Abstract
Coxiella burnetii (Cb), the causative agent of Q fever, replicates within host macrophages by modulating immune responses through poorly understood mechanisms. Long non-coding RNAs (lncRNAs) are crucial yet underexplored regulators of inflammation, particularly in Cb pathogenesis. Employing a comparative transcriptomic analysis of THP-1 macrophages infected with 16 different microbes, we dissect a core set of immune-responsive lncRNAs such as MAILR, LINC01215, PACER, and MROCKI-common to human anti-pathogen responses, and distinguish them from lncRNAs specifically altered at early (1 h) time points in individual infections. In particular, our approach identifies lncRNA CYP1B1-AS1 as specifically upregulated in a spatiotemporal manner along with CYP1B1 in cis during Cb infection. Promoter assays confirm their co-regulation via a shared bidirectional promoter, while aryl hydrocarbon receptor (AHR)-lucia luciferase and nuclear translocation assays demonstrate that Cb infection activates AHR, driving their transcription. Knockdown of CYP1B1-AS1 or CYP1B1 alone disrupts mitochondrial homeostasis, increases ROS and mitochondrial dysfunction, and exacerbates apoptosis during infection. These findings position the CYP1B1-AS1/CYP1B1 axis as a key regulator of mitochondrial homeostasis under AHR signaling, supporting an intracellular environment that benefits Cb replication. Our results highlight the critical roles of lncRNAs in immune regulation and provide a valuable resource for future lncRNA research.
Similar content being viewed by others
Introduction
Coxiella burnetii (Cb), the causative agent of Q fever, is a highly adapted obligate intracellular Gram-negative pathogen that primarily infects monocytes and macrophages. It establishes a replicative niche within lysosome-matured Coxiella-containing vacuoles (CCVs)1, a niche that is non-permissive to most intracellular pathogens2. Cb utilizes a type IVB secretion system (T4SS) to translocate effectors into host cells, thereby manipulating endocytic trafficking, immune evasion, to promote intracellular replication3,4,5. The Nine Mile strain of Cb, commonly used in research, exists in two variants: phase I (NMI, virulent), expressing a full-length lipopolysaccharide (LPS), and phase II (NMII, avirulent), characterized by truncated LPS6. Infection typically occurs via inhalation, causing acute flu-like illness, but chronic cases can lead to severe complications like endocarditis and lung fibrosis7. Long-term antibiotics are the standard treatment, highlighting the need for new therapies and diagnostics.
A hallmark of Cb infection is its ability to evade innate immune activation distinguishing it as a stealth pathogen1,8,9. In contrast, other well-characterized intracellular pathogens like Mycobacterium tuberculosis10, Legionella pneumophila11, and Salmonella enterica subsp. enterica serovar Typhimurium12, actively engage cytosolic sensors such as cGAS, NLRs, and TLRs, triggering a robust inflammatory response13,14. Despite significant research efforts, the molecular mechanisms by which Cb evade these immune activations remain poorly understood.
Long non-coding RNAs (lncRNAs) are regulatory transcripts over 200 nucleotides, and have emerged as important regulators of inflammation and immune responses15,16. Macrophages are the first line of immune defense, recognizing pathogen-associated molecular patterns (PAMPs), such as LPS, via pattern recognition receptors (PRRs) to activate central innate immune signaling pathways involving NF-κB and type I interferon pathways (IFN-I) against intracellular infections17. Recent comprehensive lncRNA transcriptome studies in human macrophages have revealed the significant involvement of lncRNAs such as MAILR, LINC01215, LUCAT1, PIRAT1, AC010980, LINC00158 and MROCKI in regulating NF-κB and IFN-I response during infection17,18,19. Other well-characterized lncRNAs, such as NEAT1, GAPLINC, HOTAIR, PACER, and LINC-COX2, are induced upon LPS stimulation, and infections caused by M. tuberculosis, S. Typhimurium, HCV and HIV-1, promoting pro-inflammatory response and pathogen restriction20,21. Despite these discoveries, most lncRNAs expressed in human immune cells remain functionally uncharacterized, likely due to their complex molecular interactions and context-dependent expression patterns, which hinder mechanistic investigation.
The involvement of lncRNAs in Cb infection has not been explored. Considering Cb’s unique immune evasion strategy and its tropism for macrophages, we hypothesized that Cb elicits a distinct lncRNA expression profile compared to other bacterial infections. To test this, we employed a comparative multi-pathogen RNA-sequencing (RNA-seq) strategy in THP-1-derived macrophages infected with 16 different microbes. This systems-level strategy identified a core set of lncRNAs commonly responsive to any pathogen confrontation (herein termed as immune-responsive lncRNAs) and distinguished them from lncRNAs uniquely altered in individual infections. Notably, this strategy identified lncRNAs specifically regulated during Cb infection, and provided insights into their temporal dynamics in host defense.
Among these, CYP1B1-AS1 emerged as a key lncRNA upregulated during Cb infection. Our in vitro studies demonstrated that CYP1B1-AS1 is co-regulated with its neighboring gene CYP1B1 via a shared bidirectional promoter. Transcriptional assays and luciferase reporter analyses revealed that this co-regulation is mediated by activation of the transcription factor (TF)-aryl hydrocarbon receptor (AHR). Functional analyses showed that silencing of CYP1B1-AS1 or CYP1B1 resulted in mitochondrial dysfunction, elevated reactive oxygen species (ROS), and apoptosis exacerbated by Cb infection. Furthermore, silencing either of the genes augmented pro-inflammatory cytokine production, implicating this axis in maintaining mitochondrial homeostasis and regulating ROS-associated inflammation.
By establishing the CYP1B1-AS1/CYP1B1 axis as an AHR-regulated, infection-responsive module, our study provides new insights into host-Coxiella interactions. Importantly, the comparative transcriptomic framework provides a scalable platform for identifying lncRNA in human macrophages and serves as a valuable resource for future investigations into immunity and infection.
Results
Identification of infection- and immune-responsive lncRNAs in human macrophages
To investigate lncRNA expression profiles across different bacterial infections and distinguish pathogen-specific responses from common macrophage immune programs, we conducted a comprehensive comparative transcriptomic analysis.
Phorbol 12-myristate-12 acetate (PMA)-differentiated THP-1 cells (THP-1 macrophages) were infected for 1 h with a panel of 16 bacterial strains, including three C. burnetii Nine Mile Phase II (Cb; BL2 variant as WT), and two attenuated mutants dotA::Tn (CbA/ΔdotA), and dotB::Tn (CbB/ΔdotB), alongside 13 additional bacterial species with varying pathogenicity and non-pathogenic controls (Supplementary Data 1). These included (a) Pathogenic bacteria: Enterohemorrhagic Escherichia coli O157 (EcT; EHEC), Enterohemorrhagic Escherichia coli O157 Δstx (nontoxigenic) (EcN; EHECΔstx), Francisella novicida U112 (Fn), Pseudomonas aeruginosa PAO1 (Pa), Staphylococcus aureus JE2 (Sa), S.Typhimurium (STm), Brucella melitensis ΔvjbR (Bmv), and (b) Opportunistic pathogen: Enterococcus faecalis (Ef), Rhizobium radiobacter (Rr), Micrococcus luteus (Ml), and (c) three non-pathogenic controls: Escherichia coli DH5α (Ec5), Listeria innocua (Li), and Bacillus subtilis P31K6 (Bs). The sample preparation, RNA sequencing, and data analysis workflow is outlined in Supplementary Fig. 1a, b.
Considering Cb’s stealth-like infection strategy and poor innate immune activation, we hypothesized that it would evoke a distinct lncRNA expression profile compared to other pathogens. To evaluate this, we implemented a three-step analysis pipeline: (a) RNA-seq data were used to identify commonly differentially expressed (DE) lncRNAs across ≥4 infections, designated as immune-responsive lncRNAs; (b) we extracted DE-lncRNAs specific to individual infections, which serve as pathogen-specific lncRNAs;(c) comparative analyses were conducted to identify DE-lncRNAs uniquely induced or absent during Cb infection at 1 h post-infection (p.i.).
We identified 2348 DE-lncRNAs (log2 fold-change (FC) ≥ 1 or ≤−1; p < 0.05) across all infection conditions compared to uninfected (Mock) controls (Supplementary Data 2). Principal component analysis (PCA) revealed 33.98% variance between mock and infected macrophages, with 14.42% variance across infection groups (Fig. 1a). An UpSet analysis (Supplementary Fig. 1c) was performed to identify immune-responsive lncRNAs and pathogen-responsive lncRNA (Fig. 1b). Differential transcriptomic profiles across all infections are illustrated in Supplementary Fig.2a–m.
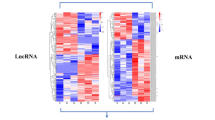
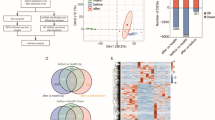
a Principal component analysis (PCA) of RNA-seq data illustrating transcriptional variance across infections, including C. burnetii Nine Mile Phase II (Cb), C. burnetii Nine Mile Phase II dotA::Tn (CbA), C. burnetii Nine Mile Phase II dotB::Tn (CbB), Escherichia coli DH5α (Ec5), enterohemorrhagic E. coli O157 (EcT/EHEC), E. coli O157Δstx (EcN/EHECΔstx), Bacillus subtilis P31K6 (Bs), Francisella novicida U112 (Fn), Pseudomonas aeruginosa PAO1 (Pa), Staphylococcus aureus JE2 (Sa), Salmonella enterica subsp. Typhimurium SL1344 (STm), Rhizobium radiobacter (Rr), Micrococcus luteus (Ml), Listeria innocua (Li), Enterococcus faecalis (Ef), and Brucella melitensis ΔvjbR (Bmv). PCA was performed using normalized RNA-seq data to assess global transcriptional variance. Infected samples are denoted by circles; mock-infected controls as triangles. Data points are color-coded by infection. b Experimental workflow for identifying common and pathogen-specific lncRNAs following infection of THP-1 macrophages. Schematics created using BioRender.com. c Bar graph summarizing the number of pathogen-specific differentially expressed (DE) lncRNAs identified across the infections analyzed. d Heatmap showing quantitative real-time PCR (RT-qPCR) validation of selected immune-regulatory lncRNAs that are either commonly regulated across infections or specifically altered during C. burnetii infection. Expression normalized to ACTB and shown relative to mock-infected controls (set to 1). lncRNAs with a log₂ fold change ≥ 1.5 or ≤ 0.5 were considered DE. Data represent mean ± SD from three independent experiments (n = 3). Source data are provided as a Source Data file.
We identified 538 immune-responsive lncRNAs (Supplementary Data 3). Additionally, we identified 19, 208, and 165 DE-lncRNAs specific to Cb, CbA, and CbB infections, respectively (Fig. 1c). No significant DE-lncRNAs were detected in Ef- or Bmv-infected transcriptomes, likely due to low infectivity or delayed host response at this early time point, and therefore, we excluded them from further analysis.
Among the 538 common immune-responsive lncRNAs, several well-characterized immune-regulatory lncRNAs were identified, including MAILR, LINC01215, LUCAT1, MROCKI, MIR155HG, MIR222HG, PACER, and EGOT (Table 1, Supplementary Data 3). These lncRNAs have recently been shown to be induced in primary human macrophages upon microbial infection or pattern recognition receptor (PRR) stimulation17,19,22,23,24. For instance, MAILR promotes TRIF-IRF3 signaling by stabilizing Optineurin, enhancing IFN-β production. LUCAT1 suppresses immune activation via its interaction with hnRNPs and STAT1. LINC01215 and MAILR are robustly induced by L. pneumophila infection and LPS exposure, while MROCKI enhances inflammatory responses by repressing GATA2 and promoting NF-κB signaling in macrophages17,23. To validate their expression, we performed quantitative real-time PCR (RT-qPCR) analysis, which confirmed that the selected immune-responsive lncRNAs were induced across bacterial infections, in agreement with the RNA-seq findings. Notably, MAILR, LINC01215, MROCKI, and PACER were significantly upregulated upon L. pneumophila infection, which served as a positive control. This observation corroborates prior reports describing their early induction (1 h post LPS-challenge) and confirms their activation in L. pneumophila-infected macrophages (Fig. 1d). Boxplot analysis summarizing their expression trends across infections is shown in Supplementary Figs. 2n–o and 3a–d.
Together, these results define a conserved set of lncRNAs representing a core transcriptional program activated in macrophages as part of the anti-pathogen response17. Importantly, these commonly induced immune-responsive lncRNAs were either dampened or not significantly expressed during Cb infection (Fig. 1d), supporting the hypothesis that Cb evades early immune activation by eliciting a distinct lncRNA transcriptional profile.
Identification of Cb-associated lncRNA profile
We identified 1692 DE genes in Cb-infected macrophages, including 748 upregulated and 944 downregulated transcripts (Supplementary Data 4a). Hierarchical clustering analysis identified 1531 DE-mRNAs and 161 DE-lncRNAs (Fig. 2a, Supplementary Data 4b). The DE-lncRNAs comprised intergenic, antisense, cis-, and trans-acting transcripts, with several annotated as novel. In parallel, 679 DE-lncRNAs were identified in macrophages infected with the attenuated CbA strain. To assess Cb strain-specific modulation, RT-qPCR analysis was performed, validating the differential modulation of the candidates that were common to Cb and CbA infections (Fig. 2b, Supplementary Fig. 4a). Cb-specific lncRNAs showed no change in expression in cells infected with CbA (ΔdotA) (Supplementary Fig. 4b). Conversely, RT-qPCR analysis of CbA-specific lncRNAs recapitulated the RNA-seq findings in CbA (ΔdotA)-infected cells (Supplementary Fig. 4c, d). LUCAT1 and PIRAT1 were also identified among the known immune regulatory lncRNAs in our panel.
a Hierarchical clustering heatmap of differentially expressed (DE) lncRNA-mRNA pairs during C. burnetii (Cb, NMII) infection. Each row represents a gene, and each column represents a biological replicate (n = 4). Color intensity indicates normalized gene expression. b RT-qPCR validation of DE lncRNAs specific to Cb and CbΔdotA (CbA) infections. Gene expression normalized to ACTB and shown relative to mock-infected controls (set to 1). Data represent the mean ± SD from three independent experiments (n = 3). Statistical test: two-way ANOVA; exact p-values (in the same order as the asterisks): lnc-DKK2: 0.035; PKP4-AS1: 0.008; GSTCD-AS1: 0.0487; DDX11-AS1: 0.0432; ENSG00000285650: 0.045; ENSG00000261668: 0.008; ENSG00000273669: 0.0087; ENSG00000260430: 0.035; LBX2-AS1: 0.0279; LINC03072: 0.008; TLR8-AS1: <0.0001; LINC01232: 0.0002; SBF2-AS1: <0.0001; LINC00942: 0.002; LINC00926: 0.008; LINC01426: 0.004; NRAD1: 0.0326; UBR5-DT: 0.0086; PIRAT1: 0.0049; CYP1B1-AS1: <0.0001. c Scatter plot of DE lncRNAs and mRNAs during Cb infection, highlighting CYP1B1-AS1 (violet), lnc-DKK2 (blue), and their target mRNAs CYP1B1 (green) and DKK2 (yellow). d Receiver operating characteristic (ROC) curve analysis displaying the diagnostic value of lnc-DKK2 and CYP1B1-AS1 in infection models, with area under the curve (AUC) values at p < 0.01 and 95% confidence intervals (CI). e, f KEGG pathway enrichment analysis of mRNAs associated with DE lncRNAs: e upregulated, f downregulated. Statistical test: one-sided Fisher’s exact test; p-values adjusted using the Benjamini–Hochberg false discovery rate (FDR) method. Bar height represents −log10(p-value) with color intensity representing the proportion of transcripts associated with each pathway. Source data are provided in the Source Data file.
