Abstract
Pancreatic ductal adenocarcinoma (PDAC) presents a therapeutic hurdle owing to its immunotherapy resistance and limited treatment options. Pyroptosis, a lytic inflammatory cell death pathway, holds promise for reversing immunosuppression in “cold” tumors, yet tumor-specific induction strategies remain unclear. Here, we employ a contrastive learning graph neural network to identify arachidonic acid (AA), an endogenous ω−6 polyunsaturated fatty acid, as a pan-cancer pyroptosis inducer operating through irreversible mitochondrial permeability transition inducing Caspase 3/GSDME activation. Crucially, a repurposed mitophagy machinery for non-autophagic functional scaffolding of Caspase 3/GSDME on autophagosomal membranes enables rapid pyroptotic execution. Pharmacological accumulation of autophagosomes amplifies AA-induced pyroptosis in PDAC organoids and patient-derived xenografts. Furthermore, AA-triggered pyroptosis, remodels immunosuppressive tumor microenvironments, boosting cytotoxic T cell infiltration and synergizing with anti-PD-1 therapy in immunocompetent models. Thus, our findings position AA as a candidate dual-function agent for pyroptosis-immune synergy, while autophagosome manipulation emerges as a strategy to potentiating strategy.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is currently one of the most intractable cancers with the lowest survival rates, which ranks third- to fourth-leading cause of cancer-related death with minimal improvement in survival rates1. Although combination chemotherapy and immunotherapy have developed over the recent years and have transformed the outlook of advanced patients with some types of tumors, high rate of relapse to chemotherapy and the near universal refractoriness of PDAC to immune checkpoint blockade (ICB) is essentially unique across all human cancer types2, highlighting the need for alternative therapeutic options.
Pyroptosis, defined as a gasdermin family protein-mediated inflammatory cell death, has emerged as a major focus in oncology research3. Unlike apoptosis, which only induces moderate tumor suppression and lacks immune-activating properties, pyroptosis exhibits dual antitumor merits: it not only suppresses malignant tumor growth more effectively but also elicits robust immunostimulatory effects that actively remodel the tumor immune microenvironment via coordinated release of damage-associated molecular patterns (DAMPs) and cytokines4,5,6, positioning such immune remodeling as a promising avenue for augmenting responses of immune “cold” tumors to immunotherapy7,8. Even chemotherapeutic drugs can induce secondary pyroptosis9, but lack the capacity for meaningful immunomodulation due to the low proportion of pyroptotic cells. Therefore, developing strategies to specifically induce pyroptosis in tumor cells has emerged as a critical issue — requiring precise control of gasdermin activation to simultaneously achieve robust tumor eradication and potentiate anticancer immunity.
Emerging evidence highlights that various kinds of lipids from a high-fat diet modulate potent regulatory effects on both inflammatory and immunomodulatory pathways in multiple cancer types10. We hypothesize that specific fatty acids may exert substantial anti-tumor efficacy through their dual immunomodulatory functions in regulating both inflammatory signaling pathways and immune cell activation states. To accelerate lipid repositioning for pyroptosis-based immunotherapy, we developed a Chemical-Disease Knowledge Graph (CDKG) that systematically integrates multi-scale evidence spanning molecular interactions to clinical manifestations from the Comparative Toxicogenomics Database (CTD)11, thus mapping the association between diseases and chemicals, including fatty acids. Subsequently, we proposed a Contrastive Learning Graph Neural Network (CLGNN) model12 to predict the interaction coefficients between fatty acids and PDAC through multi-scale graph embeddings from CDKG, followed by systematic evaluation of specific lipids exhibiting gasdermin-dependent pyroptotic activation potential.
In this study, based on the prediction results of the CLGNN model, we show that high concentration of arachidonic acid, a significant participant in the collaboration of ferroptosis induction13,14, specifically triggers acute pyroptosis in tumor cells, which is evaluated in a variety of tumor models of PDAC. We demonstrated that arachidonic acid induces mitochondrial permeability transition and mitophagy in a non-metabolism-related manner. Autophagosomes could further recruit both Caspase 3 and Gasdermin E (GSDME) by a nonautophagy-related function of mitophagy, which provides a platform for the cleavage of GSDME by pro-apoptotic factor-activated Caspase 3, thereby inducing a rapid and high proportion of pyroptosis. Cancer cell pyroptotic death upon treatment with arachidonic acid recruits T cells, increasing infiltration into the “immune-cold” microenvironment. Thus, the synergy between ICB and arachidonic acid in suppressing PDAC provides a rationale for arachidonic acid’s dual role in this combination therapy: potentially enabling tumor reduction through pyroptosis induction and functioning as an immunotherapeutic activator.
Results
CLGNN predicts therapeutic potential of PUFA and high concentration of arachidonic acid triggers pyroptosis of PDAC
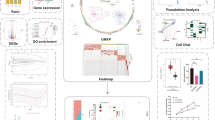
We hypothesized that selective fatty acids could treat PDAC through inducing tumor cell pyroptosis. To test this hypothesis, we proposed the CLGNN model (Fig. 1a) to detect the therapeutic potential of all typical fatty acids (Supplementary Table 1). To assess the prediction performance of the CLGNN model against baseline models, we developed CDKG that comprehensively maps the association relationships between 1329 diseases and 10,357 chemicals in CTD. Among prediction models, the finally selected CLGNN_GIN model demonstrated superior performance, as convincingly evidenced by the area under receiver operating characteristic curve (AUROC) and the area under precision recall curve (AUPR) in random data splitting experiments (Supplementary Fig. 1a). Building on the superiority in chemical-disease association prediction for therapy, the therapeutic potential of all fatty acids against PDAC was predicted (Fig. 1b). Interestingly, the predicted results revealed a significant therapeutic potential of ω−6 (pro-inflammatory) and ω−3 (anti-inflammatory) polyunsaturated fatty acids (PUFAs). Since inflammation activation is associated with pyroptosis15,16,17, we systematically investigated the pyroptosis-inducing capacity of arachidonic acid—a typical ω−6 PUFA with documented pro-inflammatory activity—in addition to its role in ferroptosis13. We administered arachidonic acid exogenously to pancreatic cancer cell lines and unexpectedly observed that concentrations ≥ 120 μM induced substantial cell death exhibiting definitive morphological features of pyroptosis—including cellular swelling and plasma membrane blebbing (Fig. 1c)— which was further confirmed by transmission electron microscopy (TEM) showing membrane perforation (Fig. 1d and Supplementary Fig. 1b). These pyroptotic characteristics were accompanied by Calcein-AM efflux (Fig. 1e, f and Supplementary Fig. 1c–e) and LDH release (Fig. 1g), phenomena consistent with non-specific membrane damage. An expansion in the Propidium Iodide (PI)+/Annexin V (AnnV)- and PI+/AnnV+ stained populations further supported that the arachidonic acid-induced cell death was pyroptosis (Fig. 1h, i and Supplementary Fig. 1f–h). Consistent with established evidence that arachidonic acid alone lacks direct ferroptosis-inducing activity18, arachidonic acid-elicited pyroptotic morphology and LDH release remained unaffected by pretreatment with either ferrostatin-1 (Fer-1, ferroptosis inhibitor), liproxstatin-1 (Lip-1, ferroptosis inhibitor), or necrosulfonamide (NSA, MLKL inhibitor for necroptosis blockade) (Supplementary Fig. 1i–m). We further assessed the cytotoxic effect of arachidonic acid across pan-cancer cell lines. As expected, treatment with arachidonic acid induced analogous acute pyroptosis effects in other kinds of cancer cell lines (Supplementary Fig. 1n, o).
a Framework of Contrastive Learning Graph Neural Network (CLGNN) model. The figure was assembled using Microsoft PowerPoint. The icons were sourced from Bioicons (https://bioicons.com/, under CC0 license) and Iconfont (https://www.iconfont.cn/, marked as free for academic use). b Scores of the therapeutic effect of common fatty acids on PDAC predicted by the CLGNN model. The polyunsaturated fatty acids were in bold. c Representative images of pyroptotic morphology changes in BxPC3, SW1990, PANC1, and Pan02 cells. Cells were stimulated with 40 or 120 μM arachidonic acid for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars: 25 μm. d Representative images of electron microscopy analysis of pyroptotic morphology changes in BxPC3 cells. Cells were stimulated with 40 or 120 μM arachidonic acid for 2 h. The disruption of the cell membrane was marked with yellow arrowheads. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars: 5 μm (Low Power Field) and 1 μm (High Power Field). e, f Representative images (e) and relative percentage of Calcein-AM release into the medium (f) in Calcein-AM release analysis in BxPC3 and SW1990 cells. Cells were stained with Calcein-AM for 30 min, then stimulated with gradient concentrations of arachidonic acid for 2 h. Yellow arrowheads mark cells exhibiting a complete loss of Calcein-AM. Cyan arrowheads mark cells exhibiting partial loss. n = 3 independent experiments, one representative image of 3 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars: 50 μm. g LDH release assay of BxPC3, SW1990, PANC1, and Pan02 cell lines treated with a gradient concentration of arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. h Flow cytometry analysis of Annexin V (AnnV)/Propidium Iodide (PI)-stained BxPC3. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Experimental design as in (c). i Proportions of AnnV-/PI+ and AnnV+/PI+ cell population cells, quantified from images as shown in (h). n = 3 independent experiments, Two-way ANOVA with Tukey’s multiple comparisons test. Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 1 and Supplementary Fig. 2. Source data are provided as a Source Data file.
Notably, we observed distinct pyroptosis-inducing properties of arachidonic acid compared to the potential secondary pyroptosis triggered by chemotherapeutic agents9. Chemotherapeutic treatments failed to induce pyroptosis-associated morphological alterations at the 4-hour time point across tested concentrations (Supplementary Fig. 2a), with no detectable Calcein-AM efflux or LDH release (Supplementary Fig. 2b, c). Prolonged 24 h exposure similarly revealed no significant pyroptotic changes – including preserved cellular morphology and absent biomarkers release (Supplementary Fig. 2a, b, d) – even when cell death was evident (Supplementary Fig. 2e). These findings demonstrated that arachidonic acid efficiently induced tumor cell-specific acute pyroptosis, contrasting mechanistically with the low-frequency secondary pyroptosis associated with conventional chemotherapy.
Caspase 3/GSDME is involved in arachidonic acid-induced pyroptosis
The Gasdermin protein family plays critical roles in pyroptosis induction19. To confirm the pyroptotic nature of arachidonic acid-induced cell death, we investigated the cleavage of two key executor proteins – Gasdermin D (GSDMD) and GSDME – which mediate pyroptotic membrane permeabilization. While GSDMD serves as a critical mediator of caspase-1/4/8/11-mediated pyroptosis20,21, Caspase 3-dependent cleavage of GSDME has been recently established9,22. High-concentration arachidonic acid treatment induced GSDME processing into its pore-forming p30 fragment (Fig. 2a and Supplementary Fig. 3a, b), with cleavage efficiency showing dose-dependent enhancement at elevated concentrations (Fig. 2b and Supplementary Fig. 3c). Indeed, cytoplasmic accumulation of cleaved GSDME significantly increased following high-concentration arachidonic acid treatment (Fig. 2c, d). Neither necroptosis nor ferroptosis inhibitors attenuated arachidonic acid-induced GSDME cleavage (Supplementary Fig. 3d, e). Furthermore, consistent with our prior findings, no alterations in ferroptosis-associated marker proteins were induced following arachidonic acid treatment (Supplementary Fig. 3f, g), indicating ferroptosis pathway independence under these experimental conditions. In contrast to GSDME cleavage, arachidonic acid failed to generate the pore formation-competent GSDMD p30 fragment but unexpectedly produced a non-functional p43 fragment (Fig. 2a and Supplementary Fig. 3h). Consistently, genetic silencing of GSDME (Supplementary Fig. 3i, j) substantially reduced arachidonic acid-induced pyroptosis, PI positive staining (Fig. 2e, f), and LDH release (Fig. 2g), whereas GSDMD knockdown (Supplementary Fig. 3k, l) failed to exert this inhibitory effect (Fig. 2h). Moreover, pretreatment with Z-DEVD-FMK, a specific Caspase 3 activation inhibitor, suppressed the cleavage of GSDME, and diminished LDH release in arachidonic acid-treated BxPC3 cells (Fig. 2i–l and Supplementary Fig. 3m).
