Abstract
Neuroimmune interactions—signals transmitted between immune and brain cells—regulate many aspects of tissue physiology1, including responses to psychological stress2,3,4,5, which can predispose individuals to develop neuropsychiatric diseases6,7,8,9. Still, the interactions between haematopoietic and brain-resident cells that influence complex behaviours are poorly understood. Here, we use a combination of genomic and behavioural screens to show that astrocytes in the amygdala limit stress-induced fear behaviour through epidermal growth factor receptor (EGFR). Mechanistically, EGFR expression in amygdala astrocytes inhibits a stress-induced, pro-inflammatory signal-transduction cascade that facilitates neuron–glial crosstalk and stress-induced fear behaviour through the orphan nuclear receptor NR2F2 in amygdala neurons. In turn, decreased EGFR signalling and fear behaviour are associated with the recruitment of meningeal monocytes during chronic stress. This set of neuroimmune interactions is therapeutically targetable through the administration of psychedelic compounds, which reversed the accumulation of monocytes in the brain meninges along with fear behaviour. Together with validation in clinical samples, these data suggest that psychedelics can be used to target neuroimmune interactions relevant to neuropsychiatric disorders and potentially other inflammatory diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data in this study have been deposited into the GEO under the super series accession number GSE262989. Bulk RNA-seq datasets of amygdala neurons were obtained from the GEO with accession numbers GSE162417 (ref. 93), GSE66345 (ref. 94), GSE130268 (ref. 95), GSE138522 (ref. 96), GSE183092 (ref. 97) and GSE151798 (ref. 98). A mouse atlas of brain structures27 was accessed at https://cells.ucsc.edu/?ds=mouse-nervous-system, and .loom files for each brain region included were downloaded with annotations from a previous study27. An atlas of amygdala cell types36 was accessed from FigShare (https://doi.org/10.6084/m9.figshare.20412573)134. Gene-expression data of purified human immune cell populations were accessed using the ImmGen Skyline RNA-seq data browser132 (https://www.immgen.org/Databrowser19/DatabrowserPage.html). Source data are provided with this paper.
Code availability
No new code was developed for this study.
References
Wheeler, M. A. & Quintana, F. J. The neuroimmune connectome in health and disease. Nature 638, 333–342 (2025).
Poller, W. C. et al. Brain motor and fear circuits regulate leukocytes during acute stress. Nature 607, 578–584 (2022).
Chan, K. L., Poller, W. C., Swirski, F. K. & Russo, S. J. Central regulation of stress-evoked peripheral immune responses. Nat. Rev. Neurosci. https://doi.org/10.1038/s41583-023-00729-2 (2023).
Cathomas, F. et al. Beyond the neuron: role of non-neuronal cells in stress disorders. Neuron 110, 1116–1138 (2022).
Fan, K.-Q. et al. Stress-induced metabolic disorder in peripheral CD4+ T cells leads to anxiety-like behavior. Cell 179, 864–879 (2019).
Cathomas, F. et al. Circulating myeloid-derived MMP8 in stress susceptibility and depression. Nature 626, 1108–1115 (2024).
Pape, K., Tamouza, R., Leboyer, M. & Zipp, F. Immunoneuropsychiatry—novel perspectives on brain disorders. Nat. Rev. Neurol. 15, 317–328 (2019).
Hodes, G. E., Kana, V., Menard, C., Merad, M. & Russo, S. J. Neuroimmune mechanisms of depression. Nat. Neurosci. 18, 1386–1393 (2015).
Wohleb, E. S., Franklin, T., Iwata, M. & Duman, R. S. Integrating neuroimmune systems in the neurobiology of depression. Nat. Rev. Neurosci. 17, 497–511 (2016).
Qing, H. et al. Origin and function of stress-induced IL-6 in murine models. Cell 182, 372–387 (2020).
Lee, H.-G., Wheeler, M. A. & Quintana, F. J. Function and therapeutic value of astrocytes in neurological diseases. Nat. Rev. Drug Discov. 21, 339–358 (2022).
Byun, Y. G. et al. Stress induces behavioral abnormalities by increasing expression of phagocytic receptor, MERTK, in astrocytes to promote synapse phagocytosis. Immunity 56, 2105–2120 (2023).
Leng, L. et al. Menin deficiency leads to depressive-like behaviors in mice by modulating astrocyte-mediated neuroinflammation. Neuron 100, 551–563 (2018).
Guayasamin, M. et al. Early-life stress induces persistent astrocyte dysfunction associated with fear generalisation. eLife https://doi.org/10.7554/eLife.99988.2 (2024).
Dudek, K. A. et al. Astrocytic cannabinoid receptor 1 promotes resilience by dampening stress-induced blood–brain barrier alterations. Nat. Neurosci. https://doi.org/10.1038/s41593-025-01891-9 (2025).
Ollivier, M. et al. Crym-positive striatal astrocytes gate perseverative behaviour. Nature 627, 358–366 (2024).
Yu, X. et al. Reducing astrocyte calcium signaling in vivo alters striatal microcircuits and causes repetitive behavior. Neuron 99, 1170–1187 (2018).
Suthard, R. L. et al. Basolateral amygdala astrocytes are engaged by the acquisition and expression of a contextual fear memory. J. Neurosci. 43, 4997–5013 (2023).
Martin-Fernandez, M. et al. Synapse-specific astrocyte gating of amygdala-related behavior. Nat. Neurosci. 20, 1540–1548 (2017).
