Abstract
Mutations in epigenetic regulators are common in bladder cancer, yet their impact on therapeutic responses remains unclear. Here, we identify that loss-of-function mutations in KDM6A, a histone demethylase altered in about 26% of advanced bladder cancers, are associated with poor survival after cisplatin chemotherapy, whereas they correlate with improved outcomes with anti-PD-1 therapy. Using CRISPR-Cas9-engineered murine and human bladder cancer models, we show that KDM6A deficiency increases formation of extrachromosomal circular DNA carrying chemoresistance loci, promoting cisplatin resistance. In parallel, KDM6A loss impairs DNA repair and rewires tumor metabolism, reducing glycolysis and lactate output. This metabolic shift diminishes histone lactylation in regulatory T cells, suppressing immunoregulatory genes and limiting expansion of PD-1hi regulatory T cells. Collectively, our findings establish KDM6A mutation as a key regulator of therapeutic responses, providing a foundation for its use in guiding precision therapy in advanced bladder cancer.
Similar content being viewed by others
Introduction
Bladder cancer is the sixth most commonly occurring cancer in the United States, with the five-year overall survival (OS) for advanced bladder cancer being less than 10%1. Cisplatin-based chemotherapy has been a mainstay of therapy for many decades; however the advent of immune checkpoint therapy (ICT) and targeted therapy has changed the current landscape of treatment for bladder cancer2,3. While the availability of different therapeutic agents such as chemotherapy, ICT and targeted therapy, either as single agents or in combination, has significantly improved outcomes, there remains a lack of biological insight into selecting and sequencing these therapies based on patient attributes to develop a personalized treatment algorithm.
The genes encoding key epigenetic regulators are frequently mutated in bladder cancer4,5. These epigenetic factors orchestrate gene expression, impacting multiple pathways governing cellular phenotype and function6,7. While the impact of mutations in epigenetic factors on initiation and progression of bladder tumorigenesis has been studied8,9,10, the mechanistic insight into the roles of these mutations in regulating response to therapeutic agents remains largely unexplored. Lysine Demethylase 6 A (KDM6A) is a commonly mutated gene in bladder cancer, and approximately 26% of patients with muscle-invasive bladder cancer harbor KDM6A mutations5. KDM6A catalyzes the demethylation of H3K27 di- and tri- methyl (H3K27me2/me3) marks which are repressive histone post-translational modifications (PTMs), thereby promoting transcriptional activation11,12.
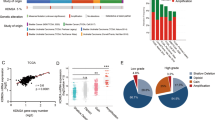
Here, we investigate how inactivating mutations in KDM6A, a frequently occurring mutation in advanced bladder cancer, regulate responses to therapeutic perturbations including chemotherapy and ICT in bladder cancer. Retrospective analyses of bladder cancer patient cohorts reveal that loss-of-function mutations in KDM6A, a histone demethylase frequently mutated in approximately 26% of advanced bladder cancer cases, are significantly associated with reduced overall survival (OS) following cisplatin-based chemotherapy. Extrachromosomal circular DNA (eccDNA) has recently emerged as a key mediator of chemoresistance across cancer types. Analysis of the TCGA-BLCA cohort13 reveals that tumors harboring KDM6A mutations contained a higher abundance of circular amplicons and upregulated the expression of POLQ14,15, a key enzyme in eccDNA biogenesis. Consistent with these observations, whole-genome sequencing (WGS) of human KDM6A-knockout bladder cancer cells demonstrate genome-wide copy number gains, particularly across chromosomes 2, 3, and 7, indicating increased regional amplifications upon KDM6A loss. Moreover, WGS identify multiple eccDNA amplicons encompassing oncogenes and chemoresistance-associated loci, including TP6316,17,18, CLDN419,20, GLI221,22,23,24, LUC7L325,26, ERCC427, and SHCBP128,29,30. Together, our data identify KDM6A loss as a key driver of eccDNA accumulation and oncogene amplification, establishing a mechanistic link between chromatin dysregulation and cisplatin resistance in bladder cancer, and highlighting KDM6A as a potential biomarker for predicting response to cisplatin-based chemotherapy.
To investigate whether KDM6A loss confers differential sensitivity to other therapeutic modalities, we analyze outcomes in patients treated with immune checkpoint blockade. In contrast to the poor response to cisplatin, patients harboring KDM6A mutations exhibit improved OS following anti-PD-1/anti-PD-L1 therapy, suggesting a context-dependent role for KDM6A in modulating therapeutic efficacy. Supporting this observation, KDM6A-mutant tumors display elevated tumor mutation burden (TMB). Mechanistic interrogation in both human and murine models reveal that KDM6A directly regulates the transcription of genes involved in mismatch repair (MMR) and double strand break repair (DSBR). Loss of KDM6A result in transcriptional repression of DNA repair programs, potentially augmenting neoantigen load and enhancing tumor immunogenicity, thereby contributing to increased responsiveness to immune checkpoint inhibitors.
In parallel, our work reveals that KDM6A is critical for tumor metabolic shift that ultimately modifies the tumor immune microenvironment in bladder cancer. Loss of KDM6A suppress glycolytic flux and decrease intratumoral lactate accumulation, thereby attenuating histone lactylation (H3K9la and H3K18la) in regulatory T cells (Tregs). This epigenetic modification results in downregulation of key immunosuppressive genes, including Foxp3, Tgfb, and Pdcd1, and impair PD-1 expression on Tregs. Consequently, the expansion of PD-1hi Tregs was limited following anti–PD-1 therapy, enhancing the effector CD8⁺ T cell-to-Treg ratio and improving anti-tumor immune responses in Kdm6a-deficient tumor-bearing mice. Building on the recognized role of lactate in Treg metabolism, our study links KDM6A-driven metabolic remodeling to histone lactylation–mediated suppression of Treg function within the tumor microenvironment.
Collectively, these findings establish KDM6A as a central epigenetic regulator integrating genomic stability, metabolic state, and immunologic context to govern therapeutic responsiveness in bladder cancer. By delineating distinct mechanisms through which KDM6A mutations mediate resistance to cisplatin-based chemotherapy and enhanced sensitivity to anti-PD-1/anti-PD-L1 therapy, this study offers a rational framework for biomarker-driven patient stratification and the development of precision therapeutic strategies in KDM6A-deficient bladder cancer.
Results
Inactivating KDM6A mutations are associated with reduced overall survival following cisplatin therapy with concurrent accumulations in extrachromosomal DNA formation
To understand how inactivating mutations in KDM6A impact responses to therapeutic perturbations in bladder cancer, we performed retrospective analyses of data from patients with resectable and advanced bladder cancer receiving cisplatin-based chemotherapy31,32. Our analysis showed that patients with KDM6A mutations (KDM6A-Mut) have reduced OS compared to those without the mutation (KDM6A-WT) (Fig. 1A). To delineate the role of KDM6A in regulating responses to cisplatin therapy, we used CRISPR-Cas9 to excise Kdm6a from the murine MB49 bladder cancer cell line and generated independent KDM6A-KO cell clones (sgKdm6a C1-C3) (Supplementary Fig. 1A). Consistent with the findings from the clinical cohorts, mice harboring sgKdm6a tumors showed resistance to cisplatin-based chemotherapy compared to mice harboring control (sgScramble) tumors (Fig. 1B, Supplementary Fig. 1C). To further investigate the differential impact of cisplatin on sgScramble versus sgKdm6a cell lines, we treated the cell lines with cisplatin in-vitro and noted significantly higher cisplatin-induced cytotoxicity in sgScramble cells compared to sgKdm6a cells (Fig. 1C, Supplementary Fig. 1D). Additionally, sgKdm6a cells demonstrated higher invasion, migration and spheroid forming potential in response to cisplatin, compared to sgScramble cells, indicating reduced sensitivity of Kdm6a-deleted murine bladder cancer cell line to cisplatin (Fig. 1D, F, Supplementary Fig. 1F, H, J). To validate the cisplatin-resistant phenotype of KDM6A-mutant murine bladder cancer cells, we extended our analysis to two human bladder cancer cell lines, RT4 and ScaBER. We performed CRISPR-Cas9 mediated knockout of KDM6A in the cell lines, with the resulting isogenic counterparts designated as K2 and B7, respectively, for subsequent analysis (Supplementary Fig. 1B). In line with our findings in murine cells, K2 and B7 cells also exhibited reduced cisplatin-induced cytotoxicity along with greater invasion, migration and spheroid formation than RT4 and ScaBER cells, respectively (Fig. 1C, E, G, Supplementary Fig. 1E, G, I, K). Importantly, we observed no change in cellular proliferation rate between the control and KDM6A knockout cells over the treatment period (Supplementary Fig. 1L).
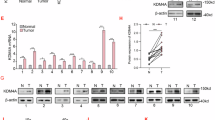
A. Kaplan-Meier plot demonstrating OS of advanced bladder cancer patients with (KDM6A-Mut) and without (KDM6A-WT) KDM6A mutation receiving platinum-based chemotherapy (n = 59 patients, KDM6A-Mut =17, KDM6A-WT = 42). Two-tailed Log-rank test was performed. B. Box-and-whisker plot illustrating weights of MB49 sgScramble and sgKdm6a Clone C1 tumors from mice treated with and without cisplatin-based chemotherapy (n = 10 mice per group). C. Box-and-whisker plots depicting percentage of Annexin+ cells in sgScramble and sgKdm6a C1(Top); and RT4 and K2(Bottom) cells, with or without 5 µM cisplatin treatment for 48 h (n = 5(Top) and n = 4(Bottom) biologically independent samples). D, E. Representative images(Left) and corresponding box-and-whisker plots(Right) showing number of invaded cells in Transwell assay for sgScamble and sgKdm6a C1(D) cells; and RT4 and K2(E) cells, treated with or without 5 µM cisplatin for 48 hours(n = 4 independent visual fields). Data are representative of two independent experiments. Scale bars included. F, G. Representative images(Left) and corresponding box-and-whisker plots(Right) comparing the diameter of sgScramble and sgKdm6a C1(F) spheroids; and RT4 and K2(G) spheroids treated with 5 µM cisplatin for 48 h (n = 7(F) and n = 8(G) biologically independent samples). Data is representative of two independent experiments. Scale bars included. H. The heatmap represent genome-wide copy-number changes in K2 and RT4 cells. The red and blue tones denote copy number amplifications and losses respectively. I. Circos plot(Left) displaying TP63 eccDNA amplicon in K2 cells, representing whole amplicon(outer red ring), start coordinate(black dot), and TP63 segment(inner blue region) within the amplicon. The scatter plots(Right) showing TP63 eccDNA loci copy number amplifications in RT4 and K2 cells. Gray dots represent probes across genomic region(x-axis), while y-axis represents the copy ratio(log2) at that genomic position for gray dots. The orange horizontal line is the segmentation line indicating true copy number state, and yellow vertical bars mark the indicated TP63 regions within the amplicon. For B-E, p-values were calculated by Two-way ANOVA test with Benjamini-Hochberg correction for multiple comparisons. For F and G, data were analyzed by two-tailed Student’s t-test. Center line marks the median, edges of the box represent the interquartile(25th-75th) percentile and whiskers represent minimum-maximum values. Source data are provided as a Source Data file.
Notably, extrachromosomal circular DNA (eccDNA) has recently emerged as a key player driving resistance to cisplatin-based chemotherapy33,34,35. Therefore, we compared eccDNA levels between sgScramble and sgKdm6a cells. Purification of eccDNA followed by rolling-circle amplification demonstrated a higher abundance of eccDNA in sgKdm6a cells (Supplementary Fig. 2A). Importantly, analysis of chromosomal structural variants in the TCGA dataset13 also demonstrated that patients harboring KDM6A mutation exhibited more circular amplicons(eccDNA) (Supplementary Fig. 2B), with consistent upregulation of POLQ expression14,15 (Supplementary Fig. 2C). Importantly, we noted that eccDNA derived from patients with KDM6A mutation selectively amplify SPP1, SHCBP1 and EGFR genes, strongly associated with cisplatin-resistance28,29,36,37,38 (Supplementary Fig. 2D). To further substantiate the role of KDM6A loss in eccDNA generation and cisplatin resistance, we performed Whole Genome Sequencing (WGS) of RT4 and KDM6A KO RT4 (K2) cell lines. We identified genome wide increase in copy numbers in various regions specifically in chromosomes 2, 3 and 7 in K2 cells (Fig. 1H). Analysis of the circular-amplicons using Circle-map showed multiple circular amplicons carrying critical genes implicated in promoting cisplatin resistance including TP63, CLDN4, GLI2, LUC7L3, ERCC4 and SHCBP1 (Fig. 1I, Supplementary Fig. 2E). Further scatter plots for the eccDNA gene loci in these chromosomes demonstrated increased copy numbers in K2 cells than RT4 cells (Fig. 1I, Supplementary Fig. 2F). Cumulatively, these findings indicate that the loss of KDM6A confers resistance to cisplatin-based chemotherapy through accumulation of eccDNA harboring oncogenes linked to cisplatin resistance and may serve as a potential biomarker of cisplatin-based chemotherapy resistance in advanced bladder cancer.
Loss of KDM6A impairs the MMR machinery and correlates with improved overall survival following immune checkpoint therapy
Given the association between KDM6A loss and cisplatin resistance, we next investigated whether KDM6A mutations impact the response to immune checkpoint therapy (ICT), a key therapeutic modality for patients with advanced bladder cancer. Analyses of the IMVigor210 cohort (N = 275) where patients with advanced bladder cancer received anti-PD-L1 therapy39 demonstrated that patients harboring the KDM6A mutation had significantly improved OS in response to anti-PD-L1 therapy compared to patients without KDM6A mutation (Fig. 2A). Analyses of another cohort40 of patients with advanced bladder cancer receiving ICT (MSK_2018, N = 148) also showed improved OS in KDM6A-Mut patients (Supplementary Fig. 3A). Further, consistent with the findings from the clinical cohorts, deletion of Kdm6a in murine bladder cancer cell line attenuated tumor growth following anti-PD-1 therapy in tumor-bearing mice (Fig. 2B, Supplementary Fig. 3B) confirming the association of inactivating KDM6A mutations with improved response to anti-PD-1 therapy.
A. Kaplan-Meier plot displaying the OS of KDM6A-Mut and KDM6A-WT patients treated with anti-PD-L1 therapy in IMVigor210 trial(n = 275 patients, KDM6A-Mut=49, KDM6A-WT = 226). Two-tailed Log-rank test was performed. B. Box-and-whisker plot illustrating weights from sgScramble and sgKdm6a C1 tumors in mice treated with and without Anti-PD-1 therapy(n = 8 mice per group). P-values were calculated by Two-way ANOVA test. C. Box-and-whisker plots representing TMB in KDM6A-WT and KDM6A-Mut patients with advanced bladder cancer, from two patient cohorts IMVigor210(n = 293, KDM6A-Mut=54, KDM6A-WT = 239) and TCGA BLCA(n = 136, KDM6A-Mut=40, KDM6A-WT = 96). Two-sided Wilcoxon test was performed. D. Somatic interaction plot depicting co-occurrence of genes in TCGA bladder cancer cohort(n = 412 patients). Two-tailed pair wise Fisher’s exact test was performed. E. ChIP-seq volcano plots demonstrating differential enrichment of KDM6A(Top) and H3K4me3(Bottom) marks in MMR pathway genes between sgScramble and sgKdm6a C1 cells with log2 ratio of fold change(log2FC) plotted against -log10 adjusted p-value(log10(FDR)). P-values were calculated using the Audic-Claverie Bayesian model with MAnorm. F. Genome browser plot demonstrating H3K4me3 peaks at Msh6 gene locus in sgScramble and sgKdm6a cells with highlighted regions indicating differential H3K4me3 enrichment. G. RNA-seq volcano plot representing differential expression of DNA MMR pathway genes between sgScramble and sgKdm6a C1 cell lines with log2 ratio of fold change(log2FC) plotted against -log10 adjusted p-value(log10(FDR)). P-values were calculated with two tailed exact test under a negative binomial distribution using EdgeR. H. Representative histograms(Left) and box-and-whisker plots(Right) showing Median Fluorescence Intensity(MFI) of MSH6 and MSH2 between sgScramble and sgKdm6a C1 cells(n = 3 biologically independent samples). One-tailed Student’s t-test was performed. I. Box-and-whisker plots showing relative expression of MSH6(Top) and MSH2(Bottom) in RT4 and K2 cells(n = 4 biologically independent samples). J. Representative electropherogram(Left) derived from Fragment Fluorescent Length Analysis(FFLA) of mBAT-64 microsatellite from sgScramble and sgKdm6a C1 tumor cells and box-and-whisker plot(Right) depicting the length of the modal peak in base pair in sgScramble and sgKdm6a C1 tumor cells(n = 7 mice per group). Data is representative of two independent experiments. For I and J, data were analyzed by two-tailed Student’s t-test. For all box-and-whisker plots, center line marks the median, edges of the box represent interquartile (25th−75th) percentile and whiskers represent minimum-maximum values. For B, D, E and G, p-values were adjusted with Benjamini–Hochberg(FDR) method. Source data are provided as a Source Data file.
