Abstract
Pancreatic ductal adenocarcinoma (PDAC) poses a serious clinical challenge, demanding further exploration of its pathogenesis and therapeutic targets for metastasis, the main cause of mortality. Here, we identify STN1, a CST complex member crucial for maintaining telomere lengths and genome stability, as a key factor in promoting PDAC metastasis. Elevated STN1 levels correlate with poor patient survival, with oncogenic protein HOXB7 as an upstream transcription factor regulating STN1. Utilizing multiple PDAC experimental models, we discover STN1’s role in promoting metastasis by functioning as an upstream factor in epithelial-mesenchymal transition (EMT). Our mechanistic evidence suggests that during transcription, STN1 binds to structurally displaced single-stranded DNA flanking the R-loop, recruiting STAT3 to activate ZEB1 transcription independent of its known telomere maintenance function. Notably, STAT3 inhibitors show enhanced efficiency in restraining metastatic potential in STN1-overexpressed PDAC cells, offering a potential therapeutic avenue for targeting metastasis in STN1-overexpressed PDAC patients facing an unfavorable prognosis.
Similar content being viewed by others
Introduction
Pancreatic cancer poses a substantial global health challenge, ranking among the leading causes of cancer-related mortality1 despite its relatively low incidence2. As patients usually exhibit few symptoms in the early stages of the disease, pancreatic cancer is commonly diagnosed at an advanced stage. Pancreatic cancer is associated with multiple genetic and epigenetic alterations as well as a complex tumor microenvironment. These properties likely account for its resistance to most therapies, including chemotherapy, radiotherapy and molecularly targeted therapy, leading to an overall 5-year survival rate of <7%1. Metastasis emerges as the predominant cause of mortality in pancreatic cancer, responsible for 90% of deaths from solid tumors3,4. Various cellular programs determine the progression of metastasis and a key process among them is the epithelial-mesenchymal transition (EMT). EMT is a reversible cellular program in which malignant cells transform from the stable and well-organized epithelial state to a more energetic mesenchymal cell state. Induced by EMT transcription factors including SNAIL, SLUG and ZEB1, EMT activation converts non-metastatic tumor cells into cells capable of invading, surviving in the bloodstream and colonizing distant organs5. The therapeutic challenges posed by metastasis, particularly the systemic ramifications of metastatic lesions6, underscore the urgent need for molecular insights and targeted interventions to improve outcomes for pancreatic cancer patients.
STN1 constitutes an essential component of the CTC1-STN1-TEN1 (CST) complex, which is recognized for its critical functions in telomere replication and genome stability7. Alongside TEN1, which exhibits a high degree of evolutionary conservation across species, CTC1 demonstrates comparatively lower conservation. The CST complex regulates telomere length, inhibiting telomerase activity by binding to telomeric 3’ overhangs and interacting with POT1-TPP18. It also acts as a DNA polymerase alpha-primase (Pol-α) cofactor for C-strand synthesis8,9. The complex is involved in genome-wide DNA stability maintenance, preferentially binding to GC-rich regions, unfolding G-quadruplex structures, and facilitating RAD51 recruitment in response to replication stress10,11. Additionally, it protects stalled forks from MRE11-induced attacks and recruits Pol-α for canonical non-homologous end-joining repair at double-strand DNA break ends8,12,13. Loss of CST components leads to PARP inhibitor resistance and severe cellular proliferative defects14. CTC1-null mice face bone marrow failure via activation of ATR-dependent G2/M checkpoint15. While CST mutations are implicated in Coats plus syndrome and dyskeratosis congenita9, their further roles in cancer remain less explored, necessitating additional investigation for potential implications in this malignancy.
In this work, we identify STN1 as a potential regulator in the progression of pancreatic cancer. We find that STN1 is highly expressed in PDAC tissues and high STN1 levels correlate with adverse patient outcomes. We further speculate that STN1 promotes metastasis capacity of pancreatic cancer cells both in vivo and in vitro, and Stn1-/+, Pdx1-creER, KrasLSL.G12D/+, Tp53fl/fl (KPC) mice reveals diminished metastatic capabilities in contrast to KPC mice. Mechanistically, STN1 recruits STAT3 to the promoter of the EMT transcription factor ZEB1 and accelerates its rate of transcription, leading to heightened EMT in pancreatic cancer cells. Finally, we demonstrate that pancreatic cancer cells overexpressing STN1 exhibit heightened sensitivity to the STAT3 inhibitors, S3I-201 and napabucasin, suggesting a promising avenue for potential therapeutic interventions targeting PDAC metastasis, particularly for patients displaying elevated STN1 expression. Our study provides important insights into the role of STN1 in pancreatic cancer metastasis and suggests an alternative therapeutic approach for individuals affected by pancreatic cancer.
Results
Elevated STN1 expression is associated with adverse survival outcomes in PDAC patients
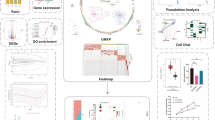
We aimed to identify candidate oncogenes driving tumor progression in PDAC using a combination of bioinformatic tools (Fig. 1a). Initially, we utilized TCGA-GTEx datasets, three large-scale Gene Expression Omnibus datasets16 consisting of 220 pairs of PDAC tumor tissues and non-tumor tissues as well as a single-cell RNA sequencing dataset17 of PDAC tumor tissues and normal tissues. Through this analysis, we identified 692 differentially expressed genes, upregulated in both tumor tissues and malignant ductal cells, with 12 genes associated with unfavorable overall survival and disease-free interval based on the TCGA datasets (Supplementary Fig. 1a, b). Among these 12 genes, all of them, except STN1, have been previously reported to play established roles in PDAC. For example, keratin 7 (KRT7) has been shown to be overexpressed in PDAC as well as associated with a worse prognosis and contribute to the immunosuppression in the PDAC microenvironment18. In addition, ADAM metallopeptidase domain 9 (ADAM9) has been shown to be overexpressed in several solid tumors include PDAC and contribute to tumor biology via proteolytic or non-proteolytic mechanisms19. These previously established roles served to validate our bioinformatic based screening approach and moreover, suggested that STN1 warranted further investigation. Consequently, we initiated our studies into the clinical significance and functional roles of STN1 in the progression of PDAC.
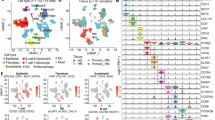
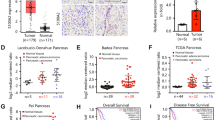
a Screening strategy for PDAC-associated genes using bioinformatics. b Analysis of TCGA data revealed a significant elevation in STN1 expression in PDAC tissues. c The bar plot illustrates the up-regulation of STN1 across all cell types in PDAC tissues. d qRT-PCR analysis demonstrated the relative levels of STN1 (normalized to ACTB) in peritumor and tumor tissues of PDAC patients (n = 55 biological replicates). e Immunoblot analysis showing STN1 protein levels in the indicated PDAC cell lines and normal pancreatic ductal cells. The assays were performed twice with similar results. f, g Representative IHC staining of STN1 protein in PDAC tissues and NATs of KPC mice. f Representative IHC images. Scale bar, left, 100 μm, right, 25 μm. g Quantification of IHC signals performed using ImageJ (n = 5 random fields from one section per group). h Representative IHC staining of STN1 protein in human PDAC tissues and matched NATs. Upper scale bar, 500 μm; lower scale bar, 100 μm. i Bar graphs showing IHC corresponding to STN1 protein levels of PDAC and NATs signals quantified using ImageJ (n = 90 biological replicates from each group). j Bar graphs showing higher STN1 protein levels in M1 versus M0 stage PDAC patients (n = 66 biological replicates for M0 and n = 24 biological replicates for M1). k Kaplan-Meier analysis conducted on 175 PDAC patients revealed a correlation between high STN1 expression and poor overall survival. OS, overall survival. l Kaplan-Meier analysis of overall survival in our PDAC patient cohort (n = 45 biological replicates from each group) demonstrating an association between up-regulation of STN1 protein and poorer survival. Quantification was performed using ImageJ. Statistical significance in (d, i) was determined using two-tailed paired t tests in GraphPad Prism. Statistical significance in (b, j) was determined using two-tailed unpaired t tests in GraphPad Prism. Statistical significance in (k, l) was determined using log rank tests. Error bars represent ± SEM. Source data are provided as a Source Data file.
