Abstract
Discovering more targets is of great importance for developing alternative interventions for tumor therapy. The roles of transmembrane protein 175 (TMEM175) in neurodegeneration diseases have been reported, however its functions in tumor immune surveillance are not known. We show that TMEM175 conditional knockout in macrophages inhibits the tumor growth and metastasis through promoting the anti-tumor immunity in the tumor microenvironment (TME), including elevated M1-like polarization, reduced M2-like polarization, and facilitated recruitment and activation of T cells and nature killer cells (NKs). The anti-tumor immunity is abrogated by caspase-1 inhibitor VX-765, anti-IL-1β, and anti-IL-18. Tmem175−/− bone marrow-derived macrophages (BMDMs) show enhanced tumor antigen cross-presentation that is further strengthened by IL-1β and IL-18. NLRP3 is robustly elicited in Tmem175−/− BMDMs by the tumor cell debris through lysosomal permeabilization and cathepsin B leakage. Finally, Tmem175−/− mice are more responsive to anti-PD-1. Our works implies TMEM175 to be a potential target for immunotherapy.
Similar content being viewed by others
Introduction
Immune surveillance in TME is crucial for the clearance of tumor cells. Diving into the mechanisms of immune surveillance could help us find more targets to propagate the anti-tumor immunity. Immunotherapy of malignant tumors have developed a constellation of therapeutic methods providing diverse choice for clinicians1. For example, anti-PD-1 antibody is commonly applied in melanoma therapy, however, drug resistance has been reported repeatedly2. Developing novel immunotherapy strategies may help improving anti-tumor immunity through alternative mechanisms.
Macrophages are a subset of crucial immune cells in TME, whose M2-like polarization and other behaviors have been shown to promote the tumor progression3,4. M2-like tumor associated macrophages (TAMs) attenuate the anti-tumor immunity through diverse mechanisms, including secretion of anti-inflammatory cytokines5, up-regulation of immune-suppressive PD-L16, and down-regulation of immune-stimulatory major histocompatibility complex class II (MHC-II)7, resulting in the immune evasion. On the contrary, reprogramming M2-like TAMs into M1-like TAMs through some interventions, such as TLR ligands8, chloroquine9, and hydroxychloroquine10, restored the recruitment and activation of T cells. Notably, the behaviors of TAMs are much more complicated beyond polarization. For instance, TAMs have been demonstrated to degrade extracellular matrix and promote the tumor invasion via secreted cathepsins and cathepsins in lysosomes4,11,12.
Lysosomes are the organelles that profoundly mediate the behaviors of macrophages in many diseases13. Macrophages intake extracellular substances through phagocytosis and transport them into lysosomes, where the low pH provides an optimal condition for nearly 60 kinds of hydrolytic enzymes13,14. The acidic luminal pH in lysosomes is maintained by the vacuolar H+-ATPase (v-ATPase) and other ion channels on lysosomal membrane. The v-ATPase hydrolyzes ATP to pump protons into lysosomes, and at the same time, cation efflux or anion influx through other ion channels is needed to balance the transmembrane potential15,16. TMEM175 is a potassium channel on lysosomal membrane, whose knockout causes incapacity to maintain acidic luminal pH during starvation17,18. Additionally, later research also illustrates the H+ permeability of TMEM17519,20. Present results have revealed that the dysfunction of TMEM175 causes Parkinson disease18,21. However researches focusing on the potential role of TMEM175 in the tumor immunity is limited. The crosstalk between lysosomal proton and calcium mediates the polarization of macrophages and anti-tumor immunity9. We thus hypothesize that TMEM175 might influence the homeostasis of lysosomal ions and regulate the behaviors of TAMs.
Dysfunction of lysosomes can activate inflammasome signal and alter the anti-tumor immunity in TME22. Inflammasome signals are composed of several kinds of intracellular sensors including, NLR family pyrin domain containing 3 (NLRP3), NLR family pyrin domain containing 1 (NLRP1), NLR family apoptosis inhibitory protein (NAIP)/NLR family CARD domain containing 4 (NLRC4), absent in melanoma 2 (AIM2), and Pyrin that recognize various danger-associated molecular patterns (DAMPs) and pathogen-associated molecular patterns (PAMPs)23. NLRP3 is the most extensively studied inflammasome sensor, partially because it can be activated by diverse stimuli, while the others can only be provoked by specific kinds of substances. Activation of NLRP3 requires signal 1 that initiates the expression of proteins in NLRP3 pathway by ligands of Toll-like receptors (TLRs) and signal 2 that engages assembly of NLRP3 inflammasomes by DAMPs and PAMPs resulting in the secretion of interleukin-1β (IL-1β) and interleukin-18 (IL-18)24. Rupture of lysosomes and leakage of lysosomal content have been shown to elicit NLRP3 signal during the pathology of atherosclerosis25. In TME, NLRP3 activation has been observed in not only tumor cells but also stromal cells26,27,28,29. Both pro- and anti-tumor effects of NLRP3 activation have been reported in massive researches26,27,30,31,32. Although many details about the relationship between NLRP3 inflammasomes and tumor progression remain fuzzy, some inhibitors of NLRP3 pathways have been developed and tested for their anti-tumor potential33,34,35. Interestingly, an NLRP3 activator, BMS-986299, was also tested in a clinical research for solid cancers (NCT03444753). More investigations about the role of NLRP3 in TME are needed for a better understanding of inflammasomes in tumors.
Here, we found that conditional knockout of TMEM175 in macrophages significantly attenuated the tumor growth of melanoma B16-F10 and lung cancer LL-2 subcutaneous tumors, as well as the lung metastasis of B16-F10. TMEM175 knockout increased the MHC-II+ M1-like macrophages and decreased the CD206+ M2-like macrophages, resulting in the fostered recruitment and activation of CD4+ T cells, CD8+ T cells, and NKs in TME. The effects of TMEM175 knockout were underpinned by the inflammasome activation, since VX-765, anti-IL-1β, anti-IL-18, and Nlrp3 silencing abrogated the anti-tumor effects. Due to the tempered lysosomal degradation, Tmem175−/− BMDMs demonstrated accentuated tumor antigen cross-presentation that was further strengthened by IL-1β and IL-18. We also found how Tmem175−/− BMDMs possessed robust inflammasome activation. The cell debris engulfed by BMDMs caused the lysosomal membrane permeabilization, resulting in the NLRP3 activation by the released cathepsin B (CTSB). During this process, the potentiated cytosolic calcium ions that might be owed to the attenuated lysosomal acidification could be the key factor facilitating the NLRP3 activation in Tmem175−/− BMDMs. Finally, the potential of clinical application of TMEM175 knockout was further verified by the result that Tmem175−/− mice showed better response to anti-PD-1 therapy.
In summary, conditional knockout of Tmem175 in macrophages demonstrates obvious anti-tumor immunity in mice. The underlying mechanisms are compromised by the cross-presentation and NLRP3 inflammasome, enhanced by the lysosomal dysfunction in Tmem175−/− BMDMs. Our findings reveal the role of TMEM175 in tumor immunity, highlighting its potential in immunotherapy. Development of interventions targeting TMEM175 is needed for alternative immunotherapy strategies in the future.
Results
TMEM175 is up-regulated in melanoma and related with M2-like TAMs
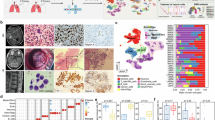
In order to investigate the role of TMEM175 in tumors, we compared the expression of TMEM175 in melanoma and paracancerous tissues. TMEM175 expression was higher in melanoma tissues than that in paracancerous tissues (Fig. 1a). In order to validate the relationship between immune-suppressive CD206+ TAMs and TMEM175 expression in different cell subsets, B16-F10 subcutaneous tumors from wild type C57BL/6J mice were analyzed by FCM. Percentages of CD206+ TAMs were positively correlated with the expression of TMEM175 in TAMs, instead of other CD45- cells (Fig. 1b, c). Thus, we decided to focus on the TMEM175 in macrophages.
a Tissue chip of paired melanoma and paracancerous tissues was stained by anti-TMEM175 to detect the expression of TMEM175 through IHC (n = 6 pairs). TMEM175 was expressed more robustly in melanoma tissues than in paracancerous tissues. Scale bars indicate 500 µm for the left panel and 50 µm for the right panel. TMEM175 positive area was calculated. b, c Wild type mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10 tumor cells (n = 18 mice). Mice were euthanized at day 18 to analyze the TME through FCM. (b) M2-like TAMs were positively correlated with the TMEM175 expression in TAMs. c M2-like TAMs were not correlated with TMEM175 expression in other CD45- cells. d, e TMEM175 expression was obviously decreased in Tmem175−/− BMDMs. Lysates of Tmem175−/− and WT BMDMs were analyzed by western blot using anti-TMEM175 antibody (n = 3 biologically independent samples). Ratios of the grayscale values of TMEM175 and β-actin were calculated. (f) Lysosomal potassium conductance in Tmem175−/− BMDMs was significantly decreased. Lysosomes from Tmem175−/− and WT BMDMs were analyzed by lysosomal voltage clamp. The current was recorded when the bath solution contained 150 mM K+ or 150 mM NMDG. Representative result of voltage clamp was displayed, and the average current intensities in +100 mV with 150 mM K+ or 150 mM NMDG were compared (n = 5 biologically independent samples). Representative results from at least two independent experiments are presented as mean ± SEM, ns denotes not significant. Linear regression was applied to calculate the relationship between the percentages of M2-like TAMs and TMEM175 expression in different cell subsets in (b, c). Statistical significances in (a, e) were determined by two-sided unpaired t-test. Statistical significances in (f) were determined by two-way ANOVA followed by Holm-Sidak’s multiple comparisons test. Source data are provided as a Source data file.
Validation of TMEM175 conditional knockout mice
Conditional knockout of TMEM175 in macrophages was achieved through cre/loxp system in C57BL/6 J mice (Supplementary Fig. 1a). Briefly, exon 5 and exon 6 were floxed (Supplementary Fig. 1a, b), then the Tmem175flox/flox mice were crossbred with Lyz2-Cre+/+ mice to generate Tmem175flox/flox Lyz2-Cre+/+ and Tmem175flox/flox Lyz2-Cre+/− mice (referred as Tmem175−/− mice for brief hereafter) (Supplementary Fig. 1c). The age-matched Tmem175flox/flox Lyz2-Cre−/− littermates were used as control, which are referred as WT mice for brief hereafter.
The abrogated TMEM175 expression in BMDMs from Tmem175−/− mice was confirmed by western blot (Fig. 1d, e). The down-regulated TMEM175 expression was also confirmed by the RNA-seq (Supplementary Data 1). Since TMEM175 is the potassium channel on the lysosomal membranes, whole-lysosomal voltage clamp was applied to confirm the attenuated potassium conductance in lysosomal membranes from Tmem175−/− macrophages. The significantly dampened cross-membrane potassium current confirmed that the function of TMEM175 channels was abrogated by the conditional knockout (Fig. 1f).
TMEM175 conditional knockout suppresses tumor growth through accentuating immune-suppressive microenvironment
To investigate the role of TMEM175 in tumor progression, we used melanoma cell line, B16-F10, to establish subcutaneous tumor models in WT and Tmem175−/− mice. We observed elevated tumor cell death indicated by the cleaved caspase-3 (Fig. 2a and Supplementary Fig. 2a), and significantly decreased tumor weights (Fig. 2b) in Tmem175−/− mice. The tumors were analyzed via flow cytometry to evaluate the recruitment and activation of diverse cell subsets in TME. M1-like MHC-II+ macrophages were augmented and M2-like CD206+ macrophages were mitigated in Tmem175−/− mice (Fig. 2c, d). Other myeloid cells, including granulocytic myeloid derived suppressor cells (G-MDSCs), monocytic myeloid derived suppressor cells (M-MDSCs), and dendritic cells (DCs), showed no significant difference between WT and Tmem175−/− mice (Supplementary Fig. 2b–d), indicating that macrophages were the mainly affected myeloid cell subsets here. We additionally evaluated the recruitment and activation of T cells and NKs in TME. Recruitment and activation of CD4+, CD8+ T cells, and NKs were significantly augmented in Tmem175−/− mice (Fig. 2e–h, Supplementary Fig. 2e, f). Expression of interferon-γ (IFN-γ) and granzyme B (GZMB) in CD4+ T cells, CD8+ T cells, and NKs was enhanced (Supplementary Fig. 2g–l).
Tmem175−/− and WT mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10 (n = 5 mice for a-h) or \(1\times {10}^{6}\) LL-2 (n = 6 mice for i–p) tumor cells. Mice were euthanized at day 18 (for B16-F10) or day 24 (for LL-2) to weigh the tumors and analyze the TME through IHC, IF, and FCM. a B16-F10 tumors were fixed, sliced, and stained by anti-cleaved caspase-3 to detect the tumor cell death through IHC. Enhanced tumor cell death was observed in tissues from Tmem175−/− mice. Scale bars indicate 50 µm. b B16-F10 tumor weights in Tmem175−/− mice were significantly decreased. c Expression of M1-like marker, MHC-II, in TAMs (gated from CD45+ CD11b+ F4/80+ cells) from B16-F10 tumors was significantly up-regulated in Tmem175−/− mice. d Expression of M2-like marker, CD206, in TAMs (gated from CD45+ CD11b+ F4/80+ cells) from B16-F10 tumors was significantly down-regulated in Tmem175−/− mice. e B16-F10 tumors were fixed, sliced, and stained by anti-CD8 and anti-NK1.1 to detect CD8+ T cells and NKs in tissues. Enhanced recruitment of CD8+ T cells and NKs was observed in tissues from Tmem175−/− mice. Scale bars indicate 50 µm. f CD4+ T cells and CD4+ CD69+ activated T cells (gated from CD3+ cells) from B16-F10 tumors were significantly augmented in Tmem175−/− mice. g CD8+ T cells and CD8+ CD69+ activated T cells (gated from CD3+ cells) from B16-F10 tumors were significantly augmented in Tmem175−/− mice. h NKs and NK1.1+ CD69+ activated NKs (gated from CD3- cells) from B16-F10 tumors were significantly augmented in Tmem175−/− mice. i LL-2 tumors were fixed, sliced, and stained by anti-cleaved caspase-3 to detect the tumor cell death through IHC. Enhanced tumor cell death was observed in tissues from Tmem175−/− mice. Scale bars indicate 50 µm. j LL-2 tumor weights in Tmem175−/− mice were significantly decreased. k Expression of MHC-II in TAMs (gated from CD45+ CD11b+ F4/80+ cells) from LL-2 tumors was significantly up-regulated in Tmem175−/− mice. l Expression of CD206 in TAMs (gated from CD45+ CD11b+ F4/80+ cells) from LL-2 tumors was not significantly changed in Tmem175−/− mice. m LL-2 tumors were fixed, sliced, and stained by anti-CD8 and anti-NK1.1 to detect CD8+ T cells and NKs in tissues. Enhanced recruitment of CD8+ T cells was observed in tissues from Tmem175−/− mice. Scale bars indicate 50 µm. n CD4+ T cells, instead of CD4+ CD69+ activated T cells (gated from CD3+ cells) from LL-2 tumors, were significantly augmented in Tmem175−/− mice. o CD8+ T cells and CD8+ CD69+ activated T cells (gated from CD3+ cells) from LL-2 tumors were significantly augmented in Tmem175−/− mice. p NKs and NK1.1+ CD69+ activated NKs (gated from CD3- cells) from LL-2 tumors were not obviously augmented in Tmem175−/− mice. Representative results from two independent experiments are presented as mean ± SEM; ns denotes not significant. Statistical significances in (b–d, f–h, j–l, and n–p) were determined by two-sided unpaired t-test. Source data are provided as a Source data file.
