Abstract
After fertilization, early embryos undergo dissolution of conventional chromatin organization, including topologically associating domains (TADs)1,2. Zygotic genome activation then commences amid unusually slow de novo establishment of three-dimensional chromatin architecture2. How chromatin organization is established and how it interplays with transcription in early mammalian embryos remain elusive. Here we show that CTCF occupies chromatin throughout mouse early development. By contrast, cohesin poorly binds chromatin in one-cell embryos, coinciding with TAD dissolution. Cohesin binding then progressively increases from two- to eight-cell embryos, accompanying TAD establishment. Unexpectedly, strong ‘genic cohesin islands’ (GCIs) emerge across gene bodies of active genes in this period. GCI genes enrich for cell identity and regulatory genes, display broad H3K4me3 at promoters, and exhibit strong binding of transcription factors and the cohesin loader NIPBL at nearby enhancers. We show that transcription is hyperactive in two- to eight-cell embryos and is required for GCI formation. Conversely, induced transcription can also create GCIs. Finally, GCIs can function as insulation boundaries and form contact domains with nearby CTCF sites, enhancing both the transcription levels and stability of GCI genes. These data reveal a hypertranscription state in early embryos that both shapes and is fostered by the three-dimensional genome organization, revealing an intimate interplay between chromatin structure and transcription.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequencing datasets have been deposited in the Gene Expression Omnibus under the accession code GSE200323. Accession codes for the raw published data used in this study are as follows: ADNP ChIP–seq of mouse ES cell, GSE97945; ribosome sequencing and RNA-seq (GSE165782), Hi-C (GSE82185, PRJCA000241), H3K4me3 ChIP–seq (GSE71434 and GSE72784), H3K27ac ChIP–seq (GSE217970 and GSE72784), H3K27me3 ChIP–seq (GSE76687), Pol II Stacc–seq (GSE135457), ATAC–seq (GSE66390) and quantitative mass spectrometry (PXD003315) of mouse early embryos; RAD21 ChIP–seq of Ctcf and Wapl double-knockout MEF, GSE76303; RAD21 ChIP–seq of RIS IMR90, GSE135093; OBOX5 Stacc–seq of 2C embryos, GSE215813; NR5A2 CUT&RUN (GSE216256) and TFAP2C CUT&RUN (GSE229740) of 8C embryos; single-cell RNA-seq of MEFs (GSE203536) and HCT116 cells (GSE144357); RAD21 ChIP–seq of zebrafish embryos, GSE84602 and GSE152744. The housekeeping gene list was retrieved from the HRT Atlas (https://github.com/bidossessih/HRT_Atlas). Source data are provided with this paper.
Code availability
All tools and software used in the analyses are publicly available, and details are described in the Methods. Custom code is available via Zenodo at https://doi.org/10.5281/zenodo.13753153 (ref. 100).
References
Hug, C. B. & Vaquerizas, J. M. The birth of the 3D genome during early embryonic development. Trends Genet. 34, 903–914 (2018).
Du, Z., Zhang, K. & Xie, W. Epigenetic reprogramming in early animal development. Cold Spring Harb. Perspect. Biol. 14, a039677 (2022).
Bonev, B. & Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 17, 661–678 (2016).
Dekker, J. & Misteli, T. Long-range chromatin interactions. Cold Spring Harb. Perspect. Biol. 7, a019356 (2015).
Van Bortle, K. & Corces, V. G. Nuclear organization and genome function. Annu. Rev. Cell Dev. Biol. 28, 163–187 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Davidson, I. F. & Peters, J. M. Genome folding through loop extrusion by SMC complexes. Nat. Rev. Mol. Cell Biol. 22, 445–464 (2021).
Banigan, E. J. & Mirny, L. A. Loop extrusion: theory meets single-molecule experiments. Curr. Opin. Cell Biol. 64, 124–138 (2020).
Nasmyth, K. & Haering, C. H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Davidson, I. F. et al. Rapid movement and transcriptional re-localization of human cohesin on DNA. EMBO J. 35, 2671–2685 (2016).
Banigan, E. J. et al. Transcription shapes 3D chromatin organization by interacting with loop extrusion. Proc. Natl Acad. Sci. USA 120, e2210480120 (2023).
Lengronne, A. et al. Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430, 573–578 (2004).
Olan, I. et al. Transcription-dependent cohesin repositioning rewires chromatin loops in cellular senescence. Nat. Commun. 11, 6049 (2020).
Heinz, S. et al. Transcription elongation can affect genome 3D structure. Cell 174, 1522–1536 (2018).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381 (2017).