a Immunoblots of GSDME, GSDMD, and Tubulin in BxPC3, SW1990, and PANC1 cells. Cells were stimulated with 40 or 120 μM arachidonic acid for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. b Immunoblots of GSDME and Tubulin in BxPC3. Cells were stimulated with 100 or 120 μM arachidonic acid for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. c Representative immunofluorescent images of cleaved-GSDME in BxPC3 and PANC1 cells. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bar: 10 μm. d Quantitative analysis of mean fluorescence intensity of cleaved-GSDME in BxPC3 and PANC1 cells. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. e Representative brightfield and Propidium Iodide (PI)-staining images of arachidonic acid treated BxPC3-NC and GSDME-knockout (KO) BxPC3 cells for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bar: 25 μm. f Quantitative analysis of PI+ cell percentage in BxPC3 cells. n = 5 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. g, h LDH release assay of BxPC3, GSDME-KO (g), and GSDMD-KD (h) BxPC3 cells treated by arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. i–l Effects of Z-DEVD-FMK on arachidonic acid-induced pyroptosis in BxPC3. Representative images (i) and quantitative analysis (j) of pyroptotic morphology changes in BxPC3 cells. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. Cleavage of GSDME and activation of Caspase 3 (k), and LDH release assay (l) in BxPC3 cells. n = 3 independent experiments, one representative image of 3 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Cells were pretreated by Z-DEVD-FMK for 6 h, and subsequently stimulated with 120 μM arachidonic acid for 2 h. m–q Mice bearing subcutaneous tumors (BxPC3-N.C. or BxPC3-GSDME-KO) received arachidonic acid (n = 7 per group). Schematic illustration. q48h, every 48 h (m). Created in BioRender. Fan, Z. (2026) https://BioRender.com/04f0qz9. Images of subcutaneous tumors (n), and the quantification of tumor volume (o, p) and weight (q) were recorded at the indicated times. n = 7, Two-way ANOVA with Tukey’s multiple comparisons test (o) and One-way ANOVA with Dunnett’s multiple comparisons test (p, q). Scale bars, 1 cm. Tubulin was used as a loading control. Densitometric quantification of cleaved GSDME ( ~ 30 kDa, lower band) and cleaved caspase 3 (17/19 kDa, lower band) normalized to loading control (Two distinct bands on the membrane). Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 3. Source data are provided as a Source Data file.
To further address the critical point raised regarding whether arachidonic acid uniquely potentiates GSDME-mediated pyroptosis beyond its ability to activate caspase-3 compared to conventional chemotherapeutic stimuli, we conducted a direct biochemical comparison. We specifically investigated potential differences in the efficiency of caspase-3 activation and subsequent GSDME cleavage between arachidonic acid and paclitaxel, moving beyond the initial assessments based on morphology and pyroptosis markers (Supplementary Fig. 2). Crucially, to enable a more meaningful comparison of caspase-3 activity levels at similar stages of cell death commitment, we selected treatment time points where both agents induced approximately 60% cell death (arachidonic acid: 120 μM for 2.5 hours; Paclitaxel: 20 μM for 24 h). At this equivalent cell death endpoint, biochemical analysis revealed comparable levels of Caspase 3 activation between arachidonic acid- and paclitaxel-treated cells, whereas GSDME cleavage was strikingly more pronounced in the arachidonic acid group (Supplementary Fig. 3n, o). These results demonstrate that despite comparable Caspase 3 activation, arachidonic acid induces not only faster Caspase 3 activation kinetics but more critically, a substantially higher efficiency of Caspase 3-dependent GSDME cleavage.
To evaluate the tumor-suppressive efficacy of arachidonic acid in vivo, xenograft models bearing either GSDME-knockout (KO) or non-targeting control (N.C.) BxPC3 cells were established (Fig. 2m). Intratumoral administration of arachidonic acid significantly suppressed tumor growth in the N.C. group compared to placebo-treated controls, as evidenced by reduced tumor volume and weight (Fig. 2n–q), concomitant with GSDME cleavage (Supplementary Fig. 3p, q). However, no discernible antitumor effects were observed in GSDME-KO tumors following arachidonic acid treatment (Fig. 2n–q). Collectively, these data demonstrate that arachidonic acid-induced tumor pyroptosis requires the Caspase 3/GSDME pathway, while its in vivo antitumor activity is primarily dependent on GSDME.
Arachidonic acid induced irreversible mPT is necessary trigger of pyroptosis
We next investigated the mechanism underlying arachidonic acid-induced pyroptosis. Mitochondria serve as pivotal organelles in cell death regulation, primarily through mitochondrial permeability alterations mediated by mitochondrial pore formation23,24. This aligns with our observations of mitochondrial swelling and cristae loss – hallmark features of mitochondrial permeability transition (mPT) – in pyroptotic cells following high-concentration arachidonic acid treatment (Fig. 3a, b, mitochondria are indicated with yellow arrows). mPT describes an abrupt increase in inner mitochondrial membrane permeability to low molecular weight solutes, mediated by the mitochondrial permeability transition pore (mPTP), which disrupts osmotic homeostasis. Unlike transient mPT flickering, a physiological phenomenon inducible by ionomycin (Supplementary Fig. 4a, b), irreversible mPT initiates divergent cellular outcomes ranging from mitophagy to apoptosis or necrosis25,26. Indeed, arachidonic acid treatment induced disruption of inner mitochondrial membrane integrity (Fig. 3c) and collapse of the mitochondrial membrane potential (Δψm) (Fig. 3d–f). Furthermore, Calcein-AM/CoCl2 co-staining with PI revealed that pyroptotic cells exhibited increased inner mitochondrial membrane permeability (Fig. 3g).
a Representative electron microscopy images of BxPC3 cells. Representative mitochondrial morphologies were marked with yellow arrowheads. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 1 μm. b Mitochondrial cristae density was represented as percent area void of cristae, quantified from images as shown in (a). n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. c–e Relative mPTP formation was detected by Calcein-AM/CoCl2 assay (c) and dissipation of Δψm by JC−1 assay (d) and Tetramethylrhodamine methyl ester perchlorate (TMRM) staining assay (e). BxPC3 were treated with 40 or 120 μM arachidonic acid for 1 h. Ionomycin and carbonyl cyanide 3-chlorophenylhydrazone (CCCP) were used as positive controls of mPT and Δψm dissipation, respectively. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. f Representative images of TMRM staining assay. Experimental design as in (e). n = 3 independent experiments, one representative image of 3 independent experiments is shown. g Representative images of Calcein-AM/CoCl2 assay and PI staining assay. Cells were treated with 40 or 120 μM arachidonic acid for 2 h. Ionomycin was used as a positive control. n = 3 independent experiments, one representative image of 3 independent experiments is shown. h–j mPT detected by Calcein-AM/CoCl2 assay (h) and dissipation of Δψm by JC−1 assay (i) and TMRM staining assay (j). BxPC3 cells were pre-treated with NIM811 (10 μM) or ER-000444793 (10 μM) for 2 h followed by 120 μM arachidonic acid treatment for 1 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. k, l Representative images (k) and quantitative analysis (l) of pyroptotic morphology changes in NIM811/ER-000444793 pre-treated BxPC3 cells. Cells were pre-treated with NIM811 (10 μM) or ER-000444793 (10 μM) for 2 h followed by 120 μM arachidonic acid treatment for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. m Immunoblots of GSDME, Caspase 3, and HSP90 in cell lysates of NIM811/ER-000444793 pre-treated BxPC3 cells. Cells were pre-treated with NIM811 (10 μM) or ER-000444793 (10 μM) for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. n LDH release assay of NIM811-/ER-000444793-pre-treated cells followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. HSP90 was used as a loading control. Densitometric quantification of cleaved GSDME (~ 30 kDa, lower band) and cleaved caspase 3 (17/19 kDa, lower band) normalized to loading control (Two distinct bands on the membrane). Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 4. Source data are provided as a Source Data file.
We further investigated whether irreversible mPT is required for arachidonic acid-induced pyroptosis. NIM81127,28,29 and ER-00044479330,31, two well-characterized inhibitors known to target mPT, were employed. Pretreatment of two specific mPT inhibitors significantly attenuated arachidonic acid-induced mPT (Fig. 3h) and Δψm collapse (Fig. 3i, j). Moreover, arachidonic acid-triggered Caspase 3/GSDME cleavage, pyroptosis, and LDH release were also inhibited upon the mPT inhibitors-pretreatment (Fig. 3k–n and Supplementary Fig. 4c). These findings demonstrate that irreversible mPT is necessary for pyroptosis during arachidonic acid treatment.
Ca2+ is responsible for irreversible mPT upon arachidonic acid treatment
Our findings have demonstrated an arachidonic acid-induced irreversible mPTP opening that leads to pyroptosis of PDAC cells upon arachidonic acid treatment (Fig. 3). However, the mechanism by which arachidonic acid facilitates irreversible mPT remains unresolved. Cyclophilin D (CypD) and Ca2+ rather than adenine nucleotide translocase (ANT) or phosphate carrier (PiC) – represent the two identified activating pathways to induce monomer deformation and destabilizes dimeric structure of ATP synthase, enabling irreversible mPT32,33. We therefore inhibited irreversible mPT in BxPC3 cells using cyclosporine A (CsA), a common mPT inhibitor that blocks CypD binding to ATP synthase. However, the CypD inhibitor CsA failed to inhibit arachidonic acid-induced mPT (Supplementary Fig. 5a–c) or pyroptosis (Supplementary Fig. 5d–h). We further applied melatonin, an F1FO-ATPase-binding inhibitor of Ca2+-activated mPTP opening34, to inhibit mPT in BxPC3 cells. Of note, melatonin significantly prevented arachidonic acid-induced mPT and Δψm collapse (Fig. 4a, b). Pretreatment with Melatonin inhibited arachidonic acid-triggered Caspase 3/GSDME cleavage, pyroptosis, and LDH release (Fig. 4c–f and Supplementary Fig. 5i). Moreover, two kinds of Ca2+ chelators were further applied. Notably, pretreatment with the fast Ca2+ chelator BAPTA-AM, which sequesters Ca2+ within sub-100 nm microdomains between channels and sensors35, effectively blocked arachidonic acid-induced mPTP formation and Δψm collapse (Fig. 4g), GSDME cleavage, and pyroptosis (Fig. 4h–k and Supplementary Fig. 5j). However, the slow Ca2+ chelator EGTA-AM showed no inhibitory effects (Fig. 4h, i, l–n and Supplementary Fig. 5k). Therefore, we demonstrate that arachidonic acid induces mPT via Ca2+, which is consistent with reported previously mPT induction pathways32,33,36, as well as the inhibitory properties of NIM811 and ER-000444793 against Ca2+-induced mPT27,28,29,30,31, and we hypothesize that the Ca2+ signaling may be spatially restricted.