We also explored whether the anti-tumor effect of TMEM175 conditional knockout was reproducible in different tumors. Tumor cell death of LL-2 was promoted (Fig. 2i and Supplementary Fig. 2m), and the tumor weights were decreased in Tmem175−/− mice (Fig. 2j). MHC-II+ M1-like macrophages were heightened, while CD206+ M2-like macrophages were not obviously changed (Fig. 2k, l). Similarly, the granulocytic myeloid derived suppressor cells, monocytic myeloid derived suppressor cells, and DCs were not the influenced myeloid cell subsets (Supplementary Fig. 2n–p). Recruitment of CD4+/CD8+ T cells in Tmem175−/− mice was potentiated; however, the percentages of NKs were not changed (Fig. 2m–p, Supplementary Fig. 2q, r). The activation of CD8+ T cells was potentiated in Tmem175−/− mice (Fig. 2o); however, the activation of CD4+ T cells and NKs was not significantly changed between the two genotypes (Fig. 2n, p). Expression of IFN-γ was significantly promoted in CD4+/CD8+ T cells and NKs from Tmem175−/− mice (Supplementary Fig. 2s-u). Additionally, the granzyme B expression in CD8+ T cells was accentuated in Tmem175−/− mice, but the granzyme B expression in CD4+ T cells and NKs was not obviously elevated (Supplementary Fig. 2v–x).
In order to further validate the importance of macrophages and CD8+ T cells in the anti-tumor immunity in Tmem175−/− mice, clodronate liposomes (Clo lipo) and anti-CD8a antibody were applied to eliminate the macrophages and CD8+ T cells, respectively. Clo lipo treatment diminished the TAMs and abrogated the inhibitory effects of TMEM175 in B16-F10 tumors (Supplementary Fig. 3a–c). The recruitment and activation of CD4+ T cells in Tmem175−/− mice were reduced by Clo lipo (Supplementary Fig. 3d, e). IFN-γ and granzyme B expression in CD4+ T cells in Tmem175−/− mice were also suppressed by Clo lipo (Supplementary Fig. 3f, g). As expected, the recruitment and activation of CD4+ T cells in Clo lipo treated WT mice were elevated to a similar level in Clo lipo treated Tmem175−/− mice (Supplementary Fig. 3d, e). More M2-like TAMs in WT mice and M1-like TAMs in Tmem175−/− mice were observed (Fig. 2c, d), thus Clo lipo tempered the anti-inflammatory effect of TAMs in WT mice and pro-inflammatory effect of TAMs in Tmem175−/− mice. Similar trends were also observed in CD8+ T cells, whose recruitment and activation were modulated to the equal levels in WT and Tmem175−/− mice by Clo lipo (Supplementary Fig. 3h, i). IFN-γ expression and granzyme B expression in CD8+ T cells were inhibited in Tmem175−/− mice instead of WT mice (Supplementary Fig. 3j, k). Anti-CD8a treatment also eradicated the differences in tumor weights between WT and Tmem175−/− mice (Supplementary Fig. 3l, m). The clearance of CD8+ T cells was validated (Supplementary Fig. 3n).
Together, these results illustrated that TMEM175 conditional knockout hampered the tumor growth through increasing M1-like macrophages, decreasing M2-like macrophages, and facilitating T cell and NK-mediated anti-tumor immunity. This anti-tumor immunity was accordant in different tumors, although some differences existed.
TMEM175 conditional knockout suppresses B16-F10 lung metastasis through activating the anti-tumor immunity
Besides subcutaneous tumor growth, we were also curious about the influence of TMEM175 conditional knockout on lung metastasis. WT and Tmem175−/− mice were intravenously injected with B16-F10 cells to establish lung metastasis models. Tmem175−/− mice demonstrated obviously fewer metastatic nodules and lower lung weights (Fig. 3a–c). Then the subsets of lung macrophages, CD11c+ CD11b- CD64+ alveolar macrophages (AMs) and CD11b+ CD24- CD64+ interstitial macrophages (IMs), were analyzed by FCM. Previous works have shown that MHC-II was down-regulated in AMs in tumor-bearing mice36 and that CD206+ AMs were increased in bronchoalveolar lavage fluid from lung cancer patients37. We observed the elevated MHC-II and mitigated CD206 expression in AMs (Fig. 3d, e). Similarly, we found the augmented MHC-IIhigh IMs (Fig. 3f), which might be superior in antigen-presentation38, as well as the reduced CD206high IMs (Fig. 3g), which show immunosuppressive phenotypes39. Another classification of IM subsets based on CD11c and MHC-II was also performed. Tmem175−/− mice possessed significantly attenuated CD11clow MHC-IIlow IM1, CD11clow MHC-IIhigh IM2, and expanded CD11c+ MHC-IIhigh IM3 (Fig. 3h–k). IM1 and IM2 are relatively similar and show immune regulatory roles, while IM3 resembles typically activated monocytes40. As expected, the recruitment, activation, and IFN-γ expression of CD4+ T and CD8+ T cells were promoted in Tmem175−/− mice (Fig. 3l–q). Notably, CD8+ T cells recognizing p15E, a tumor antigen in B16-F10, were also expanded (Fig. 3r). However, neither cDC1 nor cDC2 recruitment in lungs was obviously changed (Supplementary Fig. 4a, b), which implied the pivotal roles of macrophages in eliciting the anti-tumor immunity.
B16-F10 cells (2 × 105) were intravenously injected into WT and Tmem175−/− sex-matched mice at 6–8 weeks old (n = 6 mice). Mice were euthanized on day 21. The lungs were weighed, and the metastatic nodules were counted. Then the lungs were minced and digested to prepare the single-cell suspension for FCM analysis. a TMEM175 conditional knockout inhibited B16-F10 lung metastasis. Scale bars indicate 1 cm. b, c Tmem175−/− mice showed fewer lung metastatic nodules (b). Lung weights in Tmem175−/− mice were decreased (c). d MHC-IIhigh AMs (gated from CD45+ CD11c+ CD11b− CD64+ cells) were increased in Tmem175−/− mice. e CD206high AMs (gated from CD45+ CD11c+ CD11b− CD64+ cells) were decreased in Tmem175−/− mice. f MHC-IIhigh IMs (gated from CD45+ CD11b+ CD24− CD64+ cells) were increased in Tmem175−/− mice. g CD206high IMs (gated from CD45+ CD11b+ CD24− CD64+ cells) were decreased in Tmem175−/− mice. h Subsets of IMs (IM1, IM2, and IM3) were altered by TMEM175 conditional knockout. i CD11clow MHC-IIlow IM1 (gated from CD45+ CD11b+ CD24− CD64+ cells) were decreased in Tmem175−/− mice. j CD11clow MHC-IIhigh IM2 (gated from CD45+ CD11b+ CD24− CD64+ cells) were decreased in Tmem175−/− mice. k CD11c+ MHC-IIhigh IM3 (gated from CD45+ CD11b+ CD24− CD64+ cells) were increased in Tmem175−/− mice. CD4+ T cells (l), CD4+ CD69+ T cells (m), and CD4+ IFN-γ+ T cells (n) (gated from CD3+ cells) were increased in Tmem175−/− mice. CD8+ T cells (o), CD8+ CD69+ T cells (p), CD8+ IFN-γ+ T cells (q), and CD8+ p15E-tetramer+ T cells (r) (gated from CD3+ cells) were increased in Tmem175−/− mice. s Single cell sequencing was applied to evaluate the changes in the cell subsets and the transcriptome (n = 3 mice). Representative results from two independent experiments are presented as mean ± SEM, ns denotes not significant. Statistical significances in (b–g and i–r) were determined by two-sided unpaired t-test. Source data are provided as a Source data file.
Then we used single cell transcriptome to analyze the underlying mechanisms of the anti-tumor immunity (Fig. 3s). In accordance with the FCM results, T cells and NKs were expanded in Tmem175−/− mice (Fig. 3s, Supplementary Fig. 4c, and Supplementary Data 2). We focused on the transcriptome changes in AMs and IMs. Both AMs and IMs showed up-regulated antigen presentation pathway, which included several genes coding histocompatibility molecules such as H2-D1, H2-M3, and H2-K1 (Supplementary Fig. 5a, b and Supplementary Data 3 and 4). And this result was in accordance with our FCM results. The differently expressed genes were then analyzed by GO and KEGG enrichment (Supplementary Fig. 5c–f). In AMs, we found some up-regulated genes that have been reported to promote M1-like polarization, such as Tlr2, Irf7, Nlrp3, Malt1, Map3k1, and Nod2 (Supplementary Data 3). These genes were enriched in pro-inflammatory pathways such as NOD-like receptor signaling pathway (mmu04621), NF-kappa B signaling pathway (mmu04064), and MAPK signaling pathway (mmu04010) (Supplementary Fig. 5e and Supplementary Data 3). These genes were also enriched in GO terms that facilitate immunity and inflammation, such as immune response (GO:0006955), antigen processing and presentation of peptide antigen via MHC class I (GO:0002474), and inflammatory response (GO:0006954) (Supplementary Fig. 5c and Supplementary Data 3). In IMs, we also found some up-regulated genes that have been reported to promote M1-like polarization, such as Gbp2b, Irf7, Irf9, Gbp4, and Tlr9 (Supplementary Data 4). These genes were enriched in pro-inflammatory pathways such as NOD-like receptor signaling pathway (mmu04621), Toll-like receptor signaling pathway (mmu04620), and NF-kappa B signaling pathway (mmu04064) (Supplementary Fig. 5f and Supplementary Data 4). These genes were also enriched in GO terms that facilitate immunity and inflammation, such as immune system process (GO:0002376), antigen processing and presentation (GO:0002474), positive regulation of interleukin-1 beta production (GO:0032731), and positive regulation of NF-kappaB transcription factor activity (GO:0051092) (Supplementary Fig. 5d and Supplementary Data 4). Thus, TMEM175 knockout promoted the antigen presentation and inflammatory process, resulting in the M1-like polarization of macrophages, which could involve several different pathways, including inflammasome, TLRs, and NF-kappa B.
In summary, Tmem175−/− mice developed fewer lung metastatic nodules owing to the activated tumor immune microenvironment. Our data implied that the promoted inflammasome and cross-presentation could be the underlying mechanisms of the anti-tumor effect.
Inflammasome signal is required for the anti-tumor effects of tmem175 conditional knockout
We attempted to dissect the mechanisms of the anti-tumor immunity in Tmem175−/− mice. The single cell transcriptome analysis indicated that TLRs, NF-kappa B, and inflammasome signal were up-regulated in macrophages (Supplementary Fig. 5e, f). Indeed, TLRs and NF-kappa B are the signal 1 of NLRP3 inflammasome22. TMEM175 knockout has been reported to cause lysosomal dysfunction17,20,21. Dysfunction of lysosomes activated inflammasomes in macrophages25,41, and the activated inflammasome signal in macrophages showed anti-tumor potential31,42, which inspired us to test the role of inflammasome signal in the anti-tumor effect of TMEM175 conditional knockout mice. We validated the high expression level of caspase-1 p10 and IL-1β in B16-F10 from Tmem175−/− mice (Fig. 4a–c). VX-765 is an inhibitor of caspase-143 that is the common downstream protease of several inflammasome signals. Tumor-bearing Tmem175−/− mice and WT mice were treated with VX-765 or vehicle every day. The anti-tumor effects of TMEM175 conditional knockout were abrogated by the VX-765 treatment, which heightened the tumor weights and suppressed the tumor cell death in Tmem175−/− mice (Fig. 4d–f and Supplementary Fig. 6a). These data indicated that inflammasome signal was likely to be the key pillar of the anti-tumor effects in Tmem175−/− mice. We then used flow cytometry to explore whether the VX-765-mediated promotion of tumor growth was achieved through the deprivation of anti-tumor immunity. VX-765 reduced the high percentages of MHC-II+ M1-like macrophages in Tmem175−/− mice to the same level as WT mice, leaving CD206+ M2-like macrophages scarcely changed (Fig. 4g, h). Consistent with the curtailed M1-like polarization, recruitment and activation of CD4+ T cells and CD8+ T cells in Tmem175−/− mice were diminished to the comparable level as WT mice (Fig. 4i–l). The blunted CD8+ T cells and NKs were also illustrated by immunofluorescence (Fig. 4m and Supplementary Fig. 6b–d). We also tested the influence of inflammasome stimulator, chitosan, on B16-F10 tumor-bearing mice. Chitosan is able to activate inflammasomes in macrophages, depending on phagocytosis44. Peritoneal administration of chitosan significantly curbed the tumor weights and induced the tumor cell death in both WT and Tmem175−/− mice (Fig. 4n–p and Supplementary Fig. 6e). This effect was more remarkable in Tmem175−/− mice than in WT mice (Fig. 4n–p and Supplementary Fig. 6e). In vitro cultured Tmem175−/− macrophages were more susceptible to phagocytosis-dependent inflammasome activator, because chitosan induced more IL-1β secretion in peritoneal macrophages (PMs) and BMDMs (Fig. 4q, r). The IL-1β treatment promoted the MHC-II expression in both WT and Tmem175−/− BMDMs (Supplementary Fig. 6f), indicating the role of inflammasome signal in the M1-like polarization of macrophages. Furthermore, more caspase-1 p10 was generated in Tmem175−/− BMDMs treated by aluminium hydroxide, another phagocytosis-depended inflammasome activator45 (Fig. 4s and Supplementary Fig. 6g).
B16-F10 tumors from Fig. 2 were fixed, sliced, and stained by anti-caspase-1 p10 or anti-IL-1β (a) to detect the inflammasome activation through IHC (n = 5 biologically independent samples). Enhanced inflammasome activation was observed in tissues from Tmem175−/− mice (b, c). Scale bars indicate 50 µm. d–m Tmem175−/− and WT mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10 tumor cells (n = 5 mice). Vehicle or VX-765 solution (0.4 mg dissolved in 200 µL) was administrated every day through intraperitoneal injection since day 8. Mice were euthanized at day 18 to weigh the tumors and analyze the TME through IHC, FCM, and IF. e VX-765 elevated the tumor weights in Tmem175−/− mice, thus VX-765 abrogated the anti-tumor effect in Tmem175−/− mice. f VX-765 decreased the cleaved caspase-3 expression in Tmem175−/− mice, indicating reduced tumor cell death. Scale bars indicate 50 µm. g Expression of MHC-II in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was significantly attenuated by VX-765 in Tmem175−/− mice. h Expression of CD206 in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was not obviously altered by VX-765 in Tmem175−/− mice. i, j CD4+ T cells and CD4+ CD69+ activated T cells (gated from CD3+ cells) were significantly diminished by VX-765 in Tmem175−/− mice. k, l CD8+ T cells and CD8+ CD69+ activated T cells (gated from CD3+ cells) were significantly diminished by VX-765 in Tmem175−/− mice. m IF staining showed that VX-765 decreased the recruitment of CD8+ T cells in tumors from Tmem175−/− mice. Scale bars indicate 50 µm. n, o Tmem175−/− and WT mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10 tumor cells (n = 5 mice). Vehicle or chitosan solution (2 mg dissolved in 200 µL) was administrated every 3 days through intraperitoneal injection since day 3. Mice were euthanized at day 18 to weigh the tumors and analyze the cell death by IHC. o Tmem175−/− mice showed better response to chitosan treatment. p Chitosan promoted the expression of cleaved caspase-3 in both Tmem175−/− and WT mice. Scale bars indicate 50 µm. q, r Primary Tmem175−/− and WT BMDMs or PMs were stimulated with 500 ng/mL LPS for 4 h to activate signal 1 of NLRP3 inflammasomes, and then treated with 40 µg/mL chitosan for 5 h. Tmem175−/− BMDMs and PMs secreted more IL-1β in response to chitosan (n = 5 biologically independent samples). s Primary Tmem175−/− and WT BMDMs were stimulated with 500 ng/mL LPS for 4 h to activate signal 1 of NLRP3 inflammasomes, and then treated with 300 µg/mL aluminium hydroxide for desired period of time. Lysates of Tmem175−/− and WT BMDMs were analyzed by western blot using anti-caspase-1 antibody. Tmem175−/− BMDMs showed more enhanced cleavage of caspase-1 p10 in response to aluminium hydroxide. Representative results from two independent experiments are presented as mean ± SEM, ns denotes not significant. Statistical significances in (b, c) were determined by two-sided unpaired t-test, in (e, g–l, o, q, r) were determined by two-way ANOVA followed by Holm-Sidak’s multiple comparisons test. In (h), the CD206+ cells between Tmem175−/− and WT in VX-765 treated group were compared by two-sided unpaired t-test. Source data are provided as a Source data file. d, n Created in BioRender. Wang, Y. (2026) https://BioRender.com/yaf354h.