Jiang, Y. et al. Genome-wide analyses of chromatin interactions after the loss of Pol I, Pol II, and Pol III. Genome Biol. 21, 158 (2020).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228 (2017).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Zhang, H. Y. et al. Chromatin structure dynamics during the mitosis-to-G1 phase transition. Nature 576, 158–162 (2019).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. Elife 6, e21856 (2017).
Chen, X. et al. Key role for CTCF in establishing chromatin structure in human embryos. Nature 576, 306–310 (2019).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).
Kaaij, L. J. T., Mohn, F., van der Weide, R. H., de Wit, E. & Buhler, M. The ChAHP complex counteracts chromatin looping at CTCF sites that emerged from SINE expansions in mouse. Cell 178, 1437–1451 (2019).
Wang, W. et al. ADNP modulates SINE B2-derived CTCF-binding sites during blastocyst formation in mice. Genes Dev. 38, 168–188 (2024).
Xiong, Z. et al. Ultrasensitive Ribo-seq reveals translational landscapes during mammalian oocyte-to-embryo transition and pre-implantation development. Nat. Cell Biol. 24, 968–980 (2022).
Gassler, J. et al. A mechanism of cohesin-dependent loop extrusion organizes zygotic genome architecture. EMBO J. 36, 3600–3618 (2017).
Liu, B. et al. The landscape of RNA Pol II binding reveals a stepwise transition during ZGA. Nature 587, 139–144 (2020).
Jia, Y. W., Hartshorn, C., Hartung, O. & Wangh, L. J. Heat shock memory in preimplantation mouse embryos. Fertil. Steril. 93, 2760–2763 (2010).
Vilborg, A. et al. Comparative analysis reveals genomic features of stress-induced transcriptional readthrough. Proc. Natl Acad. Sci. USA 114, E8362–E8371 (2017).
Mahat, D. B., Salamanca, H. H., Duarte, F. M., Danko, C. G. & Lis, J. T. Mammalian heat shock response and mechanisms underlying its genome-wide transcriptional regulation. Mol. Cell 62, 63–78 (2016).
Gao, Y. et al. Protein expression landscape of mouse embryos during pre-implantation development. Cell Rep. 21, 3957–3969 (2017).
Benayoun, B. A. et al. H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell 158, 673–688 (2014).
Chen, K. et al. Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor-suppressor genes. Nat. Genet. 47, 1149–1157 (2015).
Dall’Agnese, A. & Young, R. Regulatory architecture of cell identity genes and housekeeping genes. Trends Cell Biol. 33, 1010–1013 (2023).
Ji, S. et al. OBOX regulates mouse zygotic genome activation and early development. Nature 620, 1047–1053 (2023).
Lai, F. et al. NR5A2 connects zygotic genome activation to the first lineage segregation in totipotent embryos. Cell Res. 33, 952–966 (2023).
Li, L. et al. Lineage regulators TFAP2C and NR5A2 function as bipotency activators in totipotent embryos. Nat. Struct. Mol. Biol. 31, 950–963 (2024).
Kim, E., Barth, R. & Dekker, C. Looping the genome with SMC complexes. Annu. Rev. Biochem. 92, 15–41 (2023).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Hsieh, T. H. S. et al. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell 78, 539–553 (2020).
Krietenstein, N. et al. Ultrastructural details of mammalian chromosome architecture. Mol. Cell 78, 554–565 (2020).
Meier, M. et al. Cohesin facilitates zygotic genome activation in zebrafish. Development 145, dev156521 (2018).
Wike, C. L. et al. Chromatin architecture transitions from zebrafish sperm through early embryogenesis. Genome Res. 31, 981–994 (2021).
Nakamura, R. et al. CTCF looping is established during gastrulation in medaka embryos. Genome Res. 31, 968–980 (2021).
Andreu, M. J. et al. Establishment of 3D chromatin structure after fertilization and the metabolic switch at the morula-to-blastocyst transition require CTCF. Cell Rep. 41, 111501 (2022).
Gabriele, M. et al. Dynamics of CTCF- and cohesin-mediated chromatin looping revealed by live-cell imaging. Science 376, 496–501 (2022).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Kruse, K. et al. Transposable elements drive reorganisation of 3D chromatin during early embryogenesis. Preprint at bioRxiv https://doi.org/10.1101/523712 (2019).
Niu, L. et al. Three-dimensional folding dynamics of the Xenopus tropicalis genome. Nat. Genet. 53, 1075–1087 (2021).