Cui, Y. et al. Astroglial Kir4.1 in the lateral habenula drives neuronal bursts in depression. Nature 554, 323–327 (2018).
Wheeler, M. A. et al. Droplet-based forward genetic screening of astrocyte–microglia cross-talk. Science 379, 1023–1030 (2023).
Clark, I. C. et al. Identification of astrocyte regulators by nucleic acid cytometry. Nature 614, 326–333 (2023).
Wheeler, M. A. et al. Environmental control of astrocyte pathogenic activities in CNS inflammation. Cell 176, 581–596 (2019).
Wheeler, M. A. et al. MAFG-driven astrocytes promote CNS inflammation. Nature 578, 593–599 (2020).
Sanmarco, L. M. et al. Gut-licensed IFNγ+ NK cells drive LAMP1+TRAIL+ anti-inflammatory astrocytes. Nature 590, 473–479 (2021).
Maddox, S. A., Hartmann, J., Ross, R. A. & Ressler, K. J. Deconstructing the gestalt: mechanisms of fear, threat, and trauma memory encoding. Neuron 102, 60–74 (2019).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
LeDoux, J. The amygdala. Curr. Biol. 17, R868–R874 (2007).
Wohleb, E. S., Powell, N. D., Godbout, J. P. & Sheridan, J. F. Stress-induced recruitment of bone marrow-derived monocytes to the brain promotes anxiety-like behavior. J. Neurosci. 33, 13820–13833 (2013).
Zheng, Z.-H. et al. Neuroinflammation induces anxiety- and depressive-like behavior by modulating neuronal plasticity in the basolateral amygdala. Brain Behav. Immun. 91, 505–518 (2021).
Linnerbauer, M. et al. The astrocyte-produced growth factor HB-EGF limits autoimmune CNS pathology. Nat. Immunol. 25, 432–447 (2024).
Sun, W. et al. Spatial transcriptomics reveal neuron–astrocyte synergy in long-term memory. Nature 627, 374–381 (2024).
Luo, F. et al. Modulation of proteoglycan receptor PTPσ enhances MMP-2 activity to promote recovery from multiple sclerosis. Nat. Commun. 9, 4126 (2018).
Tonks, N. K. Protein tyrosine phosphatases: from genes, to function, to disease. Nat. Rev. Mol. Cell Biol. 7, 833–846 (2006).
Won, S. Y., Lee, P. & Kim, H. M. Synaptic organizer: slitrks and type IIa receptor protein tyrosine phosphatases. Curr. Opin. Struct. Biol. 54, 95–103 (2019).
Hochgerner, H. et al. Neuronal types in the mouse amygdala and their transcriptional response to fear conditioning. Nat. Neurosci. https://doi.org/10.1038/s41593-023-01469-3 (2023).
Santiago, A. C. & Shammah-Lagnado, S. J. Efferent connections of the nucleus of the lateral olfactory tract in the rat. J. Comp. Neurol. 471, 314–332 (2004).
Menard, C. et al. Social stress induces neurovascular pathology promoting depression. Nat. Neurosci. 20, 1752–1760 (2017).
Clark, S. M., Soroka, J. A., Song, C., Li, X. & Tonelli, L. H. CD4+ T cells confer anxiolytic and antidepressant-like effects, but enhance fear memory processes in Rag2–/– mice. Stress 19, 303–311 (2016).
Rustenhoven, J. & Kipnis, J. Brain borders at the central stage of neuroimmunology. Nature 612, 417–429 (2022).
Marin-Rodero, M. et al. The meninges host a distinct compartment of regulatory T cells that preserves brain homeostasis. Sci. Immunol. 10, eadu2910 (2025).
Barron, J. J. et al. Group 2 innate lymphoid cells promote inhibitory synapse development and social behavior. Science 386, eadi1025 (2024).
Dai, W. et al. A functional role of meningeal lymphatics in sex difference of stress susceptibility in mice. Nat. Commun. 13, 4825 (2022).
McKim, D. B. et al. Sympathetic release of splenic monocytes promotes recurring anxiety following repeated social defeat. Biol. Psychiatry 79, 803–813 (2015).
Kwan, A. C., Olson, D. E., Preller, K. H. & Roth, B. L. The neural basis of psychedelic action. Nat. Neurosci. 25, 1407–1419 (2022).
Green, A. R., Mechan, A. O., Elliott, J. M., O’Shea, E. & Colado, M. I. The pharmacology and clinical pharmacology of 3,4-methylenedioxymethamphetamine (MDMA, “ecstasy”). Pharmacol. Rev. 55, 463–508 (2003).
Shao, L.-X. et al. Psilocybin induces rapid and persistent growth of dendritic spines in frontal cortex in vivo. Neuron 109, 2535–2544 (2021).
Goodwin, G. M. et al. Single-dose psilocybin for a treatment-resistant episode of major depression. N. Engl. J. Med. 387, 1637–1648 (2022).
Mitchell, J. M. et al. MDMA-assisted therapy for moderate to severe PTSD: a randomized, placebo-controlled phase 3 trial. Nat. Med. https://doi.org/10.1038/s41591-023-02565-4 (2023).
Nardou, R. et al. Psychedelics reopen the social reward learning critical period. Nature 618, 790–798 (2023).
Nardou, R. et al. Oxytocin-dependent reopening of a social reward learning critical period with MDMA. Nature 569, 116–120 (2019).
Flanagan, T. W. & Nichols, C. D. Psychedelics as anti-inflammatory agents. Int. Rev. Psychiatry 30, 363–375 (2018).
