Abstract
Regulatory T (Treg) cells expand during Mycobacterium tuberculosis (Mtb) infection and suppress T cell-mediated control. Whether Mtb actively contributes to this process is unclear. Here, using a genome-wide mutant library, we show that the expression of Mtb Rv1272c, an ATP-binding cassette transporter, increased under hypoxic conditions and promotes Mtb survival in vivo by increasing lecithin import, followed by the production and release of linoleic acid. Linoleic acid released by infected macrophages promoted surface trafficking of the immune checkpoint molecule cytotoxic T lymphocyte antigen 4 (CTLA-4) in Treg cells via the Ca²⁺ transporter ATP2a3. This in turn inhibited macrophage reactive oxygen species production and promoted Mtb survival inside macrophages. Rv1272c-induced linoleic acid further promoted Mtb immune evasion by increasing CTLA-4 surface trafficking on Treg cells in vivo. Mechanistically, linoleic acid interacts with ATP2a3 in Treg cells and promotes mitochondria-associated endoplasmic reticulum (ER) membrane formation. This facilitates ER-to-mitochondria Ca2+ transfer and depletion of Ca2+ in the ER, and triggers store-operated calcium entry, thus elevating cytosolic Ca2+ levels to increase Ca2+-dependent CTLA-4 surface trafficking in Treg cells. These findings reveal that Mtb can use a metabolite to manipulate host responses and promote its intracellular survival.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The metabolomics datasets for the analysis of Mtb-infected BMDM culture supernatant profiles are available in the MetaboLights database under accession number MTBLS12875. The CRISPR KO screening analysis data and the mass spectrometry data for comparative metabolomics analysis are provided as Supplementary Tables 1 and 2. Source data are provided with this paper.
Code availability
All analyses were done reproducibly using publicly available R scripts.
Change history
01 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41564-025-02232-z
References
Chai, Q., Wang, L., Liu, C. H. & Ge, B. New insights into the evasion of host innate immunity by Mycobacterium tuberculosis. Cell Mol. Immunol. 17, 901–913 (2020).
Cheng, H. et al. Mycobacterium tuberculosis produces D-serine under hypoxia to limit CD8+ T cell-dependent immunity in mice. Nat. Microbiol. 9, 1856–1872 (2024).
Kursar, M. et al. Cutting edge: regulatory T cells prevent efficient clearance of Mycobacterium tuberculosis. J. Immunol. 178, 2661–2665 (2007).
Cardona, P. & Cardona, P. J. Regulatory T cells in Mycobacterium tuberculosis infection. Front. Immunol. 10, 2139 (2019).
Zewdie, M. et al. Ex-vivo characterization of regulatory T cells in pulmonary tuberculosis patients, latently infected persons, and healthy endemic controls. Tuberculosis 100, 61–68 (2016).
Shafiani, S., Tucker-Heard, G., Kariyone, A., Takatsu, K. & Urdahl, K. B. Pathogen-specific regulatory T cells delay the arrival of effector T cells in the lung during early tuberculosis. J. Exp. Med. 207, 1409–1420 (2010).
Rowshanravan, B., Halliday, N. & Sansom, D. M. CTLA-4: a moving target in immunotherapy. Blood 131, 58–67 (2018).
Liu, C. W. et al. Association of tuberculosis risk with genetic polymorphisms of the immune checkpoint genes PDCD1, CTLA-4, and TIM3. PLoS ONE 19, e0303431 (2024).
Wang, P. H. et al. The dynamic change of immune checkpoints and CD14+ monocytes in latent tuberculosis infection. Biomedicines 9, 1479 (2021).
Shafiani, S. et al. Pathogen-specific Treg cells expand early during Mycobacterium tuberculosis infection but are later eliminated in response to interleukin-12. Immunity 38, 1261–1270 (2013).
Yan, M. Y., et al. A CRISPR-assisted nonhomologous end-joining strategy for efficient genome editing in Mycobacterium tuberculosis. mBio 11, e02364-19 (2020).
Vandal, O. H. et al. Acid-susceptible mutants of Mycobacterium tuberculosis share hypersusceptibility to cell wall and oxidative stress and to the host environment. J. Bacteriol. 191, 625–631 (2009).
