Abstract
Protein kinases contribute to hepatocellular carcinoma (HCC) development and immune evasion, posing major challenges for HCC management. Here we show STE20/SPS1-related proline/alanine-rich kinase (SPAK) as a candidate immune exhaustion–associated gene identified through a pooled screen of protein kinases. By integrating bioinformatic analyses, data from patient cohorts, and functional studies in mouse models and cell lines, we demonstrate that elevated expression of SPAK promotes HCC progression, enhances stemness, drives immune exhaustion, and contributes to resistance to targeted therapies. Mechanistically, SPAK phosphorylates GSK3β at Ser9, thereby inhibiting proteasome-mediated degradation of c-Jun and PD-L1. Additionally, we find that DNMT3B-dependent intragenic methylation of SPAK contributes to its high expression in HCC. Notably, the SPAK inhibitor exhibits potent inhibitory effects and synergizes with PD-1 blockade to enhance antitumor efficacy. In summary, these findings establish SPAK as a driver of oncogenesis and immune exhaustion in HCC and highlight dual inhibition as a potential therapeutic strategy.
Similar content being viewed by others
Introduction
Liver cancer is the third leading cause of cancer-related mortality, with hepatocellular carcinoma (HCC) accounting for approximately 90% of cases1,2. Over the past decade, immunotherapy has revolutionized HCC treatment, particularly through anti-PD-(L)1 therapies, which have benefited patients with advanced disease3,4,5. However, responses to these therapies vary among HCC patients, underscoring the need to identify predictive biomarkers and develop precision medicine strategies tailored to distinct tumor immune microenvironments (TIME).
A comprehensive understanding of the antitumor immune response and the interactions between HCC cells and their TIME is critical for biomarker discovery and the development of combination immunotherapies6. Previous studies have classified HCCs into inflamed and non-inflamed subtypes based on immune microenvironment characteristics. Inflamed HCCs are generally responsive to anti-PD-(L)1 monotherapy and combination therapies7,8,9,10. In contrast, non-inflamed HCCs can be further categorized into intermediate and immune-exclusion subtypes, with β-catenin signaling being the primary driver of immune exclusion11. Our previous study identified MMP9 as a promising therapeutic target for overcoming immune exclusion in HCC, as MMP9-mediated shedding of slingshot protein phosphatase 1 from CD8⁺ T cells impaired cytotoxic T-cell activation12,13. Inflamed HCCs can be further subdivided into immune-active, immune-exhausted, and immune-like subtypes. Notably, the immune-exhausted subtype is characterized by severe T-cell dysfunction, impaired cytotoxic activity, and poor prognosis, highlighting the urgent need to investigate its underlying mechanisms and therapeutic targets.
Protein kinases (PKs) play a pivotal role in HCC progression and immune evasion. Several receptor tyrosine kinases, including FGFR4, MET, and PTK7, have been shown to promote HCC growth and induce an immunosuppressive TIME by modulating PD-L1 stability or TGF-β signaling14,15,16,17. Additionally, inhibition of serine/threonine kinases such as AURKA, PKCα, and PIM2 has been reported to enhance the efficacy of anti-PD-(L)1 therapy in HCC18,19,20. Despite the presence of 514 eukaryotic PKs, the roles of most PKs in HCC remain unexplored.
In this study, we screen PKs based on high expression levels, association with poor prognosis, and enrichment in the immune-exhausted subtype using HCC databases. We identify STE20/SPS1-Related Proline-Alanine-Rich Protein Kinase (SPAK) as an immune-exhaustion-associated protein and a predictive biomarker of poor prognosis in HCC. Functionally, SPAK promotes tumorigenesis, stemness, and immune exhaustion, rendering high-SPAK-expressing HCCs resistant to sorafenib, lenvatinib, and donafenib. Mechanistically, SPAK inactivates GSK3β, leading to increased stability of PD-L1 and c-Jun. Furthermore, combined targeting of SPAK and PD-(L)1 exhibits a promising therapeutic effect in immune-exhausted HCCs with high SPAK expression.
Results
High SPAK expression is associated with poor prognosis and immune exhaustion in HCC
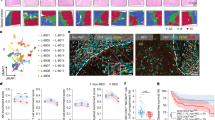
To identify protein kinases associated with immune exhaustion in HCC, we compiled a list of 514 human protein kinases from The Human Kinome21 and analyzed their expression levels in HCC and adjacent noncancerous tissues using TCGA and ICGC datasets. Differential gene expression analysis identified 47 differentially expressed kinase genes in the TCGA cohort and 18 in the ICGC cohort, with 14 overlapping kinases highly expressed in HCC (Fig. 1a and Supplementary Fig. 1a). We then examined the immune profiles of HCC samples from these cohorts based on Sia’s immune classification of HCC and assessed the expression of these 14 kinases across three immune subtypes7,8. SPAK and PTK7 emerged as differentially expressed genes (Fig. 1a). Given that immune exhaustion is characterized by T cell dysfunction, we performed a correlation analysis between kinase expression and T cell exhaustion scores, finding that SPAK exhibited the strongest correlation in both the TCGA (Pearson’s r = 0.40 vs 0.17 for PTK7, respectively) and ICGC (Pearson’s r = 0.37 vs 0.26) cohorts (Fig. 1b)22. PTK7 has been reported to regulate immune exhaustion-associated TGF-β or PD-1 signaling17, whereas no studies have reported a role for SPAK in immune evasion, particularly in HCC. Thus, we focused on investigating SPAK’s role in HCC.
a A differential expression analysis of 514 protein kinases was performed on the TCGA-LIHC (top) and ICGC-LIRI (bottom) datasets. A Venn diagram was used to visualize the overlap of upregulated genes between the two datasets. HCC samples were classified into three subtypes using NTP, and the expression of 14 genes was compared across these TME subtypes. b The correlation between T cell exhaustion scores and the expression of SPAK and PTK7 was analyzed. c wide-range data mining was employed to examine differences in SPAK mRNA expression between HCC tumor and adjacent non-tumor tissues. d IHC analysis was conducted on SPAK protein expression in a TMA containing HCC cases (n = 119 patients, scale bars, 400 μm (L), 100 μm (R)). e SPAK mRNA expression was quantified by qRT-PCR in paired fresh HCC tissue samples (n = 116 patients). f SPAK protein expression was quantified by immuneblotting in paired fresh HCC tissue samples (n = 144 patients). g Kaplan-Meier survival analysis was performed to investigate the relationship between SPAK expression and OS (n = 107 patients) as well as DFS (n = 111 patients) in HCC patients. h A forest plot was used to display the results of a multivariate analysis identifying factors associated with OS and DFS; a P values were calculated using Empirical Bayes moderated linear model and Negative binomial generalized linear model. b P values were calculated using a Spearman correlation analysis (two-tailed). d, f P values were calculated using a paired t test (two-tailed). g P values were calculated using a Kaplan–Meier test (two-tailed). h P values were calculated using a Partial Likelihood Estimation. Source data are provided as a Source Data file.
We next examined the expression pattern and functional role of SPAK in HCC. Using Sia’s inflamed HCC classification in TCGA-LIHC and ICGC cohorts, we found that SPAK expression was significantly elevated in the immune-exhausted subgroup compared to other immune-associated subgroups (Supplementary Fig. 1b, c). Consistently, mIF staining of patient samples (from Tongji cohort 1) also revealed elevated SPAK expression in tumors classified as immune-exhausted subtype, further validating this subtype-specific enrichment at the protein level (Supplementary Fig. 1d). Analyzing the TCGA PanCancer cohort revealed that SPAK was highly expressed in most cancers (Supplementary Fig. 2a). A comprehensive analysis of 92 HCC cohorts showed that 55 cohorts reported significantly increased SPAK expression in HCC tissues (Fig. 1c and Supplementary Fig. 2b). Consistently, in three independent Tongji HCC cohorts (CCGLC-002 study), SPAK was highly expressed in cancer tissues at both mRNA and protein levels (Fig. 1d–f, and Supplementary Fig. 3a). Similar findings were observed in the CPTAC proteomic cohort and TCGA-LIHC matched tumor and non-tumor tissues (Supplementary Fig. 3b, c). Further analysis of liver disease datasets (GSE9843, GSE6764, GSE114564, and GSE89377) demonstrated a gradual increase in SPAK expression from liver cirrhosis to early- and late-stage liver cancer (Supplementary Fig. 3d–g). Additionally, SPAK expression positively correlated with tumor histological grade in human clinical HCC specimens, with high SPAK expression associated with poorly differentiated tumors (Supplementary Fig. 4a, b). SPAK was also highly expressed in TCGA-iCluster1 and Hoshida S2 subtypes, which are characterized by aggressive phenotypes, activation of MAPK, AKT, and Myc signaling, and TP53 mutations23,24,25,26 (Supplementary Fig. 4c, d). Survival analysis across multiple HCC cohorts (Tongji cohort 1, TCGA-LIHC, GSE14520, and HCCDB25) indicated that high SPAK expression correlated with poorer overall survival (OS) and disease-free survival (DFS) (Fig. 1g and Supplementary Fig. 5a–c). Multivariate logistic regression confirmed that high SPAK expression was an independent predictor of postoperative OS and DFS (Fig. 1h and Supplementary Fig. 5d).
