Abstract
Posterior fossa type A (PFA) ependymoma is an unusual infantile brain tumour with few known somatic mutations, thought to be driven by epigenetic mechanisms1. PFA ependymoma has a markedly higher incidence and worse prognosis in male children than in female children2. The mechanisms that underlie these sex differences are at present unknown. Here we show that the cellular hierarchy of PFA ependymoma is less differentiated in male individuals than it is in female individuals. In the normal developing mouse hindbrain, male gliogenic progenitors are less differentiated than matched female sibling controls. To further parse the effects of chromosomal versus gonadal contributions in the male hindbrain, we used the four-core genotype mouse model3, which showed that androgen signalling, rather than sex chromosomes, prolongs hindbrain differentiation in male mice. Androgen supplementation promotes the growth of PFA ependymoma, but not that of other brain tumours. Conversely, androgen blockade diminishes both the stem-like potential and the proliferation of PFA ependymoma. We conclude that androgen signalling in both the normal developing hindbrain and PFA ependymoma is sufficient to promote growth and delay differentiation. Anti-androgen therapies represent a potential clinical avenue to target this currently untreatable childhood cancer.
Similar content being viewed by others
Main
Ependymomas are malignant brain tumours that occur throughout the central nervous system (CNS)4,5. Usually diagnosed by histology, they are now classified into at least ten clinically and molecularly distinct subtypes6. The most common is the hindbrain-derived posterior fossa type A (PFA) ependymoma, with a median age of onset of around 3 years, no effective chemotherapy and the worst prognosis, with five-year progression-free survival (PFS) rates of around 50%2. PFA ependymomas (PFA-EPNs), unlike most lethal cancers, lack clear genetic drivers, which hinders mechanistic studies and the development of therapies1,7. Most cases have no known somatic single-nucleotide variants or copy-number variants, with 1q gain being the most frequently observed somatic alteration6. A small minority of cases exhibit EZHIP missense mutations or the histone H3.3 H3K27M mutation that is also seen in diffuse midline gliomas8. PFA-EPNs exhibit CpG hypermethylation and low levels of H3K27me3, mediated by expression of the EZH2 and PRC2 inhibitor EZHIP (ref. 1) and sustained by an unusual metabolic phenotype with obligate hypoxia7. Because PFA-EPNs occur in infants and toddlers, they are hypothesized to arise from cell types that are found only in utero9,10—a notion supported by the identification, within PFA tumours, of early embryonic cell types such as gliogenic progenitors (GPs) and roof-plate-like stem cells (RPSCs)2,9,11. PFA-EPN probably arises when embryonic cells stall in differentiation, but beyond links to PRC2 and hypoxia, the mechanisms that block maturation and drive transformation remain unclear7.
Various clinical trials have shown that children with PFA-EPN are more likely to be male, and that male individuals have a significantly worse survival rate than female individuals do2. Although the underlying reasons are unclear, research in healthy children at the same age has shown that the transcriptomes of cerebellar neurons from female individuals seem to be more mature than do those from male individuals12. Male cerebella are about 10% larger than female cerebella, and the volume of the cerebellum peaks markedly later in boys than in girls13 (15.6 years versus 11.8 years).
Sex differences are well known to have key roles in oncogenesis14,15,16,17,18,19,20. Our findings reveal that androgen-dependent developmental programs in the embryonic hindbrain explain the male bias in incidence and outcome, and highlight androgen signalling as a mechanistically grounded target for therapeutic intervention.
PFA-EPNs are more common in male individuals
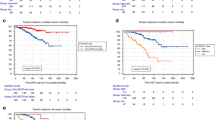
Male individuals diagnosed with PFA-EPN have a significantly worse prognosis than female individuals do, even when receiving the current best treatment (complete surgical resection and fractionated radiation) (Fig. 1a), and independently of cytogenetic markers such as chromosome 1q gain (Extended Data Fig. 1a). Incidence is also consistently higher in male than in female individuals across multiple cohorts2,6,8,21,22 (1.8–1.08, Fig. 1b). The reasons for and the mechanisms that underlie the sex differences in the incidence and prognosis of PFA-EPN are unknown.
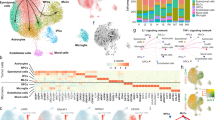
a, Sex differences in prognosis. Progression-free Kaplan–Meier survival plot for first-presentation patients with PFA-EPN (n = 300) who received gross total tumour resection and radiation. Numbers at risk are shown at the bottom. Statistical significance was calculated using a two-sided log-rank test; shaded areas indicate 95% confidence intervals. b, Literature summary of differences in the incidence of PFA-EPN between male (M) and female (F) individuals (C1–C4 denote dataset cohorts). c, Uniform manifold approximation and projection (UMAP) visualization of male and female samples in the tumour fraction from the PFA scRNA-seq cohort. d, Mean CancerSEA stemness score across the tumour fraction per sample, split by sex. e, UMAP visualization of tumour cell types in the tumour fraction from the PFA scRNA-seq cohort. f, Top three most active regulatory networks by tumour cell type. g, GP-like tumour cell fraction within the PFA sample tumour cells by sex. d,g, Female (n = 8) and male (n = 18). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles and whiskers mark 1.5× the interquartile range (IQR). P values were calculated using a two-sided Student’s t-test. h, Monocle3-inferred pseudotime trajectory of the GP-like tumour cluster split by sex. i,j, Kaplan–Meier curves of OS (i) and PFS (j) in male patients with PFA-EPN (n = 39), stratified by expression of the GP-like male-biased TF signature (low or high) in a microarray cohort treated with standard therapy (complete tumour resection and radiation). Numbers at risk are shown at the bottom. Statistical analyses were performed using a two-sided log-rank test; shaded areas indicate 95% confidence intervals.
To understand what drives the stark clinical differences between male and female patients with PFA-EPN, we analysed 26 PFA primary tumour samples (Supplementary Table 1; n = 18 male, 8 female) profiled by single-cell RNA sequencing (scRNA-seq)9,23 (Extended Data Fig. 1b). Broad cell types were annotated on the basis of cell-type markers and regulatory networks (rn) (Extended Data Fig. 1b–d). Copy-number analysis at the single-cell level was performed in samples with chromosome 1q gain, with lymphoid cells as a reference, and was used as validation for the separation of tumour cells from microenvironment cells across the clusters (Extended Data Fig. 1e). PFA samples were composed of approximately 60% tumour cells (CLU, RFX2-rn), 30% myeloid cells (SPP1, MAF-rn), 2% vascular cells (IGFBP7, SOX17-rn), 7% lymphoid cells (CD52, TBX21-rn) and 1% oligodendrocyte precursor cells (OPCs) (APOD, OLIG2-rn) (Extended Data Fig. 1f). There were no significant differences between male and female patients in the ratios of tumour cells to microenvironment cells, consistent with a model in which the differences in behaviour between male and female PFA-EPNs are mostly tumour-cell autonomous (Extended Data Fig. 1g).
Male PFA-EPNs show increased stemness
To investigate cell-autonomous sex-related factors, we focused on the tumour cells (Fig. 1c). PFA-EPNs are presumed to mirror early embryonic-stem-like neural cells, developmentally stalled during glial fate specification9. We thus compared the CancerSEA24 stemness module score between male and female patients, and found that it was higher for male individuals both at the single-cell level (Extended Data Fig. 2a) and when averaged per sample (pseudobulk) (Fig. 1d). Next, we performed clustering analysis and identified six tumour subpopulations that displayed distinct markers (Fig. 1e, Extended Data Fig. 2b,c and Supplementary Table 2), differentiation trajectories (Extended Data Fig. 2d,e) and regulatory networks (Fig. 1f). The early-to-intermediate stages of the inferred differentiation trajectory were populated by the neural stem cell (NSC)-like (CRABP1, POU4F2-rn), cycling (CENPF, EZH2-rn) and gliogenic progenitor-like (GP-like) (IGF2, HES1-rn) tumour cells, and the more differentiated stages included the hypoxic (VEGFA, TEAD4-rn), ciliated (C1orf194, PRDX5-rn) and immune-like (CD74, MAFB-rn) tumour cells. Notably, differential abundance analysis revealed a strong male-biased enrichment in cycling and GP-like tumour neighbourhoods, whereas ciliated and immune-like tumour neighbourhoods were more enriched in female individuals (Extended Data Fig. 2f). Consistently, male patients had a significantly higher proportion of GP-like tumour cells (54% versus 14%; Fig. 1g and Extended Data Fig. 2g,h), as well as showing an overall increase in cycling tumour cells (Extended Data Fig. 2i). The NSC-like cluster, although too small for statistical testing, was present in 5 out of 18 (28%) male tumours, as compared with only 1 out of 8 (12.5%) female tumours. Together, these results are consistent with a model in which PFA-EPN results from developmentally stalled stem-like cells failing to complete differentiation, with sex-related differences in the extent of the differentiation delay.
Transcriptional programs are retained in PFA-EPN
To define transcriptional programs that maintain differential progression along a differentiation axis, we compared tumour cells from male and female patients, using differential expression and regulatory network inference analysis. Male and female PFA-EPN cells share regulons with comparable activity, including EZH2, SMAD3, OLIG2 and TCF3 (Extended Data Fig. 3a and Supplementary Table 3). EZH2, the catalytic subunit of PRC2, defines the characteristic epigenetic landscape of PFA-EPN and functions as a sex-independent epigenetic driver1. SMAD3 regulates proliferation in neural progenitors25, whereas OLIG2 and TCF3 maintain glial lineage identity and stem-like potential. These shared regulatory networks delineate core transcriptional and epigenetic programs that sustain the malignant progenitor state of PFA-EPN, regardless of sex.
In addition to presenting differences in cell proportions, male GP-like tumours mapped earlier in pseudotime than female cells did, suggesting a less differentiated phenotype (Fig. 1h). We thus derived a male-biased gene signature, identifying genes with higher expression levels in male than in female GP-like cell populations (Extended Data Fig. 3b–d and Supplementary Table 3). This ‘male-biased PFA GP-like signature’ (GPlike.MbiasGene.Sig) contains early embryonic developmental genes, providing further evidence for an embryonic-stem-cell-like state in GPs from male PFA-EPN samples (Extended Data Fig. 3e). Differential regulatory network inference analysis, followed by filtering for master regulators with a significant male bias in average gene expression (Extended Data Fig. 3a,b and Supplementary Table 3), identified a second signature (GPlike.MbiasTF.Sig) for transcription factors (TFs), which also shows higher activity for early brain-development networks in male PFA-EPN samples (Extended Data Fig. 3f,g).
The male-biased signatures included several key developmental TFs, including NFIB and ZIC1, as well as PAX3, MSX1 and PBX1 (Extended Data Fig. 3f), all of which are known to be important in neural-plate and neural-crest development26,27. The signatures also included DACH1 (ref. 28) (Extended Data Fig. 3c), a regulator of neural-crest migration and differentiation through its interaction with the protein encoded by JUN. JUN (ref. 29), an immediate–early gene with a central role in cellular proliferation, was among the top marker genes for the GP-like tumour cluster (Extended Data Fig. 2b,c). Furthermore, the signatures included PAX6, a canonical marker and master regulator of radial glia30. Because PFA-EPNs are hypothesized to originate in radial glia, the male-biased activity of these regulators suggests that the transcriptional sex differences reflect normal embryonic development, rather than being acquired during transformation.
To determine whether the glial progenitor cell state associates with the overall tumour phenotype, we correlated tumour composition with the expression of male-biased gene and TF signatures in the GP-like tumour cells. In male samples, higher expression of both signatures in GP-like tumour cells correlated with an increase in the fraction of cycling cells and a decrease in the fraction of more-differentiated ciliated cells (Extended Data Fig. 3h,i). This suggests that a more stem-like state in progenitor cells supports tumour proliferation in male individuals and reflects the hierarchy of cellular differentiation in the tumour.
Finally, to assess the clinical relevance of the male-biased signatures, we compared the outcomes of patients6,23 receiving optimal treatment (gross total resection and radiotherapy), stratified by their level of expression of the GP-like male-biased signatures (Supplementary Table 1). Male patients with high expression of the GP-like male-biased gene signature had significantly worse overall survival (OS) (Extended Data Fig. 3j), and a trend towards reduced PFS (Extended Data Fig. 3k). The TF-based signature was more notable, with male patients in the high-expressing group having significantly worse OS and PFS (Fig. 1i,j). We conclude that the relative cellular immaturity of male PFA GP-like cells is relevant to the poor prognostic phenotype observed in boys with PFA-EPN.