Yu, J. et al. Structure and mechanism of a mycobacterial isoniazid efflux pump MsRv1273c/72c with a degenerate nucleotide-binding site. Nat. Commun. 16, 3969 (2025).
Martin, A. & Daniel, J. The ABC transporter Rv1272c of Mycobacterium tuberculosis enhances the import of long-chain fatty acids in Escherichia coli. Biochem. Biophys. Res. Commun. 496, 667–672 (2018).
Bomalaski, J. S., Steiner, M. R., Simon, P. L. & Clark, M. A. IL-1 increases phospholipase A2 activity, expression of phospholipase A2-activating protein, and release of linoleic acid from the murine T helper cell line EL-4. J. Immunol. 148, 155–160 (1992).
Schaffer, J. E. Fatty acid transport: the roads taken. Am. J. Physiol. Endocrinol. Metab. 282, E239–E246 (2002).
Ekim Kocabey, A. & Schneiter, R. Human lipocalins bind and export fatty acids through the secretory pathway of yeast cells. Front. Microbiol. 14, 1309024 (2023).
Cavelier, C., Lorenzi, I., Rohrer, L. & von Eckardstein, A. Lipid efflux by the ATP-binding cassette transporters ABCA1 and ABCG1. Biochim. Biophys. Acta 1761, 655–666 (2006).
Mulholland, C. V. et al. Propionate prevents loss of the PDIM virulence lipid in Mycobacterium tuberculosis. Nat. Microbiol. 9, 1607–1618 (2024).
Quigley, J., et al. The cell wall lipid PDIM contributes to phagosomal escape and host cell exit of Mycobacterium tuberculosis. mBio 8, e00148-17 (2017).
Alarcon, P. et al. Oleic and linoleic acids induce the release of neutrophil extracellular traps via pannexin 1-dependent ATP release and P2X1 receptor activation. Front. Vet. Sci. 7, 260 (2020).
Jasenosky, L. D., Scriba, T. J., Hanekom, W. A. & Goldfeld, A. E. T cells and adaptive immunity to Mycobacterium tuberculosis in humans. Immunol. Rev. 264, 74–87 (2015).
Kim, J. M., Rasmussen, J. P. & Rudensky, A. Y. Regulatory T cells prevent catastrophic autoimmunity throughout the lifespan of mice. Nat. Immunol. 8, 191–197 (2007).
Simpson, T. R. et al. Fc-dependent depletion of tumor-infiltrating regulatory T cells co-defines the efficacy of anti-CTLA-4 therapy against melanoma. J. Exp. Med. 210, 1695–1710 (2013).
Zhang, S. et al. Berbamine promotes macrophage autophagy to clear Mycobacterium tuberculosis by regulating the ROS/Ca2+ axis. mBio 14, e0027223 (2023).
Pahari, S. et al. Induction of autophagy through CLEC4E in combination with TLR4: an innovative strategy to restrict the survival of Mycobacterium tuberculosis. Autophagy 16, 1021–1043 (2020).
Silva, S. L. R. et al. Emetine induces oxidative stress, cell differentiation and NF-κB inhibition, suppressing AML stem/progenitor cells. Cell Death Discov. 10, 201 (2024).
Dudhgaonkar, S. P., Tandan, S. K., Bhat, A. S., Jadhav, S. H. & Kumar, D. Synergistic anti-inflammatory interaction between meloxicam and aminoguanidine hydrochloride in carrageenan-induced acute inflammation in rats. Life Sci. 78, 1044–1048 (2006).
Pellegrini, J. M. et al. Neutrophil autophagy during human active tuberculosis is modulated by SLAMF1. Autophagy 17, 2629–2638 (2021).
Pizzimenti, S. et al. Oxidative stress-related mechanisms in melanoma and in the acquired resistance to targeted therapies. Antioxidants 10, 1942 (2021).
Remans, P. H. et al. CTLA-4IG suppresses reactive oxygen species by preventing synovial adherent cell-induced inactivation of Rap1, a Ras family GTPASE mediator of oxidative stress in rheumatoid arthritis T cells. Arthritis Rheum. 54, 3135–3143 (2006).