To prioritize lncRNAs for functional analysis, we selected candidates transcribed from distinct or bidirectional promoters, enabling gene-specific perturbation, and those positioned proximally to protein-coding genes with potential cis-regulatory roles in Cb pathogenesis, as exemplified by lncRNAs such as lincRNA-Cox2, UMLILO, and LOUP25,26,27.
Applying these criteria, we identified lncRNAs with potential cis-regulatory functions based on their Pearson correlation coefficient (PCC ≥ 0.85, capturing both positive and negative correlations) with adjacent mRNAs. This analysis identified 55 DE lncRNA–mRNA cis-acting pairs in Cb-infected macrophages (Supplementary Data 5). UpSet plot analysis had revealed 19 and 208 DE-lncRNAs in Cb- and CbA-infected macrophages, respectively, spanning both cis- and trans-acting categories. Among infection-associated lncRNAs, PIRAT1 was upregulated specifically in Cb-infected macrophages. RT-qPCR confirmed its upregulation in all Cb infections (Figs. 1d and 2b; Supplementary Fig. 4a). PIRAT1 is a nuclear lncRNA primarily expressed in CD14⁺ monocytes, known to recruit PU.1 and inhibit its binding to S100A8/S100A9 promoters, thereby suppressing NF-κB activation. During PAMP stimulation and in severe COVID-19, PIRAT1 is downregulated, coinciding with upregulation of LUCAT1, which amplifies alarmin production via NF-κB while suppressing JAK-STAT signaling19. Interestingly, we observed upregulation of LUCAT1 only in CbA infection, but not in Cb infection, suggesting that PIRAT1-mediated immune suppression may be retained in Cb, potentially counteracting LUCAT1-driven NF-κB activation. This observation supports the known NF-κB dampening response elicited by Cb as an immune evasion strategy.
Focusing on functionally relevant cis-acting lncRNA–mRNA pairs with potential immune regulatory roles, we prioritized two candidates: CYP1B1-AS1 (ENSG00000232973)-upregulated in both Cb and CbA infections; lnc-DKK2 (ENSG00000286147)-downregulated exclusively in Cb-infected macrophages (Supplementary Data 6). A comprehensive list of all pathogen-specific lncRNAs is presented in Supplementary Data 6. Scatter plots illustrating selected differential lncRNA–mRNA pair in Cb-infection is shown in Fig. 2c. Receiver operating characteristic (ROC) curve analysis revealed high sensitivity and specificity for Cb infection, with an area under the curve (AUC) of 0.80 for CYP1B1-AS1 and 0.82 for lnc-DKK2 at 95% confidence interval (CI), p < 0.01 (Fig. 2d).
Because gene proximity often implies functional and regulatory coordination26,28,29, we performed GO and KEGG enrichment analyses on all cis-acting DE lncRNA–mRNA pairs. Kyoto encyclopedia of genes and genomes (KEGG) pathway analysis identified the top 10 pathways enriched in upregulated lncRNAs, including FOXO, PI3K-AKT, Notch signaling, and ferroptosis (Fig. 2e; Supplementary Fig. 4e–j), whereas downregulated lncRNAs were associated with NET formation, DNA repair, and cell cycle regulation (Fig. 2f; Supplementary Fig. 4e–j).
CYP1B1 encodes a mitochondrial cytochrome P450 enzyme involved in ROS production and mitochondrial function30,31. CYP1B1-AS1, located in cis to CYP1B1, may regulate these processes. lnc-DKK2 is located antisense to DKK2, implicating it in WNT signaling regulation32. Boxplots of these lncRNAs across infections confirmed their Cb-specific expression profiles (Supplementary Fig. 5a, b), which were further validated by RT-qPCR at 1 h p.i. (Fig. 1d). CYP1B1-AS1 and CYP1B1 were upregulated ~22.2-fold and ~6.6-fold, respectively, while lnc-DKK2 was downregulated ~33.3-fold (Supplementary Fig. 5c, d). Interestingly, DKK2 was upregulated ~3.56-fold at the same time point, showing a divergent expression pattern from lnc-DKK2, warranting further investigation of their temporal regulation.
This comparative multi-infection analysis enabled the identification of Cb-specific, cis-acting lncRNAs: CYP1B1-AS1 and lnc-DKK2, distinct from lncRNAs regulated by other pathogens. These findings highlight their potential roles in Cb pathogenesis and establish a framework for future functional characterization.
Spatio-temporal analysis of CYP1B1-AS1 and lnc-DKK2 during Cb infection
To further investigate the regulatory roles of these DE-lncRNAs on associated mRNAs, we analyzed their temporal expression patterns. RT-qPCR analysis of CYP1B1-AS1 and CYP1B1 exhibited similar expression trends, with both being upregulated from 1 to 72 h p.i. in THP-1 macrophages (Fig. 3a) and HeLa (Fig. 3b). In contrast, lnc-DKK2 was significantly downregulated between 1 and 48 h p.i., while DKK2 was upregulated until 24 h p.i. (Fig. 3c, d). The lack of concordant expression suggests that lnc-DKK2 does not exhibit cis-regulation of DKK2 and may regulate different targets in trans, requiring further investigation.
a RT-qPCR analysis of CYP1B1-AS1 and CYP1B1 in infected THP-1 macrophages (NMII) at the indicated time points. Statistical test: two-way ANOVA; exact p-values (in the order as asterisks): CYP1B1-AS1: 0.03, 0.009, 0.003, 0.0038, and 0.0044; CYP1B1: 0.021, 0.0098, 0.009, 0.0042, and 0.0086. b RT-qPCR analysis of CYP1B1-AS1 and CYP1B1 in infected HeLa cells. Statistical test: two-way ANOVA; exact p-values (in the order as asterisks): CYP1B1-AS1: 0.0089, 0.05, 0.0035, 0.048, and 0.036; CYP1B1: 0.029, 0.0395, 0.0027, 0.0102, and 0.033. c, d RT-qPCR analysis of lnc-DKK2 and DKK2 in infected c THP-1 macrophages and d HeLa. Statistical test: two-way ANOVA; *p < 0.05; **p < 0.01; ns, not significant, p ≥ 0.05. Exact p-values are provided in the Source data file. For a–d, expression was normalized to ACTB and shown relative to mock-infected controls (set to 1). Data represent mean ± SD from three independent experiments. e, f Bulk RNA-seq–based tissue expression profiles of e CYP1B1-AS1 and f CYP1B1 from the Genotype-Tissue Expression (GTEx v.10) database, shown as log10-transcripts per million (TPM + 1). TPMs were computed from gene models with isoforms collapsed to a single gene; no additional normalization was applied. These data are derived from a publicly available RNA-seq database; biological/technical replication is not applicable. Box plots show the median (center line), 25th and 75th percentiles (box), and whiskers at 1.5× the interquartile range (IQR). Data points outside this range are plotted as individual outliers. Tissues include brain regions (e.g., amygdala, hippocampus, cerebellum) and peripheral tissues (e.g., liver, lung, kidney, spleen, blood).
We extended this temporal analysis to four model pathogens: L. pneumophila (evolutionarily related to Cb)6, B. melitensis ΔvjbR, L. innocua, and S. Typhimurium. lnc-DKK2 expression increased at or after 48 h p.i., there was no significant correlation with DKK2 expression in cells infected with these strains, with the exception of S. Typhimurium (Supplementary Fig. 6a–h), reiterating lnc-DKK2’s potential trans-regulatory role. Considering the lack of correlation, including in Cb infection, lnc-DKK2 was not pursued further in this study. Importantly, CYP1B1-AS1 expression remained unchanged during the early infection window (1–48 h p.i.), but was upregulated from 72 to 150 h p.i. in L. pneumophila and at 150 h p.i. in the non-pathogenic control L. innocua (Supplementary Fig. 6i–l). Meanwhile, CYP1B1 was upregulated after 24 h p.i. in these strains, suggesting a potential alternative regulatory mechanism in these contexts (Supplementary Fig. 6m–p). Although these findings do not confirm the specificity of these lncRNAs across all time points, they establish a consistent cis-expression pattern between CYP1B1-AS1 and CYP1B1 in Cb infection, suggesting a coordinated role in the innate immune response. Notably, lncRNAs exhibit dynamic, temporally regulated expression, reflecting their roles in fine-tuning signaling pathways. These results support the specific induction and co-expression of CYP1B1-AS1 with CYP1B1 during the early phase of Cb infection.
To validate this co-expression pattern in physiological contexts, we analyzed tissue-specific expression using the Genotype-Tissue Expression v. 10 (GTEx)33 RNA-seq datasets. Both genes showed similar co-expression trends, with the highest expression observed in whole blood, followed by the heart, liver, spleen, and lung (Fig. 3e, f). Multigene single-cell analysis from GTEx further showed maximal expression in the heart and lung, with endothelial and fibroblast cells exhibiting high expression, followed by dendritic cells (DCs) and macrophages (Supplementary Fig. 7a–c). Although immune cell expression was lower than in stromal cells, the consistent co-expression pattern across tissues supports their functional association. As expected, CYP1B1-AS1 had lower transcript counts than CYP1B1, consistent with low basal expression and context-specific regulation characteristic of lncRNAs, particularly in immune cells34. Notably, Cb colonizes the lungs during acute infection and replicates extensively in the heart during chronic stages7. The tissue-specific expression of CYP1B1-AS1 and CYP1B1 in these infection-relevant sites makes them good candidates for further functional investigations.
CYP1B1-AS1 and CYP1B1 expression in human monocyte-derived macrophages during Cb-NMI infection
Most lncRNAs exhibit poor conservation across species, and CYP1B1-AS1 exhibited no detectable conservation, suggesting it is human-specific. To assess its clinical relevance and temporal expression, we performed RT-qPCR analysis of CYP1B1-AS1 and CYP1B1 in primary human monocyte-derived macrophages (hMDMs) infected with the virulent Cb-NMI strain or stimulated with E. coli LPS (10 ng/mL) from 1 to 150 h post-exposure. Primary CD14⁺ monocytes were isolated from peripheral blood mononuclear cells (hPBMCs), with enrichment confirmed by flow cytometry (Supplementary Fig. 8a, b).
CYP1B1-AS1 and CYP1B1 were ∼2–2.5-fold upregulated from 1 to 48 h p.i. in hMDMs, consistent with the observations from Cb-NMII infections. Although expression levels plateaued from 72 h, supporting the consistent early-phase induction pattern in primary macrophages (Fig. 4a). We also evaluated PIRAT1 and LUCAT1 in hMDMs. Consistent with their known inverse correlation, PIRAT1 was upregulated until 24 h p.i., while LUCAT1 was downregulated early but increased in the later phase (Fig. 4b). This temporal inverse correlation is consistent with their documented feedback interactions19 and may reflect Cb's strategy to fine-tune immune suppression over time.
hMDMs were either infected with Cb-NMI at an MOI of 10 or stimulated with lipopolysaccharide (LPS, 10 ng/mL). RT-qPCR analysis of (a) CYP1B1-AS1 and CYP1B1 and (b) PIRAT1 and LUCAT1 in Cb-NMI-infected hMDMs at various time points (p.i.). Statistical test: two-way ANOVA; exact p-values (in the same order as asterisks): CYP1B1-AS1: 0.0075, 0.0218, 0.0089; CYP1B1: 0.0006, 0.0458, 0.0057; PIRAT1: 0.001, 0.0037, 0.0029; LUCAT1: 0.001, 0.0035, 0.014, 0.0229; and ns, not significant, p ≥ 0.05. RT-qPCR analysis of (c) CYP1B1-AS1 and CYP1B1, d PIRAT1 and LUCAT1 in LPS-treated hMDMs. Statistical test: two-way ANOVA; exact p-values (in the same order as asterisks): CYP1B1-AS1: 0.0006, 0.0008 and 0.0178; CYP1B1: 0.0039, 0.0002, 0.0003 and 0.0122; PIRAT1: 0.0007 and 0.0051; LUCAT1: 0.0022, 0.0007, and 0.05; and ns, p ≥ 0.05. Expression dynamics of core immune-responsive lncRNAs (MAILR, LINC01215, PACER, MROCKI, MIR155HG, and MIR222HG) in response to (e) Cb-NMI infection or (f) LPS stimulation. Statistical test: two-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; ns not significant, p ≥ 0.05. Exact p-values are provided in the Source data file. For all data a–f: expression normalized to ACTB and shown as fold changes relative to uninfected or unstimulated controls (mock, set to 1). Data represent mean ± SD of three technical replicates from two independent experiments. g GTEx-heatmap showing co-expression of CYP1B1-AS1, CYP1B1, and AHR across human tissues. Values are presented as log10(TPM + 1) with color intensity representing relative gene expression.
In contrast, CYP1B1-AS1 and CYP1B1 were not induced by LPS stimulation and were instead dampened at later stages (Fig. 4c). Also, PIRAT1 was downregulated and LUCAT1 robustly upregulated following 24 h LPS exposure (Fig. 4d), consistent with their reciprocal expression under agonist-driven inflammation19. These observations reinforce the idea that PIRAT1 and LUCAT1 serve as a transcriptional switch in hMDMs, playing specific roles in broader inflammatory responses.
Among the common immune-responsive lncRNAs identified in this study, MAILR, LINC01215, and PACER were not induced at 24 h and were significantly downregulated during later phases of Cb-NMI infection. In contrast, MROCKI and MIR155HG were upregulated at 24 and 48 h p.i., but declined by 72–150 h p.i. (Fig. 4e). MROCKI’s upregulation in response to inflammation reiterates its crucial role in maintaining the functional integrity of terminally differentiated macrophages17 during infection or LPS-induced responses. Importantly, all tested immune-responsive lncRNAs were significantly induced by 24 h of LPS stimulation (Fig. 4f), consistent with their established roles in inflammation.