Sex-balanced atlas of the mouse hindbrain
To characterize the diversity of cell types in the embryonic hindbrain, their developmental course and the sex-related differences, we collected 62 mouse hindbrain samples from 8 time points for scRNA-seq profiling (n = 3 female, 3 male for embryonic day (E) 9; n = 4 female, 4 male for E10–E18) (Fig. 2a and Extended Data Fig. 4a). Expression of the X-chromosome-inactivation gene Xist and the Y-chromosome-specific gene Eif2s3y confirmed the sex assignment of embryos (Extended Data Fig. 4b). Clustering analysis revealed 41 distinct subpopulations (Fig. 2b), which we annotated on the basis of cell-cycle status and known cell-type markers (Extended Data Fig. 4c,d). Annotations were confirmed using a machine-learning approach trained on single-cell hindbrain references (Extended Data Fig. 4e). The selected time points cover all of the key stages in hindbrain development, capturing the emergence and maturation of key neuronal lineages (Fig. 2c,d). All clusters were found to contain cells from both female and male donors (Extended Data Fig. 5a,b). We further examined both the emergence and the disappearance (differentiation) of various cell types throughout embryonic hindbrain development (Extended Data Fig. 5c). No major sex differences in the timing of cell-type distribution were observed, suggesting that the overall developmental trajectory of the hindbrain is similar in both sexes.
a, Overview of methodology, data collection and analysis of single-cell transcriptomics in the mouse embryonic hindbrain. The number of cells retained after quality control at each embryonic stage is indicated below the corresponding developmental time point. uRL, upper rhombic lip. Created in BioRender; Pomada Villalbi, A. https://BioRender.com/1pyuif6 (2025). b, UMAP visualization of 420,256 mouse embryo cells. Each dot represents a single cell, colour-coded by cell type. NECs, neuroepithelial cells; IsO, isthmic organizer; NC, neural crest; RP, roof plate; RL, rhombic lip; GC, granule cell; UBC, unipolar brush cell; NTZ, nuclear transitory zone; eDCN, excitatory deep cerebellar nuclei; GABA, γ-aminobutyric acid; INs, interneurons. c, UMAP plots highlighting the expression of known lineage markers. The numbers of the cell-type clusters in each lineage are listed in the coloured boxes. d, Cells detected at each corresponding time point from E9 to E18 are highlighted in cyan in UMAP plots. e, UMAP visualization of glial lineage cells colour-coded by cell type. f, UMAP visualization of inferred lineage trajectories. g, UMAP plot showing cells colour-coded by embryonic time stage. h, UMAP visualization of predicted pseudotime, with cells colour-coded by pseudotime (dark, early; yellow, late). i, Top 20 sex-biased regulons in GPs ranked by z-scaled area under the curve (AUC) score. Male-biased regulons are shown in purple, female-biased regulons are shown in red and regulons with comparable activity between the sexes that are particularly relevant to GPs are highlighted with green circles. F, female; M, male. j, Top ten significantly enriched gene ontology (GO) biological pathways for GPs in each sex based on differentially expressed genes (DEGs). Top, male cells; bottom, female cells (three pathways are significant). Enrichment was assessed using a one-sided cumulative hypergeometric test with the g:SCS multiple-testing correction. Terms with adjusted P < 0.05 were considered significant.
Because ependymoma mirrors GPs at the top of the tumour cellular hierarchy9, we focused on glial-lineage cells, defining more granular populations by clustering (Fig. 2e and Extended Data Fig. 6a) and inferring state transitions by trajectory analysis (Fig. 2f). The inferred pseudotime was highly consistent with the developmental embryonic time (Fig. 2g,h). NSCs from E9 to E10 occupied the trajectory apex and branched into two differentiation paths: one leading to ventricular zone (VZ) progenitors (Ptf1a and Kirrel2), GPs (Ttyh1 and Fgfbp3), Bergmann glia (Gria1 and Gdf10) and astrocytes (Aldoc and Apoe); and the other leading to RPSCs (Wnt1 and Mafb) and choroid plexus (Foxj1 and Ttr). We compared the relative proportions of each cell type between sexes, across replicates and across developmental time (Extended Data Fig. 6b,c). NSCs and RPSCs were most abundant during early embryonic stages (E9–E13), and gradually diminished over time. VZ progenitors and choroid plexus cells were seen at later time points, and more differentiated cells such as Bergmann glia and astrocytes were observed from around E13. Altogether, this transcriptomic survey of the developing hindbrain provides a molecular definition of cell types and a reference of cellular transitions to investigate sex-related differences during the differentiation of the hindbrain glial lineage.
Sex-specific patterns of glial maturation
To investigate molecular mechanisms of glial differentiation in the hindbrain, we first inferred transcriptional regulatory networks (Fig. 2i, Extended Data Fig. 6d and Supplementary Table 4). For GPs, the top 20 female regulons included differentiation-associated networks, such as neuronal differentiation (Neurod1), retinoic acid-induced differentiation (Rara) and orphan nuclear receptors that are crucial for neural specification (Nr2e1). By contrast, top male regulons included neural progenitor regulatory networks (Six2 and Max), transcriptional control of stemness and proliferation (Sp1), and hormone-related or neuroendocrine pathways (Etv2, Rax and Mafk). A set of regulons had comparable activity across sexes, including Ezh2, Sox2, Sox9 and Msx1. These are well-established regulators of GPs, indicating that, although male and female GPs share core regulatory networks, they also have distinct sex-biased regulatory programs that involve key developmental TFs. We next performed differential expression analysis between males and females for each cell population. Female GPs had significantly enriched pathways related to translational initiation (Eif3j1 and Eif5b) and regulation of RNA metabolic processes (Igf2bp1 and Elavl4). By contrast, male GPs had significantly enriched pathways for neural precursor cell proliferation (Fabp7 and Ptn), Y-linked inheritance (Ddx3y and Kdm5d) and gliogenesis (Nfib and Plpp3) (Fig. 2j, Extended Data Fig. 6e and Supplementary Tables 5 and 6). Altogether, these analyses suggest that male hindbrain GPs tend to be less differentiated than their age-matched, sibling female counterparts.
Male tumours resemble progenitor cells
We showed above that male PFA-EPNs have a higher fraction of GP-like cells, and that male PFA progenitor cells as a group are less differentiated than matched female PFA progenitors. Increased expression of early-lineage genes from the GP-like cells is enriched in male tumours and correlates with a poor prognosis among male PFA-EPNs. In the normal developing embryonic mouse male hindbrain, cells in the gliogenic lineage are less differentiated than similar cells from female mice. We thus mapped PFA tumour cells to the glial lineage using three independent, complementary approaches: integration, classification and deconvolution analyses.
We first performed a direct integration of the normal embryonic hindbrain glial lineage with the PFA-EPN cells, followed by clustering (Fig. 3a,b and Extended Data Fig. 7a–d). Trajectory inference analysis verified that hierarchical relationships were maintained in the glial lineage after integration (Fig. 3c and Extended Data Fig. 7e), with cluster 1 at the apex, composed mainly of stem cells, followed by clusters 2–4, composed mainly of progenitors. On the tumour side, the NSC-like, cycling and GP-like clusters showed the strongest integration with NSCs, VZ progenitors and GPs from the normal hindbrain glial lineage, suggesting that the tumour cells retain a high degree of similarity to their embryonic state (Extended Data Fig. 7c,f–i). Next, we focused on the tumour cells, stratifying by sex. Tumour cells from male PFA-EPNs mapped mainly to the early–mid trajectory, whereas female cells were found at later time points (Fig. 3d,e), providing further evidence that female tumours are stalled at later points of glial differentiation than male tumours. Finally, we compared fractions of the newly identified clusters by sample (Fig. 3f and Extended Data Fig. 8a). Whereas no female-biased clusters were observed, male PFA-EPNs showed significantly greater integration with clusters positioned at the early stage of the pseudotime trajectory: NSCs in cluster 1 and both NSCs and GPs in cluster 3 (Fig. 3f and Extended Data Fig. 7e). These findings suggest that male PFA GP-like cells are transcriptionally more similar to earlier stages of hindbrain development, as compared with female cells.
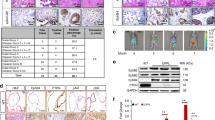
a, UMAP visualization of integrated mouse glial lineage and PFA-EPN cells. b, UMAP visualization of mouse glial lineage and PFA-EPN tumour integrated clusters. Mm indicates normal mouse glial lineage; PFA indicates PFA-EPN cells. Bglia; Bergmann glia; CP, choroid plexus. c, UMAP plot showing trajectory inference using CellRank. d, UMAP visualization of PFA-EPN cells in glial differentiation pseudotime, split by sex. e, Ridge plot visualization of male versus female PFA tumour cells in glial differentiation pseudotime. f, Clusters with significant differences in integrated cell-type proportions in PFA-EPN tumours per sample, by sex. g, CHETAH hierarchical classification tree of the glial lineage. h, CHETAH classification nodes with significant differences in the fraction of the classified cell in PFA-EPN tumours per sample, by sex. i, Glial cell types with significant differences in the BayesPrism deconvoluted fraction within the pseudobulk of PFA-EPN samples, compared between sexes. f,h,i, Female (n = 8) and male (n = 18). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles, and whiskers mark 1.5 × IQR. P values were calculated using a two-sided Student’s t-test. *P < 0.05, **P < 0.01; exact P values are provided in the Source Data.
To validate the higher resemblance of male PFA-EPNs to GPs and VZ progenitors, we first applied CHETAH31, an algorithm that generates nodes along a hierarchical tree to represent intermediate cell states for cells with high similarity to several cell types (Fig. 3g). We found a significant male bias in tumour cells classified to the stem and progenitor cell types in the glial lineage (node 2), and to an intermediate state between the gliogenic and VZ progenitors (node 4), once again demonstrating that male PFA-EPNs have higher transcriptional similarity to stem and progenitor cell states than do female tumours (Fig. 3h and Extended Data Fig. 8b). Next, we applied a deconvolution algorithm, BayesPrism32, to pseudobulk tumour samples, using the glial lineage as a reference, and obtained consistent results: the fraction of cells from the early embryonic gliogenic and VZ progenitors was higher in male tumour samples than in female ones (Fig. 3i and Extended Data Fig. 8c).
Cumulatively, these three distinct transcriptional mapping methods show that male PFA tumour cell states, compared with female ones, have a higher resemblance to earlier time points in the embryonic hierarchy of the developing hindbrain. The delayed differentiation of the male hindbrain creates a longer period of risk for the possible transformation of a normal cell into a PFA cell, and could allow the accumulation of a larger population of GPs. Both of these factors are consistent with the higher incidence of PFA-EPN in male individuals.
Androgens inhibit glial differentiation
Sexual differences in development are usually attributed to either hormonal effects (androgens) or differences in the complement of sex chromosomes (XX versus XY). Single-cell transcriptomic profiling of the developing hindbrain revealed that embryonic hindbrain GPs reach peak incidence levels at E16—a time point when gonadal testosterone production in the mouse is already underway9. To determine whether the role of sex in the dynamics of glial differentiation in the embryonic hindbrain is secondary to either the role of sex chromosomes or the action of sex hormones, we used the four-core genotypes (FCG) model3. In this genetically engineered mouse model, the testes-determining gene (Sry) is deleted from the Y chromosome (henceforth referred to as Y−), and an Sry transgene is inserted onto an autosome. The presence of the Sry gene is sufficient for testes development and gonadal testosterone production, regardless of the complement of sex chromosomes. Conversely, mice that lack Sry develop ovaries and do not produce testosterone, regardless of their sex-chromosome complement. Breeding XY−-Sry mice with wild-type XX females results in four possible genotypes: XX with ovaries (XXF), XY− with ovaries (XYF), XX-Sry with testes (XXM) and XY−-Sry with testes (XYM) (Fig. 4a). Comparisons of XX and XY mice with the same gonad type allows the contribution of sex chromosomes in a genetic model to be identified. Similarly, comparisons of mice with ovaries versus mice with testes of the same sex-chromosome complement enables the contribution of sex hormones to be determined.
a, Schematic of the FCG mouse model. b, UMAP visualization of FCG mouse cells at E16 and E18. Each dot represents a single cell, colour-coded by cell type. c,d, Significantly enriched pathways upregulated by each of the four genotypes in E16 (c) and E18 (d) GPs. e, Volcano plot of differential gene-expression analysis between XX and XY GPs at E16. Significant DEGs are indicated in red (XXF and XXM) and blue (XYF and XYM). Labelled genes represent the top ten genes per sex-chromosome type by average log2-transformed fold change (log2FC) (log2FC > 0.1, pct.diff > 0, P < 0.05). f, Volcano plot of differential gene-expression analysis between ovary GPs and testes GPs at E16. Significant DEGs are indicated in red (ovaries) and blue (testes). Astroglial genes that are enriched in ovaries, and genes associated with stem-cell population maintenance and positive regulation of Notch signalling that are upregulated in testes, are highlighted in bold (log2FC > 0.1, pct.diff > 0, P < 0.05). e,f, Differential expression was assessed using the two-sided MAST test with Benjamini–Hochberg corrections for multiple comparisons. NS, not significant. g,h, Significantly enriched pathways upregulated by the XX chromosome or the XY chromosome in E16 (g) and E18 (h) GPs. i,j, Significantly enriched pathways upregulated by ovaries or testes in E16 (i) and E18 (j) GPs. c,d,g–j, Enrichment was assessed using a one-sided cumulative hypergeometric test with the g:SCS multiple-testing correction. Terms with adjusted P < 0.05 were considered significant. c,d,i,j, Pathways of interest are highlighted in red (ovaries mice) and blue (testes mice).