a Representative images of Calcein-AM/CoCl2 assay. BxPC3 cells were pre-treated with melatonin (1 mM or 2 mM) for 2 h followed by 120 μM arachidonic acid treatment for 1 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 25 μm. b mPT detected by Calcein-AM/CoCl2 assay and dissipation of Δψm by JC-1 assay and TMRM staining assay. Experimental design as in (a). n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. c, d Representative images (c) and quantitative analysis (d) of pyroptotic morphology changes in melatonin-pre-treated BxPC3 cells. BxPC3 cells were pre-treated with melatonin (1 mM or 2 mM) for 2 h followed by 120 μM arachidonic acid treatment for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. e Immunoblots of GSDME, Caspase 3, and Tubulin in cell lysates of melatonin-pre-treated BxPC3 cells. BxPC3 cells were pre-treated with melatonin (1 mM or 2 mM) for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. f LDH release assay of melatonin-pre-treated BxPC3 cells. BxPC3 cells were pre-treated with melatonin (1 mM or 2 mM) for 2 h followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. g mPT detected by Calcein-AM/CoCl2 assay and dissipation of Δψm by JC-1 assay and TMRM staining assay. BxPC3 cells were pre-treated with 2.5–20 μM (Calcein-AM/CoCl2 assay) or 20 μM (JC-1 assay and TMRM staining assay) BAPTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 1 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. h, i Representative images (h) and quantitative analysis (i) of pyroptotic morphology changes in BAPTA-AM/EGTA-AM pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 20 μM BAPTA-AM/EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. j Immunoblots of GSDME, Caspase 3, and HSP90 in cell lysates of BAPTA-AM pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 2.5-20 μM BAPTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. k LDH release assay of BAPTA-AM pre-treated BxPC3 cells. Cells were pre-treated with 2.5–20 μM BAPTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. l mPT detected by Calcein-AM/CoCl2 assay and dissipation of Δψm by JC-1 assay and TMRM staining assay. BxPC3 cells were pre-treated with 2.5-20 μM (Calcein-AM/CoCl2 assay) or 20 μM (JC-1 assay and TMRM staining assay) EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 1 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. m Immunoblots of GSDME, Caspase 3, and GAPDH in cell lysates of EGTA-AM pre-treated BxPC3 cells. Cells were pre-treated with 2.5–20 μM EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. n LDH release assay of EGTA-AM pre-treated BxPC3 cells. Cells were pre-treated with 2.5-20 μM EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. o, p Representative immunofluorescent images (o) and quantitative analysis (p) of pCAG G-CEPIA1er and CMV-Mito4x-GCaMP6f in PANC1 cells treated with 40 or 120 μM arachidonic acid for 1 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown, mean fluorescence intensity was applied for Two-way ANOVA with Tukey’s multiple comparisons test. Scale bar: 10 μm. q mPT detected by Calcein-AM/CoCl2 assay and dissipation of Δψm by JC-1 assay and TMRM staining assay of GRP75-KD BxPC3 cells compared with N.C. group. Cells were treated with 120 μM arachidonic acid for 1 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. r, s Representative images (r) and quantitative analysis (s) of pyroptotic morphology changes in GRP75-KD and N.C. group. Cells were stimulated with 120 μM arachidonic acid for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. t Immunoblots of GSDME, Caspase 3, and Tubulin in cell lysates of GRP75-KD BxPC3 cells compared with the N.C. group. Experimental design as in (r). n = 3 independent experiments, one representative image of 3 independent experiments is shown. u LDH release assay of GRP75-KD BxPC3 cells compared with the N.C. group. Cells were stimulated with 120 μM arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. v mPT detected by Calcein-AM/CoCl2 assay and dissipation of Δψm by JC-1 assay and TMRM staining assay. BxPC3 cells were pre-treated with 5 μM Ru360 for 2 h followed by 120 μM arachidonic acid treatment for 1 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. w, x Representative images (w) and quantitative analysis (x) of pyroptotic morphology changes in Ru360 pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 5 μM Ru360 for 2 h followed by 120 μM arachidonic acid treatment for 2 h. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. y Immunoblots of GSDME, Caspase 3, and HSP90 in cell lysates of Ru360 pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 5 μM Ru360 for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. z LDH release assay of Ru360 pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 5 μM Ru360 for 2 h followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. Tubulin, HSP90, and GAPDH were used as loading controls. Densitometric quantification of cleaved GSDME ( ~ 30 kDa, lower band) and cleaved caspase 3 (17/19 kDa, lower band) normalized to loading control (Two distinct bands on the membrane). Bar graphs expressed as mean ± SEM. See also Supplementary Figs. 5–8. Source data are provided as a Source Data file.
We proposed endoplasmic reticulum (ER)-mitochondrial Ca2+ transfer (spanning tens of nanometers) as the mechanistic basis, distinct from ionomycin-induced global cytosolic Ca2+ elevation that triggers physiological mPT without cell death (Supplementary Fig. 4a, b). Therefore, we employed pCAG G-CEPIA1er (ER luminal Ca2+ indicator)37 and CMV-Mito4x-GCaMP6f (mitochondrial matrix-targeted Ca2+ indicator)38 to characterize arachidonic acid-induced interorganellar calcium transfer. Fluorescence imaging revealed simultaneous ER Ca2+ marked reduction and mitochondrial Ca2+ accumulation (Fig. 4o, p). Moreover, the Inositol 1,4,5-Trisphosphate Receptor Type 3 (IP3R3)-Glucose-Regulated Protein 75 (GRP75)-Voltage-Dependent Anion Channel 1 (VDAC1) complex, a core structural component of ER-mitochondria contact sites, mediated the ER-mitochondrial Ca2+ transfer process36,39. GRP75 knockdown (Supplementary Fig. 6a, b) reduced mPTP formation, and Δψm collapse (Fig. 4q) and subsequent pyroptotic events (Fig. 4r–u and Supplementary Fig. 6c). Following Ca2+ traverses the outer mitochondrial membrane via VDAC channels, subsequent mitochondrial matrix Ca2+ uptake – essential for mPT activation – occurs through the mitochondrial calcium uniporter (MCU) complex33,40. This MCU-dependent Ca2+ influx proved critical for arachidonic acid-induced mPTP formation and pyroptosis, as pharmacological blockade of MCU with Ru36041 abolished mPT activation (Fig. 4v), GSDME cleavage, and pyroptotic outcomes (Fig. 4w–z and Supplementary Fig. 6d). In addition, genetic ablation of MCU (Supplementary Fig. 6e, f) similarly attenuated arachidonic acid-induced mPTP formation (Supplementary Fig. 6g) and Δψm collapse (Supplementary Fig. 6h, i), GSDME cleavage, and pyroptosis (Supplementary Fig. 6j–n). These results suggest that arachidonic acid triggers pyroptosis by mediating Ca2+-induced mPT, with the source of Ca2+ likely being the rapid release of ER induced by arachidonic acid.
Given its role as a bioactive lipid13, we further investigated whether arachidonic acid metabolism via cyclooxygenase (COX), lipoxygenase (LOX), or P450 epoxygenase pathways contributes to mPT-pyroptosis induction. Heatmap analysis revealed distinct transcriptomic profiles in the high-concentration arachidonic acid treatment group (Supplementary Fig. 7a). Surprisingly, no differential expression of genes associated with arachidonic acid metabolism pathways was observed between the high-concentration group and other conditions (Supplementary Fig. 7b, c). KEGG and GO analyzes of differentially expressed genes in the high-dose arachidonic acid treatment group compared with other groups were also performed (Supplementary Fig. 7d–g). While Prostaglandin-Endoperoxide Synthase 2 (PTGS2) expression was elevated in both low- and high-concentration groups, no intergroup variation was observed (Supplementary Fig. 7c). To determine whether downstream products of arachidonic acid’s cyclooxygenase pathway mediate pyroptosis induction, we administered the COX2 inhibitor acetaminophen42. However, pretreatment failed to alter arachidonic acid-induced mPTP formation (Supplementary Fig. 8a) or pyroptotic outcomes (Supplementary Fig. 8b–f). While acyl-CoA synthetase long-chain family member 4 (ACSL4) typically catalyzes arachidonic acid esterification into phospholipids43. Of interest, knockdown of ACSL4 (Supplementary Fig. 8g, h) paradoxically slightly enhanced mPTP formation (Supplementary Fig. 8i) and pyroptosis (Supplementary Fig. 8j–n) following the arachidonic acid exposure. These results suggest that mPT-induced pyroptosis upon arachidonic acid treatment is not related to arachidonic acid metabolism or esterification.
Non-autophagic function of mitophagy induced by arachidonic acid determines mPT-triggered pyroptosis
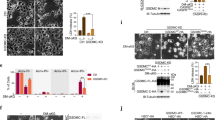
Necrosis is not the sole consequence of irreversible mPT, as noted earlier. However, we rarely observed cell death exhibiting apoptotic characteristics under arachidonic acid treatment (Fig. 1). We further investigated what determines the dominant Caspase 3/GSDME axis, distinguishing it from chemotherapy-induced Caspase 3-mediated apoptosis9 (Supplementary Fig. 2), following pro-apoptotic factor release via irreversible mPTP. Notably, TEM analysis demonstrated marked mitophagy and autolysosome formation induced by high-dose arachidonic acid treatment. (Supplementary Fig. 9a), aligning with prior studies linking mitochondrial Ca2+ overload to autophagic activation and mPTP-associated mitophagy44. Given that autophagy (including mitophagy) regulates nearly all cell death modalities45,46, and can shift from apoptosis to necroptosis in specific contexts47, we hypothesized that Ca2+-driven mitophagy might promote pyroptosis commitment. Indeed, Microtubule-associated protein 1 light chain 3-II (LC3-II) levels and mitophagic flux increased under high- but not low-concentration arachidonic acid treatment (Fig. 5a, b and Supplementary Fig. 9b, c). Furthermore, arachidonic acid significantly enhanced mitochondrial LC3 translocation (Fig. 5c, d), paralleled by reduced mitochondrial DNA (mtDNA) content (Fig. 5e). Importantly, this stress-induced mitophagy occurs independently of Caspase 3 cleavage and mPTP formation (Supplementary Fig. 9d–h), contrasting with previous research48. Conversely, BAPTA-AM blocked mitophagy (Fig. 5f, g and Supplementary Fig. 9i–k), suggesting concurrent but non-interacting Ca2+-mediated mPTP formation and mitophagy induction.
a Representative fluorescent images of BxPC3 cells with stable transfection of pLent-EF1a-cox8-eGFP-mCherry-CMV-Puro lentivirus. Cells were stimulated with 40 or 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 10 μm. b Immunoblots of LC3B, Tubulin, and HSP90 in cell lysates of arachidonic acid-treated BxPC3, SW1990, and Pan02 cells. Cells were stimulated with 40 or 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. c, d Representative immunofluorescent images (c) and quantitative analysis (d) of SW1990 cells with transfection of LC3-eGFP and MitoTracker Red staining. Cells were stimulated with 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown, Pearson’s Correlation Coefficient (PCC) and Manders’ Colocalization Coefficient (MCC) were applied for Two-way ANOVA with Tukey’s multiple comparisons test. e mtDNA expression levels in BxPC3 cells treated with 40 or 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. f Representative fluorescent images of pLent-EF1a-cox8-eGFP-mCherry-CMV-Puro lentivirus stably transfected BxPC3 cells pre-treated with BAPTA-AM/EGTA-AM. Cells were pre-treated with 20 μM BAPTA-AM/EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 10 μm. g Immunoblots of LC3B and Tubulin in cell lysates of BAPTA-AM/EGTA-AM pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 2.5–20 μM BAPTA-AM/EGTA-AM for 2 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. h Representative fluorescent images of pLent-EF1a-cox8-eGFP-mCherry-CMV-Puro lentivirus stably transfected BxPC3 cells pre-treated with chloroquine. Cells were pre-treated with 50 μM chloroquine for 24 h followed by 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 10 μm. i, j Representative images (i) and quantitative analysis (j) of pyroptotic morphology changes in chloroquine pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 50 μM chloroquine for 24 h followed by 120 μM arachidonic acid for 75 min. The pyroptotic cells were marked with yellow arrowheads. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 50 μm. k Immunoblots of GSDME, Caspase 3, LC3B, and β-actin in cell lysates of chloroquine pre-treated BxPC3 cells. Cells were pre-treated with 50 μM chloroquine for 12 or 24 h followed by 40 or 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. l LDH release assay of chloroquine pre-treated BxPC3 cells. BxPC3 cells were pre-treated with 50 μM chloroquine for 24 h followed by 40 or 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, Two-way ANOVA with Tukey’s multiple comparisons test. m Time courses of immunoblots of GSDME, Caspase 3, LC3B, and Tubulin in cell lysates. BxPC3 cells were pre-treated with 50 μM chloroquine for 24 h, followed by 120 μM arachidonic acid treatment. n = 3 independent experiments, one representative image of 3 independent experiments is shown. n Time courses of LDH release assay of chloroquine pre-treated BxPC3 cells. Experimental design as in (m). n = 3 independent experiments, Two-way ANOVA with Tukey’s multiple comparisons test. o, p Representative immunofluorescent images (o) and quantitative analysis (p) of PINK1-KD SW1990 cells with transfection of LC3-eGFP and MitoTracker Red staining. Cells were stimulated with 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown, Pearson’s Correlation Coefficient (PCC) and Manders’ Colocalization Coefficient (MCC) were applied for Two-way ANOVA with Tukey’s multiple comparisons test. q, r Representative images (q) and quantitative analysis (r) of pyroptotic morphology changes in PINK1-KD SW1990 cells. The pyroptotic cells were marked with yellow arrowheads. Cells were stimulated with 120 μM arachidonic acid for 2 h. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm.(s) Immunoblots of GSDME, Caspase 3, LC3B, and Tubulin in cell lysates of PINK1-KD SW1990 cells compared with N.C. group. Experimental design as in (q). n = 3 independent experiments, one representative image of 3 independent experiments is shown. t LDH release assay of PINK1-KD SW1990 cells compared with the N.C. group. Cells were stimulated with 120 μM arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. u Immunoblots of GSDME, Caspase 3, LC3B, and Tubulin in cell lysates of chloroquine and CCCP pre-treated BxPC3 cells. Cells were pre-treated with 50 μM chloroquine and 10 μM CCCP for 6 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. v LDH release assay of chloroquine and CCCP pre-treated BxPC3 cells. Cells were pre-treated with 50 μM chloroquine and 10 μM CCCP for 6 h followed by 120 μM arachidonic acid treatment for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. HSP90, Tubulin, β-actin, and GAPDH were used as loading controls. Densitometric quantification of cleaved GSDME (~ 30 kDa, lower band) and cleaved caspase 3 (17/19 kDa, lower band) normalized to loading control (Two distinct bands on the membrane). Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 8. Source data are provided as a Source Data file.