Present works have reported the conflict roles of inflammasome in TME. The downstream cytokines IL-1β and IL-18 could promote or suppress the anti-tumor immunity in different works28,31,32,46,47,48,49. To further validate the influences of IL-1β and IL-18 on our tumor-bearing models, we injected IL-1β and IL-18 into wild type B16-F10 subcutaneous tumors. Intratumoral administration of IL-1β and IL-18 significantly decreased tumor weights (Supplementary Fig. 7a, b). Both IL-1β and IL-18 intensified the MHC-II and IFN-γ expression in TAMs, although they did not obviously alter CD206 expression (Supplementary Fig. 7c–e). IL-18 did not significantly promote the IFN-γ expression in CD4+ T cells and the granzyme B expression in CD8+ T cells, nevertheless IL-1β and IL-18 promoted the recruitment and activation of CD4+ T and CD8+ T cells in general (Supplementary Fig. 7f–m). IL-1β also increased the recruitment, activation, IFN-γ expression, and granzyme B expression in NKs (Supplementary Fig. 7n–q). Thus the tumor immune microenvironment in wild type mice receiving IL-1β and IL-18 intratumoral administration at least partially resembled that in Tmem175−/− mice.
To ascertain the influences of IL-1β and IL-18 on anti-tumor effects by TMEM175 knockout, antibodies targeting IL-1β and IL-18 were applied. Intraperitoneal anti-IL-1β administration increased the tumor weights in Tmem175−/− mice (Fig. 5a, b). MHC-II expression in Tmem175−/− TAMs was decreased, while CD206 expression was not altered (Fig. 5c, d). Recruitment, activation, IFN-γ expression and granzyme B expression in CD4+ T cells, CD8+ T cells, and NKs in Tmem175−/− mice were mitigated by anti-IL-1β (Fig. 5e–l and Supplementary Fig. 8a–d). Similarly, anti-IL-18 abrogated the anti-tumor effects in Tmem175−/− mice (Fig. 5m, n). Percentages of M1-like TAMs in Tmem175−/− mice declined to the similar level to WT mice, while M2-like TAMs were not obviously changed (Fig. 5o, p). Recruitment, activation, IFN-γ expression and granzyme B expression in CD4+ T cells, CD8+ T cells, and NKs were suppressed by anti-IL-18 in Tmem175−/− mice (Fig. 5q–x and Supplementary Fig. 8e–h). CD8+ T/NK cell recruitment, CD4+ T/CD8+ T/NK cell activation, and granzyme B expression in CD8+ T cells were inhibited by anti-IL-18 in WT mice (Fig. 5r, u, v, x, Supplementary Fig. 8e, f), and NK cell recruitment was inhibited by anti-IL-1β in WT mice (Supplementary Fig. 8a) indicating less importance of inflammasome in the anti-tumor immunity in WT mice than that in Tmem175−/− mice. Thus increased IL-1β and IL-18 production is the key step of the anti-tumor immunity in Tmem175−/− mice.
a–l Tmem175−/− and WT mice (n = 5 mice) were treated with 100 µg/100 µL anti-IL-1β or Armenian hamster IgG isotype intraperitoneally 1 day before the subcutaneous implantation with \(1\times {10}^{6}\) B16-F10. Then same doses of IgG were injected intraperitoneally on day 6 and 13. Mice were euthanized on day 14 to weigh the tumors and analyze the TME through FCM. b The anti-tumor effects of TMEM175 knockout were suppressed by anti-IL-1β. c Expression of MHC-II in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was significantly attenuated by anti-IL-1β in Tmem175−/− mice. d Expression of CD206 in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was not obviously altered by anti-IL-1β in Tmem175−/− mice. e–h Recruitment, activation, IFN-γ expression, and granzyme B expression in CD4+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-1β in Tmem175−/− mice. i–l Recruitment, activation, IFN-γ expression, and granzyme B expression in CD8+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-1β in Tmem175−/− mice. m–x Tmem175−/− and WT mice (n = 5 mice) were treated with 100 µg/100 µL anti-IL-18 or rat IgG2a isotype intraperitoneally 1 day before the subcutaneous implantation with \(1\times {10}^{6}\) B16-F10. Then same doses of IgG were injected intraperitoneally on day 6 and 13. Mice were euthanized on day 14 to weigh the tumors and analyze the TME through FCM. n The anti-tumor effects of TMEM175 knockout were suppressed by anti-IL-18. o Expression of MHC-II in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was significantly attenuated by anti-IL-18 in Tmem175−/− mice. p Expression of CD206 in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was not obviously altered by anti-IL-18 in Tmem175−/− mice. q–t Recruitment, activation, IFN-γ expression, and granzyme B expression in CD4+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-18 in Tmem175−/− mice. Activated CD4+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-18 in WT mice. u–x Recruitment, activation, IFN-γ expression, and granzyme B expression in CD8+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-18 in Tmem175−/− mice. Recruitment, activation, and granzyme B expression in CD8+ T cells (gated from CD3+ cells) were significantly diminished by anti-IL-18 in WT mice. Representative results from two independent experiments are presented as mean ± SEM, ns denotes not significant. Statistical significances in (b–l and n–x) were determined by two-way ANOVA followed by Holm-Sidak’s multiple comparisons test. Source data are provided as a Source data file. a, m Created in BioRender. Wang, Y. (2026) https://BioRender.com/yaf354h.
Collectively, the anti-tumor effects of TMEM175 knockout mice were abrogated by VX-765, anti-IL-1β, and anti-IL-18 through restraining anti-tumor immunity, highlighting the crucial role of inflammasome signal in this process.
TMEM175 knockout augments the cross-presentation of tumor antigens
Since the tumor antigen-specific CD8+ T cells were increased in Tmem175−/− mice (Fig. 3r), we sought to compare the different cross-presentation capacities between WT and Tmem175−/− BMDMs that activate CD8+ T cells. Referring to MacNabb et al. 7 days were sufficient for priming tumor antigen specific CD8+ T cells50. B16-F10-OVA cells were implanted in wild type C57BL/6J mice and grew for 10 days to prime CD8+ T cells (Fig. 6a). Then the splenic lymphocytes from tumor bearing mice were harvested, CFSE-labeled, and cultured with WT or Tmem175−/− BMDMs with/without B16-F10-OVA cell debris for 48 h before FCM and ELISA analysis (Fig. 6a). Tmem175−/− BMDMs promoted the expansion of CD8+ T cells indicated by CFSElow CD8+ T cells (Fig. 6b, c) and increased the percentages of OVA specific CD8+ T cells (Fig. 6d). Additionally, activated CD8+ CD69+ T cells, IFN-γ secretion, and TNF secretion were all elevated in Tmem175−/− group (Fig. 6e–g). IL-1β and IL-18 supplements facilitated the proliferation or activation of CD8+ T cells in different aspects (Fig. 6b–g). Thus, knockout of TMEM175 promoted the cross-presentation signal, which could be enhanced by IL-1β and IL-18.
a–g Wild type C57BL/6 J mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10-OVA cells. On day 10, the splenic lymphocytes from tumor bearing mice were harvested, CFSE-labeled, and cultured with WT or Tmem175−/− BMDMs in the presence of IL-2 (20 U/mL) for 48 h before FCM and ELISA analysis (n = 6 biologically independent samples). During the co-culture, B16-F10-OVA cell debris (2.5 × 105/mL), IL-1β (500 pg/mL), and IL-18 (500 pg/mL) were added in the desired group. b, c Splenic lymphocytes were labeled by CFSE, thus the CFSElow cells represented the successive generations of cells. Tmem175−/− BMDMs induced more robust proliferation of CD8+ T cells (CFSElow cells gated from CD3+ CD8+ cells). IL-1β and IL-18 supplementation further promoted the CD8+ T cell proliferation in Tmem175−/− group. d Tmem175−/− BMDMs induced more CD8+ OVA-tetramer+ T cells (gated from CD3+ cells). IL-18 significantly promoted the CD8+ OVA-tetramer+ T cell expansion. e Tmem175−/− BMDMs induced more robust CD8+ CD69+ activated T cells (gated from CD3+ cells). IL-1β and IL-18 supplementation further promoted the CD8+ T cell activation in Tmem175−/− group. f Tmem175−/− BMDMs induced more IFN-γ secretion. IL-1β supplementation promoted IFN-γ secretion in WT and Tmem175−/− group. IL-18 supplementation promoted the IFN-γ secretion in WT group. g Tmem175−/− BMDMs induced more TNF secretion. IL-18 supplementation promoted the TNF secretion in WT group. h Brief illustration of the engulfment and digestion capacity examination. i Fluorescence intensities of intracellular p15E in Tmem175−/− BMDMs (gated from CD45+ CD11b+ F4/80+ cells) were significantly higher than in WT BMDMs (n = 3 biologically independent samples). j Fluorescence intensities of FITC-dextran in Tmem175−/− BMDMs (gated from CD45+ CD11b+ F4/80+ cells) were comparable with those in WT BMDMs (n = 3 biologically independent samples). k, l Cells in the co-culture system were harvested for RNA-seq to investigate the alterations in the gene expression, GO terms, and KEGG pathways (n = 3 biologically independent samples). Up-regulated GO terms by TMEM175 knockout without (k) or with (l) B16-F10-OVA cell debris treatment. m, n CD8+ T cells in the co-culture system were harvested and adopted to Cd8a−/− mice 1 day before and on day 7th after B16-F10-OVA implantation (n = 5 mice). Tumor bearing mice were euthanized on day 14th after B16-F10-OVA implantation to weigh the tumors. Representative results from two independent experiments are presented as mean ± SEM, ns denotes not significant. Statistical significances in (c–g, i, j) were determined by two-way ANOVA followed by Holm-Sidak’s multiple comparisons test. Statistical significances in (n) were determined by one-way ANOVA followed by Holm-Sidak’s multiple comparisons test. Source data are provided as a Source data file. a–m Created in BioRender. Wang, Y. (2026) https://BioRender.com/yaf354h. h Created in BioRender. Wang, Y. (2026) https://BioRender.com/69fpsu9.
There have been many researches revealing that limited lysosomal degradation of antigens favors antigen cross-presentation and presentation of APCs51,52,53. We therefore compared the intracellular tumor antigen levels between WT and Tmem175−/− BMDMs (Fig. 6h). In vitro cultured BMDMs were treated with B16-F10 cell debris or FITC-dextran for desired period of time and then analyzed by FCM. Higher levels of tumor antigen fluorescence were observed in Tmem175−/− BMDMs (Fig. 6i). However, the fluorescent intensities of FITC-dextran were comparable between WT and Tmem175−/− BMDMs (Fig. 6j). Thus, the delayed antigen degradation in Tmem175−/− BMDMs could explain the enhanced cross-presentation.
Then RNA-seq was applied to illustrate the signal changes during the cross-presentation. In the absence of B16-F10-OVA cell debris, Tmem175−/− group showed higher expression of several genes than WT group including Tlr13, Tlr8, Malt1, Irf4, Irf5, Trim29, Pcbp2, Cd9, and Sema4a that have been reported to promote T cell proliferation and activation (Supplementary Data 5). And these genes were enriched in innate immune response (GO:0045087), immune response (GO:0006955), adaptive immune response (GO:0002250), and inflammatory response (GO:0006954) (Fig. 6k and Supplementary Data 5). Although no tumor antigen exists, Tmem175−/− BMDMs showed pro-inflammatory phenotypes, thus promoting non-specific T cell proliferation and activation, which could explain the Tmem175−/− promoted CFSElow CD8+ and CD69+ CD8+ T cells without antigen stimulation in Fig. 6c, e. In B16-F10-OVA debris treated Tmem175−/− group, cell cycle (GO:0007049), cell division (GO:0051301), and mitotic spindle organization (GO:0007052) were up-regulated comparing with WT group treated with tumor antigen (Fig. 6l and Supplementary Data 6), which might be attributed to the more robust antigen specific T cell proliferation induced by Tmem175−/− BMDMs (Fig. 6c, d). We also attempted to investigate the effects of IL-1β and IL-18 on cross-presentation. Totally 7 common up-regulated genes (Cd164, Il1b, Gbp8, Iigp1, Csf2rb, Gbp4, and Gbp5) and 3 common down-regulated genes (Bcl6, Fubp1, and Atf4) between WT and Tmem175−/− group were altered by IL-1β (Supplementary Fig. 9a). However no common gene between WT and Tmem175−/− group was altered by IL-18 (Supplementary Fig. 9a). Despite this, several common altered KEGG pathway and GO terms were found to be altered by IL-1β and IL-18 (Supplementary Figs. 9b–e and 10a–d). The KEGG pathways and GO terms up-regulated by IL-1β in both WT and Tmem175−/− group included IL-17 signaling pathway (Path:04657), NOD-like receptor signaling pathway (Path:04621), cellular response to type II interferon (GO:0071346) and interleukin-1 beta production (GO:0032611) (Supplementary Figs. 9b, c and 10a, b and Supplementary Data 7 and 8). Additionally, positive regulation of canonical NF-kappaB signal transduction (GO:0043123) was up-regulated by IL-1β in WT group and regulation of canonical NF-kappaB signal transduction (GO:0043122) was up-regulated by IL-1β in Tmem175−/− group (Supplementary Data 7 and 8). Thus the common down-stream events of IL-1β in WT and Tmem175−/− group might include NF-kappaB signal, down-stream IFN-γ secretion, and possible positive feedback of IL-1β secretion. The common up-regulated KEGG pathways by IL-18 included IL-17 signaling pathway (Path:04657) and Toll-like receptor signaling pathway (Path:04625) (Supplementary Fig. 9d, e and Supplementary Data 9 and 10), indicating these two pathways to be the common down-stream events of IL-18 in WT and Tmem175−/− groups. In order to validate the down-stream events of IL-1β and IL-18, we used siRNA to silence Myd88, a known down-stream protein of IL-1R, and Csf2rb, a common DEG up-regulated by IL-1β (Supplementary Fig. 9f, g). Silencing Myd88 suppressed the effects of IL-1β and IL-18. For IL-1β, better inhibitory effects were observed in siMyd88 than that in siCsf2rb, indicating that Myd88 is the main down-stream event (Supplementary Fig. 9f–i).
To confirm the in vivo effects of the enhanced cross-presentation in Tmem175−/− BMDMs, two batches of CD8+ spleen T cells from the culture system were adopted to Cd8a−/− B16-F10-OVA tumor bearing mice (Fig. 6m). T cells cultured with Tmem175−/− BMDMs showed more obvious anti-tumor effects (Fig. 6n).
In summary, probably owing to the delayed lysosomal antigen degradation, Tmem175−/− BMDMs possess higher cross-presentation capacity, which was further enhanced by the down-stream cytokines of inflammasomes, IL-1β and IL-18.
B16-F10 cell debris activates NLRP3 inflammasomes in TMEM175 knockout BMDMs more extensively
Our previous experiments have proven that inflammasome signal was involved in the anti-tumor effects of TMEM175 conditional knockout. We then attempted to pursue the inflammasome activator in TME. We suspected that two groups of factors in TME were the potential inflammasome stimulators. One is composed of the soluble cytokines and metabolites derived from tumor cells, and the other is the dead tumor cells. Thus the conditioned medium from in vitro cultured B16-F10 was collected and filtered to remove insoluble materials, and the B16-F10 cells were harvested and repeatedly blown by syringes followed by two freeze-thaw cycles to generate cell debris. In order to test the activation of inflammasome pathway by B16-F10 supernatant and B16-F10 cell debris, LPS was applied to elicit signal 1 of inflammasomes in primary BMDMs, then the B16-F10 supernatant or B16-F10 cell debris was incubated overnight with BMDMs, from which the secreted IL-1β was detected by ELISA. The B16-F10 cell debris, instead of B16-F10 conditioned medium, provoked obvious IL-1β production more extensively in Tmem175−/− BMDMs (Fig. 7a). B16-F10 cell debris induced the secretion of IL-1β in time-dependent manner, during which the differences between WT and Tmem175−/− groups became distinct around 4 h (Fig. 7b). Decreasing FBS in the culture medium could further facilitate the IL-1β production (Supplementary Fig. 11a). We also detected the released IL-18 in cell supernatant of BMDMs after B16-F10 cell debris treatment (Fig. 7c and Supplementary Fig. 11b). Rising extracellular potassium to 40 mM almost completely inhibited the B16-F10 cell debris-mediated IL-1β production, which resembled the feature of NLRP3 inflammasomes54 implying NLRP3 to be the activated inflammasome sensor here (Fig. 7d). The enhanced recruitment and polymerization of ASC by NLRP3 in Tmem175−/− BMDMs was confirmed by co-immunoprecipitation, in which the Nigericin was used as a positive control (Fig. 7e and Supplementary Fig. 11c). The weak ASC signal in blank group could be owed to the engulfment of dead cells inevitable during the primary cell culture (Fig. 7e and Supplementary Fig. 11c). LPS alone could elicit uncanonical inflammasome signal that causes pyroptosis resulting in downstream NLRP3/ASC/caspase-1 activation55 (Fig. 7e and Supplementary Fig. 11c).