To ascertain the relative causative roles of sex hormones versus sex-chromosome complement, we performed scRNA-seq on microdissected caudal embryonic cerebella collected from litter-matched XX, XY−, XX-Sry and XY−-Sry embryos at E16 and E18, and integrated these data with the previously collected XX and XY-Sry scRNA-seq data (Fig. 4b and Supplementary Table 1). All clusters showed similar cell-type proportions across the four genotypes, with no genotype-specific clusters detected (Extended Data Fig. 9a–d). Pathway enrichment analysis across all four genotypes showed that at E16, GPs from mice with testes (XXM or XYM) were enriched for stem-cell population maintenance and negative regulation of stem-cell differentiation, compared with mice with ovaries (XXF and XYF) (Fig. 4c). At E18, mice with testes showed an enrichment of cell-proliferation-related pathways, whereas mice with ovaries exhibited an enrichment of gliogenesis and cell differentiation pathways (Fig. 4d), indicating that the presence of androgens suppresses the differentiation of GPs.
To disentangle the effects of sex chromosome and gonadal hormones on embryonic hindbrain GPs, we performed differential gene expression and pathway enrichment analysis at E16 and E18, comparing XX versus XY, as well as ovaries versus testes (Fig. 4e–j, Extended Data Fig. 9e,f and Supplementary Tables 7 and 8). XX versus XY comparisons highlighted the expected upregulation of X-linked (Xist and Tsix) and Y-linked (Ddx3y) genes (Fig. 4e and Extended Data Fig. 9e), along with metabolic pathway enrichment in XX GPs (Fig. 4g,h). At E16, GPs from mice with ovaries—independent of chromosomal sex—showed higher expression of astroglial genes (Aldoc and Aldh2) (Fig. 4f). Conversely, mice with testes showed upregulated expression of genes involved in stem-cell population maintenance (Hes1, Sox2, Kit, Bcl7b and Arid4a) and positive regulation of Notch signalling pathways (Hes1, Tgfb2, Kit and Pdcd10), as compared with mice with ovaries (Fig. 4f,i)—consistent with a model in which GPs from mice with ovaries occupy a later stage of the developmental hierarchy than do those from mice with testes. GPs from E18 mice with testes had upregulated expression of cell population proliferation pathways, compared with mice with ovaries (Fig. 4j and Extended Data Fig. 9f). The testes-upregulated pathways are crucial during the very early stages of hindbrain development to establish and maintain GP identity. Several testes-upregulated genes were also upregulated in male-biased PFA tumour subpopulations. Jun, a key proliferation-associated immediate early gene29, was among the top five highly expressed markers in male-biased GP-like tumours (Extended Data Fig. 2b). In addition, Sox2 and Hes1 showed strong regulatory network activity in male-biased cycling and GP-like PFA tumour cells (Fig. 1f). The shared sex-biased patterns of gene expression between hindbrain development and PFA-EPN support a model in which the male–female transcriptional differences in PFA-EPN reflect early embryonic programs shaped mainly by androgen exposure, rather than tumour-acquired changes. Our findings from the FCG model indicate that, during normal embryonic hindbrain development, androgens have a role in inhibiting gliogenic lineage differentiation. This androgen-driven reduction in differentiation provides a proximate explanation for the higher incidence and poorer prognosis observed in androgen-producing male individuals with PFA-EPN.
Androgens fuel PFA stemness and growth
Canonical androgen receptor (AR) signalling is initiated when testosterone or dihydrotestosterone (DHT) binds to the AR, triggering its nuclear translocation and transcriptional activity at androgen response elements. To evaluate AR activation in our system, we performed western blots on androgen-treated cell lines derived from patients with PFA-EPN (Supplementary Fig. 1). AR levels were minimal in vehicle controls but increased in testosterone- and DHT-treated male (Fig. 5a) and female (Fig. 5b) lines. Subcellular fractionation showed that AR was scarcely detectable in the cytoplasmic fraction, but was highly enriched in the nuclear fraction of both male PFA9 (Fig. 5c) and female PFA4 (Fig. 5d) cells after androgen treatment, consistent with the activation of AR signalling in androgen-treated PFA cells; normalized densities are shown in Extended Data Fig. 10a–f.
a,b, Western blots showing AR expression in PFA9 (a) and PFA4 (b) cells using whole-cell lysate. T, testosterone; Veh, vehicle. c,d, Western blots showing AR expression in PFA9 (c) and PFA4 (d) cells using subcellular protein fractionation extract. Cyt, cytoplasmic fraction; LMNB1, lamin B1; Nuc, nuclear fraction; WC, whole cell. a–d, Western blots shown are representative of n = 4 biological replicates with similar results. e, PFA-EPN cells were treated with 50 nM testosterone, oestradiol or progesterone for in vitro LDA to assess the frequency of colony-forming stem cells. n = 8 technical replicates per condition. f, PFA-EPN cell lines were cultured with testosterone for seven days. g, PFA-EPN cells were treated with 5 µM enzalutamide or MTX-23 for LDA. n = 6 technical replicates per condition. h, PFA-EPN cells were treated with 5 µM enzalutamide or MTX-23 for LDA. Cells were pre-treated with 50 nM testosterone before drug exposure. n = 8 technical replicates (control); n = 6 technical replicates (drug). e,g,h, Statistical comparisons were performed using a two-sided extreme limiting dilution analysis (ELDA) Chi-squared test. Solid lines represent maximum-likelihood estimates of clonogenic frequency from ELDA; dashed lines indicate 95% confidence intervals. i,j, PFA-EPN cell lines were pre-treated with 50 nM testosterone for seven days, followed by treatment with enzalutamide (i) or MTX-23 (j) for seven days. f,i,j, Cell counts are normalized to the vehicle control for each line. n = 2 biological replicates, each with 3 technical replicates. Data are mean ± s.d. Statistical comparisons were performed using a one-way ANOVA on replicates followed by a Tukey’s post-hoc test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; NS, not significant. Exact P values are provided in the Source Data.
To evaluate the influence of hormones on stemness potential, we performed limiting dilution analysis (LDA) using one male and one female PFA cell line cultured with 50 nM testosterone, oestradiol or progesterone (Fig. 5e). Testosterone treatment significantly increased clonogenicity relative to controls, indicating that androgen exposure enhances stem-like activity. By contrast, oestradiol and progesterone had no effect on clonogenicity. These findings show that androgen signalling promotes stemness in PFA-EPN.
To assess whether androgens are sufficient to promote the growth of PFA-EPN, we cultured cells from six PFA lines with testosterone, oestradiol or progesterone at varying concentrations for seven days. Both male-derived (PFA5, PFA7, PFA9 and PFA15) and female-derived (PFA4 and PFA6) cell lines exhibited significantly increased growth in response to testosterone, compared with vehicle controls (Fig. 5f). Maximum growth was observed at physiologically relevant testosterone levels (around 50 nM) (Extended Data Fig. 10g). By contrast, treatment with female sex hormones (oestradiol or progesterone) did not alter the growth of PFA cells compared with controls (Extended Data Fig. 10h–k). Similarly, PFA cells cultured with 50 nM testosterone or DHT showed significantly increased growth, compared with controls (Extended Data Fig. 10h,i). To determine whether this growth effect was specific to PFA-EPN, we performed the same experiments in supratentorial ependymoma (ST-EPN) and K27M-mutant diffuse midline glioma (DIPG) cell lines. Neither ST-EPN nor DIPG cells showed meaningful growth changes in response to testosterone, oestradiol or progesterone—with the exception of the DIPG004 cell line, which showed a very minor (3–5%) increase in cell growth in the presence of testosterone (Extended Data Fig. 11a–c). PFA-EPN tumours have significantly higher levels of AR than ST-EPN tumours do (Extended Data Fig. 10l), which could explain the increased growth of PFA-EPN in response to androgens. These findings support a model in which androgens, but not female sex hormones, specifically promote the growth of PFA cells.
Androgen blockade suppresses PFA stemness
To evaluate how AR inhibition affects PFA stemness, we performed LDAs on male (PFA9) and female (PFA4) PFA cell lines treated with the AR antagonist enzalutamide or the AR degrader MTX-23. Enzalutamide is a clinically approved pure AR antagonist that is used as a front-line therapy in prostate cancer and has also shown significant anti-tumour activity in glioblastoma models33,34,35. MTX-23 is a proteolysis-targeting chimera (PROTAC) that induces AR ubiquitination and degradation36,37. PFA cells were exposed to enzalutamide or MTX-23 (5 µM) for seven days, and both agents significantly reduced clonogenicity in PFA9 cells. In PFA4 cells, MTX-23 significantly decreased clonogenicity, and enzalutamide showed a strong trend towards reduction, compared with controls (Fig. 5g). To further model a hormonally active environment and analyse AR signalling directly, PFA cells were pre-treated with 50 nM testosterone for at least seven days before treatment with enzalutamide or MTX-23. Under these conditions, both inhibitors significantly reduced clonogenicity in male (PFA9) and female (PFA4) lines (Fig. 5h), demonstrating that AR blockade decreases stem-like activity in PFA-EPN.
To determine the effects of AR antagonists on the growth and maintenance of PFA-EPN, we treated six patient-derived PFA lines with either enzalutamide or MTX-23 for seven days at varying concentrations. Both inhibitors significantly reduced the growth of PFA cells (Extended Data Fig. 10m,n). By contrast, ST-EPN and DIPG cell lines showed no significant changes in growth after treatment, except for a modest reduction in DIPG007 cells at the highest enzalutamide dose of 20 µM (Extended Data Fig. 11d,e). To model a hormonally active environment and directly test AR dependency, we pre-treated PFA-EPN, ST-EPN and DIPG cells with 50 nM testosterone for at least seven days before treatment with enzalutamide or MTX-23. Under these conditions, PFA cell growth was again significantly reduced after enzalutamide or MTX-23 treatment (Fig. 5i,j), whereas ST-EPN and DIPG cells remained mostly unaffected, aside from a decrease in DIPG007 growth at the lowest enzalutamide dose of 2.5 µM (Extended Data Fig. 11f,g).
These data show that inhibition of the AR affects the fitness of PFA cells, but has minimal effects on other ependymoma or paediatric glioma cells. Our results are therefore compatible with a model in which androgens—which are normally present during embryonic hindbrain development—are sufficient to support the growth of and delay the differentiation of normal hindbrain embryonic GPs, a phenotype that is conserved in PFA-EPN (Extended Data Fig. 11h). This androgen-driven biology might underlie the higher incidence of PFA-EPN in male children, and their poorer prognosis. Given that there is a complete lack of effective chemotherapy for PFA-EPN, anti-androgen therapy could provide a much-needed new treatment approach in this patient population, and warrants testing in clinical trials.
Discussion
Male individuals have larger brains overall—and larger cerebella, specifically—than female individuals do, even after accounting for weight and height38,39,40. Sex differences are known to promote cell-autonomous as well as whole-organism phenotypes3,41,42,43, and influence the incidence and prognosis of various types of cancer44,45. Non-cell-autonomous sex-specific mechanisms that influence brain development, particularly involving the CNS microglia, are also well described46,47. Although the current study focuses on cell-autonomous mechanisms in PFA-EPN, it is possible that non-cell-autonomous mechanisms from the tumour microenvironment also have a role. Whereas most sex differences seem to be secondary to hormonal influences rather than occurring secondary to sex-chromosome dosage, phenotypes secondary to sex-chromosome dosage have also been described in the CNS3.