Our screening results revealed the elevated expression of STN1 in PDAC tumor tissues, as validated in both the TCGA-GTEx (Fig. 1b) and tissue microarray datasets (Supplementary Fig. 1c–e). Elevated STN1 was observed across all cell types of tumor tissues, with the highest expression in malignant ductal cells compared to normal pancreas (Fig. 1c, and Supplementary Fig. 1f). Intriguingly, the other component of the CST complex, CTC1, did not exhibit the same expression pattern (Fig. 1c, and Supplementary Fig. 1f). Although TEN1 had a similar expression pattern as STN1, its high expression led to a better outcome in PDAC patients, which was opposite to that of STN1 (Supplementary Fig. 1g). This observation was validated through mRNA expression analysis of 55 self-collected PDAC tumor tissues and matched adjacent normal tissues (NATs) (Fig. 1d). Levels of STN1 protein were also elevated in four human pancreatic cancer cell lines (MIA PaCa-2, PANC-1, AsPC-1 and BxPC-3) compared to a normal pancreatic ductal cell line (HPNE) (Fig. 1e). Immunohistochemical (IHC) staining in tissues of the Pdx1-cre, KrasLSL.G12D/+; Tp53fl/fl (KPC) mouse model (Fig. 1f, g) and in a tissue microarray containing 90 PDAC specimens corroborated the higher levels of STN1 protein in tumor tissues (Fig. 1h, i). Notably, patients with evidence of metastasis (M1 stage) exhibited higher STN1 levels compared to those without metastasis (M0 stage) (Fig. 1j), and STN1 protein levels were not associated with the size of the primary tumor (T stage) (Supplementary Fig. 1i), indicating a potential role of STN1 in PDAC metastasis rather than tumor growth.
In the TCGA cohort of 175 patients, a high expression level of STN1 was significantly associated with poor survival (Fig. 1k, and Supplementary Fig. 1h). This clinical relevance of STN1 expression levels was further validated in our tissue microarray, where patients with higher STN1 protein level exhibited worse outcomes (Fig. 1l). Collectively, our findings strongly implicate a clinical relevance of elevated STN1 expression in PDAC tumor tissues with poor survival, indicating its potential significance in the clinical trajectory of PDAC.
STN1 facilitates PDAC progression through promotion of metastasis
To investigate the roles of STN1 in PDAC progression, two independent shRNAs specifically targeting STN1 were designed and transfected into the four pancreatic cell lines, MIA PaCa-2, PANC-1, AsPC-1 and BxPC-3. The efficiency of knockdown and rescue was confirmed by immunoblotting and qRT-PCR analyses (Fig. 2a, and Supplementary Fig. 2a). However, altering STN1 protein levels had no discernible effect on pancreatic cancer cell proliferation (Supplementary Fig. 2b–d). Subsequently, we probed the role of STN1 in migration and invasion using Transwell cell migration assays with and without Matrigel. These experiments showed that STN1 knockdown significantly attenuated the migration and invasion capacities of pancreatic cancer cells, both of which could be restored by reintroducing STN1 using pCDH-STN1 (Fig. 2b, c, Supplementary Fig. 2e, f). A more in-depth evaluation of invasion capacity using 3D morphogenesis Matrigel culture corroborated these findings (Fig. 2d, e). The in vivo experiments using intrasplenic injection models showed the consistent results (Fig. 2f–h).
a The effectiveness of STN1 knockdown in the MIA PaCa-2, PANC-1, AsPC-1 and BxPC-3 cell lines was confirmed by immunoblotting. The assays were performed twice with similar results. b, c The motility of the indicated cells was evaluated sequentially through Transwell migration assays and invasion assays. b Representative images of migration and invasion. Scale bar, 50 μm. c Migrated cells were counted by ImageJ in five random fields per well (n = 3 biological replicates). d, e The invasion potentials of the indicated MIA PaCa-2 cells were evaluated using a 3D multicellular tumor spheroid invasion assay. d Representative images of invasion. Scale bar, 200 μm. e Analysis of the invaded area was conducted using ImageJ (n = 15 spheroids per group). f–h The impact of STN1 knockdown and rescue with STN1 as well as various mutants on tumor metastasis was investigated using an inferior hemispleen implantation model in nude mice. STN1-knockdown and -rescue stable MIA PaCa-2 cell lines were utilized in the experiment. f Representative images for the liver metastasis of the indicated mice. The arrows point to liver metastasis generated by indicated cells. g Representative hematoxylin and eosin staining for the liver metastasis of the indicated mice. Scale bar, 500 μm. h Quantification of liver metastasis in the indicated mice was conducted visually (n = 6 biological replicates per group). i The effectiveness of ZEB1 knockdown in the sh1 + STN1-OBM cells was confirmed by immunoblotting. The assays were performed twice with similar results. Statistical significance of data shown in (c–h) was determined using two-tailed unpaired t tests in GraphPad Prism. ns, no statistically significant difference. Error bars represent ± SEM. Source data are provided as a Source Data file.
Considering the association between elevated STN1 level and PDAC progression, we postulated that the metastasis-promoting ability of STN1 was dose-dependent. To test this idea, we transfected different amounts of pCMV-Tag2B-STN1 into STN1 knockdown MIA PaCa-2 stable cell lines. Gradual increases in STN1 protein levels correlated with enhanced migration and invasion abilities in MIA PaCa-2 cells (Supplementary Fig. 2g–i). Thus, our results suggest that STN1 plays a pivotal role in driving PDAC progression by promoting metastasis rather than influencing proliferation. These findings are consistent with our earlier observations that high STN1 expression levels in PDAC patients correlates with poor overall survival, emphasizing the significance of metastasis in the high mortality of PDAC.
Decrease of STN1 in KPC mice restrains PDAC metastasis
To better understand the influence of STN1 on the metastatic progression of PDAC, we generated a Stn1 knockout mouse model by application of CRISPR-Cas9 methodology. Capitalizing on the high conservation of STN1 between humans and mice, guide RNA (gRNA) was designed to induce a frameshift mutation in exon 2 of mouse Stn1 (Fig. 3a). The observed embryonic lethality in Stn1 null mice prompted an examination of Stn1 protein levels in various organs of Stn1 heterozygous (Stn1-/+) mice, revealing a significant decrease in protein level without a compensatory upregulation of Stn1 protein (Fig. 3b). We then crossed Pdx1-creER, KrasLSL.G12D/+; Tp53fl/fl (KPC) mice with Stn1-/+ mice to generate Stn1-/+; KPC mice (hereafter referred to as KPCS mice) (Fig. 3a). The progeny exhibited normal embryonic development as well as pancreas morphology. Both KPC and KPCS mice were sacrificed on the verge of death near 16 weeks after tamoxifen treatment. Like KPC mice, nearly all the KPCS mice developed pancreatic tumors but, as anticipated, immunohistochemistry confirmed substantially decreased levels of Stn1 protein in their pancreatic tumor tissues (Fig. 3c, d). However, the KPCS mice demonstrated a more favorable outcome compared to their KPC counterparts (Fig. 3e).
a Scheme of the genetic mouse models for pancreatic cancer. Up, the KPC mouse model. Below, the KPCS mouse model. b Verification of the reduction in STN1 protein levels in various tissues, including heart, liver, spleen, lung, kidney, cerebrum, large intestine and pancreas, in Stn1-/+ mice. Equal tissue weights were loaded in each lane. The assays were performed twice with similar results. c H&E staining in PDAC, liver and lung metastases. Scale bar, 200 μm. Dotted lines represent liver or lung metastasis, while solid lines indicate the adhesion between liver and tumor tissues, rather than distant metastasis. d IHC staining of STN1 in sections of PDAC tissues indicating reduced EMT in KPCS mice. Scale bar, left, 100 μm, right, 25 μm. e Kaplan-Meier survival plot comparing KPC (n = 12) and KPCS (n = 13) mice that received tamoxifen injections at 4–5 weeks of age. Survival time at x axis includes the time before tamoxifen injections. f Percentage of mice developing liver metastases in KPC (n = 12) and KPCS (n = 13) mice. g Percentage of mice developing lung metastases in the indicated mice groups. Statistical significance in (e) was determined by log rank test in GraphPad Prism. Statistical significance in (f) and (g) was determined by two-tailed Fisher’s exact test in GraphPad Prism. Source data are provided as a Source Data file.
Consistent with their longer survival times, the KPCS mice had significantly fewer liver and lung metastases than the KPC mice (Fig. 3c, f and g). Liver metastasis manifested in only 15% of KPCS mice, substantially lower than the 50% incidence observed in KPC mice (Fig. 3f). Similarly, lung metastasis occurred in just 8% of KPCS mice compared to 33% of KPC mice from our cohort (Fig. 3g). These findings strongly suggest that that STN1 plays a pivotal role in promoting metastasis, thereby accelerating the progression of PDAC.