To determine the primary cell type exhibiting increased SPAK expression in HCC, we examined mRNA and protein levels of SPAK in various human tissues using the Genotype-Tissue Expression tool and ProteomicsDB, revealing relatively low SPAK levels in normal liver tissue (Supplementary Fig. 6a, b). Single-cell RNA sequencing (scRNA-seq) data from the Human Protein Atlas further confirmed low SPAK expression in hepatocytes (Supplementary Fig. 7a). Spatial transcriptomic analysis (HCCDB-HRA000437) identified cancer cells as the primary source of SPAK expression in HCC (Supplementary Fig. 8a). Collectively, these findings suggest that high SPAK expression in HCC is associated with an aggressive phenotype, poor prognosis, and an immune-exhausted tumor microenvironment.
SPAK promotes tumorigenesis, stem cell properties and resistance to multikinase inhibitors of HCC
To investigate the role of SPAK in HCC, we generated mice with hepatocyte-specific SPAK deletion (Spakf/f; Alb-Cre) (Fig. 2a and Supplementary Fig. 9a–c), and established DEN/CCl4-induced HCC models. SPAK knockout in hepatocytes reduced hepatocarcinogenesis, accompanied by decreased expression of HCC markers (Fig. 2b, c and Supplementary Fig. 10a, b). To further investigate this, we generated shNChep mice and shSpakhep mice, and established hydrodynamic gene delivery-based mouse HCC models by introducing myr-AKT and N-RasV12 or MYC and sgP53, to resemble Hoshida S2 and TCGA-iCluster 1 HCC subtypes, and found similar results that SPAK depletion significantly attenuated oncogene-driven hepatocarcinogenesis, accompanied by decreased expression of HCC markers (Fig. 2d–f and Supplementary Fig. 10c–i). Consistently, in vitro CCK-8 and clone formation assays, as well as in vivo subcutaneous and orthotopic xenograft HCC models in nude mice, demonstrated that SPAK enhanced HCC growth (Supplementary Figs. 11a–e, 12a–d, and 13a–d). Furthermore, treatment with the selective SPAK inhibitor ZT-1a, which targets phosphorylation at serine 371 (S371)27, significantly reduced HCC growth (Supplementary Fig. 14a, b). We then generated HCC cells with SPAK S371A mutant and demonstrated that wild-type SPAK promoted orthotopic tumor growth, while the S371A mutant mitigated these effects (Supplementary Fig. 14c, d).
a Flow chart of the DEN/CCl4-induced HCC mouse model. b, c Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 1000 μm (L), 100 μm (R)), representative immunostaining (scale bars, 100 μm), tumor burden (n = 12 mice), and quantification of Ki-67 (n = 4 mice) of Spakf/f; Alb-Cre and Spakf/f mice. d Flow chart of the AKT/NRasV12-induced HCC mouse model. e, f Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 1000 μm (L), 100 μm (R)), representative immunostaining (scale bars, 100 μm), tumor burden (n = 6 mice), and quantification of Ki-67 (n = 4 mice) of pT3-shSpak and pT3-empty mice. g Analysis of a human liver development single-cell atlas revealed three cell clusters: hepatoblasts (HB), fetal hepatocytes (FH), and adult hepatocytes (AH). h UMAP of CytoTRACE 2 predicted order (more to less differentiated) and SPAK relative expression. i SPAK, hepatic progenitor and mature hepatocyte markers expression in HB/FH and AH clusters. j Correlation analysis of SPAK with stemness-related signatures. k, l Sphere formation assays to evaluate the impact of SPAK on the self-renewal capacity of HCC cells (n = 3 independent experiments, scale bar, 100 μm). m Extreme limiting dilution analysis: SPAK knockdown HCC cells were subcutaneously injected into 4-week-old male NUDE mice (n = 6 mice/group, scale bar, 5 mm). Stem cell frequency was analyzed; c, f, i, k P values were calculated using an unpaired t test (two-tailed). j P values were calculated using a Pearson correlation analysis (two-tailed). The box plots show data distribution with the overlaid box plot indicating the median, first (Q1) and third (Q3) quartiles, and the whiskers extending to the 1.5× interquartile range (IQR). Data are presented as the mean ± SD. Source data are provided as a Source Data file.
Given that liver cancer stem cells (LCSCs) were enriched in Hoshida S2 and TCGA-iCluster 1 subtypes25,26, we explore the role of SPAK in regulating stem cell properties in HCC. Correlation analyses using the HCCDB database revealed a positive association between SPAK and cancer stem cell (CSC) markers, including EpCAM, PROM1 (CD133), THY1 (CD90), SOX9, CD47, ICAM1, and KLF4, and a negative correlation with hepatic lineage genes such as HNF4A and ALB (Supplementary Fig. 15a). Consistently, EpCAM+ HCCs exhibited higher SPAK expression than EpCAM−HCCs (Supplementary Fig. 15b). To validate these findings in a human context, we analyzed a previously published scRNA-seq dataset profiling human liver development28. Cell clusters were categorized into hepatoblast (HB), fetal hepatocyte (FH), and adult hepatocyte (AH) (Fig. 2g). Using CytoTRACE29,30, we observed a progressive decline in developmental potential from HB to FH to AH. SPAK expression was enriched in the less differentiated HB/FH clusters and reduced in the mature AH cluster, and correlated with progenitor-associated but not mature hepatocyte-associated markers (Fig. 2h, i). These results suggest that SPAK is preferentially expressed in less differentiated hepatic cells. Similarly, analysis of previously published single-cell HCC datasets (GSE202642, GSE151530, GSE206325, GSE149614, GSE156625) revealed a subset of malignant hepatocytes with high SPAK expression that exhibited the greatest developmental potential (Supplementary Fig. 15c). ssGSEA analysis of the TCGA-LIHC cohort further confirmed enrichment of stem cell signatures in tumors with high SPAK expression (Fig. 2j). Overexpression of SPAK in HCC cells led to increased expression of CSC-associated markers, including EpCAM, PROM1, and SOX9, a pattern that was also observed in vivo (Supplementary Fig. 15d–g). Sphere formation assays demonstrated that SPAK overexpression enhanced spheroid-forming efficiency, whereas its depletion significantly impaired this ability (Fig. 2k, l). A limiting dilution assay in nude mice revealed that SPAK knockdown markedly reduced tumor-initiating potential and stem cell frequency (Fig. 2m).
LCSCs were reported as characteristics of drug resistance. Among first-line systemic therapies for treating advanced HCC, sorafenib and lenvatinib are approved by the US Food and Drug Administration (FDA) and National Medical Products Administration (NMPA), and donafenib is a deuterated form of sorafenib approved by NMPA. To assess whether SPAK expression affects sensitivity to these multikinase inhibitors, we conducted drug sensitivity assays and found that SPAK overexpression significantly increased resistance to sorafenib, lenvatinib, and donafenib, whereas SPAK silencing restored sensitivity (Supplementary Fig. 15h, i). Analysis of 17 HCC datasets from lenvatinib- or sorafenib-resistant cell lines, xenografts, tissues, and organoids showed that SPAK mRNA levels were significantly elevated in 76% (13/17) resistant groups (Supplementary Fig. 15j). Taken together, these data indicate that SPAK promotes hepatocarcinogenesis, growth, stem cell properties, and resistance to sorafenib, lenvatinib and donafenib in HCC.
SPAK promotes exhaustion and inhibits the cytotoxic effect of cytotoxic T cells in HCC
Building on our previous bioinformatic analyses, which revealed a significant correlation between SPAK expression and immune exhaustion in HCC (Supplementary Fig. 1a, b), we sought to investigate the underlying mechanisms driving this association. Therefore, we established subcutaneous xenograft tumor models using Hepa1-6 cells in both immunodeficient nude mice and immunocompetent C57BL/6 mice. Notably, SPAK overexpression moderately promoted tumor growth in nude mice, whereas this effect was substantially enhanced in immunocompetent C57BL/6 mice (Supplementary Fig. 16a–d), indicating a potential role of the immune system. This finding prompted further investigation into the underlying mechanisms, particularly regarding immune modulation.
To further explore the role of SPAK in the HCC immune microenvironment, we performed sample stratification based on tumor SPAK expression, dividing patients into high- and low-SPAK expression groups, and analyzed publicly available single-cell RNA-sequencing data from previously published studies (Supplementary Fig. 17a, b)31. Four distinct CD8+ T subpopulations were characterized, including proliferating CD8+ T cells, effectors_like CD8+ T cells, exhausted_like CD8+ T cells, and naive/activation CD8+ T cells. Notably, HCC cells with high expression of SPAK showed reduced subpopulation of effectors_like CD8+ T cells and increased the exhausted_like CD8+ T cells (Fig. 3a and Supplementary Fig. 17c). In addition, we examined the correlation between SPAK levels and immune-related signatures in TCGA-LIHC and ICGC-LIRI-JP cohorts22,32,33 and found that HCC with high SPAK expression were enriched by exhausted cytotoxic T cells, and high levels of exhausted immune checkpoints (Supplementary Fig. 17d, e).
a tSNE plots illustrating the distinct CD8⁺ T cell clusters from single-cell RNA-sequencing data of HCC (Mendeley Data: skrx2fz79n/1), along with the distribution of these clusters in patients with high/low SPAK expression. b Flow cytometry analysis of GZMB+IFN-γ+CD8+ T cells and PD-1+TIM-3+CD8+ T cells in AKT/NRasV12-induced HCC (n = 6 mice). c Flow cytometry analysis of GZMB+IFN-γ+CD8+ T cells and PD-1+TIM-3+CD8+ T cells in Hepa1-6-induced HCC (n = 6 mice). d, e Representative mIF staining images of CD8, Granzyme B, PD-1, and TIM-3 in the indicated mouse HCC tumor tissues (scale bars, 100 μm). f, g Quantification of GZMB+CD8+ T and PD-1+TIM-3+CD8+ T cells in indicated mouse HCC model (n = 4 mice). h Diagram illustrating the layout of the coculture experiment. i A representative histogram showing the proportion of Annexin V–positive Hepa1-6 cells. The percentage of Annexin V–positive cells in the indicated groups (n = 3 independent experiments) and the relative cytotoxic capacity of T cells were determined using the formula: Relative T-cell killing ability = [(SPAK group with T cells–without T cells) Annexin V%] / [(Control group with T cells–without T cells) Annexin V%]. j Flow cytometry analysis of GZMB+IFN-γ+CD8+ T cells and PD-1+TIM-3+CD8+ T cells in the Hepa1-6/CD8+ T-cell co-culture system (n = 3 independent experiments); b, c, f, g, i, j P values were calculated using an unpaired t test (two-tailed). The box plots show data distribution with the overlaid box plot indicating the median, first (Q1) and third (Q3) quartiles, and the whiskers extending to the 1.5x interquartile range (IQR). Data are presented as the mean ± SD. Source data are provided as a Source Data file.
