Abstract
Protein production is critically dependent on gene transcription rates, which are regulated by RNA polymerase and a large collection of different transcription factors (TFs). How these transcription factors selectively address different genes is only partially known. Recent discoveries show that the differential condensation of separate TF families through phase separation may contribute to selectivity. Here we address this by conducting phase separation studies on six TFs from three different TF families with residue-scale coarse-grained molecular dynamics simulations. Our exploration of ternary TF phase diagrams reveals four dominant sticker motifs and two orthogonal driving forces that dictate the resultant condensate morphology, pointing to sequence-dependent orthogonal molecular grammar as a generic molecular mechanism that drives selective transcriptional condensation in gene expression.
Similar content being viewed by others
Introduction
Transcription is the first step in extracting the information stored in DNA for RNA production1,2. RNA polymerase (RNAP), the enzyme responsible for RNA production, is a multi-subunit protein that must be directed to the correct gene for RNA production to start1,2,3. The mediator complex4,5 acts as a bridge between RNAP and transcription factors (TFs), while the TFs localize RNAP to the correct gene on the DNA to initiate transcription and promote elongation2,3,6,7,8,9,10,11. The bundling of DNA by histone proteins for the compact storage of DNA in chromatin bundles affects the availability of a gene for TF binding, restricting transcription to active chromatin regions2,12,13,14. The transcription rates of different genes are controlled through transcriptional (or gene) regulatory networks (TRNs)15 by the recognition of specific promoter regions on DNA by the TFs, which are located near the targeted gene in space6,16,17. Independent gene regulation requires multiple differential transcription activation pathways within a TRN to enable selective gene expression to ensure that the correct genes are transcribed by the associated TF18,19,20,21. The TRN state of a cell is cell-type specific, so upon cell differentiation the selection of the specific TRNs required for cell function leads to the selection of TFs that are essential for those specific TRNs22. This means some TFs might never be produced by the same cell and therefore may never have an opportunity to interact with TFs in the mutually exclusive TF groups. In humans around 1600 different TFs have been identified, that can be divided into around 35 different family groupings23,24, indicating the immense diversity in human TFs and the ability to establish multiple independent TRNs. Identification of the underlying TF interactions requires the collection of a vast amount of proteomics data, generated with ChIP-seq, into databases such as ChIP-Atlas25.
In recent years it has been discovered that gene transcription is not only a biochemical process; spatial localisation also plays a key role. Super-enhancer regions spatially contribute to increased expression of specific genes, with down regulation of other genes26,27,28,29,30. The formation of such super enhancer regions and the associated transcriptional regulation has been increasingly linked to principles of polymer-based phase separation (PS)18,26,28,30,31,32,33,34,35,36,37,38. PS has been identified in the recent decade to drive the formation of membraneless organelles (MLOs) in several areas of cell biology39,40,41,42. The spatial localisation of TFs through MLO formation could assist mediator-RNAP binding to the correct gene during transcription initiation5,29,43,44. Indeed condensation of the RNAP CTD45, the mediator complex44 and chromatin13,14 have been identified, which could form a critical component in the further subcompartmentalisation and organisation within super enhancer regions2,29. Post-translational modification of RNAP, through the phosphorylation of the C terminal domain, is an essential part of the transcription cycle2,45. Such modification can have an effect on RNAP-TF interactions, therefore enabling the transfer of RNAP to different TF condensates45.
Understanding polymer phase separation within a theoretical framework started with Flory46 and Huggins47, where the relative strength of homotypic and heterotypic interactions of the polymer and solvent determine whether one homogeneous phase or two heterogeneous phases exist. Higher order mixtures of polymers and solvents have been shown to display rapidly increasing complexity in the resultant systems48,49,50,51, highlighting the complex balance between competing homotypic and heterotypic interactions. PS is driven by energetically favourable multivalent interactions, that counter the entropy loss of polymer demixing, which in proteins are from well structured or low complexity domains (LCDs)28,39,44,52,53,54,55,56. Changing the residue composition in LCD systems has been shown to enable the control of the PS behaviour, including changing the topology, by the tuning of critical interactions55,57,58,59,60,61,62. The observed transition from a fully demixed state (single homogenuos condensate) to a partially demixed state (with a multiphasic core-shell topology) in multicomponent systems57,58,59 upon residue modifications show that fine topological control of condensates is possible. The protein-protein interactions present in the resultant phase also enable the formation of phase-spanning percolation networks, resulting in the concept of phase separation coupled to percolation (PSCP) in MLO42,56,63.
TFs are typically composed of two main types of domains: DNA binding domains (DBDs) and effector domains (EDs)24. TF DBDs are responsible for binding specific DNA sequences (promoter regions) near the target gene to localise the TF, whereas TF EDs control target gene expression through other mechanisms such as interactions with cofactors, enzymes and mediator, RNAP recruitment, histone modification and DNA methylation24. LCDs are common features of transcription factors5,23,64 particularly the EDs24, while the RNAP C terminal domain region also contains a LCD65,66. TF condensate formation, predominantly mediated by LCD interactions of the TF EDs, is a mechanism for transcription control that enables colocalisation of the required proteins at the correct gene5,24,29,43,67. The formation of TF condensates at SE regions, such as SP167, BRD426,43, and OCT417,35, indicates their importance in gene regulation28,52. The interaction of these LCDs can be important for RNAP binding, with the size of the RNAP C terminal domain modulating the uptake into condensates mediating the transcription rates66,68,69. Interactions of TF condensates with mediator condensates is also likely to play an important role in transcription initiation at SE regions26,44. Disruption and misregulation of TF condensates are oncogenic drivers, leading to promotion of aberrant gene transcription behaviour64,67,70,71.
Understanding the driving forces for TF condensate formation is therefore essential to understand how gene expression can be regulated, and how it can be disrupted in the case of disease. In this work we aim to identify the molecular interactions that drive selective condensate formation, by focusing on a range of transcription factors from three different transcription families (FET, SP/KLF and HNF) that have LCDs previously identified to contribute to PS32,34,72,73. It has been hypothesized that the selectivity of TFs is caused by differential phase separation propensities32,34,74,75. Here we test this hypothesis by analyzing ternary phase diagrams of six TF LCDs that undergo PS to form condensates on their own under biological conditions32,34,72.
We use coarse-grained molecular dynamics simulations at amino-acid resolution72,76,77,78,79,80 to determine the molecular interactions that drive TF selectivity. Results are presented in terms of ternary phase diagrams, residue-scale contact maps and radial density profiles. It is important to note that our ternary diagrams represent a slice of the quaternary phase space (the four species are water plus three proteins) undertaken at a constant water fraction48,49,50,51. These revealed two distinct driving forces for condensate formation, consisting of hydrophobic interactions (from aromatic and aliphatic residues) and electrostatic/cation-π interactions (from cationic residues interacting with anionic or aromatic residues). We identified four dominant amino acid types, i.e. aromatic, aliphatic, cationic and anionic residues, responsible for driving collective phase separation forming four different sticker motifs in the conceptual ’sticker-spacer’ model of polymers56. The sequence composition of the TFs enables control over the homotypic and heterotypic intermolecular interactions, and therefore condensate selectivity. The relative homotypic and heterotypic strengths result in four distinct droplet morphologies: marbled (homogeneous), coated, bimodal, or separated droplets (heterotypic interactions are significantly weaker than homotypic interactions). The ability of the LCD of RNAP (POL II) to interact with condensates of all TFs highlights its central position and points to a universal principle in which selective transcription is regulated by sequence-based homotypic and heterotypic phase separation.
Results
Selection of transcription factors
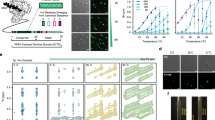
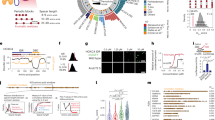
Droplet simulations were undertaken of six human TF IDRs from three different TF families: the FET family (FUS residues 2–214, EWS residues 47–266, TAF15 residues 2–205), the SP/KLF family (SP1 residues 2–507, and SP2 residues 1–524), and the HNF family (HNF1A residues 280–631) to explore the orthogonal driving forces for phase separation (see the amino acid sequences for the IDRs used in Fig. 1 and Supplementary Table 1). The FET TFs were selected due to a large body of experimental evidence of the IDR being crucial in the phase separation behaviour for FUS, EWS and TAF1532,34,73,81,82. Additional evidence also exists for SP132,34,67 and HNF1A72. SP2 was chosen as another member of the SP protein family. The IDR sequences selected correspond to those used in the experimental work of Chong et al.32,34 for FUS, EWS, SP1 and TAF15, the work of Kind et al.72 for HNF1A, and the sequence of SP2 by the use of PONDR-VSL283. We examine a large range of relative compositions of the ternary compositional space to fully describe the key motifs responsible for selective partitioning of the IDRs. To understand how the balance of homotypic and heterotypic interaction strengths varies for different members within the same family or with a different family, simulations of the seven key points on phase diagrams for different ternary combinations were conducted as shown in Table 1. Finally we conducted a study of TF-POLII (residues 1546–1790) condensate interactions. The name of the TF or POLII will be used to name the IDR fragment for brevity in the rest of the manuscript, unless explicit mention of the full length protein is made. Simulations were done at a monovalent ion concentration of 150 mM, with the maximum number of molecules set to 120 for FUS, EWS, and TAF15, 60 for SP1, SP2, and POLII, and 90 for HNF1A, so that the maximum number of amino acids per molecule type is approximately the same. PS is observed across all compositions of the TFs and POLII studied.
a FUS (residues 2–214), b TAF15 (residue 2–205), c SP1 (residues 2–507), d SP2 (residues 1-524), e EWS (residues 47–266), f HNF1A (residues 280–631), g POL II (residues 1546–1790). Residues are categorised into 5 groups: cations (R, K)- red, anions (D, E)- blue, aromatic (F, Y, W)- green, aliphatic (A, C, I, L, M, P, V)- black, hydropillic (G, N, S, H, Q, T)- white. The hydrophillic residues in the 1 bead-per-residue molecular representation are coloured differently to distinguish the proteins studied in this work: FUS (yellow), TAF15 (green), SP1 (purple), SP2 (light purple), EWS (orange), HNF1A (red), POL II (blue).
