Abstract
DNA double-strand breaks (DSBs) disrupt the continuity of the genome, with consequences for malignant transformation. Massive DNA damage can elicit a cellular checkpoint response that prevents cell proliferation1,2. However, how highly aggressive cancer cells, which can tolerate widespread DNA damage, respond to DSBs alongside continuous chromosome duplication is unknown. Here we show that DSBs induce a local genome maintenance mechanism that inhibits replication initiation in DSB-containing topologically associating domains (TADs) without affecting DNA synthesis at other genomic locations. This process is facilitated by mediators of replication and DSBs (MRDs). In normal and cancer cells, MRDs include the TIMELESS–TIPIN complex and the WEE1 kinase, which actively dislodges the TIMELESS–TIPIN complex from replication origins adjacent to DSBs and prevents initiation of DNA synthesis at DSB-containing TADs. Dysregulation of MRDs, or disruption of 3D chromatin architecture by dissolving TADs, results in inadvertent replication in damaged chromatin and increased DNA damage in cancer cells. We propose that the intact MRD cascade precedes DSB repair to prevent genomic instability, which is otherwise observed when replication is forced, or when genome architecture is challenged, in the presence of DSBs3,4,5. These observations reveal a previously unknown vulnerability in the DNA replication machinery that may be exploited to therapeutically target cancer cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data have been deposited to the GEO with accession number GSE254206. Intermediate image analysis results and statistical analysis results for the RNAi screens have been deposited at GitHub (https://github.com/CBIIT/aladjemlab-mrd-screens/tree/main).
Code availability
R code for statistical analysis has been deposited at GitHub (https://github.com/CBIIT/aladjemlab-mrd-screens/tree/main).
References
Falck, J., Mailand, N., Syljuasen, R. G., Bartek, J. & Lukas, J. The ATM–Chk2–Cdc25A checkpoint pathway guards against radioresistant DNA synthesis. Nature 410, 842–847 (2001).
Bartek, J., Lukas, C. & Lukas, J. Checking on DNA damage in S phase. Nat. Rev. Mol. Cell Biol. 5, 792–804 (2004).
Peycheva, M. et al. DNA replication timing directly regulates the frequency of oncogenic chromosomal translocations. Science 377, eabj5502 (2022).
Spies, J. et al. 53BP1 nuclear bodies enforce replication timing at under-replicated DNA to limit heritable DNA damage. Nat. Cell Biol. 21, 487–497 (2019).
Wu, J. et al. Cohesin maintains replication timing to suppress DNA damage on cancer genes. Nat. Genet. 55, 1347–1358 (2023).
Aladjem, M. I. & Redon, C. E. Order from clutter: selective interactions at mammalian replication origins. Nat. Rev. Genet. 18, 101–116 (2017).
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Cortez, D. Preventing replication fork collapse to maintain genome integrity. DNA Repair 32, 149–157 (2015).
Thomas, A. & Pommier, Y. Small cell lung cancer: time to revisit DNA-damaging chemotherapy. Sci. Transl. Med. 8, 346fs312 (2016).
Furuta, T. et al. Phosphorylation of histone H2AX and activation of Mre11, Rad50, and Nbs1 in response to replication-dependent DNA double-strand breaks induced by mammalian DNA topoisomerase I cleavage complexes. J. Biol. Chem. 278, 20303–20312 (2003).
Clouaire, T. et al. Comprehensive mapping of histone modifications at DNA double-strand breaks deciphers repair pathway chromatin signatures. Mol. Cell 72, 250–262.e256 (2018).
Fu, H. et al. Dynamics of replication origin over-activation. Nat. Commun. 12, 3448 (2021).
Arnould, C. et al. Loop extrusion as a mechanism for formation of DNA damage repair foci. Nature 590, 660–665 (2021).
Arnould, C. et al. Chromatin compartmentalization regulates the response to DNA damage. Nature 623, 183–192 (2023).
Yesbolatova, A. et al. The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice. Nat. Commun. 11, 5701 (2020).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e324 (2017).
Emerson, D. J. et al. Cohesin-mediated loop anchors confine the locations of human replication origins. Nature 606, 812–819 (2022).