Gallegos, A. M., Pamer, E. G. & Glickman, M. S. Delayed protection by ESAT-6-specific effector CD4+ T cells after airborne M. tuberculosis infection. J. Exp. Med. 205, 2359–2368 (2008).
Linsley, P. S. & Golstein, P. Lymphocyte activation: T-cell regulation by CTLA-4. Curr. Biol. 6, 398–400 (1996).
Trebak, M. & Kinet, J. P. Calcium signalling in T cells. Nat. Rev. Immunol. 19, 154–169 (2019).
Prakriya, M. & Lewis, R. S. Store-operated calcium channels. Physiol. Rev. 95, 1383–1436 (2015).
Patergnani, S. et al. Calcium signaling around mitochondria associated membranes (MAMs). Cell Commun. Signal 9, 19 (2011).
Nava Lauson, C. B. et al. Linoleic acid potentiates CD8+ T cell metabolic fitness and antitumor immunity. Cell Metab. 35, 633–650 e639 (2023).
Tovey, S. C., Sun, Y. & Taylor, C. W. Rapid functional assays of intracellular Ca2+ channels. Nat. Protoc. 1, 259–263 (2006).
Hobson, B. D. et al. Subcellular and regional localization of mRNA translation in midbrain dopamine neurons. Cell Rep. 38, 110208 (2022).
Rule, C. S., Patrick, M. & Sandkvist, M. Measuring in vitro ATPase activity for enzymatic characterization. J. Vis. Exp. 114, 54305 (2016).
Chandra, P., Grigsby, S. J. & Philips, J. A. Immune evasion and provocation by Mycobacterium tuberculosis. Nat. Rev. Microbiol. 20, 750–766 (2022).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003).
Gupta, S. et al. Rv1273c, an ABC transporter of Mycobacterium tuberculosis promotes mycobacterial intracellular survival within macrophages via modulating the host cell immune response. Int. J. Biol. Macromol. 142, 320–331 (2020).
Ambrozova, G., Pekarova, M. & Lojek, A. Effect of polyunsaturated fatty acids on the reactive oxygen and nitrogen species production by raw 264.7 macrophages. Eur. J. Nutr. 49, 133–139 (2010).
Yan, B. et al. Linoleic acid metabolism activation in macrophages promotes the clearing of intracellular Staphylococcus aureus. Chem. Sci. 13, 12445–12460 (2022).
Magdalon, J. et al. Oral administration of oleic or linoleic acids modulates the production of inflammatory mediators by rat macrophages. Lipids 47, 803–812 (2012).
Jia, L. et al. Porphyromonas gingivalis aggravates colitis via a gut microbiota–linoleic acid metabolism–Th17/Treg cell balance axis. Nat. Commun. 15, 1617 (2024).
Lin, L. et al. Oleic acid availability impacts thymocyte preprogramming and subsequent peripheral Treg cell differentiation. Nat. Immunol. 25, 54–65 (2024).
Guichard, V., et al. Calcium-mediated shaping of naive CD4 T-cell phenotype and function. eLife 6, e27215 (2017).
Robert, C. A decade of immune-checkpoint inhibitors in cancer therapy. Nat. Commun. 11, 3801 (2020).
Lazar-Molnar, E. et al. Programmed death-1 (PD-1)-deficient mice are extraordinarily sensitive to tuberculosis. Proc. Natl Acad. Sci. USA 107, 13402–13407 (2010).
Tezera, L. B., et al. Anti-PD-1 immunotherapy leads to tuberculosis reactivation via dysregulation of TNF-alpha. eLife 9, e52668 (2020).
Jayaraman, P. et al. TIM3 mediates T cell exhaustion during Mycobacterium tuberculosis infection. PLoS Pathog. 12, e1005490 (2016).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Abe, I. et al. Lipolysis-derived linoleic acid drives beige fat progenitor cell proliferation. Dev. Cell 57, 2623–2637 e2628 (2022).