Lehmann, M. L., Poffenberger, C. N., Elkahloun, A. G. & Herkenham, M. Analysis of cerebrovascular dysfunction caused by chronic social defeat in mice. Brain Behav. Immun. 88, 735–747 (2020).
Daskalakis, N. P. et al. Systems biology dissection of PTSD and MDD across brain regions, cell types, and blood. Science 384, eadh3707 (2024).
Cugurra, A. et al. Skull and vertebral bone marrow are myeloid cell reservoirs for the meninges and CNS parenchyma. Science 373, eabf7844 (2021).
Singh, V. et al. Microbiota dysbiosis controls the neuroinflammatory response after stroke. J. Neurosci. 36, 7428–7440 (2016).
Endo, F. et al. Molecular basis of astrocyte diversity and morphology across the CNS in health and disease. Science 378, eadc9020 (2022).
Kyzar, E. J., Nichols, C. D., Gainetdinov, R. R., Nichols, D. E. & Kalueff, A. V. Psychedelic drugs in biomedicine. Trends Pharmacol. Sci. 38, 992–1005 (2017).
Chiu, Y.-T. et al. A suite of engineered mice for interrogating psychedelic drug actions. Preprint at bioRxiv https://doi.org/10.1101/2023.09.25.559347 (2023).
Muir, J. et al. Isolation of psychedelic-responsive neurons underlying anxiolytic behavioral states. Science 386, 802–810 (2024).
Shultz, L. D. et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2Rγnull mice engrafted with mobilized human hemopoietic stem cells. J. Immunol. 174, 6477–6489 (2005).
Coughlan, A. M. et al. Myeloid engraftment in humanized mice: impact of granulocyte-colony stimulating factor treatment and transgenic mouse strain. Stem Cells Dev. 25, 530–541 (2016).
Srinivasan, R. et al. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes in situ and in vivo. Neuron 92, 1181–1195 (2016).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Buch, T. et al. A Cre-inducible diphtheria toxin receptor mediates cell lineage ablation after toxin administration. Nat. Methods 2, 419–426 (2005).
Xu, Z. et al. Efficient strategies for microglia replacement in the central nervous system. Cell Rep. 32, 108041 (2020).
Tomura, M. et al. Monitoring cellular movement in vivo with photoconvertible fluorescence protein “Kaede” transgenic mice. Proc. Natl Acad. Sci. USA 105, 10871–10876 (2008).
Chu, C. et al. The microbiota regulate neuronal function and fear extinction learning. Nature 574, 543–548 (2019).
Shen, Y. et al. CCR5 closes the temporal window for memory linking. Nature 606, 146–152 (2022).
Woodburn, S. C., Levitt, C. M., Koester, A. M. & Kwan, A. C. Psilocybin facilitates fear extinction: importance of dose, context, and serotonin receptors. ACS Chem. Neurosci. 15, 3034–3043 (2024).
Clark, I. C. et al. Barcoded viral tracing of single-cell interactions in central nervous system inflammation. Science 372, eabf1230 (2021).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Hunker, A. C. et al. Conditional single vector CRISPR/SaCas9 viruses for efficient mutagenesis in the adult mouse nervous system. Cell Rep. 30, 4303–4316 (2020).
Lee, J.-H. et al. Astrocytes phagocytose adult hippocampal synapses for circuit homeostasis. Nature 590, 612–617 (2020).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Nath, T. et al. Using DeepLabCut for 3D markerless pose estimation across species and behaviors. Nat. Protoc. 14, 2152–2176 (2019).
Sturman, O. et al. Deep learning-based behavioral analysis reaches human accuracy and is capable of outperforming commercial solutions. Neuropsychopharmacology 45, 1942–1952 (2020).
Gradinaru, V. et al. Molecular and cellular approaches for diversifying and extending optogenetics. Cell 141, 154–165 (2010).
He, D. et al. Disruption of the IL-33–ST2–AKT signaling axis impairs neurodevelopment by inhibiting microglial metabolic adaptation and phagocytic function. Immunity 55, 159–173 (2022).
Leites, E. P. & Morais, V. A. Protocol for the isolation and culture of microglia, astrocytes, and neurons from the same mouse brain. STAR Protoc. 5, 102804 (2024).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Laricchiuta, D. et al. Optogenetic stimulation of prelimbic pyramidal neurons maintains fear memories and modulates amygdala pyramidal neuron transcriptome. Int. J. Mol. Sci. 22, 810 (2021).
Namburi, P. et al. A circuit mechanism for differentiating positive and negative associations. Nature 520, 675–678 (2015).
Samineni, V. K. et al. Cellular, circuit and transcriptional framework for modulation of itch in the central amygdala. eLife 10, e68130 (2021).
Levitan, D. et al. Deletion of Stk11 and Fos in mouse BLA projection neurons alters intrinsic excitability and impairs formation of long-term aversive memory. eLife 9, e61036 (2020).
Knoedler, J. R. et al. A functional cellular framework for sex and estrous cycle-dependent gene expression and behavior. Cell 185, 654–671 (2022).
Yamaguchi, T. et al. Posterior amygdala regulates sexual and aggressive behaviors in male mice. Nat. Neurosci. 23, 1111–1124 (2020).