Sun, X. M. et al. Size-dependent increase in RNA polymerase II initiation rates mediates gene expression scaling with cell size. Curr. Biol. 30, 1217–1230 (2020).
Swaffer, M. P. et al. RNA polymerase II dynamics and mRNA stability feedback scale mRNA amounts with cell size. Cell 186, 5254–5268 (2023).
Boskovic, A. et al. Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev. 28, 1042–1047 (2014).
Burton, A. et al. Heterochromatin establishment during early mammalian development is regulated by pericentromeric RNA and characterized by non-repressive H3K9me3. Nat. Cell Biol. 22, 767–778 (2020).
Arias, A. M. & Hayward, P. Filtering transcriptional noise during development: concepts and mechanisms. Nat. Rev. Genet. 7, 34–44 (2006).
Zhang, B. et al. Allelic reprogramming of the histone modification H3K4me3 in early mammalian development. Nature 537, 553–557 (2016).
Wang, Q. et al. Imprecise DNMT1 activity coupled with neighbor-guided correction enables robust yet flexible epigenetic inheritance. Nat. Genet. 52, 828–839 (2020).
Xia, W. et al. Resetting histone modifications during human parental-to-zygotic transition. Science 365, 353–360 (2019).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Brind’Amour, J. et al. LTR retrotransposons transcribed in oocytes drive species-specific and heritable changes in DNA methylation. Nat. Commun. 9, 3331 (2018).
Hagemann-Jensen, M. et al. Single-cell RNA counting at allele and isoform resolution using Smart-seq3. Nat. Biotechnol. 38, 708–714 (2020).
Clift, D. et al. A method for the acute and rapid degradation of endogenous proteins. Cell 171, 1692–1706 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Percharde, M., Wong, P. & Ramalho-Santos, M. Global hypertranscription in the mouse embryonic germline. Cell Rep. 19, 1987–1996 (2017).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Abdennur, N. & Mirny, L. A. Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 36, 311–316 (2020).
Open2C et al. Cooltools: enabling high-resolution Hi-C analysis in Python. PLoS Comput. Biol. 20, e1012067 (2024).
Flyamer, I. M., Illingworth, R. S. & Bickmore, W. A. Coolpup.py: versatile pile-up analysis of Hi-C data. Bioinformatics 36, 2980–2985 (2020).
Zhou, Y. Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Tu, S. Q. et al. MAnorm2 for quantitatively comparing groups of ChIP-seq samples. Genome Res. 31, 131–145 (2021).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME Suite. Nucleic Acids Res. 43, W39–W49 (2015).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016).
Hounkpe, B. W., Chenou, F., de Lima, F. & De Paula, E. V. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res. 49, D947–D955 (2021).
Xu, S. L. Y., Grullon, S., Ge, K. & Peng, W. Q. Spatial Clustering for Identification of ChIP-Enriched Regions (SICER) to map regions of histone methylation patterns in embryonic stem cells. Methods Mol. Biol. 1150, 97–111 (2014).
Stovner, E. B. & Saetrom, P. epic2 efficiently finds diffuse domains in ChIP-seq data. Bioinformatics 35, 4392–4393 (2019).
Xia, B. et al. Machine learning uncovers cell identity regulator by histone code. Nat. Commun. 11, 2696 (2020).
Dahl, J. A. et al. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016).
Liu, B. et al. Mapping putative enhancers in mouse oocytes and early embryos reveals TCF3/12 as key folliculogenesis regulators. Nat. Cell Biol. 26, 962–974 (2024).
Chen, K. et al. DANPOS: dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res. 23, 341–351 (2013).
Zhang, X. O., Gingeras, T. R. & Weng, Z. Genome-wide analysis of polymerase III-transcribed Alu elements suggests cell-type-specific enhancer function. Genome Res. 29, 1402–1414 (2019).
Hammal, F., de Langen, P., Bergon, A., Lopez, F. & Ballester, B. ReMap 2022: a database of human, mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res. 50, D316–D325 (2022).
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. zUMIs - a fast and flexible pipeline to process RNA sequencing data with UMIs. Gigascience 7, giy059 (2018).
Choudhary, S. & Satija, R. Comparison and evaluation of statistical error models for scRNA-seq. Genome Biol. 23, 27 (2022).
Hoetker, M. S. et al. H3K36 methylation maintains cell identity by regulating opposing lineage programmes. Nat. Cell Biol. 25, 1121–1134 (2023).