Silverman, J., Takai, H., Buonomo, S. B., Eisenhaber, F. & de Lange, T. Human Rif1, ortholog of a yeast telomeric protein, is regulated by ATM and 53BP1 and functions in the S-phase checkpoint. Genes Dev. 18, 2108–2119 (2004).
Setiaputra, D. et al. RIF1 acts in DNA repair through phosphopeptide recognition of 53BP1. Mol. Cell 82, 1359–1371.e1359 (2022).
Jones, M. L., Baris, Y., Taylor, M. R. G. & Yeeles, J. T. P. Structure of a human replisome shows the organisation and interactions of a DNA replication machine. EMBO J. 40, e108819 (2021).
Baris, Y., Taylor, M. R. G., Aria, V. & Yeeles, J. T. P. Fast and efficient DNA replication with purified human proteins. Nature 606, 204–210 (2022).
Somyajit, K. et al. Redox-sensitive alteration of replisome architecture safeguards genome integrity. Science 358, 797–802 (2017).
Utani, K. et al. Phosphorylated SIRT1 associates with replication origins to prevent excess replication initiation and preserve genomic stability. Nucleic Acids Res. 45, 7807–7824 (2017).
Thakur, B. L. et al. Convergence of SIRT1 and ATR signaling to modulate replication origin dormancy. Nucleic Acids Res. 50, 5111–5128 (2022).
Lanz, M. C., Dibitetto, D. & Smolka, M. B. DNA damage kinase signaling: checkpoint and repair at 30 years. EMBO J. 38, e101801 (2019).
De Witt Hamer, P. C., Mir, S. E., Noske, D., Van Noorden, C. J. & Wurdinger, T. WEE1 kinase targeting combined with DNA-damaging cancer therapy catalyzes mitotic catastrophe. Clin. Cancer Res. 17, 4200–4207 (2011).
Dabin, J., Mori, M. & Polo, S. E. The DNA damage response in the chromatin context: A coordinated process. Curr. Opin. Cell Biol. 82, 102176 (2023).
Merigliano, C. & Chiolo, I. Multi-scale dynamics of heterochromatin repair. Curr. Opin. Genet. Dev. 71, 206–215 (2021).
Tsouroula, K. et al. Temporal and spatial uncoupling of DNA double strand break repair pathways within mammalian heterochromatin. Mol. Cell 63, 293–305 (2016).
Beck, H. et al. Cyclin-dependent kinase suppression by WEE1 kinase protects the genome through control of replication initiation and nucleotide consumption. Mol. Cell. Biol. 32, 4226–4236 (2012).
Szmyd, R. et al. Premature activation of Cdk1 leads to mitotic events in S phase and embryonic lethality. Oncogene 38, 998–1018 (2019).
Hughes, B. T., Sidorova, J., Swanger, J., Monnat, R. J. Jr. & Clurman, B. E. Essential role for Cdk2 inhibitory phosphorylation during replication stress revealed by a human Cdk2 knockin mutation. Proc. Natl Acad. Sci. USA 110, 8954–8959 (2013).
Barlow, J. H. et al. Identification of early replicating fragile sites that contribute to genome instability. Cell 152, 620–632 (2013).
Lee, J. J. et al. ERα-associated translocations underlie oncogene amplifications in breast cancer. Nature 618, 1024–1032 (2023).
Bass, T. E., Fleenor, D. E., Burrell, P. E. & Kastan, M. B. ATM regulation of the cohesin complex is required for repression of DNA replication and transcription in the vicinity of DNA double-strand breaks. Mol. Cancer Res. 21, 261–273 (2023).
Kim, S. T., Xu, B. & Kastan, M. B. Involvement of the cohesin protein, Smc1, in Atm-dependent and independent responses to DNA damage. Genes Dev. 16, 560–570 (2002).
Zagelbaum, J. et al. Multiscale reorganization of the genome following DNA damage facilitates chromosome translocations via nuclear actin polymerization. Nat. Struct. Mol. Biol. 30, 99–106 (2023).
Sanders, J. T. et al. Radiation-induced DNA damage and repair effects on 3D genome organization. Nat. Commun. 11, 6178 (2020).