Golden, S. A. et al. Basal forebrain projections to the lateral habenula modulate aggression reward. Nature 534, 688–692 (2016).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Chen, A. et al. Single-cell spatial transcriptome reveals cell-type organization in the macaque cortex. Cell 186, 3726–3743 (2023).
van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Waclaw, R. R., Ehrman, L. A., Pierani, A. & Campbell, K. Developmental origin of the neuronal subtypes that comprise the amygdalar fear circuit in the mouse. J. Neurosci. 30, 6944–6953 (2010).
Kuerbitz, J. et al. Loss of intercalated cells (ITCs) in the mouse amygdala of Tshz1 mutants correlates with fear, depression, and social interaction phenotypes. J. Neurosci. 38, 1160–1177 (2018).
Garcia-Calero, E., Martínez-de-la-Torre, M. & Puelles, L. A radial histogenetic model of the mouse pallial amygdala. Brain Struct. Funct. 225, 1921–1956 (2020).
Glendining, K. A., Fisher, L. C. & Jasoni, C. L. Maternal high fat diet alters offspring epigenetic regulators, amygdala glutamatergic profile and anxiety. Psychoneuroendocrinology 96, 132–141 (2018).
Nery, S., Fishell, G. & Corbin, J. G. The caudal ganglionic eminence is a source of distinct cortical and subcortical cell populations. Nat. Neurosci. 5, 1279–1287 (2002).
Huang, W.-C., Chen, Y. & Page, D. T. Hyperconnectivity of prefrontal cortex to amygdala projections in a mouse model of macrocephaly/autism syndrome. Nat. Commun. 7, 13421 (2016).
Stenman, J., Yu, R. T., Evans, R. M. & Campbell, K. Tlx and Pax6 co-operate genetically to establish the pallio-subpallial boundary in the embryonic mouse telencephalon. Development 130, 1113–1122 (2003).
Tosches, M. A. et al. Evolution of pallium, hippocampus, and cortical cell types revealed by single-cell transcriptomics in reptiles. Science 360, 881–888 (2018).
Boyle, M. P. et al. Cell-type-specific consequences of Reelin deficiency in the mouse neocortex, hippocampus, and amygdala. J. Comp. Neurol. 519, 2061–2089 (2011).
Ressler, K. J. et al. Post-traumatic stress disorder: clinical and translational neuroscience from cells to circuits. Nat. Rev. Neurol. 18, 273–288 (2022).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Pinho-Ribeiro, F. A. et al. Bacteria hijack a meningeal neuroimmune axis to facilitate brain invasion. Nature 615, 472–481 (2023).
Wilk, C. M. et al. Circulating senescent myeloid cells infiltrate the brain and cause neurodegeneration in histiocytic disorders. Immunity 56, 2790–2802 (2023).
Barnes, N. M. et al. International Union of Basic and Clinical Pharmacology. CX. Classification of receptors for 5-hydroxytryptamine; pharmacology and function. Pharmacol. Rev. 73, 310–520 (2021).
Gumpper, R. H. & Roth, B. L. SnapShot: psychedelics and serotonin receptor signaling. Cell 186, 232–232 (2023).
Baumann, M. H. et al. Effects of dose and route of administration on pharmacokinetics of (±)-3,4-methylenedioxymethamphetamine in the rat. Drug Metab. Dispos. 37, 2163–2170 (2009).
Lyon, R. A., Glennon, R. A. & Titeler, M. 3,4-Methylenedioxymethamphetamine (MDMA): stereoselective interactions at brain 5-HT1 and 5-HT2 receptors. Psychopharmacology 88, 525–526 (1986).
Nash, J., Roth, B., Brodkin, J., Nichols, D. & Gudelsky, G. Effect of the R(−) and S(+) isomers of MDA and MDMA on phosphotidyl inositol turnover in cultured cells expressing 5-HT2A or 5-HT2C receptors. Neurosci. Lett. 177, 111–115 (1994).
Setola, V. et al. 3,4-methylenedioxymethamphetamine (MDMA, “Ecstasy”) induces fenfluramine-like proliferative actions on human cardiac valvular interstitial cells in vitro. Mol. Pharmacol. 63, 1223–1229 (2003).
Doly, S. et al. Serotonin 5-HT2B receptors are required for 3,4-methylenedioxymethamphetamine-induced hyperlocomotion and 5-HT release in vivo and in vitro. J. Neurosci. 28, 2933–2940 (2008).
Moliner, R. et al. Psychedelics promote plasticity by directly binding to BDNF receptor TrkB. Nat. Neurosci. https://doi.org/10.1038/s41593-023-01316-5 (2023).
McClure-Begley, T. D. & Roth, B. L. The promises and perils of psychedelic pharmacology for psychiatry. Nat. Rev. Drug Discov. 21, 463–473 (2022).
Cameron, L. P. et al. Beyond the 5-HT2A receptor: classic and nonclassic targets in psychedelic drug action. J. Neurosci. 43, 7472–7482 (2023).
Nichols, D. E., Nichols, C. D. & Hendricks, P. S. Proposed consensus statement on defining psychedelic drugs. Psychedelic Med. 1, 12–13 (2023).
Schneider, K. M. et al. The enteric nervous system relays psychological stress to intestinal inflammation. Cell 186, 2823–2838 (2023).
Wheeler, M. A. et al. TNF-α/TNFR1 signaling is required for the development and function of primary nociceptors. Neuron 82, 587–602 (2014).
Glebova, N. O. & Ginty, D. D. Heterogeneous requirement of NGF for sympathetic target innervation in vivo. J. Neurosci. 24, 743–751 (2004).
Heng, T. S. P., Painter, M. W. & Immunological Genome Project Consortium. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Sanmarco, L. M. et al. Lactate limits CNS autoimmunity by stabilizing HIF-1α in dendritic cells. Nature 620, 881–889 (2023).