To further investigate the regulatory context of CYP1B1-AS1, we performed a coding–noncoding co-expression (CNC) network analysis using the guilt-by-association approach35. This revealed a set of mRNAs and lncRNAs correlated with CYP1B1-AS1 and CYP1B1 (Supplementary Data 7; Supplementary Fig. 8c), including AHR. AHR is a known TF of CYP1B136, which is induced in various respiratory viral37 and enteric bacterial infections38. Lastly, tissue-specific expression profiles from GTEx datasets revealed that these genes exhibit similar spatial expression trends in infection-relevant tissues (Fig. 4g). These data support that CYP1B1-AS1/CYP1B1 are selectively induced during Cb infection, and may function as part of AHR signaling in regulating host immune responses.
CYP1B1-AS1 is transcribed from a shared bidirectional promoter and regulates CYP1B1 in cis
CYP1B1-AS1 is located on the sense strand of human chromosome 2, and has been classified as a long non-coding RNA (lncRNA). Ribosome profiling (Ribo-seq) analysis using globally aggregated datasets from the GWIPS-viz database39 revealed weak ribosome occupancy, with a single open reading frame (ORF), c2norep36, detected and minimal ribosome-protected fragments, suggesting inefficient or non-functional translation (Supplementary Fig. 8d). Recent evidence suggests lncRNAs can contain short untranslated regions (UTRs), non-canonical initiation codons that can promote translation. To investigate this, 62 short ORFs (sORFs) were predicted within the CYP1B1-AS1 sequence (alternative initiation codon as criteria) using NCBI ORF Finder. However, BLAST analysis revealed no significant conservation to known protein sequences. To further assess CYP1B1-AS1 and its predicted sORFs for coding potential, we performed computational analysis using the coding potential assessment tool (CPAT)40 (Fig. 5a) and coding potential calculator (CPC) algorithm41 (Supplementary Data 8). The results confirmed that CYP1B1-AS1 is non-coding, except for ORF17 and ORF62, which exhibited coding capacity. To experimentally validate these findings, CYP1B1-AS1, ORF17, and ORF62 were cloned into a CMV promoter-driven 3X-FLAG-tagged pCDNA vector (pCDNA-FLAG, Supplementary Data 1), transfected into HEK293T cells, and analyzed via immunoblotting for FLAG expression. No detectable protein signal was observed for the lncRNA or its ORFs, confirming its non-coding nature (Fig. 5b, Supplementary Fig. 9a). Since lncRNA function is often linked to subcellular localization26,42, we assessed CYP1B1-AS1 using polyadenylation analysis and cellular fractionation assays. The results confirmed that CYP1B1-AS1 is a polyadenylated lncRNA (Supplementary Fig. 9b) and is equally distributed between the nuclear and cytoplasmic fractions (~1.2-fold nuclear enrichment relative to the cytoplasm, p ≥ 0.05) (Supplementary Fig. 9c).
a Coding Potential Assessment Tool (CPAT) analysis predicting low coding potential for CYP1B1-AS1. Known lncRNAs (TUG1, NEAT1, ANCR, MALAT1), and mRNAs (CYP1B1, GAPDH) serve as controls. b Immunoblot analysis of HEK293T cells transfected with pCDNA-CYP1B1-AS1. Controls: untransfected, empty vector (pCDNA-FLAG) and FLAG-tagged ANCR, CYP1B1, or GAPDH. c Dual-luciferase assay demonstrating bidirectional promoter activity (arbitrary units) of the CYP1B1-AS1 promoter (PCYP1B1-AS1), driving Firefly (F-Luc/FL) and Renilla luciferase (R-Luc/RL). pF-Luc and pR-Luc are positive controls for ACTB promoter (PACTB), and pGL4 is an empty vector. Statistical test: one-way ANOVA. Exact p-values: pF-Luc: 0.0001; pCYP1B1FL: 0.0029; pR-Luc: 0.0004; pCYP1B1RL: 0.0037. RT-qPCR showing reduced CYP1B1 expression after CYP1B1-AS1 knockdown in (d) HeLa and (e) THP-1 macrophages. RT-qPCR showing CYP1B1-AS1 expression following CYP1B1 knockdown in (f) HeLa and (g) THP-1 macrophages. Expression normalized to HPRT. KD1/KD2 represent different siRNA-mediated knockdowns or their combination (lncCYPB for CYP1B1-AS1, CYPB for CYP1B1). Statistical test d–g: one-way ANOVA; exact p-values: d ***p = 0.0002 (lncCYPB-KD1); d ***p = 0.0003 (lncCYPB-KD2); d, e ****p < 0.0001; f, g ns not significant, p ≥ 0.05. h Immunoblot and (i) densitometry analysis of CYP1B1 after CYP1B1-AS1 or CYP1B1 knockdown in HeLa cells. j Immunoblot, and (k) densitometry analysis of CYP1B1 after CYP1B1-AS1 or CYP1B1 knockdown in THP-1 macrophages. Statistical test i, k: one-way ANOVA; ns, p ≥ 0.05; ****p < 0.0001. β-Actin: the loading control. l, m mRNA decay assay measuring half-lives (t1/2) of (l) CYP1B1 and (m) HPRT in NC and lncCYPB cells after actinomycin D treatment. Decay calculated from a one-phase regression plot from 2(−ΔCT) values. All blots and data represent mean ± SD from three independent experiments (n = 3). Source data are provided as a Source Data file.
Previous studies have shown that lncRNAs, particularly those transcribed from bidirectional promoters, often regulate nearby protein-coding genes in cis26,43. Our results show CYP1B1-AS1 is antisense to CYP1B1, exhibits a similar expression pattern, and positively correlates with AHR. This prompted investigations into potential shared regulatory elements between the genes. To identify TF binding sites (TFBS) within the CYP1B1-AS1 promoter region, we used UCSC Genome Browser and Expasy Eukaryotic Promoter Database. The region upstream of the transcription start site (TSS) contained AHR motifs, as predicted by Expasy and JASPAR (Supplementary Fig. 9d–f). To confirm the bidirectional nature of the promoter, a dual-luciferase reporter assay was performed by cloning the promoter into pGL4-Luc-Rluc (Supplementary Fig. 9g). The assay demonstrated that the promoter activated both Firefly luciferase (CYP1B1-AS1 direction, pCYP1B1FL) and Renilla luciferase (CYP1B1 direction, pCYP1B1RL) by at least 2.5-fold compared to the empty vector (pGL4), confirming it as a bidirectional promoter (Fig. 5c).
Additionally, we analyzed the CYP1B1-AS1 and CYP1B1 regulatory regions upstream of the TSS using publicly available UCSC-ChIP-seq datasets. The UCSC Genome Browser track view shows enrichment of histone modifications and DNase I hypersensitivity (HS) sites at the regulatory region, indicative of an active transcriptional landscape (Supplementary Fig. 10a). Given that CpG islands are frequently found in bidirectional promoters26,44, we further analyzed the promoter region using EMBOSS CpGplot. The analysis identified a high observed-to-expected CpG ratio and a GC content above 50% (Supplementary Fig. 10b), further supporting the bidirectional transcriptional activity of this promoter.
Further, we performed knockdown using siRNAs (Supplementary Data 1) targeting CYP1B1-AS1 (lncCYPB-KD1, lncCYPB-KD2, lncCYPB) and CYP1B1 (CYPB-KD1, CYPB-KD2, CYPB). GAPDH siRNA (PC) and a negative control siRNA (NC) were used as controls. Knockdown efficiency was highest with lncCYPB (~90% in HeLa, ~67% in THP-1 macrophages) for CYP1B1-AS1 and CYPB (~90% in HeLa, ~70% in THP-1 macrophages) for CYP1B1 (Supplementary Fig. 11a–h). Knockdown of CYP1B1-AS1 significantly reduced CYP1B1 mRNA levels (~4.57-fold in HeLa, Fig. 5d; ~4.95-fold in THP-1 macrophages, Fig. 5e). In contrast, CYP1B1 knockdown had no effect on CYP1B1-AS1 expression in either HeLa (Fig. 5f) or THP-1 macrophages (Fig. 5g), indicating that CYP1B1-AS1 regulates CYP1B1 in cis, but not vice versa. Immunoblot and densitometry analysis revealed a ≥70% reduction in CYP1B1 protein levels following CYP1B1-AS1 and CYP1B1 knockdown, with maximum knockdown (~90%) observed in lncCYPB and CYPB conditions (Fig. 5h-k), confirming that this regulatory effect extends to the protein level.
We next performed an mRNA decay assay to investigate post-transcriptional regulation of CYP1B1 by CYP1B1-AS1. Cells were treated with actinomycin D to inhibit global transcription in lncCYPB and NC conditions. The knockdown of CYP1B1-AS1 did not affect the half-life of CYP1B1 or HPRT mRNA (control) over a 5 h treatment (Fig. 5l, m). These results demonstrate that CYP1B1-AS1 and CYP1B1 are co-regulated in cis via a bidirectional promoter, and that CYP1B1-AS1 does not affect CYP1B1 mRNA stability post-transcriptionally.
CYP1B1-AS1 and CYP1B1 knockdown reduce Cb replication
CYP1B1-AS1-dependent upregulation of CYP1B1 prompted us to investigate the functional relevance of these genes in Cb pathogenesis. Growth curve analyses under knockdown conditions revealed a significant reduction (~2-fold) in bacterial load in knockdown-infected cells (lncCYPB:NMII, CYPB:NMII, and dCYPB:NMII) compared to negative control (NC:NMII) and untreated-infected cells (NMII) (Fig. 6a, Supplementary Data 1). Similar results were observed in HEK293T knockdown, showing a 2-fold reduction in Cb growth by 7-day p.i. (Fig. 6b). Immunofluorescence microscopy further confirmed that both genes are necessary for CCV expansion, as LAMP-1 and Cb-stained CCVs were significantly smaller in knockdown cells, corroborating the growth curve data (Fig. 6c, d).
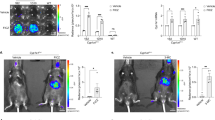
a, b Intracellular replication of C.burnetii (NMII) assessed by genomic equivalent (GE) quantification over a 7-day time course following knockdown of CYP1B1-AS1 (lncCYPB), CYP1B1 (CYPB), or both (dCYPB) and negative control siRNA-treated (NC) in (a) THP-1 macrophages (MOI = 50) and (b) HEK293T cells (MOI = 100). Additional controls include CbA-infected (ΔdotA) and uninfected cells (mock). Data represent mean ± SD from three independent experiments (n = 3). Statistical test (a-b): two-way ANOVA; exact p-values; p = 0.007 (ΔdotA); p = 0.008 (lncCYPB: NMII); p = 0.0073 (CYPB: NMII); p = 0.0091 (dCYPB:NMII); ****p < 0.0001; ns not significant, p ≥ 0.05. c Representative immunofluorescence images of THP-1 macrophages infected with C. burnetii (NMII or ΔdotA) at 5 days p.i. Nuclei (Hoechst 33342, blue), LAMP-1 (lysosomal marker, green), C.burnetii (magenta), Scale bar: 10 μm. d Quantification of Coxiella-containing vacuoles (CCVs) area using ImageJ, QuPath, and Labkit, with each LAMP-1+ bacterial compartment representing an individual CCV. Data represent median at 95% CI of 100 CCVs per repeat (n = 3); one-way ANOVA; ****p < 0.0001; ns, p ≥ 0.05. e–j ELISA quantification of cytokines in supernatants from THP-1 macrophages under indicated knockdown and infection conditions: e TNF-α, f IL-6, g IL-8, and h IL-1β, i IL-10, and j IFN-γ. For e–j, data represent mean ± SD for n = 3. Statistical test: one-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; ns, p ≥ 0.05. Exact p-values and source data are provided as a Source Data file.
To further assess their impact on Cb replication, we ectopically overexpressed CYP1B1-AS1 and CYP1B1 using pcDNA3.1/Zeo(+) constructs (pCYP1B1-AS1 and pCYP1B1, Supplementary Data 1). While no detectable change in bacterial load was observed in overexpressed-HeLa cells (Supplementary Fig. 11i), pCYP1B1 expression in HEK293T cells resulted in increased Cb replication at 7 days p.i. (Supplementary Fig. 11j), suggesting a cell-type-dependent effect. Notably, CYP1B1-AS1 overexpression neither affected Cb survival nor influenced CYP1B1 expression levels in any tested cell type (Supplementary Fig. 11k, l), suggesting that CYP1B1-AS1 does not regulate CYP1B1 in trans.
CYP1B1-AS1 and CYP1B1 suppress inflammatory cytokine responses
Cb subverts host immune signaling to limit inflammation and cytokine production8,45,46,47. We next investigated whether CYP1B1-AS1 and CYP1B1 modulate host inflammatory responses. Knockdown of either gene resulted in a significant increase in pro-inflammatory cytokines at both transcript (RT-qPCR) and protein (ELISA) levels (Fig. 6e–j, Supplementary Fig. 12a–h). Specifically, TNF-α, IL-6, and IL-8 levels were significantly elevated in knockdown-infected conditions (lncCYPB:NMII, CYPB:NMII, and dCYPB:NMII) even in mock-infected counterparts (Supplementary Figs. 12d–f and 6e–g). These results suggest that both genes suppress inflammation under basal conditions, independent of infection status. IL-1β levels exhibited variability (Fig. 6h), indicating an infection-dependent modulation of responses in knockdown conditions. In contrast, IL-10 (Fig. 6i) and IFN-γ (Fig. 6j) levels remained unchanged in knockdown conditions but exhibited infection-dependent modulation.
These findings indicate that CYP1B1-AS1 and CYP1B1 suppress inflammation under normal conditions, and their knockdown skews the cytokine profile toward a pro-inflammatory state. Knockdown-driven inflammation likely creates a less favorable niche for Cb survival, linking increased host inflammation to reduced bacterial survival.
AHR regulates CYP1B1-AS1 and CYP1B1 expression in response to Cb infection
AHR is a key regulator of immune homeostasis, primarily activated by tryptophan metabolites38, or oxidative stress, leading to transcriptional activation of cytochrome P450 genes36,48. While AHR’s role in bacterial, viral, and parasitic infections has been explored, its regulation in Cb pathogenesis remains uncharacterized.