Additionally, we noted higher tumor mutation burden (TMB) in patients with KDM6A mutations, (Fig. 2C). Since higher TMB is frequently driven by mutations in genes encoding enzymes involved in DNA MMR41, we investigated the frequency of MMR gene mutations in the TCGA bladder cancer patient cohort (N = 412). However, we did not note any cooccurrence of the KDM6A mutation with mutations in the genes associated with the MMR pathway in these patients (Fig. 2D). Notably, analyses of the HCRN dataset42 demonstrated decreased expression of several critical genes involved in the MMR pathway including MSH2 and MSH6 in patients harboring the KDM6A mutation indicating an attenuated MMR machinery in these patients (Supplementary Fig. 3C). This highlighted a distinct KDM6A-mediated pathway regulating genes associated with the MMR machinery and TMB.
Next, to investigate the mechanisms by which KDM6A regulates the expression of genes involved in the MMR pathway in bladder cancer, we performed chromatin immunoprecipitation sequencing (ChIP-seq) of sgScramble and sgKdm6a cells. We noted direct KDM6A binding to multiple MMR genes including Msh2 and Msh6, in sgScramble cells with lack of binding of KDM6A to these genes following the loss of Kdm6a in sgKdm6a cells (Fig. 2E, Supplementary Fig. 3D, F, G, I). Importantly, we also observed enrichment of H3K27me3 marks with concurrent attenuation of H3K4me3 enrichment in these genes in the absence of KDM6A (Fig. 2E, F, Supplementary Fig. 3E–I).
We used RNA-sequencing (RNA-seq) based gene expression studies to confirm the downregulation of the MMR genes in sgKdm6a cells. RNA-seq demonstrated a reduction in the expression of the MMR genes in sgKdm6a cells (Fig. 2G, Supplementary Fig. 3J), mirroring the findings from the patient cohorts, suggesting KDM6A- and H3K4me3-mediated regulation of these genes. To confirm the reduction in MMR enzyme expression at the protein level, we performed flow cytometry. This analysis revealed decreased expression of MSH2 and MSH6 in sgKdm6a cells compared to sgScramble cells (Fig. 2H). Consistent with these observations, both K2 and B7 cell lines exhibited an attenuation of MSH2 and MSH6 in both transcript and protein levels (Fig. 2I, Supplementary Fig. 4A-C). Further investigation using ChIP-qPCR in the RT4 and K2 cell lines demonstrated a reduced binding of KDM6A to specific MMR genes including MLH1 and MSH6 in K2 cells (Supplementary Fig. 4D), Additionally, we noted a reduced enrichment of the H3K4me3 in MLH1, MSH6 and MSH2 genes in absence of KDM6A (Supplementary Fig. 4E) which further solidified our data in murine sgKdm6a clones.
Disrupted MMR pathway is associated with microsatellite instability and an MSI-high phenotype43,44. Therefore, to determine the functional impact of the decreased expression of MMR genes in sgKdm6a tumor cells, we compared microsatellite instability (MSI) in sgScramble versus sgKdm6a MB49 tumors. We used fluorescent fragment length analysis (FFLA) to compare the microsatellite instability in sgScramble (N = 7) and sgKdm6a (N = 7) tumors. FFLA involves labeling DNA fragments with fluorescent dyes and analyzing their sizes and distribution via capillary electrophoresis. Importantly, analysis of electropherogram peaks revealed a significant left shift of 5 nucleotides in the modal (tallest) peak in sgKdm6a tumors compared to sgScramble tumors indicating contraction of the mBAT-64 microsatellite length and genetic instability (Fig. 2J, Supplementary Fig. 4F). Similarly, we noted a right shift of 5 nucleotides in the modal peak in K2 cells compared to RT4 cells indicating expansion of the NR21 microsatellite (Supplementary Fig. 4G, H). Cumulatively, these findings indicate that KDM6A modulates the epigenetic landscape by altering the enrichment of repressive H3K27me3 and active H3K4me3 marks at the promoter regions of DNA MMR genes thus regulating their expression and microsatellite instability in murine and human bladder cancer cells.
The MMR-deficient, MSI-h phenotype has been associated with improved responses to ICT45,46 and reduced sensitivity to platinum-based chemotherapy47,48,49,50. Therefore, attenuated MMR activity following the loss of KDM6A could account for improved response to ICT while decreasing sensitivity to cisplatin.
Loss of KDM6A impairs the double-stranded break repair pathway
In addition to reduced expression of genes involved in the MMR pathway following the loss of KDM6A, we observed lower expression of EXO1, a gene involved in DSBR in bladder cancer patients harboring KDM6A mutations42 (Supplementary Fig. 5A). Importantly, we did not observe any cooccurrence of the KDM6A mutation with mutations in the DSBR genes in these patients suggesting an independent role of KDM6A in regulating the expression of DSBR genes (Supplementary Fig. 5B).
Next, we analyzed the murine ChIP-seq data to investigate the impact of KDM6A loss on genes involved in DSBR. Our results showed a reduction in KDM6A binding in multiple genes involved in DSBR pathways including Exo1 and Lig3 in sgKdm6a cells indicating KDM6A-mediated regulation of these genes (Fig. 3A, Supplementary Fig. 5C, D, F, G). We also observed an increase in H3K27me3 (Supplementary Fig. 5E) with concurrent decrease in H3K4me3 enrichment across these genes (Fig. 3A, B, Supplementary Fig. 5C, D, F, G). RNA-seq analysis also demonstrated a reduction in the expression of these DSBR genes in sgKdm6a cells, confirming the findings from the ChIP-seq data (Fig. 3C, Supplementary Fig. 5H). Further, K2 and B7 cells demonstrated significantly lower expression of several genes involved in DSBR, compared to RT4 and ScaBER cells respectively (Supplementary Fig. 5I, J). Additionally, ChIP qPCR analysis in RT4 and K2 showed reduced KDM6A binding in multiple DSBR related genes like BAZ1B, EXO1 and LIG3 (Supplementary Fig. 5K) with simultaneous reduction in H3K4me3 enrichment at these gene promoters (Supplementary Fig. 5L). We also noted no significant differences in cell cycle distribution and cell proliferation rate between the control and the KDM6A knockout cell lines, suggesting that the reduction in DSBR gene expression is not secondary to altered proliferation or cell cycle phase distribution (Supplementary Figs. 1L, 5M).
A. ChIP-seq Volcano plots demonstrating differential enrichment of KDM6A(Left) and H3K4me3(Right) marks in DSBR pathway genes between sgScramble and sgKdm6a C1 cells, with log2 ratio of fold change(log2FC) plotted against -log10 adjusted p-value(log10(FDR). P-values were calculated using the Audic-Claverie Bayesian model with MAnorm. B. Genome browser plot demonstrating H3K4me3 peaks at Exo1 gene locus in sgScramble and sgKdm6a cells with highlighted regions indicating differential H3K4me3 enrichment. C. RNA-seq volcano plot representing differential expression of DSBR pathway genes between sgScramble and sgKdm6a C1 cells, with log2 ratio of fold change (log2FC) plotted against -log10 adjusted p-value(log10(FDR)). P-values were calculated with two tailed exact test under a negative binomial distribution using EdgeR. D. Representative images of immunohistochemical (IHC) staining and box-and-whisker plot(Right) showing percentage of EXO1 expression in sgScramble and sgKdm6a C1 tumors(n = 5 tumors per group). Data is representative of two independent experiments. E. Representative microscopy images(Left) and box-and-whisker plot(Right) depicting number of gamma-H2AX foci per cell(green-FITC and blue-DAPI/nucleus) in sgScramble and sgKdm6a C1 cells treated with and without 100 µM Etoposide followed by 4 h of recovery(n = 72). Arrows point to gamma-H2AX foci. Data is representative of two independent experiments. F. Representative histogram (Top) and box-and-whisker plots indicating the MFI of gamma-H2AX(Left) and percentage(Right) of gamma-H2AX positive E-cadherin positive tumor cells in sgScramble and sgKdm6a C1 tumors (n = 3(Left) and n = 6(Right) biologically independent samples). G. Representative images(Left) displaying comet-like appearance of DNA(vista green) and box-and-whisker plots(Right) representing the Tail DNA %, Tail Moment and Olive Moment of the corresponding comets formed in sgScramble(n = 246) and sgKdm6a C1(n = 75) cells. Arrows point to comet head and tail. Data is indicative of two independent experiments. H. Representative microscopy images(Left) and box-and-whisker plot(Right) depicting number of gamma-H2AX foci per cell(green-FITC and blue-DAPI/nucleus) in RT4 and K2 cells treated with and without 100 µM Etoposide followed by 4 h of recovery(n = 60). I. Representative images(Left) displaying comet-like appearance of DNA(vista green) and box-and-whisker plots(Right) representing the Tail DNA %, Tail Moment and Olive Moment of the corresponding comets formed in RT4(n = 106) and K2(n = 131) cells. For D, F, G, I, data were analyzed by two-tailed Student’s t-test. For E and H, data were analyzed by Two-way ANOVA. Center line marks the median, edges of the box represent the interquartile (25th-75th) percentile and whiskers represent minimum-maximum values. For A, C, E and H, p-values were adjusted with Benjamini–Hochberg(FDR) method. Source data are provided as a Source Data file.
We also assessed EXO1 protein expression levels in sgScramble and sgKdm6a tumors through immunohistochemical (IHC) staining and imaging of murine tumor sections. This revealed a significant decrease in EXO1 protein expression in sgKdm6a tumors (Fig. 3D). To elucidate the functional implications of reduction in EXO1 expression, we conducted several studies to compare genomic integrity between sgScramble and sgKdm6a cells. We first assessed the levels of Gamma(phosphorylated)-H2AX (gamma-H2AX), between sgScramble and sgKdm6a cells. Gamma-H2AX is a surrogate marker for DNA damage and forms foci at sites of DNA double-strand breaks51. The phosphorylation of H2AX at these sites is an early and sensitive indicator of DNA damage response. Confocal microscopy and flow cytometry studies revealed a significantly higher abundance of gamma-H2AX foci in both untreated and Etoposide (DNA damage inducer)-treated sgKdm6a cells compared to treated sgScramble cells (Fig. 3E, Supplementary Fig. 5N). These findings were also validated in sgKdm6a tumor samples (Fig. 3F) as well as in the human cell lines RT4 and K2 (Fig. 3H). Additionally, the COMET assay, which detects DNA damage in individual cells by measuring the migration of fragmented DNA during electrophoresis resulting in a comet-like appearance with a head (intact DNA) and tail (damaged DNA) was used to directly assess DNA breaks52. The results indicated significantly higher tail DNA percentage, olive tail moment, and tail moment in sgKdm6a cells and K2 cell with KDM6A loss, demonstrating increased DNA damage (Fig. 3G, I). Collectively, these data demonstrated that loss of KDM6A impairs DSBR, thereby compromising DNA integrity in bladder cancer cells.
KDM6A drives glycolysis and lactate production in bladder cancer cells
Previous research has shown that activation of DDR pathways can induce metabolic shifts, including enhanced glycolytic activity to support the Pentose-Phosphate-Pathway and generate nucleotide precursors53,54,55. In alignment with reduced expression of genes associated with DDR pathways in KDM6A-Mut patients, we noted attenuated expression of several genes involved in glycolysis and lactate production, including LDHA, PGAM1, and HK2 in KDM6A-Mut patients from IMvigor210 cohort along with LDHA and ENO1 from TCGA-BLCA cohort dataset5 (Fig. 4A, Supplementary Fig. 6A). Further, murine ChIP-seq data demonstrated reduced KDM6A binding, increased H3K27me3 enrichment and concurrent depletion of H3K4me3 enrichment at glycolysis associated genes in sgKdm6a cells (Fig. 4B, C, Supplementary Fig. 6B-F). Gene set enrichment analysis (GSEA) of H3K4me3 peaks differentially enriched between sgScramble and sgKdm6a cells also showed significant downregulation of the glycolysis pathway with concurrent upregulation of the oxidative phosphorylation (OXPHOS) pathway (Supplementary Fig. 6G). RNA-seq further corroborated these findings by showing decreased expression of these glycolysis genes in sgKDM6A cells compared to sgScramble cells (Fig. 4D, Supplementary Fig. 6H). Consistent decrease in expression of crucial glycolytic pathway genes HK2, LDHA and ENO2 was observed in human bladder cancer cells K2 and B7 with KDM6A excision (Fig. 4E, Supplementary Fig. 6J). ChIP-qPCR analysis in RT4 and K2 cells further demonstrated reduced KDM6A binding and decreased enrichment of H3K4me3 at the promoters of these genes in K2 cells (Supplementary Fig. 6K, L). Together, these findings indicate downregulation of the glycolysis pathway in tumor cells following Kdm6a deletion.
A. Box-and-whisker plots depicting the expression of the indicated glycolytic genes in KDM6A-Mut and KDM6A-WT patients, from the IMvigor210 cohort(n = 275 patients, KDM6A-Mut=49, KDM6A-WT = 226). Two-tailed Wilcoxon test was performed. B. ChIP-seq volcano plots demonstrating the differential enrichment of KDM6A(Left) and H3K4me3(Right) marks in glycolytic genes between sgScramble and sgKdm6a C1 cell lines, with log2 ratio of fold change (log2FC) plotted against -log10 adjusted p-value (log10(FDR)). P-values were calculated using the Audic-Claverie Bayesian model with MAnorm. C. Genome browser plot demonstrating H3K4me3 peaks at Hk2 gene locus in sgScramble and sgKdm6a cells with highlighted regions indicating differential H3K4me3 enrichment. D. RNA-seq volcano plot illustrating differentially expressed glycolytic genes between sgScramble and sgKdm6a C1 cell lines with log2 ratio of fold change (log2FC) plotted against -log10 adjusted p-value (log10(FDR)). P-values were calculated with two tailed exact test under a negative binomial distribution using EdgeR. E. Box-and-whisker plots showing relative expression of indicated genes in RT4 and K2 cells(n = 4 biologically independent samples). F. Box-and-whisker plot representing glycolytic (Top) and mitochondrial (Bottom) ATP production rate in sgScramble and sgKdm6a C1 cell lines (n = 9 biologically independent samples). G. Extracellular acidification rate (ECAR) between sgScramble and sgKdm6a C1 cell lines. Data represented as mean +/- Standard error of mean (SEM). (n = 11 biologically independent samples). H. Box-and-whisker plot representing glycolytic (Top) and mitochondrial (Bottom) ATP production rate in RT4 and K2 cell lines. I. ECAR between RT4 and K2 cells. Data represented as mean +/- SEM. (n = 5 biologically independent samples). J–K. Box-and-whisker plots indicating the Extracellular (EC) L-Lactate concentration in the supernatants of sgScramble and sgKdm6a C1 cell lines, measured following 24 h of culture (J) and in the supernatants of RT4 and K2 cells, measured following 48 h of culture (K). (n = 3 biologically independent samples). L. Box-and-whisker plot demonstrating L-lactate concentrations in the TIF normalized to corresponding tumor weights, obtained from sgScramble(n = 8) and sgKdm6a C1(n = 11) tumor bearing mice. For E, F, H, J–L, data were analyzed by two-tailed Student’s t test, center line marks the median, the edges of the box represent the interquartile (25th-75th) percentile and whiskers represent minimum-maximum values. For B and D, p-values were adjusted with Benjamini–Hochberg(FDR) method. Source data are provided as a Source Data file.