Yang, J. H., Brandao, H. B. & Hansen, A. S. DNA double-strand break end synapsis by DNA loop extrusion. Nat. Commun. 14, 1913 (2023).
Piazza, A. et al. Cohesin regulates homology search during recombinational DNA repair. Nat. Cell Biol. 23, 1176–1186 (2021).
Natale, F. et al. Identification of the elementary structural units of the DNA damage response. Nat. Commun. 8, 15760 (2017).
Dequeker, B. J. H. et al. MCM complexes are barriers that restrict cohesin-mediated loop extrusion. Nature 606, 197–203 (2022).
Xu, X., Wang, J. T., Li, M. & Liu, Y. TIMELESS suppresses the accumulation of aberrant CDC45.MCM2–7.GINS replicative helicase complexes on human chromatin. J. Biol. Chem. 291, 22544–22558 (2016).
Bantele, S. et al. Repair of DNA double-strand breaks leaves heritable impairment to genome function. Preprint at bioRxiv https://doi.org/10.1101/2023.08.29.555258 (2023).
Ochs, F. et al. Stabilization of chromatin topology safeguards genome integrity. Nature 574, 571–574 (2019).
Ghodke, I. et al. AHNAK controls 53BP1-mediated p53 response by restraining 53BP1 oligomerization and phase separation. Mol. Cell 81, 2596–2610.e2597 (2021).
Kilic, S. et al. Phase separation of 53BP1 determines liquid-like behavior of DNA repair compartments. EMBO J. 38, e101379 (2019).
Zhang, L. et al. 53BP1 regulates heterochromatin through liquid phase separation. Nat. Commun. 13, 360 (2022).
Patel, J. A. & Kim, H. The TIMELESS effort for timely DNA replication and protection. Cell. Mol. Life Sci. 80, 84 (2023).
Petropoulos, M. et al. Transcription–replication conflicts underlie sensitivity to PARP inhibitors. Nature 628, 433–441 (2024).
Chou, D. M. & Elledge, S. J. Tipin and Timeless form a mutually protective complex required for genotoxic stress resistance and checkpoint function. Proc. Natl Acad. Sci. USA 103, 18143–18147 (2006).
Katou, Y. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083 (2003).
Takahashi, N. et al. Replication stress defines distinct molecular subtypes across cancers. Cancer Res. Commun. 2, 503–517 (2022).
Patel, J. A. et al. Replisome dysfunction upon inducible TIMELESS degradation synergizes with ATR inhibition to trigger replication catastrophe. Nucleic Acids Res. 51, 6246–6263 (2023).
Bukhari, A. B. et al. Inhibiting Wee1 and ATR kinases produces tumor-selective synthetic lethality and suppresses metastasis. J. Clin. Invest. 129, 1329–1344 (2019).
Rodland, G. E. et al. Differential effects of combined ATR/WEE1 inhibition in cancer cells. Cancers 13, 3790 (2021).
Iacovoni, J. S. et al. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 29, 1446–1457 (2010).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
Fedkenheuer, M. et al. A dual role for cohesin in DNA DSB repair. Nat Commun. 16, 843 (2025).
Marchal, C. et al. Genome-wide analysis of replication timing by next-generation sequencing with E/L Repli-seq. Nat. Protoc. 13, 819–839 (2018).
Boutros, M., Bras, L. P. & Huber, W. Analysis of cell-based RNAi screens. Genome Biol. 7, R66 (2006).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Huang, J. et al. RAD18 transmits DNA damage signalling to elicit homologous recombination repair. Nat. Cell Biol. 11, 592–603 (2009).
Fu, H. et al. Mapping replication origin sequences in eukaryotic chromosomes. Curr. Protoc. Cell Biol. 65, 22.20.1–17 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Pongor, L. S. et al. BAMscale: quantification of next-generation sequencing peaks and generation of scaled coverage tracks. Epigenetics Chromatin 13, 21 (2020).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Gu, Z., Eils, R., Schlesner, M. & Ishaque, N. EnrichedHeatmap: an R/Bioconductor package for comprehensive visualization of genomic signal associations. BMC Genomics 19, 234 (2018).