Meir, Z., Mukamel, Z., Chomsky, E., Lifshitz, A. & Tanay, A. Single-cell analysis of clonal maintenance of transcriptional and epigenetic states in cancer cells. Nat. Genet. 52, 709–718 (2020).
Guang, Y. Custom scripts in “Interplay between hyper transcription and de novo establishment of 3D chromatin architecture in early mammalian embryos”. Zenodo https://doi.org/10.5281/zenodo.13753153 (2024).
Acknowledgements
We thank members of the laboratory of W. Xie for discussion and comments during the preparation of the manuscript; the animal and biocomputing facility at Tsinghua University for support; H. Yu and Z. Shi for discussion and for sharing reagents; and Q. Guo, X. Shi, Q. Wang, L. Hu and L. Dong for help with experiments. This work was supported by the National Key R&D Program of China (2021YFA1100102 to W. Xie), the National Natural Science Foundation of China (31988101 and 31830047 to W. Xie), the Postdoctoral Science Foundation of China (2022TQ0183 to K.X.) and the Tsinghua-Peking Center for Life Sciences (to W. Xie). G.Y. is supported by a postdoctoral fellowship from the Tsinghua-Peking Center for Life Sciences. W. Xie is a recipient of an HHMI International Research Scholar award and a New Cornerstone Investigator.
Author information
Authors and Affiliations
Contributions
G.Y., K.X., W. Xia and W. Xie conceived and designed the project. G.Y., K.X., W. Xia and K.Z. performed the CUT&RUN and RNA-seq experiments with the help of Q.X., L. Li, B.L., Z.D. and F.L. K.Z., G.Y., K.X. and Z.D. performed the Hi-C experiments with the help of L. Liu. K.X. collected the mouse embryos and performed the embryo injection and drug treatment experiments with the help of Q.X., Z.L., L. Liu, X.C., Q.F., F.L., W.W., L.W., F.K., C.W., H.D. and H.W. W. Xia performed the mouse ES cell differentiation experiments. G.Y. and W. Xie analysed the data with the help of K.X., W. Xia and K.Z. G.Y. and W. Xie prepared most of the figures and wrote the manuscript with help from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Validation of CTCF CUT&RUN data.
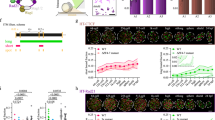
a, Immunofluorescence of CTCF in mouse 1C, 2C, 4C, 8C embryos and blastocysts. Two independent experiments reported similar results. Scale bar, 20 μm. b, UCSC browser views showing CTCF binding with replicates for each stage. c, Scatter plots showing the CTCF enrichment levels (5-kb window) between biological replicates across the genome for each developmental stage. The Pearson correlation coefficients are shown. d, Pie chart showing (left) the percentages of common and variable CTCF sites across all stages in early embryos and (right) the percentage of variable peaks that either overlap with tissue CTCF peaks or contain CTCF motifs.
Extended Data Fig. 2 Characterization of stage-specific CTCF peaks in mouse early development.
a, UCSC browser views showing CTCF signals at representative CTCF sites for 1-2C depleted, 1-8C enriched and common peaks. The expression of ADNP is also shown. b, Bar chart showing the genomic distributions of various types of CTCF peaks. c, Heatmaps showing the CTCF signals for CTCF-bound promoters in the 1-2C depleted class and the expression levels of corresponding genes. d, UCSC browser views showing the CTCF binding in WT, control and Adnp overexpressed embryos. e, Averaged CTCF signals from control or Adnp overexpressed embryos at common or 1-8C enriched CTCF sites. f, Volcano plot showing the upregulated and downregulated genes in blastocyst upon Adnp KO. Representative gene ontology terms are also shown. Adjusted P values calculated by edgeR are shown. g, Line plots showing the cumulative distributions of upregulated, downregulated and other genes in Adnp KO embryos with defined distances (x-axis) between their TSSs and nearest CTCF binding peaks in mouse 8C embryos.