We then analyzed tumor tissues from the hydrodynamic-induced HCC model and hepatic orthotopic transplant model by multicolor flow cytometry (Supplementary Fig. 18a). SPAK knockdown resulted in a slight increase in cytotoxic CD8+ Tcells abundance, as indicated by granzyme B and interferon-γ levels, whereas exhausted CD8+ T cells, marked by PD-1 and TIM-3, were significantly decreased (Fig. 3b). Conversely, SPAK overexpression led to a reduction in cytotoxic CD8+ T cells and an increase in exhausted CD8+ T cells, while overexpression of a kinase-inactive mutant (S371A) reversed these effects (Fig. 3c). These findings were further corroborated by multiplex immunofluorescence in tumor tissues from the hydrodynamic injection liver cancer model, DEN/CCl4-induced HCC model and hepatic orthotopic transplant model (Fig. 3d–g, and Supplementary Fig. 19a, b). To evaluate SPAK’s effect on T-cell-mediated cytotoxicity, we established a co-culture system of Hepa1-6 cells and CD8+ T cells (Fig. 3h). Compared with control Hepa1-6 cells (Hepa1-6-Control), Hepa1-6 cells overexpressing SPAK (Hepa1-6-SPAK) were significantly more resistant to T-cell mediated killing (Fig. 3i). Moreover, SPAK overexpression promoted CD8⁺ T cell exhaustion and impaired cytotoxic function, as indicated by elevated expression of exhaustion markers (PD-1 and TIM-3) and reduced levels of cytolytic molecules (granzyme B and interferon-γ) in CD8⁺ T cells within the co-culture system (Fig. 3j). Given the high expression of SPAK in hepatic T cells, ex vivo activated mouse CD8⁺ T cells were treated with the SPAK inhibitor ZT-1a, which did not significantly affect their cytotoxicity or exhaustion status (Supplementary Fig. 19c, d). Collectively, these findings demonstrate that SPAK induces T cell exhaustion and impairs the cytotoxic activity of cytotoxic T cells in HCC.
SPAK deactivates GSK3β and stabilizes c-Jun to facilitate HCC progression and resistance to multikinase inhibitors
To explore the underlying mechanism of SPAK in regulating the tumorigenesis and immune exhaustion of HCC, we performed phospho-kinase arrays to determine the downstream signaling cascades. Following SPAK overexpression, we found that among the top five phosphorylated proteins ranked by mean pixel density ratio, GSK3β at Serine 9 (p-GSK3βS9) exhibited the highest phosphorylation level. Additionally, phosphorylated c-Jun at Serine 63 (p-c-JunS63) was elevated, consistent with c-Jun acting as a direct downstream target of GSK3β34 (Fig. 4a, b, Supplementary Fig. 20a, and Supplementary Dataset 1). Consistently, the high SPAK expression group exhibited marked enrichment of the GSK3 and Jun pathways (Fig. 4c). Proteomic analysis of CPTAC-HCC patient samples further revealed a positive correlation between SPAK and c-Jun protein levels (Fig. 4d), and validation analyses of western blotting demonstrated that SPAK overexpression increased the levels of c-Jun, p-c-JunS63, and p-GSK3βS9 with total GSK3β levels unchanged. In contrast, phosphorylation of c-Jun at Ser243 (p-c-JunS243), a site phosphorylated by GSK3β that facilitates c-Jun ubiquitination and degradation34, was significantly reduced upon SPAK overexpression. Treatment with ZT-1a or expression of the S371A-mutant SPAK reversed this effect (Fig. 4e and Supplementary Fig. 20b). Conversely, SPAK knockdown decreased the protein levels of c-Jun, p-c-JunS63, and p-GSK3βS9, while increasing p-c-JunS243 levels (Fig. 4f and Supplementary Fig. 20c). IHC analysis of HCC tissues showed that SPAK knockout or knockdown lowered p-GSK3βS9 and c-Jun expression, while SPAK overexpression increased their levels; this effect was abolished by the S371A mutant (Supplementary Fig. 20d–g). To further confirm the regulatory effects of SPAK in GSK3β and c-Jun signaling, we generated a SPAK knockout HLF cell line using CRISPR‒Cas9 system (Fig. 4g and Supplementary Fig. 20h). Overexpression of SPAK in two independent HLF-sgSPAK clones restored the levels of c-Jun, p-c-JunS63, and p-GSK3βS9, whereas overexpression of the S371A mutant abolished these effects (Fig. 4h and Supplementary Fig. 20i).
a Representative images of human phospho-kinase array analysis in indicated PLC/PRF/5 cells. b Relative pixel density of p-GSK3βS9 and p-c-JunS63 in indicated groups (n = 2 technical replicate). c GSEA of RNA-seq results based on the TCGA-LIHC. d Public proteomic data obtained from CPTAC were used to analyze the correlation between SPAK and c-Jun (n = 165 patients). e, f Western blot analysis of the phosphorylation level of GKS3β and c-Jun in indicated HCC cells (n = 3 independent experiments). g Scheme illustrating the design of the SPAK knock-out cell lines. h Western blot analysis of the phosphorylation level of GKS3β and c-Jun in indicated HCC cells (n = 3 independent experiments). i Photographs and tumor burden of xenograft tumors generated in nude mice following inoculation with PLC/PRF/5-control, PLC/PRF/5-SPAK and PLC/PRF/5-SPAK treated with T5224 (n = 6 mice, scale bar, 5 mm). j Self-renewal capacity was evaluated using a sphere formation assay performed with PLC/PRF/5 cells (scale bar, 100 μm) (n = 3 independent experiments). k The expression levels of c-Jun in HLF cells were measured after treatment with CHX for the specified durations (n = 3 independent experiments). l The expression levels of c-Jun in HCC cells with SPAK overexpression, both in the presence and absence of MG132 treatment (n = 3 independent experiments). m, n Ubiquitination of c-Jun in SPAK-knockdown/-overexpression HCC cells (n = 3 independent experiments); d P values were calculated using a Spearman correlation analysis (two-tailed). i, j, k P values were calculated using an unpaired t test (two-tailed). The box plots show data distribution with the overlaid box plot indicating the median, first (Q1) and third (Q3) quartiles, and the whiskers extending to the 1.5× interquartile range (IQR). Data are presented as the mean ± SD. Source data are provided as a Source Data file.
Previous studies have shown that c-Jun activation drives HCC initiation, self-renewal, and resistance to sorafenib and lenvatinib35,36. To determine the role of c-Jun in SPAK-mediated HCC progression, we conducted rescue experiments using T-5224, a c-Jun inhibitor37. Western blot analysis confirmed that T-5224 did not reduce total c-Jun protein levels, consistent with its mechanism of action as a transcriptional inhibitor (Supplementary Fig. 21a). Treatment with T-5224 reversed SPAK-induced tumor formation, sphere formation, upregulation of CSC-associated markers, and drug resistance (Fig. 4i, j and Supplementary Fig. 21b, c). Next, we investigated the molecular mechanisms underlying SPAK-mediated c-Jun and p-GSK3βS9 upregulation. Protein synthesis and degradation assays showed that SPAK promoted the stability of c-Jun (Fig. 4k and Supplementary Fig. 22a). Treatment with the proteasome inhibitor MG132 counteracted the reduction in c-Jun levels induced by SPAK knockdown, suggesting that SPAK stabilizes c-Jun via a proteasome-dependent mechanism (Fig. 4l and Supplementary Fig. 22b). Consistently, c-Jun ubiquitination was inhibited by SPAK overexpression and enhanced by SPAK knockdown (Fig. 4m, n). However, when S371A was overexpressed, SPAK-inhibited ubiquitination of c-Jun was abolished (Supplementary Fig. 22c). Previous studies have established that phosphorylation of GSK3β at Ser9 inhibits its kinase activity, reducing c-Jun phosphorylation and preventing its degradation by the ubiquitin ligase FBXW734. Based on these findings, we hypothesized that SPAK upregulates c-Jun by increasing p-GSK3βS9 levels. Treatment with lithium chloride (LiCl), a GSK3β inhibitor, restored SPAK-mediated regulation of c-Jun (Supplementary Fig. 22d). Additionally, when FBXW7 levels were reduced, SPAK was still able to regulate c-Jun (Supplementary Fig. 22e, f). Furthermore, SPAK binding to c-Jun was dependent on SPAK expression levels (Supplementary Fig. 22g–j). Taken together, these results indicate that SPAK stabilizes c-Jun by interrupting GSK3β/FBXW7-mediated ubiquitination and degradation, thereby promoting HCC progression.