To delineate the functional impact of decreased expression of glycolysis genes in sgKdm6a bladder cancer cells, we compared the metabolic profiles of control and KDM6A Knockout cells using the Seahorse Extracellular Flux Analyzer-based ATP Rate Determination Assay. The data demonstrated reduced glycolytic ATP production rates in sgKdm6a cells as well as in K2 cells with KDM6A deletion compared to sgScramble and RT4 cells respectively with concurrent upregulation of mitochondrial OXPHOS (Fig. 4F-I). Given that downregulation of aerobic glycolysis attenuates lactate production, we measured lactate output in the culture supernatants of sgScramble, sgKdm6a, RT4, and K2 cells. Our results showed reduced lactate accumulation in the sgKdm6a and K2 cell lines (Fig. 4J, K). To verify whether these in-vitro findings are recapitulated in-vivo, we quantified lactate levels in the tumor interstitial fluid (TIF) of sgScramble and sgKdm6a tumors. We noted significantly lower accumulation of lactate in the sgKdm6a TIF, normalized to tumor weight (Fig. 4L, Supplementary Fig. 6I), demonstrating that the absence of KDM6A attenuates glycolysis and lactate production in bladder tumors. Importantly, the observed upregulation of the OXPHOS pathway in the cisplatin resistant sgKdm6a cells is in alignment with previous studies associating a switch from glycolysis to OXPHOS pathways with cisplatin resistance56,57. Cumulatively, these findings highlight the role of KDM6A as a major driver of metabolic pathways in bladder cancer and underscore the association between KDM6A deficiency and reduced lactate production by tumor cells.
Loss of KDM6A-mediated reduction in intratumoral lactate levels reduce Treg abundance and function
Lactate produced by tumor cells has been associated with immune suppression across various tumor types58. Consequently, we investigated how reduced lactate accumulation in sgKdm6a tumors affect the tumor immune microenvironment and response to anti-PD-1 therapy. Mass cytometry (CyTOF)-based immunophenotyping of sgScramble and sgKdm6a tumors followed by unsupervised clustering and t-SNE analyses of CD45+ immune cell subsets revealed the presence of distinct T cells, B cells, NK cells, and immune-suppressive myeloid cell clusters in the TME (Fig. 5A, Supplementary Fig. 7A). Further, this analysis revealed that sgScramble tumors, which have higher levels of intratumoral lactate, harbor a higher abundance of TGFβ + CD44 + LY6C+ immune-suppressive myeloid cells compared to sgKdm6a tumors (Fig. 5B). Additionally, sgScramble tumors showed a higher abundance of Tregs and increased expression of PD-1 in Tregs compared to sgKdm6a tumors (Fig. 5B, D). Notably, following anti-PD-1 therapy, the Treg population significantly expanded in sgScramble tumors, while this expansion was not observed in sgKdm6a tumors (Fig. 5C). This differential expansion of Tregs resulted in a higher T-effector to Treg ratio in sgKdm6a tumors, indicating a preferential pro-inflammatory shift in the tumor immune microenvironment following anti-PD-1 therapy thus improving responses to ICT (Fig. 5E, F).
A. t-SNE visualization of CD45+ immune cell subsets by CyTOF in tumor microenvironment(TME) of sgScramble and sgKdm6a C1 tumor bearing mice treated with and without anti-PD-1 (n = 4 mice per group). B. Box-and-whisker plot showing relative frequencies of intratumoral LY6C+CD44+TGFβ+ monocytes (Left) and CD3 + CD4 + FOXP3 + CD25 + LAG3 + CTLA4+ Tregs (Right) in untreated mice bearing sgScramble and sgKdm6a C1 tumors (n = 4). C. Box-and-whisker plot displaying relative frequencies of intratumoral CD3+CD4+FOXP3+ CD25+LAG3+CTLA4+ Tregs in sgScramble and sgKdm6a C1 tumor bearing mice treated with and without anti-PD-1(n = 4). D. Representative histogram(Left) and box-and-whisker plot(Right) representing MFI of PD-1 in intratumoral Tregs from mice bearing sgScramble and sgKdm6a C1 tumors(n = 4). E, F. Box-and-whisker plots depicting the ratio of intratumoral CD3+EOMES+TBET+PD1+GZMB+ T effector to CD3+CD4+FOXP3+CD25+LAG3+CTLA4+ Tregs (E) and intratumoral CD3+GZMB+TBET+LY6C+ Memory T cells to CD3+CD4+FOXP3+CD25+LAG3+CTLA4+ Tregs (F) in sgScramble and sgKdm6a C1 tumor bearing mice with and without anti-PD-1 treatment(n = 4). G. Representative histogram(Left) and box-and-whisker plot (Right) demonstrating MFI for PD-1 in in-vitro generated Tregs treated with and without 25 mM Na-La for 48 h(n = 5 biologically independent samples). H. Box-and-whisker plots indicating MFI of PD-1(Left) and TGF-β(Right) in in-vitro generated Tregs treated with supernatants from sgScramble or sgKdm6a C1 cell lines for 24 hours(n = 6 biologically independent samples). I. Box-and-whisker plot representing percentage of proliferated, anti-CD3/CD28 Dynabeads stimulated Tconv (CD4+CD25-) cells following co-culture with sgScramble or sgKdm6a C1 supernatant pre-treated Tregs (CD4+CD25+) at the indicated ratios for 4 days.(n = 4 biologically independent samples). For B data were analyzed by one-tailed and for D, G and H, data were analyzed by two-tailed Student’s t test, respectively. For C, E, F and I, p-values were calculated by Two-way ANOVA test with Benjamini-Hochberg correction for multiple comparisons. For all box-and-whisker plots, center line marks the median, edges of the box represent the interquartile (25th–75th) percentile and whiskers represent minimum-maximum values. Source data are provided as a Source Data file.
Next, to determine the impact of lactate specifically on Tregs, we performed a pHRodo-based lactic acid uptake assay which confirmed that Tregs can uptake exogenous lactic acid (La) (Supplementary Fig. 7B). This is in alignment with a previous study that demonstrated that Tregs can utilize exogenous lactate to maintain its functions59. We also noted that treatment with exogenous sodium lactate (Na-La) increased PD-1 expression in Tregs (Fig. 5G, Supplementary Fig. 7C, D). Importantly, when Tregs were treated with supernatants from sgScramble and sgKdm6a cells, those treated with sgKdm6a supernatants had lower PD-1 expression and fewer PD-1+ Tregs linked to the lower lactate levels in the sgKdm6a supernatants (Fig. 5H, Supplementary Fig. 7E, F). Together, these findings demonstrated that loss of KDM6A decreased glycolysis and lactate production in tumor cells which subsequently reduced PD-1 expression and expansion of PD-1hi intratumoral Tregs following anti-PD1 therapy thus improving the efficacy of anti-PD-1 therapy. In addition to reduction in PD-1 expression, TGFβ expression was also notably reduced in sgKdm6a supernatant treated Tregs (Fig. 5H). Production of TGFβ is one of the mechanisms through which Tregs exert their immunosuppression60. Therefore, to determine the impact of extracellular lactate levels on the immune-suppressive function of Tregs, we conducted in-vitro T cell suppression assays. We noted decreased proliferation of CD4 + CD25- T conventional cells (Tconv)when co-cultured with sgScramble supernatant-treated Tregs compared to sgKdm6a supernatant-treated Tregs indicating attenuated suppressive potential of the sgKdm6a Tregs (Fig. 5I, Supplementary Fig. 7G). Cumulatively, these findings demonstrate how tumor cell-specific KDM6A regulates the intratumoral metabolic milieu, which in turn alters the tumor immune microenvironment, including PD-1 expression in Tregs and their immune-suppressive function regulating efficacy of anti-PD-1 therapy in bladder cancer.
Loss of KDM6A in tumor cells impairs lactate mediated histone lactylation and function of regulatory T cells
We and others have demonstrated the role of lactate-derived histone lactylation in the regulation of phenotype and function of immune cell subsets including macrophages and CD8 T cells61,62. To determine lactate-mediated epigenetic changes in Tregs, we performed high-performance liquid chromatography (HPLC)–tandem mass spectrometry (MS/MS) analysis of Lys-C digested core histones isolated from murine Tregs. We noted that addition of 13C labelled Na-La increased the abundance of H3 histone lysine lactylation (Supplementary Fig. 8A). Additionally, we observed 13C incorporation in H3K18la confirming the role of exogenous lactate in driving histone lactylation in Tregs (Supplementary Fig. 8B, C). We have previously highlighted the importance of two specific histone H3 lactylation sites- H3K18la and H3K9la in regulating CD8 T cell function and phenotype62. To understand the relevance of H3K9la and H3K18la in Tregs, we performed flow-cytometry assays to determine the impact Na-La on H3K9la and H3K18la levels which demonstrated dose dependent increase in enrichment of H3K9la and H3K18la in Tregs following treatment with Na-La (Fig. 6A, Supplementary Fig. 8D-F). Further, to determine the impact of H3K9la and H3K18la on chromatin states in Tregs, we performed ChIP-seq studies followed by ChromHMM analysis. This analysis revealed that chromatin regions marked by H3K9la and H3K18la are also enriched for other transcription initiation marks including H3K4me3, H3K9ac and H3K27ac in Tregs (Supplementary Fig. 8G). Further, genomic regions enriched with H3K9la and H3K18la marks were identified as transcription start sites (TSS), TSS proximal regions and CpG islands (Supplementary Fig. 8G). Importantly, treatment with Na-La resulted in the enrichment of H3K9la and H3K18la marks at the TSS indicating a role of exogenous lactate-derived H3K9la and H3K18la in activating gene transcription (Fig. 6B). However, treatment with Na-La did not induce H3K4me3 or H3K27ac enrichment in the promoter regions, indicating selective regulation of H3K9la and H3K18la by exogenous lactate (Fig. 6B). Next, we annotated the genes showing differential enrichment of H3K9la and H3K18la following treatment with exogenous lactate (Fig. 6C). We noted H3K9la and H3K18la enrichment in multiple genes implicated in the regulation of Treg identity and function including Tgfb, Il10, Helios, Pdcd1, Lag3, Havcr2 and Irf4 in Na-La treated Tregs (Fig. 6C, Supplementary Fig. 8H). ChIP-qPCR and qPCR studies on Na-La treated Tregs also demonstrated higher H3K9la and H3K18la enrichment with concurrent increase in the expression of genes regulating Treg identity and function, including Foxp3, Pdcd1 and Tgfβ1, corroborating a role of exogenous lactate-derived H3K9la and H3K18la in regulating gene expression in Tregs (Fig. 6D, Supplementary Fig. 8I).
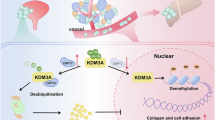
A. Box-and-whisker plots depicting the MFI of H3K9la(Top) and H3K18la (Bottom) in in-vitro generated Tregs treated with and without 25 mM Na-La for 48 h. (n = 3 biologically independent samples). B. Heatmaps demonstrating the genomic occupancy of the indicated hPTMs at the gene promoter regions (TSS ± 5 kb) in in-vitro generated Tregs treated with and without 25 mM Na-La for 48 h. C. ChIP-seq volcano plots illustrating the differential enrichment of H3K9la(Left) and H3K18la(Right) marks in the indicated Treg associated genes between in-vitro generated untreated Tregs and Tregs treated with 25 mM Na-La for 48 h, with log2 ratio of fold change (log2FC) plotted against -log10 adjusted p-value (log10(FDR). P-values were calculated using the Audic-Claverie Bayesian model with MAnorm. D. Box-and-whisker plots representing the relative expression of indicated Treg associated genes in in-vitro generated Tregs treated with and without 25 mM Na-La for 48 hours(n = 4 biologically independent samples). E. Box-and-whisker plots showing the MFI of H3K9la(Left) and H3K18la(Right) in intratumoral Tregs from sgScramble and sgKdm6a C1 tumor bearing mice(n = 10 mice per group). F. Box-and-whisker plots indicating the MFI of H3K9la(Left) and H3K18la (Right) in in-vitro generated Tregs treated with and without culture supernatants from sgScramble and sgKdm6a C1 cell lines for 24 hours(n = 3 biologically independent replictes). G. Box-and-whisker plots depicting the relative expression of indicated genes in in-vitro generated Tregs treated with sgScramble and sgKdm6a C1 culture supernatants for 24 h (n = 4 biologically independent samples). For A, D, E and G, data were analyzed by two tailed Student’s t-test. For F, p-values were calculated by one-way ANOVA test with Benjamini-Hochberg correction for multiple comparisons. Center line marks the median, edges of the box represent the interquartile (25th-75th) percentile and whiskers represent minimum-maximum values. H. Graphical summary of the findings presented in this study depicting the role of KDM6A as a critical epigenetic regulator driving genomic stability and metabolic reprogramming responsible for differential responses to chemotherapy and ICT in bladder cancer. Created in BioRender. Raychaudhuri, D. (2025) https://BioRender.com/sypcqql. Source data are provided as a Source Data file.
Next, considering the low intratumoral lactate content in sgKdm6a tumors (Supplementary Fig. 9A), we interrogated the levels of histone lactylation in intratumoral Tregs and noted significantly lower H3K9la and H3K18la in Tregs derived from sgKdm6a tumors (Fig. 6E, Supplementary Fig. 9B). Further, to directly implicate differences in lactate production by sgScramble and sgKdm6a bladder cancer cells as a major regulator of H3K9la and H3K18la in Tregs, we cultured Tregs in the presence of culture supernatants derived from sgScramble and sgKdm6a cells. Flow cytometry demonstrated significantly lower enrichment of H3K9la and H3K18la in Tregs cultured with KDM6A-deficient supernatants compared to control supernatants, linked to their respective lactate content (Fig. 6F, Supplementary Fig. 9C, D). Importantly, ChIP-qPCR studies demonstrated attenuated H3K9la and H3K18la enrichment in Foxp3, Tgfβ1 and Pdcd1 in Tregs cultured with sgKdm6a cell supernatant (Supplementary Fig. 9E). Additionally, qPCR studies confirmed reduced expression of these genes, establishing a direct association between extracellular lactate content, H3K9la/H3K18la enrichment, and gene expression including that of Pdcd1 in Tregs (Fig. 6G). Overall, these findings identified a critical role of the exogenous lactate derived histone PTM-histone lactylation in regulating the expression of critical Treg specific genes, thus regulating their function and response to immunotherapy in sgKdm6a tumors.