Easwaran, H. et al. A DNA hypermethylation module for the stem/progenitor cell signature of cancer. Genome Res. 22, 837–849 (2012).
Rajapakse, V. N. et al. CellMinerCDB for integrative cross-database genomics and pharmacogenomics analyses of cancer cell lines. iScience 10, 247–264 (2018).
Acknowledgements
The authors thank M. Kruhlak for imaging support; L. Ozbun for support with RNAi screens; S. Shema and M. Wong for sequencing support; the Biowulf, High-Performance Computing Group, Center for Information Technology (CIT), NIH, for computational support; and S. Vartak, P. Oberdoerffer and A. Xiao for critical reading of the manuscript. High-throughput imaging work was performed at the High-Throughput Imaging Facility (HiTIF), Center for Cancer Research, National Cancer Institute, NIH. Next-generation sequencing was performed at the CCR Genomics Core, Center for Cancer Research, National Cancer Institute, NIH. Confocal imaging was performed at the CCR Confocal Microscopy Core, Center for Cancer Research, National Cancer Institute, NIH. This work was supported by the Intramural Research Program of the NIH, NCI, Center for Cancer Research (ZIA BC010411 to M.I.A.). R.S. acknowledges support from the NCI Director’s Intramural Innovation Award.
Author information
Authors and Affiliations
Contributions
R.S. and M.I.A. conceptualized and designed experiments. R.S. performed all experiments and coordinated those conducted by co-authors. E.G.S. performed analysis and plotting of sequencing experiments, confocal images and created illustrations and graphs. M.F. and R.C. conceptualized the LIG4−/− AID-RAD21 AsiSI-HCT116 cell line. M.F. designed, created and validated the LIG4−/− AID-RAD21 AsiSI-HCT116 cell line and performed MRE11 ChIP–seq. H.F. performed replication timing analyses experiments in U2OS cells, DNA fibre assays and asynchronous cell cycle analysis in U2OS and AsiSI-U2OS cells. S.J. analysed NS-seq and ChIP–seq for TIMELESS, TIPIN, MRE11, pSIRT1, MCM4 and MCM2 pS139. A.D.T. analysed super-resolution and confocal images. G.P. performed RNAi screen analyses. B.L.T. performed ChIP–seq for TIMELESS, TIPIN, pSIRT1, MCM4 and MCM2 pS139 and performed replication timing analysis. C.E.R. performed NS-seq and western blots for DDR proteins. L.S.P., J.M.G., S.M. and N.A.K. performed bioinformatic analysis of analyses of replication timing, BrdU-seq and γH2AX ChIP–seq. A.R. and A.D. analysed γH2AX ChIP–seq after WEE1i. M.I.A. and R.S. wrote the paper with insights from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Kumar Somyajit and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analysis of EdU intensity ratios using irradiation and AsiSI-induced DSBs.
a, Experimental strategy for assaying EdU intensity ratios in U2OS cells using irradiation. b, (left) Representative super-resolution (SR) images showing EdU and γH2AX channels (left panels), overlay of γH2AX foci on respective channels (middle panels, green), and overlay of random mock foci on respective channels (right panels, teal) in irradiated U2OS cells. scale bar is 10 microns. (right) γH2AX intensity in irradiation-induced γH2AX foci and randomly sampled mock foci indicating the specificity of foci definitions used in the analysis strategy. p-value is from a two-sided Mann-Whitney U-test, n = 210 cells. c, (left) Representative SR images showing early, mid, and late S phase cells. Percentage of cells from asynchronous population belonging to each of these classes are shown above the images. Refer to methods for details of the S-phase segmentation. Scale bar=10 microns. (right) Violin plots for EdU intensity ratio in comparison to random foci in different S phase classes. Median value of the EdU intensity ratio is shown on top of each violin plot. A minimum of 60 cells are analyzed in each category. Two-sided Mann-Whitney U-test is used for p-value calculations. d, Schematics of the experimental workflow used for validation of the EdU intensity ratio analysis strategy. CPT stands for camptothecin (100 nM). e, (left) Cartoon depicting the process leading to increase in A/B ratio after CPT treatment. CPT introduces single-strand DNA breaks (SSBs) which are γH2AX and EdU negative. Upon replication, SSBs are converted to DSBs along with EdU incorporation, in a time-dependent manner. (right) Bar graphs showing EdU intensity ratios from CPT experiment. EdU intensity ratios using irradiation (IR) are used as a control. Mean and standard deviation are shown. The mean value of the EdU intensity ratio from three replicates is displayed against each column. A minimum of 200 cells are analyzed in each replicate. f, Schematic of the experimental strategy used for EdU/γH2AX staining and EdU intensity ratio analysis in AsiSI U2OS cells. g, (left) Representative immunofluorescence images of AsiSI-U2OS cells at 3.5 h post dThy release, showing EdU staining in all cells and proficient γH2AX induction after AsiSI activation using 4OHT (middle panel with enlarged inset view on the right panel). Scale bar is 10 microns. (right) Number of γH2AX foci in AsiSI-U2OS cells. The black bar indicates the median. A minimum of 100 cells were analyzed in each group. p-value is calculated using a two-tailed, Mann-Whitney U-test.