In the next section we start out by analyzing the FUS-SP1-TAF15 system in detail as a reference system that has been studied experimentally34.
Aromatic, aliphatic and cation-π interactions drive simple coacervation of TFs in the FUS-SP1-TAF15 system
Pure FUS, SP1 and TAF15 all exhibited strong homotypic phase separation forming stable droplets, as shown at the corners of the ternary phase diagram of Fig. 2 (with convergence data in Fig. 3). The molecular interactions driving PS can be determined by examining the molecular contacts between the amino-acids for the single component systems, shown in Fig. 4. The hydrophobic π − π interactions between tyrosine residues are the main driving force for PS in FUS (Fig. 4a, d and g). For FUS the 1D summations of interactions (Fig. 4a) show increased contacts around the locations of aromatic residues, highlighted with black dashed lines. This is clearly indicative of the ’sticker-spacer’ model56 with the tyrosine residues acting as stickers that drive phase separation and in agreement with the in vivo work of Kang et al.73. Interactions with and between the glycine, serine, and glutamine are also present in FUS, with a significant decrease in interactions around anionic residues, due to the repulsive electrostatic nature, leading to white streaks on the contact map. The contact maps for the SP1 simulation (Fig. 4b and e), show a completely different interaction profile: here aliphatic (alanine (A), isoleucine (I), leucine (L), proline (P) and valine (V) residue) hydrophobic interactions are key contributors to PS due to the high abundance of these residues in SP1, that is also reflected in the 1D summations of the interactions. The SP1 N-terminus has much lower contact frequencies due to a greater density of ionic residues, which have repulsive interactions with the aliphatic residues that constitute the majority of the SP1 molecules. This arises from the hydrophilic nature of the ionic residues, making the interactions with neutral hydrophobic aliphatic residues repulsive.
Ternary phase diagram showing the simulations undertaken in this work (orange dots) run with a total amino acid concentration of 80,000 μM. The total composition is defined relative to 120 FUS molecules, 120 TAF15 molecules, and 60 SP1 molecules, to give the percentage compositions (% FUS, % SP1, % TAF15). The end frame of 3 μs of simulation is displayed as a representative state. FUS molecules are coloured in yellow, SP1 molecules are coloured in purple, and TAF15 molecules are coloured in green. Droplets are formed in all simulations of these molecules under these concentration conditions, irrespective of composition.
a (100, 0, 0) b (0, 0, 100) c (75, 25, 0) d (75, 0, 25) e (12.5, 25, 62.5) f (33, 33, 33). Composition as a percentage is expressed by (% FUS, % SP1, % TAF15). Molecular connectivity data is sampled at 5ns intervals.
Intermolecular contacts by residue index for (a) 100% FUS, (b) 100% SP1, and (c) 100% TAF15, at 150 mM ion concentration and 300 K. The contacts are averaged in time and normalised by the number of molecules in the simulation (see section 3.2 in the SI for more details). A 1D contact profile (summation of the 2D map) is included below the contact map to show the total interactions per residue index (\({N}_{{\mathsf{contact}}}\)). The black dashed lines in (a) and (c) highlight the residues with the most contacts in FUS and TAF15 (which correspond to peaks in the 1D profiles). These are the aromatic residues in FUS driving π-π aromatic contacts, and the aromatic and cationic residues in TAF15 driving aromatic and cation-π interactions. Broad peaks are seen in (b) corresponding to the aliphatic residue stretches in SP1. Sequence information is included below the 1D summaries (copies of data displayed in a–c. Intermolecular contact map by residue type for (d) 100% FUS, (e) 100% SP1, and (f) 100% TAF15. The contact maps in (d), (e) and (f) are similar to the contact maps by residue index in (a), (b), and (c), respectively, but aggregated by residue type. A 1D contact profile (summation of the 2D map), are also included below (\({N}_{{\mathsf{contact}}}\)) together with the abundance for the residues (\({N}_{{\mathsf{residue}}}\)) shown by blue dashed lines. Intermolecular interaction summary for (g) FUS in 100% FUS (h) SP1 in 100% SP1, and (i) TAF15 in 100% TAF15 at 150 mM and 300 K. The fraction of interactions, \({{\mathsf{F}}}_{{\mathsf{int}}}\), are aggregated by type and normalised by the total number of the intermolecular interactions in (a–c) respectively. Aromatic and aliphatic interactions denote aromatic-aromatic and aliphatic-aliphatic interactions respectively. This convention is used throughout this work. Details of the contact definitions can be found in the methods.
In contrast to the purely hydrophobic driving forces for FUS and SP1, TAF15 PS has significant contributions from cation-π interactions in addition to the dominant aromatic contacts as can be seen in the contact maps in Fig. 4c and f. For TAF15 the 1D summations of interactions show increased contacts around the locations of cationic and aromatic residues indicative of cation-π interactions (black dashed lines in Fig. 4c). The N-terminus has a large number of tyrosine residues, with a relatively low number of anionic residues. The C-terminus, on the other hand, contains the majority of the cationic residues, leading to the high number of cation-π (arginine-tyrosine) contacts between the opposite ends of two TAF15 proteins. The central region contains a greater concentration of anionic residues, leading to a decrease in contacts, because it is self-repulsive and also repels the N-terminus, leading to the broad white bands. Interactions between glycine, serine and glutamine residues within TAF15 also contribute to PS due to the high abundance of these residues, but they are significantly weaker than the arginine-tyrosine interactions (see Supplementary Fig. 36C for the contact map normalised by residue abundance).
Figure 4g–i shows that the most important interactions are aromatic interactions for FUS and TAF15, and aliphatic interactions for SP1. Cation-π interactions also play a substantial role for TAF15. It is striking to see that for FUS and TAF15, despite the large difference in spatial contact maps (Fig. 4a and c based on residue index), the nature of the molecular interactions (Fig. 4g and i), and the residue-based contact maps (Fig. 4d and f) are remarkably similar, with the appearance of additional arginine-tyrosine (cation-π) contacts for TAF15.
The locations of the sticker residues within the molecules have a distinct effect on the topology and density of the single component droplets. FUS has a relatively even distribution of stickers throughout the protein allowing for the formation of relatively spherical droplets. TAF15 has a bimodal distribution of stickers located at the two termini, and a repulsive spacer region in the centre that repels other chains. This leads TAF15 to form a more open and porous structure where the droplet is formed of a collection of associating smaller TAF15 clusters (Fig. 2, density profiles in Supplementary Fig. 28). Increasing the system size (increasing the number of copies of molecules) does not change this behaviour (Supplementary Fig. 9), indicating that finite size effects do not play a role here. In all cases highly dynamic contacts are seen with contact lifetimes in the range 4-6 ns (Supplementary Fig. 10), showing that highly dynamical cross links are present42,79. Such dynamical molecular networks (Fig. 3) indicate PSCP since percolations exist within the droplets42,56,63,79.
Heterotypic interactions compete with homotypic interactions
As previously mentioned, PS is observed for all compositions, but the droplet morphologies and the molecular interactions change in nature as a second component was introduced, shown in Fig. 2. FUS-TAF15 mixtures form droplets with both proteins mixed and uniformly distributed throughout the condensate, with the droplet becoming increasingly less spherical with increasing TAF15 fraction. Whereas for FUS-SP1 mixtures the condensate is more spherical, with FUS-SP1 being well mixed at high SP1 fractions, before FUS starts to form a coating on the surface of an SP1 droplet at high FUS fractions. Interestingly, for TAF15-SP1 mixtures the condensate morphology is completely different. A singular mixed droplet is not favoured, instead the molecules are partitioned into two condensates: a TAF15 condensate and a SP1 condensate which do not interact.
Before examining the contact maps to clarify this, it is important to note that the intermolecular contact maps for the homotypic interactions of FUS, TAF15 and SP1 shown in Fig. 4 for the single component droplets are identical to the homotypic intermolecular contact maps observed in mixed systems (see Supplementary Figs. 34-60 in the SI). In this section we therefore exclusively focus on the heterotypic interactions between the two components in a system.
FUS-SP1 intermolecular interactions, shown in Fig. 5a and d, are primarily hydrophobic interactions driven by the large number of aliphatic residues in SP1 and tyrosine residues in FUS. The cation-π interactions appearing sharply in Fig. 5a are only a small part of the overall FUS-SP1 interactions with aliphatic-aromatic interactions the main FUS-SP1 intermolecular interactions (Fig. 5d and g). FUS-TAF15 intermolecular interactions, shown in Fig. 5b are driven by aromatic interactions between tyrosine residues in FUS and TAF15. Additional cation-pi contacts, from cations in TAF15 interacting with the tyrosines in FUS are also observed (Fig. 5e and h). Such cation-π interactions have previously been found to be crucial in the PS of full length FUS68,73,82, explaining why these smaller FUS and TAF15 IDR constructs exhibit a high favourability of co-condensation. It is expected that the inclusion of additional FUS domains would reduce the degree of FUS-TAF15 co-condensation due to the increased electrostatic repulsion between the FUS and TAF15, and the creation of more stable FUS condensates27,73,84. SP1-TAF15 intermolecular interactions are minimal, shown in Fig. 5c by the 1D contact summaries being two orders of magnitude smaller than for FUS-SP1 or FUS-TAF15 contacts in Fig. 5a and b. The large number of anionic residues in the centre and C-terminal of TAF15 (TAF15 has a relatively high net negative charge per residue of −12/204) only offer repulsive interactions with the large number of aliphatic and anionic (net charge per residue of −8/506) residues in SP1. This restricts favourable SP1-TAF15 interactions to much smaller chain segments, around cations in SP1, and the formation of very few contacts. The SP1-TAF15 interactions are much more localised than between SP1 and FUS, or TAF15 and FUS, or the homotypic interactions, making any mixing of the two molecules in one condensate unfavourable. By looking at the residue-based contact maps in Fig. 5d–f, the stickers and spacers of FUS and TAF15 are strikingly similar to each other and rather different from that of SP1, clearly highlighting that they originate from two different families of TFs.