Discussion
Cisplatin-based chemotherapy has been a backbone of bladder cancer treatment and was among the earliest approved therapies for the disease2,3. In recent years, therapeutic options for advanced bladder cancer have undergone significant changes, with multiple approvals of anti-PD-1/L-1 therapy, targeted therapies including enfortumab vedotin, sacituzumab govitecan, and combinations of ICT with targeted therapy or chemotherapy7,63. However, a major challenge has been identifying biological attributes to select patients for specific therapies and to develop personalized treatment algorithms. Inactivating mutations in KDM6A, a commonly mutated gene, is a major driver of bladder cancer initiation and progression. Nonsense, missense, frameshift-indel and splice-site mutations in the KDM6A gene which attenuate gene expression are frequent in patients with muscle invasive bladder cancer5,64. While KDM6A mutations correlate with poor prognosis in bladder cancer, their role in regulating therapeutic outcomes remains unknown. This study investigated the mechanistic underpinnings of how mutations in KDM6A impact sensitivity to different therapeutic perturbations. Retrospective analyses of various patient cohorts demonstrated that while mutation in KDM6A lowers OS with cisplatin-based chemotherapy, it correlates with improved OS with ICT. Based on these clinical correlations, we adopted a reverse translational strategy to garner mechanistic insights into how the loss of KDM6A impacts tumor cell intrinsic and extrinsic pathways governing response to cisplatin-based chemotherapy and ICT. Mechanistic studies using CRISPR-Cas9 mediated KDM6A deletion in murine and human bladder cancer cell lines demonstrated that KDM6A serves as an upstream regulator of key tumor-intrinsic pathways including eccDNA biogenesis and oncogene amplification, DDR pathways and metabolic pathways. Consequently, KDM6A inactivation leads to the accumulation of eccDNA driving cisplatin resistance in patients harboring KDM6A mutation. Additionally, the loss of KDM6A attenuates the DDR machinery and glycolysis pathway, which in turn shape the tumor immune microenvironment, improving the efficacy of ICT in bladder cancer (Fig. 6H).
Resistance to cisplatin-based chemotherapy has been associated with the accumulation of eccDNA in tumor cells33,34,35. Consistent with these findings, we showed that patients harboring KDM6A mutation demonstrated higher eccDNA amplicons. KDM6A deletion in human bladder cancer cell line was associated with genome wide copy number amplifications on chromosomes 2, 3, and 7. Specifically, multiple eccDNA circular amplicons carrying genes including TP6316,17,18, CLDN419,20, GLI221,22,23,24 were upregulated in the absence of KDM6A. These genes have established roles in accelerating tumor progression through mechanisms that include promoting invasiveness, metastasis, suppressing apoptosis, and mediating resistance to cisplatin. Importantly, our in vitro assays using KDM6A knockout murine and human cells also demonstrated increased invasiveness, migration, and cisplatin resistance highlighting the functional contribution of eccDNA accumulation following KDM6A loss.
Notably, previous studies have linked eccDNA amplification in tumor cells to resistance to anti-PD-1 therapy in esophageal squamous cell carcinoma65, attributing this suppression to the presence of immunomodulatory genes on eccDNA66,67. However, we did not observe the amplification of these immunomodulatory genes in eccDNA derived from patients harboring KDM6A mutation and human cell lines with KDM6A deletion. Importantly, recent studies also suggest that eccDNA may serve as a potential source of neoantigens68, warranting further investigation into its role in driving neoantigen-specific anti-tumor T cell responses and enhancing anti-PD-1 therapy efficacy. Further, our mechanistic studies using human and murine bladder cancer cell lines show that this enhanced response is driven by attenuated DDR pathways and metabolic alterations, resulting in a pro-inflammatory tumor immune microenvironment. Collectively, our findings suggest that in KDM6A-mutant bladder cancer, cisplatin resistance and improved response to anti-PD-1 therapy stem from the interplay of multiple independent pathways.
In this study, we also identified KDM6A as a critical regulator of the MMR pathway in bladder cancer. Importantly, we showed that KDM6A directly binds to and regulates the expression of multiple MMR genes. In the absence of KDM6A, the expression of these genes were significantly reduced, thereby disrupting the MMR pathway. Regulation of gene expression is governed by a dynamic interplay of different histone PTMs including H3K27me3 which represses gene transcription and H3K4me3 which creates a permissive chromatin state for transcription initiation69,70. KDM6A is known to demethylate H3K27me3 thereby initiating gene transcription. However, recent studies have also shown that knockdown of KDM6A in non-small cell lung cancer cell lines significantly reduces H3K4me3 enrichment, potentially through interactions with the histone lysine methyltransferase KMT2B71,72. Consistent with these findings, we observed that CRISPR mediated deletion of Kdm6a is associated with attenuated H3K4me3 enrichment in tumor cells. Importantly, this directionality is corroborated by orthogonal assays including ChIP qPCR and gene expression analyses in both murine and human models. Taken together, these results support a model in which KDM6A helps shape the epigenetic landscape of gene regulation, including MMR pathway genes, while acknowledging that the changes in H3K4me3 could potentially represent only one component of a broader regulatory mechanism.
We also delineated a critical role of KDM6A in regulating the DSBR machinery including homologous recombination (HR) and Non-homologous end joining (NHEJ) in bladder cancer cells. This is in alignment with previous findings in hematologic malignancies73. Tumors with defective DSBR machinery rely heavily on PARP-mediated repair for survival which makes them sensitive to PARP inhibition74,75. Therefore, future preclinical and clinical studies are warranted to delineate the efficacy of PARP inhibitors either as monotherapy or as combination therapy with ICT in patients with metastatic bladder cancer harboring KDM6A mutated tumors which are deficient in DSBR and MMR pathways. We observed that patients with advanced bladder cancer harboring KDM6A mutations exhibit decreased expression of MMR and DSBR genes. However, we did not find any correlation between KDM6A mutations and mutations in MMR and DSBR genes in these patients. These findings demonstrate that KDM6A mutations downregulate critical DNA repair pathways even in the absence of mutations in DSBR and MMR genes. Additionally, the defect in DSBR pathway due to KDM6A loss was not confounded by cell proliferation or cell cycle distribution implicating KDM6A as an independent regulator of the DSBR and MMR pathways in bladder cancer.
Among the genes downregulated upon KDM6A loss, EXO1 and LIG3 occupy critical nodes at the interface of the MMR and DSBR pathways. EXO1 governs the excision step of mismatch repair and contributes to end resection during homologous recombination76,77,78,79, while LIG3 mediates gap ligation and facilitates alternative end-joining under conditions of replicative stress80. Downregulation of these genes, together with other MMR-associated components, compromises both mismatch correction and elements of DNA double-strand break repair, cumulatively promoting genomic instability. However, the repair deficiency induced by KDM6A loss is mechanistically distinct from classical MSI-high or DSBR-defective tumors, as it arises from epigenetic transcriptional repression rather than direct mutational inactivation of canonical repair genes, defining a unique form of chromatin-driven genomic instability.
A prior study reported that KDM6A serves a critical function in promoting glycolysis and tumor progression81. Building on this concept, we established a direct link between the metabolic reprogramming induced by KDM6A loss and subsequent tumor immune modulation, specifically via attenuated glycolysis and a resulting reduction in glycolysis-derived lactate generation. Intratumoral Tregs utilize lactate as a metabolic fuel to support their immune-suppressive functions59. In alignment with this, our CyTOF- based interrogation of the tumor immune microenvironment demonstrated decreased accumulation of Tregs in sgKdm6a tumors which showed reduced intratumoral lactate accumulation. A previous study has reported that lactate plays a role in regulating Treg function through MOESIN lactylation and TGF-beta signaling82. Additionally, lactate-derived histone lactylation is a histone PTM which has been shown to regulate the phenotype and function of immune cell subsets including macrophages61. We have previously delineated the role of endogenous glycolysis-derived histone lactylation in regulating the phenotype and function of different CD8 T cell subsets62. In this study, we demonstrated that tumor-derived lactate regulates histone lactylation in Tregs. A series of experiments demonstrated the enrichment of H3K9la and H3K18la in key genes including Foxp3, Tgfb, and Pdcd1, involved in regulating Treg phenotype and function. Importantly, loss of Kdm6a-mediated reduction in intratumoral lactate resulted in attenuation of H3K9la and H3K18la in intratumoral Tregs with concurrent reduction in the expression of T-reg specific gene such as Foxp3 and Tgfβ.
Further, we noted reduced abundance of intratumoral PD-1hi T regs in mice bearing sgKDM6A tumors. A previous study indicated that lactate regulates PD-1 expression in Tregs83. Therefore, reduced abundance of intratumoral lactate in mice bearing sgKDM6A tumors possibly attenuated histone lactylation-mediated expression of PD-1 in Tregs. The expansion of PD-1hi Tregs has been linked to resistance to anti-PD-1 therapy. Failure to expand the PD-1hi T regs in mice bearing sgKDM6A tumors increased the ratio of cytotoxic CD8 T cells to Tregs with improved response to anti-PD-1 therapy highlighting the potential role of lactate mediated histone lactylation in regulating the response. Together, these findings uncovered a previously unrecognized role of histone lactylation in modulating Treg phenotype and function within the TME. Additionally, these findings delineated the role of KDM6A in regulating the intratumoral metabolic milieu thus altering the tumor immune epigenome and driving response to ICT in bladder cancer.
Overall, retrospective analysis of clinical trials and reverse translational studies using human and murine cell lines as well as syngeneic murine models provided mechanistic insight into distinct pathways by which the loss of KDM6A regulates therapeutic responses in bladder cancer (Fig. 6H). First, we demonstrated that the loss of KDM6A enhances eccDNA biogenesis and oncogene amplification driving cisplatin resistance. Second, KDM6A loss attenuates the MMR machinery leading to improved response to ICT. Third, the loss of KDM6A suppresses glycolysis and lactate production in tumor cells which attenuates histone lactylation in intratumoral Tregs. This downregulates their immune-suppressive functions, leading to a pro-inflammatory shift in the tumor immune microenvironment, driving an improved response to ICT. Together, our findings identify KDM6A as a central regulator that orchestrates diverse biological processes including eccDNA biogenesis, DNA repair, and metabolic reprogramming that collectively confer resistance to cisplatin while enhancing responsiveness to immune checkpoint therapy. Cumulatively, these results provide mechanistic insight into the utility of KDM6A mutation status for patient stratification and the development of personalized treatment algorithms in advanced bladder cancer.
Methods
Cell lines (CRISPR-Cas9 mediated deletion of Kdm6a)
For knockout murine cell line generation:
CRISPR-Cas9 technology was used to establish Kdm6a knockout (KO) lines in MB49 cells (MDACC Functional Genomics Core). Single guide RNA was subcloned into LentiCRISPR v2 vector (AddGene #52961). The following mKdm6a gRNA sequence: TAGCATTATCTGCATACCAG, TTCCTCATCACCGAAAGCGG and TTCGTAGCAGCGAACAGCCT were used. LentiCRISPR v2 gRNA was transfected into MB49 cells using the JetPrime transfection reagent (VWR, 89129-924). 2 days after transfection, 2ug/ml puromycin was used to remove non-transfected cells. Multiple single cell mKdm6a KO clones (C1-C3) were screened by western blots and genomic DNA sequencing and authenticated by ATCC mouse cell line STR testing.
For control murine cell line generation:
LentiCRISPR V2 vector only was co-transfected with pMD2.G and psPAX2-D64V into 293 T cells and the virus containing medium were collected 48 h after transfection and used for cell transduction.
Cells were transduced with lentivirus and replaced with medium containing 2ug/mL puromycin after 72 h of transduction. Cells were cultured with puromycin medium for 5 days and replaced medium every two days. Once reached confluent in 10 cm dish, cells were harvested for frozen stock.
Murine cell line maintenance: Cells were maintained in DMEM medium supplemented with 10% FBS at 37 °C and 5% CO2.
Human cell lines:
CRISPR-Cas9 mediated knockout in RT4 and ScaBER cell lines were performed following previously described protocol8. Briefly, KDM6A knockout sgRNAs (GGTATGCAGATAATGCTGAA, ACAGTTTACAGTCTGACTAC) were cloned into lentiCRISPR v2 nickase vector. Lentiviral plasmids were cotransfected with psPAX2 and pMD2.G into HEK293T cells to generate lentiviral particles and used for target cell transduction. Transduced cells bearing CRISPR vectors were cultured in a selective growth medium containing puromycin and single clones were screened for KDM6A expression by immunoblot. Clones with no detectable KDM6A expression were then selected for subsequent experiments. The RT4 and ScaBER cell lines were authenticated by STR testing.
Human cell line maintenance: RT4 and K2 were cultured in MEM media and SCaBER and B7 were cultured in DMEM media supplemented with 10% FBS at 37 °C and 5% CO2.
Mice
This study used age and sex matched 5–7-weeks old C57BL/6 mice purchased from National Cancer Institute (NCI) for experiments. Mice were housed and sustained in pathogen-free conditions, 20-25 °C, 30-70% humidity and on a 12-hour light/12-hour dark cycle at the Animal Resource Center, The University of Texas MD Anderson Cancer Center (MDACC) Institutional Animal Care and Use Committee approved all animal protocols (Protocol number 00002153-RN01). Sex was not considered in the study design. The study involves understanding the role of KDM6A in the regulation of response to therapeutic perturbations in bladder cancer. Therefore sex and gender based analyses were not performed.
Overall Survival (OS) studies
The OS was compared between bladder cancer patients with and without KDM6A mutations (KDM6A-Mut and KDM6A-WT) utilizing publicly available clinical and genomic datasets. The data for IMviogor210 study which evaluated atezolizumab in advanced bladder cancer (NCT02108652, NCT02951767) was accessed through IMvigor210CoreBiologies R package39. Additionally, the OS data for advanced bladder cancer patients treated with at least one dose ICT (atezolizumab, avelumab, durvalumab, ipilimumab, nivolumab, pembrolizumab, or tremelimumab) was retrieved from cBioPortal40. The clinical and genomic data for patients with resectable and advanced bladder cancer receiving cisplatin-based chemotherapy was retrieved from two independent studies, blca_msk_tcga_2020 and paired_bladder_2022 in cBioPortal31,32.
Survival curve for above mentioned patient data was estimated with Kaplan-Meier method using “survfit” function from “survival” R package. The survival curves were plotted using “ggsurvplot” function from “survminer” R package. The statistical analyses were conducted in R v4.1.2.
Identification of eccDNA in bladder cancer patients
We obtained previously reported amplicon intervals identified from WGS data of TCGA-BLCA cohort13. We quantified distribution of structural-class categories across patients stratified by KDM6A mutation status. Within each KDM6A stratum (WT: n = 85, Mut: n = 27), we counted samples per classification and converted counts to percentages of the stratum total. The resulting proportions for circular amplicons were visualized with barplot created using ggplot2. Amplicon intervals were annotated to proximal gene symbols using Ensemble via the biomaRT R package. The annotated amplicon intervals were manually inspected for presence of cisplatin resistant genes reported in literature. The representative figures for eccDNA amplicons in Supplementary Fig. 2D were prepared using Biorender and (Integrative Genomics Viewer) IGV84,85.
Tumor mutation burden and somatic interactions
Tumor mutation burden for patients with advanced bladder cancer was retrieved from IMvigor210 study and The Cancer Genome Atlas (Stage IV) (TCGA-BLCA, n = 136)5,39. The mutational burden was compared between patients with and without KDM6A mutations. Somatic variants summary for all bladder cancer patients in the TCGA-BLCA (n = 412) were downloaded in Mutation Annotation Format (maf) using TCGAbiolinks R package. The co-occurrence or mutual exclusivity of KDM6A mutations with mutations in MMR and DSBR-related genes was computed using the “somaticInteractions” function in maftools v2.10.05. Significance of co-occurrence and mutual exclusivity was estimated with two-tailed Pair wise Fisher’s exact test and derived P values were adjusted with Benjamini–Hochberg method (-log10(pAdj)). The somatic interactions plots were prepared in ggplot2. The statistical analyses were conducted using R v4.1.2 and GraphPad Prism 10.0.
mRNA expression levels for patient cohorts
The RNA-sequencing expression levels for genes involved in MMR and DSBR pathways were obtained from publicly available transcriptomic and genomic data at cBioPortal for advanced bladder cancer patients treated with ICT(HCRN)42. The mRNA expression levels of genes involved in glycolysis pathways were retrieved from IMvigor21039. Additionally, mRNA expression levels for genes were retrieved for TCGA-BLCA cohort were obtained cBioPortal. The mRNA expression data for TCGA-BLCA and HCRN cohorts were obtained prior to June 2024. The expression levels of gene involved in MMR, DSBR and glycolytic pathways were compared between KDM6A-Mut and KDM6A-WT. The statistical analyses were conducted using R v4.1.2 and GraphPad Prism 10.0.