SPAK deactivates GSK3β by direct binding and promoting its phosphorylation at Ser9
Given that SPAK is a serine/threonine (S/T) kinase that upregulates GSK3βS9, we speculated that SPAK directly binds to and phosphorylates GSK3β. To test this, we performed coimmunoprecipitation (co-IP) and confocal microscopic analyses, confirming the colocalization and interaction of SPAK and GSK3β in HCC cells (Fig. 5a–c and Supplementary Fig. 23a). Mutated truncation experiments revealed that amino acid regions 347-456 of SPAK interact with amino acid regions 340-434 of GSK3β (Fig. 5d–g). Structural modeling and molecular docking simulations further corroborated this interaction, revealing potential contact sites within these regions that may mediate binding (Fig. 5h). Given that GSK3β is a well-established substrate of AKT1—an upstream kinase that phosphorylates GSK3β at Ser9, thereby inhibiting its activity—we next compared the binding characteristics of SPAK and AKT1 to GSK3β. Interestingly, although AKT1 showed a slightly stronger affinity to GSK3β (KD (M) = 5.83E-09), the binding response and stability of SPAK-GSK3β complex were also notably high (KD (M) = 6.96E-09), indicating SPAK may serve as a potential functional interactor of GSK3β (Fig. 5i). Phosphorylation IP experiments confirmed that SPAK promotes the S/T phosphorylation of GSK3β (Fig. 5j), and this phosphorylation was independent of AKT1 (Supplementary Fig. 23b–e). Consistently, treatment with the AKT1 inhibitor MK2206 did not alter SPAK-induced phosphorylation of GSK3β at Ser9, further confirming that this modification occurs independently of AKT1 (Supplementary Fig. 23f). Subsequent phosphosite analysis using liquid chromatography-tandem mass spectrometry (LC-MS/MS) revealed that SPAK overexpression induced phosphorylation at serine 9 of GSK3β (Fig. 5k), and phospho-IP confirmed that SPAK-mediated S/T phosphorylation of GSK3β was abolished in cells overexpressing GSK3β serine 9 mutants (S9A) (Fig. 5l, m), and this finding was further validated by in vitro kinase assays (Fig. 5n–p). Collectively, these data suggest that SPAK directly interacts with and phosphorylates the ninth serine of GSK3β.
a, b Immunoprecipitation and western blot assays reveal the interaction between SPAK and GSK3β in HCC cells (n = 3 independent experiments). c Colocalization of SPAK and GKS3β in HCC cells detected by IF analysis (scale bars, 25 μm) (n = 3 independent experiments). d, e Development of truncations for SPAK and GKS3β. f Immunoprecipitation and western blot assays were used to demonstrate the interactions between HA-tagged truncated SPAK and FLAG-tagged GSK3β proteins in HEK293T (n = 3 independent experiments). g Immunoprecipitation and western blot assays were used to demonstrate the interactions between FLAG-tagged truncated GKS3β and HA-tagged SPAK proteins in HEK293T (n = 3 independent experiments). h The crucial amino acids for the interaction between SPAK and GKS3β were speculated using three-dimensional structures. i Surface plasmon resonance assays to evaluate and compare the relative binding affinities of AKT1 and SPAK to GSK3β. j Immunoprecipitation followed by western blot analysis was conducted to examine the phosphorylation of GSK3β in HEK293T cells transfected with FLAG-SPAK or FLAG-S371A (n = 3 independent experiments). k LC-MS/MS spectrum showing the phosphorylation of GKS3βS9. l Design of mutation of S9 sites of GKS3β. m Immunoprecipitation and western blot analyses showing the GKS3β phosphorylation in HEK293T cells transfected with FLAG-GKS3β or FLAG-S9A (n = 3 independent experiments). n Scheme displaying the procedure for kinase reaction and alkylation. o Recombinant GST-GSK3β was used in an in vitro kinase assay, where active FLAG-SPAK or FLAG-S371A proteins were incubated with GST-GSK3β to initiate the kinase reaction. S9 phosphorylation on WT GSK3β after in vitro kinase assay was determined by GSK3β-specific phospho-antibodies (n = 3 independent experiments). p S9 phosphorylation on WT GSK3β and mutant GSK3β after in vitro kinase assay was determined by GSK3β-specific phospho-antibodies (n = 3 independent experiments). Source data are provided as a Source Data file.
SPAK stabilizes PD-L1 to promote immune exhaustion of HCC
GSK3β was reported to regulate the stability of PD-L1 in cancer cells38,39,40. To explore whether SPAK regulates PD-L1 through GSK3β in HCC, we performed tissue microarray analyses of HCC and found significant positive correlations among SPAK, GSK3βS9, c-Jun and PD-L1 (Supplementary Fig. 24a, b). Notably, HCCs with high SPAK expression exhibited an immune-exhausted microenvironment, characterized by high PD-L1 expression, and low levels of GZMB+CD8+ cells (Supplementary Fig. 24c). Given that intrinsic PD-L1 expression in tumor cells has been reported to play a critical role in driving immune exhaustion in HCC15,41,42,43, analysis of previously published single-cell RNA-seq datasets further revealed that HCC cells with high CD274 expression displayed a reduced proportion of effector-like CD8+ T cells and an increased proportion of exhausted-like CD8+ T cells (Supplementary Fig. 25a). Consistently, Western blot analysis of HCC cells further confirmed that SPAK upregulated PD-L1 protein levels, an effect that was abolished by overexpression of the S371A mutant or treatment with ZT-1a (Fig. 6a–c and Supplementary Fig. 25b–g). Consistent results were obtained from flow cytometry and immunofluorescence staining of PD-L1 in HCC cells (Fig. 6d–f and Supplementary Fig. 25h). To further investigate the role of PD-L1 in SPAK-mediated immune evasion in HCC, we conducted rescue experiments using PD-L1 antibodies. Blocking PD-L1 reversed SPAK-induced reductions in cytotoxic CD8⁺ T cells, increases in exhausted CD8⁺ T cells, and tumor progression (Fig. 6g–k and Supplementary Fig. 25i). Given the weak correlation between SPAK and CD274 mRNA levels in the HCC databases, we explored post-transcriptional mechanisms of SPAK in the expression of PD-L1, and found that SPAK maintained the stability of PD-L1 in the presence of CHX (Fig. 6l, m and Supplementary Fig. 26a). Moreover, the proteasome inhibitor MG132, but not the lysosomal inhibitor chloroquine (CQ) or pan-caspase inhibitor Q-VD-OPh, reversed the SPAK-induced increase in PD-L1 levels, suggesting that SPAK regulates PD-L1 stability in a proteasome-mediated manner (Fig. 6n and Supplementary Fig. 26b). In line with this, SPAK inhibited PD-L1 ubiquitination, an effect abolished by shRNAs against SPAK, or SPAK-S371A mutant (Fig. 6o, p and Supplementary Fig. 26c), and treatment with the LiCl (GSK3β inhibitor) restored the regulatory effect of SPAK and ZT-1a on PD-L1 protein levels in HCC (Supplementary Fig. 26d, e). Previous studies have shown that β-TrCP is the ubiquitin ligase that mediates the degradation of PD-L1 by GSK3β44 thus, we hypothesized that SPAK stabilizes PD-L1 by interfering with β-TrCP-mediated degradation. Supporting this, SPAK’s regulation of PD-L1 was dependent on β-TrCP expression (Supplementary Fig. 27a, b). Furthermore, SPAK overexpression weakened the interaction between GSK3β and PD-L1, an effect reversed by S371A overexpression (Supplementary Fig. 27c, d). Conversely, SPAK knockdown enhanced the GSK3β–PD-L1 interaction (Supplementary Fig. 27e). In summary, these findings suggest that SPAK stabilizes PD-L1 by interrupting the GSK3β-/β-TrCP-mediated PD-L1 ubiquitination and degradation and thus promotes the formation of immune exhaustive microenvironment in HCC.
a PD-L1 expression levels in SPAK-overexpression HCC cells treated with or without ZT-1a (n = 3 independent experiments). b, c Western blot analysis was conducted to assess the expression levels of PD-L1 in the indicated HCC cells (n = 3 independent experiments). d PD-L1 expression on the cell surface in the indicated PLC/PRF/5 (PLC) cells was conducted using flow cytometry (n = 3 independent experiments). e PD-L1 expression on the cell surface in the indicated HLF cells was conducted using flow cytometry (n = 3 independent experiments). f Representative images of PD-L1 staining by immunofluorescence in the indicated mouse HCC tumors are shown (scale bars, 100 μm). g A schematic representation of the treatment strategy in the Hepa1-6-induced HCC model. h–k Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 100 μm), tumor burden (n = 6 mice) and flow cytometry analysis (n = 6 mice) of Granzyme B+IFN-γ+CD8+ T cells and PD-1+TIM-3+CD8+ T cells in indicated orthotopic Hepa1-6 HCC tumors. l, m The expression levels of PD-L1 in SPAK overexpression (PLC/PRF/5) and SPAK-knockdown (HLF) cells were measured after treatment with CHX for the indicated durations (n = 3 independent experiments). n The expression levels of PD-L1 in PLC/PRF/5 cells overexpressing SPAK were assessed following treatment with or without MG132, CQ and Q-VD (n = 3 independent experiments). o, p Ubiquitination of PD-L1 in SPAK-overexpression (PLC/PRF/5) and SPAK-knockdown (HLF) cells (n = 3 independent experiments); d, e, i, j, k, l, m P values were calculated using an unpaired t test (two-tailed). The box plots show data distribution with the overlaid box plot indicating the median, first (Q1) and third (Q3) quartiles, and the whiskers extending to the 1.5× interquartile range (IQR). Data are presented as the mean ± SD. Source data are provided as a Source Data file.
