri, A. et al. Cell stress in cortical organoids impairs molecular subtype specification. Nature 578, 142–148 (2020).
Zadoorian, A., Du, X. & Yang, H. Lipid droplet biogenesis and functions in health and disease. Nat. Rev. Endocrinol. 19, 443–459 (2023).
Uzquiano, A. et al. Proper acquisition of cell class identity in organoids allows definition of fate specification programs of the human cerebral cortex. Cell 185, 3770–3788.e27 (2022).
Pollen, A. A. et al. Establishing cerebral organoids as models of human-specific brain evolution. Cell 176, 743–756.e17 (2019).
Kim, D.-Y., Rhee, I. & Paik, J. Metabolic circuits in neural stem cells. Cell. Mol. Life Sci. 71, 4221–4241 (2014).
Zheng, X. et al. Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation. elife 5, e13374 (2016).
Cashikar, A. G. et al. Regulation of astrocyte lipid metabolism and ApoE secretion by the microglial oxysterol, 25-hydroxycholesterol. J. Lipid Res. 64, 100350 (2023).
Smolič, T. et al. Astrocytes in stress accumulate lipid droplets. Glia 69, 1540–1562 (2021).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Feng, Y. et al. Protocol for generating in vitro glioma models using human-induced pluripotent-or embryonic-stem-cell-derived cerebral organoids. STAR Protoc. 4, 102346 (2023).
Song, H., Stevens, C. F. & Gage, F. H. Astroglia induce neurogenesis from adult neural stem cells. Nature 417, 39–44 (2002).
Schwarts, J. P. & Wilson, D. J. Preparation and characterization of type 1 astrocytes cultured from adult rat cortex, cerebellum, and striatum. Glia 5, 75–80 (1992).
Schildge, S., Bohrer, C., Beck, K. & Schachtrup, C. Isolation and culture of mouse cortical astrocytes. JoVE J. Visualized Exp. 19, e50079 (2013).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Zhou, P. et al. Efficient and accurate extraction of in vivo calcium signals from microendoscopic video data. elife 7, e28728 (2018).
Samarasinghe, R. A. et al. Identification of neural oscillations and epileptiform changes in human brain organoids. Nat. Neurosci. 24, 1488–1500 (2021).
Amin, N. D. et al. Generating human neural diversity with a multiplexed morphogen screen in organoids. bioRxiv, https://www.biorxiv.org/content/10.1101/2023.05.31.541819v1.full (2023).
Gordon, A. et al. Long-term maturation of human cortical organoids matches key early postnatal transitions. Nat. Neurosci. 24, 331–342 (2021).
Kolberg, L. et al. g: Profiler—interoperable web service for functional enrichment analysis and gene identifier mapping (2023 update). Nucleic Acids Res. 51, W207–W212 (2023).
Franz, M. et al. GeneMANIA update 2018. Nucleic Acids Res. 46, W60–W64 (2018).
Acknowledgements
We thank members of Ma.’s lab for continuous support and insightful discussions. We thank Zhicheng Shao and Jinhong Xu for providing the human astrocytes used in this study. We would like to express our gratitude to Climb Technology Co., Ltd and Dr. Min Yao for their invaluable assistance with data mining and analysis in our proteomic profiling. The work was supported by the National Key Research and Development Program of China (2024YFA0919800) to S.M., by the National Natural Science Foundation of China (32371470) to S.M., by the National Natural Science Foundation of China (82341019) to S.M., by the Key-Area Research and Development Program of Guangdong Province (2023B0909020003) to S.M., and by the Cross-disciplinary Research and Innovation Fund of Tsinghua SIGS (No. JC2022007) to S.M. All schematics shown in the main text and Supplementary Figs. were created with biorender.com.
Author information
Authors and Affiliations
Contributions
S.M. conceived and supervised the work. H.Z. and S.M. wrote the manuscript with contributions from N.M., P. L. and Z. S. H.Z., F.Y., Z.W. and J.X. generated the figures. H.Z., J.T., Y.F., F.Y., J.X., Z.W. and M. J. performed the experiments. H.Z., F.Y. and J.X. analyzed the proteomic data. H.Z., Y.C. and C.H. analyzed the scRNA-seq data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Julia Ladewig, Alysson Muotri and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, H., Feng, Y., Tang, J. et al. Astrocyte-secreted cues promote neural maturation and augment activity in human forebrain organoids. Nat Commun 16, 2845 (2025). https://doi.org/10.1038/s41467-025-58295-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58295-3