We also sought to determine whether mitophagy is required for arachidonic acid-induced pyroptosis. To this end, chloroquine (CQ), a late-stage autophagy inhibitor that blocks degradation, was applied to block autophagic flux in BxPC3 cells. Consistent with impaired of mitophagic flux (Fig. 5h and Supplementary Fig. 9m), CQ treatment enhanced the activation of Caspase 3, cleavage of GSDME, and pyroptosis induction (Fig. 5i–l and Supplementary Fig. 9m–p). Time course analysis revealed that CQ pretreatment not only accelerated the onset but also amplified the magnitude of arachidonic acid-induced pyroptosis (Fig. 5m, n and Supplementary Fig. 9q, s). However, in contrast to the potentiating effects of CQ, genetic ablation of PTEN-induced putative kinase 1 (PINK1) (Supplementary Fig. 9t, u)- a key mediator of mitochondrial Ca2+ oscillation-dependent mitophagy initiation- effectively suppressed arachidonic acid-induced mitophagy (Fig. 5o, p) without altering mPTP opening (Supplementary Fig. 9v), and subsequently inhibited GSDME cleavage and subsequent pyroptosis (Fig. 5q–t and Supplementary Fig. 9w). We next explored the regulation of mitophagy activation on pyroptosis using Carbonyl Cyanide m-Chlorophenylhydrazone (CCCP), a mitochondrial uncoupler that induces PINK1/PARKIN-dependent mitophagy49. Unexpectedly, CCCP treatment potentiated arachidonic acid-induced pyroptosis, with maximal enhancement observed when combined with CQ – a mitophagy agonist-antagonist pair (Fig. 5u, v and Supplementary Fig. 9x, y). Neither CCCP nor CQ significantly affects mitochondrial Ca2+-induced mPTP formation (Supplementary Fig. 9z).
These data demonstrate that Ca2+-induced mitophagy participates in mPT-mediated specific pyroptosis. Crucially, this process depends on mitophagy’s non-autophagic functionality rather than autophagic cargo degradation, as CCCP-induced complete mitophagy activation would otherwise suppress arachidonic acid-induced pyroptosis if the cargo clearance were mechanistically relevant, as well as the promotion effect followed by the blockage of all stages of mitophagy.
Autophagosome formation provides a scaffold for GSDME activation and promotes arachidonic acid induced pyroptosis
Autophagy-related proteins mediating autophagy-independent functions have emerged as key regulators of regulated cell death (RCD)50,51,52. Given the contradictory effects of early- versus late- stage mitophagy modulation, we posited that mitophagy machinery might provide scaffold for pyroptosis signaling rather than execute cargo degradation. Indeed, arachidonic acid enhanced the interaction between LC3 (a core autophagosome structural component) and both Caspase 3 and GSDME (Fig. 6a and Supplementary Fig. 10a). Increased LC3 colocalization with Caspase 3 and GSDME was observed following arachidonic acid stimulation, an effect abolished by BAPTA-AM (Fig. 6b). In addition, LC3 knockdown (Supplementary Fig. 10b, c) eliminated arachidonic acid-induced Caspase 3/GSDME cleavage, pyroptosis, and LDH release (Fig. 6c–f and Supplementary Fig. 10d), whereas LC3 overexpression exerted opposing effects (Supplementary Fig. 10e–g). The E3-like activity of Autophagy-related gene 5 (Atg5) catalyzes the lipid conjugation of LC3 to phosphatidylethanolamine, essential for autophagosome formation53. Here, we found that autophagosome generation is critical for pyroptosis induction, as LC3 lipidation inhibited by knockdown of Atg5 (Supplementary Fig. 10h, i) impaired arachidonic acid-driven Caspase 3/GSDME cleavage and pyroptosis (Fig. 6g–j and Supplementary Fig. 10j) without affecting mPTP formation (Supplementary Fig. 10k). Furthermore, CCCP/CQ-enhanced GSDME cleavage and pyroptosis under arachidonic acid treatment were also abrogated upon Atg5 deletion (Fig. 6k, l and Supplementary Fig. 10l). LC3 lipidation inhibition disrupted LC3-Caspase 3/GSDME interactions (Fig. 6m, n and Supplementary Fig. 10m), while CQ strengthened these associations (Fig. 6o and Supplementary Fig. 10n). Notably, poly (ADP-ribose) polymerase 1 (PARP1, a key apoptosis effector) showed no autophagosomal binding despite activation (Supplementary Fig. 10o). LC3 overexpression similarly enhanced Caspase 3/GSDME interactions during arachidonic acid exposure (Supplementary Fig. 10p, q). Collectively, these findings demonstrate that autophagosome formation scaffolds Caspase 3/GSDME recruitment to enable rapid pyroptosis.
a Immunoprecipitation analysis of GSDME, Caspase 3, and LC3 from lysates of BxPC3 cells. Cells were treated with 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. b Representative immunofluorescent images and quantitative analysis of GSDME, Caspase 3, and LC3 in 120 μM arachidonic acid treated BxPC3 cells for 1.5 h with/without 20 μM BAPTA-AM pre-treatment. Scale bar: 2 μm. n = 3 independent experiments, one representative image of 3 independent experiments is shown, Pearson’s Correlation Coefficient (PCC) and Manders’ Colocalization Coefficient (MCC) were applied for Two-way ANOVA with Tukey’s multiple comparisons test. c, d Representative images (c) and quantitative analysis (d) of pyroptotic morphology changes in LC3-KD BxPC3 cells. Cells were stimulated with 120 μM arachidonic acid for 2 h. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. e Immunoblots of GSDME, Caspase 3, LC3, and Tubulin in cell lysates of LC3-KD BxPC3 cells compared with the N.C. group. Experimental design as in (c). n = 3 independent experiments, one representative image of 3 independent experiments is shown. f LDH release assay of LC3-KD BxPC3 cells compared with the N.C. group. Cells were stimulated with 120 μM arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. g, h Representative images (g) and quantitative analysis (h) of pyroptotic morphology changes in Atg5-KD BxPC3 cells. Cells were stimulated with 120 μM arachidonic acid for 2 h. n = 5 independent experiments, one representative image of 5 independent experiments is shown, One-way ANOVA with Dunnett’s multiple comparisons test. Scale bars, 25 μm. i Immunoblots of GSDME, Caspase 3, LC3, and Tubulin in cell lysates of Atg5-KD BxPC3 cells compared with N.C. group. experimental design as in (g). n = 3 independent experiments, one representative image of 3 independent experiments is shown. j LDH release assay of Atg5-KD BxPC3 cells compared with the N.C. group. Cells were stimulated with 120 μM arachidonic acid for 4 h. n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. k Immunoblots of GSDME, Caspase 3, LC3, and HSP90 in cell lysates of chloroquine and CCCP pre-treated Atg5-KD BxPC3 cells compared with the N.C. group. Cells were pre-treated with 50 μM chloroquine and 10 μM CCCP for 6 h followed by 120 μM arachidonic acid treatment for 2 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. l LDH release assay of Atg5-KD BxPC3 cells compared with the N.C. group. Experimental design as in (k). n = 3 independent experiments, One-way ANOVA with Dunnett’s multiple comparisons test. m Immunoprecipitation analysis of GSDME, Caspase 3, and LC3 from lysates of Atg5-KD BxPC3 cells compared with the N.C. group. Cells were treated with 120 μM arachidonic acid for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. n Representative immunofluorescent images and quantitative analysis of GSDME, Caspase 3, and LC3 in 120 μM arachidonic acid treated Atg5-KD BxPC3 cells compared with the N.C. group. Scale bar: 2 μm. n = 3 independent experiments, one representative image of 3 independent experiments is shown, Pearson’s Correlation Coefficient (PCC) and Manders’ Colocalization Coefficient (MCC) were applied for Two-way ANOVA with Tukey’s multiple comparisons test. o Immunoprecipitation analysis of GSDME, Caspase 3, and LC3 from lysates of chloroquine pre-treated BxPC3 cells. Cells were pre-treated with 50 μM chloroquine for 24 h followed by 120 μM arachidonic acid treatment for 1.5 h. n = 3 independent experiments, one representative image of 3 independent experiments is shown. (p, q) Representative fluorescent images (p) and quantitative analysis (q) of Calcein-AM and PI staining in 200 μM arachidonic acid treated PDAC organoid with/without chloroquine pre-treated. n = 6 independent experiments, one representative image of 6 independent experiments is shown, mean fluorescence intensity was applied for Two-way ANOVA with Tukey’s multiple comparisons test. Scale bars, 50 μm. r Longitudinal imaging analysis of Calcein-AM and PI staining in 200 μM arachidonic acid-treated PDAC organoid with/without chloroquine pre-treated. The cell membrane swelling was marked with yellow arrowheads. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 50 μm. s Cell viability of 200 μM arachidonic acid treated PDAC organoid with/without chloroquine pre-treated. Bar graphs expressed as mean ± SEM (n = 3). t, y BxPC3 derived subcutaneous xenograft (n = 7 per group), and patient derived xenograft (PDX) (n = 4 per group) models received chloroquine (CQ) and arachidonic acid (AA) combined treatment. Schematic illustration. q24h, every 24 h (t). Created in BioRender. Fan, Z. (2026) https://BioRender.com/04f0qz9. Images of subcutaneous tumors (u), and the quantification of tumor volume (v and x) and weight (w and y) were recorded at the indicated times. n = 7 independent experiments (CDX), Two-way ANOVA with Tukey’s multiple comparisons test for Tumor volume curve (v) and One-way ANOVA with Dunnett’s multiple comparisons test for Change in Tumor volume (v) and Tumor weight (w). n = 4 independent experiments (PDX), Two-way ANOVA with Tukey’s multiple comparisons test for Tumor volume curve (x) and One-way ANOVA with Dunnett’s multiple comparisons test for Change in Tumor volume (x) and Tumor weight (y). Scale bars, 1 cm. HSP90 and Tubulin were used as loading controls. Densitometric quantification of cleaved GSDME (~ 30 kDa, lower band) and cleaved caspase 3 (17/19 kDa, lower band) normalized to loading control (Two distinct bands on the membrane). Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 9. Source data are provided as a Source Data file.