Over the course of human life, the hypothalamic–pituitary–gonadal (HPG) axis undergoes activation during three distinct periods: first, in fetal development; second, during the early postnatal window known as mini-puberty; and third, at the onset of puberty48,49. Male human fetuses start to produce testosterone around eight weeks after fertilization, with maximum testosterone levels between week 10 and week 20 (ref. 50). Female human fetuses are also exposed to low levels of androgens, produced by the fetal adrenal glands, as well as the maternal adrenal glands, ovaries and fat tissues50. The duration and hormonal patterns of mini-puberty differ markedly between the sexes. In male individuals, testosterone production peaks at around 1–3 months of age, and declines to prepubertal levels by 4–6 months as the HPG axis becomes quiescent. In female individuals, oestradiol levels rise and remain elevated for a longer period, in some cases persisting up to 12–24 months48. This neonatal surge of androgens in male babies is referred to as the neonatal surge. Androgens have been shown to increase the proliferation of dorsal glutamatergic progenitors in human forebrain organoids51. Notably, oestrogens and progestins are not known to have similar effects on the human developing brain51.
The high levels of in utero androgens in male fetuses could modulate the maturation of the glial lineage, offering a longer window for possible malignant transformation, and could therefore account for the increased incidence of PFA-EPN in male individuals. The neonatal surge of androgens in male infants overlaps with the timing of presentation of patients with PFA-EPN, which is usually in the first three years of life. Indeed, it seems likely that the neonatal surge of androgens in male babies promotes the growth and prevents the differentiation of PFA cells during the first year of life. The neonatal surge and/or other sources of ongoing androgen production could therefore explain the worse prognosis of male individuals with PFA-EPN. Androgens seem to be both sufficient and necessary for PFA growth, because PFA tumours grow better in the presence of androgens, and their growth is inhibited by AR blockade.
The phenotypic and teleological reasons behind the promotion of hindbrain growth by androgens are not clear. We note the profound effect of androgens on the incidence and prognosis of hindbrain PFA-EPN, but not the highly similar ST-EPN. Notably, PFA-EPNs—but not ST-EPNs—respond to androgens in vitro. The hindbrain is the brain centre for both suckling and swallowing in mammals, and increased or advanced control of suckling or swallowing could provide a nutritional advantage for male newborns over their female siblings52. Perinatal androgen signalling might favour early functional advantages in hindbrain development, but concomitantly predispose to malignant transformation in susceptible GPs, particularly in the hindbrain.
The current therapy for children with PFA-EPN is maximal safe surgery followed by local external beam radiation. Chemotherapy is not part of the standard of care at present, because, despite numerous randomized, prospective controlled trials, there is currently no evidence that chemotherapy increases PFS. The fact that very few somatic mutations are associated with PFA-EPN has hampered the development of any rational targeted therapies for this under-served patient group. Androgen blockade could provide a treatment avenue that fills this gap. A trial of androgen blockade merits consideration—using the caution appropriate in the context of a paediatric population—and could eventually lead to the development of an effective targeted therapy for infants with PFA-EPN.
Methods
Animal experiments
All animal work in this study was approved by the Animal Care Committee of The Centre for Phenogenomics (TCP) (AUP 25-0100H) and by the Institutional Animal Care and Use Committee of the Center for Comparative Medicine (CCM) at Baylor College of Medicine (AN-9134 and AN-9136). All mice housed at TCP were maintained under standard housing conditions with a 12-h light–12-h dark cycle (lights on at 07:00; lights off at 19:00), an ambient temperature of 21–23 °C and relative humidity of 40–60%. All mice housed at CCM were maintained under standard housing conditions with a 12-h light–12-h dark cycle (lights on at 06:00; lights off at 18:00), an ambient temperature of 20–22 °C and relative humidity of 30–70%. All mice had ad libitum access to food and water. Timed breeding of C57BL/6J mice was set up, and whole litters were collected from eight independent embryonic time points: E9, E10, E11, E12, E13, E14, E16 and E18. Sexes for all embryos were determined from tail clippings using PCR protocols. Tissues from two female and two male littermates were selected at each time point to undergo dissociation.
FCG experiments
The FCG mice were originally provided by J. Danska, and additional mice were purchased from The Jackson Laboratory (strain 039108). The strain was maintained on a C57BL/6J background. XY mice with testes (XY−-Sry) were bred with wild-type C57BL/6J females to generate the four types of offspring. Caudal cerebellum was collected from E16 and E18 embryos. Genotypes for all embryos were determined from tail clippings using PCR protocols. Cerebella from one XX embryo with ovaries (XX), one XX embryo with testes (XX-Sry), one XY embryo with ovaries (XY−) and one XY embryo with testes (XY−-Sry) were collected at each time point for tissue dissociation.
DNA extraction
DNA was extracted from mouse tissue using the QIAGEN DNeasy Blood and Tissue Kit (QIAGEN, 69506) according to the manufacturer’s instructions.
PCR reactions
Genomic DNA was amplified with the following primer pairs: SX_F (5′-GATGATTTGAGTGGAAATGTGAGGTA-3′), SX_R (5′-CTTATGTTTATAGGCATGCACCATGTA-3′), SRY_F (5′-AGCCCTACAGCCACATGATA-3′) and SRY_R (5′-GTCTTGCCTGTATGTGATGG-3′). X and Y chromosomes were distinguished by PCR as previously described53. In brief, SX_F and SX_R amplified the pseudoautosomal Sly and Xlr genes on the Y and X chromosome, respectively. The presence or absence of the Sry gene in FCG offspring was determined using the SRY_F and SRY_R primers. PCR reactions were performed in a final volume of 20 µl with 1.5 mM MgCl2, 200 µM dNTPs, 0.5 µM primers, 0.4 µl of Phire Hot Start II DNA Polymerase (Thermo Fisher Scientific, F-122L) and the following PCR parameters: initial denaturation at 98 °C for 30 s, 30 cycles with denaturation at 98 °C for 5 s, annealing at 60 °C for 5 s and extension at 72 °C for 5 s, followed by final extension at 72 °C for 1 min. PCR products were analysed on a 1.5% agarose gel with a 100-bp DNA ladder (FroggaBio, DM001-R500) and visualized with ethidium bromide.
Patient-derived PFA-EPN tissues
All PFA-EPN tissues used in this study were obtained with properly informed consent of patients. All experimental procedures were performed in accordance with the Research Ethics Boards (REB 1000055059) at The Hospital for Sick Children.
Tissue handling and dissociation
Fresh patient tumours were collected at the time of surgical resection. Tumour tissue was mechanically dissociated followed by collagenase-based enzymatic dissociation as previously described9. Whole hindbrain was dissected from E9, E10, E11 and E12 embryos; one incision was made between the boundary of the midbrain and hindbrain, and a second between the hindbrain and spinal cord. Whole cerebellum was dissected from E13 and E14 embryos. Caudal cerebellum was collected from E16 and E18 embryos; rostral structures were removed from the cerebellum. All embryonic mouse dissections were performed using Dumont fine forceps (Fine Science Tools, no. 5) under a Leica stereoscope. All mouse tissue was dissociated using the Papain Dissociation System (Worthington Biochemical Corporation, LK003153) according to the manufacturer’s instructions. Cells were filtered using a 40-µm strainer and resuspended in phosphate-buffered saline with 0.1% bovine serum albumin (BSA).
scRNA-seq library preparation and sequencing
Single-cell suspensions were assessed with a trypan blue (Wisent, 609-130-EL) count. We aimed to load 10,000–16,000 cells per sample using the Chromium Controller in combination with the Chromium Single Cell 3’ V3 and V3.1 Gel Bead and GEM Kits (10X Genomics). Individual cells were partitioned into gel beads-in-emulsion (GEMs), followed by reverse transcription of barcoded RNA and cDNA amplification. Individual single-cell libraries with indices and Illumina P5/P7 adapters were generated with the Chromium Single Cell 3’ Library kit and Chromium Multiplex kit. The libraries were sequenced on an Illumina Novaseq6000 or Illumina NovaSeq X Plus sequencer.
Primary cell culture
All samples used in this study were obtained with properly informed consent of patients. All experimental procedures were performed in accordance with the Research Ethics Boards at The Hospital for Sick Children. Patient-derived PFA-EPN (MDT-PFA4, MDT-PFA5, MDT-PFA6, MDT-PFA7, MDT-PFA9 and MDT-PFA15) and ST-EPN (ST1 and ST4) cell lines were established as previously described7. PFA-EPN and ST-EPN cell lines were cultured in EPN-C medium, which is defined as NeuroCult NS-A basal medium (human)(STEMCELL Technologies, 05750), supplemented with N-2 (Thermo Fisher Scientific, 17502048), B27 (Thermo Fisher Scientific, 12587010), 2 mM GlutaMAX (Thermo Fisher Scientific, 35050061), 100 U ml−1 penicillin (Thermo Fisher Scientific, 15240062), 100 µg ml−1 streptomycin (Thermo Fisher Scientific, 15240062), 0.25 µg ml−1 Gibco amphotericin B (Thermo Fisher Scientific, 15240062), 150 µg ml−1 BSA (Sigma-Aldrich A8412), 2 µg ml −1 heparin (Sigma-Aldrich, H3393), 10 ng ml−1 EGF (Sigma-Aldrich, E9644) and 10 ng ml−1 FGF (PeproTech, 100-18B). Laminin-coated Primaria plates (Corning, 353803) were prepared by treating plates with poly-l-ornithine (Sigma-Aldrich, P4957) for 30 min, followed by treatment with laminin (Sigma-Aldrich, P4957). Cells were seeded on laminin-coated plates, cultured in 1% hypoxia (1% oxygen, 94% nitrogen and 5% carbon dioxide), and the medium was replaced every three days.
DIPG cell lines (DIPG004 and DIPG007) were maintained in Matrigel Corning-coated plates with DIPG medium (equal parts of Neurobasal A and Dulbecco’s modified Eagle’s medium with F12 (DMEM/F12) supplemented with B27 (without vitamin A), HEPES buffer (10 mM), MEM sodium pyruvate solution (1 mM), MEM non-essential amino acids (1×), GlutaMAX-I supplement (1×), antibiotic–antimycotic (1×), heparin (2 μg ml−1), human-EGF (20 ng ml−1), human-bFGF (20 ng ml−1), PDGF-AA (10 ng ml−1) and PDGF-BB (10 ng ml−1)).
Hormone treatment experiments
Testosterone (Sigma-Aldrich, T1500), DHT (Sigma-Aldrich, A8380), progesterone (Sigma-Aldrich, P8783) and oestradiol (Sigma-Aldrich, E1024) were reconstituted in high-purity ethanol (VWR, TS61509-5000). All of the cell culture models, PFA-EPN, ST-EPN and DIPG cells were seeded (1 × 106 cells) in laminin-coated Primaria plates. Twenty-four hours after seeding, cells were treated with the indicated hormones and incubated for seven days. For those seven days, each cell line was treated with fresh medium replacement with exogenous hormone(s) every 48 h. Ethanol alone was used as vehicle control. After seven days of treatment, treated and control cells were lifted using accutase (Thermo Fisher Scientific, A1110501) and cell counts (proliferation) and viability were measured using a Countess 3 Automated Cell Counter (Thermo Fisher Scientific). Cell counts were normalized to the vehicle control for each sample. Each treatment group was assessed in two biological replicates, each with two or three technical replicates. A one-way ANOVA followed by a Tukey’s post-hoc test using GraphPad Prism (v.10.2.0) software was performed to assess differences in cell growth.
Western blots
PFA cells were cultured for 24 h with vehicle, 50 nM or 100 nM testosterone or 50 nM or 100 nM DHT to assess the nuclear response of ARs. Total cell lysates from three million cells were prepared using RIPA buffer with added protease inhibitors (Sigma-Aldrich, 11836170001). To obtain nuclear and cytoplasmic fractions, cell pellets from three million cells were resuspended in NE-PER Nuclear and Cytoplasmic Extraction Reagents (78833) as per the manufacturer’s protocol.
Protein concentration was measured using a Pierce BCA Protein Assay (Thermo Fisher Scientific, A55865). Fifty micrograms of whole-cell lysate or fractionated lysate were subjected to SDS–PAGE on a 7% polyacrylamide gel. The resulting protein was semi-dry transferred to a polyvinylidene difluoride (PVDF) membrane (Bio-Rad, 1704272) using the Trans-Blot Turbo Transfer System (Bio-Rad, 1704150), and blocked using 5% skimmed milk in TBST (20 mM Tris, 150 mM NaCl and 0.1% Tween 20, pH 7.5). Membranes were incubated with the following primary antibodies overnight: GAPDH (Abcam, ab9484, mouse, 1:600; Abcam, ab9485, rabbit, 1:2,500); lamin B1 (Abcam, ab16048, rabbit, 1:1,000); and AR (Cell Signaling Technology, D6F11, rabbit, 1:1,000). Membranes were then washed with TBST three times, incubated with anti-rabbit, HRP-linked antibody (Cell Signaling Technology, 7074S, goat, 1:3,000) or anti-mouse, HRP-linked antibody (Cell Signaling Technology, 96714S, goat, 1:40,000) and washed with TBST three times. Membranes were incubated with HRP substrate (Thermo Fisher Scientific, WP20005) for 1 min, before imaging on a Li-COR Odyssey Fc Imaging System.