Western Blot
KDM6A expression was accessed in all the murine and human cell lines. Briefly, cells were lysed using freshly prepared lysis buffer (2%SDS, 10 mM Tris pH 7.5, supplemented with 1X protease inhibitors) by frequent vortexing and incubated on ice for 20 min. This was followed by centrifugation at 17000 × g for 2 min using QIAshredder and the protein in the collected supernatant was quantified using the Pierce BCA Protein Assay kit according to the manufacturer’s protocol. Samples were mixed with 2X reducing Laemmli buffer and incubated at 95 °C for 10 min. 15 μg of samples were loaded into a PROTEAN TGX gel and underwent electrophoresis. After electrophoresis, proteins were transferred to an Immobilon membrane for 1 h. Then, the membrane with the transferred protein was blocked for 1 h in EveryBlot Blocking buffer, then incubated overnight at 4 °C in 1:2000 of anti-KDM6A primary antibody diluted in EveryBlot Blocking Buffer. The following morning, the membrane was washed 5 times in PBST before being incubated with 1:2000 of anti-Rabbit IgG HRP linked secondary antibody diluted in EveryBlot Blocking Buffer for 1 h. Lastly, the membrane was washed in PBST six times followed by chemiluminescent imaging in a BioRad ChemiDoc imaging instrument.
In-vivo tumor model and tumor processing
sgScramble and sgKdm6a C1 cells were collected from culture via trypsinization and then washed with PBS. C57BL/6 mice were injected subcutaneously in the right flank with 2 ×105 sgScramble or sgKdm6a C1 cells resuspended in 100 μl of PBS. The tumor volume was monitored on day 3, 4, 7 and 10 using digital calipers post-tumor inoculation. On Day 11 post tumor implantation, mice were sacrificed, and the tumors were isolated and weighed. The maximum tumor size for subcutaneous tumors permitted by the Institutional Animal Care and Use Committee (IACUC protocol no. 00002153-RN01) at The University of Texas MD Anderson Cancer Center (MDACC) is 15 mm. However, mice were sacrificed, and tumors were isolated on Day 11, prior to reaching this size limit.
Collected tumors were transferred to Eppendorf tubes containing FBS-free media supplemented with 0.66 mg/ml of Liberase and 20 mg/ml of DNAse. Next, tumors were chopped using scissors, and the sample tubes were transferred to a thermal shaker for tumor digestion at 37 °C at 1000 rpm for 30 min. Following tumor digestion, a single-cell suspension was obtained by mushing tumors through a 70 μm strainer. The single-cell suspension was subsequently washed in PBS at 200 × g for 5 min at 4 °C, the cell pellet was subjected to RBC lysis by resuspending in ACK Lysis Buffer and then washed in PBS before proceeding to downstream assays. For some experiments, the cell pellet was resuspended in freezing media comprising of 90% FBS and 10% DMSO before storage at −80 °C for downstream use.
Anti-PD-1 in-vivo treatment regimen
Tumor-bearing mice were injected intraperitoneally with 200 μg, 100 μg, and 100 μg of anti-PD1 diluted in 100 μl PBS on Days 3, 6 and 9 post-tumor implantations. The tumor volume was monitored on day 3, 4, 7 and 10 using digital calipers post-tumor inoculation and was calculated using the formula, Tumor Volume = (Length × Width2)/2. The tumors were isolated, weighed, and subjected to tumor processing on Day 11 post-tumor implantation as described previously. The maximum tumor size for subcutaneous tumors permitted by the Institutional Animal Care and Use Committee (IACUC protocol no. 00002153-RN01) at The University of Texas MD Anderson Cancer Center (MDACC) is 15 mm. However, mice were sacrificed, and tumors were isolated on Day 11, prior to reaching this size limit.
In-vivo cisplatin-based chemotherapy regimen
Tumor-bearing mice were injected intraperitoneally with 2.5 mg/kg of gemcitabine plus 6 mg/kg of cisplatin on Days 1, 4 and 7 post-tumor implantations. The tumor volume was monitored on day 3, 4, 7 and 10 using digital calipers post-tumor inoculation and was calculated using the formula, Tumor Volume = (Length × Width2)/2. Tumors were isolated, weighed and subjected to tumor processing on Day 11 post-tumor implantation as described previously.
Murine splenocyte isolation and in-vitro Treg generation
Spleens were harvested from C57BL/6 mice, mushed and single-cell suspension was obtained by passing through a 70 µm strainer. The resulting single-cell suspension was treated with ACK Lysis Buffer to lyse the RBCs. The cell counts were obtained using an automated cell counter before proceeding to Magnetic Assisted Cell Sorting (MACS) of naïve CD4 T cells.
Naïve CD4 T cells were magnetically isolated using the Murine Naïve CD4 T Cell Isolation Kit, as per manufacturer’s guidelines. Briefly, splenocytes were resuspended in MACS buffer (1X PBS supplemented with 0.5% BSA and 2 mM EDTA) on ice. Naïve CD4 Biotin Antibody Cocktail was added, mixed thoroughly, and incubated at 4 °C for 15 min. Next, Anti-Biotin Microbeads were added to the samples, mixed thoroughly, and incubated on ice for an additional 15 min. CD44 Microbeads were then added, followed by a final incubation for 20 min on ice. The samples were subsequently washed by centrifugation at 200 × g for 5 min at 4 °C in surplus MACS Buffer. The resulting cell pellets were resuspended in MACS buffer, passed through LS columns and the naïve CD44- CD4 T cells were collected in the flow-through.
Subsequently, to generate Tregs, naïve CD4 T cells (1-1.5 ×106 cells per well) were stimulated with 3 μg/ml of anti-CD3, 2 μg/ml of anti-CD28, 10 ng/ml of murine recombinant IL-2, and 10 ng/ml of recombinant mouse TGF-β in complete RPMI media and plated in 24-well plates. The cells were then incubated for 5 days at 37 °C with 5% CO2.
Genomic DNA isolation for Whole Genome Sequencing (WGS)
Genomic DNA was extracted from 2 ×106 RT4 and K2 cells using QIAamp DNA Micro Kit, according to the manufacturer’s instructions. For cell lysis, the cell pellet was resuspended in buffer ATL, proteinase K and buffer AL was added and pulse-vortexed for 15 s, followed by 10 min incubation at 56 °C in a thermo shaker. After incubation, 100% ethanol was added, pulse-vortexed for 15 s and incubated at room temperature for 3 min. The lysates were then transferred to QIAamp MinElute columns and centrifuged at 6000 × g for 1 min at room temperature, followed by washing with buffer AW1 and AW2. Then, the columns were centrifuged at 20,000 × g to remove excess buffer, nuclease-free water was added, incubated for 5 min at room temperature and centrifuged at 20,000 × g to elute the genomic DNA. The concentration of genomic DNA was measured with a Qubit Flex Fluorometer using the dsDNA HS Assay kit according to the manufacturer’s instruction. 500 μg of DNA with an DNA Integrity Number value > 7 from each condition was sent to the MDACC Advanced Technology Genomics Core for library preparation and WGS sequencing on the Illumina NovaSeq instrument.
Whole Genome Sequencing data analysis
The raw paired-end FASTQ reads were generated from whole genome sequencing of RT4 and K2 cells. We trimmed the raw sequences with Trimmomatic v. 0.39 to remove adapters and low-quality bases. The reads were mapped to human reference genome (hg19) using BWA-MEM of Burrows-Wheeler Aligner(BWA) v0.7.17-r1188 and mapped “bam” reads were generated. Further processing including duplicate removal, sorting and indexing was performed with SAMTOOLS v1.1386. We estimated copy number variants for each sample using CNVkit and genome wide heatmap showing copy number ratio was generated with “cnvkit.py heatmap” module. As our study was focused on identification of eccDNA, we used Circle-Map v.1.1.2 to identify circular amplicons87. All soft clipped, hard clipped and discordant reads were extracted to a new “bam” file using “ReadExtractor” module and “realignment” module was executed with default parameters of Circle-Map. We used standard threshold of minimum 2 split reads and circle score above 50 to obtain robust set of eccDNA. Circlize v 0.4.15, Biorender and ggplot2 were used to generate circular representations of identified eccDNA amplicons and copy number variation for specific regions between different groups were visualized with “cnvkit.py scatter” module.
ChIP Sequencing
ChIP assays were performed using pooled samples from MB49 sgScramble and two biological clones of sgKdm6a (C1, C2) cell lines, as well as in-vitro generated Tregs, treated with or without 25 mM Na-La for 48 h, using the MAGnify Chromatin Immunoprecipitation System kit as per the manufacturer’s instructions. Briefly, cells were incubated with 37% formaldehyde for 10 min at room temperature to crosslink the chromatin. Then to stop the crosslinking reaction, the samples were treated with 1.5 M glycine and incubated for 5 min at room temperature. Next, the samples were washed thrice with ice-cold PBS through centrifugation at 200 × g at 4 °C for 10 min. The resulting cell pellets were then resuspended in lysis buffer supplemented with protease inhibitors and subjected to sonication to fragment the DNA into 150-300 kb fragments. For every immunoprecipitation reaction, 10 μg of specific antibodies as detailed in the Figure legends and Supplementary Table 1, were first coupled with Protein A/G Dynabeads and then added to the sheared samples and incubated overnight at 4 °C.The subsequent day, unbound antibodies were removed by washing thrice with Immunoprecipitation (IP) Buffer 1 and twice with IP Buffer 2. The crosslinking was then reversed through heat treatment, using the Reverse crosslinking buffer supplemented with Proteinase K. Next, the target DNA was purified using the DNA purification magnetic beads resuspended in DNA purification buffer and incubated at room temperature for 30 min. The bead-bound DNA was then washed twice with DNA wash buffer to remove any residual contaminants. Finally, the purified target DNA was eluted out by incubating the bead-bound DNA with DNA elution buffer at 55 °C for 20 min in a thermocycler. The immunoprecipitated DNA was then quantified with the dsDNA HS Assay kit using the Qubit Flex Fluorometer. 10 μg of DNA from each condition was sent to the MDACC ATGC for library preparation and sequencing which was carried out using the Illumina NextSeq500 instrument.
ChIP Sequencing data analysis
The quality of ChIP-seq FASTQ sequences generated as described above, were assessed using FastQC v0.11.9, followed by mapping by bowtie2 v2.3.5.188 with mouse reference genome mm10. The bam files obtained from mapping were further processed using SAMBLASTER v0.1.2689 and SAMTOOLS v1.1386, for duplicate removal, sorting and indexing. Further, SAMBAMBA v0.6.690 was used to normalize the bam files per read counts by performing random sampling. The ChIP-seq signal enrichment over “Input control” background was identified using Model based analysis of ChIP-seq (MACS)2 v2.2.891. The peaks with significant p values (<0.05) were considered for further annotation with using CHIPseeker v1.30.392 and clusterProfiler93 packages in R v4.3.2. The quantitative pairwise comparisons of different datasets were performed using Manorm v1.1.494 using both peak (bed) and read (bam) coordinates for respective sample. The significant enrichment of target genes was estimated of the basis of M value which describes the log2 fold change and plotted with ggplot2. The differential peaks for specific genes were visualized using Integrative Genomics Viewer (IGV)84,85. Differential pathways enriched among datasets were identified using GSEA v4.2.395. The bigwig files were generated from aligned bam reads with “bamCoverage” function of deepTools v3.5.196 and profile plots for specific genes were plotted using the computeMatrix and plotProfile programs of deepTools. We obtained differential peaks for set of genes involved in MMR, DSBR and glycolysis to prepare consolidated BED for each pathway between groups. Signal intensity over these regions was quantified with deepTools using computeMatrix. The resulting matrix was visualized with plotProfile to obtain average signal profiles for consolidated set of genes involved in MMR, DSBR and glycolysis for different groups. For profile plots of individual genes, we extracted the differential peaks mapping to genes involved in MMR, DSBR and glycolysis as BED regions for each gene individually (reference-point mode, TSS, +/−1kb flanking regions). Signal intensity over these regions was quantified with deepTools using computeMatrix (reference-point mode, TSS, +/−1kb flanking regions). The resulting matrix was visualized with plotProfile to obtain average signal profiles for signals for genes of interest for different groups. The coverage (bw) data were used to generate heatmaps showing the distribution of promoters (TSS, +/−5 kb) in Treg with and without sodium lactate treatment for different histone post-translational using “computeMatrix” and “plotHeatmap” functions of deepTools. The genome wide annotation of aligned reads was performed using ChromHMM v1.2497 which applies multivariate hidden Markov model (HMM) to assign states by modeling combinatorial presence and absence of each mark. The reads were subjected to “BinarizeBam” followed by “LearnModel” using mm10 assembly study enrichment and functional annotation of each mark in 7-state model.
RNA sequencing
MB49 sgScramble and two biological clones of sgKdm6a (C1, C2) cells were collected from culture by trypsinization, washed once in PBS, and then snap-frozen in liquid nitrogen before total RNA extraction performed by the MDACC Biospecimen Extraction Core Facility. A minimum of 120 ng of extracted RNA with an RNA Integrity Number value > 7 was sent to the MDACC ATGC for stranded, paired-end mRNA sequencing in a NextSeq500 instrument.
RNA sequencing data analysis
For RNA-seq analysis, the paired-end raw FASTQ sequences were subjected to quality control with FastQC v0.11.9, followed by adapter removal using Trimmomatic v0.39 78. The trimmed fastq reads were mapped to mm10 reference genome using STAR v2.7.3a 79 and resulting bam reads were sorted and indexed with SAMTOOLS v1.13. The gene counts for each sample were estimated with HTseq v2.0.2. The raw counts from HTSeq were imported into R and analyzed with edgeR to compare different groups. Count matrices were assembled and labeled accordingly, then converted to DGElist. CPM were computed for quality checks and dispersion was visualized with plotBCV. As this comparison did not include biological replicates, differential expressions were assessed with edgeR’s exact test with a fixed biological coefficient of variation as recommended by published protocol98. We adjusted the resulting P-values obtained from differential gene expression analysis between groups for multiple testing using the Benjamini–Hochberg false discovery rate (FDR). The volcano plots showing differential gene expression between groups were plotted with ggplot2 in R.
Isolation of CD4+CD25+ Tregs and CD4+CD25− Tconv cells
Murine CD4+CD25+ Tregs, and CD4+CD25− Tconv cells were magnetically isolated using murine CD4+CD25+ Regulatory T Cell Isolation Kit, according to the manufacturer’s protocol. Briefly, cells were resuspended in MACS buffer and CD4+CD25+ Regulatory T Cell Biotin-Antibody Cocktail was added and incubated on ice for 15 min. Next, the Anti-Biotin Microbeads were added, incubated at 4 °C for 15 min, following which the samples were washed in surplus MACS buffer by centrifugation. The resulting cell pellet was passed through the LD columns, flow through was collected and centrifuged at 200 × g for 5 min at 4 °C. The cell pellet was then resuspended in MACS buffer, CD25-PE antibody was added, mixed well, and incubated for another 15 min on ice. The samples were incubated for an additional 15 min with Anti-PE Microbeads and washed in excess MACS buffer by centrifugation. The pellets were resuspended and run through the MS Columns. CD4+CD25− Tconv cells were collected in the flow-through, while CD4+CD25+ Tregs were eluted from the column using a plunger.