HOXB7 is an upstream transcription factor of STN1
We next investigated the mechanism underlying the elevated expression of STN1 in PDAC tumor tissues. Utilizing JASPAR, we employed a strategy to predict potential transcription factors capable of binding to the promoter region of STN1, with a specific focus on factors overexpressed in PDAC patients and co-expressing with STN1. Using this approach, we identified four potential upstream transcription factors-SNAI2, PPARD, HOXB7 and ZBTB7A, which exhibited elevated expression in PDAC tumor tissues and displayed a correlation with STN1 expression levels in PDAC patients (Fig. 4a, and Supplementary Fig. 3a, b). Notably, the elevated expression of three of these transcription factors (HOXB7, ZBTB7A, and SNAI2) was associated with a poorer clinical outcome (Fig. 4b, and Supplementary Fig. 3c). To experimentally validate their roles in STN1 transcriptional regulation, stable knockdown cell lines targeting the four transcription factors were generated in PANC-1 cells. The efficacy of SNAI2, PPARD, HOXB7 and ZBTB7A knockdown as well as STN1 mRNA levels were determined by qRT-PCR, revealing that the reduction of HOXB7 and ZBTB7A markedly attenuated STN1 mRNA levels to a much greater extent than SNA12 and PPARD (Fig. 4c, Supplementary Fig. 3d).
a Analysis of TCGA data revealed significantly elevated HOXB7 expression in tumor tissues from PDAC patients. The boxplot represents log2-transformed transcripts per million (TPM) of PDAC patients and normal controls from TCGA and GTEx. The center line marks the median and two hinges indicate the 25th and 75th percentile, with whiskers extending to 1.5 × interquartile range (IQR) from hinges. The method for differential analysis is one-way ANOVA. T, tumor tissues. N, normal tissues. n = 183 samples for tumor tissues and n = 167 samples for normal tissues. b Kaplan-Meier analysis of overall survival of PDAC patients in TCGA indicates an association between high HOXB7 expression and poorer outcomes. c HOXB7 knockdown in PANC-1 cells reduces STN1 mRNA (n = 3 biological replicates per group). d JASPAR-predicted HOXB7 binding motifs in the STN1 promoter. e HOXB7 overexpression activates STN1 promoter in luciferase assays (n = 3 biological replicates per group). PSA-N/PSA-N-HOXB7 plasmids were co-transfected with a Renilla luciferase plasmid and pGL3 plasmids carrying the STN1 promoter or STN1 promoter with a mutation at the HOXB7-binding motif. f ChIP-qPCR confirms HOXB7 binding to the STN1 promoter (n = 3 biological replicates per group). The ACTB loci was used as a negative control which HOXB7 cannot bind. g Correlation analysis demonstrated a relationship between the HOXB7 and STN1 mRNA levels in tumor tissues of PDAC patients, as measured by qRT-PCR (n = 55 samples). h Immunoblot analysis shows HOXB7 knockdown reduces STN1 protein. The assays were performed twice with similar results. i, j Rescue experiments were conducted in STN1-knockdown PANC-1 cells using HA-tagged HOXB7, and cell motility was assessed through Transwell migration and invasion assays. (i) Representative image. Scale bar, 50 μm. j Quantification of migrated cells performed using ImageJ in five random fields per well (n = 3 biological replicates). Statistical significance in (b) was determined using log rank test. Statistical significance in (c–j) was determined using two-tailed unpaired t tests in GraphPad Prism. ns, no statistically significant difference. Error bars represent ± SEM. Source data are provided as a Source Data file.
Increased expression of HOXB7 significantly enhanced the transcriptional activity of our STN1-luciferase reporter construct, and the enhancement could be abrogated by mutation of the HOXB7-binding motif in the STN1 promoter (from 5’-GTCACTTACC-3’ to 5’-AGAAGCCTTG-3’) (Fig. 4d and e). Results from ChIP-qPCR assays also confirmed the direct binding of HOXB7 to the STN1 promoter region (Fig. 4f). Moreover, analysis of cDNA from 55 human PDAC tissues in our cohort revealed co-expression of HOXB7 with STN1 (Fig. 4g). Reduction of HOXB7 could also cause a decrease in protein level of STN1 (Fig. 4h). Conversely, overexpression of ZBTB7A had no discernible effect on the transcriptional activity of the STN1-luciferase reporter construct (Supplementary Fig. 3e). HOXB7 is a member of the HOX gene family of transcription factor and has been implicated in the progression and unfavorable outcome of various tumor types by targeting downstream genes in oncogenic hierarchies20. The pro-metastatic effect of HOXB7 overexpression was markedly attenuated in STN1-knockdown cells relative to control cells, indicating that STN1 acts as one of the functional downstream effectors in HOXB7-mediated oncogenic signaling (Fig. 4i, j). Taken together, these findings showed that HOXB7 is an upstream transcription factor of STN1, suggesting its elevated expression in PDAC patients may contribute to the increased expression of STN1.
STN1 promotes PDAC metastasis by facilitating the process of EMT
We aimed to elucidate the mechanistic involvement of STN1 in facilitating the metastatic processes of PDAC. Utilizing RNA-seq profiling, we observed substantial downregulation of the EMT pathway in STN1-knockdown PANC-1 cells, accompanied by marked increases in epithelial marker genes and decreases in mesenchymal markers and EMT-associated transcription factors (EMT-TFs) (Fig. 5a). Given the complexity of metastasis—which involves malignant cell dissemination through the circulatory system—and the established role of EMT in this process, we investigated STN1 potential contribution to EMT. Subsequent gene set enrichment analysis (GSEA) corroborated these findings (Fig. 5b, and Supplementary Fig. 4). Validation was performed through immunostaining of EMT markers in STN1-knockdown PDAC cells and tumor tissues from KPC and KPCS mice. Specifically, STN1 knockdown in PANC-1, AsPC-1 and BxPC-3 cells led to an inhibition in EMT process, which could be reversed upon the reintroduction of STN1. In MIA PaCa-2 cells, decreased STN1 expression resulted in reduced vimentin expression, while E-cadherin levels exhibited no significant changes, potentially attributable to the high methylation of the E-cadherin promoter in MIA PaCa-2 cells21 (Fig. 5c). Furthermore, the reduction of STN1 in KPC mice also led to a decrease in the epithelial marker E-cadherin and an increase in the mesenchymal marker vimentin (Fig. 5d–f). Collectively, these data suggest that STN1 promotes PDAC metastasis by facilitating the progression of EMT.
a Heatmap illustrating a significant upregulation of epithelial marker genes and a concurrent downregulation of mesenchymal markers and epithelial-mesenchymal transition transcription factors (EMT-TFs) in STN1-knockdown PANC-1 cells (n = 3 biological replicates per group). b Gene Set Enrichment Analysis (GSEA) plot demonstrates the enrichment of the curated EMT gene set in the transcriptome of STN1-knockdown PANC-1 cells. c Immunoblot analysis of the epithelial marker E-cadherin and the mesenchymal marker vimentin in STN1-knockdown and -rescue stable cell lines. The assays were performed three times with similar results. d Immunoblot analysis of ZEB1 and EMT markers in the pancreases of KPC and KPCS mice. The assays were performed twice with similar results. e IHC staining of ZEB1, E-cadherin and vimentin in consecutive sections of PDAC tissues indicating reduced EMT in KPCS mice. Scale bar, left, 100 μm, right, 25 μm. f Quantification of IHC E-cadherin and vimentin signals performed using ImageJ (n = 5 random fields from one section per group). Source data are provided as a Source Data file.
STN1 recruits STAT3 to facilitate transcription of the EMT-TF ZEB1 in PDAC cells
To elucidate the mechanistic underpinnings of STN1 involvement in PDAC metastasis, we initially assessed the expression of EMT-TFs in STN1 knockdown cell lines as well as KPC and KPCS tumor tissues using qRT-PCR. The reduction of STN1 correlated with decreased mRNA and protein expression of the EMT-TF ZEB1 in STN1 knockdown cell lines and KPCS mouse tumor tissues (Fig. 6a–c, Fig. 5d, and Supplementary Fig. 5a), whereas the other EMT-TFs (SNAI1/SNAI2) exhibited no statistically significant correlation with STN1 levels. Consistent co-expression of ZEB1 and STN1 was further confirmed in qRT-PCR analysis of tumor tissues from our cohort of 55 PDAC patients (Fig. 6d). Previous studies reported that ZEB1 is a key factor of PDAC metastasis, and deletion of Zeb1 in KPC mice is sufficient to inhibit metastasis22. Consequently, we wondered whether STN1 might facilitate ZEB1 transcription. Indeed, dual luciferase reporter assays revealed that STN1 was able to increase the transcriptional activity of the ZEB1-luciferase reporter construct (Fig. 6e, f, and Supplementary Fig. 5b). We then stably overexpressed STN1 in MIA PaCa-2 and PANC-1 cells (Supplementary Fig. 5c) and performed ChIP-qPCR assays. These experiments showed that STN1 could substantially bind to the ZEB1 promoter region (Fig. 6g). We took advantage of the known ACTB loci that STN1 cannot bind11 as a negative control and found that STN1 was not enriched at this site. In addition, reintroducing ZEB1 led to an increase in the invasion and migration capacity of the STN1 knockdown MIA PaCa-2 cell line (Supplementary Fig. 5d–f). These findings raised the possibility that STN1 could promote PDAC metastasis by facilitating the transcription of ZEB1.