Intermolecular contact map by residue index for (a) FUS with SP1 (50%, 50%), (b) FUS with TAF15 (50%, 50%), and (c) SP1 with TAF15 (50%, 50%) at 150 mM and 300 K. For the definitions of the different contact types see the caption of Fig. 4 and section 3.2 of the SI. The 1D contact profiles denote a summation of the 2D map of the corresponding molecules. The black dashed lines highlight the key residues: (a) the aromatic residues in FUS with cationic and aromatic residues in SP1, (b) aromatic residues in FUS with the aromatic and cationic residues in TAF15, and (c) the cationic residues in SP1 with the aromatic residues in TAF15. Sequence information is included below the 1D summaries (copies of data displayed in a–c). Intermolecular contact map by residue type for (d) FUS with SP1 (50%, 50%), (e) FUS with TAF15 (50%, 50%), and (f) SP1 with TAF15 (50%, 50%) at 150 mM and 300 K. Intermolecular interaction summary for (g) FUS-SP1 interactions in (50%, 50%, 0%), (h) FUS-TAF15 interactions in (50%, 0%, 50%), and (i) SP1-TAF15 interactions in (0%, 50%, 50%) at 150 mM and 300 K. The fraction of interactions, \({{\mathsf{F}}}_{{\mathsf{int}}}\), are aggregated by type and normalised by the total number of the intermolecular interactions in (a–c) respectively.
The normalised number of interactions between different species provides a method for comparison (full data for all simulations can be found in Supplementary Tables 9-13 and Supplementary Figs. 6-8 in the SI). The single component mixtures all have comparable numbers of homotypic interactions for FUS (0.015), SP1 (0.016), and TAF15 (0.014) with TAF15 forming slightly more open condensates. In the two component mixtures we see the emergence of a difference in the relative strengths of the homotypic and heterotypic interactions. Three possible cases exist for binary mixtures. In the first case the heterotypic interactions are stronger than the homotypic interactions (FUS-TAF15), then mixing of the two species in a marbled condensate is preferred (see (75,0,25) in Fig. 2 and Supplementary Fig. 6B). In the second case when the homotypic interactions are both stronger than the heterotypic interactions (SP1-TAF15), then a bimodal/separated condensate system is preferred (see (0, 50, 50) in Fig. 2 and Supplementary Fig. 6C). For the third case the homotypic and heterotypic interactions are of a comparable strength (FUS-SP1), leading to a coated (or core-shell) condensate structure (see (75, 25,0) in Fig.2 and Supplementary Fig. 6A).
Ternary systems display complex condensate morphology driven by interaction orthogonality
Moving into the centre of the FUS-SP1-TAF15 phase diagram we now see the competition between the attractive groups in the different molecules for interactions in the resultant ternary condensates. A combination of the previously observed morphologies is now seen, as shown in Fig. 6. Attractive interactions of FUS with both SP1 and TAF15 are opposed by the repulsive interactions between SP1 and TAF15 in the formation of a single heterogeneous condensate (see radial density profiles in Supplementary Figs. 28-29). Competition between SP1 and TAF15 for the interactions with FUS leads to a complex, composition-dependent behaviour of the droplets. In high FUS fraction compositions a single droplet is observed with SP1 and TAF15 localised at different ends of the condensate such that SP1-TAF15 contacts are minimised. At low FUS fractions a singular large condensate is lost, and instead separate droplets are formed containing only TAF15 with FUS inclusions or SP1 with FUS inclusions, see e.g., the (25, 50, 25) and (12.5, 12.5, 75) compositions. This transition to separate droplets occurs at 50% SP1 for SP1 rich systems, whereas TAF15 rich systems require 75% TAF15 for analogous behaviour.
The percentage compositions are described in the image. The end frame of 3 μs of simulation is displayed as a representative state. Composition as a percentage is shown next to the droplets in brackets: (% FUS, % SP1, % TAF15). Droplets are formed in all simulations of these molecules under these concentration conditions, irrespective of composition. The relative interaction strengths are displayed on this diagram for the homotypic and heterotypic contacts. The definition of this quantity can be found in section 4.7.3.
The intermolecular contact maps for the three-component mixtures do not yield any extra information compared to the contact maps from the two-component mixtures. The intermolecular contact maps for all residue pairings exhibit the same interaction patterns, with only the absolute magnitude of each contact map changing (see Supplementary Figs. 34-60 in the SI). From the convergence data (Fig. 3) it can be deduced that at all times percolations exist within both homotypic as well as heterotypic droplets. The intermolecular interactions in the TF condensates, however, are not at all permanent; they are highly dynamic with contact lifetimes on the order of 5 ns (see Supplementary Fig. 10) such that the cross-links between TFs are transient in nature, similar to FG-nucleoporin condensates79. The nature of this phase state can best be categorized as viscoelastic: on one hand the continuous percolations suggest elastic properties, while their transient nature is indicative of a fluid, with mostly spherical morphologies and the ability of two droplets to merge into one42,56,63,79,85,86.
In the ternary mixtures (Fig. 6) the competition for interactions is dominated by the competition for the strongest heterotypic interactions. The strongest heterotypic interaction is a function of composition, with FUS-TAF15 the strongest for the majority of compositions, and FUS-SP1 interactions the strongest at 75% SP1 content, and SP1-TAF15 the weakest for all compositions (Supplementary Fig. 6). The dominance of the FUS-TAF15 interactions drive the formation of FUS-TAF15 marbled condensates, since it out-competes the other heterotypic interactions. Since the FUS-SP1 interactions are also strong, but weaker than SP1-SP1 interactions this promotes the FUS to coat the SP1 droplet. This results in a complex bimodal structure where a FUS-TAF15 marbled droplet is partially coating a SP1 droplet. The intriguing behaviour shown in the simulations is that if sufficient FUS is present, both SP1 and TAF15 are incorporated into the same bimodal droplet even though there interactions are very unfavourable (<0.001). At lower FUS compositions the ability of FUS to provide sufficient interactions to shield TAF15 from SP1 in the same droplet is reduced and eventually leads to the fragmentation of the droplets as in the binary SP1-TAF15 systems. A characteristic feature of the ternary interactions in Supplementary Fig. 6 is that the homotypic interactions of SP1 and TAF15 can increase with droplet composition. This is particularly strong for SP1 in high TAF15 fraction mixtures and TAF15 in high SP1 fraction mixtures (Supplementary Figs. 6(D)-S7(F)), indicating that the presence of the other proteins has a crowding effect through repulsive protein-protein interactions.
We identify four different residue types that can behave as stickers: aromatic, aliphatic, anionic, cationic. The table in Supplementary Fig. 5A shows the mean interaction propensity calculated for all 34 FUS-SP1-TAF15 systems studied (shown by the orange dots in Fig. 2, data in Supplementary Figs. 3 and 4 in the SI). This shows that cation-π interactions are the strongest interaction type, with 20% of all theoretically possible interactions being formed. The second most important interaction type are the aliphatic interactions which have a greater uncertainty than the cation-π interactions. Aromatic interactions are the third most important interaction, with a similar uncertainty to cation-π. Electrostatic and aliphatic-aromatic interactions provide the smallest relative contribution to the total. From these mean interaction propensities in the table in Supplementary Fig. 5A, we compute the possible fraction of interactions we expect to see in any FUS-SP1-TAF15 stochiometry as shown in Supplementary Fig. 5B–F. In Supplementary Fig. 5B, aromatic-aliphatic interactions are seen across all compositions, with fewer present in higher FUS/TAF15 compositions. Aromatic interactions occur most frequently in FUS and TAF15 rich mixtures. Cation-π and electrostatic interactions occur most frequently at large TAF15 fractions, due to the lower fraction of cationic residues in SP1 and FUS sequences. Aliphatic interactions decrease with decreasing SP1 fractions, since SP1 contains the majority of the aliphatic residues in the sequences investigated. This allows the interactions to be grouped into three main classes: SP1 preferred interactions (aliphatic), FUS/TAF15 preferred interactions (aromatic) and TAF15 preferred interaction (cation-π and electrostatic).
Universality of TF selectivity
In this section we extend the analysis to also include the TFs EWS, SP2 and HNF1A. These TFs also undergo PS on their own (Supplementary Figs. 11-16)32,72. The IDR of EWS contains a high aromatic content similar to FUS and TAF15, but with more aliphatic residues (Fig. 1 and Supplementary Fig. 18A). This leads to EWS interactions to be driven by aliphatic contacts from the higher alanine and proline content, and aromatic (tyrosine-tyrosine) contacts (Supplementary Fig. 18G). A similarly high aliphatic residue content in SP2 as SP1 leads to the same dominance of aliphatic interactions in driving SP2 condensate formation (Supplementary Fig. 18H). These aliphatic interactions in SP2 are similarly distributed along the entire length of the SP2 IDR in the same manner as SP1 (Fig. 1 and Supplementary Fig. 18B). HNF1A contains a high aliphatic residue content (Fig. 1 and Supplementary Fig. 18C), which leads to the dominance of leucine, valine, alanine and proline interactions (Supplementary Fig. 18F). It is interesting to see that the sticker-spacer paradigm is valid for FUS, TAF15 and EWS, in contrast to SP1, SP2 and HNF1A where it is the more uniform attraction between the aliphatic residues that drive PS.