Western blot images were quantified using ImageJ (v.1.54g; National Institutes of Health; NIH)54. Integrated density values were measured for all bands using a consistent region of interest (ROI) across samples. AR band density was normalized to the housekeeper band. Statistical significance was evaluated using the Wilcoxon test performed on four biological replicates. P < 0.05 was considered statistically significant.
AR inhibition
The PROTAC-based AR inhibitor MTX-23 (MedChemExpress, HY-148771) and enzalutamide (MedChemExpress, HY-70002) were dissolved in dimethyl sulfoxide (DMSO; Sigma-Aldrich, D8418), and DMSO was used as a vehicle control. PFA-EPN cells were seeded at a density of 3,000 cells per well in 96-well Primaria plates (Corning), because PFA-EPN is a slow-growing tumour and requires a higher initial density to establish growth, whereas ST-EPN and DIPG cells were seeded at 1,000 cells per well in the same plates. Twenty-four hours after seeding, cells were treated with enzalutamide or MTX-23 for seven days with varying concentrations. Medium with the indicated dose of MTX was replaced every three days. Cell viability was measured using either a Countess 3 Automated Cell Counter (Thermo Fisher Scientific) or the CyQUANT Direct Cell Proliferation Assay (Thermo Fisher Scientific, C7026), as per the manufacturer’s protocol at end-point. Cell counts were normalized to the vehicle control for each sample. Each treatment group was assessed in two biological replicates, each with three technical replicates. A one-way ANOVA followed by a Tukey’s post-hoc test using GraphPad Prism (v.10.2.0) software was performed to assess differences in cell growth.
Colony formation assay and LDAs
For colony formation assays, PFA cells were dissociated using accutase (Thermo Fisher Scientific, A1110501) and stained with DAPI (Sigma-Aldrich, D9542) to assess viability. Single viable cells were sorted into a 96-well plate format and cultured under hypoxic conditions (1% O2). Serial dilutions of PFA cells were performed, ranging from a seeding density of 1,024 cells per well to 32 cells per well. Six cell doses, each with at least six technical replicates, were tested. To evaluate the effects of hormone treatments, cells were plated in the presence of vehicle control or 50 nM of testosterone, oestradiol or progesterone. To assess the effects of AR antagonists, cells were plated with vehicle control or 5 µM enzalutamide or MTX-23. To examine the effects of AR antagonists in a hormonally active environment, cells were pre-treated with 50 nM testosterone for at least seven days, followed by plating in the presence of vehicle control or 5 µM enzalutamide or MDT-23. Plates were incubated for one to two weeks to allow for colony formation and were subsequently scored as ‘positive’ (colony present) or ‘negative’ (no colony). Media and hormone treatments were replenished every three days. Statistical analysis of limiting dilution data was performed using the ELDA55 R package statmod (v.1.5.0).
Survival analysis
A previously reported22 multicentre cohort of 599 PFA-EPNs was re-analysed to assess the effect of biological sex specifically. PFS was determined using the Kaplan–Meier method and P values were determined using the two-sided log-rank test. Univariable and multivariable Cox proportional hazard regression was used to estimate hazard ratios including 95% confidence intervals. All statistical analyses were performed using the R statistical environment (v.4.3.2), with R packages survival (v.3.5-7) and ggplot2 (v.3.4.4).
scRNA-seq analysis of human PFA-EPN data
Published PFA-EPN cohorts
In addition to our newly generated scRNA-seq PFA database, we added published scRNA-seq PFA 10X Genomics samples from EGAS00001003170 (ref. 9) and GSE125969 (ref. 23). The samples’ copy-number statuses and ependymoma subgroups were classified on the basis of methylation arrays.
Copy-number analysis
The copy-number status of samples in the scRNA-seq PFA cohort was determined using methylation arrays. EPIC and Infinium Human Methylation 450K BeadChip arrays were analysed and combined using the minfi (v.1.34.0) package, with conversion with combineArrays to 450K. The converted samples were combined with normal brain controls (GSE74193, samples from infants and children, 450K BeadChip array)56,57. Using conumee (v.1.18.0) the combined intensities were extracted and normalized on the basis of the controls with a linear regression model58. Segments were calculated with a conumee wrapper based on the DNAcopy package, using the following parameters: significance threshold of 0.001; 50,000 permutations; and a minimum of 5 markers for a changed segment (https://bioconductor.org/packages/release/bioc/html/DNAcopy.html). Methylation-derived copy-number calls were then compared to inferCNV (v.1.4.0)59 calls based on the single-cell data at the single-cell levels compared with reference clusters from the lymphoid cell types60.
Alignment and preprocessing
scRNA-seq samples from the PFA cohort were aligned to the human genome using CellRanger (v.3.1.0)61 to human reference hg19 (v.3.0.0) with default settings. CellRanger-filtered count matrices per sample were preprocessed individually with the Seurat pipeline and filtered further (nFeature_RNA > 400, percent.mt < 3/4 quartile+ 3 times interquartile, genes expressed > 3 cells) and doublets were identified by DoubletFinder (v.2.0.3)62. Filtered samples were then combined with the published cohort (GSE125969) with Seurat. The biological batch effect arises from inherent biological differences between individual PFA tumour samples. Our dataset consists of three distinct cohorts of PFA tumour samples. To account for batch effects within and across these cohorts, we grouped the PFA samples into three separate groups and applied Harmony to correct for batch-related variations within each group. Specifically, we provided two variables for Harmony’s batch correction: sample identity (to address differences between individual samples) and group identity (to correct for batch effects across the three cohorts). Given that we anticipate batch effects both at the individual sample level and across sample groups, we set θ = 1 for both variables to achieve a balanced correction. Single-cell data were normalized with the Seurat SCTransform (v.0.3.2)63 workflow and regularized negative binomial regression with mitochondrial percentage and difference between the G2M and S phase scores regressed out64,65. Principal components to use for nearest-neighbour analysis and UMAP dimension reduction were selected using a quantitative elbow plot approach (https://hbctraining.github.io/scRNA-seq/lessons/elbow_plot_metric.html).
Cell-type annotations
Clusters were identified in the integrated object with the default Seurat Louvain algorithm and cluster markers were identified with the MAST (v.1.25.2)66 test with a 0.25 log fold change threshold, 0.1% expression threshold and minimum 10% expression in cluster. Top markers were selected by log fold change. Clusters were annotated on the basis of published markers and matching to built-in SingleR (v.1.0.1)67 databases and the mouse glial lineage. Tumour and microenvironment separation was validated on the basis of inferCNV analysis of PFA samples with chromosome 1q gain, with lymphoid cells as a reference.
Tumour cell-type trajectory inference
To assist with annotations of tumour subgroups and their hierarchy from stem-like to differentiated, trajectory inference for tumour cells was performed with Monocle3 (v.1.3.1)68,69. The normalized data from Seurat were imported to Monocle3 for UMAP dimensional reduction and cell clustering with default parameters. Root cells (with three standard deviations over the median) were selected on the basis of the CancerSEA (v.0.1.0) database stemness signature score. Next, a principal graph was generated through the UMAP using the learn_graph function, depicting the path through development70. The graph was subsequently used to order cells along the developmental program as pseudotime.
Cell composition analysis
Sex differences in cluster proportions were analysed by dividing the number of cells in the cell type of interest by the total number of cells in the full sample or the sample tumour fraction. Next, a Student’s t-test was used to analyse differences in cell-type fraction between male and female samples.
Differential abundance analysis
Differential abundance between male and female PFA tumour cells was assessed using miloR71 (v1.2.0). A k-nearest neighbour (k-NN) graph was constructed with the buildGraph function from the first 30 Harmony dimensions (d = 30), connecting each cell to its 30 nearest neighbours (k = 30). Differential abundance testing was performed using the testNhoods function, with all other parameters set to default.
Differential expression and pathway analysis
Differential expression between male and female samples within clusters was performed with the MAST test with a log2 fold change threshold of 0.25 and a minimum expression threshold of 10%. To avoid samples with large cell counts, our sex-biased gene lists were further filtered for genes that also had a significant sex bias between samples in average expression across the tumour fraction (AverageExpression function from Seurat). Next, we performed pathway analysis with the gprofiler2 (v.0.2.1)72 package with a multiple-testing g:SCS threshold of 0.1. Average expression of the male-biased gene signature by cell was visualized with the Seurat ModuleScore function.
TF analysis
Raw counts from the combined PFA object were used as input for the single-cell regulatory network inference and clustering (SCENIC) Python pipeline (pySCENIC, v.0.10.3)73,74. GENIE3 was used to infer gene co-expression networks within each cell and combined with the human RcisTarget database included with SCENIC to identify TF targets among the genes. Next, TF networks were binarized as ‘on/1’ or ‘off/0’ with automatic AUCell thresholds. Top regulatory networks by cell type were selected on the basis of fold change after filtering for networks with at least 10% binary difference in expression between clusters. Sex-biased transcriptional networks were selected from the top 20 networks with the highest relative difference in activity by sex within clusters, then filtered for networks with significant sex bias between samples in TF average gene expression across the tumour fraction (AverageExpression function from Seurat). Average expression of the male-biased TF signature by cell was measured with the Seurat ModuleScore function and compared between samples across the tumour fraction with the AverageExpression function.
Microarray analysis and survival
Published datasets generated using Affymetrix HG-U133 Plus 2 chip microarrays (GSE125861 and GSE64415) were used for survival analysis6,23. Only primary PFA samples that received both radiation and gross total resection with full survival data were kept to account for variation in prognosis due to treatment. Analysis was performed as previously described23, with robust multichip average normalization and a pipeline with the affy (v.1.66.0), affyPLM (v.1.64.0) and simpleaffy (v.2.64.0) R packages and the hgu133plus2.db database75,76. For scoring the male-biased gene signatures, the R package jetset (v3.4.0) was used for selecting the optimal probes for each gene. Then, the average of the rma counts was taken for each signature, and samples were divided into low and high groups by the median value77. OS and PFS were correlated with signature expression by the Kaplan–Meier estimator and plotted with the survival (v.3.3-1) and survminer (v.0.4.9) R packages with default parameters (https://cran.r-project.org/web/packages/survminer/index.html).
Comparison of AR expression in bulk RNA-seq data
The Stand Up to Cancer bulk RNA-seq dataset (EGAD00001006046) was published previously7. Primary PFA-EPN and ST-EPN tissue samples were integrated and normalized with the DESeq2 pipeline, and log-transformed counts of AR were compared with a two-sided Student’s t-test.
scRNA-seq analysis of mouse data
Alignment of raw reads
CellRanger (10X Genomics) pipelines (v.3.0.2 and v.5.0.1) were used to process the raw sequencing data for the mouse developing hindbrain and the FCG data, respectively. The raw base call (BCL) files were demultiplexed into FASTQ files with default parameters. The FASTQ files for mouse developing hindbrain data and the FCG data were then aligned to the reference mouse genome mm10 (v.3.0.0) and mm10 (v.2020-A) (10X Genomics), respectively. The count function was used to generate raw gene–barcode count matrices. It is worth noting that, to eliminate batch effects introduced by the technical process, the male sample and the female sample from normal development at E16 and E18 were reprocessed using the same alignment pipeline as was used for the FCG dataset. Alignment quality control metrics are provided in Supplementary Table 1.
Quality control and normalization
Quality control was performed on individual samples before their integration and normalization. Low-quality and outlier cells were identified and excluded from the datasets. We considered low-quality cells as those with fewer than 500 genes expressed, and cells with mitochondrial gene-expression fractions greater than 3/4 quartile + 3 times interquartile. Low-abundance genes were also excluded from the datasets as genes expressed in fewer than ten cells. Outlier cells were identified and excluded as cells with number of genes and unique molecular identifier (UMI) counts greater than 3/4 quartile + 3 times interquartile and less than 1/4 quartile − 1.5 interquartile. Doublets were detected and filtered out using the R package DoubletFinder (v.2.0.3); the doublet rate used the multiplex rate from 10X Genomics. Clusters with high mitochondrial gene-expression fraction but low number of genes expressed and UMI counts were considered as low-quality populations and thus manually removed from further analysis. Individual samples were merged and normalized together using SCTransform (v.0.3.5) using the parameter variable.features.n = 3000 and regressing unwanted variance associated with mitochondrial and cell-cycle content.
Cell-cycle analysis
We used cell-cycle phase-specific annotations to characterize the cell-cycle status of each individual cell64. We assigned a score to each cell on the basis of its expression of S and G2/M phase markers using the CellCycleScoring function from the R package Seurat (v.4.3.0)65. These marker sets should be anticorrelated in their expression levels, and cells that do not express either are probably not actively cycling and are likely to be in G1 phase.