Flow cytometry
Flow cytometry was conducted on the cells as mentioned in the figure legend.
2-3 ×106 cells were briefly rinsed in FACS buffer (PBS supplemented with 5% FBS) by centrifugation at 200 × g at 4 °C for 5 min. After washing, the cells were incubated on ice for 15 min in a blocking buffer containing murine, bovine, hamster, rat, and rabbit serum along with 25 μg/mL 2.4G2 (Fc block) antibody in PBS. The cells were then stained with the surface antibody cocktail as detailed in the Supplementary Table 1 for 30 min at 4 °C. After, two subsequent washes with FACS buffer, the cells were fixed and permeabilized on ice for 45 min. Following another wash with 1X permeabilization buffer, the cells were incubated in the intracellular antibody cocktail as mentioned in Supplementary Table 1 for 20 min at room temperature. Finally, the samples were washed with FACS buffer, fixed with 1% paraformaldehyde. The samples were then acquired using a BD LSRFortessa flow cytometer and FlowJo v10.10 software was used for analysis of the flow cytometry data.
Annexin V based cytotoxicity assay
To evaluate cisplatin-induced cytotoxicity, between MB49 sgScramble, RT4 and SCaBER cells with their corresponding KDM6A KO clones, cells were treated with or without 5 µM cisplatin for 48 h. Following that the cells were harvested and stained with Annexin V-Pacific Blue diluted in 1X Annexin Binding Buffer for 15 min at room temperature. The stained cells were then washed with 1X Annexin Binding Buffer and acquired immediately on the BD LSR Fortessa flow cytometer and analyzed using FlowJo v10.10 software.
Spheroid generation assay
To compare the spheroid formation ability between MB49 sgScramble, RT4 and SCaBER cells with their corresponding KDM6A KO clones in presence of cisplatin, cells were allowed to form spheroids on top of agarose. Briefly, 100 µl of 1.5% molten agarose was transferred to each well of a 96-well flat-bottom plate and allowed to solidify. Then 2 ×104 cells resuspended in 200 µl of complete media containing 5 µM cisplatin were transferred to the top of the agarose. To allow spheroid formation, the plates were centrifuged at 220 × g for 10 min and incubated for 48 h at 37 °C. After 48 h, spheroids were imaged in a EVOS FL imaging system under a 4X objective. The ImageJ software was used to measure the diameter of the spheroids using the scale in the microscopy images as reference.
Wound-healing assay
To compare the migration ability of MB49 sgScramble, and SCaBER with their respective KDM6A KO clones in presence of cisplatin, wound-healing assay was performed. Briefly, 2 ×105 cells were seeded into each well of a 24-well plate and incubated until cells reached confluency. Using a sterile 200 µl pipette tip a scratch was made through the middle of the well and imaged at 0 h in a EVOS FL imaging system under a 4X objective. The cells were then treated with or without 5 µM cisplatin in serum free media followed by 48 h incubation and imaged as described previously. The images were analyzed in ImageJ Software using the Wound healing size tool to determine the wound closure percentage. The wound closure was calculated as the percentage reduction in scratch area between 0 and 48 h.
Cell Transwell Invasion assay
To determine cellular invasiveness, the MB49 sgScramble, RT4 and SCaBER with their respective KDM6A KO clones were subjected to starvation by culturing in serum free media for 24 h. Following serum starvation, 5 ×104 cells were seeded in FBS free media, either with or without 5 µM cisplatin, onto the transwell inserts with 8μm pore membrane coated with Matrigel (1:8 dilution in serum free media). The inserts were then placed into wells of 24-well plate containing 30% FBS media. Following 48 h incubation, the invaded cells on the lower side of the membrane were washed with PBS and fixed in 4% PFA for 7 min at RT. The membranes were then excised out using scalpel, mounted on a glass slide using VECTASHIELD mounting media containing DAPI and imaged using the EVOS FL imaging system at either 10X or 20X magnification. The cell numbers were quantified by counting 4 independent visual fields using the ImageJ Software.
Cell proliferation assay
1 ×106 cells of interest were stained with CellTrace™ CFSE according to the manufacturer’s instructions. Briefly, cells were harvested and resuspended in 1 ml PBS, then 1 µl of 5 mM CFSE dye was added, and the cells were incubated in dark at RT for 20 min. Following incubation, the cells were resuspended in complete media and incubated for another 5 min. Cells were then centrifuged and washed again with complete medium to remove any excess dye. At 0 h, a baseline measurement of CFSE intensity was acquired in BD LSRFortessa flow cytometer in FITC channel. The cells were then cultured for an additional 48 h, after which they were acquired again in flow cytometer. FlowJo v10.10 software was employed to analyze the differences in the CFSE dilution which is used as a measure of cell proliferation.
Extrachromosomal circular DNA isolation and rolling circle amplification
Nuclei extraction was performed on 7 million sgScramble and sgKdm6a C1 cells per reaction, using CUTANA nuclei extraction buffer supplemented with spermidine and Halt protease inhibitor. The samples were incubated for 10 min on ice, and then centrifuged at 600 × g for 5 min at 4 °C. Following nuclei extraction, lysis and circular DNA isolation was performed using the QIAprep spin miniprep kit according to the manufacturer’s instructions. Briefly, the nuclei were resuspended in P1 buffer, following which proteinase K was added to the cell suspension. The samples were incubated for 17 h on a 50 °C heating block with agitation at 700 rpm. Then, buffer P2 was added to the samples and mixed thoroughly by inversion. To stop the lysis reaction, buffer N3 was added and mixed thoroughly by inversion. The sample was spun at 17,900 × g for 1 min. The supernatant was collected, added in the QIAprep spin column, and spun at 17,900 × g for 1 min. The spin column was washed using buffer PB, and spun at 17,900 × g for 1 min followed by a wash with buffer PE and spun at 17,900 × g for 1 min. An additional spin was performed to remove residual wash buffer, then the column was placed on a fresh 1.5 mL Eppendorf tube. Nuclease free water was added to the column and left to stand for 5 min at room temperature and eluted by centrifugation at 17,900 × g for 1 min.
Subsequently the isolated DNA was subjected to linear DNA using the Plasmid safe ATP dependent DNase kit at 37 °C for 1 h and the reaction was deactivated at 70 °C for 30 min, as per the manufacturer’s instruction. Next, the sample was purified using the AMPure XP Reagent according to the manufacturer’s instructions.
Subsequently the purified DNA was subjected to rolling circle DNA amplification. Briefly, a denaturation mix was prepared using the phiDNA polymerase kit according to the manufacturer’s instructions. The sample was amplified for 21 h at 30 °C, then inactivated for 10 min at 65 °C. The PCR products were visualized via electrophoresis on a 1% agarose gel.
Immunohistochemistry (IHC)
IHC staining was performed on formalin-fixed, paraffin-embedded tissue sections of sgScramble and sgKdm6a C1 tumors to detect EXO1 levels. sgScramble and sgKdm6a C1 tumors were isolated as described previously, fixed in 10% formalin for 4 h and sent to MDACC Research Histology Core Laboratory (RHCL) for sectioning and staining. Fixed tumors were embedded in paraffin and sectioned at 4-micron thickness. Next, sections were deparaffinized, hydrated and antigen retrieved with BOND Epitope Retrieval Solution 1 followed by washing with BOND wash buffer. For staining BOND Polymer Refine Detection kit was used, sections were blocked with hydrogen peroxide for 15 min, stained with rabbit anti-mouse EXO1 antibody at a 1:200 dilution for 30 min, followed by staining with anti-rabbit Poly-HRP-IgG secondary antibody for 15 min. Slides were visualized using 3′-3-Diaminobenzidine substrate as a chromogen, followed by hematoxylin counterstaining for 3 min. Slides were scanned into Aperio eSlide manager for visualization. The quantification of the Exo1 expression was performed using the color deconvolution 2 plugin on the ImageJ software.
Microsatellite Instability detection
To detect Microsatellite instability, the following markers were used- mBAT64, mBAT24, mBAT37 and mBAT59 for the mouse genome and NR21, NR27, NR24, BAT25 and BAT26 for the human genome. These microsatellite markers were PCR amplified from the respective genomes, and the size of the marker was analyzed by fluorescent fragment length analysis (FFLA). Briefly, sgScramble and sgKdm6a C1 tumors were processed as described previously to obtain sgScramble and sgKdm6a C1 single cell suspensions, while RT4 and K2 cells were harvested via trypsinization. Genomic DNA was extracted from 2 ×106 cells using QIAamp DNA Micro Kit as previously detailed. These microsatellite markers were PCR amplified from 20 ng genomic DNA with primers as detailed in the Supplementary Table 1, using the Type-it Microsatellite PCR Kit as per the manufacturer’s instructions. PCR conditions were as follows for murine microsatellite markers: initial activation at 95 °C for 5 min; followed by 27 cycles of 95 °C for 30 s, 60 °C for 90 s and 72 °C for 30 s; then a final extension step at 60 °C for 30 min. For human microsatellite markers, the following PCR conditions were used: initial activation at 95 °C for 15 min; followed by 35 cycles of 95 °C for 30 s, 55 °C for 90 s and 72 °C for 30 s; then a final extension step at 72 °C for 10 min. The size and distribution of the PCR amplicons were determined by capillary electrophoresis performed on a 3730 Genetic Analyzer instrument followed by analysis of data using the Peak Scanner software. The microsatellite-based amplicons were sent to MDACC ATGC for FFLA. The amplicons were mixed with formamide and Liz 500 size standard in a 96-well PCR plate, loaded on the mentioned instrument and run for 45 min. The results (.fcs files) were imported to Peak Scanner software, size standard and analysis method were set to GS500 LIZ and Sizing Default PP, respectively and analyzed. The modal peaks of the microsatellite markers in the electropherogram were identified between 75-195 bp. The modal peak position in base pairs of mBAT-64 was compared between sgScramble and sgKdm6a C1 tumor cells and the modal peak position for NR21 was compared between RT4 and K2 cells.
Gamma-H2AX assays
To detect dsDNA breaks in cell lines, presence of gamma-H2AX foci were detected using flow cytometry or microscopy-based studies.
For flow cytometry experiments, sgScramble and sgKdm6a C1 cells were seeded in a 24-well plate and incubated overnight at 37 °C. The following day, cells were treated with 100 µM Etoposide for 1 h followed by 4 h recovery or no recovery. Cells resuspended in PBS were transferred to a 96-well round bottom plate, centrifuged at 900 × g for 5 min at 4 °C and resuspended in fixation and permeabilization buffer for 40 min on ice. Following fixation and permeabilization, the cells were centrifuged at 900 × g for 5 min at 4 °C and the cell pellet was subsequently stained with Anti-Phospho-Histone H2A.X (Ser 139) primary antibody for 20 min at room temperature. After incubation, cells were washed with PBS by centrifugation at 1400 × g for 5 min at 4 °C and subsequently stained with anti-mouse FITC-conjugated secondary antibody for 15 min in dark at room temperature. Then, the cells were washed with PBS and fixed with 1% paraformaldehyde. The samples were acquired using a BD LSR Fortessa flow cytometer and analyzed using FlowJo v10.10 to determine the percentage of gamma-H2AX positive cells.
For microscopy experiments, 1 ×106 sgScramble, sgKdm6a C1, RT4, and K2 cells were seeded on poly-D-lysine coated coverslips and incubated for 24 h at 37 °C. After the cells properly adhered, they were treated as described previously, then fixed with 3.7% paraformaldehyde for 20 min at room temperature, permeabilized with 1% Triton X-100 for 10 min at room temperature and subsequently blocked for 30 min with blocking buffer (1% BSA in PBS) at room temperature. After blocking, the cells were incubated with the H2AX primary antibody as mentioned previously for 90 min at room temperature, washed with PBST (0.05% Tween-20 in PBS) and then incubated with anti-mouse secondary antibody in the dark for 1 h at room temperature. Then, the cells were washed with PBST followed by DAPI staining in the dark for 15 min. To remove excess dye cells were washed with PBS and cover slips were mounted on to slides using mounting media. The slides were imaged using the Leica SP8 confocal system or the Zeiss LSM880 confocal system at the MDACC Advanced Microscopy Core (AMC) and the raw image files were imported to ImageJ software for analysis. The gamma value was corrected uniformly within each comparison group and the number of distinctly visible gamma-H2AX foci were counted manually for each cell.
Gamma-H2AX assay of in-vivo tumor cells
sgScramble and sgKdm6A C1 tumors were processed as described previously to obtain sgScramble and sgKdm6a C1 tumor single cell suspensions. 3 ×106 sgScramble and sgKdm6a C1 tumor cells were transferred to a 96-well round bottom plate, centrifuged at 900 × g for 5 min at 4 °C and the cell pellet was stained with PB conjugated anti-CD45+ antibody followed by 15 min incubation at room temperature. The cells were washed with PBS followed by fixation and permeabilization and then stained with H2AX primary antibody and anti-mouse secondary antibody as described previously. The cells were then washed with PBS and incubated with PE conjugated tumor cell marker anti-E-Cadherin antibody for 15 min at room temperature. Cells were washed, fixed, acquired and analyzed as described previously to determine the MFI of gamma-H2AX in tumor cells and percentage of gamma-H2AX positive tumor cells.
Comet assay
To compare DNA damage in MB49 sgScramble and RT4 cell lines with their respective KDM6A KO cell lines, comet assay kit was used for single cell gel electrophoresis and DNA staining according to the manufacturer’s protocol. 2 × 105 cells were seeded in a 24-well plate and incubated overnight at 37 °C. The following day, cells were treated with 100 µM Etoposide for 2 h followed by 1 h recovery. Following the treatment, cells were collected, resuspended in cold 1X PBS to achieve a final of 500 cells/μl and mixed with molten comet agarose at 37 °C in the ratio of 1:10 (v/v). A base layer of agarose was prepared on comet slides and allowed to solidify, then the cell-agarose mixture was transferred on top of the base layer. Following solidification, alkaline lysis of cells was performed by immersing the slides in pre-chilled lysis buffer for 45 min at 4 °C in the dark, followed by a 45-minute immersion in pre-chilled alkaline solution. Then single cell alkaline electrophoresis was performed by placing the slides in an electrophoresis chamber containing cold alkaline electrophoresis solution, at 1 volt/cm for 25 min at 310 mA. The slides were then washed three times with cold autoclaved water, followed by cold 70% ethanol for 5 min. Slides were allowed to air dry at room temperature and stained with Vista green DNA dye for 15 min in dark at room temperature. Excess dye was removed, and slides were imaged using Leica SP8 confocal system at the MDACC Advanced Microscopy Core. The raw image files were imported to ImageJ software, and gamma value was corrected uniformly within each comparison group for further analysis. Using the OpenComet plugin, profile analysis was done with background correction off for comet finding and head finding. The output of the analysis provided the corresponding tail DNA percentage, tail moment and olive moment of the comets.
Cell cycle analysis
Cell population in each cell cycle stage was determined by using the Vybrant DyeCycle Violet DNA binding Stain according to the manufacturer’s instruction. Briefly, 2 ×105 cells were incubated with 10 µM of the DNA binding dye in complete media at 37 °C in dark for 30 min. The cells were then directly acquired in BD LSRFortessa flow cytometer using the BV421 channel. FlowJo v10.10 software was used for analysis of the flow cytometry data. The univariate cell cycle model using the Watson (Pragmatic) algorithm in FlowJo was used to analyze the distribution of cells across different cell cycle stages based on their DNA content.