To evaluate its role, Cb-infected macrophages were treated with 6-Formylindolo [3,2-b] carbazole (FICZ, AHR agonist), at concentrations of 200 or 400 nM. RT-qPCR analysis revealed a 2-fold increase in CYP1B1-AS1 expression in FICZ-treated-infected cells (Fig. 7a). Concurrently, RT-qPCR (Supplementary Fig. 13a), immunoblotting (Fig. 7b), and densitometry analysis (Fig. 7c) showed at least 5-fold upregulation in CYP1B1 transcripts and protein upon infection. While no significant difference in CYP1B1 protein levels was observed between the FICZ-treated control and the FICZ-treated, NMII-infected group at 24 h p.i., a ~2-fold increase in expression was detected in the infected group at 48 and 72 h p.i. at 48 and 72 h p.i. (Fig. 7b, c).
a RT-qPCR analysis of CYP1B1-AS1 in C. burnetii-infected (NMII) THP-1 cells, with or without AHR agonist FICZ (200 or 400 nM). Expression normalized to HPRT and compared to mock. Statistical test: two-way ANOVA; ****, p < 0.0001. Representative immunoblot (b) and densitometry (c) of CYP1B1 protein following NMII infection and/or FICZ treatment. β-Actin: loading control. Statistical test: two-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001. d AHR nuclear translocation assessed by fractionation and immunoblotting after infection and FICZ (400 nM) treatment. Lamin B1 (nuclear) and α-Tubulin (cytoplasmic) were used as controls. Samples were from the same experiment, processed in parallel (n = 3). Nuc-nuclear fraction; Cyto-cytoplasmic fraction; WCL-whole cell lysate. e Densitometry quantification of AHR in Nuc vs. WCL, normalized to Lamin B1. One-way ANOVA; **p = 0.002; ****p < 0.0001. f Densitometry quantification of AHR in Nuc vs. Cyto (normalized to α-Tubulin). One-way ANOVA; ***p = 0.0003; ****p < 0.0001. g AHR-luciferase reporter assay in HepG2-AHR-Lucia cells infected with NMII and treated with FICZ (400 nM) or AHR antagonist CH-223191 (10 μM). Luminescence was measured over 150 h p.i., and normalized to mock. NC = endotoxin-free water; CH223 = antagonist control. Two-way ANOVA; exact p-values: 0.0005 (FICZ-1h); 0.0003 (NMII-1h); 0.009 (NMII-FICZ-1h); <0.0001. RT-qPCR analysis of CYP1B1-AS1 and CYP1B1 after AHR knockdown (AHR-KD) in (h) HeLa and i THP-1. Expression normalized to HPRT and compared to negative control siRNA (NC)-treated cells. h, i Two-way ANOVA; exact p-values: ***p = 0.0002; ****p < 0.0001. Immunoblot and densitometry analysis of CYP1B1 protein following AHR knockdown in j, k HeLa and l, m THP-1. β-Actin: loading control. k, m: one-way ANOVA; exact p-values: *p = 0.0109 (k); *p = 0.0159 (m); ns, p ≥ 0.05. All data represent mean ± SD; n = 3. Source data are provided as a Source Data file.
To investigate AHR activation dynamics, subcellular fractionation was performed in Cb-infected macrophages, followed by immunoblot analysis of whole-cell lysates (WCL), cytoplasmic, and nuclear fractions at 24 h p.i. (Fig. 7d–f). A significant decrease in cytoplasmic AHR upon FICZ treatment and infection indicated degradation of inactive cytoplasmic AHR. In contrast, nuclear AHR levels remained relatively stable, supporting the mechanism by which AHR translocates to the nucleus upon activation and is retained there, while cytoplasmic AHR is selectively degraded49.
A HepG2-AHR-Lucia luciferase reporter assay was employed to quantify AHR activation. Immunofluorescence microscopy confirmed the presence of bacteria within Lamp-1+ CCVs at 3 days p.i. (Supplementary Fig. 13b), validating HepG2 as a suitable model for downstream analyses. HepG2-AHR-Lucia luciferase reporter cells were treated with FICZ (400 nM) or CH-223191 (10 µM, AHR antagonist) before Cb infection. FICZ significantly increased luciferase activity (p < 0.0001), confirming AHR activation, while CH-223191 treatment markedly reduced activity (p < 0.0001), demonstrating assay specificity (Fig. 7g). Importantly, Cb infection (NMII) alone significantly induced AHR compared to the mock (p < 0.001). Although CH-223191-treated infected cells (CH223: NMII) exhibited reduced AHR activation, their luciferase levels remained higher than uninfected controls (p < 0.0001).
Time-course RT-qPCR in HepG2 reporter cells revealed a ≥50-fold upregulation of CYP1B1 and CYP1B1-AS1 following infection (Supplementary Fig. 13c). To evaluate whether AHR activation required functional T4SS, luciferase induction was compared between NMII and ΔdotA mutants. No significant difference in activation was observed between these strains, nor upon FICZ or CH-223191 treatment (Supplementary Fig. 13d), demonstrating AHR activation is T4SS-independent.
To determine whether AHR is essential for CYP1B1-AS1 and CYP1B1 induction, we silenced AHR (AHR-KD) in HeLa and THP-1 macrophages, achieving ~90% reduction in expression (Supplementary Fig. 14a–f). RT-qPCR showed ≥10-fold reduction in CYP1B1-AS1 and CYP1B1 expression in AHR-KD cells compared to NC-treated controls (Fig. 7h, i). Immunoblot analysis further confirmed a ~ 10-fold decrease in CYP1B1 protein levels following AHR-KD (Fig. 7j–m). Neither CYP1B1-AS1 nor CYP1B1 expression was rescued in AHR-KD cells following infection (Supplementary Fig. 14g, h), confirming that AHR is necessary for their transcriptional induction. Furthermore, AHR-KD macrophages exhibited ~3-fold reduced bacterial load compared to NMII-infected controls (Supplementary Fig. 14i).
Collectively, these findings demonstrate that canonical AHR signaling is activated as a host response to Cb infection, leading to AHR nuclear translocation and the transcriptional activation of downstream targets CYP1B1-AS1 and CYP1B1 from a shared promoter.
CYP1B1-AS1 and CYP1B1 regulate mitochondrial function and ROS
CYP1B1 has been previously linked to increased ROS generation and disruption of mitochondrial homeostasis31. We hypothesized that the increased inflammation from CYP1B1-AS1 and CYP1B1 silencing in THP-1 macrophages is due to disrupted mitochondrial functions and ROS production. To assess this, we measured ROS accumulation by DCFDA (total ROS) and MitoSOX (mitochondrial ROS) using flow cytometry. While Cb infection modestly increased ROS levels, as expected for macrophage responses to pathogens, both total ROS (Fig. 8a, b) and mitochondrial ROS (Fig. 8c, d) were significantly elevated in resting lncCYPB, CYPB, and dCYPB and infected cells at 24 h.
Flow cytometry and metabolic assays were performed on THP-1 macrophages with siRNAs targeting CYP1B1 (CYPB), CYP1B1-AS1 (lncCYPB), or both (dCYPB), and C. burnetii-infected (NMII) conditions at 24 h p.i. a, b Total ROS production assessed by flow cytometry, measured as mean fluorescence intensity (MFI) of DCFDA-stained cells compared to mock. Statistical test: one-way ANOVA; ****p < 0.0001; **p = 0.006 (NMII); **p = 0.0021 (NC: NMII). c, d Mitochondrial ROS production assessed by flow cytometry, measured as MFI of MitoSOX™ Red-stained cells. e, f Mitochondrial membrane potential assessed by flow cytometry, measured as MFI of MitoProbe™ TMRM-stained cells. For d, f; Statistical test: one-way ANOVA; ****p < 0.0001. g Oxygen consumption rate (OCR) measured using an Agilent Seahorse Metabolic Analyzer to evaluate mitochondrial respiration. h Maximal respiration was measured across knockdown and controls. Statistical test: two-way ANOVA; ****p < 0.0001. All data represent mean ± SD from three independent experiments (n = 3). Source data are provided as a Source Data file.
Since elevated mitochondrial ROS can disrupt mitochondrial function, leading to altered membrane potential50, we measured mitochondrial membrane potential using tetramethylrhodamine methyl ester (TMRM). TMRM intensity was ~3-fold reduced in knockdown cells in both resting and infection states (Fig. 8e, f), suggesting mitochondrial membrane depolarization. To confirm that ROS generation originated from dysfunctional mitochondria rather than mitochondrial loss, we performed Mitotracker Green staining, which showed no significant changes in mitochondrial mass (Supplementary Fig. 15a, b).
To evaluate whether metabolic reprogramming could contribute to mitochondrial ROS and dysfunctions in CYP1B1-AS1 and CYP1B1-silenced cells, we assayed their oxygen consumption rate (OCR) and extracellular acidification rate (ECAR). Basal, spare capacity and maximal respiration were significantly elevated in silenced cells, in both mock and infected conditions (Fig. 8g, h, Supplementary Fig. 15c, d). The results suggest that silenced-macrophages increased glycolytic capacity (ECAR) during infection compared to the uninfected cells (Supplementary Fig. 15e). However, these cells fail to undergo Warburg shift51, as they continue to rely on oxidative phosphorylation (OXPHOS) (Fig. 8g, Supplementary Fig. 15c). This inability of silenced-cell mitochondria to metabolically reprogram from OXPHOS to glycolysis imposes additional stress on the electron transport chain, thereby contributing to elevated ROS levels.
In summary, CYP1B1-AS1 and CYP1B1 are essential for maintaining mitochondrial functions and ROS balance. Their silencing results in increased ROS and inflammation, impairing the macrophage’s ability to undergo metabolic reprogramming during infection. These data suggest that upregulation of CYP1B1-AS1 and CYP1B1 during Cb infection represents a host response to counteract mitochondrial ROS and inflammation, favoring intracellular bacterial survival.
CYP1B1-AS1 and CYP1B1 regulate mitochondrial ROS and inhibit apoptosis
Mitochondrial ROS and inflammation are well-established triggers for cell death50,52. To investigate the lncRNA and gene’s role in modulating cell death, we measured Annexin-V and Propidium Iodide (PI) accumulation using flow-cytometry (Supplementary Fig. 16a–d). lncCYPB, CYPB, and dCYPB knockdown exhibited increased Annexin-V accumulation (early-apoptosis). Notably, upon infection, silenced-cells displayed significantly higher proportions of Annexin-V+ and Annexin-V+PI+ (late apoptosis) populations at 24 h compared to the controls (Fig. 9a–c). By 48 h p.i., these cells progressed significantly towards late apoptosis (Fig. 9d, e), with lncCYPB:NMII, CYPB:NMII, and dCYPB:NMII showing markedly increased cell death (Fig. 9f). These findings suggest that CYP1B1-AS1 and CYP1B1 are critical for maintaining mitochondrial function and ROS balance, and that their silencing sensitizes host cells to apoptosis, which is further exacerbated by Cb infection.
a–f Flow cytometry analysis of apoptosis during CYP1B1-AS1 (lncCYPB), CYP1B1 (CYPB), dual knockdown (dCYPB), negative control (NC), and C. burnetii-infected (NMII) conditions at 24 h and 48 h p.i. Cells were stained with propidium Iodide (PI) and FITC-Annexin-V, at 24 h and 48 h p.i. to quantify populations of early apoptotic (Annexin-V+ PI−; Q1), late apoptotic (Annexin-V+ PI+; Q2), dead (PI+; Q3), and non-apoptotic (Annexin-V− PI−; Q4) cells. Percentage of (a) early apoptotic, b late apoptotic, c dead cells at 24 h p.i. Percentage of (d) early apoptotic, e late apoptotic, f dead cells at 48 h p.i. For a–f, data represent mean ± SD from three independent experiments (n = 3). Statistical test: one-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; ns not significant, p ≥ 0.05. Exact p-values are provided in the Source Data file. g Proposed model of CYP1B1-AS1-mediated regulation of CYP1B1 during C. burnetii (Cb) infection. During Cb infection, aryl hydrocarbon receptor (AHR) is activated and translocates into the nucleus, where it transcriptionally induces both CYP1B1-AS1 and CYP1B1 from a shared bidirectional promoter. The activation of CYP1B1-AS1 influences CYP1B1 expression through a cis-regulatory mechanism within the local genome, thereby modulating CYP1B1 transcript levels. This enhances turnover of CYP1B1, a mitochondria-enriched cytochrome P450 enzyme involved in maintaining redox homeostasis. CYP1B1 modulates reactive oxygen species (ROS), suppresses ROS-driven inflammation, and inhibits apoptosis, thereby facilitating an intracellular environment favorable for Cb replication. Schematic was created using BioRender.com.
To further elucidate the mechanism linking CYP1B1-AS1 and CYP1B1 upregulation during Cb infection to ROS regulation and apoptosis, we overexpressed 3XFLAG-CYP1B1 (pcDNA-CYP1B1, Supplementary Data 1) in HEK293T cells and infected them with Cb. Cells transfected with 3XFLAG-GFP (pcDNA-eGFP) and only pcDNA-CYP1B1 overexpressed cells served as immunoprecipitation (IP) controls. This system was selected for two reasons: (a) CYP1B1-AS1 overexpression did not induce CYP1B1 expression in trans and (b) to identify proteins specifically enriched upon CYP1B1 upregulation during infection.
Mass spectrometry (MS) analysis identified CYP1B1-associated protein profiles in control versus infected samples (Supplementary Data 9). Functional enrichment revealed that CYP1B1-associated proteins were predominantly involved in ROS regulation and negative regulation of apoptosis (Supplementary Fig. 16e, Supplementary Data 10 and 11). Additionally, CYP1B1 overexpression was linked to several key immune and stress response pathways, including TNF signaling, p38/MAPK, mTOR, AMPK, and IL-17 (Supplementary Fig. 17a–d), suggesting broader roles in host-pathogen interaction.
In summary, our results demonstrate that Cb infection induces AHR, which transcriptionally upregulates CYP1B1-AS1 and CYP1B1 in cis. Regulation of ROS and cell death represents a key immune evasion strategy. Our findings suggest the AHR/CYP1B1-AS1/CYP1B1 axis reprograms ROS and inflammatory responses of the cell during Cb infection (Fig. 9g), facilitating an intracellular niche conducive to bacterial survival.
Discussion
The functional characterization of lncRNAs remains a major challenge, hindered by factors such as complex genomic context, variable subcellular localization, overlap with protein-coding genes, and stimulus-dependent expression patterns. In this study, immune-responsive lncRNAs commonly induced by PAMP exposure were identified based on defined selection criteria, aiming to capture DE transcripts with potential regulatory roles in macrophages during infection.
We identified a well-characterized subset of lncRNAs with known functions in human macrophage responses to infection and LPS stimulation (Table 1, Supplementary Data 3). Among these, MAILR, MROCKI, PACER, and LINC01215 represent established LPS-responsive lncRNAs and were commonly induced across several infections, consistent with recent literature17,23. Additional immune-regulatory lncRNAs, including MIR222HG, MIR155HG, LINC01465, LUCAT1, and EGOT, were induced in at least four infection models. Schmerer et al. 17 recently developed SMyLR (rna-lab.org/smylr), an atlas of lncRNA regulation and interactomes in activated primary macrophages, revealing that these lncRNAs modulate ~16% of LPS-responsive mRNAs, notably within TLR and IFN-α pathways17.