Rossi, A. M. & Taylor, C. W. Reliable measurement of free Ca2+ concentrations in the ER lumen using Mag-Fluo-4. Cell Calcium 87, 102188 (2020).
Tekguc, M., Wing, J. B., Osaki, M., Long, J. & Sakaguchi, S. Treg-expressed CTLA-4 depletes CD80/CD86 by trogocytosis, releasing free PD-L1 on antigen-presenting cells. Proc. Natl Acad. Sci. USA 118, e2023739118 (2021).
Acknowledgements
We thank B. Li (Shanghai Jiao-Tong University) for critical reading of the paper. We thank the members of the B.G. laboratory for discussions and technical assistance. This project was supported by grants from National Natural Science Foundation of China (32030038, 32188101, 918423003 and 31730025 to B.G.; 32394010 and 32394012 to Z.J., 82470001, 82270006, 82070007 to H.Y., 82122029, 82071776 to L.W.), the National Key R&D Program of China (2021YFA1300902 to H.Y.), Chinese National Program on Key Basic Research Project (2017YFA0505900 to B.G.), Science Project of Shanghai Municipal Health Commission (202040028 to Z.J.), the Fundamental Research Funds for the Central Universities (22120210624 and 22120240345 to Z.J.), CAMS Innovation Fund for Medical Sciences (2023-I2M-2-001 to Y.S.), and the Science and Technology Innovation Action Plan (STCSM, 24YF2735400 to H.C.).
Author information
Authors and Affiliations
Contributions
H.C., S. Li, H.L., Z.J., Y.S., H.Y. and B.G. designed the study. S. Li, H.C. and H.L. performed most of the experiments and analysed the data. M.Y., J.W., X.H., S. Liu, Y.S. and H.Y. constructed the H37Rv genome-wide CRISPR KO mutant library. S. Li, H.Y., J.H., Y.Y., X.C. and J.X.W. constructed the H37Rv KO strain, performed metabolomics analysis and detected the level of linoleic acid. P.C., Y.C., R.Z., L.W., L.Q., C.G.F. and Q.Z. assisted with the preparation of the paper. S. Li, H.C., X.H., J.W., Z.L., Y.C. and Z.J. performed mouse infection experiments. The Mtb strains were stored by X.H. All authors discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks William Jacobs, Ye Zheng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Rv1272c induces linoleic acid export from Mtb-infected macrophages.
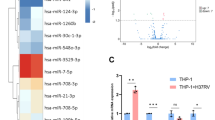
(a) Distribution of designed sgRNAs in the Mtb H37Rv genome-wide clustered regularly interspaced short palindromic repeats (CRISPR) knockout (KO) mutant library. (b) C57BL/6 mice were infected with CRISPR KO library. At 4 weeks after infection, lung homogenates were plated on 7H10 agar to recover surviving bacteria. Genes required for survival in mice but not for growth in vitro were analyzed. (c) Variation of all mutants with sgRNAs targeting rv1272c were analyzed by comparing sgRNA abundance in infected mice to the original screening library. The dashed lines indicate the values of fold change that are equal to 2 and 0.5. The red and blue bars represent Mtb mutants with corresponding sgRNA that can and cannot survive in mice respectively. (d) Immunoblot (IB) of cell lysates from Mtb H37Rv incubated under aeration and hypoxia. Levels of Rv1272c and SigA proteins were quantified using ImageJ and grayscale ratio of Rv1272c/SigA were indicated below blots. (e) IB of cell lysates of indicated Mtb strains. Levels of Rv1272c and SigA proteins were quantified using ImageJ and grayscale ratio of Rv1272c/SigA were indicated below blots. (f) C57BL/6 mice were aerosol-infected with indicated Mtb strains (~200 c.f.u. per mouse). Bacterial loads in lung tissues at 1 day post infection were assayed by c.f.u. (g) Impact factor histogram showing metabolic pathways of metabolites in indicated Mtb-infected bone marrow-derived macrophages (BMDMs) culture supernatants. The enrichment significance was determined through hypergeometric testing with FDR correction. (h, i) Linoleic acid (LA) concentration in the cell lysis (h) or culture supernatants (i) of Mtb-infected BMDMs for 24 h. (j, k) Linoleic acid concentration in bacterial lysis (j) or culture supernatants (k) of Mtb incubacted at normal condition for indicated days. (l, m) Assay of Lecithin concentration in lysis (l) or culture supernatants (m) of Mtb incubated with or without Lecithin (100 μM) for 14 days. (n) Schematic representation of experimental design for Fig. 1l, m. Data in d-f, h-k, l, m represent one experiment with three independent biological replicates (n= 3); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (f, h, i) and Two-way ANOVA with Tukey’s multiple comparisons test (j, k, l, m) were used for statistical analyses.