Clustering analysis, visualization and annotation
Clustering analysis was performed with the R package Seurat (v.4.3.0). Linear dimensionality reduction was performed using principal component analysis (PCA) to determine statistically significant principal components. We considered the possibility of a biological batch effect arising from inherent biological differences between individual mouse samples rather than technical variability. A data harmonization method known as Harmony (v.0.1.1)78 with parameter theta = 0 was applied to remove sample-specific biological differences, addressing variations specific to cell types and states. The clusters of cells were identified by a shared nearest neighbour (SNN) modularity optimization-based clustering algorithm from Seurat. We then visualized these clusters using UMAP embedding. Cell types were assigned by manual annotation using known cell-type marker genes and computation of DEGs using the FindAllMarkers function in the Seurat package. The non-default parameters of the FindAllMarkers method are MAST test, minimum log fold change threshold of 0.1 and genes that are detected in a minimum of 10% of cells. DEG markers for each cluster are provided in Supplementary Table 2.
Validation of cell-type annotation in scRNA-seq data
Cell-type predictions in the mouse developing hindbrain data were performed using CoRAL (v.4.0.1; https://github.com/fungenomics/CoRAL), an annotation pipeline in which an ensemble of reference-based annotation tools are used to compute a weighted consensus mapping score to label individual cells.
We used a prenatal mouse cerebellar developmental atlas9 as a reference (five developmental samples from E10 to E18; n = 34,382 cells), which we reprocessed and refined to obtain more granular labels as follows. For each sample, RNA libraries were scaled using Seurat v.4.3.065 to 10,000 UMIs per cell and log-normalized. UMI counts and mitochondrial content were regressed from normalized gene counts, and the residuals were z-scored gene-wise. Dimensionality reduction was performed using PCA on the top 2,000 most variable features, keeping the top 30 principal components as input for clustering with an SNN algorithm (resolution 1). This resulted in 119 clusters, which were labelled on the basis of age, cluster number and cell class information from the original publication9. Cell class was assigned on the basis of the proportion of the original cell type in the new cluster: if the proportion of one original cell type was greater than 75% in the new cluster, that label was used; otherwise, a label consisting of two prominent cell types was used. Twenty clusters were manually reannotated on the basis of marker-gene expression.
Eight different annotation tools were benchmarked and trained on this reference: ACTINN79, scClassify80, SciBet81, singleCellNet82, SingleR67, Symphony83, Seurat and Support Vector Machines. Benchmarking was performed by fivefold cross validation using the average of F1 scores across labels as readout. The predictions of each of these methods were combined into a weighted consensus score using the CAWPE algorithm84, in which the weights for each method are based on their individual benchmarking performance. The final label in the query dataset is assigned to the label with the maximum consensus score.
Pseudotime cell-state trajectory reconstruction
Pseudotime analysis was performed using the R package Monocle3 (v.1.3.1). The preprocessed and normalized data from Seurat were imported to Monocle3 for UMAP dimensional reduction and cell clustering with default parameters. A principal graph was generated through the UMAP using the learn_graph function, depicting the path through development. The graph was subsequently used to order cells along the developmental program as pseudotime using Hes3 expression greater than 1 in E9 cells at the start of the program. To recapitulate the developmental trajectory in the glial lineage, cell-state transition directions were inferred using the Python package CellRank (v.1.4.0)85. CellRank’s PseudotimeKernel function was used to compute directed transition probabilities on the basis of a k-NN graph and Monocle3’s pseudotime ordering. The compute_transition_matrix function was used to compute a cell–cell transition matrix with parameter threshold_scheme = ‘soft’, nu = 0.5.
Reconstruction of gene regulatory and TF networks
The activity of specific TFs in each cell type was inferred in the mouse glial lineage cells using the package pySCENIC (v.0.10.3), implemented in Python (v.3.8.2). Co-expression modules between TFs and putative target genes were estimated by GENIE3. We then performed cis-regulatory motif analysis using RcisTarget, and indirect targets lacking motif-binding sites were removed. The resulting regulatory module (regulon) activities in each cell were then scored and binarized with AUCell. Regulon activity was computed independently in male and female cells of the glial lineage and subsequently integrated on the basis of shared regulons for visualization. The top sex-biased regulons were ranked by z-scaled AUC score. Inferred regulons and their activity across clusters by sex in the dataset are provided in Supplementary Table 4.
Differential expression and pathway enrichment analysis
To perform functional enrichment analysis of the sex-biased gene list in the glial lineage, DEGs between male and female within clusters were identified using the FindMarkers function from Seurat (v.4.3.0), with the parameters MAST test, minimum log fold change threshold 0.1 and genes detected in a minimum of 10% of cells. To ensure that the sex-biased DEGs were not driven by unequal numbers of cells from males and females, an equal number of cells from both sexes was used. Specifically, we kept all of the cells for the sex that had fewer cells. For the sex with more cells, we randomly selected the same number of cells as that used in the sex with fewer cells. For the differential expression comparisons of sex hormones (mouse with ovary versus mouse with testis), sex chromosomes (XX versus XY) and four genotypes (XXF, XYF, XXM and XYM) in the FCG dataset, we used the same pipeline as that described for the developing glial lineage. However, genes that were detected in a minimum of 5% of cells were analysed with the FindMarkers function, with the aim of recovering genes that were highly expressed, but in few cells. The resulting DEGs (Supplementary Tables 5 and 7) were filtered to include only those with an average log2 fold change greater than 0.1, P < 0.05 and an absolute difference in the fraction of expressing cells between comparison groups greater than 0. The filtered DEGs were ranked by average log2 fold change and subsequently analysed using gprofiler2 (v.0.2.1) for pathway enrichment, with multiple-testing correction applied using the g:SCS algorithm (Supplementary Tables 6 and 8).
Integration of PFA tumour data with the single-cell developmental glial lineage
Human genes from the PFA tumour fraction were converted to mouse genes with the convert_human_to_mouse_symbols function from the nichenetr package86. Raw counts from shared genes in PFA tumour and mouse glial lineage were merged and normalized with SCTransform with regression of mitochondrial percentage and difference between the G2M and S phase scores. Cross-species integration was performed with the Seurat Fast integration using the reciprocal PCA (RPCA) workflow on normalized counts (https://satijalab.org/seurat/articles/integration_rpca.html). Cells were integrated in an unsupervised manner using the 3,000 most variable features. Our dataset includes a similar number of male and female tumour cells, ensuring that the selection of the 3,000 variable genes was not biased toward either sex. The primary batch effect in this integration arises owing to differences between the two species: the human PFA tumour dataset and the mouse developmental dataset. To correct for this batch effect, RPCA identifies anchors between the two datasets using the FindIntegrationAnchors() function. In this approach, one dataset is projected into the PCA space of the other, ensuring that the integration process respects mutual nearest-neighbour relationships across species. This mutual neighbourhood constraint helps to identify biologically relevant anchors while minimizing technical artefacts. A key parameter in this process is k.anchor, which defines the number of nearest neighbours considered when selecting anchors between datasets. Increasing k.anchor enhances the alignment strength by incorporating a broader set of neighbours during anchor selection. For our integration, we set k.anchor = 20 to optimize the alignment between the PFA tumour and developing mouse datasets, ensuring a more accurate cross-species comparison. Principal components to use for nearest-neighbour analysis and UMAP dimension reduction were selected using a quantitative elbow plot approach as before, after clustering with the Seurat Louvain algorithm. Clusters were annotated on the basis of their composition of previously annotated mouse and PFA tumour clusters. Markers were identified with the MAST test, 0.25 log fold change threshold, 0.1% expression threshold and minimum 10% expression in cluster Top markers were selected by log fold change. Sex differences in integration were analysed in a similar manner to differences in tumour composition by comparing ratios of integrated cell type in the tumour fraction between male and female samples. Pseudotime and trajectory analysis were performed using the same workflow as for the mouse glial lineage, with ordering initiated from E9 cells with SOX3 expression greater than 1, because HES3 was not detected in this dataset after human–mouse gene conversion.
Deconvolution of PFA tumours with the single-cell mouse developmental glial lineage
For BayesPrism (v.2.2.2) deconvolution, the tumour fraction in each sample was converted to pseudobulk with the Seurat AverageExpression function. Human genes were converted to mouse and raw average counts of shared genes were used for the analysis. A glial lineage reference was constructed by keeping only human and mouse shared genes in the glial lineage raw count matrix, then filtering them for Ribosomal, XY-linked and non-protein-coding genes. The top 300 markers for each glial cluster were identified with the MAST test with a log fold change threshold of 0.6 and at least 5% expression in cluster. Deconvolution was then performed with default BayesPrism parameters. For CHETAH (v.1.18.0) deconvolution, a reference was constructed from the full mouse glial lineage Seurat object and ribosomal genes were removed as suggested in the CHETAH workflow. PFA tumour cells were matched on the basis of 500 genes selected by CHETAH, with a 0.1 threshold and other parameters at default settings. Fractions of deconvoluted cell types and nodes were compared between male and female samples with a Student’s t-test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated in the current study have been deposited in the Gene Expression Omnibus (GEO), European Genome-phenome Archive (EGA) and CNGB Sequence Archive (https://db.cngb.org/cnsa/): scRNA-seq data of mouse developmental hindbrain (GSE262208 and GSE311731), scRNA-seq of the mouse FCG model (GSE262207 and GSE311730) and scRNA-seq of PFA tumour samples (EGAS50000000507 and CNP0005408). Previously published datasets analysed in this study are available in the EGA and GEO databases: PFA scRNA-seq cohorts (EGAS00001003170 and GSE125969), PFA Affymetrix HG-U133 Plus 2 chip microarray cohorts (GSE125861 and GSE64415) and PFA-EPN and ST-EPN bulk RNA-seq data (EGAD00001006046). Source data are provided with this paper.
Code availability
No custom code was used in this study. All analyses were performed using open-source algorithms, with their applications outlined in the Methods. Additional information on the use of these algorithms is available from the corresponding authors upon request.
References
Mack, S. C. et al. Epigenomic alterations define lethal CIMP-positive ependymomas of infancy. Nature 506, 445–450 (2014).
Ramaswamy, V. et al. Therapeutic impact of cytoreductive surgery and irradiation of posterior fossa ependymoma in the molecular era: a retrospective multicohort analysis. J. Clin. Oncol. 34, 2468–2477 (2016).
De Vries, G. J. et al. A model system for study of sex chromosome effects on sexually dimorphic neural and behavioral traits. J. Neurosci. 22, 9005–9014 (2002).
Pajtler, K. W. et al. The current consensus on the clinical management of intracranial ependymoma and its distinct molecular variants. Acta Neuropathol. 133, 5–12 (2017).
Saleh, A. H. et al. The biology of ependymomas and emerging novel therapies. Nat. Rev. Cancer 22, 208–222 (2022).
Pajtler, K. W. et al. Molecular classification of ependymal tumors across all CNS compartments, histopathological grades, and age groups. Cancer Cell 27, 728–743 (2015).
Michealraj, K. A. et al. Metabolic regulation of the epigenome drives lethal infantile ependymoma. Cell 181, 1329–1345 (2020).
Pajtler, K. W. et al. Molecular heterogeneity and CXorf67 alterations in posterior fossa group A (PFA) ependymomas. Acta Neuropathol. 136, 211–226 (2018).
Vladoiu, M. C. et al. Childhood cerebellar tumours mirror conserved fetal transcriptional programs. Nature 572, 67–73 (2019).
Taylor, M. D. et al. Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8, 323–335 (2005).
Johnson, R. A. et al. Cross-species genomics matches driver mutations and cell compartments to model ependymoma. Nature 466, 632–636 (2010).
Ament, S. A. et al. A single-cell genomic atlas for maturation of the human cerebellum during early childhood. Sci. Transl. Med. 15, eade1283 (2023).
Tiemeier, H. et al. Cerebellum development during childhood and adolescence: a longitudinal morphometric MRI study. Neuroimage 49, 63–70 (2010).
Xiao, T. et al. Hallmarks of sex bias in immuno-oncology: mechanisms and therapeutic implications. Nat. Rev. Cancer 24, 338–355 (2024).
Bayik, D. et al. Myeloid-derived suppressor cell subsets drive glioblastoma growth in a sex-specific manner. Cancer Discov. 10, 1210–1225 (2020).
Lee, J. et al. Sex-biased T-cell exhaustion drives differential immune responses in glioblastoma. Cancer Discov. 13, 2090–2105 (2023).
Kwon, H. et al. Androgen conspires with the CD8+ T cell exhaustion program and contributes to sex bias in cancer. Sci. Immunol. 7, eabq2630 (2022).