Extended Data Fig. 2 Dynamics of DNA replication upon DSB induction.
a, Western blots depicting changes to various checkpoint phospho-modifications after DSB induction in AsiSI-U2OS and U2OS cells. The 50 Gy dose is not included in western blotting experiments as it is out of the dynamic range for most of the proteins tested. Representative blots from three independent experiments are shown. Blots for internal loading controls are performed along with each individual western. Representative loading control (H3) is shown. b, Representative EdU intensity plots of asynchronous AsiSI-U2OS cells with and without DSB induction (left) and asynchronous U2OS cells after various doses of irradiation (right). EdU was incubated for 30 min post 4OHT treatment and 20 min post irradiation for AsiSI-U2OS and U2OS, respectively. Representative of two independent experiments. c, Cell cycle profiles of U2OS cells from 0 h and 3.5 h post dThy release with or without DSB induction via AsiSI activation using 4OHT treatment. d, (top) Representative image of a DNA fiber labeled with DNA (blue), IdU (green), and CldU (red) taken from dThy released, AsiSI-induced U2OS cells. IdU treatment is initiated at 3 h post dThy release. (bottom) Graphs showing global replication fork speed in AsiSI-U2OS cells and U2OS cells after DSB induction using AsiSI activation and irradiation, respectively. A high dose of 50 Gy is used as a positive control. The blue bar indicates the median. A minimum of 120 fibers are analyzed in each condition. p-values are based on a two-tailed, Mann-Whitney U-test.
Extended Data Fig. 3 Analysis of replication timing and S-phase progression in AsiSI-U2OS cells.
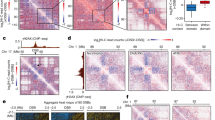
a, Schematics of the experimental flow to assay replication timing in U2OS cells. b, (top) Integrative Genomics Viewer (IGV) screenshot of the replication timing in U2OS cells and its subcategorization into early, early-mid, mid-late, and late zones. Early and late regions are color-coded in blue and red, respectively. (bottom) Table showing the classification of all 80 AsiSI sites according to their replication timing. c, Kinetics of S-phase progression of AsiSI-U2OS cells released from double-Thymidine (dThy)-mediated G1/S cell cycle arrest. Cells are released with or without DSB induction by the 4OHT-controlled AsiSI enzyme. The shaded region indicates the time frame for BrdU and EdU incubation for sequencing and imaging, respectively. Each time point is an average of 3 biological repeats and error bars indicate standard deviation. d, (top) Aggregate plots for BrdU and γH2AX signals with or without DSB induction centered on AsiSI sites. (bottom) Aggregate plots and heatmaps of γH2AX and BrdU showing log2 fold change across all 80 AsiSI sites upon DSB induction. AsiSI-accessible sites in AsiSI-U2OS cells are taken from11.