Primary BMDMs from Tmem175−/− and WT mice were stimulated with 500 ng/mL LPS for 4 h to activate signal 1 of NLRP3 inflammasomes, and then treated with B16-F10 supernatant (25% volum) or cell debris (2.5 × 105/mL) for desired period of time, during which VX-765, CA-074, or KCl were added, whose final concentrations were 10 µg/mL, 100 µM, and 40 mM respectively. a, b The culture medium was analyzed by ELISA. B16-F10 cell debris, instead of supernatant, induced obviously more potentiated IL-1β secretion from Tmem175−/− than from WT BMDMs (a) (n = 3 biologically independent samples). BMDMs were treated with B16-F10 supernatant or B16-F10 cell debris overnight in (a). The differences between Tmem175−/− and WT BMDMs became distinct after 4 h (b) (n = 6 biologically independent samples). c Total protein in the culture medium was concentrated by trichloroacetic acid for western blot analysis (n = 3 independent experiments). Tmem175−/− BMDMs released more IL-18 than WT BMDMs. d Treatment of KCl, VX-765, or CA-074 significantly inhibited the IL-1β secretion (n = 3 biologically independent samples). Cells were treated by B16-F10 cell debris for 8 h. e Interaction between ASC and NLRP3 was confirmed by CO-IP in the B16-F10 cell debris treated BMDMs. Nigericin (10 µM) was used as positive control (n = 3 independent experiments). Cells were treated by B16-F10 cell debris or Nigericin for 4 h. f Tmem175−/− and WT BMDMs were harvested for electron microscopy observation (n = 3 biologically independent samples). B16-F10 cell debris caused lysosomal permeabilization. Scale bars indicate 500 nm. g Tmem175−/− and WT BMDMs treated with LPS and B16-F10 cell debris were fixed and permeabilized. Then, cells were stained by cathepsin B rabbit mAb, cathepsin D rabbit mAb, or cathepsin L rabbit mAb (n = 3 biologically independent samples). The spots of cathepsin B, cathepsin D, and cathepsin L diffused in the cytoplasm by cell debris treatment. Scale bars indicate 10 µm. h CA-074, an inhibitor of cathepsin B, suppressed the ASC-NLRP3 interaction and the cathepsin B-NLRP3 interaction (n = 3 independent experiments). Cells were treated with B16-F10 cell debris for 4 h. i In vitro cultured WT and Tmem175−/− BMDMs were transfected with siRNA targeting Nlrp3 or control siRNA. Then the BMDMs were adopted to wild type mice 1 day before and on day 7th after B16-F10 implantation (n = 5 mice). Tumor bearing mice were euthanized on day 14th after B16-F10 implantation to weigh the tumors. Representative results from two independent experiments are presented as mean ± SEM; ns denotes not significant. Statistical significances in (a, b, d, i) were determined by two-way ANOVA followed by Holm-Sidak’s multiple comparisons test. Source data are provided as a Source data file. i Created in BioRender. Wang, Y. (2026) https://BioRender.com/yaf354h.
The next question is how the inflammasomes were activated. Untreated WT and Tmem175−/− BMDMs were analyzed by RNA-seq to explore whether any pro-inflammasome genes were altered in resting state. One up-regulated gene and 25 down-regulated genes were found (Supplementary Data 1), none of which was related with inflammasome strongly. And these genes were not significantly enriched in inflammasome related GO term or KEGG pathway (Supplementary Fig. 11d and Supplementary Data 1). These results indicated that the Tmem175−/− BMDMs did not show pro-inflammasome phenotype in resting state and that post transcription mechanisms might exist during the inflammasome activation. Since the chitosan and aluminium hydroxide elicited inflammasome through lysosomal destabilization44,45, we observed the morphological changes of the lysosomes through electron microscope. B16-F10 cell debris was engulfed into lysosomes leading to lysosomal membrane permeabilization featured with the discontinuous high density lysosomal membranes (Fig. 7f). Leakage of lysosomal contents, including cathepsin B, has been proven to stimulate NLRP3 inflammasome during the progression of arteriosclerosis and multiple myeloma25,41. The lysosomal membrane permeabilization was confirmed by the results that round, and light spots of CTSB, CTSD, and CTSL became fuzzy and dim after B16-F10 cell debris treatment (Fig. 7g and Supplementary Fig. 11e–g). The activation of NLRP3 and production of IL-1β were dependent on the interaction between cathepsin B and NLRP3, which was abrogated by CA-074, a specific cathepsin B inhibitor (Fig. 7d, h, Supplementary Fig. 11h, i).
To further validate the role of NLRP3 in the anti-tumor effects of TMEM175 knockout, we silenced NLRP3 in BMDMs through siRNA and adopted these cells into WT B16-F10 tumor bearing mice. As expected, silencing NLRP3 expression in Tmem175−/− BMDMs abrogated the anti-tumor effects (Fig. 7i and Supplementary Fig. 11j, k).
Previous work has demonstrated that lysosomal rupture activates NLRP3 signal through not only cathepsin B but also reactive oxygen species (ROS)41. We tested the level of ROS in B16-F10 cell debris treated BMDMs. B16-F10 cell debris significantly increased intracellular ROS, and the highest ROS level was found in Tmem175−/− BMDMs (Supplementary Fig. 12a). Inhibiting ROS through acetylcysteine (N-Acetyl-L-cysteine, NAC) mitigated the activation of NLRP3 inflammasome and IL-1β secretion (Supplementary Fig. 12b–d).
In summary, B16-F10 cell debris was engulfed by the BMDMs, resulting in the lysosomal membrane permeabilization and NLRP3 activation, and stronger inflammasome activation was observed in Tmem175−/− BMDMs. These results explained the strong inflammasome signal in Tmem175−/− mice.
Calcium ions are involved in the promoted inflammasome activation in TMEM175 knockout BMDMs
We tried to interrogate how TMEM175 knockout promoted the NLRP3 inflammasome activation. Previous research reported that knockout of TMEM175 suppressed the acidification of lysosomes during starvation17. Through pH sensitive lysosomal probe, we found that after B16-F10 cell debris treatment, lysosomal pH in Tmem175−/− BMDMs was higher than that in WT BMDMs (Supplementary Fig. 13a) indicating a hampered lysosomal acidification in Tmem175−/− BMDMs by cell debris treatment. Elevated lysosomal pH has been reported to induce the efflux of lysosomal calcium9,56. Lysosomal calcium concentration determined by rhod-dextran illustrated less lysosomal calcium in Tmem175−/− BMDMs after B16-F10 cell debris treatment (Supplementary Fig. 13b). Additionally, cytosolic calcium determined by Fluo-4-AM indicated higher cytosolic calcium level in Tmem175−/− BMDMs treated by B16-F10 cell debris (Supplementary Fig. 13c). Inhibiting the lysosomal acidification in WT BMDMs by BafA1 promoted the efflux of lysosomal calcium and NLRP3 activation (Supplementary Figs. 12b–d and 13b, c). Thus, we speculated that the hampered lysosomal acidification could explain the enhanced lysosomal calcium efflux and the elevated cytoplasm calcium in Tmem175−/− BMDMs treated with cell debris.
Cytosolic calcium promoted the NLRP3 activation through calcium-sensing receptor or mitochondrial damage57,58, but it was fuzzy whether the direct interaction between calcium and NLRP3 existed. Chelating cytosolic calcium via BAPTA-AM before B16-F10 cell debris treatment significantly impeded the IL-1β production and ASC polymerization (Supplementary Fig. 13d–f), which suggested the higher cytosolic calcium to be the reason of the more intensified inflammasome activation in Tmem175−/− BMDMs. The cytosolic calcium was required for the direct interaction between NLRP3 and ASC, because both BAPTA-AM administration before B16-F10 cell debris treatment and EDTA-Na2 administration in cell lysate during CO-IP hampered the interaction between NLRP3 and ASC (Supplementary Fig. 13e, f). Additionally, calcium ions might also participate in the interaction between NLRP3 and CTSB (Supplementary Fig. 13g, h). On the contrary, inhibiting CTSB did not obviously alter the cytosolic calcium level (Supplementary Fig. 13c), indicating the lysosomal calcium efflux to be the up-stream event of CTSB activation.
We additionally investigated the role of CTSB, cytosolic calcium ions, and ROS in the anti-tumor immunity through inhibitors. The anti-tumor effects of TMEM175 knockout were blunted by the treatment of CA-074 and BAPTA-AM (Supplementary Fig. 14a, b). M1-like polarization of TAMs in Tmem175−/− mice was reversed by CA-074 and BAPTA-AM, and more obvious effects were observed in BAPTA-AM treatment (Supplementary Fig. 14c, d). Similarly, recruitment and activation of CD4+ T cells were curbed by CA-074 and BAPTA-AM in Tmem175−/− mice, while IFN-γ expression and granzyme B expression were only repressed by BAPTA-AM (Supplementary Fig. 14e–h). Recruitment, activation, IFN-γ expression, and granzyme B expression of CD8+ T cells were restrained by CA-074 and BAPTA-AM in Tmem175−/− mice (Supplementary Fig. 14i–l). Thus, cytosolic calcium elevation seemed to be a more crucial factor than CTSB activation for the pro-inflammatory phenotypes in TMEM175 knockout. Comparing with the distinctly inhibitory effects of BAPTA-AM, the ROS scavenger, NAC, showed partial inhibitory effects on the anti-tumor immunity. NAC administration through drinking water59,60 increased the tumor weights, decreased the M1-like TAMs, and elevated the M2-like TAMs in Tmem175−/− mice (Supplementary Fig. 14m–p). These results are in accordance with previous reports60,61. NAC partially inhibited the recruitment and activation of CD4+ T cells, but it did not obviously alter the IFN-γ and granzyme B expression in CD4+ T cells in Tmem175−/− mice (Supplementary Fig. 14q–t). Recruitment, activation, IFN-γ expression, and granzyme B expression of CD8+ T cells were suppressed by NAC in Tmem175−/− mice (Supplementary Fig. 14u–x). These effects were not obvious in NAC treated WT mice, which could be owed to the low level of ROS activation in WT mice. Although NAC inhibited the inflammasome activation in WT BMDMs in vitro, the inhibitory effects of NAC on immune microenvironment in vivo could only be clearly observed in mice with high basal ROS activation level. Our in vivo research demonstrated that the cytosolic calcium seems to be the key factor, although CTSB and ROS are also involved in the NLRP3 activation.
We also tested whether calcium was involved in the PYD-PYD interaction of ASC and NLRP3 through molecular dynamics simulation. The PYD-PYD interaction complex was used for the molecular dynamics simulation in the existence of ten calcium ions for 100 ns. Eight of the ten calcium ions demonstrated obvious interaction with the amino-acid residues through hydrogen bonds (Supplementary Fig. 15a). The PYD-PYD interaction complex reached a stable state after 20 ns (Supplementary Fig. 15b). The radius of gyration (Rg), solvent accessible surface area (SASA), and distance between two PYD domains of the PYD-PYD interaction complex decreased during the 100 ns simulation, indicating that the PYD domains of ASC and NLRP3 get closer to each other with the help of calcium ions (Supplementary Fig. 15c-e). The detailed interactions between each calcium ion and amino-acid residues were shown (Supplementary Fig. 15f). Thus calcium ions facilitated the PYD-PYD interaction of ASC and NLRP3 through direct interactions.
Together, the augmented NLRP3 activation in Tmem175−/− BMDMs could be owed to the elevated cytosolic calcium, which might result from the suppressed lysosomal acidification during cell debris treatment. Although other factors, such as ROS and cathepsins, may also participate in this process, the interaction between lysosomal protons and calcium ions could at least partially explain the elevated NLRP3 activation.
TMEM175 conditional knockout mice show better therapeutic responses to anti-PD-1 therapy
Knowing that TMEM175 knockout ameliorated the immune-suppressive microenvironment, we attempted to interrogate whether it could be combined with other immunotherapy methods. Since T cells were vigorously recruited into TME in Tmem175−/− mice, we chose an anti-PD-1 antibody. Tumor-bearing Tmem175−/− and WT mice were treated with anti-PD-1 or isotype antibodies every 3 days. Tmem175−/− mice showed promoted tumor cell death and decreased tumor weights in response to anti-PD-1 antibody, while WT mice were seldom relieved by the anti-PD-1 therapy (Fig. 8a, b and Supplementary Fig. 16a). This result could be explained by the sparse infiltration of T cells in the tumors of WT mice. MHC-II+ M1-like macrophages were increased by the anti-PD-1 therapy both in Tmem175−/− and WT mice (Fig. 8c), which coincided with previous report62. On the contrary, CD206+ M2-like macrophages were not obviously altered by the anti-PD-1 therapy (Fig. 8c). The recruitment of CD4+ T cells, CD8+ T cells, and NKs was significantly elevated by the anti-PD-1 therapy in Tmem175−/− mice, while NKs in WT mice were scarcely altered by the anti-PD-1 (Fig. 8d-h, Supplementary Fig. 16b, c). Anti-PD-1 antibody promoted the activation of CD4+/CD8+ T cells both in Tmem175−/− mice and WT mice (Fig. 8f, g), however no matter whether anti-PD-1 antibody was administrated, the recruitment and activation of CD4+/CD8+ T cells in Tmem175−/− mice were higher than that in WT mice (Fig. 8f, g). Additionally, recruitment and activation of NKs were selectively expanded in Tmem175−/− mice (Fig. 8h).
Tmem175−/− and WT mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10 tumor cells (n = 5 mice). Isotype or anti-PD-1 antibody (200 µg dissolved in 200 µL saline) was administrated every 3 days through intraperitoneal injection since day 3. Mice were euthanized at day 18 to weigh the tumors and analyze the TME through IHC, FCM, and IF. a Anti-PD-1 therapy promoted the tumor cell death, indicated by the cleaved caspase-3, in Tmem175−/− mice, while the tumor cell death in WT mice was only slightly changed. Scale bars indicate 50 µm. b Tumor weights in Tmem175−/− mice were reduced by anti-PD-1, while WT mice showed a negligible response to anti-PD-1 therapy. c Expression of M1-like marker, MHC-II, in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was significantly elevated by anti-PD-1 therapy in both Tmem175−/− and WT mice. Tmem175−/− mice showed higher percentages of M1-like macrophages, no matter whether anti-PD-1 was administered. Expression of M2-like marker, CD206, in TAMs (gated from CD45+ CD11b+ F4/80+ cells) was not obviously altered by anti-PD-1. d, e IF showed that anti-PD-1 promoted the recruitment of CD8+ T cells and NKs in tumors from Tmem175−/− mice. Scale bars indicate 50 µm. f CD4+ T cells and CD4+ CD69+ activated T cells (gated from CD3+ cells) were significantly elevated by anti-PD-1 in both Tmem175−/− and WT mice. Tmem175−/− mice showed higher percentages of CD4+ T cells, and CD4+ CD69+ activated T cells, no matter whether anti-PD-1 was administered. g CD8+ T cells and CD8+ CD69+ activated T cells (gated from CD3+ cells) were significantly elevated by anti-PD-1 in both Tmem175−/− and WT mice. Tmem175−/− mice showed higher percentages of CD8+ T cells, and CD8+ CD69+ activated T cells, no matter whether anti-PD-1 was administered. h NK1.1+ cells and NK1.1+ CD69+ activated NKs (gated from CD3- cells) were significantly augmented by anti-PD-1 antibody in Tmem175−/− mice, while NKs and activated NKs in WT mice did not show obvious change. No matter whether anti-PD-1 antibody was administered, percentages of NKs and activated NKs in Tmem175−/− mice were higher than those in WT mice. Representative results from two independent experiments are presented as mean ± SEM; ns denotes not significant. Statistical significances in (b, c, and f–h) were determined by two-way ANOVA followed by Tukey’s multiple comparisons test. The tumor weights between isotype-treated and anti-PD-1 antibody-treated Tmem175−/− mice in (b), the percentages of M2-like macrophages and NKs between isotype-treated WT and Tmem175−/− mice in (c, h) were compared by two-sided unpaired t-test. Source data are provided as a Source data file. a Created in BioRender. Wang, Y. (2026) https://BioRender.com/yaf354h.