Haupt, S., Caramia, F., Klein, S. L., Rubin, J. B. & Haupt, Y. Sex disparities matter in cancer development and therapy. Nat. Rev. Cancer 21, 393–407 (2021).
Rubin, J. B. The spectrum of sex differences in cancer. Trends Cancer 8, 303–315 (2022).
Arnold, A. P., Klein, S. L., McCarthy, M. M. & Mogil, J. S. Male–female comparisons are powerful in biomedical research — don’t abandon them. Nature 629, 37–40 (2024).
Witt, H. et al. Delineation of two clinically and molecularly distinct subgroups of posterior fossa ependymoma. Cancer Cell 20, 143–157 (2011).
Baroni, L. V. et al. Ultra high-risk PFA ependymoma is characterized by loss of chromosome 6q. Neuro Oncol. 23, 1360–1370 (2021).
Gillen, A. E. et al. Single-cell RNA sequencing of childhood ependymoma reveals neoplastic cell subpopulations that impact molecular classification and etiology. Cell Rep. 32, 108023 (2020).
Yuan, H. et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 47, D900–D908 (2019).
Tapia-González, S., Muñoz, M. D., Cuartero, M. I. & Sánchez-Capelo, A. Smad3 is required for the survival of proliferative intermediate progenitor cells in the dentate gyrus of adult mice. Cell Commun. Signal. 11, 93 (2013).
Boudjadi, S., Chatterjee, B., Sun, W., Vemu, P. & Barr, F. G. The expression and function of PAX3 in development and disease. Gene 666, 145–157 (2018).
Fraser, J. et al. Common regulatory targets of NFIA, NFIX and NFIB during postnatal cerebellar development. Cerebellum 19, 89–101 (2020).
Kim, Y. K., Lee, H., Ismail, T., Kim, Y. & Lee, H. S. Dach1 regulates neural crest migration during embryonic development. Biochem. Biophys. Res. Commun. 527, 896–901 (2020).
Udou, T., Hachisuga, T., Tsujioka, H. & Kawarabayashi, T. The role of c-Jun protein in proliferation and apoptosis of the endometrium throughout the menstrual cycle. Gynecol. Obstet. Invest. 57, 121–126 (2004).
Götz, M., Stoykova, A. & Gruss, P. Pax6 controls radial glia differentiation in the cerebral cortex. Neuron 21, 1031–1044 (1998).
de Kanter, J. K., Lijnzaad, P., Candelli, T., Margaritis, T. & Holstege, F. C. P. CHETAH: a selective, hierarchical cell type identification method for single-cell RNA sequencing. Nucleic Acids Res. 47, e95 (2019).
Chu, T., Wang, Z., Pe’er, D. & Danko, C. G. Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat. Cancer 3, 505–517 (2022).
Zalcman, N. et al. Intracranial assessment of androgen receptor antagonists in mice bearing human glioblastoma implants. Int. J. Mol. Sci. 25, 332 (2024).
Liang, D. et al. Ferroptosis surveillance independent of GPX4 and differentially regulated by sex hormones. Cell 186, 2748–2764 (2023).
Bishop, J. L. et al. The master neural transcription factor BRN2 is an androgen receptor-suppressed driver of neuroendocrine differentiation in prostate cancer. Cancer Discov. 7, 54–71 (2017).
Lee, G. T. et al. Effects of MTX-23, a novel PROTAC of androgen receptor splice variant-7 and androgen receptor, on CRPC resistant to second-line antiandrogen therapy. Mol. Cancer Ther. 20, 490–499 (2021).
Hung, C.-L. et al. Targeting androgen receptor and the variants by an orally bioavailable Proteolysis Targeting Chimeras compound in castration resistant prostate cancer. EBioMedicine 90, 104500 (2023).
Ritchie, S. J. et al. Sex differences in the adult human brain: evidence from 5216 UK biobank participants. Cereb. Cortex 28, 2959–2975 (2018).
Giedd, J. N. et al. Quantitative magnetic resonance imaging of human brain development: ages 4-18. Cereb. Cortex 6, 551–559 (1996).
Raz, N., Gunning-Dixon, F., Head, D., Williamson, A. & Acker, J. D. Age and sex differences in the cerebellum and the ventral pons: a prospective MR study of healthy adults. Am. J. Neuroradiol. 22, 1161–1167 (2001).
Wallen, K. Hormonal influences on sexually differentiated behavior in nonhuman primates. Front. Neuroendocrinol. 26, 7–26 (2005).
McCarthy, M. M., Nugent, B. M. & Lenz, K. M. Neuroimmunology and neuroepigenetics in the establishment of sex differences in the brain. Nat. Rev. Neurosci. 18, 471–484 (2017).
Li, F. et al. Sex differences orchestrated by androgens at single-cell resolution. Nature 629, 193–200 (2024).
Clocchiatti, A., Cora, E., Zhang, Y. & Dotto, G. P. Sexual dimorphism in cancer. Nat. Rev. Cancer 16, 330–339 (2016).
Yang, W. et al. Sex differences in GBM revealed by analysis of patient imaging, transcriptome, and survival data. Sci. Transl. Med. 11, eaao5253 (2019).
Lenz, K. M., Nugent, B. M., Haliyur, R. & McCarthy, M. M. Microglia are essential to masculinization of brain and behavior. J. Neurosci. 33, 2761–2772 (2013).
Guneykaya, D. et al. Transcriptional and translational differences of microglia from male and female brains. Cell Rep. 24, 2773–2783 (2018).
Lanciotti, L., Cofini, M., Leonardi, A., Penta, L. & Esposito, S. Up-to-date review about minipuberty and overview on hypothalamic–pituitary–gonadal axis activation in fetal and neonatal life. Front. Endocrinol. 9, 410 (2018).
Rohayem, J., Alexander, E. C., Heger, S., Nordenström, A. & Howard, S. R. Mini-puberty, physiological and disordered: consequences, and potential for therapeutic replacement. Endocr. Rev. 45, 460–492 (2024).
Christine Knickmeyer, R. & Baron-Cohen, S. Fetal testosterone and sex differences. Early Hum. Dev. 82, 755–760 (2006).
Kelava, I., Chiaradia, I., Pellegrini, L., Kalinka, A. T. & Lancaster, M. A. Androgens increase excitatory neurogenic potential in human brain organoids. Nature 602, 112–116 (2022).
Moore, T. & Haig, D. Genomic imprinting in mammalian development: a parental tug-of-war. Trends Genet. 7, 45–49 (1991).
McFarlane, L., Truong, V., Palmer, J. S. & Wilhelm, D. Novel PCR assay for determining the genetic sex of mice. Sex. Dev. 7, 207–211 (2013).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Hu, Y. & Smyth, G. K. ELDA: Extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Jaffe, A. E. et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat. Neurosci. 19, 40–47 (2016).
Daenekas, B. et al. Conumee 2.0: enhanced copy-number variation analysis from DNA methylation arrays for humans and mice. Bioinformatics 40, btae029 (2024).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Kumar, M., Bowers, R. R. & Delaney, J. R. Single-cell analysis of copy-number alterations in serous ovarian cancer reveals substantial heterogeneity in both low- and high-grade tumors. Cell Cycle 19, 3154–3166 (2020).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Finak, G. et al. MAST: A flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2022).
Peterson, H., Kolberg, L., Raudvere, U., Kuzmin, I. & Vilo, J. gprofiler2—an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res. 9, 709 (2020).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
Heber, S. & Sick, B. Quality assessment of Affymetrix GeneChip data. OMICS 10, 358–368 (2006).
Wilson, C. L. & Miller, C. J. Simpleaffy: a BioConductor package for Affymetrix quality control and data analysis. Bioinformatics 21, 3683–3685 (2005).
Li, Q., Birkbak, N. J., Gyorffy, B., Szallasi, Z. & Eklund, A. C. Jetset: selecting the optimal microarray probe set to represent a gene. BMC Bioinformatics 12, 474 (2011).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Ma, F. & Pellegrini, M. ACTINN: automated identification of cell types in single cell RNA sequencing. Bioinformatics 36, 533–538 (2020).
Lin, Y. et al. scClassify: sample size estimation and multiscale classification of cells using single and multiple reference. Mol. Syst. Biol. 16, e9389 (2020).
Li, C. et al. SciBet as a portable and fast single cell type identifier. Nat. Commun. 11, 1818 (2020).
Tan, Y. & Cahan, P. SingleCellNet: a computational tool to classify single cell RNA-seq data across platforms and across species. Cell Syst. 9, 207–213 (2019).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
Andrade, A. F. et al. Immune landscape of oncohistone-mutant gliomas reveals diverse myeloid populations and tumor-promoting function. Nat. Commun. 15, 7769 (2024).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Acknowledgements
M.D.T. is a CPRIT Scholar in Cancer Research (CPRIT, RR220051) and is the Cyvia and Melvyn Wolff Chair of Pediatric Neuro-Oncology, Texas Children’s Cancer and Hematology Center. M.D.T. is supported by the NIH (R01NS106155, R01CA159859 and R01CA255369), the National Cancer Institute (NCI; 2P50CA127001-16), the Pediatric Brain Tumour Foundation, the Terry Fox Research Institute, the Canadian Institutes of Health Research, the Cure Search Foundation, the Matthew Larson Foundation, Meagan’s Walk, the SWIFTY Foundation, the Victory Foundation, the Ependymoma Research Foundation (ERF), the STOPbraintumors Foundation, the Brain Tumour Charity, Genome Canada, Genome BC, Genome Quebec, the Ontario Research Fund, Worldwide Cancer Research, the V Foundation for Cancer Research and the Ontario Institute for Cancer Research funded by the government of Ontario. M.D.T. was also supported by a Canadian Cancer Society Research Institute Impact grant, a Cancer Research UK Brain Tumour Award, a Stand Up To Cancer (SU2C) St Baldrick’s Pediatric Dream Team Translational Research grant (SU2C-AACR-DT1113) and SU2C Canada Cancer Stem Cell Dream Team research funding (SU2C-AACR-DT-19-15) provided by the government of Canada through Genome Canada and the Canadian Institutes of Health Research, with supplementary support from the Ontario Institute for Cancer Research through funding provided by the government of Ontario. SU2C is a program of the Entertainment Industry Foundation, administered by the American Association for Cancer Research. M.D.T. was previously a Garron Family Chair in Childhood Cancer Research at the Hospital for Sick Children and the University of Toronto. C.L.K. is supported by a Canadian Institutes of Health Research (CIHR) grant (PJT-190271) and the Fonds de Recherche du Québec-Santé (FRQS) salary award. J.Z. is supported by the Andrew McDonough B+ (Be Positive) Foundation. K.A.M. is supported by the Matthew Larson Foundation (IronMatt), the Stanley M. Marks Cancer Research Fund, the V Foundation for Cancer Research and a UPMC Hillman Cancer Center core grant (NCI, P30CA047904). Q.R.L. is supported by the NIH (R01NS140460). F.Y.L. is supported by the Chance for Hope Foundation. A.S. is supported by Baylor College of Medicine Comprehensive Cancer Training Program, CPRIT RP210027. Data were analysed on the HPC cluster managed by the Biostatistics and Informatics Shared Resource and supported by NIH S10 Shared Instrument grant S10-OD032185, NCI P30-CA125123 and institutional funds from the Dan L Duncan Comprehensive Cancer Center and Baylor College of Medicine. Data analyses were also enabled by compute and storage resources provided by the Digital Research Alliance of Canada. We acknowledge the Robert Connor Dawes Foundation for funding the EPCAM series of meetings and accelerating the pace of ependymoma research across the globe. We thank A. Pomada Villalbi and B. J. Pakuts for graphic design and figure editing, and S. E. Archer for technical writing assistance.