We next evaluated whether CQ-enhanced autophagosome accumulation potentiates arachidonic acid-induced pyroptosis in PDAC models. In PDAC organoids, arachidonic acid induced pyroptosis (evidenced by Calcein-AM loss and PI staining), markedly enhanced by CQ pretreatment (Fig. 6p, q). Longitudinal imaging analysis of PDAC organoids maintained at identical spatial coordinates demonstrated accelerated pyroptosis under combination therapy (Fig. 6r). Furthermore, a more rapid decline in cell viability was concurrently observed following combination treatment (Fig. 6s). In BxPC3 xenografts, combination therapy (Fig. 6t) significantly reduced tumor volume/weight (Fig. 6u–w) with concomitant GSDME cleavage (Supplementary Fig. 10r, s). Comparable efficacy occurred in PDAC patient-derived xenografts (PDX) (Fig. 6u, x, y). Therefore, these data established that autophagosome accumulation scaffolds Caspase 3/GSDME interactions to potentiate arachidonic acid-driven pancreatic cancer pyroptosis.
Arachidonic acid induced pyroptosis recruits immune cells infiltration and synergizes with anti-PD-1 in immunocompetent PDAC models
PDAC, as the most representative “cold” tumor, is resistant to most therapies including ICB54. The observation that acute pyroptotic tumor cell death stimulates inflammatory responses in the tumor microenvironment and mobilizes potent anticancer immunity suggests that arachidonic acid may activate immunotherapy in pancreatic cancer6,7,17. Supporting this, arachidonic acid administration significantly reduced tumor volume and weight in the immunocompetent C57BL/6 Pan02 pancreatic cancer model (Fig. 7a–e and Supplementary Fig. 11a), while markedly increasing infiltration of cytotoxic cells and CD4+ T cells in tumors (Fig. 7f–j). Increased NK cell infiltration was also observed following arachidonic acid treatment (Supplementary Fig. 11b, c). Concurrently, we observed decreased infiltration of immunosuppressive cells, including Treg cells (Fig. 7k, l) and reduced M2 marker (CD163)-positive cells (Fig. 7m), though myeloid-derived suppressor cell (MDSC) infiltration showed an unexpected increase following arachidonic acid treatment (Supplementary Fig. 11d). Thus, arachidonic acid-induced pyroptosis partially mediates remodeling of the immune microenvironment. Notably, no significant adverse effects were detected in major organs of treated mice (Supplementary Fig. 11e, f), indicating the favorable tolerability of arachidonic acid. Although immune checkpoint blockade, particularly anti-programmed cell death protein 1 (PD1) therapy, has revolutionized cancer treatment, its efficacy remains limited in PDAC55,56, primarily due to the immunosuppressive tumor microenvironment that restricts T lymphocyte infiltration and activation57. Here, when a single course of arachidonic acid pretreatment preceded anti-PD-1 therapy (Fig. 7n), we observed enhanced tumor volume reduction and weight loss compared to monotherapy groups, whereas the Pan02 model showed no response to anti-PD-1 alone (Fig. 7o–r). These results demonstrate that pyroptosis-induced inflammation synergizes with checkpoint blockade to potentiate antitumor immunity.
a–e Mice bearing subcutaneous tumors (Pan02) received arachidonic acid (n = 5 per group). Schematic illustration. q48h, every 48 h (a). Created in BioRender. Fan, Z. (2026) https://BioRender.com/04f0qz9. Images of subcutaneous tumors (b), and the quantification of tumor volume (c, d) and weight (e) were recorded at the indicated times. Cells were stimulated with 120 μM arachidonic acid for 2 h. n = 5, Two-way ANOVA with Tukey’s multiple comparisons test (c) and Unpaired two-tailed Student’s t test (d, e). Scale bars, 1 cm. f Representative immunofluorescent images of CD3+, CD4+, and CD8+ T cells in arachidonic acid treated subcutaneous Pan02 tumors in C57 mice. n = 3 independent experiments, one representative image of 3 independent experiments is shown. Scale bars, 50 μm. g Flow cytometry analysis of CD3+, CD4+, and CD8+ T cells in arachidonic acid treated subcutaneous Pan02 tumors in C57 mice. n = 3 independent experiments, one representative image of 3 independent experiments is shown. h–j Proportion analysis of CD3+ (h), CD4+ (i), and CD8+ (j) T cells, quantified from images as shown in (g). Experimental design as in Fig. 7a. n = 3 independent experiments, Unpaired two-tailed Student’s t test. k–m Proportion analysis of Treg (k, l) and CD163+ Macs (m). Experimental design as in Fig. 7a. n = 5 independent experiments, Unpaired two-tailed Student’s t test. n–r Mice bearing subcutaneous tumors (Pan02) received anti-PD-1 and arachidonic acid combined treatment. (n = 6 per group). Schematic illustration. q48h, every 48 h (n). Created in BioRender. Fan, Z. (2026) https://BioRender.com/04f0qz9. Images of subcutaneous tumors (o), and the quantification of tumor volume (p, q) and weight (r) were recorded at the indicated times. n = 6, Two-way ANOVA with Tukey’s multiple comparisons test (p) and One-way ANOVA with Dunnett’s multiple comparisons test (q, r). Scale bars, 1 cm. Bar graphs expressed as mean ± SEM. See also Supplementary Fig. 10.
Discussion
Despite growing recognition of pyroptosis as a promising strategy for cancer treatment9,58 and an immunogenic cell death mechanism6,7, achieving efficient induction remains a critical challenge in therapeutic development. Our study addresses this unmet need through computational prediction and multi-model validation, establishing the previously unrecognized capacity of ω-6 PUFA arachidonic acid as a potent pyroptosis inducer with pan-cancer efficacy, which is distinctive from conventional chemotherapy-induced apoptosis and secondary pyroptosis (Fig. 1 and Supplementary Figs. 1, 2). Mechanistically, we demonstrate that the non-autophagic function of mitophagy50 mediates arachidonic acid-induced mPT to selectively trigger acute pyroptosis, thereby providing a theoretical basis for synergistic therapies. Crucially, our integrative approach employing PDAC organoids, PDX, cell line-derived xenograft (CDX) models, and immunocompetent syngeneic models provides translational relevance rarely achieved in single-model studies, while resolving key mechanistic questions about pyroptosis regulation. This finding expands the therapeutic repertoire for PDAC beyond traditional cell death modalities while addressing the critical need for immunotherapy-sensitizing strategies in this recalcitrant malignancy.
A remarkable aspect suggested by our research is the elucidation of mitophagy’s non-degradative function in pyroptosis execution. Unlike canonical mitophagy that eliminates damaged mitochondria, mitophagic machinery is repurposed to scaffold caspase 3/GSDME activation. Arachidonic acid triggers both irreversible mPTP formation and autophagosome formation, which was previously demonstrated to be involved in kinds of necrotic cell death with plasma membrane disruption, including ferroptosis in ways other than autophagy59,60,61,62(Figs. 3–5). Crucially, autophagosomes generated during this process serve as physical platforms for recruiting and activating caspase 3/GSDME, significantly contribute to mPT-induced specific pyroptosis rather than autophagic flux (Figs. 5, 6), which potentially parallel the previously reported necroptosis-promoting mechanisms51,52. This mechanistic divergence explains why pharmacological (CQ and CCCP) and genetic (PINK1, Atg5 knockout) perturbations of mitophagy flux paradoxically enhance pyroptosis (Figs. 5, 6)—a hallmark of non-autophagic function of mitophagy.
Translational implications are preclinically underscored by the synergistic tumor suppression achieved through chloroquine combination therapy (Fig. 6)—a promising strategy applicable to ongoing clinical trials with autophagy modulators. The observed immune microenvironment remodeling mediated by pyroptosis, characterized by cytotoxic T cell infiltration and PD-1 checkpoint sensitization (Fig. 7), which is consistent with prior research15,16,63,64, suggesting arachidonic acid’s dual activity: direct tumoricidal effects and immune microenvironment modulation in preclinical settings.
Collectively, our work establishes arachidonic acid as a potent and selective pyroptosis inducer capable of remodeling immunosuppressive tumor microenvironments and synergizing with ICB in preclinical PDAC models (Fig. 8). Pan-cancer efficacy in vitro suggests broad pyroptosis-inducing potential, while the discovery of autophagosomal membrane scaffolding—a non-canonical mechanism enabling caspase 3/GSDME recruitment and activation—represents a meaningful shift in understanding pyroptosis regulation. These findings propose strategies to address two key challenges: (1) the lack of tumor-selective pyroptosis inducers with rapid action kinetics, and (2) the mechanistic ambiguity surrounding mitochondrial permeability transition (mPT)-mediated cell fate decisions.
Created in BioRender. Fan, Z. (2026) https://BioRender.com/w8r37is.
Despite these advances, translational challenges remain. Although the therapeutic potential of arachidonic acid in cancer treatment, current systemic administration paradigms demonstrate suboptimal therapeutic efficacy due to inadequate tumor targeting and off-site biodistribution. This limitation underscores the urgent need to develop clinically translatable delivery platforms—such as tumor-responsive nanoparticle formulations or ligand-engineered exosomes—capable of achieving spatially controlled enrichment of arachidonic acid within malignant tissues while circumventing systemic exposure. Notably, while arachidonic acid treatment reduces immunosuppressive elements like Tregs and M2 macrophages, we paradoxically observed increased MDSC infiltration—a cell population known to exert pro-tumorigenic immunosuppressive effects. This compensatory response may partially counteract the immunotherapeutic benefits, suggesting that combinatorial strategies targeting MDSC recruitment or function could further enhance immune microenvironment remodeling. Furthermore, the molecular architecture of autophagosomal membranes facilitating pyroptosis-related protein interactions remains undefined, necessitating advanced proteomic and structural analyzes. Intriguingly, given chloroquine’s dual role in enhancing pyroptosis (this study) and potentiating ICB efficacy65, future investigations should explore whether combined arachidonic acid/chloroquine treatment amplifies immunogenic cell death cascades in “cold” tumors.
Thus, our results will stimulate a bridge of interest in exploring the feasibility of endogenous metabolites to activate pyroptosis-immune synergy. The non-autophagic repurposing of mitophagy revealed here opens new avenues for targeting autophagosome biogenesis in pyroptotic cancer therapy—a strategy transcending traditional autophagy modulation paradigms.
Methods
Contrastive learning graph neural network
An extensive screening of 10,533 chemicals using CTD, identifying 103,232 known chemical-disease associations across 3309 diseases. Then, the Chemical-Disease Knowledge Graph (CDKG) are developed to assess the prediction performance of our proposed model against baseline models. In this process, to alleviate the sparsity of the knowledge graph, diseases with less than 10 associations are further excluded in this paper. The resulting CDKG comprehensively maps the association relationships between 1329 diseases and 10,357 chemicals. Graph neural networks (GNNs), which can capture graph structure information and node attributes through local message aggregation and propagation mechanisms, have attracted significant attention in the bioinformatics community. To achieve information passing between chemical and disease nodes in CDKG and mitigate graph sparsity, we proposed CLGNN, a graph neural network model based on self-supervised contrastive learning. The CLGNN predictor module consists of a GNN optimized on the relationships in the CDKG. Moreover, we introduced a self-supervised contrastive learning task that enables robust node embedding learning without relying on explicit labels, simultaneously serving as a regularization technique in a graph learning framework to enhance the model’s generalization capabilities. Finally, the neural network was performed to predict the strength of interaction between query chemicals and diseases.