Intragenic hypermethylation of SPAK contributed to its high expression in HCC
To investigate the mechanisms underlying high SPAK expression in HCC, we first analyzed the frequency of SPAK alterations in the TCGA-LIHC cohort, and found only three patients with SPAK mutations (V206L, R353I, and G144) and low-level gains (roughly four copies) of SPAK allele were detected in the TCGA-LIHC cohort of 366 patients (Supplementary Fig. 28a, b). Subsequently, we explored the role of epigenetic modifications, particularly DNA methylation, in regulating SPAK expression. Analysis of TCGA-LIHC methylome data revealed 37 CpG sites within the SPAK gene: six in the promoter region, two in the first exon, and 29 in the gene body (Supplementary Fig. 28c). Notably, the methylation levels of CpG sites within the gene body were significantly higher than those in the promoter region (Fig. 7a). Recent studies reported that gene body methylation in cancer cells can promote active transcription45,46,47. Furthermore, we observed a positive correlation between the overall methylation level of the SPAK gene and its expression (Supplementary Fig. 28d). We also found that SPAK is highly expressed in HCCs with high CpG island methylator phenotype (CIMP-H) subtype, which is characterized by extensive CpG island methylation and poor prognosis, as previously described by our group (Supplementary Fig. 28e)48. When analyzing individual CpG sites, most gene body methylation sites showed a strong positive correlation with SPAK expression, whereas a few promoter region sites exhibited a negative correlation (Fig. 7a). Additionally, the majority of CpG sites in the gene body were associated with poor prognosis in HCC patients (Fig. 7b).
a The correlation between the methylation level of the CpG sites of SPAK gene and its expression level. b Most of the CpG sites in the gene body are correlated with the overall survival (OS) of HCC patients, while only one site in the promoter region is related to OS (TCGA-LIHC, n = 371 patients). c SPAK protein levels in HCC cells treated with 5-aza-dC for the indicated concentrations (n = 3 independent experiments). d SPAK mRNA levels in HCC cells treated with 5-aza-dC for the indicated concentrations (n = 3 independent experiments). e SPAK protein levels in HCC cells treated with DNMTs inhibitors (n = 3 independent experiments). f, g SPAK mRNA levels in HCC cells treated with DNMTs inhibitors (n = 3 independent experiments). h SPAK protein levels in indicated HCC cells treated with or without si-DNMT3B transfection (n = 3 independent experiments). i SPAK mRNA levels in indicated HCC cells treated with or without si-DNMT3B transfection (n = 3 independent experiments). j Tracks showing the DNMT3B and H3K36me3 occupancy at gene body region of SPAK based on analysis of ChIP seq from the ENCODE (ENCSR156CWW and ENCSR000AMB). k Tracks showing the DNMT3B occupancy at gene body region of SPAK based on analysis of CUT&Tag sequencing data performed in PLC/PRF/5 and HLF cells; a P values were calculated using a Pearson correlation analysis (two-tailed). b P values were calculated using a Cox proportional hazards regression model. d, f, g, i P values were calculated using an unpaired t test (two-tailed). Data are presented as the mean ± SD. Source data are provided as a Source Data file.
To further determine whether SPAK expression is regulated by its gene body methylation, we treated HCC cell lines (PLC/PRF/5 and HLF) with DNA methyltransferase inhibitor 5-aza-2’-deoxycytidine (5-aza-dC) and found this treatment significantly reduced both mRNA and protein levels of SPAK in HCC cells (Fig. 7c, d). We then investigated specific DNA methyltransferases (DNMTs) involved in this process, and found that Nanaomycin A, a DNMT3B inhibitor, significantly reduced SPAK expression in PLC/PRF/5 and HLF cells, whereas DNMT1 and DNMT3A inhibitors (GSK-3685032 and DNMT3A-IN-1, respectively) had no effect (Fig. 7e–g). Furthermore, DNMT3B knockdown via shRNA transfection also downregulated SPAK expression in HCC cell lines (Fig. 7h, i). We analyzed the co-expression of SPAK and DNMT3B in 92 HCC datasets and found a positive correlation across multiple datasets (Supplementary Fig. 29a), indicating that DNMT3B plays a key role in SPAK gene body methylation. Previous studies have shown that H3K36me3 facilitates DNMT3B binding and promotes de novo DNA methylation within actively transcribed gene bodies in cancer cells45,49. By examining the ChIP-seq profiles of hepatoma cells (HepG2) in the ENCODE, we found that H3K36me3 and DNMT3B are enriched on the SPAK gene body (Fig. 7j). To further confirm this, genome-wide CUT&Tag analysis demonstrated significant overlap of DNMT3B peaks on the SPAK gene body in HLF and PLC/PRF/5 cells (Fig. 7k and Supplementary Fig. 29b). Taken together, these findings suggest that DNMT3B-dependent hypermethylation of the SPAK gene body promotes its expression in HCC.
Combined targeting SPAK and PD-1 showed enhanced efficacy in HCC
To further validate the upregulation of SPAK and its association with the immune therapy response in HCC, we analyzed SPAK expression levels in a spatial transcriptome dataset. Our results showed that SPAK was highly expressed in HCC cells from responders to anti-PD-1 therapy but weakly expressed in adjacent normal tissues and non-responders (Fig. 8a)31. Similar findings were observed in another dataset of HCC patients who received immunotherapy (Supplementary Fig. 30a). Additionally, in a retrospective analysis of a prospective cohort of patients undergoing anti-PD-1 neoadjuvant therapy (Tongji cohort 4), SPAK expression was significantly higher in samples from responders (Fig. 8b). These results uncovered SPAK level as a potential predictor of response to ICIs in HCC.
a Representative tissue region maps and expression analysis of SPAK levels from a spatial transcriptome dataset of ICI responders and non-responders (Mendeley Data: skrx2fz79n). b Representative abdominal MRI images, representative IHC staining and quantitative IHC analysis of SPAK levels performed on human HCC tissue obtained before and after treatment of ICI responders (n = 7 patients) and non-responders (n = 12 patients) (scale bars, 100 μm). c A schematic representation of the treatment strategy in the DEN/CCl4-induced HCC model. d, e Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 1000 μm), and tumor burden (n = 5 mice) from the indicated DEN/CCl4-induced HCC model. f A schematic representation of the treatment strategy in the AKT/NRasV12-induced HCC model. g, h Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 1000 μm), and tumor burden (n = 5 mice) from the indicated AKT/NRasV12-induced HCC model. i A schematic representation of the treatment strategy in the Hepa1-6-induced HCC model. j, k Gross images of livers (scale bars, 5 mm), representative H&E staining (scale bars, 200 μm), and tumor burden (n = 6 mice) from the indicated Hepa1-6-induced HCC model. l, m Total flux curve and bioluminescence images of mice from the indicated Hepa1-6-induced HCC model for the indicated days (n = 6 mice); a, b, e, h, k, m P values were calculated using an unpaired t test (two-tailed). The box plots show data distribution with the overlaid box plot indicating the median, first (Q1) and third (Q3) quartiles, and the whiskers extending to the 1.5× interquartile range (IQR). Source data are provided as a Source Data file.
Inspired by these observations, we investigated whether improving the efficacy of anti-PD-1 therapy in HCC could be achieved by targeting the immunosuppressive interactions between tumor cells and CD8+ T cells through SPAK inhibition. To this end, we established preclinical HCC mouse models using DEN induction, hydrodynamic injection of myrAKT and N-RasV12, and orthotopic xenograft models with Hepa1-6-luc cells. We then administered anti-PD-1 antibodies, the SPAK inhibitor ZT-1a, or a combination of both. Notably, the combination therapy significantly enhanced and prolonged antitumor responses, as evidenced by gross morphology, luciferase flux, H&E staining, liver weight, and the liver-to-body weight ratio (Fig. 8c–m). Together, our study highlights the critical role of SPAK in HCC progression and identifies a key signaling axis that can be pharmacologically targeted, offering a promising avenue for combination therapy in HCC patients.
Discussion
SPAK belongs to the GCK-VI subfamily of the STE20 kinase family50,51 and has been implicated in promoting the proliferation and metastasis of various cancers, including NSCLC, cholangiocarcinoma, breast cancer and HCC52,53,54,55. However, its role in oncogenic processes beyond proliferation and metastasis remains largely unexplored. Recent studies suggest that SPAK facilitates homologous recombination repair following DNA damage in pancreatic cancer56. In this study, we demonstrated that SPAK promotes the tumorigenesis, stem cell characteristics, and resistance to tyrosine kinase inhibitors (TKIs) including sorafenib, lenvatinib and donafenib of HCC by stabilizing c-Jun. The AP-1 transcription factor c-Jun has been implicated in HCC tumorigenesis, stemness, and TKIs resistance35,36,57,58,59. Our findings demonstrate that SPAK enhances TKI resistance by regulating c-Jun through GSK3β. Specifically, SPAK binds to and phosphorylates GSK3β at serine 9, inhibiting the E3 ubiquitin ligase FBXW7 from mediating c-Jun degradation. These findings elucidate a crucial molecular mechanism by which SPAK drives both HCC progression and therapeutic resistance. Furthermore, our data suggest that high SPAK expression may serve as a biomarker for sorafenib and lenvatinib resistance in HCC, highlighting the need for biomarker-guided clinical trials.