Extended Data Fig. 2 Linoleic acid exports from Mtb-infected macrophages through ATP-binding cassette transporter G1 (ABCG1).
(a-g) qPCR analysis of Fabp4 (a), Fabp5 (b), Fatp1 (c), Fatp2 (d), Obp2a (e), Abca1 (f) and Abcg1 (g) mRNA from BMDMs transfected with specific targeting siRNA or control siRNA. (h) Assay of linoleic acid concentration in the culture supernatants of Mtb-infected BMDMs transfected with Abcg1 specific siRNA or control siRNA for 24 h. (i) Assay of linoleic acid concentration in the culture supernatants of Mtb-infected BMDMs transfected with specific siRNA targeting Fabp4, Fabp5, Fatp1, Fatp2, Abca1, Obp2a, or control siRNA for 24 h. (j-m) qPCR analysis of Fabp4 (j), Fabp5 (k), Fatp1 (l) and Fatp2 (m) mRNA from BMDMs infected with Mtb H37Rv or H37RvΔRv1272c strains for 24 h. (n) Schematic representation of mice experimental design for Fig. 1n–p. C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of indicated Mtb strains and treated with or without linoleic acid. (o, p) C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of indicated Mtb strains and treated with or without linoleic acid. Linoleic acid concentrations in lung homogenate supernatants (o) and serum (p) were analyzed at 4 weeks post infection. Data in a-m, o, p represent one experiment with three independent biological replicates (n = 3); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (a-g, j-m, o, p) and Two-way ANOVA with Tukey’s multiple comparisons test (h, i) were used for statistical analyses.
Extended Data Fig. 3 TLC analysis of phthiocerol dimycocerosate (PDIM) production in Mtb strains.
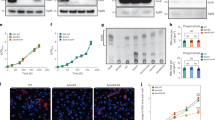
TLC analysis of PDIM production by Mtb H37Rv, H37RvΔRv1272c and H37Rv(ΔRv1272c + Rv1272c) strains. Three hundred micrograms of total lipid was loaded in each lane. The plate was resolved in a mobile phase of 9:1 petroleum ether-diethyl ether. Lipid spots were revealed by charring. Data of a represents one experiment with three independent biological replicates (n = 3).
Extended Data Fig. 4 Rv1272c or linoleic acid promotes Mtb survival in vivo via inhibiting adaptive immunity.
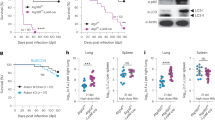
(a-d) WT mice and Rag1−/− mice were aerosol-infected with ~200 c.f.u. per mouse of indicated Mtb strains and treated with or without linoleic acid. After 4 weeks of infection, histopathology of lung sections from infected mice was assessed via acid-fast staining (a; scale bar = 100 μm [upper] and 20 μm [lower]), H&E staining (scale bar=1 mm) (b), bacterial c.f.u. (c) and quantified inflammatory areas (d). #1–3 indicate three representative lung sections. Quantified inflammatory areas are shown in sections. Data in a-d represent one experiment with three independent biological replicates (n = 3); mean ± s.e.m. Two-way ANOVA with Tukey’s multiple comparisons test (c, d) were used for statistical analyses.
Extended Data Fig. 5 Rv1272c or linoleic acid promotes the intramacrophage survival of Mtb via Treg cells.