Overall, conditional knockout of TMEM175 in macrophages improved the effects of anti-PD-1 therapy through promoting the recruitment and activation of T cells and NKs, implying a promising application of the interventions targeting TMEM175 in immunotherapy. The figure summarizing the immune mechanisms involved in the Tmem175−/− BMDMs has been presented (Supplementary Fig. 17a). The FCM gating strategies in our work have been presented (Supplementary Fig. 18a, b and Supplementary Fig. 19a).
Discussion
One of the present strategies for tumor therapy is to stimulate the immune surveillance. Immunotherapy has been successfully applied in cancer therapy, however, resistance to these therapeutic methods has been reported63,64. Possible strategies to overcome the resistance include finding novel markers and combining the novel interventions with the present therapeutic methods. Through in vivo and in vitro experiments, we substantiated that the anti-tumor immunity of TMEM175 knockout was underpinned by the strengthened inflammasome activation and cross-presentation. Delayed antigen degradation in Tmem175−/− BMDMs accentuated the cross-presentation, resulting in the expanded CD8+ T cells. Tumor cell debris was phagocytosed into the lysosomes of macrophages, causing lysosomal membrane permeabilization. Released lysosomal cathepsins, such as cathepsin B, activated NLRP3, resulting in the IL-1β secretion. At the same time, NLRP3 stimulation in Tmem175−/− BMDMs was facilitated by the higher cytosolic calcium that might be attributed to the suppressed lysosomal acidification. Finally, knockout of TMEM175 enhanced the therapeutic effects of anti-PD-1 antibodies through promoting the immune surveillance in TME. Our work implied that TMEM175 in TAMs might be a promising intervention target for future immunotherapy. Moreover, the effects of interventions targeting TMEM175 in TAMs might be further expanded if other therapeutic methods are added.
In our work, knockout of TMEM175 suppressed the acidification of lysosomes during the LPS + B16-F10 cell debris stimulation, which was in accordance with some of the previous reports17,18. However, Hu et al. reported that knockout of TMEM175 decreased the lysosomal pH in resting state20. It might be because that the influence of TMEM175 KO on lysosomal pH is depended on the status of cells. In resting state, TMEM175 knockout yielded more acidic lysosomes17,18,20,21,65, however during challenges such as starvation17,18,21 and cell debris treatment (in our work), TMEM175 knockout inhibited the lysosomal acidification. The inhibitory effects of TMEM175 knockout on the lysosomal acidification become obvious when pressures exist, such as starvation, engulfing dead cells, and other stimulations. It is noteworthy that TMEM175 could interact with different proteins (such as AKT21 and LAMP1/265) causing conformational changes resulting in the altered K+ or H+ conductance21,65. The in vitro lysosomal patch-clamp electrophysiology may not be sufficient to illustrate the physical function of TMEM175 in cells, because it requires some pre-treatment and dissection of lysosomes from cells. Mass spectrum might be helpful to explore the interaction between TMEM175 and other molecules in different cells and different status. Imaging of ion flux across the lysosomal membranes by isotopes is a possible approach to investigate the physical function of TMEM175.
The activation of inflammasomes by components in TME has been reported in many researches. In breast cancer, DAMPs in tumor cell necrotic fluid stimulated NLRP326. The DAMPs and other molecules activating inflammasomes in TME include ROS66, TNF67, Ca2+ influx68, extracellular ATP32, and various chemotherapy agents29,69,70,71. Previous research found that gemcitabine and 5-FU activated NLRP3 via lysosomal permeabilization and release of cathepsin B29. And the cathepsin B has been validated as a NLRP3 activator in other researches68,72,73. In our work, engulf of tumor cell debris caused lysosomal permeabilization and release of cathepsin B into cytoplasm. Additionally, present works demonstrate that cathepsin B activated NLRP3 indirectly through ROS, K+ efflux, and Ca2+ influx68,73 or direct interaction29. We confirmed that the direct interaction between cathepsin B and NLRP3 existed in our in vitro model. The cell debris-mediated lysosomal permeabilization might be a common mechanism of inflammasome activation in different tumors, which could explain the similar anti-tumor effects in our different tumor models. We did not observed significant differences in lysosomal permeabilization between Tmem175−/− and WT BMDMs via immunofluorescence, however this result might need to be further validated.
Extracellular calcium activates NLRP3 through calcium-sensing receptor74,75. However the relationship between intracellular calcium and NLRP3 seems fuzzy. Extracellular ATP promoted the Ca2+ release from endoplasmic reticulum leading to NLRP3 activation through mitochondrial damage-derived ROS and mtDNA58,76. On the contrary, chelating intracellular Ca2+ suppressed NLRP3 instead of AIM2 or NLRC457. However present results did not illustrate whether the direct interaction between cytosolic calcium ions and NLRP3 inflammasome exists. Our data proved that Ca2+ is necessary for the recruitment and polymerization of ASC by NLRP3, and the calcium might be mobilized from the lysosomes more efficiently in Tmem175−/− BMDMs. Our in vivo data further confirmed that Ca2+ might be a more important factor when comparing with CTSB and ROS.
Maturation and secretion of IL-1 family members are the downstream events of inflammasome activation. In some researches the production of IL-1β was recognized as a marker of M1-like polarization of macrophages10,77. There are many delicate elaborations about the anti-tumor effects of IL-1β and IL-18 through stimulation of NK cells and CD8+ T cells31,32,46,47. However, reverse results demonstrated that IL-1β and IL-18 strengthened tumor development through promoting angiogenesis78 and recruitment of immune-suppressive cells, including myeloid-derived suppressor cells (MDSCs), M2-like TAMs, regulatory B cells, and regulatory T cells28,48,49,79. A possible hypothesis is that the effects of IL-1β/IL-18 signals are affected by other signals in different cell subsets. If IL-1β/IL-18 signals encounters IL-6/STAT3 signal, MDSCs could be extensively recruited80. On the contrary, if IL-1β/IL-18 signals are accompanied by antigen presentation in antigen presenting cells, anti-tumor immunity could be augmented32, which is exactly the mechanism shown in our research that TMEM175 knockout promoted the inflammasome activation and antigen cross-presentation at the same time.
Our research work highlighted that inhibiting TMEM175 might be a potential strategy for immunotherapy. Although a broad K+ channel inhibitor, 4-aminopyridine, has been reported to inhibit TMEM17581, selective TMEM175 inhibitor is needed for future research and clinical practices. A recent article has published two TMEM175 inhibitors82. These agents might be applied in tumor immunotherapy; however, two impediments exist in the future research work. First, the inhibitory efficacy of one compound might differ between human and mouse TMEM175 protein. Thus, how to select an optimized model for the drug screening in the next step could be a challenge. Second, it is reasonable that specific delivery of TMEM175 inhibitors to TAMs is necessary for an ideal therapeutic effect. How to selectively deliver the TMEM175 inhibitors to TAMs could be another challenge.
For the limitations of the study, we have outlined the mechanisms of TMEM175 knockout in promoting anti-tumor immunity through lysosomal permeabilization and cathepsin B leakage-induced NLRP3 inflammasomes. We also unraveled that the efflux of lysosomal calcium facilitated ASC-NLRP3 interaction. However, knockout of TMEM175 might influence anti-tumor immunity through autophagy inhibition and ROS generation, whose details need further investigation.
In summary, TMEM175 conditional knockout in macrophages suppresses the tumor growth of B16-F10 and LL-2, as well as the metastasis of B16-F10, through amplifying anti-tumor immunity. These effects rely on the intensified cross-presentation and tumor cell debris-mediated inflammasome activation, which is further enhanced by the increased cytosolic calcium in TMEM175 knockout macrophages. The mechanisms of TMEM175 knockout-mediated anti-tumor immunity have been shown in Supplementary Fig. 17a. Thus, TMEM175 is a potential target for future immunotherapy. Better results could be achieved if TMEM175-targeting therapy is accompanied by other therapy approaches.
Methods
Mouse breeding and generation of TMEM175 conditional knockout mice
All animal studies in this work were approved by the Animal Ethics Committee of Sichuan University (Chengdu, Sichuan, China). Our experiments are in accordance with the NIH Guide for the Care and Use of Laboratory Animals. Mice were housed in specific pathogen-free (SPF) conditions with 22–23 °C, 45–55% relative humidity and a 12–12 h light/dark cycle with free food and water supplies. In this work, experimental and control animals were co-housed. All animals used were euthanized with carbon dioxide in accordance with approved animal protocols.
Tmem175flox/wt mice were created by Nanjing Biomedical Research Institute of Nanjing University. A pair of sgRNAs was designed and validated to target wild type Tmem175. The sequences of sgRNA were 5′-end: GGAGTTTTAACTGTTCAGG and 3′-end: CATTATAGCCCTGCATACTTGG. Referring to the sgRNA, a donor vector carrying exon 5 and exon 6 encircled by two loxP sequences was constructed. The donor vector was injected into mouse oosperm, then the E5-E6 of the wild type Tmem175 gene was replaced by the loxp-E5-E6-loxp through homologous recombination with the help of Cas9 system to generate Tmem175flox/wt mice. The targeting strategy and loxp-E5-E6-loxp sequence have been shown in Supplementary Fig. 1. Tmem175flox/wt mice mated to generate Tmem175flox/flox mice, which mated with Lyz2-Cre+/+ mice (Jackson Lab, stock number: 004781) and then self-crossed to generate Tmem175flox/flox Lyz2-Cre+/+ and Tmem175flox/flox Lyz2-Cre+/- mice. Both Tmem175flox/flox Lyz2-Cre+/+ and Tmem175flox/flox Lyz2-Cre+/- mice were regarded as TMEM175 conditional knockout mice (Tmem175−/− mice). The Tmem175flox/flox Lyz2-Cre−/− littermates were used as control (WT mice). Every WT or Tmem175−/− mouse was identified by PCR before being included in our experiments. To eliminate inevitable mutations during the self-fertilization, the 15th generation of Tmem175flox/flox Lyz2-Cre+/+ mice mated with wild type Tmem175wt/wt Lyz2-Cre−/− C57BL/6 J mice, and the offspring self-crossed to generate a novel line of Tmem175−/− mice. Both previous and novel lines of WT and Tmem175−/− mice were used in our experiments.
The PCR primers (Tsingke biotechnology) used in the identification of our mice include: Tmem175 F: 5′- CCCATAACTCCTATTAGCATATCAAC-3′, Tmem175 R: 5′- GGCTGGGCTAGGTATATTCAATC-3′, (WT = 195 bp, flox = 285 bp); Lyz2-Cre A: 5′-CCCAGAAATGCCAGATTACG-3′, Lyz2-Cre B: 5′-TTACAGTCGGCCAGGCTGAC-3’, Lyz2-Cre AB: 5′- CTTGGGCTGCCAGAATTTCTC-3′, (WT = 350 bp, Mut = ~700 bp).
Cd8a−/− mice were purchased from Jackson Lab (stock number: 002665), and the genotypes were identified according to the protocol provided by Jackson Lab. The PCR primers (Tsingke biotechnology) used in the identification of Cd8a−/− mice include: oIMR1098: 5′-GACCTGGTATGTGAAGTGTTGG-3′; oIMR1099: 5′-ACATCACCGAGTTGCTGATG-3′; oIMR6828: 5′-CATAGCGTTGGCTACCCGTG-3′ (WT = 265 bp, Mut = 343 bp).
Both male and female mice were included in our experiments. Sex-matched WT and Tmem175−/− mice at 6–8 weeks old were included in the in vivo and in vitro experiments. The oligonucleotide sequences have been summarized in Supplementary Data 11.
Cell culture
Mouse melanoma cell line (B16-F10) and Lewis lung cancer cell line (LL-2) were purchased from ATCC. B16-F10, B16-F10-OVA, and LL-2 cells were cultured in DMEM medium (Gibco, C11009955CP) supplied with 10% FBS (Gibco, 10091148) and 100 U/mL penicillin + 100 µg/mL streptomycin (Beyotime, C0222). G418 (0.4 mg/mL) was applied in the B16-F10-OVA culture. Mouse BMDMs were extracted from tibias and femurs. The bone marrow was treated by red blood cell lysis, filtered by a 70 µm cell strainer (Falcon, CLS352350), and cultured in RPMI-1640 medium (Gibco, C11875500BT) supplied with 10% FBS (Gibco, 10091148), 100 U/mL penicillin + 100 µg/mL streptomycin (Beyotime, C0222), and 20 ng/mL M-CSF (Novoprotein, CB34) for 3 days. Then the culture medium was discarded, and the cells were cultured in fresh medium for an additional 2–3 days. Afterward, the suspension cells were removed, and the adherent cells were applied in our experiments. For mouse peritoneal macrophages, humanely euthanized mice were intraperitoneally injected with 5 mL cold saline. The ascites was extracted after gentle massage on abdomen. The peritoneal cells were centrifuged and cultured in RPMI-1640 medium (Gibco, C11875500BT) supplied with 10% FBS (Gibco, 10091148) and 100 U/mL penicillin + 100 µg/mL streptomycin (Beyotime, C0222) for 2 h to allow the adherence of macrophages and remove the suspension cells. All the cells were maintained at 37 °C and 5% CO2.
Electrophysiology
Whole-organelle voltage-clamp recording was applied according to previously reported methods17,83. Macrophages were seeded onto coverslips for 48-well plates. Lysosomes were enlarged by 5 µM vacuolin-1 (Sigma, 673000) treatment for 2–3 h. The enlarged lysosomes were dissected out with a glass pipette. Then another fire-polished glass pipette was applied to form the gigaseal formation with the lysosomes. After the pipette tips touch the lysosomes, slight suction through a plastic pipette connected to the pipette holder might help with the gigaseal formation. When the current decreased to less than 10 pA at 5 mV, or the seal resistance increased to around 8–20 GΩ, a successful gigaseal was formed. Then ZAP pulses (0.1–50 ms, −200 to −1000 mV voltage pulses) were used to rupture the part of the membrane in the glass pipette tips and to establish whole-lysosomal mode. Successful establishment of whole-lysosomal configuration was indicated by the decrease in seal resistance to 20–100 MΩ. Whole-organelle voltage-clamp recording of current from −100 mV to 100 mV was performed with Axon patch-clamp 700B (Molecular Devices). The bath solution contained 145 mM K-methanesulfonate, 5 mM KCl, and 10 mM HEPES (pH 7.2); alternatively, 150 mM NMDG-methanesulfonate was used to replace the potassium. The pipette solution contained 145 mM K-methanesulfonate, 5 mM KCl, and 10 mM MES (pH 5.5).
Animal subcutaneous tumor models
WT and Tmem175−/− sex-matched mice at 6–8 weeks old were used in our experiments. B16-F10 and LL-2 subcutaneous tumor models were used in our research. Mice were subcutaneously injected with \(1\times {10}^{6}\) tumor cells. For VX-765 treatment, VX-765 (Selleck, S2228) was administrated at the 8th day after inoculation. Vehicle (containing 5% DMSO, 40% PEG400, 5% Tween 80, and 50% saline) or VX-765 solution (0.4 mg dissolved in 200 µL) was administrated every day through intraperitoneal injection. For clodronate liposomes treatment, mice were intravenously injected with the clodronate liposomes (YEASEN Biotechnology, 40337ES08) or control liposomes (YEASEN Biotechnology, 40338ES08) at a dose of 1 mg/20 g body weight at day −1, 6, and 13. For anti-CD8a (Bio X Cell, BE0061), anti-IL-1β (Bio X Cell, BE0246), and anti-IL-18 (Bio X Cell, BE0237) treatment, mice were intraperitoneally injected with 100 µg/100 µL antibody or isotype control (rat IgG2b isotype (Bio X Cell, BE0090) for anti-CD8a, Armenian hamster IgG isotype (Bio X Cell, BE0091) for anti-IL-1β, and rat IgG2a isotype (Bio X Cell, BE0089) for anti-IL-18) diluted with saline at day −1, 6, and 13. For CA-074 and BAPTA-AM treatment, mice were intratumorally injected with CA-074 (MCE, HY-103350) or BAPTA-AM (MCE, HY-100545) or vehicle (containing 10% DMSO, 40% PEG400, and 50% saline) at a dose of 200 µg/20 g body weight at days 7, 10, and 13. For anti-PD-1 therapy, 200 µg anti-PD-1 antibody (Bio X Cell, BE0146) or rat IgG2a isotype (Bio X Cell, BE0089) dissolved in 200 µL saline was administered every 3 days through intraperitoneal injection, and the 1st treatment was administrated at the 3rd day after tumor cell inoculation. For chitosan treatment, vehicle (containing 2% acetic acid and 98% saline) or chitosan (Sigma, 448869) solution (2 mg dissolved in 200 µL) was administered every 3 days through intraperitoneal injection, and the 1st treatment was administrated at the 3rd day after tumor cell inoculation. For NAC treatment, 1 g/L NAC (MCE, HY-B0215) was administrated in drinking water since day 759,60.