Next we extend the analysis to binary and ternary systems, following the combinations shown in Table 1. Analysis of the different interaction types of the FUS-TAF15-SP1 system (Supplementary Fig. 6), the contact maps (Fig. 5, Supplementary Figs. 22-28) and the ternary phase diagrams (Figs. 2, 6 and Supplementary Figs. 12-17) of all systems studied, it can be concluded that the selectivity of phase separation is related to two major groupings of interactions (shown in Fig. 7a). The first grouping of hydrophobicity-controlled interactions encompasses aliphatic, aromatic and aliphatic-aromatic interactions that utilise stickers that can interact with other residues of the same type. The second grouping consists of cation-controlled interactions which encompasses cation-π and electrostatic interactions. Here sticker residues belong to different residue types, so the stickers in the interactions must be heterogeneous, i.e. cationic, anionic and aromatic residues. Interestingly, aromatic and anionic residues both have attractive interactions with cationic residues, but anionic residues have repulsive interactions with aromatic or other anionic residues. This categorisation of interactions highlights the molecular mechanism to achieve selectivity of phase separation. By exploiting the orthogonal molecular driving forces the relative homotypic and heterotypic interaction strengths can be tuned through sequence composition. These orthogonal interactions lead to four characteristic binary droplet morphologies: marbled (homogeneous) droplets (such as the SP1-SP2 system) with heterotypic interactions larger than homotypic interactions (Fig. 7b), coated (core-shell) droplets (such as the EWS-TAF15 system) with heterotypic interactions of comparable strength to the homotypic interactions (Fig. 7c), bimodal droplets (such as the HNF1A-TAF15 system) with heterotypic interactions smaller than homotypic interactions (Fig. 7d), separated droplets (such as the SP1-TAF15 system) with heterotypic interactions significantly smaller (almost negligible) than homotypic interactions (Fig. 7e).
a Amino acid sticker types and interactions (as published in84 Fig. 6a, used under CC BY 4.0 license [http://creativecommons.org/licenses/by/4.0]). The favourable interactions (black) promote condensate formation, whereas unfavourable interactions (red) oppose condensate formation. Sticker interactions fall into two orthogonal molecular interactions: 1) Hydrophobic interactions (aliphatic, aromatic and aliphatic-aromatic contacts), and 2) Cationic interactions (cation-π and electrostatic). b–e Different condensate morphologies for the four cases of relative homotypic and heterotypic interaction strengths, illustrated with example (50%-50%) mixtures from simulations undertaken in this work. Corresponding radial density profiles can be found in Supplementary Figs. 28 and 31.
The universality of condensate selectivity can be nicely depicted by discussing the phase diagrams (of Table 1) relative to the reference system of FUS-SP1-TAF15. The first contains the substitution of FUS by EWS. The additional aliphatic residues in EWS lead to some important changes (Supplementary Fig. 11) when compared to the FUS-SP1-TAF15 triangle (Figs. 2 and 6). Of EWS, SP1, and TAF15 the homotypic interactions of EWS are the strongest (0.025), also stronger than FUS (0.015). Moreover, the additional heterotypic aliphatic interactions of EWS with SP1 (Supplementary Fig. 22) results in heterotypic EWS-SP1 interactions that are stronger than SP1 homotypic interactions, but weaker than EWS homotypic interactions and causes the formation of a binary SP1-coated EWS droplet (Fig. 7c). The additional aliphatic residue content of EWS compared to FUS leads to more repulsion with the ionic residues in TAF15, resulting in decreased heterotypic interactions of 0.01-0.02 (Supplementary Fig. 22B), leading to a more patchy binary interaction with TAF15 compared to FUS and a distinct separation from TAF15 in the 3-component system (Fig. 8a and Supplementary Fig. 11).
a EWS-SP1-TAF15, b FUS-SP2-TAF15, c SP1-SP2-TAF15, d FUS-HNF1A-TAF15, e HNF1A-SP1-TAF15, f HNF1A-SP1-SP2. All simulations were run with a total amino acid concentration of 80,000 μM. The end frame of 3 μs of simulation is displayed as a representative state. A segment is not displayed to reveal the internal droplet structure. FUS molecules are coloured in yellow, EWS molecules are coloured in orange, SP1 molecules are coloured in purple, SP2 molecules are coloured in light purple, HNF1A molecules are coloured in red, and TAF15 molecules are coloured in green. Corresponding radial density profiles can be found in Supplementary Fig. 33.
The replacement of SP1 by SP2 leads to the FUS-SP2-TAF15 triangle in Fig. 8b and Supplementary Fig. 12. SP2 has an increased cationic residue content compared to SP1. These cationic residues in SP2 are clustered in three main locations around residue 250, 400 and in the C terminal end (above residue 500), see Fig. 1. These clusters are sufficient to enable the formation of strong electrostatic and cation-π interactions with TAF15 (Supplementary Fig. 21C), and the formation of marbled SP2-TAF15 condensates (Supplementary Fig. 12). All the heterotypic interactions in this triangle are stronger than homotypic interactions (Supplementary Fig. 8), resulting in the formation of a three-component marbled droplet (Fig. 8b and Supplementary Fig. 12). If we then replace FUS by SP2 the phase diagram of SP1-SP2-TAF15 in Supplementary Fig. 13 is created. The large number of aliphatic residues in both SP1 and SP2 dominate the heterotypic interactions leading to the formation of marbled condensates (see contact maps in Supplementary Fig. 21B). The heterotypic SP1-SP2 interactions are much stronger than the homotypic interactions (Supplementary Fig. 8B). Intriguingly, the strong heterotypic interactions of SP2 with both SP1 and TAF15 are able to overcome the repulsive SP1-TAF15 heterotypic interactions (like FUS in the FUS-SP1-TAF15 droplets). In Fig. 8c the equal contribution ternary SP1-SP2-TAF15 mixture results in marbled SP1-SP2 condensates forming around a disk of TAF15, with SP2 acting as a glue at the interface to reduce contact between SP1 and TAF15.
Three distinct cases of ternary phase diagrams are considered with HNF1A. The first case (Supplementary Fig. 14) is HNF1A with two FET proteins (FUS and TAF15). The heterotypic HNF1A-TAF15 interactions are unfavourable, producing a bimodal condensate with a minimal interfacial region between the HNF1A and TAF15 condensates. This is in contrast to the marbled FUS-HNF1A and FUS-TAF15 condensates where favourable heterotypic interactions are present in the binary mixtures. In the ternary mixture (Fig. 8d), FUS-HNF1A heterotypic contacts dominate over those of FUS-TAF15, leading to formation of a HNF1A condensate coated in FUS with a TAF15 condensate interacting with FUS on a shared interface. The second case (Supplementary Fig. 15) is HNF1A with one FET (TAF15) and one SP/KLF (SP1) protein. Here SP1-HNF1A heterotypic interactions are the strongest giving rise to the marbled HNF1A-SP1 binary condensates. In the ternary mixture the weak HNF1A-TAF15 heterotypic interactions lead to the association of a TAF15 homotypic domain to the marbled HNF1A-SP1 droplet (Fig. 8e). The third case (Supplementary Fig. 16) of HNF1A with two SP/KLF family TFs (SP1 and SP2) contains three heterotypic interactions that are more favourable than homotypic interactions (Supplementary Fig. 8). Therefore a marbled three component droplet is observed for this system driven by the large number of aliphatic residues in all three components.
The HNF1A IDR from the HNF family contains a high number of aliphatic residues, with a small number of aromatic and ionic residues, meaning it more closely resembles the IDRs of the SP/KLF family proteins than the FET proteins. Aliphatic-aromatic interactions dominate the heterotypic interactions of HNF1A with FUS and TAF15 (Supplementary Figs. 22C and 23C). The heterotypic interactions of HNF1A with SP1 and SP2 are dominated by aliphatic residue contacts (see contact maps in Supplementary Figs. 23A and B).
Pol II interaction promiscuity overcomes interaction orthogonality
The function of transcription factors is to direct RNAP to the required gene for transcription initiation. The C-terminal domain of RNAP II (referred to as POL II) is known to be able to undergo PS66,68,69,87. A key question exists: how can POL II circumvent the orthogonality of the interactions displayed by the transcription factors, such that it can reach all required genes to initiate and undertake transcription? The answer lies in the POL II amino acid sequence, which is based on a heptad repeat unit containing the consensus sequence YSPTSPS65. POL II homotypic interactions are dominated by aromatic tyrosine contacts (see contact maps in Supplementary Fig. 19). The use of aromatic interactions in a low charge IDR provides the opportunity to maximise the propensity for co-condensation with any TF, as shown in Fig. 9, where POL II was found to undergo condensation with all TFs in this study (FUS, EWS, HNF1A, SP1, SP2, and TAF15), overcoming the potential barrier posed by the orthogonal interactions used by different TF families (radial density profiles in Supplementary Figs. 30 and 32). This is achieved by heterotypic hydrophobic interactions of POL II with all TFs (Supplementary Figs. 24 and 25). The ability for POL II to enter any TF condensate means that the local TF condensate environment formed in the nucleus plays a crucial role in orchestrating POL II localisation to the correct genes for transcription.
Simulations of 60 POL II molecules (centre) and 30 POL II with: 60 FUS (top left), 60 TAF15 (centre left), 60 EWS (bottom left), 30 SP1 (top right), 30 SP2 (centre right), and 45 HNF1A (bottom right). All simulations were run with a total amino acid concentration of 80,000 μM. The end frame of 3 μs of simulation is displayed as a representative state. POL II molecules are coloured in blue, FUS molecules are coloured in yellow, EWS molecules are coloured in orange, SP1 molecules are coloured in purple, SP2 molecules are coloured in light purple, HNF1A molecules are coloured in red, and TAF15 molecules are coloured in green. Droplets are formed in all simulations of these molecules under these concentration conditions, irrespective of composition. See radial density profiles in Supplementary Figs. 30 and 32).