Zeisel, A. Amy_FC_allcells_with_metadata_31-Jul-2022.txt. Figshare https://doi.org/10.6084/m9.figshare.20412573 (2022).
Acknowledgements
M.A.W. acknowledges support from the NIMH, NIDA and NINDS (R01MH130458, R01MH132632, R01DA061199 and R00NS114111), the Boston Claude D. Pepper Older Americans Independence Center and the Brigham Research Institute. F.J.Q. acknowledges support from grants NS102807, ES02530, ES029136 and AI126880 from the NIH; RG4111A1 and JF2161-A-5 from the NMSS; RSG-14-198-01-LIB from the American Cancer Society; and PA-1604-08459 from the International Progressive MS Alliance. V.K.K. acknowledges support from NIH grants R01AG080992 and R01AI139536. I.M.C. acknowledges support from grants R01AI168005 and R01DK127257 from the NIH; the Chan–Zuckerberg Initiative; the Jackson–Wijaya Fund; and the Burroughs Wellcome Fund. E.N.C. acknowledges a Research Supplement to Promote Diversity in Health-Related Research from the NINDS (R00NS114111-S1) and the Brigham Research Institute. C.M.P. was supported by the National Multiple Sclerosis Society (FG-2307-42209) and the Mayer Foundation. M.K. was supported by a Leopoldina Research Fellow Grant of the German Academy of Sciences. S.S.D. was supported by a Banting Postdoctoral Fellowship from the Canadian Institutes for Health Research. J.-H.L. was supported by the Basic Science Research Program funded by the NRF of Korea/Ministry of Education (2022R1A6A3A03071157) and by a long-term postdoctoral fellowship funded by the Human Frontier Science Program (LT0015/2023-L). H.-G.L. was supported by the Basic Science Research Program through the NRF funded by the Ministry of Education (2021R1A6A3A14039088) and The Gene Lay Institute of Immunology and Inflammation of Brigham and Women’s Hospital, Massachusetts General Hospital and Harvard Medical School (FP-0034491). D.F. was supported by the Gemeinnützige Hertie-Stiftung foundation. Illustrations in Figs. 1a,d,h, 2c,d and 4b,g,k,n and Extended Data Figs. 1a,f, 2g and 7d,f were created using BioRender (https://www.biorender.com). We thank L. Ding and staff at the Neurotechnology Studio at Brigham and Women’s Hospital for equipment access; F. Pinho-Ribeiro for discussions on meningeal immune cells; M. Lee and staff at the Gene Lay Institute Genomics Platform for equipment access; Z. Houston, R. Gessner and K. Quick for discussions related to IVIS imaging; R. Krishnan for assistance with flow cytometry; L. Gaffney for assistance with figure illustrations; C. Jennings for feedback on the manuscript text; staff at STOmics, Eve Technologies, Watershed Bioinformatics and Via Foundry; staff at the NIH NeuroBioBank for providing human amygdala samples from healthy individuals and from patients with MDD; staff at the NIDA Drug Supply Program and the Research Triangle Institute for providing psilocybin and MDMA; and the patients and their families for agreeing to participate in this study.
Author information
Authors and Affiliations
Contributions
E.N.C., J.L., C.M.P., J.C., C.F.A., M.K., W.M.W., G.G., M.Y., T.H.H., S.S.D., L.Y., C.R.G.L.d., J.-H.L., L.D., D.F., A.M.S., H.-G.L., O.A. and M.A.W. performed experiments, analysed data and/or interpreted data. S.G. aided with interpretation of data involving psychedelics. S.G., S.M.S., I.M.C., V.K.K. and F.J.Q. contributed specific reagents or methods, interpreted data and provided input on the experiments. E.N.C., J.L. and M.A.W. wrote the manuscript with input from all co-authors. M.A.W. directed and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Scott Russo, Marco Colonna and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Molecular and behavioral changes after 18-days restraint stress.
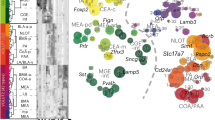
a, Schematic of behavioral paradigm. Tube restraint for 7, 12, or 18 days, followed by cued fear acquisition, with fear retrieval conducted the next day in a novel context with only cue presentation. b, Acquisition of fear behavior in mice exposed to 7-days (n = 10), 12-days (n = 5), or 18-days (n = 15) of stress or controls (n = 14). Two-way repeated measures ANOVA. c, Time spent freezing in conditioned fear behavior across trials. n = 10 mice per group. Two-way repeated measures ANOVA. d, Time in open/closed arm of elevated plus maze (EPM) in mice exposed to 18-days of stress (n = 20) or controls (n = 9). Unpaired t-test. e, Cytokines detected by ELISA in the plasma of mice exposed to 18-days of restraint stress (n = 4) or controls (n = 5). Two-way ANOVA. f, Changes in corticosterone levels in 18-day stressed mice or control mice, measured at 30-min (n = 5 per group) and 6-h (control, n = 5; 18d-stress, n = 4) after final session of restraint stress concluded. Unpaired t-test. g, UMAP plot of Aldh1l1tdTomato/+ cells from control (n = 3) and 18-day stress (n = 3) mice. h, UMAP plot of Aldh1l1tdTomato/+ cells from individual mice. i, Scatterplots of astrocyte (top row) or control (bottom row) markers. j, Pseudotime violin plot of captured Aldh1l1tdTomato/+ cells. Unpaired t-test. k, Prediction by Qiagen IPA of glucocorticoid receptor as a transcriptional regulator of cluster 1 Aldh1l1tdTomato/+ cells. l-o, Heatmap (l), UMAP (m,n), and violin plot (o) of RNA velocity data associating Aldh1l1tdTomato/+ cell clusters to stages of the pseudotime trajectory. p, Dot plot of Egfr expression in each cluster. q, Plots of pathways in cluster 0 (left) and cluster 2 (right) cells. Schematics (a,f) were created using BioRender (https://biorender.com). Data shown as mean ± SEM.