For the administration of IL-1β and IL-18 in wild type mice, intratumoral injections of 100 ng IL-1β (Novoprotein, C042) or 100 ng IL-18 (Novoprotein, CK06) dissolved in 50 µL saline were implemented every day from day 8 to day 17.
For the adoption of in vitro cultured T cells, spleen lymphocytes from B16-F10-OVA tumor bearing mice were harvested and cultured with WT or Tmem175−/− BMDMs in the existence of B16-F10-OVA for 48 h. Then the CD8+ T cells were isolated by immunomagnetic negative selection (STEMCELL TECHNOLOGIES, 19853) and injected intravenously into Cd8a−/− mice (\(1\times {10}^{6}\) cells per mouse) 1 day before B16-F10-OVA implantation and seven days after B16-F10-OVA implantation. Cd8a−/− mice were subcutaneously injected with \(1\times {10}^{6}\) B16-F10-OVA tumor cells.
For the adoption of BMDMs, in vitro cultured WT or Tmem175−/− BMDMs were transfected with siRNA (GenePharma) targeting Nlrp3 (sense: 5′-CCAGGAGAGAACCUCUUAUTT-3′ anti-sense: 5′-AUAAGAGGUUCUCUCCUGGTT-3′) or control siRNA via Lipofectamine RNAiMAX (Thermo Fisher, 13778030). One day after transfection, the BMDMs were harvested and injected intravenously into WT B16-F10 tumor bearing mice (\(1\times {10}^{6}\) cells per mouse) at 1 day before B16-F10 implantation and 7 days after B16-F10 implantation. The oligonucleotide sequences have been summarized in Supplementary Data 11.
Tumor sizes were monitored by vernier calipers every 2 days. The largest tumor should not exceed 2 cm at the largest diameter according to the ethics committee; otherwise, the mouse should be euthanized. At the end of our experiments, the mice were euthanized with carbon dioxide. The tumors were removed and weighed.
B16-F10 lung metastatic model
B16-F10 cells (\(2\times {10}^{5}\)) were intravenously injected into WT and Tmem175−/− sex-matched mice at 6–8 weeks old. Referring to the experience in our laboratory, B16-F10 lung metastatic nodules do not grow excessively on day 21, which minimizes animal suffering. Mice were euthanized with carbon dioxide on day 21. The lungs were weighed, and the metastatic nodules were counted.
Single-cell RNA sequencing analysis
Lung tissues of the B16-F10 lung metastatic WT and Tmem175−/− mouse models were used in the single cell transcriptome analysis. Lungs were minced and digested to create single cell suspension. Then the ratio of live cells was confirmed to be around 90%. Single-cell RNA sequencing (scRNA-seq) libraries were constructed via the 10x Chromium Single Cell 5′ platform (10X Genomics), referring to the manufacturer’s protocol. Briefly, single-cell suspensions were added onto a Chromium microfluidic chip in order to generate single-cell Gel Beads in Emulsion. Then the scRNA-seq libraries were prepared through the Chromium Single Cell 5′ Reagent Kit v2 (10X Genomics). Library quality was assessed by the Agilent 2100 Bioanalyzer system (Agilent Technologies), and the sequencing was performed by the Illumina NovaSeq X Plus platform (Illumina) with a sequencing depth of higher than 50,000 reads per cell.
The acquired sequencing data were aligned to the mouse reference genome (GRCm39) and processed through CellRanger (version 9.0.0, https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/what-is-cell-ranger). The gene expression matrix obtained from the CellRanger pipeline was filtered and normalized via the Seurat R package (version 5.2, https://cran.r-project.org/web/packages/Seurat/index.html)84. Cells were included if they met the following criteria: (i) >200 genes and <4500 genes; (ii) <5% of mitochondrial gene expression in the UMI counts. After removing low-quality cells, the gene expression matrices were normalized to the total UMI counts per cell and transformed to a natural log scale. All datasets from the same group were integrated through the “FindIntegrationAnchors” and “IntegrateData” functions in Seurat. We identified “anchors” among all the individual datasets in the same group with multiple canonical correlation analyses (CCA) that have been implemented by the “FindIntegrationAnchors” function. These “anchors” were applied in the “IntegrateData” function to create a batch-corrected expression matrix of all the cells, which allowed the cells from different datasets to be integrated and analyzed.
The integrated data were scaled to a unit variance and zero mean, and the dimensionality was reduced by principal component analysis (PCA). A shared nearest neighbor (SNN) graph was constructed based on the distances in PCA space (top 1:50 principal components) with the “FindNeighbors” function, in which the k-nearest neighbors (KNN) of each cell were determined. Then this KNN graph was used to construct the SNN graph by calculating the neighborhood overlaps (Jaccard index) between each cell and its KNN. The Louvain algorithm was used to iteratively group the proximal cells together with the “FindClusters” function, with a resolution of 1. UMAP analysis was performed through RunUMAP (dims = 1:47) functions. Cell type annotation was performed via Azimuth based on Azimuth Mouse Lung CellRef (https://app.lungmap.net/app/azimuth-mouse-lung-cellref-seed)85 in company with marker-based annotations.
The “FindMarkers” function in Seurat was applied to find the markers for alveolar macrophages and interstitial macrophages. Genes that were significantly differentially expressed between the Tmem175−/− and WT groups (|log2FC| ≥ 0.5, Padj ≤ 0.01) were included in the functional enrichment analysis, which has been implemented by KOBAS. The P values of differently expressed genes were calculated by Wilcoxon rank-sum test and adjusted by Bonferroni correction. The enrichment analysis of comprehensive functions, GO terms, and KEGG pathways. The P values of GO and KEGG analysis were calculated by Fisher’s exact test and adjusted by Benjamini–Hochberg procedure.
Flow cytometry
For the analysis of TME, tumor tissues from euthanized mice were cut up and digested in DMEM medium supplied with 0.5 mg/mL collagenase I (Gibco, 17018029), 1 mg/mL collagenase VI (Gibco, 17104019), and 1 U/mL DNase I (KeyGEN BioTECH, KGA1506-1) for 30 min (for B16-F10) or 1 h (for LL-2). For lung metastatic models, the lungs were cut up and digested in DMEM medium supplied with 1 mg/mL collagenase I (Gibco, 17018029), 0.5 mg/mL collagenase VI (Gibco, 17104019), and 1 U/mL DNase I (KeyGEN BioTECH, KGA1506-1) for 1 h. The suspensions were filtered through 70 µm strainers (Falcon, CLS352350) and treated by red blood cell lysis. For lung metastatic models, cells were stained by H-2Kb MuLV p15E Tetramer (MBL, TS-M507) on ice for 1 h. Then cells were stained by antibodies and LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit (Thermo Fisher, L34975) for 30 min on ice. Alternatively, dead cells might also be stained with DAPI (Beyotime, C1005) for 5 min. For intracellular staining, the cells were fixed and permeabilized by Cytofix/Cytoperm Fixation/Permeabilization Kit (BD Biosciences, 554714) after surface antigen staining. Then the cells were stained by the desired antibodies for 1 h on ice. The cells were analyzed by ACEA NovoCyte (ACEA Biosciences).
For the analysis of intracellular antigen degradation, WT and Tmem175−/− BMDMs (\(1\times {10}^{6}\)) were treated with B16-F10 cell debris (2.5\(\times\)105/mL) for desired time (12, 24, and 48 h). At the same time, FITC-dextran (MCE, HY-128868E) (0.2 mg/mL) was incubated with WT and Tmem175−/− BMDMs (\(1\times {10}^{6}\)) for 48 h. At the end of the treatment, the cells were harvested and stained by desired antibodies and LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit (Thermo Fisher, L34975). Then the cells were fixed and permeabilized by Cytofix/Cytoperm Fixation/Permeabilization Kit (BD bioscience, 554714) for intracellular p15E staining. The cells were analyzed by ACEA NovoCyte (ACEA Biosciences).
The following antibodies were used: TMEM175 rabbit pAb (Proteintech, 19925-1-AP), FeLV p15e Antibody (Santa Cruz Biotechnology, sc-65622), FITC anti-mouse CD45 (BD Biosciences, 553080), PerCP-Cy5.5 anti-mouse CD45 (BioLegend, 103132), BV421 anti-mouse CD45 (BioLegend, 103133), BV510 anti-mouse CD45 (BioLegend, 103137), BV650 anti-mouse CD45 (BioLegend, 103151), BV785 anti-mouse CD45 (BioLegend, 103149), PE anti-mouse CD206 (BioLegend, 141706), APC anti-mouse CD206 (BioLegend, 141708), FITC anti-mouse CD206 (BioLegend, 141704), APC anti-mouse F4/80 (BioLegend, 123116), PE anti-mouse F4/80 (BioLegend, 123110), FITC anti-mouse F4/80 (BioLegend, 123108), BV421 anti-rabbit IgG (BD Biosciences, 565014), BV650 anti-mouse CD11b (BioLegend, 101239), FITC anti-mouse CD11b (BioLegend, 101206), PE-Cy7 anti-mouse CD11b (BioLegend, 101216), BV510 anti-mouse CD11b (BioLegend, 101245), BV421 anti-mouse CD11b (BioLegend, 101235), APC anti-mouse CD11b (BioLegend, 101212), PerCP-Cy5.5 anti-mouse CD11b (BioLegend, 101228), BV711 anti-mouse-I-A/I-E (BioLegend, 107643), FITC anti-mouse-I-A/I-E (BioLegend, 107606), BV650 anti-mouse-I-A/I-E (BioLegend, 107641), PE-Cy7 anti-mouse-I-A/I-E (BioLegend, 107630), PE-Cy7 anti-mouse-CD11c (BD Biosciences, 558079), FITC anti-mouse-CD11c (BioLegend, 117306), BV510 anti-mouse CD11c (BioLegend, 117337), APC anti-mouse-Ly6G (BD Biosciences, 560599), PE anti-mouse-Ly6G (BioLegend, 127608), BV421 anti-mouse-Ly6C (BioLegend, 128031), APC anti-mouse-Ly6C (BioLegend, 128016), FITC anti-mouse-CD8a (BD Biosciences, 553031), BV421 anti-mouse CD8a (BioLegend, 100737), BV510 anti-mouse CD8a (BioLegend, 100751), PE anti-mouse CD8a (BD Biosciences, 553033), PE anti-mouse-IFN-γ (BioLegend, 505808), BV421 anti-mouse-IFN-γ (BioLegend, 505829), PE-Cy7 anti-mouse-IFN-γ (BioLegend, 505826), PerCP-Cy5.5 anti-mouse CD3 (BioLegend, 100218), PE anti-mouse CD3 (BD Biosciences, 555275), BV421 anti-mouse CD3 (BioLegend, 100227), PE-Cy7 anti-mouse CD3 (BioLegend, 100220), PE anti-mouse granzyme B (BioLegend, 372208), PE-Cy7 anti-mouse granzyme B (BioLegend, 372214), AF647 anti-mouse granzyme B (BioLegend, 515406), BV421 anti-mouse CD69 (BioLegend, 104527), APC anti-mouse CD69 (BioLegend, 104514), PE anti-mouse CD69 (BD Biosciences, 553237), PE-Cy7 anti-mouse CD69 (BioLegend, 104512), FITC anti-mouse CD69 (BD Biosciences, 553236), PerCP-Cy5.5 anti-mouse CD69 (BD Biosciences, 561931), BV510 anti-mouse CD4 (BioLegend, 100553), FITC anti-mouse CD4 (BioLegend, 100406), PE anti-mouse CD4 (BioLegend, 100512), APC anti-mouse CD4 (BioLegend, 100516), PerCP-Cy5.5 anti-mouse CD4 (BD Biosciences, 550954), BV421 anti-mouse CD4 (BioLegend, 100543), BV650 anti-mouse NK1.1 (BioLegend, 108735), APC anti-mouse NK1.1 (BioLegend, 108710), BV421 anti-mouse NK1.1 (BioLegend, 108731), FITC anti-mouse CD64 (BioLegend, 139316), APC anti-mouse CD24 (BioLegend, 121435), BV711 anti-mouse CD103 (BioLegend, 138506), APC goat anti-mouse IgG (BioLegend, 405308).
The gating strategies of our FCM have been shown in Supplementary Figs. 18 and 19. The antibodies have been summarized in Supplementary Data 11.
Immunohistochemistry and immunofluorescence
Tissue chip of paired melanoma and paracancerous tissues was purchased from WEIAOBIO (Catalog: ZL-MEL963). All uses of human material have been approved. The IRB approval (approval number: LLS M-15-01) has been provided by the ethics committee of Shanghai Zhuoli Biotechnology Co., Ltd, whose general distributor is the WEIAOBIO. Tumor tissues from humanely euthanized mice were fixed in 4% paraformaldehyde, dehydrated, embedded in paraffin, sectioned at 3 µm, hydrated, and underwent antigen retrieval.
For immunohistochemistry, the sections were pre-treated with 3% H2O2 to remove endogenous peroxidase activity. Then the cell slides were blocked with goat serum and incubated with desired antibody and detect antibody successively. The cell slides were incubated with DAB substrate, counterstained by hematoxylin, dehydrated, and mounted. The following antibodies were used: TMEM175 antibody (Proteintech, 19925-1-AP), cleaved caspase-3 (Asp175) antibody (Cell Signaling Technology, 9661L), caspase-1 p10 antibody (SANTA CRUZ, sc-56036), IL-1β (3A6) Mouse antibody (Cell Signaling Technology, 12242S), HRP anti-rabbit IgG antibody (ServiceBio, GB23303), and HRP anti-mouse IgG antibody (ServiceBio, G1214). For immunofluorescence, the slides were incubated with NK1.1 mouse mAb (Thermo Fisher, 14-5941-82) and CD8 rat mAb (Abcam, ab25478) overnight. Then the slides were incubated with AF488 anti-rat IgG (Thermo Fisher, A-11006) and Cy3 anti-mouse IgG (Thermo Fisher, M30010). Finally, the slides were incubated with DAPI (Beyotime, C1005) and mounted. The TMEM175 positive area, caspase-1 p10 positive area, IL-1β positive area, cleaved caspase-3 positive area, CD8-AF488 positive area, and NK1.1-Cy3 positive area were measured by ImageJ software. The antibodies have been summarized in Supplementary Data 11.
Cross-presentation assay
B16-F10-OVA cells were implanted in wild type C57BL/6J mice and grew for 10 days to prime CD8+ T cells. Then the mice were humanely euthanized to collect the spleens, from which the lymphocytes were separated by Mouse lymphocyte separation medium (DAKEWE, DKW33-R0100). The lymphocytes (\(1\times {10}^{6}\)) were labeled by CFSE kit (Thermo Fisher) and then cultured with BMDMs (\(1\times {10}^{6}\)) with IL-2 (20 U/mL) (Novoprotein, CK24) and B16-F10-OVA cell debris (2.5\(\times\)105/mL) for 48 h with or without IL-1β (500 pg/mL) and IL-18 (500 pg/mL). After 48 h, the cells were harvested and stained by H-2Kb OVA Tetramer (MBL, TS-5001-1C), antibodies, and LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit (Thermo Fisher, L34975). The cells were analyzed by ACEA NovoCyte (ACEA Biosciences). The culture medium was collected for ELISA assay.