POLII undergoes phosphorylation during the transition from the initiation to elongation stage of transcription2,45. This change leads to a change in the TFs and cofactors that interact with POLII. To model the change in charge on POLII the mutation of all (141) serine residues to aspartic acid residues is used (sequence POLIIcharged, in Supplementary Table 1). The increase in negative charge leads to the loss of simple coacervation for POLII (radial density profile in 31), and also the lack of solubility in FUS, EWS, TAF15, SP1, and HNF1A (see Supplementary Figs. 17 and 32 in the SI). Partial solubility of POLIIcharged is still observed for SP2 (Supplementary Figs. 17 and 33). The change in charge of POLII also leads to a significant reduction in the number of contacts (see contact maps in Supplementary Figs. 20, 26 and 27). This shows that post translational modification can be used to tune the condensation of RNAP, promoting RNAP to leave the TF condensates that can aid transcription initiation and transition towards RNA elongation45.
Discussion
We explored transcriptional condensates consisting of six TFs from three different families. We focused on different spatial levels of molecular interactions by analysing (i) residue scale contact maps (ii) the relative strengths of aliphatic, aromatic, aliphatic-aromatic, electrostatic, and cation-π interactions (iii) the aggregated overall homotypic and heterotypic interaction strength between the molecules and (iv) radial density profiles. We have found that four sticker motifs56 exist that feature two orthogonal driving forces causing PS in ternary TF systems, shown in Fig. 7a. The competitive interactions driving PS within the ternary TF systems can be ranked by their relative strength (the table in Supplementary Fig. 6A): cation-π are the strongest, followed by aliphatic, then aromatic, and finally aliphatic-aromatic and electrostatic are the weakest. The sticker motifs trigger hydrophobic interactions (aromatic, aliphatic-aromatic, and aliphatic interactions) between Y, A, L, V, P residues, or cation-π and electrostatic interactions of R and K with Y, D, or E, to drive PS (Fig. 7a).
PSCP of the single component droplets is driven by homotypic intermolecular interactions (Fig. 4 and Supplementary Fig. 18): PS of FUS and EWS is driven by aromatic interactions, TAF15 by electrostatic, cation-π and (mostly) aromatic interactions, and SP1, SP2, and HNF1A by aliphatic interactions. The contact maps we present highlight the key residue interactions responsible for the observed phase separation- mutations of these sites are expected to alter the PS behaviour. For instance, in our previous work on HNF1A72 we carried out deletion mutations which showed there were still sufficient interactions in other regions to promote PS, exemplifying redundancy. Such a perturbation analysis would be an interesting topic for future work. The two component mixtures feature competition of homotypic with heterotypic intermolecular interactions (Fig. 5 and Supplementary Figs. 21-27) leading to the identification of four distinct droplet morphologies in binary systems where both species can undergo phase separation: 1) heterotypic interactions being stronger than homotypic interactions leads to a single marbled condensate (Fig. 7b), 2) heterotypic interactions being comparable to homotypic interactions leads to a single coated condensate (Fig. 7c), 3) heterotypic interactions being weaker than homotypic interactions leads to bimodal condensates (Fig. 7d) that have a shared interface. 4) heterotypic interactions being substantially weaker than homotypic interactions leads to two separate condensates (Fig. 7e). These morphologies are also represented in the ternary systems studied, featuring selective phase separation (as summarised in Fig. 8). The ability of TF IDRs to undergo selective PS when combined with TF-DNA interactions through specific DNA binding domain interactions contributes to the selective gene localisation of such condensates.
Since one of the orthogonal driving forces for phase separation involves ionic residues, the localized partitioning of transcriptional condensates at the DNA is sensitive to the local electrostatic environment. To explore this we have studied the effect of ion concentration in one specific ternary system that has also been investigated experimentally: FUS-TAF15-SP134. Our results showed that with increasing ion concentration the strength of FUS-TAF15 interactions grows with a corresponding weakening of FUS-SP1 interactions in ternary mixtures, making SP1 inclusion in the condensates less favourable (see SI section 5 and Supplementary Fig. 7). In living cells, this interaction orthogonality is expected to be even larger, due to crowding or modified local electrostatic environments (for example the presence of large amounts of negatively charged DNA), enhancing the heterotypic FUS-TAF15 interactions at the expense of heterotypic interactions with SP1. This was indeed shown in the in vivo work of Chong and coworkers32,34, where FUS-TAF15 condensates were observed, with SP1 forming separate condensates. FUS-TAF15 and HNF1A-SP1 interactions have also been recorded in ChIP-seq data deposited in ChIP-Atlas25, however no data is present for other TF pairs examined in this work. The lack of ChIP-seq data could be due to no interactions being present to detect, as in the case of TAF15 with SP1, or insufficient sampling. The six TF IDRs studied in this work have provided significant insight into the mechanisms of TF condensation, yet these IDRs represent a subset of the entire proteins. Inclusion of the DNA/RNA binding domains of the TFs is expected to accentuate the selectivity of the interactions seen in the chosen TF IDRs due to the additional binding of DNA/RNA57,61,62,82,88. It is important to note that in the current work we focused on one specific LCD as used in previous experimental studies. However, some TFs have more than one IDR bridged by folded domains, which add additional multivalent interactions as driving forces for phase separation67,73,82. The mutual exclusion of SP1 and TAF15 in condensates through repulsion from the anionic residues also indicates the role anionic species could play in determining condensate structure and percolation through crowding effects. This is of particular interest for exploration in the DNA rich nuclear environment where TFs function, which would be an interesting topic for future studies. It should be noted that in this work we focus on the interactions of FET proteins with two SP/KLF family proteins and one HNF family protein, which almost always includes a FET protein. Therefore, exploration of the specificity in the molecular grammar and ensemble types using other physiological combinations of the 1600 human TFs would provide opportunities for future work to validate the condensate morphologies described in the current work.
The fact that both orthogonal driving forces for phase separation involve aromatic residues enables POL II to interact with any TF to enable co-condensation. Co-condesation of POLII with TFs (Fig. 9) enables the co-localisation in the vicinity of the gene promoter ready for transcription initiation. This provides a simple mechanism that aids in enzyme partitioning to active transcription sites where increased TF concentrations are present, and aids in the pre-assembly of RNAP-mediator complexes5,44. The condensates enable the RNAP to dock to the correct chromatin location through interaction of its IDR with TFs and at the same time facilitate the active enzymatic site to attach to the DNA and initiate transcription. The principle of sequence-dependent selective TF co-condensation together with the promiscuous binding of RNAP, provides a universal mechanistic basis for selective gene transcription. Subsequent POLII phosphorylation then enables the relocation of the RNAP complex to start the process of RNA elongation, leaving the TF condensate which promoted the transcription initiation45. Additionally the condensation of the mediator complex44 indicates that a complex network of percolated condensates may be present in cells with the variable miscibility of components (as seen in Fig. 6) contributing to the internal spatial organisation of super enhancer regions.
The wide range of protein density within condensates (SI section 10, Supplementary Figs. 28-33) further increases the complexity when the solubility of enzymes and other cofactors needed for transcription are considered. The uptake of various cofactors into the TF condensates are governed by enthalpic and entropic driving forces. The entropic penalty of confinement within a condensate is typically conteracted by intermolecular interactions which provide an enthalpic driving force for inclusion (as in condensate formation). The porosity of TF condensates will contribute to the selectivity of cofactor uptake based on the available molecular interactions (enthalpic contribution) and the displacement of water/rearrangement of TF (entropic contribution) within the condensate to enable cofactor uptake. Comparison of binary and ternary condensates in Figs. 6–8 and Supplementary Figs. 12-17 highlight how the addition of an additional species can lead to a wide array of different droplet morphologies when all components are able to undergo simple coacervation. When we also consider cofactors which must be recruited by TFs to enter condensates, then the surface topologies of TF condensates provide an additional layer of selectivity, with only TFs present on the droplet surface able to directly interact with cofactors. The percolated networks formed within the condensates allow for the transmission of information through the droplets, potentially creating a route for the transfer of information between adjacent condensates if a topological change is detected during the rearrangement of the transcriptional machinery (such as the unwinding of chromatin fibres)29.
All-in-all, our results might have several implications. The orthogonal molecular grammar provides generic molecular insights in understanding TF phase separation in selectively controlling gene transcription rates. It introduces the viewpoint that molecular interactions might be evolutionary converged to a state in which subtle differences in molecular grammar might steer different genetic programs. We anticipate that this will lead to increased research activity in the broader scope of proteomic exploration of interaction orthogonality in all existing TF families in human cells. This might also lead to exciting insights on disease mutations dysregulating epigenetic modifications as e.g., in many cancers71. Beyond gene expression, orthogonal molecule grammar might also provide a universal mechanistic basis for the cell to organise targeted (enzymatic) reactions in its highly crowded intracellular environment through transient compartmentalization. Moreover, the understanding of orthogonal interactions used in cells could also be of interest to scientists seeking inspiration from nature in the design of nanoscale soft-matter systems, such as the development of nanoscale reactors89 or containers for targeted drug delivery90. In closing, we believe that these insights will provide an important step forward in our understanding of transcriptional selectivity and might also have profound implications in the broad field of condensate biology in both health and disease.
Methods
Protein sequences
The exact sequences for the LCDs used for the work, as shown in Fig. 1, are contained in Supplementary Table 1 of the supporting information.
The 1 bead per amino acid (1BPA) molecular dynamics model
In this work we use the 1BPA model that was previously developed for the study of intrinsically disordered proteins in the nuclear pore complex (NPC)76,77. It has been used extensively to model the behaviour of the disordered nucleoporin regions which fill the center of the yeast NPC and provide a selective barrier to the transport of cargo between the cytoplasm and nucleoplasm78,79,91. In this work an updated 1BPA model is developed (1BPA-2.1) that extends the applicability domain beyond the intrinsically disordered domains of yeast nucleoporins, by incorporating a greater training set for parameterisation (collating experimental data from55,92,93,94,95), and further improves the performance of the previously used 1BPA-2.0 model72,80. The full details of the force field are given in the ESI.