Extended Data Fig. 2 Amygdala astrocytes regulate chronic stress responses.
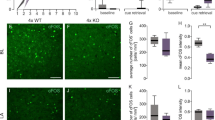
a, Analysis of astrocyte Egfr expression across brain regions using a previously annotated mouse brain single-cell atlas27. One-way ANOVA, Fisher post-test. b, Quantification and confocal images of EGFR expression in astrocytes across multiple brain regions. n = 4 per group. Yellow arrows indicate EGFR+S100β+ cells. Unpaired t-test. c, Analysis by ELISA of cytokines from control (n = 4) or 7-day stressed (n = 10) mice. Two-way ANOVA. d,e, Validation of DeepLabCut forced swim model (d) and quantification (e) of time spent immobile in forced swim. n = 5 per control group, n = 10 for 7-day restraint. Unpaired t-test. f, Quantification (left) and staining (right) validating EGFR KD. n = 6 images from n = 3 mice per group. Unpaired t-test. g, Schematic and bar graph of elevated plus maze (EPM) analysis in mice with astrocytic Egfr knockdown via AAV vector delivery to the basolateral amygdala. Mice were subjected to 7-days restraint stress prior to testing in the EPM (sgRosa26, n = 4; sgEgfr, n = 5). Unpaired t-test. h, Validation of astrocyte cell type markers detected by bulk RNA-seq of amygdala astrocytes. n = 7 mice. i, Analysis of transcription factor motifs detected in sgEgfr (n = 3) versus sgRosa26 (n = 4) astrocytes. j, Plots of PTPR family member expression in cells of each cluster from Aldh1l1tdTomato/+ cells analyzed by scRNA-seq. saCas9, S. aureus Cas9; NS, not significant. FDR, false discovery rate. RPKM, Reads Per Kilobase of transcript per Million mapped reads. Schematic (g) was created using BioRender (https://biorender.com). Data shown as mean ± SEM.
Extended Data Fig. 3 Screening neuron transcription factors involved in fear behavior.
a, Validation of stereotaxic targeting of BLA neurons by immunostaining, using an AAV5 delivering hSyn-driven EGFP. b, Cell type markers for neurons versus other amygdala cell types in samples analyzed in these studies. n = 7 mice. c, Principal component analysis of control amygdala neuron samples collected in this study versus previously published studies. n = 4 mice. d, Pathways upregulated in bulk RNA-seq data of amygdala neurons from astrocyte-targeted sgEgfr (n = 3) relative to controls (n = 3). e, qPCR analysis of the indicated genes in neuron-astrocyte co-cultures following viral transduction with either sgPtprs under the GfaABC1D promoter, sgSlitrk2 under the hSyn promoter, or sgRosa26 controls. n = 5 per group. One-way ANOVA, Dunnett post-test. f,g, Freezing behavior of mice exposed to 7-days of restraint stress following injection of AAV delivering saCas9 under the hSyn promoter and targeting sgRosa26, sgCebpg, or sgEts1. n = 5 mice per group. Two-way repeated measures ANOVA. h, GSEA plot comparing bulk RNA-seq data of amygdala neurons targeting sgNr2f2 (n = 3) or sgRosa26 (n = 4) in neurons. i, Pie chart showing distribution of detected ATAC-seq motifs in neurons isolated from neuron-targeting sgNr2f2 (n = 4) or sgRosa26 (n = 4) mice. j, Quantification of BLA coverage by excitatory neurites as a function of Nr2f2 KD. Cortical notching was used to differentiate between hemispheres. n = 6 images per group. One-tailed unpaired t-test. RPKM, Reads Per Kilobase of transcript per Million mapped reads. NS, not significant. Data shown as mean ± SEM.
Extended Data Fig. 4 Chronic stress and fear conditioning combinatorially regulate neural activity and astrocyte transcriptional signatures.
a, Representative images and bar graph of cFos signals in the BLA of unstressed and stressed mice exposed to 18-days restraint stress, and/or fear conditioning, and/or a control context (control, n = 4; fear conditioning, n = 7; 18-day stress, n = 4; 18-day stress with fear conditioning, n = 6). Yellow arrows indicate masked cFos signals. One-way ANOVA, Holm-Sidak post-test. b-e, Bar plots of bulk RNA-seq data comparing pathways detected in amygdala astrocytes in stressed versus unstressed conditions, with and without fear conditioning. n = 5 mice per group. f, Expression levels of Slitrk2 in neurons of control (n = 4 mice) or fear conditioning (n = 5 mice). One-tailed unpaired t-test. Data shown as mean ± SEM.