Extended Data Fig. 4 D compartment and local inhibition of DNA replication.
a, IGV screenshot of the BrdU-seq and γH2AX ChIP-seq sequencing tracks with or without DSB induction via AsiSI activation by 4-Hydroxytamoxifen (4OHT), relative to D compartment. Arrowhead shows one of the AsiSI sites on chromosome 17 (chr17:5,390,222). 2D coordinates of both AsiSI and non-AsiSI D compartments are visible. D compartment coordinates are obtained from14. D comp stands for D compartment. b, Violin plots showing relative log2 fold changes of γH2AX ChIP-seq signal in D compartments with or without AsiSI sites. Shuffle controls are derived from non-D compartment regions of chr1 (p-arm), chr17 and chrX. p-values are calculated using two-sided Wilcoxon rank sum exact test.
Extended Data Fig. 5 Role of cohesin in DNA replication following DSB induction.
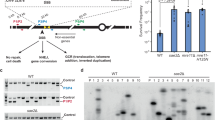
a, Representative flow cytometry plot depicting depletion of mClover-RAD21 after RAD21 depletion via the addition of IAA (auxin). b,c, Area of the γH2AX foci (b) and the number of γH2AX per cell (c) for the control and IAA-treated HCT116 cells. Error bars indicate standard error from 3 biological replicates. A minimum of 61 cells are analyzed in each replicate. p-value is based on Student’s two-tailed t-test. d-f Characterization and analysis of replication in LIG4−/− AID-RAD21 AsiSI-HCT116 cells. d, Cell cycle analysis of asynchronous LIG4−/− AID-RAD21 AsiSI-HCT116 cells after DSB induction, RAD21 depletion, or both. Representative EdU-DNA 2D flow cytometry plots are on the left and a graph depicting quantification of various cell cycle phases is on the right. e, Workflow of the dThy experiment and flow cytometry histogram plot depicting depletion of mClover-RAD21 after the addition of IAA. Analyses were performed 1.5 h after release from dThy. f, Cell cycle analysis of dThy-released cells with or without RAD21 depletion and DSB induction. Representative flow cytometry plots are on the left and a graph showing the quantification of early-S phase on the right. Results of two independent repeats. g, BrdU heatmaps showing log2 fold change after DSB induction in control and RAD21-depleted cells. Enzyme-accessible AsiSI sites in HCT116 cells defined using MRE11 ChIP-seq. h, (top) Violin plots of EdU intensity ratios after siRNA-mediated knockdown of NIPBL, WAPL and RAD21. Median value of the ratio is shown on top of each violin. Irradiation dose is 2 Gy. p-values are calculated using two-tailed, Mann-Whitney U-test. A minimum of 50 cells are analyzed from 2 independent repeats using 3D SR microscopy. (bottom) Western blots showing knockdown efficiency of the siRNAs used. i, A cartoon representing the model of DNA replication in DSB chromatin. Cohesin-mediated loop extrusion is critical for DSB containment and thereby keeps replication away from DSB chromatin. When RAD21 is depleted, cohesin is no longer functional, and loop domains are dissolved. DSB chromatin is no longer “insulated” and is vulnerable to incoming DNA replication.
Extended Data Fig. 6 Identification of MRD factors.
a, Workflow of the high-throughput siRNA screening pipeline used to identify MRD factors. b, Dot plots depicting the Z-score derived from A/B ratio of EdU intensity after the primary screen. Genes involved in HR, NHEJ, and MMEJ are color-coded and highlighted, in addition to MRD factors confirmed using a secondary screen (blue color). The Z-score rank is based on two independent repeats of the screen. c, (top) EdU ratio (A/B) analysis using the negative hits with representative genes from HR (BRCA1), NHEJ (LIG4 & DNA-PK), and MMEJ (LIG3) repair pathways using 3D SR microscopy. Median value of the ratio is shown on top of each violin. DNA-PK is inhibited using the inhibitor KU-0060648 (10 µM) for 1 h prior to irradiation and EdU staining. siRNAs are used for the rest of the genes. (bottom) Western blots confirming the gene knockdown using an independent set of siRNAs against selected negative hits from the primary screen. d, (top) EdU ratio (A/B) analysis of the primary hits using 3D SR microscopy. Median value of the ratio is shown using continuous black line in the violin plot and is written on top of each violin. (bottom) Western blots confirming the gene knockdown using an independent set of siRNAs against selected hits from the primary screen. Red arrowheads indicate the correct band corresponding to the protein of interest if multiple bands are present. T-LESS denotes TIMELESS. H3 internal control is run separately. e, List of validated MRD factors after the primary and secondary analyses.