Extended Data Fig. 3 Validation of SMC3 CUT&RUN data.
a, Immunofluorescence of SMC3 in mouse 1C, 2C and 8C embryos. Embryos were permeabilized after fixation. Scale bar, 20 μm. Two independent experiments reported similar results. b, Immunofluorescence of RAD21 in mouse 1C, 2C, 8C and blastocysts. Embryos were permeabilized prior to fixation. Scale bar, 20 μm. Two independent experiments reported similar results. c, The translation levels of RAD21, SMC1A, STAG1 and STAG2 in early embryos. d, UCSC browser views showing SMC3 binding with replicates for each stage. e, Scatter plots showing the SMC3 enrichment levels (5-kb window) between biological replicates across the genome for each developmental stage. The Pearson correlation coefficients are shown. f, Averaged directionality index scores at different quartiles of CTCF sites with respect to the cohesin traveling distances. g, The CTCF sites were pre-selected to ensure comparable enrichment of CTCF signals in different quartiles, as validated by the box plots showing CTCF signals at CTCF sites from different quartiles (indicated by colors) with respect to the cohesin traveling distances. A total of 12,003 peaks were analyzed. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × IQR. P values calculated by one-sided Mann-Whitney U-test are also shown.
Extended Data Fig. 4 Relationship of GCIs and transcription.
a, Heatmaps showing the SMC3, Pol II and CTCF signals at distal CTCF sites (promoter sites removed, left) and all genes (right). b, UCSC browser views (top) and SMC3 signals (bottom) at GCI genes identified in Ctcf Wapl DKO MEF13, RAS-induced senescence IMR90 cells17 and mouse 8C embryos. c, (Left) Schematic model showing the experimental design of heat shock treatment in 8C embryos. (Right) UCSC browser views showing the total RNA, Pol II and SMC3 signals with or without heat shock. d, UCSC browser views showing the Pol II and SMC3 signals with or without heat shock at the Nr5a2 locus. e, (Left) Spike-in normalized Pol II signals at GCI genes in 8C embryos with or without heat shock. (Right) Spike-in normalized SMC3 signals at the CTCF sites (promoter CTCF sites removed) or at GCI genes in 8C embryos with or without heat shock.
Extended Data Fig. 5 Hypertranscription in mouse early embryos.
a, Averaged signals of Pol II and SMC3 at the top 500 transcribed genes in early embryos and somatic tissues/cell lines. b, Validation of ERCC spike-in RNA-seq data using different numbers of mESCs and embryos (n = 4 independent experiments). Dashed lines indicate the expected fold changes between samples using different cell numbers. c, Jitter plot combined with boxplot showing the ERCC normalized reads per kilobase (RPK) for top 500 newly activated genes during embryo development and retinoic acid (RA) induced differentiation. Inset shows magnified view when lowering Y-scale. Each dot in the jitter plot represents one gene. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × IQR. d, Jitter plot combined with boxplot showing the ERCC normalized reads per kilobase (RPK) for top 500 newly activated genes during embryo development (replicate 2), RA induced differentiation (replicate 2), and BMP4/Activin A induced differentiation. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × IQR.
Extended Data Fig. 6 GCI genes are enriched for embryo-specific regulatory genes.
a, UCSC browser views showing SMC3 signals at representative GCI and HE gene locus in mouse early embryos. b, Scatter plots showing the distribution of HE and GCI genes with respect to their expression levels and ranks in mouse 8C embryos. c, Heatmaps showing the gene expression and protein37 levels of HE-specific, HE-GCI common and GCI-specific genes in mouse early embryos. d, Boxplot showing the Log2 expression fold change of GCI (n = 993) and HE genes (n = 1,000) after 4 or 8 h of DRB treatment. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × IQR. P values calculated by one-sided Mann-Whitney U-test are also shown. e, UCSC browser views showing H3K4me3 and H3K27ac distribution at the promoters of representative GCI and HE genes in mouse 8C embryos. f, Venn diagram showing the overlap between housekeeping genes and GCI genes. P values calculated by hypergeometric test. g, UCSC browser views showing SMC3 and Pol II signals on representative genes for HK-specific, HK-GCI common and GCI-specific classes. The related RNA and protein levels at corresponding stages are also plotted on the right side. h, Heatmaps showing the gene expression and protein37 levels of HK-specific, HK-GCI common and GCI-specific genes in mouse early embryos and somatic tissues.
Extended Data Fig. 7 GCI genes exhibit strong binding of transcription factors and NIPBL in nearby regulatory elements in early embryos.
a, Line plots showing the cumulative distributions of GCI, HE (embryo) or top Pol II (Ser2) genes (other cell types), expressed and all genes with defined distances (x-axis) between their TSSs and nearest distal key transcription factor binding peaks in mouse 8C embryos and various cell lines. b, UCSC browser views showing SMC3, Pol II33, NR5A242, TFAP2C43 and NIPBL binding signals at the eukaryotic translation initiation factor (Eif) gene family in mouse 8C embryo. *OBOX5 binding data41 are available only in 2C embryos. c, Heatmaps showing the binding signals of NIPBL, OBOX5, NR5A2, and TFAP2C in distal accessible chromatin regions in 8C embryos (2C for OBOX5).