Extended Data Fig. 5 Analysis of the chronically stressed brain by Stereo-seq.
a, H&E and nuclei staining of coronal brain sections used in Stereo-seq. b, RIN number of samples used in Stereo-seq analyses. n = 3 per group. Unpaired t-test. c, Rarefaction curve of sequencing depth for each sample used. d, Abundance of RNA molecules detected per spot at a resolution of bin200, meaning a 100 µm2 spot. Please note that Stereo-seq transcriptomic data were analyzed using cellbin analysis, which maintains a resolution of 500-nm. e, Representative tissue architecture for sections used. f, Representative mID counts in an 18-day stressed mouse at cellbin resolution (500-nm). RIN, RNA integrity number. Data shown as mean ± SEM.
Extended Data Fig. 6 Determination of amygdala cell types and states by Stereo-seq.
a, Marker genes used in differentiating the amygdala from other brain areas. b, Representative ROI used (green) to select the amygdala (left) and StereoMap neighborhood-based spatial clustering analysis at bin200 resolution (right). c, Quality control data per cell of mID counts (left), unique genes expressed (middle), and percent of mitochondrial reads (right) within the selected amygdalae. d,e, Projection of sample distribution by cluster (d) and fraction of cell type within each sample (e). n = 16,496 cells. f, Marker genes of cell types detected in the amygdala. g-j, Subclustered cell types detected in the amygdala k, RNA velocity analysis of myeloid cell clusters. l, Plot of microglial or monocyte marker genes in microglial cells detected by Stereo-seq. m, Total counts of selected cytokine genes in analyzed amygdala samples. MSN, medium spiny neurons. CeA, central amygdala. BLA, basolateral amygdala. ROI, region of interest. ND, not detected. Data shown as mean ± SEM.
Extended Data Fig. 7 Mobilization of immune cells in response to chronic stress.
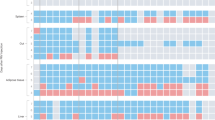
a-c, FACS analysis of immune cell types in meninges (a), dCLN (b), and spleen (c) in mice exposed to 0-, 7-, or 18-days of restraint stress. n = 10 per group for meninges and spleen 18-day stress, n = 5 otherwise. Unpaired t-test. d,e, Schematic of photoconversion in the spleen of Kaede mice (left) and number of photoconverted cells (right) (d); percentage of photoconverted monocytes in the meninges, dCLN, bone marrow, and spleen of stressed and unstressed mice (e). n = 3 mice per group. Unpaired t-test. f, Schematic and FACS analysis of monocyte adoptive transfer via i.v. injection in stressed and control NSG mice. (Unstressed mice (vehicle, n = 5); 7-day stressed mice (vehicle, n = 5; monocyte, n = 10). Kruskal-Wallis test, with Dunn post-test. g, Time spent in open arms versus closed arms in EPM. (Unstressed mice (vehicle, n = 4); 7-day stressed mice (vehicle, n = 5; monocyte, n = 9). Unpaired t-test. h, FACS analysis of CCR2+ pro-inflammatory meningeal monocytes following i.c.m. injection of α-CCR2 or isotype control. n = 5 per group. Unpaired t-test. i, Experimental schematic and FACS validation of meningeal and splenic monocyte depletion in Ccr2DTR/+ mice. Unpaired t-test j, Bar graph of immune cell numbers in mice administered DTx (n = 8) or vehicle (n = 6). Unpaired t-test. k, Immunostaining of Iba1+ microglia in Ccr2DTR/+ mice treated with DTx (n = 11 images from 3 mice) or vehicle (n = 12 images from 4 mice). Note absence of EGFP (driven by Ccr2 promoter) in microglia. Unpaired t-test. l, Time spent in open arms versus closed arms in EPM in Ccr2DTR/+ mice treated with DTx (n = 6) or vehicle (n = 4). Unpaired t-test. m, EGFR transcriptional network (left) and pathway analysis from bulk RNA-seq of amygdala astrocytes (right) from Ccr2DTR/+ mice with or without monocyte depletion after 18-days of restraint stress. Fisher’s exact test. n, Dorsal and ventral images of a mouse brain 2-h after i.c.m. injection of 3% Evans blue dye or 1X PBS. o, Confocal images of coronal brain sections showing Evans blue dye signal in the amygdala. p, Secondary-only negative control of biotinylation signal using mice injected i.c.m. with 1X PBS. q, Expression of IL-1R1 in S100β+ astrocytes in 18-day stress and control groups. n = 17 cells per group. Unpaired t-test. FOV, field of view. NS, not significant. Schematics (d,f) were created using BioRender (https://biorender.com). Data shown as mean ± SEM.
Extended Data Fig. 8 Regulation of the immune response by psychedelics.
a, Expression of marker genes in meningeal monocytes. n = 6 mice per group. b, Normalized gene expression data obtained from ImmGen ultra-low-input bulk RNA-seq across several progenitor, circulating, and tissue-resident myeloid cell types in mice collated in the database. c,d, qPCR analysis of primary splenic CD11b+ cells treated with the given compounds for 6-h. n = 6 per group, except for the compound-only group (n = 5). Mann-Whitney t-test. e, qPCR analysis of gene expression in primary CD11b+ splenic cells treated with or without 10 µM psilocybin (left; n = 6 per group) 10 µM MDMA racemate (right; n = 6 per group, except for the MDMA-only group (n = 5)) or 10 ng/mL LPS for 6-h. One-way ANOVA, Mann-Whitney t-test. f,g Fear acquisition (f) and freezing behavior (g) from mice exposed to 18-days restraint stress or controls. All mice were untreated during fear acquisition and are grouped according to future treatment during fear extinction. Two hours before fear extinction mice were treated with the given compounds. n = 5 (control), n = 10 (vehicle), n = 5 (psilocybin), and n = 4 (MDMA). Two-way repeated measures ANOVA. h, GSEA plots from bulk RNA-seq data of meningeal monocytes exposed to 18-days of restraint stress and treated with 1 mg/kg psilocybin or 10 mg/kg MDMA racemate. n = 3 mice per group. i, qPCR analysis of primary CD11b+ splenic cells treated with indicated compounds for 6-h. n = 6 per group, except for the psilocybin-only group (n = 5). One-way ANOVA, Dunnett post-test. j, Volcano plot of differentially expressed genes from bulk RNA-seq of meningeal monocytes after 18-days of restraint stress and treatment with 1 mg/kg psilocybin or vehicle. n = 3 per group. RPKM, Reads Per Kilobase of transcript per Million mapped reads. Data shown as mean ± SEM.