Cell lines
Human PDAC cell lines PANC1, SW1990, BxPC3, human CCA (Cholangiocarcinoma) cell line RBE, human HCC cell line HepG2, human LC cell line A549, human BLCA cell line T24, human cervical cancer cell line Siha, human glioma cell line A172, and mouse PDAC cell line Pan02 were obtained from the RIKEN Bioresource Center (Riken Cell Bank, Japan). Human CRC cell line SW480 were kindly granted by Prof. Quan Wang from the First Hospital of Jilin University. All cell lines were authenticated using short tandem repeat (STR) analysis and tested for mycoplasma contamination, and referenced against databases from the Chinese Academy of Sciences and the American Type Culture Collection. The human cell lines PANC1, HepG2, SW480, MCF-7, Siha, A172, A549 and mouse PDAC cell line Pan02 were cultured in Dulbecco’s Modified Eagle Medium (DMEM)-high glucose (Gibco) supplemented with 10% fetal bovine serum (FBS), 100 IU penicillin and 100 mg/mL streptomycin (Bio Basic Inc.). BxPC3, RBE, and T24 cells were cultured in RPMI1640 supplemented with 10% FBS. SW1990 cells were maintained in DME/F-12 1:1 supplemented with 10% FBS. All cell lines in our laboratory are routinely tested for mycoplasma contamination, and the cells used in this study were negative for mycoplasma. After culture, cancer cells were washed extensively in serum-free medium and incubated in medium supplemented with 0.1% fatty acid-free BSA before arachidonic acid treatment. Cells were treated with arachidonic acid (different concentrations) for 2–4 h to induce pyroptosis, unless otherwise specifically described.
Animal models
Female nude mice (BALB/c, 18–22 g, 4–6 weeks old) and C57BL/6 N mice were obtained from the Beijing Vital River Laboratory Animal Technology (Beijing, China), and maintained in a 12 h light/dark cycle with free access to food and water. All mice were specific pathogen-free and quarantined for one week before starting the experiments. All animal experiments were approved by the Animal Ethics Committee of the First Hospital of Jilin University (acceptance number: 20241010-010).
For the subcutaneous xenograft or syngeneic models, 5 × 106 BxPC3 cells or 5 × 106 Pan02 cells were suspended in 100 μL of PBS and then injected into the anterior flanks of nude mice (BxPC3 cells) or C57BL/6 N (Pan02 cells). For PDAC patient-derived xenografts (PDX) models, passage 2-3 PDX models were used to evaluate the efficiency of various treatment regimens for PDAC. When the tumor long diameter reached approximately 0.8–1.0 cm, mice were randomly divided into groups with 4 mice per group. Tumor diameters were measured using calipers. Mice were randomly assigned to specific treatment groups at the beginning of treatment and were euthanized at the indicated time points, and tumors or other organs were collected.
For arachidonic acid treatment, mice received treatment with either vehicle (PBS, 100 μL per mouse) or arachidonic acid (peroxide free) (C20H32O2) (Cayman Chemical, 90010.1) (200 mg/kg), administered via intratumoural injections every other day for 10 days unless otherwise indicated. For chloroquine treatment, mice received intraperitoneal injection of chloroquine (MedChemExpress, H-17589A-500mg) solution in PBS at 60 mg kg−1 or PBS every day starting day 7 after tumor cell implantation. For immune checkpoint blockade experiments, mice received intraperitoneal injection of anti-mouse PD-1 antibody (100 μg, clone RMP1-14, BE0146) or rat IgG2a isotype control (100 μg, clone 2A3, BE0089) every three days for four times. All antibodies used in in vivo experiments were obtained from BioXCell.
PDAC patients and ethics
This study utilized specimens from 6 patients with PDAC undergoing surgery for organoid culture and PDX models building. All patients underwent surgical resection at the First Hospital of Jilin University. Sample selection was based on the following criteria: (1) Patients who underwent curative-intent resection with negative surgical margins, (2) Patients with well-preserved paraffin-embedded tumor tissues, comprehensive follow-up information, and complete medical records, (3) Patients who survived more than one month postoperatively, (4) Patients without a history of other malignant tumors. All patients gave informed consent for specimen acquisition and data analysis. The Ethics Committee of the First Hospital of Jilin University approved and oversaw the research protocol (24K330-001).
Graph Neural Networks for Knowledge Graph Encoding
For encoding Graph Neural Networks for Knowledge Graph, the CDKG can be viewed as an undirected graph \(G=\left(V,E,X\right)\), where \(V\) denotes the set of nodes encompassing molecular entities (chemicals and diseases), and \(E\) represents the set of edges in which there is an interaction between the two entities in \(V\). \(X\in {R}^{\left|V\right|\times d}\) is the feature matrix, where each node in \(V\) is encoded as a predefined \(d\)-dimensionality as an attribute vector. Specifically, the knowledge graph can be mathematically represented through its adjacency matrix \(A\in {R}^{\left|V\right|\times \left|V\right|}\). Then, GNN leverage both graph structure and node attributes to derive a high-level, compact node representation, denoted as \(H\in {R}^{\left|V\right|\times m}\) where \({h}_{v}\) symbolizes the potential representation of a specific entity \(v\). The GNN employs an iterative approach to update node representations by aggregating information from neighboring nodes. Following \(l\) iterations, the hidden state of node \(v\) can be represented as:
where \(N(v)\) denotes the set of nodes connected to entity \(v\), and \({AGG}\) refers to the aggregation function of layer \(l\). \({\sigma }^{\left(l\right)}\) denotes the combination function (e.g., summation, a neural network, etc.) of layer \({l}\). \({h}_{v}^{(l)}\) represents the node embedding of layer \(l\), and \({h}_{v}^{\left(0\right)}=X\left[v,:\right]\).
For chemical-disease association strength prediction, this module takes as inputs chemical \(i\) embedding \({h}_{i}\) and disease \(j\) embedding \({h}_{j}\) learned by the GNN, then computes the probability of their interactions \({p}_{{ij}}\). To fully integrate the embedded features of the entities, we perform multiple operation decoders to model the relationship between chemicals and diseases in the following process:
where \(\odot\) denotes element-wise multiplication, \({||}\) represents vector concatenation. The \({MLP}\) classifier is composed of a two-layer fully connected neural network. The corresponding supervised loss can be formulated as:
where \({y}_{{ij}}\) is the true association relationship between chemical \(i\) and disease \(j\). \(T\) is the training set of node pairs.
For self-supervised contrastive learning, the aim of this paper is to design a chemical-disease association prediction framework for a sparse knowledge graph. Therefore, to address this challenge, CLGNN introduces a self-supervised learning task designed to enhance the extraction of chemical and disease embeddings. Following DGI66, the self-supervised auxiliary task aims to maximize the mutual information between a local patch (i.e., node-level representation) and its global context (i.e., graph-level representation). We model the self-supervised learning task using a binary cross-entropy (BCE) loss function that compares positive pairs (each consisting of a local patch and global information) against negative pairs (each consisting of a local patch and global information of the corrupted graph), as follows:
where \({h}_{i}\) and \({\widetilde{h}}_{j}\) represent the node local embedding representations of the original and corrupted knowledge graphs, respectively. The global representation \(c\) is calculated as \(c={MLP}({sigmoid}(\frac{1}{\left|V\right|}{\sum }_{i=1}^{\left|V\right|}{h}_{i}))\). The discriminator \(D(\cdot )\), implemented as a bilinear layer, assigns probability scores to pairs of local patches and global context. Following DGI, the corrupted knowledge graph is generated by randomly shuffling features through the corruption function as follows:
To some extent, the contrastive learning mechanism effectively maximizes the mutual information between original global information and local patches, thereby enhancing CLGNN’s discriminative capabilities in node representation learning. Finally, CLGNN combines supervised prediction task and self-supervised learning task into an overall loss function for training:
where \(\lambda\) is a trade-off hyperparameter that modulates the relative importance of different loss components.
Cytotoxicity Assays (LDH release assay)
Pyroptosis was analyzed by detecting the activity of LDH released into cell culture supernatants using the CytoTox 96 Non-Radioactive Cytotoxicity Assay Kit (Promega) according to the manufacturer’s protocol.
siRNA Transfection
Gene knockdown was achieved using small interfering RNA. SiRNA was transfected into cells with Lipofectamine 3000 (L3000001, Invitrogen) according to instructions. Briefly, 500 μL of Opti-MEM medium, 3–5 μL of Lipofectamine 3000 and 20 μM siRNA were premixed per well in a 6-well plate. BxPC3 or SW1990 cells were plated at a density of 5 × 105 per well, the optimal transfection time was assessed by the maximal silencing efficiency evidenced from western blot analysis, and cells were used for assays 48 h after transfection. The sequences of siRNA were listed in the Supplementary Table 2.
Plasmid transfection
Gene overexpression was achieved using plasmids, which originated from JTSBIO Co., Ltd (Wuhan, China). BxPC3 cells were seeded in a 6-well plate at a density of 3 × 105/well and plasmids were transfected into cells with Lipofectamine 3000/P3000 following the manual’s instructions. Briefly, 2 μg plasmid, 5 μL P3000 and 5 μL Lipofectamine 3000 were mixed into 500 μL Opti-MEM medium in order, then added up to 2 mL with RPMI1640 medium before transferring to 6-well plate. 6 h after transfection, the cell culture was replaced with RPMI1640 medium containing 10% FBS, and cells were used for assays 48 h after transfection.
CRISPR/Cas9 targeted GSDME-KO cell lines construction was conducted as previously described67. Briefly, a target-specific sgRNA targeting the GSDME gene was designed and synthesized. The sgRNA and Cas9 expression plasmid were cotransfected into the target cells using a lipid-based transfection reagent. After 48 hours, cells were subjected to puromycin selection to obtain stably transfected cells. Single-cell clones were then isolated by limiting dilution and expanded. Western blot analysis was performed to confirm the knockout of the GSDME. The sequence of the plasmid was listed in the Supplementary Table 2.
Generation of the lentiviral system
The pLent-EF1a-cox8-eGFP-mCherry-CMV-Puro lentivirus was originated from WZ Biosciences (Shandong province, China). BxPC3 cells were seeded in 24-well plates at a density of 1 × 105 cells/ well 24 h prior to transduction to achieve 40–60% confluence. Polybrene (6–8 μg/mL, Sigma-Aldrich) was added to enhance transduction efficiency by neutralizing electrostatic repulsion between viral particles and cell membranes. A multiplicity of infection (MOI) gradient (0, 1, 2, 3, 5, 10) was tested using serial dilutions of lentiviral supernatant. After 4 h of incubation in serum-free medium, the viral suspension was replaced with complete medium containing 10% fetal bovine serum (FBS). Fluorescence microscopy was performed at 48–72 h post-transduction to assess eGFP/mCherry expression. The optimal MOI (MOI = 20) was selected based on > 80% fluorescent-positive cells with minimal cytotoxicity. At 24 h post-transduction, cells were subjected to puromycin (1.5 μg/mL, Invitrogen) selection. The puromycin concentration was predetermined via a kill curve assay, where untransduced BxPC3 cells were completely eliminated within 48 h at this concentration. The selection medium was refreshed every 2–3 days for 10–14 days until resistant colonies formed. To ensure monoclonality, individual colonies were isolated using cloning rings and expanded in 6-well plates. Stable clones were validated for sustained cox8-eGFP-mCherry expression via fluorescence microscopy. The loss of eGFP fluorescence indicates the occurrence of mitophagic flux.
Co-immunoprecipitation
Protein samples were prepared following Western blotting protocols until loading buffer addition. For studying the interaction between GSDME, Caspase 3, LC3, and PARP1, each 1 mg protein sample was incubated with 2 μg of the primary antibody at 4 °C overnight. The rProtein G MagPoly beads were then added and incubated at 4 °C for 4 h. Subsequently, sediments were collected post-centrifugation at 20,000 × g and 4˚ C for 15 min and washed three times with lysis buffer and then subjected to western blotting.
Flow cytometry
For Annexin-V/PI staining analysis, 2 × 106 cells of each group in 6-well plate were collected with 0.25% trypsin without EDTA (T1350, Solarbio), centrifuged at 600 rpm for 5 min and washed pellet once with PBS. Cells were resuspended with 200 μL 1 × binding buffer and stained with PI and Annexin-V from Annexin V-FITC/PI Apoptosis Detection Kit (HY-K1073, MCE) for 15–20 min at room temperature. Cells were collected and underwent flow cytometry analysis (BD Fortessa). Using FlowJo software, cells were categorized into Annexin-V-PI-, Annexin-V+PI-, Annexin-V-PI+, and Annexin-V+PI+ cells, with the percentages of the four parts compared across groups. Experiments were conducted in triplicate.