Notably, we provide evidence that SPAK contributes to an immune-exhaustive tumor microenvironment by stabilizing PD-L1 in HCC. We found that SPAK interacts with and phosphorylates GSK3β, thereby inhibiting PD-L1 degradation. Consistently, in murine immunocompetent HCC models, anti-PD-(L)1 antibodies showed antitumor effects on SPAK-expressing HCCs. Moreover, HCC patients with high SPAK expression demonstrated enhanced sensitivity to anti-PD-(L)1 based immunotherapy. These results suggest that SPAK-stabilized PD-L1 contributed to the maintenance of exhaustive TIME in HCC. Among the currently approved first-line systemic therapies for HCC, anti-PD-(L)1 therapies, rather than sorafenib, lenvatinib, or donafenib, may be more effective for patients with high SPAK expression60.
Our study reveals that SPAK promotes HCC progression by stabilizing both c-Jun and PD-L1. Although PD-L1 blockade alone elicited significant anti-tumor effects in the SPAK-overexpressing model, these results may be context-dependent. Considering that SPAK also activates additional oncogenic pathways, including c-Jun, we propose that direct SPAK inhibition, either alone or in combination with PD-(L)1 blockade, could offer a more effective therapeutic approach, particularly for HCC tumors with high SPAK expression. To test this hypothesis, we evaluated the efficacy of SPAK kinase inhibitors in preclinical HCC models. A recent study identified ZT-1a as a potent and selective inhibitor of SPAK kinase activity by suppressing its phosphorylation at serine 37127. In this study, we demonstrate that ZT-1a showed inhibitory effects on the progression of HCC in preclinical rodent HCC models, and when combined with a PD-(L)1 antibodies, ZT-1a achieves synergistic therapeutic effects. Consistently, a recent proteomics study investigating various post-translational modification events in HCC indicates that SPAK-S371 phosphorylation is notably elevated in cancerous tissues61. In addition, an integrative proteogenomics study of analyzing CPTAC data across 10 cancer types (HCC was not included) identified SPAK-S371 as a potential targetable hyperactivation event in clear cell renal cell carcinoma and lung adenocarcinoma (LUAD)62. Together, these findings suggest that targeting SPAK kinase activity represents a promising therapeutic approach for HCC and warrants further investigation in a pan-cancer context.
Methylation of CpG islands (CGIs) in gene promoter regions results in gene silencing in cancer s63, whereas gene body methylation—more abundant across the genome—may be associated with increased or reduced gene expression, or alternative transcript products64. Our previous studies demonstrated that deficiency of tumor suppressors due to promoter methylation promotes the progression of HCC, and we identified of the CIMP in HCC and found that HCC with CIMP-High showed poor prognosis and immunosuppressive phenotypes48. While prior research has primarily focused on promoter methylation, the role of gene body methylation in HCC remains poorly understood. This study provides evidence linking increased SPAK expression to the methylation of its gene body. Previous research has suggested that such epigenetic modifications play a role in the regulation of oncogene activity. Based on these findings, we propose that the hypermethylation of the gene body could act as a potential predictor, although not the exclusive factor, for elevated SPAK expression in HCC. In addition, in this study, we demonstrated that high levels of SPAK, either detected by mRNA, immunostaining, or specific hypermethylated intragenic CGIs from tumor tissue, could predict poor prognosis of HCC. One drawback of biopsy analysis, whether based on DNA or RNA content assessment or immunostaining techniques, is the inherent variability in sampling. In contrast, liquid biopsy offers a promising solution to the challenges associated with tissue biopsy, as it is expected to provide a more comprehensive molecular profile of a patient’s cancer. Future research will investigate the potential of utilizing intragenic hypermethylation of SPAK in circulating cell-free DNA as a prognostic biomarker for HCC with poor outcomes.
Previous studies discovered that DNMTs, including DNMT1, 3A, and 3B, recruited to the methylated intragenic DNA by distinct mechanisms, permit transcription of associated genes49,65,66. In this study, after excluding the contribution of DNMT1 and 3A to the expression of SPAK, we found that H3K36me3 and DNMT3B associate with methylated DNA in the gene body and enhance the expression of SPAK. A previous study reported that SPAK could be induced by SP-1 in HCC53. Taken together, these results suggested that SPAK was induced by both transcriptional factors and epigenetic mechanisms in HCC. Besides regulating transcription levels of genes, another consequence of intragenic methylation is to produce methylation-regulated, tissue- and cell type-specific alternative transcripts67, and N-terminally truncated alternative SPAK transcripts were reported in the nephron, with very low expression seen in other tissues, including the liver68. It is interesting to explore the existence, role and regulatory mechanism of alternative transcripts of SPAK in HCC.
This study has several limitations. First, considering SPAK is a serine/threonine kinase, we employed a Human Phospho-Kinase Array Kit to identify its downstream effectors in HCC, whereas molecules not represented on the array were not examined. Secondly, the potential kinase-independent effect of SPAK in HCC was not explored. Thirdly, although previous studies have shown that WNT1 promotes HCC progression via the SPAK/NKCC1 axis69, our study did not further investigate the potential connection between this pathway and the SPAK/p-GSK3βSer9/c-Jun/PD-L1 signaling cascade. Fourth, the relevance of SPAK-mediated GSK3β phosphorylation may depend on cellular context, including the level of AKT activation, which varies among HCC tumors. In tumors with strong AKT activation, AKT may dominate GSK3β phosphorylation, whereas SPAK-dependent regulation may be more pronounced in tumors with lower AKT activity. Lastly, our data suggest that SPAK may promote immune evasion in part through PD-L1–dependent mechanisms, while additional pathways contributing to immune resistance cannot be excluded. Finally, we performed preclinical models and demonstrated the antitumor effects of the SPAK kinase inhibitor (ZT-1a), either in single use or in combination with anti-PD-(L)1 antibodies, in HCC. However, currently, there is no SPAK kinase inhibitor that has been approved by the FDA or NMPA for cancer treatment, and further clinical trials are necessary to validate the efficacy and safety of therapeutic strategies targeting SPAK in HCC.
In summary, our study provides insights into the role of SPAK in the tumorigenesis, stem cell properties, resistance to TKIs and immune exhaustion of HCC (Fig. 9). Additionally, we identified a mechanism by which SPAK regulates c-Jun and PD-L1 stability via phosphorylate and deactivate GSK3β. Furthermore, we carried out preclinical investigations to confirm the effects of the SPAK kinase inhibitor in HCC. These findings highlight the potential of SPAK as a predictive biomarker for the response to approved systemic therapies, including sorafenib, lenvatinib and anti-PD-(L) 1-based therapies, in HCC patients, and suggest that targeting SPAK combined with ICB therapy offers a promising therapeutic strategy for treating HCC.
DNMT3B-mediated hypermethylation of the SPAK gene body promotes the high expression of SPAK in HCC. SPAK directly phosphorylates GSK3β and inhibits the degradation of PD-L1 and c-Jun. Overall, the combination of monoclonal antibodies targeting the immune checkpoint PD-1 with the SPAK inhibitor ZT-1a might constitute a promising therapeutic strategy for treating HCC. A schematic diagram was designed using Adobe Illustrator.
Methods
Ethics statement
The research adhered to the ethical standards of the 1975 Declaration of Helsinki and the International Conference on Harmonization’s Good Clinical Practice guidelines. Approval was granted by the Ethics Committee of Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (Wuhan, China), under approval No. TJ-IRB20210924. Clinical data were collected from HCC patients who underwent curative hepatectomy at Tongji Hospital. All participants provided written informed consent.
All animal care and research in this study were conducted in accordance with the guidelines for the care and use of experimental animals of the National Institutes of Health and approved by the ethics committee of Tongji Hospital, Huazhong University of Science and Technology (No. TJH-202106010). In accordance with the Laboratory animal—Guideline for ethical review of animal welfare (GB/T35892-2018), the committee requires that tumor burden in animals not exceed 10% of body weight and that average tumor diameter remain below 20 mm. Throughout the study, tumor sizes in all mice adhered to these guidelines. All mice used were male.
Study design
This study aimed to identify protein kinases that could be targeted to impede the progression and immune evasion of HCC. Additionally, it sought to evaluate the potential of targeting the identified protein kinase (SPAK), either alone or in combination with anti-PD-(L)1 antibodies, in preclinical HCC models. To achieve this, we performed transcriptomic and methylomic analyses, as well as clustering using nearest template prediction algorithms and immune-related signatures, applied to large-scale human HCC datasets, including bulk RNA-seq, scRNA-seq, spatial transcriptomics, and microarray data. Multiple models were used to investigate SPAK’s biological role in tumor progression and immune evasion, such as carcinogen-induced liver cancer, Sleeping Beauty transposon-mediated liver cancer, xenograft mouse models, flow cytometry, multiplex immunofluorescence, sphere assays, and in vitro coculture systems. We also evaluated the therapeutic efficacy of SPAK inhibitors combined with anti-PD-1 antibodies in murine HCC models. Microarray analysis was employed to investigate the pathological relevance of the SPAK/p-GSK3β/c-Jun&PD-L1 signaling axis. To explore the mechanisms underlying SPAK’s function, we utilized molecular and biochemical techniques, including kinase antibody arrays, in vitro protein kinase assays, CUT&Tag, and mass spectrometry. For the animal studies, mice were randomly assigned to treatment groups, with investigators blinded to group assignments. Experimental endpoints were predefined before study initiation.