Extended Data Fig. 7 Validation of MRD factors.
a, (top) Experimental strategy for assaying the effect of transcription on EdU intensity ratios. (bottom) Representative images showing cumulative 5-ethynyluridine (EU) staining in control and actinomycin D-treated U2OS cells. b, Dot plots displaying EU (left) and EdU (right) intensity after actinomycin D treatment. c, Violin plot showing A/B ratio in U2OS cells after actinomycin D treatment. Median value of the ratio is shown on top of each violin. 66 and 77 cells are analyzed from 2 independent repeats using 3D SR microscopy for control and actinomycin D treatment. Irradiation dose is 2 Gy. p-values in b and c are calculated using two-tailed, Mann-Whitney U-test. d, (top) Flowchart of the experimental strategy to test the role of 53BP1 as an MRD factor using BrdU-seq. (bottom) IGV screenshot of the BrdU sequencing from dThy-released AsiSI-U2OS cells. AsiSI site chr1:89,458,598 is shown using the arrowhead. The boxed region indicates an AsiSI-containing TAD. e, Violin plots depicting log2 fold change in BrdU upon DSB induction in DSB-TADs and random-TADs in si-control and si-53BP1 treated AsiSI-U2OS cells. Two independent experiments are used. p-value is from two-sided paired Wilcoxon signed rank test. si-RIF1 samples are not used for quantification due to total loss of replication timing after siRNA treatment. f, g, Assessment of DSB-induced RIF1 foci in the absence of 53BP1. f, Representative confocal images of irradiation-induced RIF1 foci in control and si-TP53BP1 treated cells. si-RIF1 is used as positive control. Irradiation is 2 Gy. Inset is enlarged on the bottom panel with scale bar of 10 microns. g, Dot plots showing the distribution of number of RIF1 foci/cell in control and siRNA treatments. A minimum of 250 cells are analyzed in each condition. p-value is from two-sided Wilcoxon unpaired test.
Extended Data Fig. 8 Mapping DSB-proximal suppression of replication origins.
a, IGV screenshots of the NS-seq track, ChIP-seq tracks for γH2AX, TIPIN and TIMELESS with or without DSB induction (as described in Fig. 3a). D compartment (D comp) coordinates are shown for screenshots from chr1 and 17 (screenshots on top). DSB-TAD coordinates are shown for all four screenshots. DSB location is marked using an arrowhead in each screenshot. Highlighted regions in yellow are shown as zoomed. Identical scaling is used between – and + DSB samples of all the sequencing tracks. b, Heatmaps showing the binding of pSIRT1 (T530), MCM2-pS139 and MCM4 on origins from synchronously replicating AsiSI-U2OS cells. All heatmaps are centered and arranged based on NS-seq peaks with >10 RPKM in -DSB sample. The corresponding aggregate plot is on top of each heatmap. c, Graph showing total nuclear intensity of γH2AX in U2OS cells at 20- and 40-min post 2 Gy irradiation. n.s. is not significant based on two-sided Mann-Whitney U-test.
Extended Data Fig. 9 Control of replication origin by MRD factor WEE1 kinase.