(a) C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of indicated Mtb strains. Percentages of CD45+Ly6G+ cells were measured using FCM. (b-g) C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of indicated Mtb strains. IFN-γ, TGF-β and IL-10 concentrations in serum (b, d, f) and lung homogenate supernatants (c, e, g) were analyzed at 4 weeks post infection. (h) C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of Mtb H37Rv and administered with linoleic acid for 4 weeks. Mean Fluorescence Intensity (MFI) of membrane CTLA-4 levels were measured in CD4+ Tregs in lung tissues. (i) Schematic representation of mice experimental design for Fig. 2j, k. (j) Schematic representation of Mtb-infected BMDMs and CD4+ Tregs co-culture experimental design for Fig. 2m, n. (k) After 1 day of infection with Mtb strains, BMDMs were co-incubated with CD4+ Treg cells for 1 day, and percentage of Foxp3+CD4+ T cells was measured in CD4+ T cells. (l) After 1 day of infection with Mtb H37Rv strains, BMDMs were co-incubated with CD4+ Treg cells treated with or without linoleic acid for 24 h, percentage of Foxp3+CD4+ T cells was measured in CD4+ T cells. (m) After 1 day of infection with indicated Mtb strains (MOI = 5), BMDMs were co-incubated with CD4+ Treg cells treated with or without linoleic acid for 1 day, MFI of membrane CTLA-4 levels in CD4+ Tregs were measured by FCM. (n, o) C57BL/6 mice were aerosol-infected with approximately 200 c.f.u. per mouse of Mtb H37Rv and administered with linoleic acid for 4 weeks. MFI of membrane (n) and total (o) CTLA-4 levels were measured in CD4+ Tregs in spleen tissues. Data in a-h, k-o represent one experiment with at least three independent biological replicates (a,c-d,f-h,k-o, n = 3; b,e, n = 4); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (a-h, k, l, n, o) and Two-way ANOVA with Tukey’s multiple comparisons test (m) were used for statistical analyses. Panel j created with BioRender.com.
Extended Data Fig. 6 Linoleic acid promotes mycobacteria survival via CTLA-4 of Tregs in vivo.
(a) Naïve CD4+ T cells were stimulated with anti-CD3 and anti-CD28 antibodies in the presence of IL-2 and TGF-β for 3 days and treated with or without linoleic acid (100 μM). The percentage of Foxp3+CD4+ T cells was measured in CD4+ T cells. (b-d) Naïve CD4+ T cells were stimulated with anti-CD3 and anti-CD28 antibodies in the presence of IL-2 and TGF-β for 3 days and treated with or without linoleic acid (100 μM or 200 μM). MFI of total (b) and membrane (c) CTLA-4 levels of CD4+ Tregs were measured by FCM. (d) Immunofluorescence data shows Hoechst (Blue), membrane (Green), and CTLA-4 (Magenta) staining in CD4+ Tregs. Below, pixel intensity plot for dashed line; Scale bar = 1 μm. (e-g) C57BL/6 mice treated with/without anti-CTLA-4 antibodies (500 μg/mouse) on 4, 8, 12, 16, 20, and 24 days post-infection were aerosol-infected with approximately 200 c.f.u. per mouse of indicated Mtb strains and treated with or without linoleic acid via drinking water. After 4 weeks of infection, histopathology (scale bar=1 mm) of lung sections was assessed via H&E staining. #1–3 indicate three representative lung sections (e). Quantified inflammatory areas in the lung tissues of mice infected with Mtb strains were analyzed at 4 weeks post infection (f). The percentage of Foxp3+CD4+ T cells was measured in CD4+ T cells (g). Data in a-g represent one experiment with three independent biological replicates (n = 3); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (a-c, g) and Two-way ANOVA with Tukey’s multiple comparisons test (f) were used for statistical analyses.
Extended Data Fig. 7 Rv1272c induced linoleic acid promotes CTLA-4 surface trafficking on Tregs to inhibit cytosol ROS and facilitate the survival of Mtb in macrophages.