a, b qRT-PCR of EMT-TFs in STN1-knockdown/rescue MIA PaCa-2 cells and KPC/KPCS mouse pancreases (n = 3 biological replicates per group). c Immunoblot confirmed ZEB1 protein reduction in STN1-knockdown cells. The assays were performed twice with similar results. d Positive correlation between STN1 and ZEB1 mRNA levels in 55 PDAC patients. e, f Luciferase reporter assays show STN1-dependent ZEB1 promoter activation (n = 3 biological replicates per group). g ChIP-qPCR reveals STN1 binding to ZEB1 promoter with the ACTB locus serving as a negative control (n = 3 biological replicates per group). h Silver staining of proteins pulled down by HA-STN1 in PANC-1 cells. i Immunoprecipitation confirms STN1-STAT3 interaction. The assays were performed twice with similar results. j ChIP-qPCR assay shows STN1 enables STAT3 promoter recruitment (n = 3 biological replicates per group). The JASPAR-predicted genomic site in ZEB1 promoter that could not be bound by STAT3 was employed as a negative control in this experiment. k 293 T cells co-transfected with pCMV-Tag2B or pCMV-Tag2B-STN1, pGL3 carrying the ZEB1 promoter, Renilla luciferase plasmid, and treated with 100 μM STAT3 inhibitor S3I-201 or DMSO for 24 h. The activity of the ZEB1 promoter was detected by luciferase reporter assay (n = 3 biological replicates per group). l Scheme of the structure of STN1 and its deletion mutants (Created in BioRender. Dong, D. (2024) https://BioRender.com/h09c753). STN1-OBD, STN1 mutant with OB-fold deletion; STN1-wHTH1D, STN1 mutant with wHTH1 domain deletion. STN1-wHTH2D, STN1 mutant with wHTH2 domain deletion. STN1-OBM, STN1 mutant with point mutations in the OB domain exhibiting reduced telomere binding affinity. (m) Immunoprecipitation of HA-tagged wild-type full-length STN1 or deletion mutants targeting specific domains of STN1 with Flag-tagged STAT3. The assays were performed three times with similar results. n Immunoblot analysis of ZEB1 protein expression in STN1-knockdown and STN1-knockdown cell lines rescued with various mutants. The assays were performed twice with similar results. Statistical significance in (a–k) was determined using two-tailed unpaired t tests in GraphPad Prism. ns, no statistically significant difference. Error bars represent ± SEM. Source data are provided as a Source Data file.
Prior studies have demonstrated that STN1 is a single-strand DNA binding protein and is not able to bind double strand DNA7, implying that it is not possible to act as a transcription factor to regulate transcription. Based on this assumption, we hypothesized that STN1 might act by recruiting other transcription factors to the ZEB1 promoter. S-Tag-STN1 pull-down assays were carried out to identify potential transcription factors binding to STN1 (Fig. 6h, Supplementary Data 1). We were aimed to detect transcription factors which are reported to bind to ZEB1 promoter, promoting transcription of ZEB1 as well as process of EMT. In the list of proteins identified by subsequent mass spectrometry, we noticed STAT3, which is a member of the STAT family of transcription factors and well-known to induce EMT-mediated metastasis23, directly bind to the ZEB1 promoter in pancreatic cancer cells24.The interaction between STN1 and STAT3 was confirmed by co-immunoprecipitation assays (Fig. 6i, Supplementary Fig. 5g, h). ChIP-qPCR results from STN1 knockdown and rescued cells indicated that the decrease in STN1 correlated with a diminished binding affinity of STAT3 to the ZEB1 promoter (Fig. 6j). Data of Cleavage Under Tagets and Tagmentation (CUT& Tag) also showed that STN1 and STAT3 could bind to the same region in ZEB1 promoter (Supplementary Fig. 5i). Furthermore, the impact of STN1 on ZEB1-luciferase reporter activity was evaluated under STAT3 inhibition conditions using the STAT3 inhibitor S3I-201(Fig. 6k). These experiments demonstrated that inhibition of STAT3 eliminated the ability of STN1 to increase the ZEB1-luciferase reporter activity.
To investigate whether the functional role of STN1 in transcriptional processes relies on its binding affinity to single-strand DNA, we constructed a series of deletion mutants targeting specific domains of human STN1. These mutants included an OB-fold deletion mutant (STN1-OBD, disrupting the oligonucleotide/oligosaccharide-binding domain critical for ssDNA recognition and interaction), a wHTH1 domain deletion mutant (STN1-wHTH1D, ablating the first winged helix-turn-helix motif), a wHTH2 domain deletion mutant (STN1-wHTH2D, removing the second winged helix-turn-helix domain), an OB-fold and wHTH2 domain deletion mutant (STN1-wHTH1) (Fig. 6l). Co-immunoprecipitation assays revealed that deletion of OB fold and wHTH2 domain of STN1 had no effect on its interaction with STAT3, but deletion of wHTH1 domain was able to disrupt its binding to STAT3. Even a single wHTH1 domain could bind to STAT3(Fig. 6m). The IP results indicating that STN1 binds to STAT3 through its wHTH1 domain. Reintroduction of STN1-wHTH2D and STN1-wHTH1 could effectively rescue expression of ZEB1, while reintroduction of STN1-OBD as well as STN1-wHTH1D did not have such effect (Fig. 6n). Subsequent reintroduction of STN1-wHTH2D in STN1-knockdown cells significantly reversed the decline in migration and invasion abilities caused by reduced STN1, while reintroduction of STN1-OBD had no effect on promotion of metastasis both in vitro and in vivo (Supplementary Fig. 5j–l, Fig. 2f–i). Together, these observations suggest that STN1 participates in PDAC metastasis and the transcriptional activation of ZEB1 through both its protein-binding and ssDNA-binding affinity.
To better understand how STN1 participates in transcription through its ssDNA-binding affinity, we considered the dynamics of transcriptional processes. In the course of transcription, RNA-DNA hybridization gives rise to a specialized chromatin structure known as an R-loop. This structure induces replication fork collision and Pol-II blocking, posing risks to productive transcriptional elongation and contributing to pathogenesis25. Thus, we hypothesized that during the transcription process, STN1 may bind to the displaced ssDNA beside an R-loop, resolving G4 structures to inhibit R-loop formation, as well as recruiting STAT3 to activate ZEB1 transcription. The Electrophoretic Mobility Shift Assay (EMSA) showed that STN1 might bind to the displaced ssDNA beside the R-loop localized in the STAT3-binding region in ZEB1 promoter, and STAT3 could bind to this region via interaction with STN1 at R-loop (Supplementary Fig. 6a and b), raising a possibility that STN1 and STAT3 might participate ZEB1 transcription via this important architecture.
We also tested whether the well-known function of STN1 in telomere maintenance participate in the phenotype. Firstly, we examined telomere length in the pancreas of KPC and KPCS mice. There was no difference in telomere length between KPC and KPCS mice (Supplementary Fig. 6c). We then performed Telomere Dysfunction Induced Foci (TIF) analysis to determine the role reduction of STN1 in telomere protection. The result showed that decrease of STN1 leads to telomere dysfunction both in pancreatic cancer cell lines (Supplementary Fig. 6d and e) and pancreas of KPC mice (Supplementary Fig. 6f and g). We constructed a mutant (STN1-OBM) with three substituted residues (W89A, R139L, Y141A) that affect telomere ssDNA binding while still preserving other STN1 functions26 (Fig. 6l). In order to further confirm the role played by both mechanisms (ZEB1 transcription and telomere dysfunction) in the process of STN1-promoting metastasis, we utilized intrasplenic injection models to test the metastasis capability of scramble, shSTN1-1, sh1-rescue, sh1 + STN1-OBD, sh1 + STN1-OBM, sh1 + STN1-OBM+shZEB1 (Fig. 2f–h).We observed that rescue with STN1 could significantly rescue the metastasis ability of MIA PaCa-2 cells, while rescue with STN1-OBD had no effect on metastasis promotion of shSTN1 cells. Rescue with STN1-OBM (which has no function in telomere maintenance) could partially rescue the metastasis ability, indicating that both ssDNA-binding ability (in order to activate transcription of ZEB1) and inhibition of telomere dysfunction of STN1 contribute to its phenotype of metastasis promotion. The conclusion can be proved that when both functions are blocked, sh1 + STN1-OBM+shZEB1 cells have the same metastasis ability as the shSTN1 cells (Fig. 2f–i). In summary, these results provide evidence that STN1 recruits STAT3 to facilitate the transcription of the EMT-TF ZEB1 in PDAC cells and promotes metastasis through EMT activation as well as telomere maintenance.