For the siRNA silencing, in vitro cultured WT or Tmem175−/− BMDMs were transfected with siRNA (GenePharma) targeting Myd88 (sense: 5′-GAAGCGACUGAUUCCUAUUTT-3′ anti-sense: 5′-AAUAGGAAUCAGUCGCUUCTT-3′) or Csf2rb (sense: 5′-GCCUGCAUUGGGAAACUCATT-3′ anti-sense: 5′-UGAGUUUCCCAAUGCAGGCTT-3′) via Lipofectamine RNAiMAX (Thermo Fisher, 13778030). The oligonucleotide sequences have been summarized in Supplementary Data 11.
In vitro stimulation of inflammasomes
B16-F10 tumor cell debris was the main inflammasome stimulator in our experiment. The concentration of cell debris was 2.5\(\times\)105/mL. The harvested cells were repeatedly blown by syringes followed by two freeze-thaw cycles to generate cell debris. BMDMs (\(1\times {10}^{6}\)) were activated by 500 ng/mL LPS (Sigma, L2880) for 4 h, then the medium was discarded, and the BMDMs were treated with cell debris for a desired period of time. During the treatment of cell debris, VX-765, CA-074, or KCl were added, whose final concentrations were 10 µg/mL, 100 µM, and 40 mM respectively. For positive control, 10 µM Nigericin (MCE, HY-127019) was added after LPS stimulation. For calcium chelation, cells were incubated with 20 µM BAPTA-AM (MCE, HY-100545) for 30 min after the LPS treatment. For ROS inhibition, cells were incubated with 10 mM NAC (MCE, HY-B0215) during the cell debris treatment. For Bafilomycin A1 treatment, cells were incubated with 1 µM Bafilomycin A1 (MCE, HY-100558) during the cell debris treatment. For the treatment of chitosan, BMDMs or peritoneal macrophages were activated by 500 ng/mL LPS (Sigma, L2880) for 4 h and then treated with 40 µg/mL chitosan (Sigma, 448869) for 5 h. For the treatment of aluminium hydroxide (Invivogen, vac-alu-50), BMDMs were activated by 500 ng/mL LPS (Sigma, L2880) for 4 h and then treated with 300 µg/mL aluminium hydroxide for a desired period of time. For the treatment of B16-F10 supernatant, BMDMs were activated by 500 ng/mL LPS (Sigma, L2880) for 4 h and then treated with 25% volume of filtered B16-F10 supernatant overnight.
ELISA
Cell culture supernatants were analyzed by Mouse IL-1 beta Uncoated ELISA Kit (Invitrogen, 88-7013A-88), Mouse TNF Uncoated ELISA Kit (Invitrogen, 88-7324-88), and Mouse IFN gamma “Femto-HS” High Sensitivity Uncoated ELISA Kit (Invitrogen, 88-8314-88) according to the manufacturer’s instructions.
Western blot
For the western blot analysis of cell lysis, BMDMs were lysed by RIPA (Beyotime, P0013B) and boiled with 1 \(\times\) SDS buffer (Beyotime, P0015L) for 5 min at 95 °C. Protein lysates were separated by SDS-PAGE and transferred to PVDF membranes (Millipore, IPVH00010). The membranes were probed by desired antibodies.
For the western blot analysis of cell supernatant, total proteins in the cell supernatant were precipitated by an equal volume of 10% trichloroacetic acid (TCA). The protein precipitants were washed by cold acetone three times. Then precipitants were subjected to air-drying for 5 min. The precipitants were dissolved by alkaline 1 \(\times\) SDS buffer (Beyotime, P0015L) plus 200 mM DTT (Thermo Fisher Scientific, R0861) and then boiled for 5 min at 95 °C. The grayscale values of TMEM175, β-actin, caspase-1 p10, IL-18 p24, ASC polymer, and cathepsin B were measured by ImageJ software. If the immunoblotting included β-actin, the results were presented as the ratio of the target protein/β-actin; otherwise, the grayscale values of the target protein were presented directly.
The following antibodies were used: TMEM175 mouse mAb (Proteintech, 68172-1-Ig), caspase-1 rabbit mAb (Abcam, 3345-1), IL-18 mouse mAb (LSBio, LS-C179849), β-actin mouse mAb (Santa Cruz Biotechnology, sc-47778), anti-rabbit IgG (HRP) (Thermo Fisher Scientific, 31460), anti-mouse IgG (HRP) (Thermo Fisher Scientific, 31430), and anti-rabbit IgG-Fc (HRP) (SinoBiological, SSA003). The antibodies have been summarized in Supplementary Data 11.
Co-immunoprecipitation
After the desired treatment, BMDMs were lysed by RIPA. Protein lysates were incubated with 1 µg/mL rabbit IgG isotype (Cell Signaling Technology, 3900S) and 20 µL Protein A + G Agarose (Beyotime, P2012) at 4 °C for 30 min to avoid non-specific combination. The lysates were centrifuged to remove the agarose beads and incubated with 1 µg/mL NLRP3 rabbit mAb (Cell Signaling Technology, 15101S) at 4 °C overnight, during which 5 mM EDTA-Na2 was added to the desired group. For negative control, the lysates were incubated overnight with 1 µg/mL isotype rabbit IgG. Afterward, the lysates were incubated with 20 µL Protein A + G Agarose (Beyotime, P2012) at 4 °C for 3 h. The beads were washed 3 times with RIPA. Finally, the beads were re-suspended in 1\(\,\times\) SDS buffer and boiled for 5 min at 95 °C.
The following antibodies were used: Cathepsin B rabbit mAb (Cell Signaling Technology, 31718S), ASC mouse mAb (Santa Cruz Biotechnology, sc-514414), NLRP3 rabbit mAb (Cell Signaling Technology, 15101S), Rabbit IgG isotype (Cell Signaling Technology, 3900S), anti-rabbit IgG (HRP) (Thermo Fisher Scientific, 31460), anti-mouse IgG (HRP) (Thermo Fisher Scientific, 31430), and anti-rabbit IgG-Fc (HRP) (SinoBiological, SSA003). The antibodies have been summarized in Supplementary Data 11.
Transcriptome analysis
Tmem175−/− and WT BMDMs were cultured, harvested, and lysed in TRIzol (Invitrogen, 15596018CN). Novogene Bioinformatics Institute, Beijing, China, performed the RNA extraction and sequencing. The raw data were processed by adapter trimming, and reads were filtered by trim galore version 0.5.0 to ensure data quality. The reads were mapped to Mus_musculus.GRCm38.dna.toplevel.fa and annotated with Mus_musculus.GRCm38.98.gtf.
To identify differentially expressed genes between Tmem175−/− and WT groups, fold change (FC) was calculated. The cut-off number of log2FC was set to be 1. The P values were calculated by likelihood ratio test, and adjusted by Benjamini–Hochberg procedure. The differentially expressed genes were further annotated using Gene Ontology and Kyoto Encyclopedia of Genes and Genomes database. The P values of GO and KEGG analysis were calculated by Fisher’s exact test and adjusted by Benjamini–Hochberg procedure.
The RNA-seq of the co-culture system was performed by DNBSEQ-T7 (MGI Tech). Clean reads were obtained by removing the adaptor sequences, reads with >5% ambiguous bases, and low-quality reads containing more than 20% of bases with qualities of <13. The clean reads were aligned to mouse genome (version: GRCh38 NCBI) using hisat2. The DESeq algorithm was used to filter the differentially expressed genes (Fold Change>1.1 or <0.9, FDR < 0.05). The P values were calculated by Negative binomial distribution test, and adjusted by Benjamini–Hochberg procedure. For GO enrichment analysis, GO annotations were downloaded from NCBI, UniProt, and the Gene Ontology. Significant GO categories were identified by Fisher’s exact test, and the p-value was corrected by FDR through Benjamini–Hochberg procedure. Significant KEGG pathways were selected by Fisher’s exact test, and the significance thresholds were defined by p-value and FDR through Benjamini–Hochberg procedure.
Electron microscopy
BMDMs were harvested, fixed with 3% glutaraldehyde, and re-fixed with 1% osmium tetroxide. The pellets were dehydrated with a series of acetone and embedded in Ep812. The ultrathin sections were cut by a diamond knife and stained by uranyl acetate and lead citrate. Images were taken by JEM-1400-FLASH Transmission Electron Microscope.
Lysosomal pH determination
The determination of lysosomal pH was performed using LysoSensor Yellow/Blue DND-160 (Invitrogen, L7545) according to manufacturers’ instructions. BMDMs were seeded in non-transparent 96-well plates. BMDMs were incubated with 5 µM LysoSensor Yellow/Blue DND-160 for 5 min after cell debris stimulation. Then the cells were washed twice by phenol red-free 1640 (Gibco, 11835030). The plates were read by Microplate Reader (PerkinElmer). Two fluorescence intensities, 329 nm excitation/440 nm emission and 384 nm excitation/540 nm emission, were collected. The ratio of 440 nm/540 nm was calculated for every well. We created a standard curve for pH calculation with a series of solutions with known pH at 3, 4, 5, 6, and 7. The LysoSensor Yellow/Blue DND-160 fluorescents were collected, and the 440 nm/540 nm ratio of each solution was calculated and used to create the standard curve. The values of lysosomal pH were calculated by referring to the standard curve.
Lysosomal calcium determination
The determination of lysosomal calcium determination was performed using AF488-dextran (Invitrogen, D34682) and rhod-dextran (AAT Bioquest, 20451) according to previous report86. Briefly, BMDMs were seeded in non-transparent 96-well plates and incubated with 0.1 mg/mL rhod-dextran and 0.2 mg/mL AF488-dextran overnight. The cells were then washed twice and stimulated with LPS and B16-F10 tumor cell debris as mentioned above in phenol red-free 1640 (Gibco, 11835030). The plates were read by a Microplate Reader (PerkinElmer). At the same time, standard wells with known calcium concentration were used as control. Two fluorescence intensities, 543 nm excitation/560 nm emission and 488 nm excitation/530 nm emission, were collected. The lysosomal calcium concentrations were calculated using the following equations: \([{{Ca}}^{2+}]={K}_{d}\times \frac{{S}_{f2}}{{S}_{b2}}\times \frac{(R-{R}_{\min })}{({R}_{\max }-R)}\). R was the ratio of 560 nm/530 nm for a single well at a certain time point; Rmin was the lowest 560 nm/530 nm ratio; Rmax was the highest 560 nm/530 nm ratio of a saturating calcium concentration. Sf2 and Sb2 showed fluorescence of 530 nm in the absence and presence of calcium, respectively. Kd is the dissociation constant determined by the standard curve with pH at 5, which equals to the concentration of calcium at which 560 nm fluorescence of rhod-dextran reached 50% brightness. In the desired wells, 1 µM Bafilomycin A1 (MCE, HY-100558) was added during the cell debris treatment.
Cytoplasmic calcium determination
The determination of cytoplasmic calcium was performed using Fluo-4-AM (Invitrogen, F14201). Briefly, BMDMs were seeded in non-transparent 96-well plates. BMDMs were incubated with 2 µM Fluo-4-AM for 30 min after cell debris stimulation. Then the cells were washed twice in phenol red-free 1640 (Gibco, 11835030). The plates were read by a Microplate Reader (PerkinElmer). Fluorescence intensity 480 nm excitation/525 nm emission was collected. The fluorescence of Fluo-4 was positively related to the concentration of cytoplasmic calcium; thus, the final results were presented as the ratio of 525 nm fluorescence in each group to the average 525 nm fluorescence in the blank group. In the desired wells, 1 µM Bafilomycin A1 (MCE, HY-100558) or 100 µM CA-074 (MCE, HY-103350) was added during the cell debris treatment.
Confocal microscope
BMDMs were seeded in glass slips and treated by LPS and B16-F10 tumor cell debris mentioned above. The cells were fixed and permeabilized by Cytofix/Cytoperm Fixation/Permeabilization Kit (BD bioscience, 554714). Then the cells were incubated with the desired antibodies overnight and the desired fluorescent secondary antibodies at room temperature. For cathepsin B rabbit mAb (Cell Signaling Technology, 31718S), the fluorescent secondary antibodies were Cy3 goat anti-rabbit IgG (Abcam, ab98416). For cathepsin D mAb (HUABIO, ET1608-49) and cathepsin L mAb (HUABIO, HA722063), the fluorescent secondary antibodies were AF488 goat anti-rabbit IgG (Thermo Fisher Scientific, A-11034). Nuclei were stained with DAPI (Beyotime, C1005). Imaging was performed using laser scanning confocal microscope (Olympus), and the images were analyzed using CellSens Dimension Software (Olympus).
ROS detection
The determination of ROS was performed using ROS probe, H2DCFDA (Invitrogen, D399). Briefly, BMDMs were seeded in non-transparent 96-well plates. BMDMs were incubated with 1 µM H2DCFDA for 30 min after cell debris stimulation. Then the cells were washed twice in phenol red-free 1640 (Gibco, 11835030). The plates were read by a Microplate Reader (PerkinElmer). Fluorescence intensity 488 nm excitation/520 nm emission was collected. The fluorescence intensity was positively related to the concentration of ROS.
Molecular dynamics simulation
The model of PYD domain from ASC was downloaded from protein data bank (PDB: 2N1F). The model of PYD domain from NLRP3 was built by Swiss-model87. The GMQE was 0.88, and the QMEANS was 0.81\(\pm\)0.05, indicating a reliable structure generation. The molecular docking was performed using HDock88, in which the docking score was 176.33 kcal/mol. The molecular docking result was used as the initiating conformation in the molecular dynamics simulation. Gromacs2019.689 was applied, and Amber14SB was selected as protein force field. The water model was established by TIP3P. Ten calcium ions were randomly added, and the sodium ion/chloride system was founded. The particle-mesh Ewald algorithm was used for electrostatic calculations, in which the system was minimized via steepest descent algorithm for 50,000 steps. The cut-off distance of both Coulomb force and the van der Waals radii was 1.4 nm. Canonical ensemble (NVT) and constant-temperature, constant-pressure ensemble (NPT) were adopted. Molecular dynamics simulation was performed for 100 ns at normal temperature and pressure. The results were displayed by PyMOL 2.5.190 and LigPlot 2.191.
Statistical analysis
Results from two or three independent experiments are presented as average values ± SEM. The number of biological replicates is shown in the figure legends. All statistical analyses were performed by linear regression, two-sided unpaired t-test (for 2 groups), one-way ANOVA (for 3 groups), or two-way ANOVA (for ≥ 2 variables or ≥ 4 groups) using GraphPad Prism 8 (GraphPad Prism Software Inc., La Jolla). The statistical methods to calculate the statistical significances were indicated in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for all the charts and graphs are available in the Source Data file whenever possible. The RNA seq and single cell RNA seq data generated in this work have been deposited in the GEO databases under the GSE248359, GSE291434, and GSE291346. Source data are provided with this paper.
References
Kennedy, L. B. & Salama, A. K. S. A review of cancer immunotherapy toxicity. CA Cancer J. Clin. 70, 86–104 (2020).
Shui, I. M. et al. Baseline and post-treatment biomarkers of resistance to anti-PD-1 therapy in acral and mucosal melanoma: an observational study. J. Immunother. Cancer 10, e004879 (2022).
Ruffell, B. & Coussens, L. M. Macrophages and therapeutic resistance in cancer. Cancer Cell 27, 462–472 (2015).
Madsen, D. H. et al. Tumor-associated macrophages derived from circulating inflammatory monocytes degrade collagen through cellular uptake. Cell Rep. 21, 3662–3671 (2017).
Ruffell, B. et al. Macrophage IL-10 blocks CD8+ T cell-dependent responses to chemotherapy by suppressing IL-12 expression in intratumoral dendritic cells. Cancer Cell 26, 623–637 (2014).
Zhu, Z. et al. PD-L1-mediated immunosuppression in glioblastoma is associated with the infiltration and M2-polarization of tumor-associated macrophages. Front. Immunol. 11, 588552 (2020).
Wang, B. et al. Transition of tumor-associated macrophages from MHC class II(hi) to MHC class II(low) mediates tumor progression in mice. BMC Immunol. 12, 43 (2011).
Müller, E. et al. Both type I and type II interferons can activate antitumor M1 macrophages when combined with TLR stimulation. Front. Immunol. 9, 2520 (2018).