Original 1BPA potential
As a starting point for the parameterisation in this work the 1BPA-cp version is used96,97 (referred to as 1BPA-1.0 in this work). The bonded potential, ϕb (equation (1)), consists of three components: bonding (ϕbond, equation (2)), bending (ϕbend) and torsion (ϕtorsion) components. The bonding potential, ϕbond is a simple harmonic potential where k = 8038 kJ mol−1nm−2 and b = 0.38 nm. An iterative Boltzmann inversion was used to fit Ramachandran data to generate the bending and torsion potentials, as described in76. During this analysis, the presence of glycine or proline residues was found to create distinctive Ramanchandran plots, which resulted in the definition of 6 different angular potentials and nine torsional potentials, described in Supplementary Table 2.
There are three components that constitute the non-bonded potential, ϕnb (equation (3)), of the 1BPA model: hydrophobic interactions (ϕhp, equation (4)), cation-π interactions (ϕcp, equation (5)) and electrostatic interactions (ϕel, equation (6)).
where \({\epsilon }_{ij}={\epsilon }_{{{\rm{hp}}}}\sqrt{{({\epsilon }_{i}{\epsilon }_{j})}^{\alpha }}\), σ = 0.6 nm, ϵrep = 10 kJ mol−1, α = 0.27, ϵhp = 13.0 kJ mol−1 and ϵi, ϵj are residue specific hydrophobicities. The cation-π interactions are represented by:
where ϵcp,ij is the energy of the cation-π interaction, and σcp = 0.45 nm is the radius used for cation-π interactions. The parameterisation of ϵcp,ij was undertaken by Jafarinia et al.96. The electrostatic interactions are represented by:
where Ss = 80 and z = 0.25 nm. The Debye screening coefficient, \(\kappa={\left({\epsilon }_{0}{\epsilon }_{r}{k}_{b}T\left.\right)/2{N}_{A}{e}^{2}I\right)}^{-0.5}\), where ϵ0 is the permittivity of free space, ϵr is the permittivity of water, kb is Boltzmann’s constant, NA isAvogadro’s number, e is the unit charge of an electron, T = 300 K and I equal to the experimental ion concentration (150 mM) unless specified otherwise.
Parameterisation data
Data originally used in the 1BPA parameterisation for FG-Nups is from Yamada et al.92, which contains the hydrodynamic radii (Rh) data for the FG-Nups of yeast. Additional data from the work of Dignon and coworkers for the HPS model94 and the Mpipi model from Joseph et al.93 contains experimental Rg information on a broader set of IDPs. Additional data on hnRNPA variants from Bremer et al.55 and human Nups from Kapinos et al.95 were incorporated in the reparameterisation. This information is included in Supplementary Table 8. The total number of IDPs with single molecule data now totals 66 molecules, with a wider variance in charged residue content.
Updated model parameters in the 1BPA-2.0 and 1BPA-2.1 models
The updates to the basic 1BPA-1.0 model96,97 are included in the following tables detailing the changes made: Supplementary Table 3 for changes to global parameters, Supplementary Table 4 for hydrophobicity changes, Supplementary Table 5 for aromatic residue hydrophobicity changes, Supplementary Table 6 for cation-π interaction changes, and Supplementary Table 7 for the neutral hydrophobic potential pairs. These parameters were found to reduce the mean error in δ(Rg/h), while maintaining performance on the Yamada FG-Nups data92 used in the original parameterisation.
Droplet simulation protocol
To be able to explore the behaviour of the transcription condensates we used droplet formation simulations to be able to look at the internal structure. We start by forming a condensed phase droplet at the start of the simulation, which is then inserted into an empty dilute phase. If PS is favoured, this droplet structure should remain stable throughout the subsequent simulation; if unstable this droplet would break up into a dilute phase of monomers. This is done because self-assembly and clustering of individual monomers into phase separated condensates can be a slow process to observe.
Molecules are placed in the initial cubic simulation box (using a random initial conformation), with their centre of mass placed upon a regular grid, with a small buffer region to avoid overlap between molecules. A temperature of 300 K and a timestep of 20 fs are used for all simulations in this work. For equilibration of the droplet, energy minimisation on the initial configuration is used (energy tolerance of 1 kJ mol−1 nm−1), then 50 ns NVT Langevin dynamics simulations (Nosé-hoover thermostat with τt = 100 ps), followed by 500 ns NPT Langevin dynamics (Nosé-hoover thermostat with τt = 100 ps and a Berendsen barostat with τp = 10 ps, 1 bar reference pressure and a compressiblity of 4.5 × 10−5 bar−1). The end state of the NPT equilibration step is inserted into a new periodic box with a volume chosen to give a total residue density of 80,000 μM, after recentering on the center of mass and after the molecules have been unwrapped through the previous periodic boundary conditions. A second energy minimisation step is applied in the new simulation box to relax the molecules after expansion of the box (energy tolerance of 1 kJ mol−1 nm−1). A final 3 μs NVT production run (Nosé-hoover thermostat with τt = 100 ps) is used for data collection. The trajectory is sampled every 5 ns to determine whether convergence was reached (see methods for details on molecular connectivity graph creation and convergence testing). Convergence information for a selection of systems is shown in Fig. 3.
Simulation analysis using a molecular connectivity graph
To assess condensate stability in the droplet simulations, a molecular connectivity graph, consisting of nodes and edges, is used to describe the clustering and assess the time evolution of the molecular network during the simulation. Each simulation frame can be described by a connectivity graph. In the graph representation each node corresponds to a unique molecule, with an edge between the nodes denoting the number of non-zero interactions (if zero interactions are present no edge is added). Within an edge information can be stored about the number and type of contacts between molecules, based on residue categorisation. An interaction was determined to be present using a cutoff of 0.7 nm between 1BPA residues. The cutoff of 0.7 nm represents the bead diameter (0.6 nm) plus an additional 0.1 nm buffer to account for thermal fluctuations, as previously explored in Ghavami et al.91. Computation and processing of the contact matrix for a simulation frame allows the creation of the molecule graph. This process is shown in Supplementary Fig. 2.
Readouts from a molecular connectivity graph
A molecular connectivity graph computed for a simulation can be used to get several different readouts about a simulation system. Information on the size of clusters can be found by identifying the discrete sub-graphs of nodes with only internal connections (no contacts outside of the cluster). Through assessment of the time evolution of the number of clusters, grouped based on number of molecules, and the number of molecules in clusters within specific size categories it is possible to determine whether the system has reached equilibrium. After convergence is established it is possible to examine the distribution of components within the different clusters. The percentage of species i in a cluster is computed using Ni∈C/(NframesNi) where Ni∈C is the number of molecules of i in a cluster of size C.
Contact map definition
From the system graph creation process, contact matrices for all molecule combinations, over all frames are computed. This information is also useful for understanding the specific residues which drive condensation. As such, this information is aggregated during graph computation. Contact maps can be divided into two categories: intramolecular contact maps (interactions between particles in the same molecule copy), corresponding to the diagonal sub-matrices (Iii) in Supplementary Fig. 2, and intermolecular contact maps (interactions between particles of two different molecules), corresponding to the off-diagonal sub-matrices (Iij) in Supplementary Fig. 2. Since individual molecules of the same type are indistinguishable at the macroscale, only unique combinations of molecules need to be stored in different matrices, such that a single contact map for each intermolecular pairing and intramolecular contact map is computed. These maps contain the data for all copies of the same molecule pairs and for all timeframes studied. The contact maps by particle index (Figs. 3a–c and 4a–c in manuscript and Supplementary Figs. 34-60 parts A,D,G- if D or G present) display the contact information as a contact probability: the contact information is normalised by the number of frames, Nframes and (\({({N}_{i}{N}_{j})}^{0.5}\)) where Ni and Nj are the number of copies of molecule types i and j in the simulation.
When the contact information is presented based on residue type in a molecule (Figs. 4d–f and 5d–f in manuscript and Supplementary Figs. 34-60 parts B,E,H- if E or H present), a summation over the information in the contact maps by particle index to group interactions by residue type (equivalent to a matrix reduction operation). The presentation of normalised residue contact maps (Supplementary Figs. 34-60 parts C,F,I- if F or I present) applies a second normalisation step to the contact map by residue type of \({({N}_{i,r1}{N}_{j,r2})}^{0.5}\), with Nj,r1 the number of residue r1 in molecule i and Nj,r2 the number of residue r2 in molecule j.
Interaction summary information
Data on the distribution of interactions between different interaction types is also extracted from the contact maps for a system. The contact data is aggregated by summation over the contact maps and grouped into five categories of attractive interactions: aromatic, aliphatic, aliphatic-aromatic, cation-π, electrostatic. To compute the fraction of interactions, Fint, between two species (Figs. 3d–f and 4d–f in manuscript) the number of interactions is divided by the total interactions between the species. The fraction of interactions, Fint, for a system (Supplementary Figs. 3 and 4) is the sum of the interactions from all contact maps (intramolecular and intermolecular) in the system divided by the total number of all interactions in the system. The theoretical upper bound of the number of interactions that are possible between two classes of residue types is NiNj, where Ni and Nj are the number of residues of type i and j, respectively, in the simulation. By normalising the number of actual interactions by the upper bound NiNj it is possible to compute the mean interaction propensity for a ternary mixture. This was done for FUS-SP1-TAF15 mixtures to generate the data in Supplementary Fig. 5. From these mean interaction propensities in the table in Supplementary Fig. 5A, we compute the possible fraction of interactions we expect to see in any FUS-SP1-TAF15 stochiometry by multiplying the interaction propensities in the table of Supplementary Fig. 5A with the number of residue types in the composition, as shown in Supplementary Fig. 5B-F.