STAT3 inhibitors exhibited enhanced efficacy in restraining the metastatic potential of PDAC cells with elevated STN1 expression
Observing reduced transcription activity of the ZEB1-luciferase reporter in STN1-overexpressed cells with S3I-201 treatment (Fig. 6k), we hypothesized that PDAC cells overexpressing STN1 might exhibit heightened sensitivity to STAT3 inhibitor treatment. To test this idea, we assessed the migration and invasion capacities of MIA PaCa-2 and PANC-1 cells stably overexpressing STN1 and in the presence or absence of the STAT3 inhibitors, S3I-201 or napabucasin. The latter drug has already been tested in a Phase III Clinical Trial (NCT0183621) for advanced colorectal cancer27. The results obtained demonstrate that the increased STN1 expression in both cell lines enhances the invasion and migration capabilities in cells treated with vehicle (DMSO) alone, consistent with our earlier conclusion that STN1 promotes PDAC metastasis. However, in the presence of STAT3 inhibitors, a reduction in the invasion and migration capacity of both STN1-overexpressing cell lines and the PCDH-Ctrl group compared to vehicle treated cells was observed (Fig. 7a–c). These results are in line with reports indicating that S3I-201 and napabucasin can inhibit tumor cell proliferation and metastasis. However, STN1-overexpressed cells exhibited a more substantial decrease in invasion and migration compared to the PCDH-Ctrl group under STAT3 inhibitor treatment (Fig. 7c, d). Furthermore, data from a 3D spheroid invasion assay indicated a more pronounced decrease in the invasion ability of STN1-overexpressing cells treated with STAT3 inhibitors (Fig. 7e, f). Treatment of STN1-knockdown pancreatic cells could also result in a bigger decline in ZEB1 expression and EMT process than that of control cells (Fig. 7g). These results suggest that STAT3 inhibitors exhibit enhanced efficiency in restraining the metastatic potential of PDAC cells with elevated STN1 expression, thus STAT3 inhibitors may represent a promising therapeutic avenue for targeting metastasis in PDAC patients with heightened STN1 expression who face an unfavorable prognosis.
a–d PDAC cells overexpressing STN1 or control cells were treated with STAT3 inhibitors (100 μM S3I-201 or 2 μM napabucasin), or DMSO for 12 h. Transwell migration and invasion assays were performed to assess the motility of the cells as indicated. a Representative images for the migration and invasion of the indicated cells treated with S3I-201 or DMSO. Scale bar, 50 μm. b Representative images for the migration and invasion of the indicated cells treated with napabucasin or DMSO. Scale bar, 50 μm. c Quantification of migrated cells in five random fields per well performed using ImageJ (n = 3 biological replicates per group). d Percentage decrease in cell counts after treatment with STAT3 inhibitors (n = 15 random fields from 3 biological replicates per group). Left, S3I-201. Right, napabucasin. e, f 3D multicellular tumor spheroid invasion assays of STN1 overexpressing and control MIA PaCa-2 cells treated with STAT3 inhibitors. e Representative images of invasion assays. Scale bar, 200 μm. f Quantification of the invading area using ImageJ (n = 10 spheroids per group). g Immunoblot analysis of ZEB1 protein expression in STN1-overexpression cells treated with STAT3 inhibitors. The assays were performed twice with similar results. Statistical significance in (c–f) was determined using two-tailed unpaired t tests in GraphPad Prism. ns, no statistically significant difference. Error bars represent ± SEM. Source data are provided as a Source Data file.
Discussion
Pancreatic cancer’s high mortality and resistance to most conventional treatment1 drive search for novel oncogenes. Our study employed bioinformatic tools to sort for potential oncogenes and identified STN1 as a potential upstream of EMT and regulator in PDAC progression. Furthermore, we found that STN1-overexpressing PDAC cells display heightened sensitivity to STAT3 inhibitors, which suggests a potential therapeutic intervention for PDAC patients with elevated STN1 expression and poor prognoses, offering a potential treatment option for a specific subset of PDAC patients.
We first utilized multiple bioinformatic tools to identify STN1 as a candidate regulator in pancreatic cancer progression, whose overexpression in PDAC tumor tissues and correlation with poor prognosis suggested its significance. While mutations in STN1 and CTC1 are linked to Coats plus syndrome and dyskeratosis congenita1,28,29, recent studies have demonstrated the relevance of CST complex subunits, including STN1, in human cancer30,31,32,33,34. Our study shed light on the clinical significance of STN1 in PDAC. Functional investigations revealed that STN1 promotes PDAC metastasis in a dose-dependent manner. We also identified HOXB7 as a potential upstream transcription factor for STN1, whose elevated expression in PDAC patients may contribute to the increased expression of STN1.
While the CST complex and its subunits are well documented for their roles in telomere maintenance and genome stability7,11,35,36,37,38,39,40, their involvement in transcriptional regulation remains less explored. Our experimental evidence demonstrated that STN1 promotes transcription of ZEB1 via recruiting STAT3, and the interaction between STN1 and STAT3 occurred through the wHTH1 motif of STN1. Notably, the reintroduction of the point mutant STN1-OBM, characterized by its inability to bind telomere ssDNA but retaining its affinity for genomic ssDNA, markedly restored the metastatic capabilities of STN1-knockdown PDAC cells, in contrast to the ineffectiveness of reintroducing the deletion mutant STN1-OBD. These observations further establishes that STN1 role in PDAC metastasis is distinct from its better-known function in telomere maintenance, emphasizing a reliance on its ssDNA-binding affinity. In the course of transcription, the RNA-DNA hybridization structure, known as an R-loop, is able to induce replication fork collision and Pol-II blocking, posing risks to productive transcriptional elongation and contributing to pathogenesis25. R-loop hotspot regions are typically mapped to areas exhibiting a GC skew, including promoter regions characterized by high G + C contents25, where STN1 displays a predilection for binding41. Furthermore, once initiated, R-loop elongation is facilitated by the G-quadruplex (G4) secondary structure on the non-template strand of DNA25, while STN1 has been reported to prevent G4 accumulation and resolve G4 structures42. We showed that STN1 could bind to the displaced ssDNA beside the R-loop in STAT3-binding region of ZEB1 promoter, and STAT3 could bind to the indicated R-loop via interaction with STN1. Thus, during the transcription process, STN1 may bind to the displaced ssDNA beside an R-loop, resolving G4 structures to inhibit R-loop formation, as well as recruiting STAT3 to activate ZEB1 transcription (Fig. 8). Thus, we have elucidated STN1 involvement in transcription through its dual capabilities of protein-binding and ssDNA-binding affinity, ultimately contributing to the induction of EMT in PDAC.
Created in BioRender. Dong, D. (https://BioRender.com/q05s064.). HOXB7 upregulates STN1 in PDAC tumors. The elevated STN1 then binds to structurally displaced single-stranded DNA adjacent to R-loops in the EMT-TF ZEB1 promoter region. This facilitates STAT3 recruitment, which promotes transcription of ZEB1, thereby activating epithelial-mesenchymal transition and metastatic progression.
STAT3 is widely regarded as a promising therapeutic target for STAT3-related diseases, including cancer, autoimmune as well as inflammatory diseases43. Several small molecule inhibitors of STAT3 have been tested in clinical trials and some of them show encouraging results43. We found that STN1-overexpressing cells exhibited a more pronounced reduction in migration and invasion capacity than controls. Thus, we present STAT3 inhibitors, drugs that are still under testing in clinical trials, as a potential therapeutic strategy to directly target metastasis in PDAC patients with elevated STN1 who bearing severe metastasis and a poor prognosis.
Methods
Our research complies with all relevant ethical regulations, approved by Institutional Animal Care and Use committee (IACUC) of Peking University Health Science Center (LA2017142) and Ethics Committee of Peking University First Hospital (No. 2019-147).
Identification of potential oncogenes for PDAC via data mining
Differential genes between tumors and normal tissues in microarray datasets were determined using paired t test with P value < 0.05. All microarray datasets are deposited in the National Center for Biological Information Gene Expression Omnibus repository (accession codes: GSE28735, GSE16515, GSE15471). Differential genes between TCGA tumor samples and GTEx normal samples were determined using t test with P value < 1 × 10−10 and log2 fold change > 1. Differential genes between malignant cells and normal epithelial in scRNA-seq datasets were identified using the FindMarkers function in R package Seurat with P values < 1 × 10-10, log2 fold change > 0.2 and percent of expressed cells in either population > 20%. Survival analysis was performed via Cox regression with age, sex, and clinical stage as covariates of each cancer type. Kaplan-Meier curve was plotted to show the survival difference using R package survival and survminer.
Cell lines, antibodies, and reagents
All human cell lines (HEK293T, HPNE, MIA PaCa-2, PANC-1, BxPC-3 and AsPC-1) used in this study were obtained from the American Type Culture Collection (CRL-3216, CRL-4023, CRL-1420, CRL-1469, CRL-1687, CRL-1682) and subjected to STR locus analysis for authentication, as well as regular screening for mycoplasma contamination. BxPC-3 cells were cultured in RPMI 1640 (Gibco, C11875500BT), and other cells were maintained in DMEM (Gibco, C11995500BT), supplemented with 10% fetal bovine serum (FBS, PAN) at 37 °C in an atmosphere of 5 % CO2 in air. Primary antibodies used for immunoblotting, immunoprecipitations, ChIP and immunohistochemistry are listed in Supplementary Table 1.