Several immune-regulatory lncRNAs identified in our study align with known roles in pathogen sensing and macrophage activation. EGOT, an established marker in HPV infection and a prognostic indicator in HNSCC53, is inducible by TNF-α, LPS was induced by various bacterial strains in this study, including EHEC, B. subtilis, and L. innocua, highlighting its PAMP-responsive expression in macrophages. EGOT is regulated by RIG-I and PKR during HCV infection and modulates inflammation via the PI3K/AKT, MAPK, and NF-κB pathways24, although it suppresses ISG expression through an undefined mechanism54. Similarly, MIR155HG is highly expressed in pro-inflammatory macrophages, promotes M1 polarization55, enhances IFN-β production via the JAK-STAT pathway, and supports antiviral responses56,57 through miR-155-5p. MIR222HG, another lncRNA with antiviral functions, facilitates M1 polarization by sponging miR-146a-5p to activate TRAF6 for activating the IKK/IκB/p65 signaling axis58. It produces miR-22-3p, which modulates HDAC4-MEF2C signaling to influence myoblast differentiation59.
Interestingly, LUCAT1, which is known to exhibit a Yin–Yang regulatory relationship with PIRAT1 in human myeloid cells during infection and LPS response17,19,22, was consistently upregulated across several infections. Although PIRAT1 did not conform to the established pattern at 1 h time point in our RNA-seq dataset, our time-course analysis revealed that PIRAT1 was specifically induced in Cb-NMI-infected macrophages, while LUCAT1 was downregulated, supporting their temporally staggered expression. These findings align with prior studies showing that these lncRNAs typically display inverse expression dynamics ≥4 h post-LPS stimulation or during advanced stages of severe COVID-19. Together, this reinforces the notion that lncRNA-mediated immune regulation is temporally stratified, enabling these transcripts to orchestrate distinct layers of innate immune response over time.
Our multi-pathogen RNA-seq approach aimed to distinguish lncRNAs uniquely associated with pathogenesis from those broadly activated as part of the macrophage immune response. This strategy identified a set of Cb-specific DE lncRNAs, among which lnc-DKK2 and CYP1B1-AS1 were prioritized for further investigation. CYP1B1-AS1 was selected for functional validation based on its syntenic relationship with CYP1B1 and its coordinated spatiotemporal expression across multiple cell lines, hMDMs, and GTEx data.
Our RNA-seq analysis identifies a previously uncharacterized regulatory axis involving CYP1B1-AS1 and CYP1B1 in the context of Cb infection. This axis, integrated within AHR signaling, modulates ROS-induced inflammation and infection-exacerbated apoptosis. Notably, CYP1B1-AS1 in cis, as its knockdown resulted in a concordant decrease in both CYP1B1 transcript and protein levels, indicating a direct regulatory relationship. Like other immune-associated lncRNAs, CYP1B1-AS1 may also function via multiple mechanisms15. For instance, lincRNA-Cox2 regulates PTGS2 in cis via an enhancer-like function, while also modulating distal immune gene networks in trans27. Our cytokine profiling and mass spectrometry data align with a similar model, suggesting that CYP1B1-AS1 may also participate in divergent regulatory pathways.
One of the key finding of this study is the activation of the AHR signaling in response to Cb infection. AHR is a ligand-activated TF that drives macrophage polarization60 and suppresses inflammatory cytokines such as TNF-α, IL-6, IL-12 and IL-1736,38,61,62. A limitation of our study is that specific ligand responsible for AHR activation during Cb infection remains to be identified.
Since no significant differences were observed in AHR activation between the wild-type and DotA-deficient strain, we propose that AHR activation is a T4SS-independent host response, potentially triggered by endogenous tryptophan metabolites, LPS, or non-T4SS bacterial effectors. AHR is well documented to be activated by gut- and lung microbiota-derived metabolites38,63,64, LPS62, which can inhibit IL-17 signaling65, and promote an immunosuppressive environment that may support Cb replication45. Elucidating the specific ligand(s) and possible role of T4SS-independent effectors in this context will be a focus of future studies.
The present study positions AHR as a central regulatory node, linking transcriptional activation of CYP1B1-AS1 and CYP1B1, contributing to immune suppression and redox balance in infected macrophages45. Functionally, silencing of CYP1B1-AS1 or CYP1B1 alone induced a hyperinflammatory response, marked by elevated levels of TNF-α, IL-6, and IL-1β, cytokines known to be actively dampened by Cb to avoid immune clearance8,66,67,68. TNF-α, in particular, has been shown to limit Cb infection both locally and systemically69,70. The observed increase in these cytokines corresponded with the significant reduction in intracellular bacterial load in knockdown cells.
Furthermore, these cytokines are known to synergize with ROS in triggering apoptosis during infection71,72,73. Our data demonstrate that silencing CYP1B1-AS1 or CYP1B1 leads to elevated ROS levels and mitochondrial dysfunction, as evidenced by Seahorse profiling. Knockdown cells showed increased OXPHOS activity and an inability to undergo a Warburg shift, suggesting mitochondrial stress and energy imbalance. Considering mitochondria are a primary source of ROS, this dysfunction likely predisposes cells to apoptosis. MS data further revealed enrichment of pathways associated with CYP1B1-mediated intrinsic apoptosis, strengthening the mechanistic link, ROS regulation, and infection-exacerbated cell death. Taken together, our findings support a model in which loss of the CYP1B1-AS1/CYP1B1 axis leads to increased inflammation and mitochondrial damage, ultimately sensitizing cells to apoptosis and limiting Cb’s intracellular survival.
Overall, this study expands the emerging paradigm of lncRNA-mediated immune regulation in infection by showing how the CYP1B1-AS1/CYP1B1 axis creates a controlled redox environment that benefits Cb replication. Through AHR-dependent transcriptional regulation, this axis fine-tunes host immunity, limits damaging inflammation, and preserves mitochondrial integrity50, all of which are essential for successful intracellular replication.
Importantly, our findings introduce mitochondrial immunity and AHR signaling as critical yet underexplored components of the Coxiella-host interaction. These insights enable future studies investigating the ligands driving AHR activation and the broader role of lncRNA- and mitochondria-based immune regulation in Coxiella pathogenesis. The discovery of an AHR-driven CYP1B1-AS1/CYP1B1 axis specifically activated during Cb infection, and its role in supporting bacterial survival, highlights potential RNA- and pathway-based therapeutic targets for Q fever.
Methods
Ethics statement
All experiments were conducted in accordance with relevant ethical regulations and guidelines at Texas A&M University Health Science Center, USA. Human PBMCs (Lonza, #CC-2703) were commercially obtained from Lonza, a certified provider of de-identified human cells. PBMCs were derived from one 24-year-old male and one 47-year-old female healthy donor, as specified by the lot information provided by the vendor. The use of commercially purchased, fully de-identified PBMCs does not constitute human subjects research and is therefore exempt from institutional review board (IRB) approval under federal regulations (45 CFR 46.102(l)) and the Texas A&M University Human Research Protection Program SOP (HRP-093). No additional ethical approval was required for their use in this study. Sex was not considered a variable in the study design or analysis, and no sex-based or other demographic subgroup analyses (e.g., race or ethnicity) were performed.
C. burnetii strains and culture conditions
All Cb mutant strains used in the study were derived from Nine Mile phase II (NMII), strain RSA439 (Cb), clone 4 (Supplementary Data 1). Strains were cultured for 7 days in ACCM-2 (Sunrise Science Products) at pH 4.75, 37 °C in 2.5% O2 and 5% CO2 in a tri-gas incubator (New Brunswick™, Eppendorf), and stored as described previously74. For Cb Nine Mile phase I (NMI) clone 7 (RSA493) culture, the strain was grown in embryonated yolk sacs, and purified using density-gradient centrifugation as described previously7. Experiments with NMI were performed in a biosafety level 3 (BSL3) facility at Texas A&M University Health Science Center, USA.
Bacterial strains and culture conditions
The bacterial strains used in this study included Ec5, EcT, EcN, Bs, Fn, Pa, Sa, STm, Rr, Ml, Li, Ef and Bmv (Supplementary Data 1). Sa, Bmv, Pa and Fn were grown for 20 h (overnight) in Tryptic soy broth (TSB; Millipore-Sigma) medium at 37 °C and 220 rpm in a shaker-incubator (MaxQ™ Thermo Scientific). Ef and Li were cultured under identical conditions in Brain Heart Infusion (BHI; Millipore-Sigma) broth. EcT, EcN, STm, Ec5, Bs, Li, Rr and Ml were cultured in Luria-Bertini (LB; MP Biomedicals) medium at 37 °C, 220 rpm. Lp was cultured in buffered charcoal yeast extract (BCYE; Millipore-Sigma) broth with legionella growth supplements at 37 °C and 220 rpm overnight. All bacterial strains were sub-cultured to early logarithmic phase (OD₆₀₀ ∼ 0.6) prior to use in mammalian cell infection assays.
Mammalian cell culture
THP-1 monocytes (ATCC, #TIB-202) were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium (Cytiva HyClone™), with 10% fetal bovine serum (FBS; Avantor) to confluence in 5% CO2 incubator (Nuaire). The cells were then seeded in an appropriate cell culture dish and treated with 100 ng/mL PMA (InvivoGen, #tlrl-pma) to differentiate into macrophages75. After 24 h of PMA treatment, the medium was removed, and cells were allowed a 24 h resting phase before experiments. HEK293T (ATCC, #CRL-3216) and HeLa (ATCC, # CCL-2) cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Sigma-Aldrich) with 10% FBS to confluence in a 5% CO2 incubator for subsequent experiments. These cell lines were not re-authenticated in our laboratory.
Mammalian cell infections for RNA-seq and transcript analysis
For RNA-seq, Cb inoculum was prepared from frozen stocks to infect THP-1 macrophages at MOI of 50 for 1 h. Infected cells were centrifuged at 500 × g for 10 min to synchronize bacterial uptake74. Cells were collected 1 h p.i., snap-frozen in liquid nitrogen, and processed for RNA isolation and sequencing.
For spatio-temporal gene expression studies, THP-1 macrophages and HeLa were infected with Cb and other bacterial strains at MOI of 50. For Lp, Bmv, Li and STm, infections, cells were washed with RPMI 1 h p.i., and treated with 100 μg/mL gentamicin (Sigma-Aldrich) for 1 h to eliminate extracellular bacteria. Cells were then maintained in antibiotic-free medium and harvested at the indicated time points for downstream RNA extraction and RT-qPCR analysis.
RNA sequencing and data analysis
THP-1 macrophages were infected with 16 different strains as described above. Total RNA was isolated using Trizol Reagent (Invitrogen, #15596026) followed by purification with the Direct-zol RNA Miniprep Plus Kit (Zymo Research, # R2072), according to the manufacturer’s instructions. RNA quality and quantity were assessed using the 4200 TapeStation system (Agilent Technologies) prior to library preparation.
RNA sequencing and library preparation were performed by the Texas A&M AgriLife Genomics and Bioinformatics Service. Libraries were prepared from total RNA using the TruSeq Stranded Total RNA-Seq Library Kit (Illumina) with rRNA depletion, and cDNA was synthesized using random primers. Paired-end libraries (2 × 150-bp) were constructed according to the manufacturer’s protocol and sequenced on the Illumina NovaSeq6000 platform.
RNA-seq data processing and alignment
Raw reads in FASTQ format were processed using FastQC to remove adapters, poly-N stretches, and low-quality reads, yielding high-quality clean reads. Clean FASTQ data were mapped to the Homo sapiens reference genome (GRCh38.p12) using STAR (v. 2.5.4b) with the corresponding GENCODE annotation file (gencode.v30.chr_patch_hapl_scaff.annotation.gff3). Mapping yielded 35.9–83.4 million aligned reads per sample, representing 73–87% alignment efficiency. The uniquely mapped reads were used for gene-level quantification, with read counts assigned to genes only if they overlapped a single gene annotation. Both ends of the paired-end reads were checked for overlaps.
RNA-seq data analysis
The raw RNA-seq expression data were then pre-processed by filtering out the transcripts with count per million (CPM) below 1 (low-expressed transcripts). Filtered data were then normalized using the DESeq function implemented in the DESeq2 R package (v. 1.36.0), adjusting for known biological treatment conditions of the pathogen-infected samples76. A generalized linear model (GLM) design matrix was used to incorporate treatment conditions (mock vs. infected) and to adjust for biological covariates, enabling the model to distinguish true treatment effects from confounding factors. Following the instructions from DESeq2, we then performed Variance stabilizing transformation (VST)77 using the VST function to stabilize variance across the mean count data. Differential expression analysis was performed using the DESeq2 R package. Statistical significance was determined using Benjamini–Hochberg’s correction to control the false discovery rate (FDR). Genes with an adjusted p-value < 0.05 and log2 fold change (log2FC) > 1 or <−1 were considered differentially expressed. The normalized expression matrix (pre-processed/raw read counts after aforementioned normalization) was used to generate the principal component analysis (PCA) plot. All visualizations, including box plots, volcano plots, and hierarchical clustering heatmaps, were generated using R v. 3.6.0 (www.r-project.org).
Prediction and annotation of lncRNA targets
The human lncRNA catalog used in this study was extracted from the GENCODE human gene annotation database. Identified lncRNAs were reclassified as lincRNA, antisense lncRNA, or sense lncRNA based on their annotated biotypes in the GENCODE catalog. A list of anti-sense or sense-overlapping lncRNAs was filtered using manually annotated gene annotations from GENCODE. Antisense or sense-overlapping lncRNAs were filtered using manually curated annotations from GENCODE. To identify putative cis-associated targets, adjacent protein-coding genes located within 5 kb of each lncRNA were selected based on genomic proximity. Correlation analysis was performed to assess co-expression between lncRNAs and their neighboring genes. Pearson correlation coefficients (PCC) were calculated using the cor() function in R (v. 3.6.0).
Identification of DE-lncRNA in Cb infection
A list of DE-lncRNAs for each strain was generated and compared to those identified in Cb infection. An UpSet plot from the concatenated list of DE-lncRNAs in all strains was generated using the R package UpSetR (v. 1.4.0), to visualize strain-specific and shared DE-lncRNAs across all infections.
Functional enrichment and pathway analysis
To investigate the functional relevance of DE lncRNA-mRNA pairs and proteins identified from MS analysis, GO (http://www.geneontology.org/) and KEGG enrichment analyses were performed. Enrichment analysis was conducted using a one-sided Fisher’s Exact Test. p-values were adjusted for multiple testing using the Benjamini–Hochberg method, and an adjusted p-value ≤ 0.05 was considered statistically significant. Enrichment results are reported as −log10(p-value), and overlap sizes (number of matching genes or proteins per term) were visualized in bar plots.