Seahorse assay (ATP Real-time Rate Assay)
To compare the ATP production rates from glycolytic and mitochondrial pathways between murine MB49 sgScramble and sgKdm6a C1 cell lines as well as between human RT4 and its KDM6A KO Clone K2, the Seahorse XF Real-Time ATP rate assay was performed following the manufacturer’s instructions by the MDACC Metabolomics Core. The cells were seeded overnight in the XF-96 well-plate. The assay was performed using an XF-96 Analyzer which measured the extracellular acidification rate (ECAR) under basal conditions as well as after sequential injections of 1.5 μM oligomycin and 0.5 μM Rotenone/ Antimycin A. Real-time rates of ATP production were recorded and assessed using the Wave software.
Extracellular lactate measurement assay
To distinguish differences in concentration of L-Lactate produced, culture supernatants were collected from murine MB49 sgScramble and sgKdm6a C1 cell lines after 24 h of culture, and from human RT4 and its KDM6A KO Clone K2 cells following 48 h of culture.
To compare the intratumoral concentration of L-Lactate in sgScramble and sgKdm6a C1 tumors, tumor interstitial fluid (TIF) was collected by resting tumors on a 40 μm nylon filter, then centrifuging the sample tubes at 106 × g for 10 min at 4 °C. The collected TIF samples were transferred to separate Eppendorf tubes, snap-frozen in liquid nitrogen, and then stored at −80 °C before assaying.
TIF and culture supernatant samples were processed for extracellular lactate quantification using the L-Lactate Assay Kit. Briefly, samples were subjected to deproteination using cold 0.5 M MPA on ice for 5 min. After deproteination, samples were centrifuged at 10,000 × g for 5 min at 4 °C. Next, the supernatant was collected, transferred to a new tube, and neutralized using potassium carbonate and centrifuged at 10,000 × g for 5 min at 4 °C, after which the supernatant was collected. The assay plating was conducted following the manufacturer’s instructions, and the absorbance was read using a BioTek Synergy H1 microplate reader at 535 nm.
Mass cytometry (CyTOF) for immune profiling of tumor microenvironment
Tumor samples processed and stored as previously described were revived in complete DMEM media. Then, 3 × 106 cells were collected from each sample and washed with PBS, resuspended in 194Pt monoisotopic cisplatin at a final concentration of 5 μM in FACS buffer and incubated for 3 min at room temperature. Subsequently, samples were washed three times with FACS buffer followed by fixation and permeabilization for 15 min at room temperature using Maxpar barcode fixation and permeabilization buffer. Then, samples were labeled using palladium barcoding, according to the manufacturer’s protocol, and incubated for 30 min at room temperature. After labeling, samples were washed three times in FACS buffer and surface staining was performed by resuspending samples in the antibody mixture (outlined in Supplementary Table 1) prepared in blocking buffer at 4 °C for 30 min. After surface staining, samples were washed twice with FACS buffer and then subjected to fixation and permeabilization for 1 h at 4 °C. Next, samples were washed twice in permeabilization buffer and resuspended in the intracellular antibody mixture prepared in permeabilization buffer (described in Supplementary Table 1) for 30 min at 4 °C. After washing with FACS three times, samples were fixed using 1.6% paraformaldehyde in PBS supplemented with 100 nM iridium nucleic acid intercalator and left overnight at 4 °C. The following day, samples were washed with PBS three times and resuspended in nuclease-free water before acquisition using a Helios mass cytometer. We analyzed the CyTOF data using Premessa, FlowCore, and FlowSOM packages as described in a previously published protocol99.
pHrodo red assay for lactic-acid uptake
Tregs were collected from culture and 1 ×106 cells were plated per well in a 96 well plate, then washed with 20 mM HEPES in PBS. After washing, cells were resuspended in the pHrodo staining solution prepared according to manufacturer’s instructions in 20 mM HEPES and incubated at 37 °C for 30 min. Following staining, cells were washed twice in 20 mM HEPES. Following the washes, lactic acid was added to each sample well at final concentrations of 5 mM and 15 mM and then incubated at 37 °C for 30 min, following which samples were acquired on a BD LSR Fortessa flow cytometer and analyzed using FlowJo v10.10.
In-vitro suppression assay
Tregs were generated in-vitro, magnetically sorted for CD4+CD25+ Tregs as described previously and then treated with the supernatant of sgScramble or sgKdm6a C1 cell lines for 36 h. For Tconv cells, single cell suspensions of murine splenocytes were obtained and magnetically isolated for CD4+CD25− Tconv cells as previously outlined.
Tconv and Tregs were labelled with Cell Trace Violet (CTV) and Far Red (FR) respectively according to the manufacturer’s protocol. Tconv (5 × 104) cells were co-cultured with Tregs (5 × 104) at varying ratios (1:1,2:1 and 1:2, Tconv:Treg ratio) and stimulated using anti-mouse CD3/CD28 Dynabeads and 10 ng/ml of murine recombinant IL-2 per the manufacturer’s guidelines. The cells were co-cultured in 96 well round bottom plates with 200 μl RPMI for 4 days at 37 °C and 5% CO2. On Day 4, cells were acquired using the BD LSR Fortessa. FlowJo v10.10 software was employed to analyze the differences in the CTV dilution which is used as a measure of Tconv cell proliferation in response to co-culture with Tregs pre-treated with sgScramble supernatant or sgKdm6a supernatant.
Isolation and enrichment of histones for high-performance liquid chromatography (HPLC)–tandem mass spectrometry (MS/MS)
Tregs were generated in-vitro as described above and then treated with or without 25 mM 13C labelled Na-La for 48 h. Following treatment, histones were isolated using the Active Motif Histone Purification Mini Kit according to the manufacturer’s instructions. Briefly cells were harvested and washed by centrifugation at 200 × g for 5 min at 4 °C. The pellet was resuspended in pre-chilled extraction buffer and incubated overnight at 4 °C with end-to-end rotation. The subsequent day, the samples were centrifuged at maximum speed for 5 min at 4 °C, the supernatant containing the crude histones was collected and neutralized with 5X Neutralization Buffer. The neutralized histone samples were then loaded onto a spin column and centrifuged at 2000 × g for 5 min at 4 °C. The column was washed thrice with Histone wash buffer and the histones were eluted out in the flow through by centrifugation at 2000 × g for 5 min at 4 °C with the elution buffer. Next, the eluted histones were precipitated using 4% perchloric acid by overnight incubation at 4 °C. The subsequent day, samples were centrifuged at maximum speed at 4 °C for an hour. The pelleted histones were then washed twice with cold 4% perchloric acid followed by two washes with ice-cold acetone (with 0.2% HCl) and pre-chilled acetone. The pellets were air-dried for 15 min and then resuspended in 100 mM tetraethylammonium bromide (pH 8) by incubation for 15 min at room temperature with periodic vortexing.
The extracted histones were then quantitated using the Pierce BCA Assay Kit as outlined previously. 1000 µg of purified histones were then subjected to Lys-C digestion overnight at 1:50 (Lys-C to histone) ratio following the manufacturer’s guidelines. The digested samples were subsequently reduced and alkylated using 5 mM Dithiothreitol at 37 °C for an hour and 11 mM Iodoacetamide for 50 min at room temperature respectively.
The Lys-C digested histones were then immunoprecipitated using a Pan anti-Kla antibody using Pierce Classic IP Kit following the manufacturer’s protocol. 10 µg of Pan anti-Kla antibody was incubated overnight with 1000 µg of digested histones at 4 °C with end-to-end rotation. The next day, Pierce Protein A/G Agarose slurry solution was added into the Pierce Spin Column and centrifuged for 1 min at 1000 × g. Then the antibody/ digested peptide mixture was added onto the Protein A/G Plus Agarose in the spin column and incubated over end-over-end rotation for 3.5 h. The column was then washed thrice using 1 × 0.025 M TBS Buffer and once with 1X Conditioning Buffer. The immune complex was eluted out of the column following incubation with low-pH elution buffer for 15 min and then centrifugation at 1000 × g for 1 min at 4 °C. The enriched peptides were then quantitated by BCA assay as described previously and sent to the MDACC Proteomics Core for LC–MS/MS analysis.
LC–MS/MS Analysis
For the analysis of histone lactylation, vacuum-dried tryptic peptides were dissolved in solvent A (0.1% formic acid in water) and directly loaded onto a nano reverse-phase C18 column (75 μm ID × 25 cm L, 1.7 μm particle size). Peptides were separated using a linear gradient of solvent A (0.1% formic acid in water) and solvent B (0.1% formic acid in acetonitrile). The gradient was as follows: 0% to 10% solvent B over 2 min, 10% to 60% solvent B over 30 min, 60% to 100% solvent B over 1 min, and held at 100% solvent B for 11 min. The eluate was electrosprayed into an Orbitrap Astral Mass Spectrometer (Thermo Fisher Scientific) operating in positive ion mode, with an electrospray voltage of 2 kV and an ion transfer tube temperature of 280 °C, flowrate 400 nl/min. Data were acquired in DDA mode with a fixed cycle time of 2 s. Full MS scans were performed in the range of 350 to 1500 m/z at a resolution of 240,000, with an AGC target of 500% and a maximum injection time of 100 ms. Precursor ion selection width was set to 2 Th. Peptide fragmentation was carried out using HCD with a normalized collision energy of 27%. Fragment ion scans were recorded at a resolution of 80,000, with a maximum injection time of 30 ms and a normalized AGC target of 300%. Dynamic exclusion was enabled and set to 15 s.
For data analysis, the MS/MS spectra were processed using Proteome Discoverer v2.4 with a mouse protein sequence database from UniProt (downloaded November 2023, 17,184 entries). Peptide searching was performed with Sequest HT using the following parameters: Protease- Trypsin, allowing up to two missed cleavages, minimum peptide length of 5, a precursor mass tolerance of 10 ppm, and a product ion tolerance of 0.6 Da and applied score cutoff-none. Carboxyamidomethylation of cysteine was set as a fixed modification, while oxidation of methionine and lactylation of lysine were set as variable modifications. A total of eight samples were analysed. These included two technical replicates of in-vitro generated untreated Tregs and two technical replicates of Tregs labeled for 48 h with 25 mM 13C sodium L-lactate. For each experimental condition, two corresponding input controls (5% of the peptide collected prior to IP) were also included.
ChIP-qPCR
ChIP assay was performed as previously detailed on the following samples- Human RT4 and K2 cells using antibodies against KDM6A and H3K4me3 and in-vitro generated, CD4+CD25+ sorted Tregs treated with and without 25 mM Na-La for 48 h or treated with supernatants from sgScramble or sgKdm6a C1 cells for additional 24 h against anti- H3K9la and H3K18la antibodies (detailed in Supplementary Table 1). For each condition, 100 ng of immunoprecipitated DNA was used as input for real-time PCR (qPCR) to evaluate the gene expression utilizing the SYBR Green Master Mix according to the manufacturer’s instructions and using the primer sequences listed in Supplementary Table 1.
RNA Isolation, cDNA preparation and qPCR
RT4 with its KDM6A KO K2; SCaBER with its KDM6A KO B7 and in-vitro generated, CD4+CD25+ sorted Tregs treated with and without 25 mM Na-La for 48 h or treated with the supernatant of sgScramble and sgKdm6a C1 cells for 24 h were lysed using the TriZol reagent and one-fifth volume of chloroform was added. After vigorous mixing and a 15-minute incubation at room temperature the samples were centrifuged at 12,000 × g for 15 min at 4 °C. The aqueous layer on top was carefully collected and incubated with equal volume of isopropanol for 10 min at room temperature. Following this, the samples were centrifuged at 12,000 × g for 15 min at 4 °C. The pellets were then washed with 75% ethanol by centrifugation. The resulting RNA pellets were air-dried, resuspended in nuclease-free water and quantified using a Nanodrop spectrophotometer. Subsequently, cDNA synthesis was performed by reverse transcribing 1 µg of RNA using the Superscript III cDNA synthesis kit, according to the manufacturer’s instructions containing reverse transcriptase (RT) buffer, dNTP mix, RT Random primers, and Reverse transcriptase enzyme. 1000-2000ng of cDNA was then utilized for downstream qPCR analysis to measure gene expression using the SYBR Green Master Mix according to the manufacturer’s instructions and the primer sequences detailed in Supplementary Table 1.
Statistics & Reproducibility
R V4.0.2 and GraphPad Prism software V10 were used for statistical analyses. The specific statistical tests used have been indicated in the respective figure legends.
Ethical Statement
The study complies with all relevant ethical regulations, and all animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC protocol no. 00002153-RN01) at The University of Texas MD Anderson Cancer Center (MDACC).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ChIP-seq, RNA-seq and WGS data supporting this study have been deposited in Sequence Read Archive (SRA) under accession code PRJNA1173195. The mouse reference genome (mm10) coordinates are available in GENCODE. The human reference genome (hg19) coordinates are available in GENCODE. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE100 partner repository with the dataset identifier PXD065234. The clinical and genomic data for patients with resectable and advanced bladder cancer receiving cisplatin-based chemotherapy is available in cBioPortal as blca_msk_tcga_2020 (https://www.cbioportal.org/study/summary?id=blca_msk_tcga_2020) and paired_bladder_2022 (https://www.cbioportal.org/study/summary?id=paired_bladder_2022). The amplicon intervals for The Cancer Genome Atlas (TCGA) cohort are available in Supplementary Table 1 of a previously reported study (https://www.nature.com/articles/s41588-020-0678-2#Sec20). The overall survival, tumor mutation burden and mRNA expression data for IMviogor210 study is available as IMvigor210CoreBiologies R package (http://research-pub.gene.com/IMvigor210CoreBiologies/). The clinical and genomic data for MSK_2018 cohort is available in cBioPoral as tmb_mskcc_2018 (http://www.cbioportal.org/study?id=tmb_mskcc_2018). The tumor mutations burden and mRNA expression data for TCGA-BLCA cohort is available at cBioPortal (https://www.cbioportal.org/study/summary?id=blca_tcga_pub_2017). Somatic variants summary for bladder cancer patients is available at TCGA. The genomic and mRNA expression data for HCRN cohort is available in cBioPortal with identifier blca_bcan_hcrn_2022 (https://www.cbioportal.org/study/clinicalData?id=blca_bcan_hcrn_2022). The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
Code availability
The ChIP–seq, RNA-seq and WGS analyses were performed with open-source algorithms as described in the Methods.
References
Siegel, R. L., Kratzer, T. B., Giaquinto, A. N., Sung, H. & Jemal, A. Cancer statistics, 2025. CA Cancer J Clin 75, 10–45 (2025).
Alifrangis, C., McGovern, U., Freeman, A., Powles, T. & Linch, M. Molecular and histopathology directed therapy for advanced bladder cancer. Nat Rev Urol 16, 465–483 (2019).
Felsenstein, K. M. & Theodorescu, D. Precision medicine for urothelial bladder cancer: update on tumour genomics and immunotherapy. Nat Rev Urol 15, 92–111 (2018).
Wolff, E. M., Liang, G. & Jones, P. A. Mechanisms of Disease: genetic and epigenetic alterations that drive bladder cancer. Nat Clin Pract Urol 2, 502–510 (2005).
Robertson, A. G. et al. Comprehensive Molecular Characterization of Muscle-Invasive Bladder Cancer. Cell 171, 540–556.e525 (2017).
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome - biological and translational implications. Nat Rev Cancer 11, 726–734 (2011).
Tran, L., Xiao, J. F., Agarwal, N., Duex, J. E. & Theodorescu, D. Advances in bladder cancer biology and therapy. Nat Rev Cancer 21, 104–121 (2021).
Qiu, H. et al. KDM6A Loss Triggers an Epigenetic Switch That Disrupts Urothelial Differentiation and Drives Cell Proliferation in Bladder Cancer. Cancer Res 83, 814–829 (2023).