Human tissue samples
The cDNA of PDAC and NATs was from 55 PDAC patients at the Peking University First Hospital from September 2018 to November 2023. The tissue microarray containing samples from 90 PDAC patients was purchased from Shanghai Outdo Biotech.
Plasmids and cloning strategies
The plasmids pCMV-tag-2b, pLKO.1-puro, pSA-N, pCDH-CMV and pGL3-Basic were purchased from Addgene. CDS of STN1, STAT3 and ZEB1 were inserted into these plasmids.
Mice
The BALB/c nude mice used in this study were obtained from Charles River Laboratory (Beijing, China). For the liver metastasis model, 6-week-old male BALB/c nude mice were anesthetized with tribromoethanol and 3 × 106 MIA PaCa-2 cells suspended in 150 μl PBS were injected into the inferior hemispleen (n = 6 biological replicates per group). To prevent tumor cell leakage and breeding, the spleen was ligated immediately after injection. The mice were euthanized after 12 weeks and necropsied for observation of visible metastasis lesions in the liver.
Pdx1-creER, KrasLSL_G12D/+; Tp53fl/+ (KPC) mice were a gift of Dr. Ming Cui from Peking Union Medical College and were kept on a C57BL/6 background. Stn1-/+ mice were generated using a CRISPR/Cas9 system. The Cas9 mRNA and sgRNA targeting exon 2 of mouse Stn1 were generated using a mMESSAGE mMACHINE T7 ULTRA Transcription Kit (AM1345; Thermo Fisher) and a MEGAshortscript T7 Transcription Kit (AM1354; Thermo Fisher), respectively. After injection into zygotes of C57BL/6 mice, the zygotes were transplantated into pseudopregnant ICR mice. The offspring were genotyped and the disruption of Stn1 detected by sequencing using the following PCR primers: 5’-CCTGAACCCCTGTGTTACCC-3’ and 5’- ATCTTTGAGGGTCAGGGTGC-3’. One founder mouse was obtained but Stn1-/- mice were embryonic lethal. Heterozygous (Stn1-/+) mice were back crossed with C57BL/6 mice for 5 generations before manipulating to avoid the off-target effects of the CRISPR/Cas9 system. KPC mice was crossed with Stn1-/+ mice to generate KPCS (Stn1-/+KPC) mice.
To induce Cre recombinase expression in KPC and KPCS mice, tamoxifen (T5648, Sigma-Aldrich) was dissolved in corn oil (20 mg/ml) and administered via intraperitoneal injection to 4-5-week-old mice at 100 mg/kg once daily for six consecutive days. Both male and female KPC/KPCS mice were used throughout the study, as differences in sex were not observed. In our animal protocol approved by Institutional Animal Care and Use committee (IACUC) of Peking University Health Science Center, the standard tumor endpoint is 1.2 centimeters, we affirm that the maximum tumor size and burden in our protocol is not surpassed.
Bulk RNA-seq
Total RNA was isolated from cultured cells using TRIzol (Invitrogen, 10296010). After quality check, samples were sent to library preparation using the NEB Next Ultra RNA Library Prep Kit for Illumina (NEB, USA, E7530L). High-throughput sequencing was performed using the Illumina HiSeq X Ten platform with 150 bp paired-end reads according to the manufacturer’s instructions. FastQC was used for quality control of the sequencing data. Reads were aligned to the human (GRCh38) reference genome using STAR44 (v2.7.2b) and assigned to genes using featureCounts45 (v2.0.1). Differentially expressed genes between experimental groups were identified using DESeq246 (v1.38.3).
Histology, histopathology
Tissues were fixed in 10% formalin, dehydrated, embedded in paraffin, and then sectioned (4 μm) and stained with Mayer’s hematoxylin and eosin solution G (HE). Images were captured using a Axio Vision 2.05 imaging system (Zeiss) and processed with Adobe Photoshop.
Immunohistochemistry
Tissue sections were deparaffinized and rehydrated and then 3% (v/v) hydrogen peroxide in methanol was used to block endogenous peroxidase activity. Antigen retrieval was performed by incubating slides in 1 mM EDTA buffer (pH 9.0) in a microwave oven. Sections were then incubated with 5% BSA for 1 h to prevent nonspecific staining followed by incubation with primary antibodies overnight. After incubation with secondary antibodies, target proteins were detected by diaminobenzidine (DAB) staining and then counterstained with hematoxylin. Images were acquired using a Nikon ECLIPSE Ci microscope.
The IHC results were analyzed using ImageJ. Briefly, the colors were separated using the “Color Deconvolution 2” plugin (accessible via Image - Color - Color Deconvolution 2), with DAB selected as the stain. The resulting DAB-positive image was converted to an 8-bit grayscale format. The threshold was then adjusted to isolate the positive regions, and the positive area was quantified by selecting Analyze - Measure.
Luciferase reporter assays
HEK293T cells were transfected with combination of indicated protein expression and various pGL3 firefly luciferase reporter plasmids as well as Renilla luciferase plasmid. Cells were harvested 48 h after transfection and lysed with Passive Lysis Buffer (Promega). Luciferase activity was measured using the Dual Luciferase Assay System (Promega).
The STN1 and ZEB1 promoter were chosen by 1000 bp before transcription start site (TSS) and 500 bp after TSS (about 1500 bp) and generated by PCR. The regions were listed in Supplementary Table 4.
RNA isolation and quantitative RT-PCR
Total RNA was isolated using TRIzol (Invitrogen, 10296010) according to manufacturer’s protocol and reversed transcribed into cDNA using a commercial kit (TransGen Biotech, AE341-02). Quantitative RT-PCR was performed with TransStart® Green qPCR SuperMix (TransGen Biotech, AQ101). The mRNA expression levels of target genes were normalized to those of GAPDH or Gapdh mRNA. The primers used for qPCR are listed in Supplementary Table 2.
S-Tag pull-down assays and mass spectrometry
STN1-overexpressing PANC-1 cells were harvested and lysed in HEPS-KOH IP buffer [50 mM Tris-HCl (pH 7.4), 200 mM NaCl, 10% (v/v) glycerol, 0.5% (v/v) NP40, 1 mM EDTA] freshly supplemented with protease inhibitors. Equal amounts of protein were incubated with S-protein agarose (Novagen, 69704-4) for 4 h at 4 °C. Precipitates were washed four times in HEPS-KOH wash buffer [50 mM Tris-HCl (pH 7.4), 100 mM NaCl, 1% (v/v) glycerol, 0.1% (v/v) NP40, 1 mM EDTA]. Aliquots of the samples were incubated at 100 °C for 15 min and then loaded onto NuPAGE 4 ~ 12% gradient gels (Invitrogen, NP0321BOX) and visualized by silver staining (Pierce Silver Stain Kit, 24612).
The remaining samples (n = 1 for STN1 overexpressing and control group) were analyzed by mass spectrum analysis to identify the potential interacting proteins. Proteins were digested by endoproteinase Trypsin (Promega) following the manufacturer’s instructions. Peptides were separated by online reversed-phase nanoscale capillary liquid chromatography (Easy-nLC 1,000, Thermo Scientific). The data dependent mass spectra were acquired with the LTQ-Orbitrap Elite mass spectrometer (Thermo Scientific) equipped with a nanoelectrospray ion source (Thermo Scientific). Raw files were searched by Proteome Discoverer (Version 1.4.1.14, Thermo Scientific) against the UniProt Human database. Search parameters were set as follows: enzyme with Trypsin; up to two missed cleavages; carbamidomethyl cysteine as fixed modification; methionine oxidation as variable modifications. The false discovery rates were set at 0.01.
Co-immunoprecipitation
Cells were harvested and lysed in HEPS-KOH IP buffer [50 mM Tris-HCl (pH 7.4), 200 mM NaCl, 10% (v/v) glycerol, 0.5% (v/v) NP40, 1 mM EDTA] with protease inhibitors. Cell lysates (1000 μg) were incubated with S-protein agarose (Novagen, 69704-4) for 4 h and precipitates washed for 4 times in Heps-KOH Wash buffer [50 mM Tris-HCl (pH 7.4), 100 mM NaCl, 1% (v/v) glycerol, 0.1% (v/v) NP40, 1 mM EDTA] and subjected to Western blot to evaluate protein interaction.
Transwell migration and invasion assays
Transwell migration assays were performed in a 24-well Boyden chamber (6.5 mm diameter, 8.0 μm) (Corning, 3422). 2.5 × 105 MIA PaCa-2 or PANC-1 cells in 200 μl serum-free DMEM were seeded into the upper chamber, with the lower chamber filled with 600 μl DMEM/30% (v/v) FBS (PAN). After 36 h, the upper chambers were gently removed and the migrated cells on the lower side of the membrane were fixed with methanol for 30 min and then stained with 0.1% crystal violet (Solarbio Life Sciences, G1063). The cells remaining on the upper side of the membrane were gently rubbed with cotton swabs. Migrated cells were calculated by ImageJ in five random fields per well. For in vitro invasion assays, the chambers were pre-coated with Matrigel matrix (Corning, 356234), diluted 1:20 in serum-free DMEM.