Coding-non-coding co-expression (CNC) network
A CNC network was constructed to evaluate the correlation between DE-lncRNAs and mRNAs, as described previously35. DE-mRNAs associated with the GO terms and KEGG pathways were selected, and the median value was calculated if the coding genes had several transcripts. PCCs were calculated between CYP1B1-AS1 and CYP1B1. A correlation was considered significant if the absolute PCC was ≥0.9. The CNC network was constructed and visualized using Cytoscape (v. 3.10.2). A partial view of the network was manually curated for data representation. In the visualized networks, positive correlations are depicted as solid lines, while negative correlations are shown as dashed lines.
Polyadenylation assay
Total RNA from THP-1 cells was extracted using Trizol Reagent and the Direct-zol RNA Miniprep Plus Kit. cDNA was synthesized using the iScript Select cDNA Synthesis Kit (Bio-Rad, #1708897) with oligo-dT primers26. The resulting cDNA was used for PCR amplification of CYP1B1-AS1, NEAT1 (lncRNA control), CYP1B1, and ACTB (mRNA control) using PrimeSTAR HS DNA Polymerase with GC Buffer (Takara Bio, # R044B) and gene-specific primers (Supplementary Data 1). PCR products were resolved on a 2% agarose gel containing ethidium bromide and visualized using the ChemiDoc Imaging System (Bio-Rad).
Cellular localization assay
Cytoplasmic and nuclear RNA fractions from THP-1 cells were obtained using the PARIS kit (Invitrogen, #AM1921), following the manufacturer’s instructions. RNA from each fraction was extracted and reverse-transcribed to cDNA for quantitative PCR (RT-qPCR) analysis of CYP1B1-AS1. Expression levels were normalized to ACTB and compared relative to the cytoplasmic fraction26 to determine subcellular distribution.
Protein coding potential assay
To evaluate the translational potential of CYP1B1-AS1, ribosome profiling (Ribo-seq) analysis was conducted using the GWIPS-viz database (gwips.ucc.ie), utilizing global aggregate tracks compiled from multiple studies. Short open reading frames (sORFs) within CYP1B1-AS1 were predicted using NCBI ORF Finder (https://www.ncbi.nlm.nih.gov/orffinder/), and their non-coding potential was assessed using both the Coding Potential Assessment Tool (CPAT) (http://lilab.research.bcm.edu/) and Coding Potential Calculator 2 (CPC2) (https://cpc2.gao-lab.org/index.php).
Synthetic transcripts of CYP1B1-AS1, DANCR, CYP1B1, GAPDH, and predicted ORFs (ORF17 and ORF62) were obtained from Integrated DNA Technologies (IDT) and cloned into the pcDNA-FLAG vector using Gibson Assembly (NEB, #E2611L). Primer sequences used for construct generation are listed in Supplementary Data 1. HEK293T cells were transfected with the constructs using Lipofectamine 3000 (Invitrogen, #L3000015) according to the manufacturer’s instructions. FLAG-tag expression was assessed by western blotting using an anti-FLAG mouse monoclonal antibody (Invitrogen, #MA1-91878), 24 h post-transfection.
Promoter region and TF prediction, and bidirectional promoter dual luciferase assay
Promoter regions of CYP1B1-AS1 were identified using the UCSC Genome Browser and the EPD new promoter database. ChIP-seq signals for active promoter marks, including H3K4me3 and H3K27Ac, were analyzed across multiple publicly available datasets.
The region between CYP1B1-AS1 and CYP1B1, upstream of the transcription start site (+1), was screened for promoter regions, TF binding sites (TFBS), and CpG/methylation islands, as described previously78. A 600 bp promoter fragment upstream of CYP1B1-AS1 (PCYP1B1-AS1) was cloned into the pGL4-Luc-Rluc dual-reporter vector (Addgene #64034). HEK293T cells were seeded in 96-well plates (Corning) and transfected with 100 ng of plasmid DNA per well (n = 5) using Lipofectamine 3000. Luciferase activity was measured using the Renilla luciferase assay kit (Promega, #E2810) and firefly luciferase assay kit (Promega, #E1500) on a BioTek Cytation 5 Reader.
siRNA mediated knockdown of CYP1B1-AS1, CYP1B1 and AHR
Knockdown of CYP1B1, CYP1B1-AS1, and AHR was performed in HeLa, THP-1 macrophages, and HEK293T cells using Lipofectamine RNAiMAX transfection reagent (Invitrogen, #13778150), following the manufacturer’s protocol. Silencer Select Pre-Designed siRNAs (Thermo Fisher Scientific, #4392421) targeting the respective genes were used (Supplementary Data 1). Silencer Select Negative Control No. 1 siRNA (Thermo Fisher Scientific, #4390844) and GAPDH Positive Control siRNA (Thermo Fisher Scientific, #4390850) served as negative control (NC) and positive control (PC), respectively. Combinations of siRNAs were used to generate the following knockdown groups: lncCYPB, CYPB, dCYPB, and AHR-KD.
To investigate cis-regulation of CYP1B1 by CYP1B1-AS1, knockdowns were performed in THP-1 macrophages and HeLa cells. Cells were seeded in six-well plates and transfected with 75 pM siRNAs. After 48 h, cells were harvested in Trizol Reagent for RNA extraction and M-PER lysis buffer (Invitrogen, #78501) for protein extraction. CYP1B1-AS1 and CYP1B1 expression were quantified by RT-qPCR, normalized to HPRT, and compared to NC-treated cells. Protein lysates were analyzed by western blot using an anti-CYP1B1 antibody [EPR14972]—C-terminal Rabbit monoclonal antibody (Abcam, #ab185954) and anti-AHR (D5S6H) Rabbit mAb (Cell Signaling, #83200S). Densitometry analysis was performed to quantify CYP1B1 and AHR expression, normalized to β-actin (Abcam, #ab8227) as a loading control.
CYP1B1-AS1 and CYP1B1 expression levels in AHR-KD cells were analyzed by RT-qPCR and western blot. For RT-qPCR, gene expression was normalized to HPRT and compared to NC-treated controls. For the western blot, protein lysates were analyzed for CYP1B1 expression and compared to untreated (UT) and NC-treated cells.
Over-expression of CYP1B1-AS1 and CYP1B1
CYP1B1-AS1 and CYP1B1 were cloned into a pcDNA3.1/Zeo (+) vector (Invitrogen, #V86020) to generate pCYP1B1-AS1 and pCYP1B1 constructs (Supplementary Data 1) using Gibson Assembly Master Mix. HEK293T and HeLa cells were transfected using Lipofectamine™ 3000 according to the manufacturer’s instructions. Total RNA was isolated 24 h post-transfection, and RT-qPCR analysis was performed to assess CYP1B1 expression, normalized to HPRT and compared to untransfected controls.
mRNA stability assay
CYP1B1 mRNA stability was indirectly evaluated by analyzing its half-life26,79. Briefly, lncCYPB and NC HeLa cells were treated with 5 μg/ml actinomycin D (Gibco). RNA was isolated at 1 h intervals for 5 h post-treatment using Trizol Reagent for RT-qPCR analysis. The ΔCt of CYP1B1 and HPRT was calculated by subtracting the Ct value at each time point from the value at 0 h (t0). Relative mRNA abundance was determined using the 2(−ΔCt) method. The data were then fitted using non-linear regression curve fitting (one-phase decay, ordinary-fit) to calculate the half-lives at 95% CI.
RNA isolation, cDNA synthesis and RT-qPCR analysis
Cells were harvested at the indicated time points in Trizol Reagent. Total RNA was extracted using the Direct-zol RNA Miniprep Kit prior to cDNA synthesis. RT-qPCR was performed using SsoAdvanced Universal SYBR® Green Supermix (Bio-Rad, #1725271) on a CFX Opus 384 Real-Time PCR System (Bio-Rad, #12011452). All reactions were run in technical triplicate, and gene expression was normalized to endogenous controls ACTB or HPRT, as specified. Fold change in expression was calculated by dividing the normalized expression in the test sample by that of the control, which was set to 1.0 unless otherwise noted. Genes with a fold change ≥ 1.5 were considered upregulated, while those with a fold change ≤ 0.5 were considered downregulated. For genes with expression values ≤ 0.5 relative to the mock control, the degree of downregulation was expressed as the inverse (1/expression ratio) to clearly represent the magnitude of change.
GE determination for infection and growth curve analysis
The concentration of axenically grown Cb strains was determined by isolating bacterial DNA using GenElute Bacterial Genomic DNA Kits (Sigma-Aldrich, #NA2120). A standard curve was generated from known concentrations of Cb genomic DNA, and absolute quantification of the strains was performed using Taqman probes, com1 gene-specific primers, and TaqMan Fast Advanced Master Mix (Applied Biosystems, #4444557) on a QuantStudio™ 6 Flex (Applied Biosystems, #4485691).
THP-1 macrophages and HEK293T cells were seeded in 24-well plates and treated with siRNA (25 pM) to generate knockdowns. Infections were performed at an MOI of 50 or 100 (as indicated) at 48 h siRNA post-treatment. A second round of siRNA treatment was administered 3 days p.i. to maintain knockdown. Genomic DNA was isolated from infected cells at the indicated time p.i. using the High Pure PCR Template Preparation Kit (Roche, #11796828001). Purified DNA was quantified using RT-qPCR as previously described80. GE counts were compared to non-treated cells infected with Cb (NMII) and CbA (ΔdotA). For overexpression growth curves, cells were transfected with pCYP1B1-AS1 and pCYP1B1, followed by infection 24 h post-transfection. GE quantification was performed by RT-qPCR in comparison to empty vector-transfected controls.
For Cb GE determination in AHR-KD THP-1 macrophages, cells were infected with C. burnetii until 7 days p.i. DNA was isolated and quantified by RT-qPCR as described above. GE values were compared to non-treated NMII- and ΔdotA-infected controls.
Indirect immunofluorescence microscopy
THP-1 cells were seeded on coverslips in 24-well plates, treated with siRNA, and infected as described. At 5 days p.i., cells were fixed with 4% paraformaldehyde (PFA). Coverslips were permeabilized with 0.1% saponin, blocked with 2% BSA and stained with anti-C. burnetii guinea pig sera (James Samuel Laboratory, TAMHSC)74, anti-LAMP-1 mouse antibody (Abcam, #ab25630), and Hoechst 33342 (Thermo Scientific, #62249) at dilutions of 1:1000, 1:250, and 1:10,000, respectively. Secondary antibodies, anti-guinea pig Alexa Fluor 647 (Invitrogen, #A-21450) and goat anti-mouse Alexa Fluor 488 (Invitrogen, #A-11001), were used at 1:1000 dilutions. The stained coverslips were imaged using an Olympus FV3000 scanning confocal microscope. Quantitative measurements of CCV were performed using ImageJ (NIH; Java v. 1.8.0_322; IJ. get v. 2.16.0/1.54p) with Qupath plug-in (v. 0.5.1), and Labkit algorithms, with each LAMP-1+ bacterial compartment representing an individual CCV.
Cytokine analysis
THP-1 cells were seeded, treated with siRNA, and infected as described earlier. Supernatants were collected 24 h p.i. from infected and mock controls. The supernatants were analyzed by Eve Technologies’ Human Focused 15-Plex Discovery Assay (Millipore-Sigma) on the Luminex 200 system, following the manufacturer’s protocol. Cytokine concentrations were measured in biological triplicates, and results were reported as the average of samples in pg/ml.
For mRNA analysis, infected and control cells were collected at 12 h p.i., using Trizol reagent, and RNA was isolated for RT-qPCR analysis. Gene expression was normalized to HPRT and compared to mock.
AHR agonist treatment
THP-1 macrophages were treated with FICZ (Sigma-Aldrich, #SML1489) for 1 h. Media containing FICZ was removed, and cells were infected with Cb (NMII) at an MOI of 50. Infected cells were collected at 24, 48, and 72 h p.i. using Trizol Reagent for RNA isolation and M-PER lysis buffer for western blot analysis. CYP1B1-AS1 expression levels were analyzed by RT-qPCR, normalized to HPRT and compared to the mock control. For the western blot, CYP1B1 protein expression in infected cells was compared to control using an anti-CYP1B1 rabbit polyclonal antibody. Densitometry was performed to calculate the expression of CYP1B1, normalized to β-Actin as a loading control.
AHR nuclear translocation assay
THP-1 macrophages were seeded in 10 cm culture plates, treated with FICZ (400 nM), and infected with Cb (NMII). Nuclear and cytoplasmic proteins were isolated using NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific, #78833) according to the manufacturer’s instructions. For western blot analysis, AHR protein expression in infected cells was compared to controls in whole-cell lysates, cytoplasmic, and nuclear fractions. Anti-α-Tubulin and (Abcam, #ab7291) and anti-Lamin B1 antibody (Abcam, #ab16048) served as cytoplasmic and nuclear controls, respectively. Densitometry analysis was performed to quantify AHR expression, normalized to whole-cell lysates and the cytoplasmic fraction.
HepG2-based AHR luciferase reporter assay
HepG2 Lucia AHR cells (InvivoGen, #HPGL-AHR), stably expressing a dioxin-responsive luciferase reporter, were treated with FICZ or CH-223191 for 1 h. Cells were then infected with Cb at an MOI of 100 or left uninfected (Mock). Luciferase induction levels were measured from the media supernatant using the Quanti-Luc 4 Lucia/Gaussia reagent (InvivoGen, #rep-qlc4lg1) and BioTek Cytation 5 Reader. Induction levels were compared to mock over a time course of up to 150 h p.i.
To confirm HepG2 Lucia reporter cell infection, immunofluorescence imaging of Cb-infected HepG2 cells was performed as described previously. Imaging was conducted at 3 days p.i. using an Olympus FV3000 scanning confocal microscope. In parallel, CYP1B1-AS1 and CYP1B1 expression levels were measured in the HepG2 reporter cells after infection. RNA was harvested at time points, from 1 to 150 h p.i., and analyzed by RT-qPCR.
Human monocyte-derived macrophages
Human PBMCs were resuspended in 1X MojoSort Buffer (BioLegend, #480017), and monocytes were enriched from PBMCs using the MojoSort Human CD14+ Monocyte Isolation Kit (BioLegend, #480047) via negative selection. Following enrichment, monocyte purity was assessed by flow cytometry, comparing CD14+ monocytes in PBMCs and post-enriched cells. Cells were stained with FITC anti-human CD14 mouse monoclonal antibody [HCD14] (BioLegend, #325603) and 7-AAD (BioLegend, #420403) for purity and viability assessment. The gating strategy used for this analysis is provided in Supplementary Fig. 18a.