Rachakonda, P. S. et al. TERT promoter mutations in bladder cancer affect patient survival and disease recurrence through modification by a common polymorphism. Proc Natl Acad Sci USA 110, 17426–17431 (2013).
Glaser, A. P., Fantini, D., Shilatifard, A., Schaeffer, E. M. & Meeks, J. J. The evolving genomic landscape of urothelial carcinoma. Nat Rev Urol 14, 215–229 (2017).
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007).
Lan, F. et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature 449, 689–694 (2007).
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat Genet 52, 891–897 (2020).
Paulsen, T. et al. MicroDNA levels are dependent on MMEJ, repressed by c-NHEJ pathway, and stimulated by DNA damage. Nucleic Acids Res 49, 11787–11799 (2021).
Rose, J. C. et al. Disparate Pathways for Extrachromosomal DNA Biogenesis and Genomic DNA Repair. Cancer Discov 15, 69–82 (2025).
Prieto-Garcia, C. et al. Inhibition of USP28 overcomes Cisplatin-resistance of squamous tumors by suppression of the Fanconi anemia pathway. Cell Death Differ 29, 568–584 (2022).
Hira, A., Zhang, J. & Kadakia, M. P. TIP60 enhances cisplatin resistance via regulating DeltaNp63alpha acetylation in SCC. Cell Death Dis 15, 877 (2024).
Mendoza-Rodriguez, M. G. et al. IL-1beta Inflammatory Cytokine-Induced TP63 Isoform ∆NP63alpha Signaling Cascade Contributes to Cisplatin Resistance in Human Breast Cancer Cells. Int J Mol Sci 20, 270 (2019).
Breed, C. et al. Ovarian Tumor Cell Expression of Claudin-4 Reduces Apoptotic Response to Paclitaxel. Mol Cancer Res 17, 741–750 (2019).
Stewart, J. J. et al. Proteins associated with Cisplatin resistance in ovarian cancer cells identified by quantitative proteomic technology and integrated with mRNA expression levels. Mol Cell Proteomics 5, 433–443 (2006).
Zhu, W. et al. GLI2 inhibits cisplatin sensitivity in gastric cancer through DEC1/ZEB1 mediated EMT. Cell Death Dis 16, 204 (2025).
Wang, Q. et al. Hedgehog-Gli2 Signaling Promotes Chemoresistance in Ovarian Cancer Cells by Regulating MDR1. Front Oncol 11, 794959 (2021).
Tang, Y. A. et al. Hypoxic tumor microenvironment activates GLI2 via HIF-1alpha and TGF-beta2 to promote chemoresistance in colorectal cancer. Proc Natl Acad Sci USA 115, E5990–E5999 (2018).
Jia, Y. et al. The role of GLI-SOX2 signaling axis for gemcitabine resistance in pancreatic cancer. Oncogene 38, 1764–1777 (2019).
Nishii, Y. et al. CROP/Luc7A, a novel serine/arginine-rich nuclear protein, isolated from cisplatin-resistant cell line. FEBS Lett 465, 153–156 (2000).
Hou, Y. et al. Proteomics Identifies LUC7L3 as a Prognostic Biomarker for Hepatocellular Carcinoma. Curr Issues Mol Biol 46, 4004–4020 (2024).
Huang, D. et al. A highly annotated database of genes associated with platinum resistance in cancer. Oncogene 40, 6395–6405 (2021).
Qi, G. et al. SHCBP1 promotes cisplatin resistance of ovarian cancer through AKT/mTOR/Autophagy pathway. Apoptosis 30, 83–98 (2025).
Sun, Y., Pan, H., He, Y., Hu, C. & Gu, Y. Functional roles of the SHCBP1 and KIF23 interaction in modulating the cell-cycle and cisplatin resistance of head and neck squamous cell carcinoma. Head Neck 44, 591–605 (2022).
Zou, A. et al. SHCBP1 promotes cisplatin induced apoptosis resistance, migration and invasion through activating Wnt pathway. Life Sci 235, 116798 (2019).
Clinton, T. N. et al. Genomic heterogeneity as a barrier to precision oncology in urothelial cancer. Cell Rep 41, 111859 (2022).
Pietzak, E. J. et al. Genomic Differences Between “Primary” and “Secondary” Muscle-invasive Bladder Cancer as a Basis for Disparate Outcomes to Cisplatin-based Neoadjuvant Chemotherapy. Eur Urol 75, 231–239 (2019).
Lin, C. et al. Encoding gene RAB3B exists in linear chromosomal and circular extrachromosomal DNA and contributes to cisplatin resistance of hypopharyngeal squamous cell carcinoma via inducing autophagy. Cell Death Dis 13, 171 (2022).
Pal Choudhuri, S. et al. Acquired Cross-Resistance in Small Cell Lung Cancer due to Extrachromosomal DNA Amplification of MYC Paralogs. Cancer Discov 14, 804–827 (2024).
Nguyen, D. D. et al. The interplay of mutagenesis and ecDNA shapes urothelial cancer evolution. Nature 635, 219–228 (2024).
Tang, H., Chen, J., Han, X., Feng, Y. & Wang, F. Upregulation of SPP1 Is a Marker for Poor Lung Cancer Prognosis and Contributes to Cancer Progression and Cisplatin Resistance. Front Cell Dev Biol 9, 646390 (2021).
Chen, X. et al. SPP1 inhibition improves the cisplatin chemo-sensitivity of cervical cancer cell lines. Cancer Chemother Pharmacol 83, 603–613 (2019).
Hsu, S. C., Miller, S. A., Wang, Y. & Hung, M. C. Nuclear EGFR is required for cisplatin resistance and DNA repair. Am J Transl Res 1, 249–258 (2009).
Mariathasan, S. et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51, 202–206 (2019).
von Loga, K. et al. Extreme intratumour heterogeneity and driver evolution in mismatch repair deficient gastro-oesophageal cancer. Nat Commun 11, 139 (2020).
Damrauer, J. S. et al. Collaborative study from the Bladder Cancer Advocacy Network for the genomic analysis of metastatic urothelial cancer. Nat Commun 13, 6658 (2022).
Lipkin, S. M. et al. MLH3: a DNA mismatch repair gene associated with mammalian microsatellite instability. Nat Genet 24, 27–35 (2000).
Chung, J. et al. DNA Polymerase and Mismatch Repair Exert Distinct Microsatellite Instability Signatures in Normal and Malignant Human Cells. Cancer Discov 11, 1176–1191 (2021).
Maio, M. et al. Pembrolizumab in microsatellite instability high or mismatch repair deficient cancers: updated analysis from the phase II KEYNOTE-158 study. Ann Oncol 33, 929–938 (2022).
Grothey, A. Pembrolizumab in MSI-H-dMMR Advanced Colorectal Cancer - A New Standard of Care. N Engl J Med 383, 2283–2285 (2020).
Aebi, S. et al. Loss of DNA mismatch repair in acquired resistance to cisplatin. Cancer Res 56, 3087–3090 (1996).
Silva, M. M., Rocha, C. R. R., Kinker, G. S., Pelegrini, A. L. & Menck, C. F. M. The balance between NRF2/GSH antioxidant mediated pathway and DNA repair modulates cisplatin resistance in lung cancer cells. Sci Rep 9, 17639 (2019).
Sawant, A., Kothandapani, A., Zhitkovich, A., Sobol, R. W. & Patrick, S. M. Role of mismatch repair proteins in the processing of cisplatin interstrand cross-links. DNA Repair (Amst) 35, 126–136 (2015).
Tuninetti, V. et al. Retrospective Analysis of the Correlation of MSI-h/dMMR Status and Response to Therapy for Endometrial Cancer: RAME Study, a Multicenter Experience. Cancers (Basel) 15, 3639 (2023).
Kuo, L. J. & Yang, L. X. Gamma-H2AX - a novel biomarker for DNA double-strand breaks. In Vivo 22, 305–309 (2008).
Olive, P. L. & Banath, J. P. The comet assay: a method to measure DNA damage in individual cells. Nat Protoc 1, 23–29 (2006).
Chatzidoukaki, O., Goulielmaki, E., Schumacher, B. & Garinis, G. A. DNA Damage Response and Metabolic Reprogramming in Health and Disease. Trends Genet 36, 777–791 (2020).
Milanese, C. et al. DNA damage and transcription stress cause ATP-mediated redesign of metabolism and potentiation of anti-oxidant buffering. Nat Commun 10, 4887 (2019).
Sobanski, T. et al. Cell Metabolism and DNA Repair Pathways: Implications for Cancer Therapy. Front Cell Dev Biol 9, 633305 (2021).
Matassa, D. S. et al. Oxidative metabolism drives inflammation-induced platinum resistance in human ovarian cancer. Cell Death Differ 23, 1542–1554 (2016).
Xu, Y. et al. ABT737 reverses cisplatin resistance by targeting glucose metabolism of human ovarian cancer cells. Int J Oncol 53, 1055–1068 (2018).
De Martino, M., Rathmell, J. C., Galluzzi, L. & Vanpouille-Box, C. Cancer cell metabolism and antitumour immunity. Nat Rev Immunol 24, 654–669 (2024).
Watson, M. J. et al. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature 591, 645–651 (2021).
Batlle, E. & Massague, J. Transforming Growth Factor-beta Signaling in Immunity and Cancer. Immunity 50, 924–940 (2019).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Raychaudhuri, D. et al. Histone lactylation drives CD8(+) T cell metabolism and function. Nat Immunol 25, 2140–2151 (2024).
Dyrskjot, L. et al. Bladder cancer. Nat Rev Dis Primers 9, 58 (2023).
Chen, X. et al. Significance of KDM6A mutation in bladder cancer immune escape. BMC Cancer 21, 635 (2021).
Wang, S. et al. Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer. Nature communications 15, 1515 (2024).
Lv, W. et al. Spatial-Temporal Diversity of Extrachromosomal DNA Shapes Urothelial Carcinoma Evolution and Tumor-Immune Microenvironment. Cancer Discov. 15, 1225–1246 (2025).
Bailey, C. et al. Origins and impact of extrachromosomal DNA. Nature 635, 193–200 (2024).
Su, S. Beneath the Surface: Neoantigens beyond Chromosomal DNA Mutations. Cancer Discov 14, 2066–2070 (2024).
Akkers, R. C. et al. A hierarchy of H3K4me3 and H3K27me3 acquisition in spatial gene regulation in Xenopus embryos. Dev Cell 17, 425–434 (2009).
Lim, P. S., Li, J., Holloway, A. F. & Rao, S. Epigenetic regulation of inducible gene expression in the immune system. Immunology 139, 285–293 (2013).
Leng, X. et al. Histone 3 lysine-27 demethylase KDM6A coordinates with KMT2B to play an oncogenic role in NSCLC by regulating H3K4me3. Oncogene 39, 6468–6479 (2020).
Chen, Z., Qi, Y., Shen, J. & Chen, Z. Histone demethylase KDM6A coordinating with KMT2B regulates self-renewal and chemoresistance of non-small cell lung cancer stem cells. Transl Oncol 37, 101778 (2023).
Boila, L. D. et al. KDM6 demethylases integrate DNA repair gene regulation and loss of KDM6A sensitizes human acute myeloid leukemia to PARP and BCL2 inhibition. Leukemia 37, 751–764 (2023).
Lee, J. M., Ledermann, J. A. & Kohn, E. C. PARP Inhibitors for BRCA1/2 mutation-associated and BRCA-like malignancies. Ann Oncol 25, 32–40 (2014).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Goellner, E. M. et al. Identification of Exo1-Msh2 interaction motifs in DNA mismatch repair and new Msh2-binding partners. Nat Struct Mol Biol 25, 650–659 (2018).
Roy, M. et al. EXO1 promotes the meiotic MLH1-MLH3 endonuclease through conserved interactions with MLH1, MSH4 and DNA. Nat Commun 16, 4141 (2025).
Yan, S. et al. ZGRF1 promotes end resection of DNA homologous recombination via forming complex with BRCA1/EXO1. Cell Death Discov 7, 260 (2021).
Bolderson, E. et al. Phosphorylation of Exo1 modulates homologous recombination repair of DNA double-strand breaks. Nucleic Acids Res 38, 1821–1831 (2010).
Lu, G. et al. Ligase I and ligase III mediate the DNA double-strand break ligation in alternative end-joining. Proc Natl Acad Sci USA 113, 1256–1260 (2016).
Yang, R. et al. Homeobox A3 and KDM6A cooperate in transcriptional control of aerobic glycolysis and glioblastoma progression. Neuro Oncol 25, 635–647 (2023).
Gu, J. et al. Tumor metabolite lactate promotes tumorigenesis by modulating MOESIN lactylation and enhancing TGF-beta signaling in regulatory T cells. Cell Rep 39, 110986 (2022).
Kumagai, S. et al. Lactic acid promotes PD-1 expression in regulatory T cells in highly glycolytic tumor microenvironments. Cancer Cell 40, 201–218.e209 (2022).
Robinson, J. T. et al. Integrative genomics viewer. Nat Biotechnol 29, 24–26 (2011).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14, 178–192 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Prada-Luengo, I., Krogh, A., Maretty, L. & Regenberg, B. Sensitive detection of circular DNAs at single-nucleotide resolution using guided realignment of partially aligned reads. BMC Bioinformatics 20, 663 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
Faust, G. G. & Hall, I. M. SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505 (2014).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137 (2008).
Wang, Q. et al. Exploring Epigenomic Datasets by ChIPseeker. Curr Protoc 2, e585 (2022).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Shao, Z., Zhang, Y., Yuan, G. C., Orkin, S. H. & Waxman, D. J. MAnorm: a robust model for quantitative comparison of ChIP-Seq data sets. Genome Biol 13, R16 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550 (2005).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 44, W160–W165 (2016).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat Protoc 12, 2478–2492 (2017).
Chen, Y., Lun, A. T. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res 5, 1438 (2016).
Goswami, S. et al. Myeloid-specific KDM6B inhibition sensitizes glioblastoma to PD1 blockade. Nat Cancer 4, 1455–1473 (2023).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res 53, D543–D553 (2025).
Acknowledgements
This research is supported by the James P. Allison Institute Assistant Member Fund (S.G.), the MD Anderson Physician Scientist Award (S.G.), and NIH-R01836 Merit Award (R37 CA279192-01) (S.G.). We thank the MDACC Functional Genomics Core for generating the sgScramble and sgKdm6a cell lines (Cancer Center Support Grant P30CA016672), MDACC Research Histology Core Laboratory (CCSG P30CA016672) for the immunohistochemistry assay, MDACC Advanced Microscopy core (NIH grant S10 RR029552) for the comet assay and gamma-H2AX assay imaging services and the MDACC Metabolomics core (CCSG Grant NIH P30CA016672) for the Seahorse XFe96 Flux Analyzer service. We acknowledge the MDACC Proteomics core (UTMDACC NIH grants S10OD012304-01) for the LC-MS/MS study and the MDACC ATGC core (grant CA016672) for sequencing studies. S.G. and P. S are James P. Allison Institute members.
Author information
Authors and Affiliations
Contributions
P. Singh, R.D, B.C, S.M and D.R. designed and performed the experiments, analyzed the data, and wrote the manuscript. A.J.T, A.M, J.H and Y.L assisted in in vitro experiments. Y.X and P.L. performed the LC-MS/MS analysis. B.L, P.P, P.S, K.A and A.K.J provided scientific input. S.G. developed the project, designed experiments, analyzed data, wrote the manuscript and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, P., D’Rozario, R., Chakraborty, B. et al. Loss of KDM6A-mediated genomic instability and metabolic reprogramming regulates response to therapeutic perturbations in bladder cancer. Nat Commun 17, 1382 (2026). https://doi.org/10.1038/s41467-025-68132-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68132-2