3D multicellular tumor spheroid invasion assays
Cell suspensions (1.5 × 104 cells/well in 250 μl) were plated in 96-well ULA round bottom dishes (Corning, 7007) and incubated under standard culture conditions for 96 h to generated multicellular tumor spheroids (MTSs). The MTSs were transferred to 96-well dishes coated with hydrogel matrices constructed by 1% type I collagen. The 96-well dishes were incubated at 37 °C for 20 min to make the hydrogel matrices polymerizing and overlaid with 200 μl standard culture medium. The medium was exchanged every other day. An inverted microscope was used to observe the invasion of the MTSs in hydrogel matrices for 4 days and the invading area was quantified using ImageJ.
Chromatin immunoprecipitation assays (ChIP) -qPCR
MIA PaCa-2 or PANC-1 cells were crosslinked with 1% formaldehyde for 15 min and then the cross-linking was quenched by 0.125 M glycine for 5 min. Cells were collected in 15 mL PBS containing 0.1% BSA and centrifugated at 402 × g for 3 min and then lysed in Lysis Buffer [5 mM PIPES, 85 mM KCl, 0.5% NP40] with freshly added protease inhibitors. Lysates were centrifugated and resuspended in MNase buffer [10 mM Tris-HCl (pH 7.5), 0.25 M sucrose, 75 mM NaCl] with protease inhibitors, followed by sonicated and precleaned with fish sperm DNA and IgG. Sonicated chromatin was incubated with anti-HA (Sigma-Aldrich, H3663) or anti-FLAG (Sigma-Aldrich, F3165) antibody at 4 °C overnight. Protein A/G PLUS-Agarose (Santa Cruz, sc-2003) was added to the mixture and incubated for 4 h at 4 °C. The chromatin bound beads were washed sequentially with TSE1 [1% (v/v) Triton X-100, 0.1% SDS, 2 mM EDTA, 150 mM NaCl, 20 mM Tris-HCl (pH 8.1)] with protease inhibitors, TSE2 [1% (v/v) Triton X-100, 0.1% SDS, 2 mM EDTA, 500 mM NaCl, 20 mM Tris-HCl (pH 8.1)] with protease inhibitors, TSE3 [0.25 M LiCl, 1% (v/v) NP40, 1% deoxycholate, 1 mM EDTA, 10 mM Tris-HCl (pH 8.8)] and TE buffer [10 mM Tris-HCl (pH 8.8), 1 mM EDTA]. The chromatin was eluted in elution buffer (1% SDS, 0.1 M NaHCO3) and then reverse crosslinked at 65 °C overnight. ChIP DNA was treated with 8 μL protease K (20 mg/mL) at 56 °C for 1 h and was then purified by ZYMO ChIP DNA Clean & Concentrator™ (ZYMO, D5205). The purified ChIP DNA was quantified by qPCR. The primers used in the ChIP-qPCR assays are listed in Supplementary Table 2.
CUT & Tag sequencing
CUT & Tag libraries were generated using MIA PaCa-2 transfected with pCMV-Tag2B-STN1 or pCMV-Tag2B-STAT3 and carried out according to the protocol of TransNGS® CUT&Tag Library Prep Kit for Illumina® (TransGen Biotech, KP172-01). Briefly, 100,000 cells were digested into single cells, resuspended in PBS, binded by magnetic beads and incubated with anti-FLAG (Cell Signaling Technology, 14793S) or IgG (TransGen Biotech). Cells were then incubated with secondary antibody and pAG-Tn5, subsequently collected by magnetic beads. After DNA transposition in incubated at 37 °C for 1 h and incubated at 70 °C for 10 min to terminate the transposition reaction, DNA extraction was performed using CUT & Tag DNA Clean Beads. Library amplification was performed using PCR and the number of amplification cycles was 11. The libraries were subjected to next-generation sequencing on an Illumina NovaSeq platform.
Telomere length measurement
Genome DNA was generated from pancreas of KPC and KPCS mice. Telomere length was then measured by qPCR with oligonucleotide primers against the telomeric repeats and a reference single-copy genomic gene 36B4. The primers used for qPCR are listed in Supplementary Table 2.
Telomere Dysfunction Induced Foci (TIF) analysis
Tissue sections of pancreas from KPC and KPCS mice were deparaffinized and rehydrated. Antigen retrieval was performed by incubating slides in 1 mM EDTA buffer (pH 9.0) in a microwave oven. Sections were then incubated with 5% BSA for 1 h to prevent nonspecific staining followed by incubation with anti-53BP1 (Cell Signaling Technology, 4937S) overnight. After incubation with Alexa Fluor 488-conjugated goat anti-rabbit antibodies for 45 minutes at room temperature, slides were washed with PBST and fixed with 4% paraformaldehyde for 20 min. The slides were then incubated with a Cy3-labeled PNA telomere repeat probe (Panagene, (5’-CCCTAA-3’)3) applied in hybridization mix (per 100 ml, mix 70 ml of freshly deionized formamide, 15 ml PNA buffer (80 mM Tris-Cl pH 8, 33 mM KCl, 6.7 mM MgCl2, 0.0067% Triton X-100), 10 ml 25 mg/ml acetylated BSA, 5 ml 10 mg/ml PNA probe). Sections were denatured at 83 °C at room temperature for 4 min and incubated in the dark overnight. Slides were then stained with DAPI and mounted in antifade reagent. Confocal images were obtained with a Nikon A1 microscope.
For TIF at cells, scramble and shSTN1 MIA PaCa-2 were fixed with 4% paraformaldehyde and dehydrated. Cells were then hybridized with the PNA telomere probe and anti-53BP1 antibody. Images were acquired using a Nikon A1 microscope.
Electrophoretic mobility shift assay (EMSA)
The R-loop Region was chosen based on the binding site of STAT3 at ZEB1 promoter. The oligo sequences used in the experiment were listed in supplementary Table 2. Duplex DNA of the R-loop was prepared by annealing DNA-oligo-1 and DNA-oligo-2 via incubating at 95 °C for 5 min in a thermal cycler and slowly cooling to 21 °C at 0.1 °C/s. RNA of the R-loop was generated by annealing Oligo for transcription-1 and Oligo for transcription-2 and subsequent transcription by T7 High Yield Transcription Kit (Beyotime, R7017S) with biotin-labeled UTP. Duplex DNA and RNA were mixed in assembly buffer (90 mM Tris pH 7.5, 90 mM Borate, 10 mM MgCl2) and incubated at room temperature for 2 h.
The R-loop was incubated with purified Flag-STN1 and Flag-STAT3. R-loop generated by unlabeled RNA was used as a competitor. Reactions were subjected to PAGE using native gels and then transferred to nylon membranes. The biotin labeled RNA was detected using SA-HRP and ECL.
Statistical analysis
Prism GraphPad software v9.0.2 was used for statistical analyses. Each experiment was performed in triplicate, and the data are shown means ± SEM, unless otherwise stated. A two-tailed unpaired t-test was employed to determine the statistical significance of the both the in vitro and in vivo experiments. Chi-square Fisher’s exact tests were used to determine the statistical significance of the metastasis ratio, macro metastasis ratio and ratio of animals with metastasis in multiple organs, while a two-tailed unpaired t-test was used to determine values of other characters of KPC and KPCS mice. Associations of STN1 expression with the four potential upstream transcription factors in PDAC patients and normal groups were analyzed by the Pearson correlation test. Associations of STN1 expression levels with PDAC patient outcomes in TCGA were analyzed by Kaplan-Meier analysis with the log-rank test. P values of less than 0.05 were considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. Microarray data were obtained from Gene Expression Omnibus (GSE16515, GSE28735, GSE15471), The TCGA-GTEx Toil re-computed expression data were downloaded from the UCSC Xena website (https://xenabrowser.net/).The plots of correlation analysis of HOXB7, ZBTB7A, SNAI2, and PPARD mRNA levels with STN1 mRNA levels in PDAC patients from TCGA were generated using Gepia (http://gepia.cancer-pku.cn/). Clinical information of TCGA patients was obtained from PanCanAtlas (https://gdc.cancer.gov/about-data/publications/pancanatlas). ScRNA datasets were obtained from previous study by Peng et al. 17. The original photographs of immunoblots and acrylamide gels are provided in the source data file. The RNA-seq data reported in this work is available in the Gene Expression Omnibus under the accession number GSE254968. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD062518 and the potential proteins identified in the S-Tag pulldown assays were listed in Supplementary Data 1. The CUT&Tag sequencing data is available in the Gene Expression Omnibus under the accession number GSE282970. Primers used in this study are listed in Supplementary Table 2. The shRNAs used in this study are listed in Supplementary Table 3. The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Kleeff, J. et al. Pancreatic cancer. Nat. Rev. Dis. Prim. 2, 16022 (2016).