Patients and specimens
This study utilized a tissue microarray consisting of 119 paired primary HCC tissue samples from patients who underwent curative resection at Tongji Hospital, Huazhong University of Science and Technology, between February 2012 and April 2014 (referred to as the Tongji cohort 1). Cohort 1 comprised 119 pairs of HCC tissues (pairs consisting of tumor tissue and adjacent tissue from the same patient). The cohort included 99 men and 20 women, with 61 patients older than 50 years and 58 patients aged 50 years or younger. Overall survival (OS) was defined as the time from treatment to death from any cause or the last recorded observation, while disease-free survival (DFS) was the period from randomization until disease recurrence or death due to HCC progression. To assess SPAK mRNA expression in HCC tissues, samples were randomly selected from an additional cohort of 116 patients who also underwent curative resection at the same hospital between February 2017 and January 2020 (Tongji cohort 2). Cohort 2 comprised 116 pairs of HCC tissues (pairs consisting of tumor tissue and adjacent tissue from the same patient). The cohort included 102 men and 14 women, with 72 patients older than 50 years and 44 patients aged 50 years or younger. To evaluate SPAK protein levels, another set of specimens was obtained from 144 patients who underwent curative resection during the same period (Tongji cohort 3). Cohort 3 comprised 144 pairs of HCC tissues (pairs consisting of tumor tissue and adjacent tissue from the same patient). The cohort included 120 men and 24 women, with 98 patients older than 50 years and 46 patients aged 50 years or younger. All HCC samples were verified by pathologists at Tongji Hospital. No local or systemic treatments were administered prior to surgery, and no anticancer therapies were given before disease recurrence after surgery. Additionally, a group of 19 HCC patients who received PD-1 monoclonal antibody therapy (tislelizumab) at Tongji Hospital between January 2020 and December 2022 was included (Tongji cohort 4). Cohort 4 comprised 19 HCC tissues. The cohort included 15 men and 4 women, with 14 patients older than 50 years and 5 patients aged 50 years or younger.
Data from all HCC patients across these cohorts were retrospectively analyzed as part of the CCGLC-002 study (Registered at clinicaltrials.gov: NCT05520801), a longitudinal cohort investigation focused on the clinicopathological features, treatment strategies, and prognosis of hepatobiliary cancer patients. This study was led by the Chinese Collaborative Group of Liver Cancer (CCGLC) and the Chinese Chapter of the International Hepato-Pancreato-Biliary Association, and conducted across Tongji Hospital’s main campus and three branches in Wuhan, China.
Statistical analysis
Statistical analyses were performed using GraphPad Prism 9 (GraphPad, CA, USA) and R software (version 4.4.0, R Foundation for Statistical Computing). Continuous variables following a normal distribution were compared using Student’s t test, while the Mann–Whitney U test was applied for non-normally distributed data. Survival analysis was conducted using the Kaplan–Meier method. Pearson’s correlation test was used to assess associations between continuous variables. A P < 0.05 was considered statistically significant. Results are presented as mean ± SD.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD062063. The CUT&Tag data generated in this study have been deposited in the Genome Sequence Archive database (https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA037512) under the accession code PRJCA037512. The RNA-seq data of the HCC cohorts in TCGA-LIHC were retrieved from the TCGA data hub (https://tcga-data.nci.nih.gov/tcga/). The ICGC-LIRI-JP was obtained from the ICGC data hub (https://dcc.icgc.org/). Transcriptomic profiles, including microarray and RNA-seq data from 92 HCC cohorts, were analyzed using the web-based Integrative HCC Gene Analysis (IHGA) platform (https://www.hccdatasph.cn/app/ihga). The human HCC methylome data were collected from the TCGA data hub (https:// cancergenome.nih.gov/). The CpG sites on the SPAK gene were obtained from the DNMIVD. The survival analysis of the CpG sites on the SPAK gene in HCC was evaluated using the MethSurv web tool (https://biit.cs.ut.ee/methsurv/). Furthermore, a correlation between the CpG methylation level and gene expression in HCC was obtained via the Shiny Methylation Analysis Resource Tool (SMART) (http://www.bioinfo-zs.com/smartapp/). The ChIP-Seqs were obtained from the ENCODE (https://www.encodeproject.org/experiments/ENCSR156CWW/ and https://www.encodeproject.org/experiments/ENCSR000AMB/) under accession codes ENCSR156CWW and ENCSR000AMB. Additionally, scRNA-seq for HCC immune microenvironment analyses, spatial transcriptomic, and immunotherapy-related datasets were retrieved from Mendeley Data (https://data.mendeley.com/datasets/skrx2fz79n/1) under accession code skrx2fz79n/1 and the European Genome-Phenome Archive (https://ega-archive.org/studies/EGAS00001005477) under accession code EGAS00001005477. The scRNA-seq data profiles of human liver development are obtained from http://collections.cellatlas.io/liver-development. The HCC scRNA-seq data profiles HCC developmental potential are obtained from GEO (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE202642, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE151530, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE206325, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE149614, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE156625) under accession codes GSE202642, GSE151530, GSE206325, GSE149614, and GSE156625. The gene sets used for GSEA were collected from the Molecular Signatures Database (https://www.gsea-msigdb.org/gsea/msigdb). Proteomic data of HCC patients were downloaded from the National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) (https://proteomics.cancer.gov/programs/cptac). The publicly available datasets used in this study were provided in Supplementary Dataset 2. Source data are provided with this paper.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Vogel, A., Meyer, T., Sapisochin, G., Salem, R. & Saborowski, A. Hepatocellular carcinoma. Lancet 400, 1345–1362 (2022).
Rimassa, L., Finn, R. S. & Sangro, B. Combination immunotherapy for hepatocellular carcinoma. J. Hepatol. 79, 506–515 (2023).
Qin, S. et al. Atezolizumab plus bevacizumab versus active surveillance in patients with resected or ablated high-risk hepatocellular carcinoma (IMbrave050): a randomised, open-label, multicentre, phase 3 trial. Lancet 402, 1835–1847 (2023).
Llovet, J. M. et al. Lenvatinib plus pembrolizumab versus lenvatinib plus placebo for advanced hepatocellular carcinoma (LEAP-002): a randomised, double-blind, phase 3 trial. Lancet Oncol. 24, 1399–1410 (2023).
Hänggi, K. & Ruffell, B. Cell death, therapeutics, and the immune response in cancer. Trends Cancer 9, 381–396 (2023).
Sia, D. et al. Identification of an immune-specific class of hepatocellular carcinoma, based on molecular features. Gastroenterology 153, 812–826 (2017).
Montironi, C. et al. Inflamed and non-inflamed classes of HCC: a revised immunogenomic classification. Gut 72, 129–140 (2023).
You, W. et al. Extending inflamed-class signature to predict immune checkpoint inhibitor-based combination therapy in hepatocellular carcinoma. Gut 74, 679–681 (2024).
Li, B., Cao, Y., Li, Y., Cheng, C. & Yu, D. Letter to the editor: the inflamed subclass predicts immunotherapy response—external validations. Gut gutjnl, 2022–328130 (2022).
Cai, G. et al. A Novel hepatocellular carcinoma specific hypoxic related signature for predicting prognosis and therapeutic responses. Front. Immunol. 13, 997316 (2022).
Cai, N. et al. Targeting MMP9 in CTNNB1 mutant hepatocellular carcinoma restores CD8(+) T cell-mediated antitumour immunity and improves anti-PD-1 efficacy. Gut 73, 985–999 (2024).
Barcena-Varela, M. & Berraondo, P. Targeting metalloproteases is a promising strategy to enhance immunotherapy responses by overcoming immune exclusion in hepatocellular carcinoma. Gut 73, 890–891 (2024).
Xie, M. et al. FGF19/FGFR4-mediated elevation of ETV4 facilitates hepatocellular carcinoma metastasis by upregulating PD-L1 and CCL2. J. Hepatol. 79, 109–125 (2023).
Yi, C. et al. Lenvatinib targets FGF receptor 4 to enhance antitumor immune response of anti-programmed cell death-1 in HCC. Hepatology 74, 2544–2560 (2021).
Li, H. et al. MET inhibitors promote liver tumor evasion of the immune response by stabilizing PDL1. Gastroenterology 156, 1849–1861.e13 (2019).
Wong, T. et al. Protein tyrosine kinase 7 (PTK7) promotes metastasis in hepatocellular carcinoma via SOX9 regulation and TGF-β signaling. Cell. Mol. Gastroenterol. Hepatol. 15, 13–37 (2023).
Wang, X. et al. Aurora A kinase inhibition compromises its antitumor efficacy by elevating PD-L1 expression. J. Clin. Investig. 133, e161929 (2023).
Chaib, M. et al. Protein kinase C delta regulates mononuclear phagocytes and hinders response to immunotherapy in cancer. Sci. Adv. 9, eadd3231 (2023).
Wang, J. C. et al. PIM2 expression induced by proinflammatory macrophages suppresses immunotherapy efficacy in hepatocellular carcinoma. Cancer Res. 82, 3307–3320 (2022).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Cancer, G. A. R. N. E. a. w. e. & Network, C. G. A. R. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341.e23 (2017).
Hoshida, Y. et al. Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res. 69, 7385–7392 (2009).
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Prim. 7, 6 (2021).
Calderaro, J., Ziol, M., Paradis, V. & Zucman-Rossi, J. Molecular and histological correlations in liver cancer. J. Hepatol. 71, 616–630 (2019).
Zhang, J. et al. Modulation of brain cation-Cl(-) cotransport via the SPAK kinase inhibitor ZT-1a. Nat. Commun. 11, 78 (2020).
Wesley, B. T. et al. Single-cell atlas of human liver development reveals pathways directing hepatic cell fates. Nat. Cell Biol. 24, 1487–1498 (2022).