The propensity for a specific interaction type is the sum of the interactions from all contact maps (intramolecular and intermolecular) in the system divided by the theoretical upper limit for the number of interactions, NiNj, where Ni, Nj are the number of residues of type i and j that form the interaction. The mean propensity (Fig. 7a) is the average propensity from all systems at 150mM ion concentration (34 compositions). To be able to generate the interaction triangles displaying Fint for all compositions (Fig. 7b–f), the mean propensity in Fig. 7a is multiplied by NiNj in the system (to get the expected number of interactions for the given composition), and divided by the expected total number of interactions (0.026NtotNtot).
Intermolecular contact propensity between two molecules of different types (in Fig. 6, Supplementary Figs. 6-8, 11-16 and Supplementary Tables 9-19) is computed by summation of the interactions in the corresponding intermolecular contact map for the species involved. To enable comparison between different compositions normalisation by the theortical upp NiLiNjLj is used, where Ni and Nj are the number of copies of molecule types i and j, and Li and Lj are the number of simulation residues per molecule of i and j, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Supplementary information contains the protein sequences, 1BPA-v2.1 model parameters and a comparison to 1BPA-v1.0 and 1BPA-v2.0, simulation images from large scale simulations, molecular interaction data, contact lifetimes, ternary phase diagrams for the additional ternary systems, intermolecular contact maps for systems discussed in the main text. Source Data are provided with this paper for the main manuscript figures and Supplementary Figs. 1, 3-5, 6-10, 28-33. The high resolution images for Supplementary Figs. 18-27, and 34-60 can be found in the source data. The source data for Supplementary Figs. 6, 18-27, and 34-60 (additional contact maps) can be provided by the corresponding author upon request after extraction from the simulation trajectories. The simulation inputs (including forcefield files compatible with GROMACS) can be found on github [https://github.com/Onck-group/transcriptioncondensatesimulations] or Zenodo [https://doi.org/10.5281/zenodo.14836530]. The corresponding simulation trajectories are also available upon request due to the large total volume of data (over 2 TB). Source data are provided with this paper.
Code availability
All MD simulations were run with the GROMACS 2019.6 package. Analysis was undertaken using MDAnalysis v2.3.0, with matplotlib v3.6.2 used for plotting data and VMD v1.9.3 for simulation visualisation. The simulation inputs (including forcefield files compatible with GROMACS) can be found on github [https://github.com/Onck-group/transcriptioncondensatesimulations] or Zenodo [https://doi.org/10.5281/zenodo.14836530].
References
Cramer, P. Organization and regulation of gene transcription. Nat. 573, 45–54 (2019).
Roeder, R. G. 50+ Years of Eukaryotic Transcription: an Expanding Universe of Factors and Mechanisms. Nat. Struct. Mol. Biol. 26, 783–791 (2019).
Barba-Aliaga, M., Alepuz, P. & Pérez-Ortín, J. E. Eukaryotic RNA Polymerases: The Many Ways to Transcribe a Gene. Front. Mol. Biosci. 8, 1–8 (2021).
Allen, B. L. & Taatjes, D. J. The mediator complex: a central integrator of transcription. Nature Reviews Molecular Cell Biology 16, 155–166 (2015).
Richter, W. F., Nayak, S., Iwasa, J. & Taatjes, D. J. The mediator complex as a master regulator of transcription by rna polymerase ii. Nature Reviews Molecular Cell Biology 23, 732–749 (2022).
Andersson, R. Promoter or enhancer, what’s the difference? Deconstruction of established distinctions and presentation of a unifying model. BioEssays 37, 314–323 (2015).
Browning, D. F. & Busby, S. J. W. The regulation of bacterial transcription initiation. Nat. Rev. Microbiol. 2, 57–65 (2004).
Browning, D. F. & Busby, S. J. W. Local and global regulation of transcription initiation in bacteria. Nat. Rev. Microbiol. 14, 638–650 (2016).
Carter, R. & Drouin, G. Structural differentiation of the three eukaryotic RNA polymerases. Genomics 94, 388–396 (2009).
Hahn, S. Structure and mechanism of the RNA polymerase II transcription machinery. Nat. Struct. Mol. Biol. 11, 394–403 (2004).
Duttke, S. H. C. Evolution and diversification of the basal transcription machinery. Trends Biochem. Sci. 40, 127–129 (2015).
Rippe, K., Papantonis, A. Rna polymerase ii transcription compartments: from multivalent chromatin binding to liquid droplet formation? Nat. Rev. Mol. Cell Biol. 0123456789 https://doi.org/10.1038/s41580-021-00401-6 (2021).
Watson, M. & Stott, K. Disordered domains in chromatin-binding proteins. Essays Biochem. 63, 147–156 (2019).
Palikyras, S. & Papantonis, A. Modes of phase separation affecting chromatin regulation. Open biol. 9, 190167 (2019).
Emmert-Streib, F., Dehmer, M. & Haibe-Kains, B. Gene regulatory networks and their applications: Understanding biological and medical problems in terms of networks. Front. Cell Dev. Biol. 2, 1–7 (2014).
Shin, Y. et al. Liquid nuclear condensates mechanically sense and restructure the genome. Cell 175, 1481–149113 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–185516 (2018).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Fang, X. et al. Global transcriptional regulatory network for Escherichia coli robustly connects gene expression to transcription factor activities. Proc. Natl. Acad. Sci. USA 114, 10286–10291 (2017).
Fang, L. et al. GRNdb: Decoding the gene regulatory networks in diverse human and mouse conditions. Nucleic Acids Res. 49, 97–103 (2021).
Iacono, G., Massoni-Badosa, R. & Heyn, H. Single-cell transcriptomics unveils gene regulatory network plasticity. Genome Biol. 20, 1–20 (2019).
Zhang, S. et al. Inference of cell type-specific gene regulatory networks on cell lineages from single cell omic datasets. Nat. Commun. 14, 3064 (2023).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Soto, L. F. et al. Compendium of human transcription factor effector domains. Molecular Cell 82, 514–526 (2022).
Zou, Z., Ohta, T., Miura, F. & Oki, S. Chip-atlas 2021 update: a data-mining suite for exploring epigenomic landscapes by fully integrating chip-seq, atac-seq and bisulfite-seq data. Nucleic Acids Res. 50, 175–182 (2022).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, 3958 (2018).
Wang, X., Cairns, M. J. & Yan, J. Super-enhancers in transcriptional regulation and genome organization. Nucleic Acids Res. 47, 11481–11496 (2019).
Sabari, B. R., Dall’Agnese, A. & Young, R. A. Biomolecular Condensates in the Nucleus. Trends Biochem. Sci. 45, 961–977 (2020).
Zhang, J. et al. Super enhancers—functional cores under the 3d genome. Cell Proliferation 54 https://doi.org/10.1111/cpr.12970 (2021).
Silva Pinheiro, E., Preato, A. M., Petrucci, T. V. B., Santos, L. S. D. & Glezer, I. Phase-separation: a possible new layer for transcriptional regulation by glucocorticoid receptor. Front. Endocrinol. (Rome, Italy) 14, 1160238 (2023).
Peng, L., Li, E. M. & Xu, L. Y. From start to end: Phase separation and transcriptional regulation. Biochim. Biophys. Acta, Gene Regul. Mech. 1863, 194641 (2020).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, 2555 (2018).
Wei, M. T. et al. Nucleated transcriptional condensates amplify gene expression. Nat. Cell Biol. 22, 1187–1196 (2020).
Chong, S. & Mir, M. Towards Decoding the Sequence-Based Grammar Governing the Functions of Intrinsically Disordered Protein Regions. J. Mol. Biol. 433, 166724 (2021).
Shrinivas, K. et al. Enhancer features that drive formation of transcriptional condensates. Mol. Cell 75, 549–5617 (2019).
Sharp, P.A., Chakraborty, A.K., Henninger, J.E., Young, R.A. Rna in formation and regulation of transcriptional condensates https://doi.org/10.1261/rna.078997.121 (2022).
Chen, Q. et al. Enhancer rnas in transcriptional regulation: recent insights. Front. Cell Dev. Biol. 11 https://doi.org/10.3389/fcell.2023.1205540 (2023).
Demmerle, J., Hao, S., Cai, D. Transcriptional condensates and phase separation: condensing information across scales and mechanisms. Nucleus 14 https://doi.org/10.1080/19491034.2023.2213551 (2023).
Brangwynne, C. P. et al. Germline P Granules Are Liquid Droplets That Localize by Controlled Dissolution/Condensation. Science 324, 1729–1732 (2009).
Hyman, A. A., Weber, C. A. & Jülicher, F. Liquid-liquid phase separation in biology. Annu. Rev. Cell Dev. Biol. 30, 39–58 (2014).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Han, X. et al. Roles of the brd4 short isoform in phase separation and active gene transcription. Nat. Struct. Mol. Biol. 27, 333–341 (2020).
Zamudio, A. V. et al. Mediator condensates localize signaling factors to key cell identity genes. Mol. Cell 76, 753–7666 (2019).
Changiarath, A. et al. Promoter and gene-body rna-polymerase ii co-exist in partial demixed condensates. biorxiv https://doi.org/10.1101/2024.03.16.585180 (2024)
Flory, P. J. Thermodynamics of high polymer solutions. J. Chem. Phys. 10, 51–61 (1942).
Huggins, M. L. Solutions of long chain compounds. J. Chem. Phys. 9, 440–440 (1941).
Oh, S. Y. & Bae, Y. C. Liquid-liquid equilibria for ternary polymer mixtures. Chem. Phys. 379, 128–133 (2011).
Thomas, S., Durand, D., Chassenieux, C. & Jyotishkumar, P. Handbook of Biopolymer-Based Materials : From Blends and Composites to Gels and Complex Networks, p. 875. Wiley-VCH, ??? (2013)
Liu, D. et al. Phase behavior and interfacial tension of ternary polymer mixtures with block copolymers. RSC Advances 11, 38316–38324 (2021).