Wong, H. H. & Lemoine, N. R. Pancreatic cancer: molecular pathogenesis and new therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 6, 412–422 (2009).
Nguyen, D. X. & Massague, J. Genetic determinants of cancer metastasis. Nat. Rev. Genet 8, 341–352 (2007).
Steeg, P. S. Targeting metastasis. Nat. Rev. Cancer 16, 201–218 (2016).
Dongre, A. & Weinberg, R. A. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 20, 69–84 (2019).
Steeg, P. S. Tumor metastasis: mechanistic insights and clinical challenges. Nat. Med 12, 895–904 (2006).
Rice, C. & Skordalakes, E. Structure and function of the telomeric CST complex. Comput Struct. Biotechnol. J. 14, 161–167 (2016).
Lyu, X., Sang, P. B. & Chai, W. CST in maintaining genome stability: beyond telomeres. DNA Repair (Amst.) 102, 103104 (2021).
Lim, C. J. et al. The structure of human CST reveals a decameric assembly bound to telomeric DNA. Science 368, 1081–1085 (2020).
Bhattacharjee, A., Wang, Y., Diao, J. & Price, C. M. Dynamic DNA binding, junction recognition and G4 melting activity underlie the telomeric and genome-wide roles of human CST. Nucleic Acids Res. 45, 12311–12324 (2017).
Chastain, M. et al. Human CST Facilitates genome-wide RAD51 recruitment to GC-rich repetitive sequences in response to replication stress. Cell Rep. 16, 1300–1314 (2016).
Lyu, X. et al. Human CST complex protects stalled replication forks by directly blocking MRE11 degradation of nascent-strand DNA. EMBO J. 40, e103654 (2021).
Mirman, Z. et al. 53BP1-RIF1-shieldin counteracts DSB resection through CST- and Pol alpha-dependent fill-in. Nature 560, 112 (2018).
Barazas, M. et al. The CST complex mediates end protection at double-strand breaks and promotes PARP inhibitor sensitivity in BRCA1-deficient cells. Cell Rep. 23, 2107–2118 (2018).
Gu, P. et al. CTC1 deletion results in defective telomere replication, leading to catastrophic telomere loss and stem cell exhaustion. EMBO J. 31, 2309–2321 (2012).
Li, Z. et al. BZW1 facilitates glycolysis and promotes tumor growth in pancreatic ductal adenocarcinoma through potentiating eIF2alpha phosphorylation. Gastroenterology 162, 1256–1271 e1214 (2022).
Peng, J. et al. Single-cell RNA-seq highlights intra-tumoral heterogeneity and malignant progression in pancreatic ductal adenocarcinoma. Cell Res 29, 725–738 (2019).
Song, J. et al. Construction of a novel model based on cell-in-cell-related genes and validation of KRT7 as a biomarker for predicting survival and immune microenvironment in pancreatic cancer. BMC Cancer 22, 894 (2022).
Oria, V. O., Lopatta, P. & Schilling, O. The pleiotropic roles of ADAM9 in the biology of solid tumors. Cell Mol. Life Sci. 75, 2291–2301 (2018).
Errico, M. C., Jin, K., Sukumar, S. & Care, A. The widening sphere of influence of HOXB7 in solid tumors. Cancer Res 76, 2857–2862 (2016).
Winter, J. M. et al. Absence of E-cadherin expression distinguishes noncohesive from cohesive pancreatic cancer. Clin. Cancer Res 14, 412–418 (2008).
Krebs, A. M. et al. The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nat. Cell Biol. 19, 518–529 (2017).
Sadrkhanloo, M. et al. STAT3-EMT axis in tumors: modulation of cancer metastasis, stemness and therapy response. Pharm. Res 182, 106311 (2022).
Liu, M. et al. ZIP4 increases expression of transcription factor ZEB1 to promote integrin alpha3beta1 signaling and inhibit expression of the gemcitabine transporter ENT1 in pancreatic cancer cells. Gastroenterology 158, 679–692 e671 (2020).
Kim, A. & Wang, G. G. R-loop and its functions at the regulatory interfaces between transcription and (epi)genome. Biochim Biophys. Acta Gene Regul. Mech. 1864, 194750 (2021).
Bhattacharjee, A., Stewart, J., Chaiken, M. & Price, C. M. STN1 OB fold mutation alters DNA binding and affects selective aspects of CST function. PLoS Genet 12, e1006342 (2016).
Flebbe, H. et al. Targeting STAT3 Signaling facilitates responsiveness of pancreatic cancer cells to chemoradiotherapy. Cancers (Basel) 14 (2022).
Anderson, B. H. et al. Mutations in CTC1, encoding conserved telomere maintenance component 1, cause Coats plus. Nat. Genet 44, 338–342 (2012).
Simon, A. J. et al. Mutations in STN1 cause Coats plus syndrome and are associated with genomic and telomere defects. J. Exp. Med 213, 1429–1440 (2016).
Han, P. et al. Association of SNPs in the OBFC1 gene and laryngeal carcinoma in Chinese Han male population. Int J. Clin. Oncol. 24, 1042–1048 (2019).
Luo, Y. M. et al. CTC1 increases the radioresistance of human melanoma cells by inhibiting telomere shortening and apoptosis. Int. J. Mol. Med. 33, 1484–1490 (2014).
Gyorffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 123, 725–731 (2010).
Nguyen, D. D. et al. Deficiency in mammalian STN1 promotes colon cancer development via inhibiting DNA repair. Sci. Adv. 9, eadd8023 (2023).
Feng, X. et al. CTC1-STN1 terminates telomerase while STN1-TEN1 enables C-strand synthesis during telomere replication in colon cancer cells. Nat. Commun. 9, 2827 (2018).
Miyake, Y. et al. RPA-like Mammalian Ctc1-Stn1-Ten1 Complex Binds to Single-Stranded DNA and Protects Telomeres Independently of the Pot1 Pathway. Mol. Cell 36, 193–206 (2009).
Gu, P. et al. CTC1-STN1 coordinates G- and C-strand synthesis to regulate telomere length. Aging Cell 17, e12783 (2018).
Giraud-Panis, M. J., Teixeira, M. T., Geli, V. & Gilson, E. CST meets shelterin to keep telomeres in check. Mol. Cell 39, 665–676 (2010).
Wu, P., Takai, H. & de Lange, T. Telomeric 3’ overhangs derive from resection by Exo1 and Apollo and fill-in by POT1b-associated CST. Cell 150, 39–52 (2012).
Wang, F. et al. Human CST has independent functions during telomere duplex replication and C-strand fill-in. Cell Rep. 2, 1096–1103 (2012).
Stewart, J. A., Wang, Y., Ackerson, S. M. & Schuck, P. L. Emerging roles of CST in maintaining genome stability and human disease. Front. Biosci. -Landmark 23, 1564–1586 (2018).
Hom, R. A. & Wuttke, D. S. Human CST prefers G-rich but not necessarily telomeric sequences. Biochemistry 56, 4210–4218 (2017).
Zhang, M. et al. Mammalian CST averts replication failure by preventing G-quadruplex accumulation. Nucleic Acids Res. 47, 5243–5259 (2019).
Dong, J., Cheng, X. D., Zhang, W. D. & Qin, J. J. Recent update on development of small-molecule stat3 inhibitors for cancer therapy: from phosphorylation inhibition to protein degradation. J. Med Chem. 64, 8884–8915 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank H. Chang for help with plasmid construction and M. Cui for gift of KPC mice. The manuscript was polished by ChatGPT 3.5. Figure 6l and Fig. 8 were created in BioRender [https://BioRender.com] with Publication License. This study is supported by grants to Y. Yin and L. L including the National Key Research and Development Program of China (2021YFA1300601, Y. Yin), National Natural Science Foundation of China (82030081 to Y. Yin and 82273439 to H. L).
Author information
Authors and Affiliations
Contributions
Y.Y., H.L., D.D. and Z.Z. conceived the project and designed the major experiments. D.D. conducted the experiments. Z.Z. collected datasets and performed computational analysis. M. Zhu constructed the Stn1-/+ mice. Z. Hou, J. Gong and M. Chen helped with the animal experiments. X. Zhao conducted the mass spectrometry. A. Yan provided the cDNA of PDAC tumor tissues and adjacent normal tissues. D.D. and Z.Z. wrote the manuscript, Y. Yin and H.L. thoroughly revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Robert Eferl and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dong, D., Zhou, Z., Zhu, M. et al. STN1 facilitates metastasis by promoting transcription of EMT-activator ZEB1 in pancreatic cancer. Nat Commun 16, 7815 (2025). https://doi.org/10.1038/s41467-025-63083-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63083-0
This article is cited by
-
tRNA-derived fragment tRF-24 drives CELF1 phase separation to promote oncogenic splicing in esophageal squamous cell carcinoma
Journal of Experimental & Clinical Cancer Research (2025)