Kang, M. et al. Improved reconstruction of single-cell developmental potential with CytoTRACE 2. Nat. Methods 22, 2258–2263 (2025).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Liu, Y. et al. Identification of a tumour immune barrier in the HCC microenvironment that determines the efficacy of immunotherapy. J. Hepatol. 78, 770–782 (2023).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to the exclusion of T cells. Nature 554, 544–548 (2018).
Tamborero, D. et al. A pan-cancer landscape of interactions between solid tumors and infiltrating immune cell populations. Clin. Cancer Res. 24, 3717–3728 (2018).
Wei, W., Jin, J., Schlisio, S., Harper, J. W. & Kaelin, W. G. Jr The v-Jun point mutation allows c-Jun to escape GSK3-dependent recognition and destruction by the Fbw7 ubiquitin ligase. Cancer Cell 8, 25–33 (2005).
Xiang, D. M. et al. Oncofetal HLF transactivates c-Jun to promote hepatocellular carcinoma development and sorafenib resistance. Gut 68, 1858–1871 (2019).
Yang, H. et al. Pharmacogenomic profiling of intra-tumor heterogeneity using a large organoid biobank of liver cancer. Cancer Cell 42, 535–551.e8 (2024).
Wang, J. et al. A positive feedback loop of OTUD1 and c-Jun driven by leptin expedites stemness maintenance in ovarian cancer. Oncogene 44, 1731–1745 (2025).
Yamaguchi, H., Hsu, J. M., Yang, W. H. & Hung, M. C. Mechanisms regulating PD-L1 expression in cancers and associated opportunities for novel small-molecule therapeutics. Nat. Rev. Clin. Oncol. 19, 287–305 (2022).
Cha, J. H., Chan, L. C., Li, C. W., Hsu, J. L. & Hung, M. C. Mechanisms controlling PD-L1 expression in cancer. Mol. Cell 76, 359–370 (2019).
Li, C. W. et al. Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nat. Commun. 7, 12632 (2016).
Wu, M., Xia, X., Hu, J., Fowlkes, N. W. & Li, S. WSX1 acts as a tumor suppressor in hepatocellular carcinoma by downregulating neoplastic PD-L1 expression. Nat. Commun. 12, 3500 (2021).
Zhang, Y. et al. HKDC1 promotes tumor immune evasion in hepatocellular carcinoma by coupling the cytoskeleton to STAT1 activation and PD-L1 expression. Nat. Commun. 15, 1314 (2024).
Zhang, J. et al. Cyclin D-CDK4 kinase destabilizes PD-L1 via cullin 3-SPOP to control cancer immune surveillance. Nature 553, 91–95 (2018).
Kornepati, A., Vadlamudi, R. K. & Curiel, T. J. Programmed death ligand 1 signals in cancer cells. Nat. Rev. Cancer 22, 174–189 (2022).
Yang, X. et al. Gene body methylation can alter gene expression and is a therapeutic target in cancer. Cancer Cell 26, 577–590 (2014).
Wang, Y. W. et al. ITPKA gene body methylation regulates gene expression and serves as an early diagnostic marker in lung and other cancers. J. Thorac. Oncol. 11, 1469–1481 (2016).
Arechederra, M. et al. Hypermethylation of gene body CpG islands predicts high dosage of functional oncogenes in liver cancer. Nat. Commun. 9, 3164 (2018).
Li, G. et al. Development and validation of a CIMP-associated prognostic model for hepatocellular carcinoma. EBioMedicine 47, 128–141 (2019).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Johnston, A. M. et al. SPAK, a STE20/SPS1-related kinase that activates the p38 pathway. Oncogene 19, 4290–4297 (2000).
Gagnon, K. B. & Delpire, E. Molecular physiology of SPAK and OSR1: two Ste20-related protein kinases regulating ion transport. Physiol. Rev. 92, 1577–1617 (2012).
Qiu, Z. et al. STK39 promotes breast cancer invasion and metastasis by increasing SNAI1 activity upon phosphorylation. Theranostics 11, 7658–7670 (2021).
Zhang, C. et al. STK39 is a novel kinase contributing to the progression of hepatocellular carcinoma by the PLK1/ERK signaling pathway. Theranostics 11, 2108–2122 (2021).
Hao, X. et al. STK39 enhances the progression of Cholangiocarcinoma via PI3K/AKT pathway. iScience 24, 103223 (2021).
Chen, J. et al. Knockdown of STK39 suppressed cell proliferation, migration, and invasion in hepatocellular carcinoma by repressing the phosphorylation of mitogen-activated protein kinase p38. Bioengineered 12, 6529–6537 (2021).
Xu, Y. et al. STK39-mediated amplification of γ-H2A.X promotes homologous recombination and contributes to PARP inhibitor resistance. Nucleic Acids Res. 52, 13881–13895 (2024).
Eferl, R. et al. Liver tumor development. c-Jun antagonizes the proapoptotic activity of p53. Cell 112, 181–192 (2003).
Min, L. et al. Liver cancer initiation is controlled by AP-1 through SIRT6-dependent inhibition of survivin. Nat. Cell Biol. 14, 1203–1211 (2012).
Villarruel-Melquiades, F. et al. Activator protein-1 in liver pathology: from precancerous diseases to hepatocellular carcinoma. Hepatoma Res. 10, 51 (2024).
Wu, Y., Wakil, A., Salomon, F. & Pyrsopoulos, N. Issue on combined locoregional and systemic treatment for hepatocellular carcinoma. Hepatoma Res. 9, 6 (2023).
Lao, Y. et al. Deciphering a profiling based on multiple post-translational modifications functionally associated regulatory patterns and therapeutic opportunities in human hepatocellular carcinoma. Mol. Cancer 23, 283 (2024).
Savage, S. R. et al. Pan-cancer proteogenomics expands the landscape of therapeutic targets. Cell 187, 4389–4407.e15 (2024).
Nishiyama, A. & Nakanishi, M. Navigating the DNA methylation landscape of cancer. Trends Genet. 37, 1012–1027 (2021).
Wang, Q. et al. Gene body methylation in cancer: molecular mechanisms and clinical applications. Clin. Epigenetics 14, 154 (2022).
Weinberg, D. N. et al. The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature 573, 281–286 (2019).
Quarto, G. et al. Fine-tuning of gene expression through the Mettl3-Mettl14-Dnmt1 axis controls ESC differentiation. Cell S0092-8674, 01422–01423 (2025).
Maunakea, A. K. et al. Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature 466, 253–257 (2010).
Mercado, C. J., Wang, X., Grimm, P. R., Welling, P. A. & Chang, Y. C. Identification and characterization of alternative STK39 transcripts within human and mouse kidneys reveals species-specific regulation of blood pressure. Physiol. Rep. 8, e14379 (2020).
Sie, Z. L. et al. WNK1 Kinase Stimulates Angiogenesis to Promote Tumor Growth and Metastasis. Cancers 12, 575 (2020).
Acknowledgements
The authors thank the flow cytometer provided by the Experimental Medical Research Center of Tongji Hospital (Wuhan, China), and thank Prof. Weimin Wang (Dean of the Department of Immunology, Tongji Medical College, HUST) for providing technical support for flow cytometric analyses. The authors also thank Prof. Jiahuai Han (School of Life Sciences, Faculty of Medicine and Life Sciences, Xiamen University) for presenting the CDS and plasmid. In addition, the authors thank Jonathan Wei and Jimmy Zhan (Cyagen, Suzhou, China) for their assistance in the generation of genetically modified mice, Nanjing Vazyme Biotech Co., Ltd (Nanjing, China) for providing technical guidance for the CUT&Tag DNA library construction, Annoroad Gene Technology Co., Ltd (Beijing, China) for providing CUT&Tag library sequencing analysis and SpecAlly Life Technology Co., Ltd (Wuhan, China) for providing technical support for the LC-MS/MS analysis. This research was funded by the National Natural Science Foundation of China (82303188 to Y.P., 82472970 and 82273441 to Z.D., 82400870 to C.Y. and 82303516 to Y.H.), the first level of the public health youth top talent project of Hubei province (2022SCZ051 to Z.D.), the Tongji Hospital (HUST) Foundation for Excellent Young Scientists (2020YQ05 and 24-2KYC13057-05 to Z.D.), and Shenzhen Key Medical Discipline Construction Fund (No.SZXK015 to J.W.).
Author information
Authors and Affiliations
Contributions
Y.P. and Z.D. developed the study design and made substantial contributions to the manuscript writing. Y.P. and C.Z. conducted the experiments and performed statistical analyses. Y.H., R.C., J.W., and Haisen L. were responsible for bioinformatics analysis and data visualization. Y.P., C.Y., C.S., and M.H. contributed to the in vitro experiments. Y.P., C.Z., C.Y., X.L., F.L., and M.W. contributed to the in vivo experiments. P.L., L.X., H.Z., and Huifang L. contributed to the collection of clinical specimens and provided clinical pathology evaluations. Z.D., B.Z., Z.L., and J.L. analyzed and interpreted the study results. Z.D., B.Z., W.Z., and X.C. were responsible for conceiving, designing, and overseeing the study, offering critical feedback on the manuscript. All authors have read and approved the final manuscript. Y.P. and Z.D. are responsible for the overall content of the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pan, Y., Zeng, C., He, Y. et al. Targeting SPAK suppresses progression and averts an immune exhaustive microenvironment in hepatocellular carcinoma. Nat Commun 17, 1414 (2026). https://doi.org/10.1038/s41467-025-68156-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68156-8