Human monocyte-derived macrophages (hMDMs) were differentiated from purified monocytes by culturing them in complete RPMI 1640 supplemented with 10% FBS, HEPES, penicillin/streptomycin (Thermo Scientific), and 10 ng/mL recombinant human GM-CSF (R&D Systems, #215-GM-050). On day 3, the culture medium was replaced with fresh RPMI 1640 containing 10 ng/mL GM-CSF, and cells were harvested on day 6.
hMDM infection with C. burnetii NMI
Differentiated hMDMs were seeded in 24-well plates at a density of 2 × 10⁵ cells per well in complete RPMI 1640 without antibiotics for RT-qPCR analysis. Cells were infected with Cb (NMI) at an MOI of 10 or treated with LPS (10 ng/mL; Millipore-Sigma, #L2637) as a positive control. At the indicated time points, RNA was harvested by lysing cells in Trizol reagent. The expression levels of target genes were quantified using ACTB as the endogenous control. Fold-change in gene expression upon NMI infection or LPS treatment was determined by normalizing transcript expression to uninfected controls (Mock, normalized to 1).
Flow cytometry analysis
The assays were performed in THP-1 macrophages at the indicated time p.i., using LSR Fortessa X20 flow cytometer (BD Biosciences). To measure intracellular ROS, cells were stained with 20 µM DCFDA/H₂DCFDA (Abcam, #ab113851) according to the manufacturer’s protocol and re-suspended in 1X buffer (supplied with kit) with 4% FBS. ROS levels were measured at an excitation/emission of 488/535 nm.
For mitochondrial ROS and mass assessment, cells were stained with 5 μM MitoSOX Mitochondrial Superoxide Indicators (Invitrogen, #M36005) and 200 nM Mitotracker Green dye (Invitrogen, #M46750) manufacturer’s protocol. Mitochondrial ROS and mass were measured at excitation/emission of 510/580 and 488/516 nm, respectively. Mitochondrial membrane potential was assessed using the 20 nM MitoProbe TMRM assay (Invitrogen, #M20036) as per the protocol and analyzed at 561/585 nm.
Apoptosis assay was assessed using FITC-Annexin-V Apoptosis Detection Kit with PI (BioLegend, #640914) following the manufacturer’s instructions. Annexin-V and PI signals were measured using 488 and 610/10 nm lasers, respectively. The gating strategies for these assays are provided in Supplementary Figs. 18b, c and 19.
Seahorse metabolic assays
Seahorse XF Mito Stress Test Kits (Agilent, #103015-100) and cartridges were prepared according to Agilent’s protocols and analyzed using an Agilent Seahorse XF 96-well analyzer. THP-1 macrophages were transfected with siRNA 48 h before infection at a density of 2 × 10⁴ cells per well (n = 3). Control and knockdown conditions were either mock-infected or Cb (NMII)-infected. Cells were processed 24 h p.i., following the manufacturer’s instructions, and were analyzed using the analyzer.
Immunoprecipitation (IP), mass spectrometry and analysis
FLAG-IP- HEK293T cells were seeded in 10 cm dish and transfected with 14 µg of either pcDNA-CYP1B1 or pcDNA-eGFP (Supplementary Data 1). For the infection group, one set of pcDNA-CYP1B1-transfected cells at 24 h post-transfection was infected with Cb at an MOI of 100. The experiment was conducted in biological duplicates (n = 2). Cells were harvested 48 h post-transfection and lysed in ice-cold RIPA buffer (Thermo Scientific, #89900) supplemented with Halt Protease and Phosphatase Inhibitor Cocktail (Thermo Scientific, #78442). Lysates were centrifuged at 12,000×g at 4 °C for 10 min, and supernatants were incubated with anti-FLAG M2 beads (Sigma-Aldrich, #A2220), and IP was performed per manufacturer’s instructions, followed by SDS–PAGE on a commercial pre-cast gel (Bio-Rad, #4569034). Sample bands were excised and sent for LC-MS/MS analysis.
Samples (n = 2) were digested overnight with trypsin (Pierce) following reduction and alkylation with DTT and iodoacetamide (Sigma-Aldrich). Following solid-phase extraction cleanup with an Oasis HLB µelution plate (Waters), the resulting peptides were reconstituted in 10 μL of 2% (v/v) acetonitrile (ACN) and 0.1% trifluoroacetic acid in water. 5 μL of this was injected onto an Orbitrap Eclipse mass spectrometer (Thermo) coupled to an Ultimate 3000 RSLC-Nano liquid chromatography system (Thermo). Samples were injected onto a 75 μm i.d., 75-cm long EasySpray column (Thermo), and eluted with a gradient from 0% to 28% buffer B over 90 min. Buffer A contained 2% (v/v) ACN and 0.1% formic acid in water, and buffer B contained 80% (v/v) ACN, 10% (v/v) trifluoroethanol, and 0.1% formic acid in water. The mass spectrometer operated in positive ion mode with a source voltage of 2.5 kV and an ion transfer tube temperature of 300 °C. MS scans were acquired at 120,000 resolution in the Orbitrap, and up to 10 MS/MS spectra were obtained in the Orbitrap for each full spectrum acquired using higher-energy collisional dissociation (HCD) for ions with charges 2–7. Dynamic exclusion was set for 25 s after an ion was selected for fragmentation.
Raw MS data files were analyzed using Proteome Discoverer v. 3.0 (Thermo), with peptide identification performed using Sequest HT searching against the human reviewed protein database from UniProt (downloaded November 15, 2022, 20,327 entries). Fragment and precursor tolerances of 10 ppm and 0.6 Da were specified, and three missed cleavages were allowed. Carbamidomethylation of Cys was set as a fixed modification, with oxidation of Met set as a variable modification. The false-discovery rate (FDR) cutoff was 1% for all peptides. Peptide peak intensities were used as peptide abundance values.
Enriched proteins from CYP1B1 IP were compared, with background proteins filtered using the CRAPome 2.0 database. Protein–protein interactions were analyzed using STRING (v. 12) and Cytoscape (v. 3.10.2), and a partial network of proteins interacting with CYP1B1 was manually constructed for data representation.
Western blot and antibodies
For western blot analysis, cell lysates were resolved on Bio-Rad pre-cast SDS–PAGE gels and transferred to polyvinylidene difluoride (PVDF) membranes (Millipore-Sigma, #IPFL00010). The following primary antibodies were used at the listed dilutions: anti-CYP1B1 antibody [EPR14972] - C-terminal Rabbit monoclonal antibody (1:1000), anti-DYKDDDDK (FLAG) Tag mouse Monoclonal Antibody (FG4R) (1:1000), anti-beta Actin antibody-Loading Control-Rabbit polyclonal (1:5000), anti-GAPDH mouse monoclonal antibody [6C5] (1:3000; Abcam, #ab8245), anti-AHR (D5S6H) Rabbit mAb (1:1000), anti-alpha Tubulin antibody [DM1A]-Loading Control-Mouse Monoclonal (1:5000), anti-Lamin B1 antibody-Nuclear Envelope Marker-rabbit polyclonal antibody (1:1000), and FITC anti-human CD14 mouse monoclonal antibody [HCD14] (for PBMCs and monocytes;1:50). For detection, the secondary antibodies used were: IRDye 680RD goat anti-mouse IgG (LI-COR Biosciences, #926-68070), IRDye 680RD goat anti-rabbit IgG (LI-COR Biosciences, #926-68071), and IRDye 800CW goat anti-rabbit IgG (LI-COR Biosciences, #926-32211), each at a 1:10,000 dilution. Blots were imaged using a LI-COR Odyssey Fc imager. Densitometry analysis was performed using ImageJ software and normalized to the respective internal control (β-Actin or α-Tubulin) band. The full uncropped blots are provided in the Source Data file.
Statistical analysis
All data are representative of atleast three independent experiments (n = 3), unless otherwise specified in the figure legends. Post-acquisition analysis of flow cytometry data was performed using FlowJo v.10 software (BD Biosciences, USA). Statistical significance was calculated using either a two-tailed unpaired Student’s t-test, one-way ANOVA with Sidak’s post hoc test, one-way ANOVA with Dunnett’s post hoc test, or two-way ANOVA with Tukey’s, Sidak’s, or Dunnett’s multiple comparisons test. All statistical analyses were performed using GraphPad Prism v.10 software (San Diego, USA).
Key reagents, software, and equipment
All reagents, software, and equipment used in this study, along with their identifiers, are listed in Supplementary Data 1.
Resource availability
Further requests for resources and reagents should be directed to and will be fulfilled by the lead contact, James E. Samuel (jsamuel@tamu.edu).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA-seq data generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject accession code PRJNA1228654. Processed RNA-seq data are available at the Gene Expression Omnibus (GEO) database under accession code GSE292317, and are linked to the same BioProject. The FLAG-immunoprecipitation mass spectrometry (FLAG-IP-MS) data are available in the MassIVE database (a ProteomeXchange partner repository) under accession code MSV000096725. All datasets are publicly accessible without restriction. Additional data supporting the findings of this study are provided in Supplementary Information, Supplementary Datasets, and Source Data files. Source data are provided with this paper.
Code availability
This study does not report original code.
References
van Schaik, E. J., Chen, C., Mertens, K., Weber, M. M. & Samuel, J. E. Molecular pathogenesis of the obligate intracellular bacterium Coxiella burnetii. Nat. Rev. Microbiol. 11, 561–573 (2013).
Bird, L. E. et al. Coxiella burnetii manipulates the lysosomal protease cathepsin B to facilitate intracellular success. Nat. Commun. 16, 3844 (2025).
Larson, C. L., Beare, P. A., Howe, D. & Heinzen, R. A. Coxiella burnetii effector protein subverts clathrin-mediated vesicular trafficking for pathogen vacuole biogenesis. Proc. Natl Acad. Sci. USA 110, E4770–E4779 (2013).
Luhrmann, A., Nogueira, C. V., Carey, K. L. & Roy, C. R. Inhibition of pathogen-induced apoptosis by a Coxiella burnetii type IV effector protein. Proc. Natl Acad. Sci. USA 107, 18997–19001 (2010).
Steiner, S., Meir, A. & Roy, C. R. Coxiella burnetii encodes an LvgA-related protein important for intracellular replication. Cell. Microbiol. 23, e13331 (2021).
Howe, D., Shannon, J. G., Winfree, S., Dorward, D. W. & Heinzen, R. A. Coxiella burnetii phase I and II variants replicate with similar kinetics in degradative phagolysosome-like compartments of human macrophages. Infect. Immun. 78, 3465–3474 (2010).
Gregory A. E., van Schaik E. J., Russell-Lodrigue K. E., Fratzke A. P. & Samuel J. E. Coxiella burnetii intratracheal aerosol infection model in mice, guinea pigs, and nonhuman primates. Infect. Immun. 87, e00178-19 (2019).
Burette, M. et al. Modulation of innate immune signaling by a Coxiella burnetii eukaryotic-like effector protein. Proc. Natl Acad. Sci. USA 117, 13708–13718 (2020).
Duncan-Lowey, J., Crabill, E., Jarret, A., Reed, S. C. O. & Roy, C. R. The Coxiella burnetii effector EmcB is a deubiquitinase that inhibits RIG-I signaling. Proc. Natl Acad. Sci. USA 120, e2217602120 (2023).
Watson Robert, O. et al. The cytosolic sensor cGAS detects Mycobacterium tuberculosis DNA to induce type I interferons and activate autophagy. Cell Host Microbe 17, 811–819 (2015).
Monroe, K. M., McWhirter, S. M. & Vance, R. E. Identification of host cytosolic sensors and bacterial factors regulating the type I interferon response to Legionella pneumophila. PLoS Pathog. 5, e1000665 (2009).
Xu, L. et al. Salmonella induces the cGAS-STING-dependent type I interferon response in murine macrophages by triggering mtDNA release. mBio 13, e03632–03621 (2022).
Boxx, G. M. & Cheng, G. The roles of type I interferon in bacterial infection. Cell Host Microbe 19, 760–769 (2016).
Jastrab, J. B. & Kagan, J. C. Strategies of bacterial detection by inflammasomes. Cell Chem. Biol. 31, 835–850 (2024).
Statello, L., Guo, C. J., Chen, L. L. & Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 22, 96–118 (2021).
Arunima, A., van Schaik, E. J. & Samuel, J. E. The emerging roles of long non-coding RNA in host immune response and intracellular bacterial infections. Front. Cell. Infect. Microbiol. 13, 1160198 (2023).
Schmerer, N. et al. A searchable atlas of pathogen-sensitive lncRNA networks in human macrophages. Nat. Commun. 16, 4733 (2025).
Aznaourova, M. et al. Noncoding RNA MaIL1 is an integral component of the TLR4–TRIF pathway. Proc. Natl Acad. Sci. USA 117, 9042–9053 (2020).
Aznaourova, M. et al. Single-cell RNA sequencing uncovers the nuclear decoy lincRNA PIRAT as a regulator of systemic monocyte immunity during COVID-19. Proc. Natl Acad. Sci. USA 119, e2120680119 (2022).
Agliano, F., Rathinam, V. A., Medvedev, A. E., Vanaja, S. K. & Vella, A. T. Long noncoding RNAs in host–pathogen interactions. Trends Immunol. 40, 492–510 (2019).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013).
Agarwal, S. et al. The long non-coding RNA LUCAT1 is a negative feedback regulator of interferon responses in humans. Nat. Commun. 11, 6348 (2020).
Aznaourova, M. et al. Noncoding RNA MaIL1 is an integral component of the TLR4-TRIF pathway. Proc. Natl Acad. Sci. USA 117, 9042–9053 (2020).
Barriocanal, M. et al. Long noncoding RNA EGOT responds to stress signals to regulate cell inflammation and growth. J. Immunol. 206, 1932–1942 (2021).
Lopes, M. et al. Co-expression analysis of lncRNA and mRNA identifies potential adipogenesis regulatory non-coding RNAs involved in the transgenerational effects of tributyltin. PLoS ONE 18, e0281240 (2023).
Zhou, H. et al. LncRNA-MAP3K4 regulates vascular inflammation through the p38 MAPK signaling pathway and cis-modulation of MAP3K4. FASEB J. 35, e21133 (2021).
Montano, C., Sergio, C., Eric, M., Sol, K. & Carpenter, S. Identification and functional characterization of lncRNAs involved in human monocyte-to-macrophage differentiation. RNA Biol. 21, 1115–1127 (2024).
Semon, M. & Duret, L. Evolutionary origin and maintenance of coexpressed gene clusters in mammals. Mol. Biol. Evol. 23, 1715–1723 (2006).
Chen, W. H., de Meaux, J. & Lercher, M. J. Co-expression of neighbouring genes in Arabidopsis: separating chromatin effects from direct interactions. BMC Genom. 11, 178 (2010).