a, Dot plot depicting the Z-score based on EdU intensity ratios using a kinome library siRNA screen. The screening strategy is like the one described in Extended Data Fig. 6a. Top three hits are marked. The Z-score rank is based on two independent repeats. CHEK1 was not selected for downstream assays due to the pan-nuclear staining pattern of γH2AX after knockdown (data not shown). b, (top) Western blot for PKMYT1 and WEE1 proteins after 72 h of siRNA knockdown in U2OS cells. Different sets of siRNAs are used compared to the primary screen. H3 is used as a loading control. (bottom) Violin plots showing EdU ratio (A/B) elucidated from SR microscopy after siRNA knockdown. Mean EdU intensity ratios are shown on top of each violin. A minimum of 100 cells were analyzed. Representative of two biological replicates. c, Representative EdU cell cycle plots of WEE1i-treated U2OS cells. DMSO-treated cells are used as control. Hyper-replication fraction is marked using a dotted line with the percentage and standard deviation. The numbers are from three independent experiments. d,e, DNA combing experiments in DMSO or WEE1i-treated (1 h) U2OS cells. d, Representative images of DNA fibers labeled with DNA (blue), IdU (green), and CldU (red) taken from DMSO and WEE1i-treated cells. IdU treatment is initiated at 1 h post-DMSO or WEE1i treatment. A minimum of 78 fibers are analyzed in each condition. Scale bar is 50 microns. e, Graphs showing quantitative analyses of the DNA combing assay comparing DMSO or WEE1i-treated U2OS cells. Replication fork speed (left), inter-origin distance (center) and number of origins per field (right) are shown. p-values from two-sided Wilcoxon signed rank. p = 2.2e-16 was displayed when p-value was equal or smaller than 2.2e-16. f, Violin plots depicting log2 fold change in BrdU after WEE1i treatment across the genome classified according to replication timing. Control AsiSI-U2OS cells are shown. Replication timing regions are taken from Extended Data Fig. 2b. Median values are provided on top of the violins. All p-values from two-sided Wilcoxon signed rank test are <2.2e-16. Representative of 2 independent experiments. g,h, Analysis of genomic distribution of γH2AX following WEE1i. Violin plots depicting log2 fold change in γH2AX after WEE1i treatment across the genome, classified according to (g) replication timing domains, and (h) euchromatin (H3K4me3) vs heterochromatin (H3K27me3) domains. Histone marks are from74. Median values are shown on top of the violins. p-values from Wilcoxon signed rank test between early vs. late and H3K4me3 v.s H3K27me3 are <2.2e-16. Results are averaged from 2 independent experiments. i, γH2AX and EdU cell cycle flow cytometry plots of U2OS cells treated with DMSO or WEE1i for 1 h. γH2AX-positive fraction is gated and is shown in red color with the average percentage and standard deviation. Hyper-replication fraction is shown using a dotted line. This plot is representative of three independent experiments.
Extended Data Fig. 10
a, Representative western blots for TIMELESS and TIPIN in chromatin extracts from U2OS cells. DMSO or WEE1i (10 µM) treatment is for 1 h before irradiation (2 Gy). H3 is used as loading control. Loading control-normalized, relative fold change is shown below the blots. Representative of three biological replicates. b, Western blots for TIPIN in chromatin extracts from hTERT RPE-1, AG04446 and hTERT IMR90 cells. DMSO or WEE1i (10 µM) treatment is for 1 h before irradiation (2 Gy). H3 is used as loading control. Relative fold change is shown as numbers below the blot. Representative of two biological replicates. c, Western blots for TIMELESS and TIPIN in chromatin extracts of U2OS cells with combinations of CDK1 (RO-3306), CDK2 (PF-06873600) and WEE1 inhibitors. Inhibitor treatment time is 1 h. CDK1, CDK2 and WEE1 inhibitors are used at a concentration of 10, 5 and 10 µM, respectively. Normalized relative fold change is shown below the blots. Representative of two biological replicates. d, Expression of TIMELESS-TIPIN complex correlates with replication stress burden in tumors. Scatter plots for testing the correlation between Repstress score53 and the expression of TIMELESS and TIPIN in small-cell lung cancer (SCLC) patients and cancer cell line encyclopedia (CCLE) data set. These plots are generated using Tumorminer and Cellminer for patient and CCLE data sets, respectively75.
Supplementary information
Supplementary Figures (download PDF )
Supplementary Figs. 1 and 2
Supplementary Table 1 (download XLSX )
Results of the RNAi screen using human DNA damage response and replication (Extended Data Fig. 6b) and human kinome (Extended Data Fig. 9a) siRNA libraries. All values are average of two biological repeats.
Rights and permissions
About this article
Cite this article
Sebastian, R., Sun, E.G., Fedkenheuer, M. et al. Mechanism for local attenuation of DNA replication at double-strand breaks. Nature 639, 1084–1092 (2025). https://doi.org/10.1038/s41586-024-08557-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08557-9