For the tumor immune microenvironment analysis, the mice were sacrificed, and the tumors were collected for flow cytometry analysis after one-time arachidonic acid intratumoural injection. Tumor tissues were finely chopped and digested in HBSS (Gibco) with DNAase I, C9407, Collagenase Type II, and Collagenase Type IV at 37 °C until most of the tissue is dissolved. The suspension was then filtered through a 70 μm cell strainer and centrifuged. Approximately (1-2) × 106 cells were suspended in 100 μl of PBS, incubated with antibodies at 4 °C for 30 min, and washed twice with PBS. 7-AAD was added to the samples 10 minutes before measurement to exclude dead cells. Surface staining of cells was performed using fluorescence-conjugated antibody labels (see Supplementary Table 2 for a complete list of antibodies). Stained samples were detected by an LSR Fortessa flow cytometer (BD Biosciences, San Jose, California) or Cytek® Aurora (Cytek® Biosciences, Fremont, California), and data were analyzed with Flowjo 10.6.2 (TreeStar, Inc., Ashland, OR). Gating strategy is shown in Supplementary Fig. 11g.
Mitochondrial membrane potential assay
The change of mitochondrial membrane potential was assayed by using the JC-1 mitochondrial membrane potential assay kit (C2006, Beyotime Biotechnology). In brief, 1 × 105/well BxPC3 were seeded into a 96-well plate and treated as indicated in the figure legends. Culture medium was replaced with 100 μL JC-1 probe solution and incubation/wash according to instructions. JC-1 staining was detected by a fluorescent reader (Molecular Devices, Sunnyvale, CA) under Ex490/Em530 nm for monomer and Ex525/Em590 nm for polymer (aggregate). The ratio of aggregates/monomers suggested the dissipation of mitochondrial transmembrane potential, and data were relative to vehicle/control. CCCP was applied as a positive control, inducing JC-1 depolymerization.
Mitochondrial permeability transition pore (MPTP) detection
Cells were seeded into a 96-well plate at a density of 1 × 105/well and treated as indicated in the figure legends. MPTP opening was determined following the instructions of the Mitochondrial Permeability Transition Pore Assay Kit (K239-100, Biovision) with minor changes. Briefly, calcein-AM solution was diluted to 500 nM with solution buffer, and the cells were washed once with wash buffer and incubated with 50 μL calcein-AM solution for 20 min at 37 °C, then added with 50 μL CoCl2 solution and incubated for another 10 min to rule out excessive calcein-AM. The plate was read under Ex490/Em515 nm with a fluorescent reader (Molecular Devices, Sunnyvale, CA). The decline rate of fluorescent value indicated the degree of MPTP opening, and the percent of each group was relative to the vehicle/control. 1 μM Ionomycin was used as a positive control.
Tetramethylrhodamine methyl ester (TMRM) staining analysis
Cells were seeded into a 96-well plate at a density of 1 × 105/well and treated as indicated in the figure legends. TMRM solution was diluted to 20 μM with solution buffer, and the cells were washed once with wash buffer and incubated with 50 μL TMRM solution for 20 min at 37 °C. The plate was read under Ex525/Em590 nm with a fluorescent reader (Molecular Devices, Sunnyvale, CA). The decline rate of fluorescent value indicated the degree of mitochondrial transmembrane potential dissipation, and the percent of each group was relative to vehicle/control. 1 μM CCCP was used as a positive control.
Microscopy imaging and immunofluorescence
To observe cell morphology changes, (2-4) × 104 cells were plated in cover-glass bottom dishes and treated as indicated in the figure legends. For Calcein-AM staining, cells were stained with 10 μM Calcein-AM at 37 °C for 30 min before indicated treatment. For propidium iodide (PI) staining, cells were washed twice with PBS and incubated with 200 μL 1 × binding buffer containing 10 μM propidium iodide (HY-D0815, MCE) for 30 min at room temperature. For TMRM staining, cells were stained with 50 μM TMRM at 37 °C for 30 min after the indicated treatment. Cells were finally washed thrice with PBS. Cell images were acquired on fluorescence microscope.
For Immunofluorescence analysis, cells were cultured to 50% density in 24-well polystyrene plates, then fixed with 4% formaldehyde and permeabilized with 0.2% Triton X-100 for 15 minutes following three PBS washes. Following blocking with 1% BSA, cells were incubated overnight with the primary antibody at 4 °C. After being washed three times with PBS, cells were incubated with secondary antibodies for 1 hour at room temperature. Subsequently, the cells were washed three times with PBS and stained with DAPI before being analyzed using a fluorescence microscope or confocal laser scanning microscopy. The catalog number of the phalloidin is provided in the Key Resources Table.
For paraffin-embedded tissues staining, immunofluorescence staining was performed as described above. After antigen retrieval and antigen blocking, the sections were incubated with diluted primary antibodies at 4 °C overnight. Then the sections were incubated with secondary antibodies at room temperature for 2 h. Finally, the sections were mounted in an anti-fade mounting medium with DAPI.
For organoid staining, organoids were cultured in 24-well polystyrene plates and stained with 10 μM Calcein-AM at 37 °C for 30 min before arachidonic acid treatment. At the end time point of arachidonic acid treatment, 10 μM PI staining was performed for 30 min. The organoid was observed under a fluorescence microscope at the indicated time points. Immunofluorescence colocalization analysis was performed using the Plot Profile function in ImageJ.
Spatiotemporal organellar calcium assay
For organelle-specific Calcium imaging with genetically encoded indicators, PANC1 cells were transfected with CMV-Mito4x-GCaMP6f (mitochondrial Ca2+; Addgene #127870) or pCAG G-CEPIA1er (ER Ca2+; Addgene #105012) using Lipofectamine 3000 (L3000015, Thermo Fisher) (2 μg each plasmid/well) separately for 72 h. Transfected cells were treated with 40 or 120 μM arachidonic acid for 60 min. Subsequently, the cells were washed three times with PBS and stained with DAPI before being analyzed using the confocal laser scanning microscopy with excitation at 488 nm wavelength. Mean fluorescence intensity was applied for quantitative analysis. Bar graphs expressed as mean ± SEM (n = 3).
Transmission electron microscopy (TEM)
TEM analysis was conducted as previously described68. First, cells were loaded on parafilm, and then a 300 mesh copper grid (Agar Scientific Ltd., Stansted, UK) was placed over the drop for 2 min. After the excess liquid was removed by blotting with filter paper, the grid was negatively stained with 2% phosphotungstic acid (PTA) for 2 min and examined at 80 kV with a JEM-1200 EXII TEM (JEOL, Ltd., Tokyo, Japan).
Quantitative Real-Time PCR (qPCR)
Total DNA was isolated from cells by column purification with PureLink™ Genomic DNA Mini Kit (K182000, Invitrogen). Quantitative PCR (qPCR) was performed on DNA using Fast SYBR Green Master Mix (4385618, Thermo Fisher Scientific) on a StepOnePlus Real-Time PCR System (Thermo Fisher Scientific). Gene expression was quantified using the following primers:
Human mtDNA-Fw: 5’-AGGACAAGAGAAATAAGGCC-3’
Human mtDNA-Rev: 5’-TAAGAAGAGGAATTGAACCTCTGACTGTAA-3’
Human β-Actin-Fw: 5’- ACCAACTGGGACGACATGGA -3’
Human β-Actin-Rev: 5’- GGTCTCAAACATGATCTGGGTCAT -3’
Fold changes in mtDNA expression were calculated by the ΔΔCt method using β-Actin as an endogenous control. All fold changes are expressed normalized to the untreated control.
Immunoblotting
Cells or tissue were washed in cold PBS and lysed in 1× RIPA lysis buffer (Beyotime, P0013B) with 1% PMSF and 1% protein phosphatase inhibitor cocktail. Lysates were incubated on ice for 30 min and cleared by centrifugation at 15,000 × g for 15 minutes. Protein concentration was quantified using a BCA protein assay kit (Thermo Fisher, 23225). 30 μg total protein was mixed with sample buffer (NCM, WB2001) and denatured at 95 °C for 10 min. The sample was separated by 10% SDS-PAGE and transferred to a PVDF membrane (Millipore, IPVH00010). Membranes were blocked with 5% w/v nonfat dry milk and incubated with primary antibodies overnight at 4 °C and HRP-conjugated secondary antibodies (Thermo Fisher Scientific) for 2 h at room temperature. All antibodies used for incubation were diluted with a universal antibody diluent (New Cell & Molecular Biotech). Signal was detected using Clarity and Clarity Max Western ECL Blotting Substrates (Bio-Rad) and captured using ChemiDoc Imaging System (Bio-Rad). Protein band intensities were quantified using densitometric analysis. Band densities were measured with ImageJ. Local background subtraction was applied to all regions of interest (ROIs). Target protein signals were normalized to the corresponding loading control for each sample. Data represent the mean ± SD from 3 independent biological replicates. Detailed antibody sources are listed in the Supplementary Table 2.
Patients-derived PDAC organoids generation
PDAC tissues were finely chopped and digested in HBSS (Gibco) with DNAase I, C9407, Collagenase Type II, and Collagenase Type IV at 37 °C until most of the tissue is dissolved. The suspension was then filtered through a 100 μm cell strainer and centrifuged. The pellet was resuspended in cold organoid culture medium and mixed at a 1:2 ratio with Matrigel (Corning) to achieve a density of 2000 cells per 40 μL, prior to seeding in a 24-well culture plate. After solidification, the organoid culture medium was added to each well, and the organoids were incubated at 37 °C with 5% CO2 in a humidified environment.
RNAseq analysis
BxPC3 cells were stimulated with 40 or 120 μM arachidonic acid for 1 h, and subsequently lysed with TRIzol™ reagent (Invitrogen) for RNA extraction. mRNA expression levels underwent hierarchical clustering. DEGs between control, low-dose and high-dose arachidonic acid-treatment groups were identified using R package limma69, with |log2FC | >2 and adjusted P value < 0.05. GO and KEGG enrichment analyzes of DEGs, and GSEA of arachidonic acid metabolism related genes, were performed with R package clusterProfiler70. The R package simplifyEnrichment was used to streamline and visualize GO function enrichment results. Visualization was primarily done with the R package ggplot2, unless specifically indicated otherwise.
Statistics and reproducibility
No statistical methods were used to predetermine sample size. For cell-based experiments, biological triplicates were performed in each single experiment in general, unless otherwise stated. Animal experiments were performed in BALB/c nude and C57BL/6 mice. Animals were randomized into different groups after tumor cell inoculation; 4−6 mice were used for each group, unless otherwise indicated. Animals that failed to develop tumors were excluded from the analysis. Statistical analysis was performed using GraphPad Prism8 software (GraphPad Software, Inc.). Statistical significance between two groups was determined by a Two-tailed Student’s t test. For three or more groups, statistical significance was determined by One-way or Two-way ANOVA analysis, unless otherwise indicated in each figure legend. p-values of < 0.05 were considered significant, otherwise indicated as nonsignificant (ns). Graphs and all statistical analysis were performed in GraphPad Prism 8.0 software. Flow cytometry analysis were carried out using Flowjo 10.6.2 (TreeStar, Inc., Ashland, OR). Image-pro was used to calculate the area void of cristae (%); the area of mitochondria and cristae were traced respectively; area void of cristae (%) = (area of mitochondria - area of cristae) / area of mitochondria; a minimum of 15 mitochondria was analyzed per group (n = 3).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The disease and chemical data are available at the Comparative Toxicogenomics Database (https://ctdbase.org/). The raw sequence data of arachidonic acid-treated cells have been deposited in the GEO database with accession number GSE283340. The remaining data supporting the findings of this study are available within the Source Data file. Source data are provided in this paper.
Code availability
The CLGNN code is available at (https://github.com/chutongjia/CLGNN)71. The code supporting the current study is available from the lead contact upon request.
References
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
Bear, A. S., Vonderheide, R. H. & O’Hara, M. H. Challenges and opportunities for pancreatic cancer immunotherapy. Cancer Cell 38, 788–802 (2020).