Bansal, S. et al. Mitochondrial targeting of cytochrome P450 (CYP) 1B1 and its role in polycyclic aromatic hydrocarbon-induced mitochondrial dysfunction. J. Biol. Chem. 289, 9936–9951 (2014).
Falero-Perez, J., Song, Y. S., Sorenson, C. M. & Sheibani, N. CYP1B1: a key regulator of redox homeostasis. Trends Cell Mol. Biol. 13, 27–45 (2018).
Zhang, Y. et al. Dickkopf-2 knockdown protects against classic macrophage polarization and lipid loading by activation of Wnt/beta-catenin signaling. J. Cardiol. 78, 328–333 (2021).
Consortium, T. G. et al. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Atianand, M. K. & Fitzgerald, K. A. Long non-coding RNAs and control of gene expression in the immune system. Trends Mol. Med. 20, 623–631 (2014).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Gutierrez-Vazquez, C. & Quintana, F. J. Regulation of the Immune Response by the Aryl Hydrocarbon Receptor. Immunity 48, 19–33 (2018).
Yamada, T. et al. Constitutive aryl hydrocarbon receptor signaling constrains type I interferon-mediated antiviral innate defense. Nat. Immunol. 17, 687–694 (2016).
Lamas, B., Natividad, J. M. & Sokol, H. Aryl hydrocarbon receptor and intestinal immunity. Mucosal Immunol. 11, 1024–1038 (2018).
Michel, A. M. et al. GWIPS-viz: development of a ribo-seq genome browser. Nucleic Acids Res. 42, D859–D864 (2013).
Wang, L. et al. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 41, e74 (2013).
Kang, Y.-J. et al. CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res. 45, W12–W16 (2017).
Carlevaro-Fita, J. & Johnson, R. Global positioning system: understanding long noncoding RNAs through subcellular localization. Mol. Cell 73, 869–883 (2019).
Ponjavic, J., Oliver, P. L., Lunter, G. & Ponting, C. P. Genomic and transcriptional co-localization of protein-coding and long non-coding RNA pairs in the developing brain. PLoS Genet. 5, e1000617 (2009).
Seila, A. C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Clemente T. M. et al. Coxiella burnetii Blocks Intracellular Interleukin-17 Signaling in Macrophages. Infect. Immun. 86, e00532-18 (2018).
Mahapatra, S. et al. Coxiella burnetii employs the Dot/Icm type IV secretion system to modulate host NF-kappaB/RelA activation. Front. Cell. Infect. Microbiol. 6, 188 (2016).
Fischer, N. L. et al. Coxiella burnetii effector interacts with the host PAF1 complex and suppresses the innate immune response. Preprint at bioRxiv https://doi.org/10.1101/2022.04.20.488957 (2022).
Ikuta, T. et al. Nucleocytoplasmic shuttling of the aryl hydrocarbon receptor. J. Biochem. 127, 503–509 (2000).
Davarinos, N. A. & Pollenz, R. S. Aryl hydrocarbon receptor imported into the nucleus following ligand binding is rapidly degraded via the cytosplasmic proteasome following nuclear export. J. Biol. Chem. 274, 28708–28715 (1999).
Weindel, C. G. et al. Mitochondrial ROS promotes susceptibility to infection via gasdermin D-mediated necroptosis. Cell 185, 3214–3231.e3223 (2022).
Kelly, B. & O’Neill, L. A. J. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 25, 771–784 (2015).
Tait S. W. & Green, D. R. Mitochondrial regulation of cell death. Cold Spring Harb Perspect. Biol 5, a008706 (2013).
Kolenda, T. et al. lncRNA EGOT is the marker of HPV infection and a prognostic factor for HNSCC patients. Biomedicines 13, 798 (2025).
Carnero, E. et al. Long noncoding RNA EGOT negatively affects the antiviral response and favors HCV replication. EMBO Rep. 17, 1013–1028 (2016).
Li, N., Liu, Y. & Cai, J. LncRNA MIR155HG regulates M1/M2 macrophage polarization in chronic obstructive pulmonary disease. Biomed. Pharmacother. 117, 109015 (2019).
Rai, K. R. et al. MIR155HG plays a bivalent role in regulating innate antiviral immunity by encoding long noncoding RNA-155 and microRNA-155-5p. mBio 13, e02510–e02522 (2022).
Maarouf, M. et al. Identification of lncRNA-155 encoded by MIR155HG as a novel regulator of innate immunity against influenza A virus infection. Cell. Microbiol. 21, e13036 (2019).
Wen, S. et al. MIR222HG attenuates macrophage M2 polarization and allergic inflammation in allergic rhinitis by targeting the miR146a-5p/TRAF6/NF-κB axis. Front Immunol. 14, 1168920 (2023).
Li, R. et al. Long non-coding RNA Mir22hg-derived miR-22-3p promotes skeletal muscle differentiation and regeneration by inhibiting HDAC4. Mol. Ther. Nucleic Acids 24, 200–211 (2021).
Hezaveh, K. et al. Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity 55, 324–340.e328 (2022).
Domínguez-Acosta, O. et al. Activation of aryl hydrocarbon receptor regulates the LPS/IFNγ-induced inflammatory response by inducing ubiquitin-proteosomal and lysosomal degradation of RelA/p65. Biochem. Pharmacol. 155, 141–149 (2018).
Masuda, K. et al. Aryl hydrocarbon receptor negatively regulates LPS-induced IL-6 production through suppression of histamine production in macrophages. Int. Immunol. 23, 637–645 (2011).
Pinto, C. J. G., Ávila-Gálvez, M. Á, Lian, Y., Moura-Alves, P. & Nunes dos Santos, C. Targeting the aryl hydrocarbon receptor by gut phenolic metabolites: a strategy towards gut inflammation. Redox Biol. 61, 102622 (2023).
Giovannoni, F. & Quintana, F. J. SARS-CoV-2-induced lung pathology: AHR as a candidate therapeutic target. Cell Res. 31, 1–2 (2021).
Li, H. et al. The aryl hydrocarbon receptor is a novel negative regulator of interleukin-17-mediated signaling and inflammation in vitro. FEBS Lett. 593, 952–961 (2019).
Osbron, C. A., Lawson, C., Hanna, N., Koehler, H. S. & Goodman, A. G. Caspase-8 activity mediates TNFα production and restricts Coxiella burnetii replication during murine macrophage infection. Infect. Immun. 92, e00053–00024 (2024).
Benoit, M., Barbarat, B., Bernard, A., Olive, D. & Mege, J.-L. Coxiella burnetii, the agent of Q fever, stimulates an atypical M2 activation program in human macrophages. Eur. J. Immunol. 38, 1065–1070 (2008).
Cunha, L. D. et al. Inhibition of inflammasome activation by Coxiella burnetii type IV secretion system effector IcaA. Nat. Commun. 6, 10205 (2015).
Jansen A. F. M. et al. Viable Coxiella burnetii induces differential cytokine responses in chronic Q fever patients compared to heat-killed Coxiella burnetii. Infect. Immun. 86, e00333-18 (2018).
Honstettre, A. et al. Dysregulation of cytokines in acute Q fever: role of interleukin-10 and tumor necrosis factor in chronic evolution of Q fever. J. Infect. Dis. 187, 956–962 (2003).
Karki, R. et al. Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 184, 149–168.e117 (2021).
Bertheloot, D., Latz, E. & Franklin, B. S. Necroptosis, pyroptosis and apoptosis: an intricate game of cell death. Cell. Mol. Immunol. 18, 1106–1121 (2021).
Morgan, M. J. Liu Z-g. Crosstalk of reactive oxygen species and NF-κB signaling. Cell Res. 21, 103–115 (2011).
Case, E. D. R. et al. Primary murine macrophages as a tool for virulence factor discovery in Coxiella burnetii. Microbiol. Spectr. 10, e0248421 (2022).
Beare, P. A. et al. Dot/Icm type IVB secretion system requirements for Coxiella burnetii growth in human macrophages. mBio 2, e00175–00111 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Tibshirani, R. Estimating transformations for regression via additivity and variance stabilization. J. Am. Stat. Assoc. 83, 394–405 (1988).
Tang, L. et al. FOXO1-regulated lncRNA CYP1B1-AS1 suppresses breast cancer cell proliferation by inhibiting neddylation. Breast Cancer Res. Treat. 202, 397–408 (2023).
Ratnadiwakara, M. & Anko, M. L. mRNA stability assay using transcription inhibition by actinomycin D in mouse pluripotent. Stem Cells Bio Protoc. 8, e3072 (2018).
Gregory, A. E. et al. Soluble antigens derived from Coxiella burnetii elicit protective immunity in three animal models without inducing hypersensitivity. Cell Rep. Med. 2, 100461 (2021).
Liu, W., Tan, S., Bai, X., Ma, S. & Chen, X. Long non-coding RNA LINC01215 promotes epithelial–mesenchymal transition and lymph node metastasis in epithelial ovarian cancer through RUNX3 promoter methylation. Transl. Oncol. 14, 101135 (2021).
Gu, X., Chen, H., Li, R. & Guo, D. Roles of mir155hg and TNF-α in evaluation of prognosis of patients with systemic lupus erythematosus. J. Med. Biochem. 43, 265–272 (2024).
Mohamed, N. R., El-Fattah, A. L. A., Shaker, O. & Sayed, G. A. The diagnostic and predictive potential of lncRNA CASC2 targeting miR-155 in systemic lupus erythematosus patients with nephritis complication. Sci. Rep. 14, 30537 (2024).
Wood C. D. et al. Enhancer control of microRNA miR-155 expression in Epstein-Barr virus-infected B cells. J. Virol. 92, e00716-18 (2018).
Lu, F. et al. Epstein-Barr virus-induced miR-155 attenuates NF-kappaB signaling and stabilizes latent virus persistence. J. Virol. 82, 10436–10443 (2008).
Bogerd, H. P., Kennedy, E. M., Whisnant, A. W. & Cullen, B. R. Induced packaging of cellular microRNAs into HIV-1 virions can inhibit infectivity. mBio 8, https://doi.org/10.1128/mbio.02125-02116 (2017).
Sarkar, N. et al. Expression of microRNA-155 correlates positively with the expression of Toll-like receptor 7 and modulates hepatitis B virus via C/EBP-β in hepatocytes. J. Viral Hepat. 22, 817–827 (2015).
Peng, L. et al. Lipopolysaccharide facilitates immune escape of hepatocellular carcinoma cells via m6A modification of lncRNA MIR155HG to upregulate PD-L1 expression. Cell Biol. Toxicol. 38, 1159–1173 (2022).
Peng, L., Chen, Z., Chen, Y., Wang, X. & Tang, N. MIR155HG is a prognostic biomarker and associated with immune infiltration and immune checkpoint molecules expression in multiple cancers. Cancer Med. 8, 7161–7173 (2019).
Sokhanvar, Z., Elikaei, A. & Sharifi, Z. Evaluation of miR-222 expression in HBV Infected patients in comparison with HDV and HBV Co-infected patients. Avicenna J. Med. Biotechnol. 13, 223–225 (2021).
Lodge, R. et al. Host MicroRNAs-221 and -222 inhibit HIV-1 entry in macrophages by targeting the CD4 viral receptor. Cell Rep. 21, 141–153 (2017).
Luna Joseph, M. et al. Hepatitis C virus RNA functionally sequesters miR-122. Cell 160, 1099–1110 (2015).
Zhang, L., Li, C. & Su, X. Emerging impact of the long noncoding RNA MIR22HG on proliferation and apoptosis in multiple human cancers. J. Exp. Clin. Cancer Res. 39, 271 (2020).
Zhang Q. et al. The long noncoding RNA ROCKI regulates inflammatory gene expression. EMBO J. 38, e100041 (2019).
Ilott, N. E. et al. Long non-coding RNAs and enhancer RNAs regulate the lipopolysaccharide-induced inflammatory response in human monocytes. Nat. Commun. 5, 3979 (2014).
Cheng, Y., Liang, Y., Tan, X. & Liu, L. Host long noncoding RNAs in bacterial infections. Front. Immunol.15 (2024).
Tamgue O., Chia J. E., Brombacher F. Triptolide modulates the expression of inflammation-associated lncRNA-PACER and lincRNA-p21 in Mycobacterium tuberculosis-infected monocyte-derived macrophages. Front. Pharmacol. 12, 618462 (2021).
Chini, A., Guha, P., Malladi, V. S., Guo, Z. & Mandal, S. S. Novel long non-coding RNAs associated with inflammation and macrophage activation in human. Sci. Rep. 13, 4036 (2023).
Liu, Y. et al. LncRNA-PACERR induces pro-tumour macrophages via interacting with miR-671-3p and m6A-reader IGF2BP2 in pancreatic ductal adenocarcinoma. J. Hematol. Oncol. 15, 52 (2022).
Liu, Y. et al. The CTCF/LncRNA-PACERR complex recruits E1A binding protein p300 to induce pro-tumour macrophages in pancreatic ductal adenocarcinoma via directly regulating PTGS2 expression. Clin. Transl. Med. 12, e654 (2022).
Acknowledgements
The authors thank the TIGSS-Molecular Genomics Core at Texas A&M University for RNA-sequencing analysis, and Robbie Moore of the Texas A&M College of Medicine Flow Cytometry & Cell Sorting Facility for assistance with flow cytometric analysis. We also acknowledge the Integrated Microscopy and Imaging Laboratory at the Texas A&M College of Medicine (RRID:SCR_021637) and the UT Southwestern Proteomics Core for mass spectrometry analysis. We are grateful to Kristin Patrick for critical input and review of the manuscript. Schematics were created using BioRender.com. The work was supported by the Defense Threat Reduction Agency (HDTRA1210002, J.E.S.) and the National Institutes of Health (R01AI090142, J.E.S.). The views expressed in this article are those of the authors and do not represent the official policy or position of the U.S. Department of Defense or the U.S. Army.
Author information
Authors and Affiliations
Contributions
J.E.S., E.J.v.S., P.d.F., and A.A. conceived the study. J.E.S., A.A., and E.J.v.S. designed the methodology. A.A., S.N., S.M.B., S.D.C., A.P., and E.D.R.C performed the experiments. A.A., S.N., D.K., and X.Q. were responsible for data analysis. J.E.S. supervised the project. A.A. wrote the original draft of the manuscript. J.E.S., A.A., E.J.v.S., P.d.F., and X.Q. reviewed and edited the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Arunima, A., Niyakan, S., Butler, S.M. et al. CYP1B1-AS1 regulates CYP1B1 to promote Coxiella burnetii pathogenesis by inhibiting ROS and host cell death. Nat Commun 16, 7493 (2025). https://doi.org/10.1038/s41467-025-62762-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62762-2