Chen, D. et al. Chloroquine modulates antitumor immune response by resetting tumor-associated macrophages toward M1 phenotype. Nat. Commun. 9, 873 (2018).
Li, Y. et al. Hydroxychloroquine induced lung cancer suppression by enhancing chemo-sensitization and promoting the transition of M2-TAMs to M1-like macrophages. J. Exp. Clin. Cancer Res. 37, 259 (2018).
Vasiljeva, O. et al. Tumor cell–derived and macrophage-derived cathepsin B promotes progression and lung metastasis of mammary cancer. Cancer Res. 66, 5242–5250 (2006).
Gocheva, V. et al. Abstract LB-379: IL-4 induces cathepsin protease activity in tumor-associated macrophages to promote cancer growth and invasion. Genes Dev. 24, 241–255 (2010).
Zhang, Z. et al. Role of lysosomes in physiological activities, diseases, and therapy. J. Hematol. Oncol. 14, 79 (2021).
Bonam, S. R., Wang, F. & Muller, S. Lysosomes as a therapeutic target. Nat. Rev. Drug Discov. 18, 923–948 (2019).
Forgac, M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 8, 917–929 (2007).
Xiong, J. & Zhu, M. X. Regulation of lysosomal ion homeostasis by channels and transporters. Sci. China Life Sci. 59, 777–791 (2016).
Cang, C., Aranda, K., Seo, Y. J., Gasnier, B. & Ren, D. TMEM175 is an organelle K(+) channel regulating lysosomal function. Cell 162, 1101–1112 (2015).
Jinn, S. et al. TMEM175 deficiency impairs lysosomal and mitochondrial function and increases alpha-synuclein aggregation. Proc. Natl. Acad. Sci. USA 114, 2389–2394 (2017).
Zheng, W. et al. pH regulates potassium conductance and drives a constitutive proton current in human TMEM175. Sci. Adv. 8, eabm1568 (2022).
Hu, M. et al. Parkinson’s disease-risk protein TMEM175 is a proton-activated proton channel in lysosomes. Cell 185, 2292–2308 e2220 (2022).
Wie, J. et al. A growth-factor-activated lysosomal K(+) channel regulates Parkinson’s pathology. Nature 591, 431–437 (2021).
Zhang, Z., Li, X., Wang, Y., Wei, Y. & Wei, X. Involvement of inflammasomes in tumor microenvironment and tumor therapies. J. Hematol. Oncol. 16, 24 (2023).
Chauhan, D., Vande Walle, L. & Lamkanfi, M. Therapeutic modulation of inflammasome pathways. Immunol. Rev. 297, 123–138 (2020).
Chan, A. H. & Schroder, K. Inflammasome signaling and regulation of interleukin-1 family cytokines. J. Exp. Med. 217, e20190314 (2020).
Sheedy, F. et al. CD36 coordinates NLRP3 inflammasome activation by facilitating the intracellular nucleation from soluble to particulate ligands in sterile inflammation. Nat. Immunol. 14, 812–20 (2013).
Ershaid, N. et al. NLRP3 inflammasome in fibroblasts links tissue damage with inflammation in breast cancer progression and metastasis. Nat. Commun. 10, 4375 (2019).
Weichand, B. et al. S1PR1 on tumor-associated macrophages promotes lymphangiogenesis and metastasis via NLRP3/IL-1b. J. Exp. Med. 214, 2695–2713 (2017).
Das, S., Shapiro, B., Vucic, E. A., Vogt, S. & Bar-Sagi, D. Tumor cell-derived IL1beta promotes desmoplasia and immune suppression in pancreatic cancer. Cancer Res. 80, 1088–1101 (2020).
Bruchard, M. et al. Chemotherapy-triggered cathepsin B release in myeloid-derived suppressor cells activates the Nlrp3 inflammasome and promotes tumor growth. Nat. Med. 19, 57–64 (2013).
Deng, Q. et al. NLRP3 inflammasomes in macrophages drive colorectal cancer metastasis to the liver. Cancer Lett. 442, 21–30 (2019).
Dupaul-Chicoine, J. et al. The Nlrp3 inflammasome suppresses colorectal cancer metastatic growth in the liver by promoting natural killer cell tumoricidal activity. Immunity 43, 751–763 (2015).
Ghiringhelli, F. et al. Activation of the NLRP3 inflammasome in dendritic cells induces IL-1beta-dependent adaptive immunity against tumors. Nat. Med. 15, 1170–1178 (2009).
Chen, L. et al. Blockage of the NLRP3 inflammasome by MCC950 improves anti-tumor immune responses in head and neck squamous cell carcinoma. Cell. Mol. Life Sci. CMLS 75, 2045–2058 (2018).
Zhou, T. et al. IL-18BP is a secreted immune checkpoint and barrier to IL-18 immunotherapy. Nature 583, 609–614 (2020).
Voigt, C. et al. Cancer cells induce interleukin-22 production from memory CD4(+) T cells via interleukin-1 to promote tumor growth. Proc. Natl. Acad. Sci. USA 114, 12994–12999 (2017).
Wang, D. H. et al. Progression of EGFR-mutant lung adenocarcinoma is driven by alveolar macrophages. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 23, 778–788 (2017).
Hu, X., Gu, Y., Zhao, S., Hua, S. & Jiang, Y. Increased IL-10+CD206+CD14+M2-like macrophages in alveolar lavage fluid of patients with small cell lung cancer. Cancer Immunol. Immunother. CII 69, 2547–2560 (2020).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, eaau0964 (2019).
Schyns, J. et al. Non-classical tissue monocytes and two functionally distinct populations of interstitial macrophages populate the mouse lung. Nat. Commun. 10, 3964 (2019).
Shi, T., Denney, L., An, H., Ho, L. P. & Zheng, Y. Alveolar and lung interstitial macrophages: definitions, functions, and roles in lung fibrosis. J. Leukoc. Biol. 110, 107–114 (2021).
Hofbauer, D. et al. beta(2)-microglobulin triggers NLRP3 inflammasome activation in tumor-associated macrophages to promote multiple myeloma progression. Immunity 54, 1772–1787.e1779 (2021).
Chai, D. et al. Absent in melanoma 2-mediating M1 macrophages facilitate tumor rejection in renal carcinoma. Transl. Oncol. 14, 101018 (2021).
Flores, J. et al. Caspase-1 inhibition alleviates cognitive impairment and neuropathology in an Alzheimer’s disease mouse model. Nat. Commun. 9, 3916 (2018).
Bueter, C. L. et al. Chitosan but not chitin activates the inflammasome by a mechanism dependent upon phagocytosis. J. Biol. Chem. 286, 35447–35455 (2011).
Hornung, V. et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat. Immunol. 9, 847–856 (2008).
Timperi, E. et al. IL-18 receptor marks functional CD8+ T cells in non-small cell lung cancer. Oncoimmunology 6, e1328337 (2017).
Li, X. Y. et al. Targeting CD39 in cancer reveals an extracellular ATP- and inflammasome-driven tumor immunity. Cancer Discov. 9, 1754–1773 (2019).
Tengesdal, I. W. et al. Targeting tumor-derived NLRP3 reduces melanoma progression by limiting MDSCs expansion. Proc. Natl. Acad. Sci. USA 118, e2000915118 (2021).
Nakamura, K. et al. Dysregulated IL-18 is a key driver of immunosuppression and a possible therapeutic target in the multiple myeloma microenvironment. Cancer cell 33, 634–648.e635 (2018).
MacNabb, B. W. et al. Dendritic cells can prime anti-tumor CD8(+) T cell responses through major histocompatibility complex cross-dressing. Immunity 55, 982–997.e988 (2022).
Lu, Z. et al. Tumor factors stimulate lysosomal degradation of tumor antigens and undermine their cross-presentation in lung cancer. Nat. Commun. 13, 6623 (2022).
Savina, A. et al. NOX2 controls phagosomal pH to regulate antigen processing during crosspresentation by dendritic cells. Cell 126, 205–218 (2006).
Delamarre, L., Pack, M., Chang, H., Mellman, I. & Trombetta, E. S. Differential lysosomal proteolysis in antigen-presenting cells determines antigen fate. Science 307, 1630–1634 (2005).
Munoz-Planillo, R. et al. K(+) efflux is the common trigger of NLRP3 inflammasome activation by bacterial toxins and particulate matter. Immunity 38, 1142–1153 (2013).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Christensen, K. A., Myers, J. T. & Swanson, J. A. pH-dependent regulation of lysosomal calcium in macrophages. J. Cell Sci. 115, 599–607 (2002).
Lee, G. S. et al. The calcium-sensing receptor regulates the NLRP3 inflammasome through Ca2+ and cAMP. Nature 492, 123–127 (2012).
Murakami, T. et al. Critical role for calcium mobilization in activation of the NLRP3 inflammasome. Proc. Natl. Acad. Sci. USA 109, 11282–11287 (2012).
Sayin, V. I. et al. Antioxidants accelerate lung cancer progression in mice. Sci. Transl. Med. 6, 221ra215 (2014).
Wiel, C. et al. BACH1 stabilization by antioxidants stimulates lung cancer metastasis. Cell 178, 330–345.e322 (2019).
Piskounova, E. et al. Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527, 186–191 (2015).
Rao, G. et al. Anti-PD-1 induces M1 polarization in the glioma microenvironment and exerts therapeutic efficacy in the absence of CD8 cytotoxic T cells. Clin. Cancer Res. 26, 4699–4712 (2020).
Bagchi, S., Yuan, R. & Engleman, E. G. Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu. Rev. Pathol. 16, 223–249 (2021).
Shah, N. N. & Fry, T. J. Mechanisms of resistance to CAR T cell therapy. Nat. Rev. Clin. Oncol. 16, 372–385 (2019).
Zhang, J. et al. Lysosomal LAMP proteins regulate lysosomal pH by direct inhibition of the TMEM175 channel. Mol. Cell 83, 2524–2539.e2527 (2023).
Chen, Q. et al. BRCA1 deficiency impairs mitophagy and promotes inflammasome activation and mammary tumor metastasis. Adv. Sci. 7, 1903616 (2020).
Yang, D. et al. LFG-500, a novel synthetic flavonoid, suppresses epithelial-mesenchymal transition in human lung adenocarcinoma cells by inhibiting NLRP3 in inflammatory microenvironment. Cancer Lett. 400, 137–148 (2017).
Xu, Y. et al. Mycoplasma hyorhinis activates the NLRP3 inflammasome and promotes migration and invasion of gastric cancer cells. PLoS ONE 8, e77955 (2013).
Sauter, K. A., Wood, L. J., Wong, J., Iordanov, M. & Magun, B. E. Doxorubicin and daunorubicin induce processing and release of interleukin-1beta through activation of the NLRP3 inflammasome. Cancer Biol. Ther. 11, 1008–1016 (2011).
Wong, J., Tran, L. T., Magun, E. A., Magun, B. E. & Wood, L. J. Production of IL-1beta by bone marrow-derived macrophages in response to chemotherapeutic drugs: synergistic effects of doxorubicin and vincristine. Cancer Biol. Ther. 15, 1395–1403 (2014).
Voloshin, T. et al. Blocking IL1beta pathway following paclitaxel chemotherapy slightly inhibits primary tumor growth but promotes spontaneous metastasis. Mol. Cancer Ther. 14, 1385–1394 (2015).
Tang, T. T. et al. Hydroxychloroquine attenuates renal ischemia/reperfusion injury by inhibiting cathepsin mediated NLRP3 inflammasome activation. Cell Death Dis. 9, 351 (2018).
Lunov, O. et al. Amino-functionalized polystyrene nanoparticles activate the NLRP3 inflammasome in human macrophages. ACS Nano 5, 9648–9657 (2011).
Jager, E. et al. Calcium-sensing receptor-mediated NLRP3 inflammasome response to calciprotein particles drives inflammation in rheumatoid arthritis. Nat. Commun. 11, 4243 (2020).
Rossol, M. et al. Extracellular Ca2+ is a danger signal activating the NLRP3 inflammasome through G protein-coupled calcium sensing receptors. Nat. Commun. 3, 1329 (2012).
Horng, T. Calcium signaling and mitochondrial destabilization in the triggering of the NLRP3 inflammasome. Trends Immunol. 35, 253–261 (2014).
Genin, M., Clement, F., Fattaccioli, A., Raes, M. & Michiels, C. M1 and M2 macrophages derived from THP-1 cells differentially modulate the response of cancer cells to etoposide. BMC Cancer 15, 577 (2015).
Kolb, R. et al. Obesity-associated NLRC4 inflammasome activation drives breast cancer progression. Nat. Commun. 7, 13007 (2016).
Lu, F. et al. NLRP3 inflammasome upregulates PD-L1 expression and contributes to immune suppression in lymphoma. Cancer Lett. 497, 178–189 (2021).
Tengesdal, I. W. et al. Tumor NLRP3-derived IL-1beta drives the IL-6/STAT3 axis resulting in sustained MDSC-mediated immunosuppression. Front. Immunol. 12, 661323 (2021).
Oh, S., Stix, R., Zhou, W., Faraldo-Gomez, J. D. & Hite, R. K. Mechanism of 4-aminopyridine inhibition of the lysosomal channel TMEM175. Proc. Natl. Acad. Sci. USA 119, e2208882119 (2022).
Oh, S. et al. Discovery of selective inhibitors for the lysosomal Parkinson’s disease channel TMEM175. J. Am. Chem. Soc. 33, 23230–23239 (2024).
Chen, C. C. et al. Patch-clamp technique to characterize ion channels in enlarged individual endolysosomes. Nat. Protoc. 12, 1639–1658 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Morgan, A. J., Davis, L. C. & Galione, A. Imaging approaches to measuring lysosomal calcium. Methods Cell Biol. 126, 159–195 (2015).
Kiefer, F., Arnold, K., Kunzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009).
Yan, Y., Zhang, D., Zhou, P., Li, B. & Huang, S. Y. HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 45, W365–W373 (2017).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Mooers, B. H. M. Shortcuts for faster image creation in PyMOL. Protein Sci. Publ. Protein Soc. 29, 268–276 (2020).
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011).
Acknowledgements
We thank Guobo Shen (West China Hospital, Sichuan University) for helpful discussion in our experiment. Fangfang Wang (Institute of hematology, West China Hospital, Sichuan University) for the help in FCM analysis. The illustrations in Fig. 6h and Supplementary Fig. 17a were created with https://BioRender.com. The illustrations of mice, bones, and spleens in Figs. 4d, n, 5a, m, 6a, m, 7i, and 8a, Supplementary Figs. 1c, 3a, l, 7a, 9f, 14a, m were created with https://BioRender.com. The figures are licensed under CC BY 4.0, and the licenses are indicated in the related legends. This work was supported by 1.3.5 project for disciplines of excellence from West China Hospital of Sichuan University (ZYGD23038, X.W.), the National Key Research and Development Program of China (2024YFC2310700, X.W.), the National Natural Science Foundation of China (No. 82303752, Z.Z. and No. 82303723, T.L.), the China Postdoctoral Science Foundation (2023M732466, Z.Z.), the Natural Science Foundation of Sichuan Province (2024NSFSC1948, Z.Z.).
Author information
Authors and Affiliations
Contributions
Ziqi Zhang performed the experiments, analyzed the data, and wrote the manuscript. Xue li, Tianqi Lu, Zhenfei Bi, Jingyun Yang, Jingwen Luo, Aqu Alu, and Weiqi Hong helped with the in vivo experiments and FCM analysis of TAMs. Siyuan Chen, Lei Chen, Xuejiao Han, Yuan Cheng, Xuemei He, Chenjing Zhu, Manni Wang, and Yang Wang helped with the in vitro experiments including transcriptome data analysis. Lu Li, Hong Lei, Jie Lan, and Qingfang Li provided some key technical support for both in vivo and in vitro experiments. Jia Geng provided technical support specifically in lysosomal voltage clamp. Yuquan Wei provided conceptual advice and helped with the manuscript preparation. Xiawei Wei conceived the project, designed the experiments, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declar no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., li, X., Lu, T. et al. Deficiency of lysosomal TMEM175 in myeloid macrophages exerts anti-tumor immunity via inflammasome and cross-presentation pathway. Nat Commun 17, 2770 (2026). https://doi.org/10.1038/s41467-026-69546-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69546-2