(a) Mtb-infected BMDMs (MOI = 5) were pretreated with indicated inhibitors for 24 h and then co-cultured with Tregs treated with isotype control IgG or anti-CTLA-4 blocking antibody (40 µg/mL) for 1 day, intracellular survival of the indicated Mtb strains in BMDMs infected for 2 days were determined using c.f.u. assay. (b) After 1 day of infection with indicated Mtb strains (MOI = 5), BMDMs were co-incubated with CD4+ ESAT6-specific Treg cells treated with or without linoleic acid for 24 h, MFI of membrane CTLA-4 levels in CD4+ Tregs were measured by FCM. (c) After 1 day of infection with indicated Mtb strains (MOI = 5), BMDMs were co-incubated with CD4+ ESAT6-specific Treg cells treated with or without linoleic acid for 24 h. MFI of ROS levels in BMDMs were measured by FCM. Data in a-e represent one experiment with at least three independent biological replicates (a, n = 4; b-c, n = 3); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (b) and Two-way ANOVA with Tukey’s multiple comparisons test (a, c) were used for statistical analyses.
Extended Data Fig. 8 Linoleic acid promotes the formation of mitochondria-associated ER membranes (MAMs).
Naïve CD4+ T cells were stimulated with anti-CD3 and anti-CD28 antibodies in the presence of IL-2 and TGF-β for 3 days and treated with or without linoleic acid (100 μM). Immunofluorescence data shows Hoechst (Blue), Mito-Traker (Green), and ER-Traker (Magenta) staining in CD4+ Tregs. Right, pixel intensity plot for dashed line. Scale bar = 1 μm. Data of a represents one experiment with three independent biological replicates (n = 3).
Extended Data Fig. 9 Linoleic acid interacts with ATP2a3.
(a) Representative mass spectrum of interacting proteins for linoleic acid identified by Liquid Chromatography Mass Spectrometry (LC-MS)/MS analysis. (b) Naïve CD4+ T cells were stimulated with anti-CD3 and anti-CD28 antibodies in the presence of IL-2 and TGF-β for 3 days and treated with DMSO or FITC-linoleic acid (100 μM). Immunofluorescence data shows Hoechst (Blue), linoleic acid-FITC (Green), and ATP2a3 (Magenta) staining in CD4+ Tregs. Scale bar = 1 μm. (c) Direct interaction of linoleic acid with ATP2a3 was detected using pull-down assay in vitro. (d) Predicted potential binding sites between linoleic acid and ATP2a3 using AlphaFold. (e) qPCR analysis of Atp2a3 mRNA from CD4+ Tregs transfected with Atp2a3 specific siRNA or control siRNA. (f) The ATP2a3 protein levels of splenic CD4+Foxp3+ Treg cells isolated from Foxp3cre (Cre) and Foxp3creAtp2a3fl/+(CKO) mice. Data in b, c, e, f represent one experiment with three independent biological replicates (n = 3); mean ± s.e.m. Two-tailed unpaired Student’s t-tests (e) was used for statistical analyses.
Extended Data Fig. 10 Summary diagram.
Hypoxia-induced Mtb Rv1272c enhanced the import of lecithin, leading to the generation of linoleic acid (LA) and subsequent release of LA from Mtb-infected macrophages through ATP Binding Cassette Subfamily G Member 1 (ABCG1). The released LA directly interacted with ATP2a3 and enhanced the formation of MAMs, which facilitated Ca²⁺ transfer from ER to mitochondria via the mitochondrial calcium uniporter (MCU), leading to ER Ca²⁺ depletion in Tregs. Reduced ER Ca²⁺ subsequently triggered store-operated calcium entry (SOCE), thus elevating cytosolic Ca²⁺ levels and Ca2+-dependent CTLA-4 surface trafficking, which reduced ROS levels in macrophages and promoted mycobacterial intracellular survival. Figure created with BioRender.com.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1.
Supplementary Tables (download XLSX )
Supplementary Tables 1–5.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Fig. 6 and Extended Data Figs. 1, 3 and 6 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, H., Li, S., Liu, H. et al. Mycobacterium tuberculosis-derived linoleic acid increases regulatory T cell function to promote bacterial survival within macrophages. Nat Microbiol 10, 2949–2965 (2025). https://doi.org/10.1038/s41564-025-02140-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02140-2