Extended Data Fig. 8 GCIs form contact domains with nearby CTCF sites.
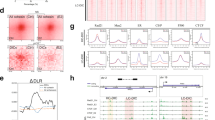
a, Heatmaps showing the insulation scores around GCI genes in mouse early embryos19,20 in the absence or presence of α-amanitin (20 h and 45 h, starting from 1C) or DRB (4 h, starting from 8C), and mESCs. b-c, Heatmaps showing the chromatin interaction frequencies at example GCI genes Gata6 (b) and Zbtb10 (c) in mouse early embryos in the absence or presence of α-amanitin (20 h and 45 h starting from 1C)19,20. Corresponding CTCF binding, SMC3 binding, RNA expression and insulation scores are also shown. CTCF and SMC3 CUT&RUN for α-amanitin treated embryos was only performed for 45 h group. Green arrows refer to the chromatin boundaries at Gata6 or Zbtb10, with position also indicated by dashed lines in all cell types. Orange arrows indicate the valleys of insulation scores at Gata6 or Zbtb10.
Extended Data Fig. 9 GCI-CTCF domains and the directions of CTCF motifs and GCI genes.
a, (Left) Schematic model showing the regions used for pile-up interaction heatmaps. (Right) Pile-up interaction heatmaps showing the interactions between randomly shuffled regions and CTCF binding sites (Methods). The dashed circles indicate interactions between regions near CTCF and regions beyond the shuffled sites. b, (Top) Schematic models showing the classification of GCI-CTCF pairs according to directions of the CTCF motifs and the GCI genes. Stripes originating from the CTCF sites are represented as red bars. (Bottom) Aggregated interaction heatmaps showing the interactions between GCI genes and CTCF sites with different combinations of directions in 8C embryos in the presence (starting from 1C) or absence of α-amanitin. c, 1D Micro-C profiles to show the phased nucleosome occupancy around the CTCF binding sites in 8C embryos.
Extended Data Fig. 10 GCIs form contact domains with nearby CTCF and facilitate both transcription amplitude and stability of GCI genes.
a, (Left) Immunofluorescence of RAD21 in IgG or RAD21 Trim-Away 2C embryos. Scale bar, 20 μm. (Right) Embryo morphology of IgG or RAD21 Trim-Away embryos at the 8C stage. Scale bar, 50 μm. Two independent experiments reported similar results. b, Immunofluorescence of RAD21 in control, Rad21 KD and Triple KD 8C embryos. Scale bar, 10 μm. Two independent experiments reported similar results. c, Spike-in normalized signals of SMC3 at CTCF and GCI genes (left) and CTCF at CTCF binding sites (right) in control, Rad21 KD and Triple KD 8C embryos. d, Principal component analysis (PCA) of transcriptomes of mouse early embryos and cohesin KD embryos. e, Heatmaps showing normalized insulation scores around GCI genes in control, Rad21 KD and Triple KD 8C embryos. f, Normalized Pol II signals at GCI genes (left) and non-GCI genes (right) in control, Rad21 KD and Triple KD 8C embryos. g, Gene set enrichment analysis (GSEA) showing the enrichment of GCI genes in Rad21 KD and Triple KD 8C embryos. P values were generated by permutation tests. h, Gene expression changes upon cohesin knockdown. Dots represent biological replicates. i, Embryo morphology of control, Rad21 KD and Triple KD embryos after 120 h of hCG injection. Scale bar, 100 μm. j, Box plots showing the transcription variation of GCI genes (n = 993), HE genes (n = 1,000), top Ser2 Pol II genes (n = 1,000) and all expressed genes (n = 7,875) in mouse 2-8C embryos and other cell types97,98. Center line, median; box, 25th and 75th percentiles; whiskers, 1.5 × IQR. P values calculated by one-sided Mann-Whitney U-test are shown.
Supplementary information
Supplementary Tables 1 and 2 (download XLSX )
Supplementary Table 1: The number of embryos or cells used in various experiments; the number of mouse embryos or cells used in CUT&RUN, Hi-C and RNA-seq. Supplementary Table 2: siRNA sequences used in cohesin KD experiment; siRNAs targeting Nipbl, Smc3 and Rad21 are included.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, G., Xu, K., Xia, W. et al. Establishment of chromatin architecture interplays with embryo hypertranscription. Nature 646, 208–217 (2025). https://doi.org/10.1038/s41586-025-09400-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09400-5