Bot, A. & Venema, P. Phase behavior of ternary polymer mixtures in a common solvent. ACS Omega 8, 28387–28408 (2023).
Sabari, B. R. Biomolecular Condensates and Gene Activation in Development and Disease. Dev. Cell 55, 84–96 (2020).
Fare, C.M., Villani, A., Drake, L.E. & Shorter, J. Higher-order organization of biomolecular condensates. Open Biol. 11 https://doi.org/10.1098/rsob.210137 (2021).
Murthy, A. C. et al. Molecular interactions underlying liquid-liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Bremer, A. et al. Deciphering how naturally occurring sequence features impact the phase behaviours of disordered prion-like domains. Nat. Chem. 14, 196–207 (2022).
Choi, J. M., Holehouse, A. S. & Pappu, R. V. Physical Principles Underlying the Complex Biology of Intracellular Phase Transitions. Annu. Rev. Biophys. 49, 107–133 (2020).
Kaur, T. et al. Sequence-encoded and composition-dependent protein-rna interactions control multiphasic condensate morphologies. Nat. Commun. 12, 872 (2021).
Chew, P. Y., Joseph, J. A., Collepardo-Guevara, R. & Reinhardt, A. Thermodynamic origins of two-component multiphase condensates of proteins. Chem. Sci. 14, 1820–1836 (2023).
Chew, P. Y., Joseph, J. A., Collepardo-Guevara, R. & Reinhardt, A. Aromatic and arginine content drives multiphasic condensation of protein-rna mixtures. Biophys. J. 123, 1342–1355 (2024).
Rekhi, S. et al. Expanding the molecular language of protein liquid-liquid phase separation. Nature Chemistry 16, 1113–1124 (2024).
Welles, R. M. et al. Determinants that enable disordered protein assembly into discrete condensed phases. Nat. Chem. 16, 1062–1072 (2024).
Rana, U. et al. Asymmetric oligomerization state and sequence patterning can tune multiphase condensate miscibility. Nat. Chem. 16, 1073–1082 (2024).
Farag, M., Borcherds, W.M., Bremer, A., Mittag, T., Pappu, R.V. Phase separation of protein mixtures is driven by the interplay of homotypic and heterotypic interactions. Nat. Commun. 14 https://doi.org/10.1038/s41467-023-41274-x (2023).
Lee, J., Cho, H. & Kwon, I. Phase separation of low-complexity domains in cellular function and disease. Exp. Mol. Med. 54, 1412–1422 (2022).
Zaborowska, J., Egloff, S. & Murphy, S. The pol ii ctd: New twists in the tail. Nat. Struct. Mol. Bio. 23, 771–777 (2016).
Sawicka, A. et al. Transcription activation depends on the length of the RNA polymerase II C-terminal domain. EMBO J. 40, 1–17 (2021).
Shan, L. et al. Sp1 undergoes phase separation and activates rgs20 expression through super-enhancers to promote lung adenocarcinoma progression. Proc. Natl Acad. Sci. USA 121 https://doi.org/10.1073/pnas.2401834121 (2024).
Murthy, A. C. et al. Molecular interactions contributing to fus sygq lc-rgg phase separation and co-partitioning with rna polymerase ii heptads. Nat. Struct. Mol. Bio. 28, 923–935 (2021).
Flores-Solis, D. et al. Driving forces behind phase separation of the carboxy-terminal domain of rna polymerase ii. Nat. Commun. 14 https://doi.org/10.1038/s41467-023-41633-8 (2023).
Zuo, L. et al. Loci-specific phase separation of FET fusion oncoproteins promotes gene transcription. Nat. Commun. 12 https://doi.org/10.1038/s41467-021-21690-7 (2021).
Ahn, J. H. et al. Phase separation drives aberrant chromatin looping and cancer development. Nature 595 (January 2020) https://doi.org/10.1038/s41586-021-03662-5 (2021).
Kind, L. et al. Structural properties of the hnf-1a transactivation domain. Front. Mol. Biosci. 10 https://doi.org/10.3389/fmolb.2023.1249939 (2023).
Kang, J., Lim, L., Lu, Y. & Song, J. A unified mechanism for llps of als/ftld-causing fus as well as its modulation by atp and oligonucleic acids. PLOS Biol. 17, 3000327 (2019).
Hirose, T., Ninomiya, K., Nakagawa, S., Yamazaki, T. A guide to membraneless organelles and their various roles in gene regulation. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-022-00558-8 (2022)
Mann, R. & Notani, D. Transcription factor condensates and signaling driven transcription. Nucleus (Austin, Tex.) 14, 2205758 (2023).
Ghavami, A., Giessen, E. & Onck, P. R. Coarse-Grained Potentials for Local Interactions in Unfolded Proteins. J. Chem. Theory Comput. 9, 432–440 (2013).
Ghavami, A., Veenhoff, L. M., Giessen, E. & Onck, P. R. Probing the Disordered Domain of the Nuclear Pore Complex through Coarse-Grained Molecular Dynamics Simulations. Biophys. J. 107, 1393–1402 (2014).
Fragasso, A. et al. A designer FG-Nup that reconstitutes the selective transport barrier of the nuclear pore complex. Nat. Commun. 12, 2010 (2021).
Dekker, M., Giessen, E.V., Onck, P.R. Phase separation of intrinsically disordered fg-nups is driven by highly dynamic fg motifs. Proc. Natl. Acad. Sci. USA 120 https://doi.org/10.1073/pnas.2221804120 (2023).
Heesink, G. et al. Exploring intra- and inter-regional interactions in the idp α-synuclein using smfret and md simulations. Biomacromolecules 24 https://doi.org/10.1021/acs.biomac.3c00404 (2023).
Schwartz, J. C., Wang, X., Podell, E. R. & Cech, T. R. Rna seeds higher-order assembly of fus protein. Cell Rep. 5, 918–925 (2013).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like rna binding proteins. Cell 174, 688–69916 (2018).
Peng, K., Radivojac, P., Vucetic, S., Dunker, A. K. & Obradovic, Z. Length-dependent prediction of protein intrinsic disorder. BMC Bioinformatics 7, 208 (2006).
Driver, M. D., Postema, J. & Onck, P. R. The effect of dipeptide repeat proteins on fus/tdp43-rna condensation in c9orf72 als/ftd. The Journal of Physical Chemistry B 128, 9405–9417 (2024).
Jafarinia, H., Giessen, E. & Onck, P. R. Phase separation of toxic dipeptide repeat proteins related to c9orf72 als/ftd. Biophys. J. 119, 843–851 (2020).
Qian, D. I. et al. Dominance analysis to assess solute contributions to multicomponent phase equilibria 121 https://doi.org/10.1073/pnas (2024).
Boehning, M. et al. Rna polymerase ii clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Bio. 25, 833–840 (2018).
Alshareedah, I., Moosa, M. M., Pham, M., Potoyan, D. A. & Banerjee, P. R. Programmable viscoelasticity in protein-rna condensates with disordered sticker-spacer polypeptides. Nat. Commun. 12, 6620 (2021).
Cao, S. et al. Dipeptide coacervates as artificial membraneless organelles for bioorthogonal catalysis. Nat. Commun. 15, 39 (2024).
Garabedian, M. V. et al. Designer membraneless organelles sequester native factors for control of cell behavior. Nat. Chem. Biol. 17, 998–1007 (2021).
Ghavami, A., Van der Giessen, E. & Onck, P. R. Sol-gel transition in solutions of FG-Nups of the nuclear pore complex. Extreme Mech. Lett. 22, 36–41 (2018).
Yamada, J. et al. A Bimodal Distribution of Two Distinct Categories of Intrinsically Disordered Structures with Separate Functions in FG Nucleoporins. Mol. Cell. Proteomics 9, 2205–2224 (2010).
Joseph, J. A. et al. Physics-driven coarse-grained model for biomolecular phase separation with near-quantitative accuracy. Nat. Comput. Sci. 1, 732–743 (2021).
Dignon, G. L., Zheng, W., Kim, Y. C., Best, R. B. & Mittal, J. Sequence determinants of protein phase behavior from a coarse-grained model. PLOS Comput. Biol. 14, 1005941 (2018).
Kapinos, L. E., Schoch, R. L., Wagner, R. S., Schleicher, K. D. & Lim, R. Y. H. Karyopherin-Centric Control of Nuclear Pores Based on Molecular Occupancy and Kinetic Analysis of Multivalent Binding with FG Nucleoporins. Biophys. J. 106, 1751–1762 (2014).
Jafarinia, H., Giessen, E. V. & Onck, P. R. Molecular basis of c9orf72 poly-pr interference with the β-karyopherin family of nuclear transport receptors. Sci. Rep. 12, 21324 (2022).
Fragasso, A. et al. Transport receptor occupancy in nuclear pore complex mimics. Nano Res. 15, 9689–9703 (2022).
Acknowledgements
We thank H.G.S. Arends and J. Postema for discussions. We thank the oLife COFUND project for funding MDD. The COFUND project oLife has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 847675. We thank the Center for Information Technology of the University of Groningen for their support and for providing access to the Peregrine and Habrok high performance computing clusters. This work made use of the Dutch national e-infrastructure with the support of the SURF Cooperative using grant no. EINF-3233 (MDD) and EINF-5917 (MDD).
Author information
Authors and Affiliations
Contributions
M.D.D. and P.R.O. designed the research. M.D.D. carried out all the simulations, analysed the data, and wrote the manuscript. P.R.O. reviewed and edited the article and supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Driver, M.D., Onck, P.R. Selective phase separation of transcription factors is driven by orthogonal molecular grammar. Nat Commun 16, 3087 (2025). https://doi.org/10.1038/s41467-025-58445-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58445-7
This article is cited by
-
Intrinsically disordered domains expand the CAR T cell toolbox
Nature Chemical Biology (2025)