Extended Data Fig. 9 Psychedelic control of the immune response across tissues.
a-c, FACS analysis of major cell types in the meninges (a), dCLN (b), and spleen (c) in mice exposed to 18 days of restraint stress and treated with vehicle (n = 5), 1 mg/kg psilocybin (n = 5), or 10 mg/kg MDMA racemate (n = 5). Unpaired t-test. d, Expression levels of serotonergic receptors in CD45− cells within the meninges (Htr2a, n = 5; Htr2b, n = 5, Htr2c, n = 4; Ntrk2, n = 7). e, Representative images and quantification of vascular fluorescent signals following tail vein injection in unstressed (n = 13) and stressed mice treated with vehicle (n = 5), 1 mg/kg psilocybin (n = 5), or 10 mg/kg MDMA (n = 4). One-way ANOVA, Holm-Sidak post-test. f, Bar graph and FACS plots showing the number of monocytes in the meninges of mice treated for 6 h with 30 mg/kg nifedipine and psilocybin (n = 10 or n = 5 for vehicle or nifedipine groups, respectively) or MDMA (n = 9 or n = 3 for vehicle or nifedipine groups, respectively). Unpaired t-test. g, Quantification of plasma corticosterone in controls or mice exposed to 18-days of restraint stress and treated with vehicle, 1 mg/kg psilocybin, or 10 mg/kg MDMA racemate. n = 9 mice per group. One-way ANOVA, Tukey post-test. h, Representative images showing validation of chemical sympathectomy by whole mount TH immunostaining of the mouse kidney. i, FACS analysis of splenic (left) and meningeal (right) cells in mice that underwent 18-days of restraint stress followed by chemical sympathectomy, followed by treatment with the given compounds (vehicle, n = 5; psilocybin, n = 4; MDMA, n = 4). Two-way ANOVA, Dunnett post-test (left) or unpaired t-test (right). j, qPCR analysis in primary astrocytes at baseline (left: Il1r1, n = 4; Htr2a, n = 4; Htr2b, n = 5, Htr2c, n = 6; Ntrk2, n = 6) or after a 6-h treatment with 10 µM psilocybin, 10 µM MDMA, or vehicle (right, n = 6 per group). One-way ANOVA per gene. k, Ptprs expression level by qPCR in primary astrocytes treated with indicated compounds for 6-h (vehicle, n = 9; psilocybin, n = 9; MDMA, n = 6). Unpaired t-test. Data shown as mean ± SEM.
Extended Data Fig. 10 Control analyses of human samples.
a, Normalized expression of genes detected by ultra-low-input bulk RNA-seq from human immune cell populations in the blood from the ImmGen human immune system cell atlas. b, Pathways detected by GSEA in primary human monocytes treated for 6-h with 100 ng/mL LPS with or without 10 µM psilocybin or 10 µM MDMA racemate. n = 3 per group. c,d, Demographics of patient samples used for snRNA-seq. n = 6 per group. Unpaired t-test (c) or Fisher’s exact test (d). e, Violin plots of cell type markers for cells detected in the amygdala by snRNA-seq. f, Separation by disease status of samples used in snRNA-seq. g, Pathway plots of cluster 1 and 6 astrocytes from snRNA-seq. h, qPCR analysis of EGFR expression in human amygdala. n = 6 per group. One-tailed unpaired t-test. i, Representative confocal images and bar graphs showing EGFR expression in control and MDD patient amygdala astrocytes. n = 7-10 images and n = 3 patients per group. Unpaired t-test. j, Violin plot of NR2F2 and SLITRK2 expression in excitatory neurons. In violin plots, Solid horizontal line: median; white box: interquartile range (25th–75th percentile); whiskers: min and max values; dashed line: mean. Data shown as mean ± SEM.
Supplementary information
Supplementary Information (download PDF )
Supplementary Data 1–5 and legends to Supplementary Tables 1–22.
Supplementary Tables (download ZIP )
Supplementary Tables 1–22.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chung, E.N., Lee, J., Polonio, C.M. et al. Psychedelic control of neuroimmune interactions governing fear. Nature 641, 1276–1286 (2025). https://doi.org/10.1038/s41586-025-08880-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08880-9
This article is cited by
-
Circuit-informed modulation of traumatic memory in PTSD: integrating extinction, suppression, and reconsolidation
Molecular Psychiatry (2026)
-
Astrocytes enable amygdala neural representations supporting memory
Nature (2026)
-
Psychedelics elicit their effects by 5-HT2A receptor-mediated Gi signalling
Nature (2026)
-
Connection and communication between the nervous and immune systems
Nature Reviews Immunology (2025)
-
Psychedelics target neuroimmune interactions to limit fear
Cell Research (2025)