Shi, J., Gao, W. & Shao, F. Pyroptosis: gasdermin-mediated programmed necrotic cell death. Trends Biochem. Sci. 42, 245–254 (2017).
Zhang, Z. et al. Gasdermin E suppresses tumour growth by activating anti-tumour immunity. Nature 579, 415–420 (2020).
Zhou, Z. et al. Granzyme A from cytotoxic lymphocytes cleaves GSDMB to trigger pyroptosis in target cells. Science 368, https://doi.org/10.1126/science.aaz7548 (2020).
Hou, J., Hsu, J. M. & Hung, M. C. Molecular mechanisms and functions of pyroptosis in inflammation and antitumor immunity. Mol. Cell 81, 4579–4590 (2021).
Wang, Q. et al. A bioorthogonal system reveals antitumour immune function of pyroptosis. Nature 579, 421–426 (2020).
Gao, W., Wang, X., Zhou, Y., Wang, X. & Yu, Y. Autophagy, ferroptosis, pyroptosis, and necroptosis in tumor immunotherapy. Signal Transduct. Target. Ther. 7, 196 (2022).
Wang, Y. et al. Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature 547, 99–103 (2017).
Fritsche, K. L. The science of fatty acids and inflammation. Adv. Nutr. 6, 293s–301s (2015).
Davis, A. P. et al. Comparative toxicogenomics database (CTD): update 2023. Nucleic Acids Res. 51, D1257–d1262 (2023).
Sheng, N. et al. 2023 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). (2023).
Wang, D. & Dubois, R. N. Eicosanoids and cancer. Nat. Rev. Cancer 10, 181–193 (2010).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linkingmetabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Yu, P. et al. Pyroptosis: mechanisms and diseases. Signal Transduct. Target. Ther. 6, 128 (2021).
Hsu, S. K. et al. Inflammation-related pyroptosis, a novel programmed cell death pathway, and its crosstalk with immune therapy in cancer treatment. Theranostics 11, 8813–8835 (2021).
Xia, X. et al. The role of pyroptosis in cancer: pro-cancer or pro-“host”? Cell Death Dis. 10, 650 (2019).
Liao, P. et al. CD8(+) T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 40, 365–378 (2022).
Broz, P., Pelegrín, P. & Shao, F. The gasdermins, a protein family executing cell death and inflammation. Nat. Rev. Immunol. 20, 143–157 (2020).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Rogers, C. et al. Cleavage of DFNA5 by caspase-3 during apoptosis mediates progression to secondary necrotic/pyroptotic cell death. Nat. Commun. 8, 14128 (2017).
Flores-Romero, H., Dadsena, S. & García-Sáez, A. J. Mitochondrial pores at the crossroad between cell death and inflammatory signaling. Mol. Cell 83, 843–856 (2023).
Glover, H. L., Schreiner, A., Dewson, G. & Tait, S. W. G. Mitochondria and cell death. Nat. Cell Biol. 26, 1434–1446 (2024).
Baines, C. P. et al. Loss of cyclophilin D reveals a critical role for mitochondrial permeability transition in cell death. Nature 434, 658–662 (2005).
Bonora, M., Giorgi, C. & Pinton, P. Molecular mechanisms and consequences of mitochondrial permeability transition. Nat. Rev. Mol. Cell Biol. 23, 266–285 (2022).
Waldmeier, P. C., Feldtrauer, J. J., Qian, T. & Lemasters, J. J. Inhibition of the mitochondrial permeability transition by the nonimmunosuppressive cyclosporin derivative NIM811. Mol. Pharmacol. 62, 22–29 (2002).
Vanderhaeghe, S. et al. A pathogenic mutation in the ALS/FTD gene VCP induces mitochondrial hypermetabolism by modulating the permeability transition pore. Acta Neuropathol. Commun. 12, 161 (2024).
Sheng, X. et al. Host cyclophilin A facilitates SARS-CoV-2 infection by binding and stabilizing spike on virions. Signal Transduct. Target. Ther. 8, 459 (2023).
Briston, T. et al. Identification of ER-000444793, a Cyclophilin D-independent inhibitor of mitochondrial permeability transition, using a high-throughput screen in cryopreserved mitochondria. Sci. Rep. 6, 37798 (2016).
Huang, J. C. et al. HB-EGF induces mitochondrial dysfunction via estrogen hypersecretion in granulosa cells dependent on cAMP-PKA-JNK/ERK-Ca(2+)-FOXO1 pathway. Int. J. Biol. Sci. 18, 2047–2059 (2022).
Pinke, G., Zhou, L. & Sazanov, L. A. Cryo-EM structure of the entire mammalian F-type ATP synthase. Nat. Struct. Mol. Biol. 27, 1077–1085 (2020).
Bauer, T. M. & Murphy, E. Role of mitochondrial calcium and the permeability transition pore in regulating cell Death. Circ. Res. 126, 280–293 (2020).
Algieri, C. et al. Melatonin rescues cell respiration impaired by hypoxia/reoxygenation in aortic endothelial cells and affects the mitochondrial bioenergetics targeting the F(1)F(O)-ATPase. Redox Biol. 82, 103605 (2025).
Eggermann, E., Bucurenciu, I., Goswami, S. P. & Jonas, P. Nanodomain coupling between Ca²⁺ channels and sensors of exocytosis at fast mammalian synapses. Nat. Rev. Neurosci. 13, 7–21 (2011).
Giorgi, C., Marchi, S. & Pinton, P. The machineries, regulation and cellular functions of mitochondrial calcium. Nat. Rev. Mol. Cell Biol. 19, 713–730 (2018).
Hirabayashi, Y. et al. ER-mitochondria tethering by PDZD8 regulates Ca(2+) dynamics in mammalian neurons. Science 358, 623–630 (2017).
Ashrafi, G., de Juan-Sanz, J., Farrell, R. J. & Ryan, T. A. Molecular tuning of the axonal mitochondrial Ca(2+) uniporter ensures metabolic flexibility of neurotransmission. Neuron 105, 678–687 (2020).
Rimessi, A. et al. Interorganellar calcium signaling in the regulation of cell metabolism: A cancer perspective. Semin. Cell Dev. Biol. 98, 167–180 (2020).
Dia, M. et al. Reduced reticulum-mitochondria Ca(2+) transfer is an early and reversible trigger of mitochondrial dysfunctions in diabetic cardiomyopathy. Basic Res. Cardiol. 115, 74 (2020).
Liu, C. et al. MCU Upregulation overactivates mitophagy by promoting VDAC1 dimerization and ubiquitination in the hepatotoxicity of cadmium. Adv. Sci. 10, e2203869 (2023).
Hinz, B., Cheremina, O. & Brune, K. Acetaminophen (paracetamol) is a selective cyclooxygenase-2 inhibitor in man. Faseb J. 22, 383–390 (2008).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Rimessi, A. et al. Perturbed mitochondrial Ca2+ signals as causes or consequences of mitophagy induction. Autophagy 9, 1677–1686 (2013).
Baechler, B. L., Bloemberg, D. & Quadrilatero, J. Mitophagy regulates mitochondrial network signaling, oxidative stress, and apoptosis during myoblast differentiation. Autophagy 15, 1606–1619 (2019).
Kang, R., Zeh, H. J., Lotze, M. T. & Tang, D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 18, 571–580 (2011).
Liu, S., Yao, S., Yang, H., Liu, S. & Wang, Y. Autophagy: Regulator of cell death. Cell Death Dis. 14, 648 (2023).
Rodriguez-Enriquez, S., Kim, I., Currin, R. T. & Lemasters, J. J. Tracker dyes to probe mitochondrial autophagy (mitophagy) in rat hepatocytes. Autophagy 2, 39–46 (2006).
Narendra, D., Tanaka, A., Suen, D. F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Galluzzi, L. & Green, D. R. Autophagy-independent functions of the autophagy machinery. Cell 177, 1682–1699 (2019).
Basit, F., Cristofanon, S. & Fulda, S. Obatoclax (GX15-070) triggers necroptosis by promoting the assembly of the necrosome on autophagosomal membranes. Cell Death Differ. 20, 1161–1173 (2013).
Goodall, M. L. et al. The autophagy machinery controls cell death switching between apoptosis and necroptosis. Dev. Cell 37, 337–349 (2016).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008).
O’Reilly, E. M. et al. Durvalumab with or without tremelimumab for patients with metastatic pancreatic ductal adenocarcinoma: a phase 2 randomized clinical trial. JAMA Oncol. 5, 1431–1438 (2019).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv324 (2016).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Hou, J. et al. PD-L1-mediated gasdermin C expression switches apoptosis to pyroptosis in cancer cells and facilitates tumour necrosis. Nat. Cell Biol. 22, 1264–1275 (2020).
Jahreiss, L., Renna, M., Bittman, R., Arthur, G. & Rubinsztein, D. C. 1-O-Hexadecyl-2-O-methyl-3-O-(2’-acetamido-2’-deoxy-beta-D-glucopyranosyl)-sn-glycerol (Gln) induces cell death with more autophagosomes which is autophagy-independent. Autophagy 5, 835–846 (2009).
Zhu, Y. et al. Loss of WIPI4 in neurodegeneration causes autophagy-independent ferroptosis. Nat. Cell Biol. 26, 542–551 (2024).
Hofius, D. et al. Autophagic components contribute to hypersensitive cell death in Arabidopsis. Cell 137, 773–783 (2009).
Imagawa, Y., Saitoh, T. & Tsujimoto, Y. Vital staining for cell death identifies Atg9a-dependent necrosis in developmental bone formation in mouse. Nat. Commun. 7, 13391 (2016).
Wu, J., Wang, L. & Xu, J. The role of pyroptosis in modulating the tumor immune microenvironment. Biomark. Res. 10, 45 (2022).
Du, T. et al. Pyroptosis, metabolism, and tumor immune microenvironment. Clin. Transl. Med. 11, e492 (2021).
Yamamoto, K. et al. Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I. Nature 581, 100–105 (2020).
Velickovic, P. W. F., Hamilton, W. L., Lio, P., Bengio, Y. & Hjelm, R. D. in International Conference on Learning Representations. (2019).
Joung, J. et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Wei, F. et al. Exosomes derived from gemcitabine-resistant cells transfer malignant phenotypic traits via delivery of miRNA-222-3p. Mol. Cancer 16, 132 (2017).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Chu, T., Sheng, N. & Wei, F. Code for Arachidonic acid induces pyroptosis via mitophagy’s nonautophagic function and acts as an PDAC immunotherapy activator. Zenodo https://doi.org/10.5281/zenodo.17935355(2025).
Acknowledgements
We are grateful to Prof. Quan Wang and Dr. Heshi Liu from the First Hospital of Jilin University for providing the human CRC cell line SW480. F.W. received support from the National Natural Science Foundation of China (No. 82473314, 82273276). Z.L. received support from the National Natural Science Foundation of China (No. 82403380)
Author information
Authors and Affiliations
Contributions
T.C. and F.W. conceived the idea and designed the experiments. T.C., Z.L., H.L. (Huan Liu), and W.M. conducted most experiments and analyzed data; N.S. and Y.L. assisted in CLGNN model training and data analysis; T.C., H.L. (Huan Liu), and Y.Z. assisted in FACS analysis; T.C., C.W., and Y.Z. assisted in bioinformatics analysis; T.C., Z.L., J.Z. (Jian Zhang), H.Y., K.D., F.L., and J.Z. (Jianpeng Zhou) assisted in animal experiments; T.C., X.L., and J.X. assisted in qPCR; T.C., J.Z. (Jian Zhang), S.L., and X.P. assisted in TEM analysis. T.C. and H.L. (Han Liu) assisted in pathological analysis; T.C., Z.L., W.M., Z.F., and F.W. contributed to the interpretation of the results. T.C. and F.W. edited the manuscript. F.W. supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Olivier Peulen and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chu, T., Liu, Z., Liu, H. et al. Arachidonic acid induces pyroptosis via a non-autophagic function of mitophagy and enhances immunotherapy in a PDAC model. Nat Commun 17, 1545 (2026). https://doi.org/10.1038/s41467-025-68267-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68267-2