Author information
Authors and Affiliations
Contributions
M.D.T. led the study. C.L.K. led and supervised the bioinformatics analyses. K.A.M. contributed to and provided advice on cell line validation experiments. J.Z., W.O. and A.R. designed, performed and analysed most of the experiments. R.D.G., R.L.G., H.N., K.K., C.R., D.P., E.Y.W., N.A., K.M. and H. Whetstone contributed to experimental, statistical and bioinformatics analysis and/or interpretation. P.B., A.S., X.W., M.C.V. and W.O. contributed to tissue isolation of mouse embryos and scRNA-seq library preparation. V.S.-M.L., F.G.-S., M.M.K.-S., B.L. and D.S. contributed to western blot validations. N.Y., D.M., K.B.N.R., R.G.R.M., R.N.C. and R.Y. contributed to cell line validations. S.P., A.W.E., L.Z.H., C.P., S.H., H.M. and H. Lakkis contributed to bioinformatics analysis. H. Su, N.M., N.H., Y.W., H. Wang, T.D. and J.P. contributed to mouse experiments. Z.L., M.X., L.K.D., H. Li, W.Z. and Q.R.L. contributed to scRNA-seq preparation of PFA tumours. R.S. contributed to copy-number analysis. K.N. optimized 10X Genomics scRNA-seq protocols. J.L. and J.D.L. provided advice on tumour sex differences. S.K.R., Y.P., Y.T., F.Y.L., G.A., M.G., R.M.F., M.C., B.W., Y.L., D.D.D.C., Z.Z., Y.M., W.H., J.K., D.G.T., T.J., H. Liu, H. Suzuki, J.N.R., L.D.S. and N.J. provided advice and helped design the study. F.N. and C.D. provided advice, reagents and equipment. C.E., C.-H.G.L., S.V., P.H.S. and A.D. provided patient tumour material and helped design the study. V.R. contributed to clinical analysis of patient samples. A.B.H. provided advice and contributed to data analysis and interpretation. J.Z., K.A.M., C.L.K. and M.D.T. jointly supervised the project. J.Z., W.O., A.R., K.A.M., C.L.K. and M.D.T. prepared the figures and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sex differences in prognosis and cell populations in PFA-EPN.
a, Progression-free Kaplan–Meier survival plot for all patients with primary PFA-EPN (n = 599), chromosome 1q balanced patients (n = 484), and chromosome 1q gain patients (n = 114). Statistical significance was calculated using two-sided log-rank test, shaded areas indicate 95% confidence intervals. b, UMAP visualization of broad tumour and microenvironment cell types in the PFA single-cell cohort. c, Top 3 markers of broad cell types by average log2FC. DEGs were assessed using the two-sided MAST test with Benjamini–Hochberg corrections for multiple comparisons (log2FC > 0.25, min.prc.exp > 0.1, adjusted P < 0.05). d, Top 3 most active regulatory networks of broad cell types from SCENIC analysis. e, Copy-number variation analysis by cell in tumour and microenvironment clusters with lymphoid cells as a reference within PFA samples with chromosome 1q gain. f, PFA sample composition of broad cell types. g, Fraction of broad cell types in PFA samples by sex.
Extended Data Fig. 2 PFA tumours from male patients are more stem-like and have a higher fraction of progenitor-like cells.
a, UMAP visualization of CancerSEA stemness module score expression across the tumour fraction split by sex. b, Top 5 cluster markers of tumour cell types by average log2FC. c, Top 20 GP-like tumour markers by average log2FC. b,c, DEGs were assessed using the two-sided MAST test with Benjamini–Hochberg corrections for multiple comparisons (log2FC > 0.25, min.prc.exp > 0.1, adjusted P < 0.05). d, Monocle3-inferred pseudotime trajectory of tumour cells across different tumour cell clusters. e, UMAP visualization of tumour pseudotime trajectory across PFA tumour cells using Monocle3. f, Beeswarm plot of the distribution of log fold change across sex in neighbourhoods containing cells from different cell-type clusters. Differential abundance neighbourhoods at FDR 10% are coloured. g, Tumour cell-type distribution between PFA tumour cells by sex. h, Tumour cell-type fraction within the PFA sample tumour cells by sex. i, Tumour cell-type fraction within the PFA sample tumour and microenvironment cells by sex. h,i, Female (n = 8) and male (n = 18). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles, and whiskers mark 1.5× the IQR. P values were calculated using a two-sided Student’s t-test. *P < 0.05; NS, not significant. Exact P values are provided in the Source Data.
Extended Data Fig. 3 Progenitor-like tumour cells in male PFA-EPNs upregulate transcriptional regulatory networks retained from early embryonic development and are associated with worse survival outcomes.
a, Sex-biased regulons in GP-like tumours. Regulon activity was computed independently in male and female PFA cells and subsequently integrated based on shared regulons for visualization. Male-biased regulons are shown in blue, the top 20 female-biased regulons are shown in red, and regulons with comparable activity between sexes that are relevant to GPs are highlighted in black. b, Genes and regulatory network master regulators identified by single-cell differential expression and regulatory network analysis showing significant male-biased pseudobulk expression per sample. Female (n = 8) and male (n = 18). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles, and whiskers mark 1.5× the IQR. P values were calculated using a two-sided Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001. Exact P values are provided in the Source Data. c, Dot plot visualization of male-biased gene expression in the GP-like tumour cluster by sex. F, female; M, male. d, UMAP visualization of average expression of the male-biased GP-like tumour gene signature in PFA tumours from females versus males. e, Top 5 pathways enriched in the male-biased GP-like tumour gene signature. Enrichment was assessed using a one-sided cumulative hypergeometric test with g:SCS multiple-testing correction. Terms with adjusted P < 0.05 were considered significant. f, Dot plot visualization of male-biased regulatory networks (SCENIC) in the GP-like tumour cluster by sex. F, female; M, male. g, UMAP visualization of average expression of the male-biased GP-like tumour TF signature in tumours from females versus males. h,i, Correlation of male-biased gene (h) and TF (i) signature expression in the GP-like cluster with tumour cell-type fractions in male patients with PFA-EPN from scRNA-seq cohort (n = 18). j,k, Kaplan–Meier curves of OS (j) and PFS (k) in male patients with PFA-EPN (n = 39), stratified by expression of the GP-like tumour male-biased gene signature in a microarray cohort treated with standard therapy (complete tumour resection and radiation). Statistical analyses were performed using two-sided log-rank test, shaded areas indicate 95% confidence intervals.
Extended Data Fig. 4 Quality control of scRNA-seq data and sampling overview of the sex-balanced developmental transcriptional atlas of the embryonic mouse hindbrain.
a, Number of cells per replicate organized by embryonic time stage. b, Scatter plot of scaled expression of the female-specific non-coding RNA Xist and the Y-chromosome-specific gene Eif2s3y, which shows the sex assignment for female (red) and male (blue) cells. c, UMAP visualization of cell-cycle phase. d, Dot plot depicting marker genes for all cell types identified. e, Cell-type prediction using CoRAL with a previously published scRNA-seq dataset of the developing mouse cerebellum9 as reference.
Extended Data Fig. 5 Number and proportion of cell types by sex in the sex-balanced developmental transcriptional atlas of the embryonic mouse hindbrain.
a, Stacked bar plot showing cell-type composition by replicate. Each bar represents an individual biological replicate, with the cell-type fraction shown as stacked segments. Female samples are displayed in red and male samples in blue. Dark separator lines within each bar distinguish individual replicates. b, Bar plot showing the total number of male (cyan) and female (pink) cells for each annotated cell type. c, Fraction of each cell type across developmental time in male and female combined dataset (upper panel), in males only (bottom left panel) and females only (bottom right panel).
Extended Data Fig. 6 Sex differences in the embryonic mouse hindbrain glial lineage.
a, Dot plot depicting marker genes for all cell types identified in the glial lineage. b, Cell-type composition across developmental time points by sex in the hindbrain glial lineage. Stacked area shows the fractional abundance of each cell type at embryonic days E9 to E18, separated by sex. Each colour indicates a distinct cell type within either females or males. F, female; M, male. c, Cell fraction of each replicate by cell type. Each point represents a replicate; females are in red, males are in cyan. Lines represent the average of each sex. F, female; M, male. d, Top 20 sex-biased regulons in each cell type from the glial lineage, ranked by Z-scaled AUC score. F, female; M, male. e, Significantly enriched pathways per cluster by sex based on DEGs. Enrichment was assessed using a one-sided cumulative hypergeometric test with g:SCS multiple-testing correction. Terms with adjusted P < 0.05 were considered significant.
Extended Data Fig. 7 Unsupervised integration of PFA tumour cells and the mouse glial lineage.
a, UMAP visualization of PFA tumour clusters following RPCA integration with glial lineage. b, UMAP visualization of glial lineage clusters following RPCA integration with PFA tumours. c, Distribution of original PFA tumour and mouse glial lineage cell types across integrated clusters. d, Top 3 marker genes per cluster by average log2FC in the integrated dataset. DEGs were assessed using the two-sided MAST test with Benjamini–Hochberg corrections for multiple comparisons (log2FC > 0.25, min.pct > 0.1, adjusted P < 0.05). e, Ridge plot visualization of integrated clusters across glial developmental pseudotime trajectory (Monocle 3). f, Pie visualization of glial lineage and PFA tumour proportions in integrated clusters. g, Pie visualization of glial lineage and PFA tumour proportions in integrated clusters by sex. h, Pie visualization of glial lineage cell-type proportions in integrated clusters. i, Pie visualization of PFA tumour cell-type proportions in integrated clusters.
Extended Data Fig. 8 Sex differences in PFA tumour fractions on the basis of hierarchical classification, deconvolution and integration with the developing glial lineage.
a, Integrated cell-type proportion in PFA tumour cells per sample by sex. b, Fraction of CHETAH matched cells across glial cell types and intermediate nodes for PFA tumour cells by sample per sex. c, BayesPrism deconvoluted glial lineage cell-type proportion in PFA tumour fraction pseudobulk per sample by sex. a–c, Female (n = 8) and male (n = 18). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles, and whiskers mark 1.5× the IQR. P values were calculated using a two-sided Student’s t-test. *P < 0.05, **P < 0.01; NS, not significant. Exact P values are provided in the Source Data.
Extended Data Fig. 9 Effects of sex chromosomes and hormones on cell-type composition and transcriptional profiles in embryonic mouse cerebellar cells of the FCG dataset.
a,b, Stacked bar plot depicting the number of cells per cell type and cell-type fraction by sex chromosomes, sex hormones and FCG at E16 (a) or E18 (b). Each bar represents an individual biological replicate, with the cell-type fraction shown as stacked segments. Dark separator lines within each bar distinguish individual replicates. c,d, Dot plot depicting marker genes for all cell types identified for E16 (c) or E18 (d). e, Volcano plot of the differential gene expression analysis between XX and XY E18 GPs. Significant DEGs are indicated in red (XXF & XXM) and blue (XYF & XYM). Labelled genes represent the top 10 genes per sex-chromosome type by average log2FC. f, Volcano plot of the differential gene expression analysis between ovaries and testes E18 GPs. Significant DEGs are indicated in red (ovaries) and blue (testes). Genes involved in response to hormone and peptide hormone pathways that are upregulated in testes are highlighted in bold. e,f, Differential expression was assessed using the two-sided MAST test with Benjamini–Hochberg corrections for multiple comparisons (log2FC > 0.1, pct.diff > 0, P < 0.05).
Extended Data Fig. 10 Androgens promote stemness and growth of PFA cells in vitro.
a,b, Quantified western blots of the female (F) PFA4 (a) and male (M) PFA9 (b) cell lines showing AR expression normalized to vehicle in whole-cell lysates. c,d, Quantified western blots of the female (F) PFA4 (c) and male (M) PFA9 (d) cell lines showing AR expression normalized to vehicle in subcellular protein fractionation extracts. e,f, Box plot showing normalized AR expression relative to vehicle in the female (F) PFA4 (e) and male (M) PFA9 (f) cell lines, measured in cytoplasmic and nuclear fractions across four biological replicates. n = 4 per group. Data are presented as mean ± s.e.m. P values were calculated using a two-sided Wilcoxon test. *P < 0.05; NS, not significant. Exact P values are provided in the Source Data. g, Images of cell culture confluency of (F) PFA4, (M) PFA5, and (M) PFA9 after being cultured with vehicle control or testosterone at different concentrations for 7 days. Scale bars, 100 μm. h,i, PFA male cell lines (PFA5 (h) and PFA9 (i)) were cultured with 50 nM or 100 nM of sex hormones (testosterone, DHT, progesterone or oestradiol) for 7 days. Cell counts were normalized to vehicle control for each line. n = 2 biological replicates, each with 2 technical replicates. j,k, Male PFA cell lines (PFA5, PFA7, PFA9, and PFA15) and female PFA cell lines (PFA4, PFA6) were cultured with oestradiol (j) or progesterone (k) for 7 days at concentrations ranging from 10 nM to 1 uM. Cell counts were normalized to vehicle control for each line. l, Box plot of expression levels of AR in ST ZFTA fusion-EPN or PFA-EPN bulk RNA-seq patient samples. ST ZFTA-EPN (n = 4), PFA-EPN (n = 14). In box plots, the centre line denotes the median, box hinges indicate the 25th and 75th percentiles, and whiskers mark 1.5× the IQR. P values were calculated using a two-sided Student’s t-test. m,n, Male PFA cell lines (PFA5, PFA7, PFA9, and PFA15) and female PFA cell lines (PFA4, PFA6) were cultured with the AR inhibitor enzalutamide (m) or MTX-23 (n) at concentrations ranging from 2.5 µM to 20 µM for 7 days. Cell counts were normalized to the vehicle control for each line. j,k,m,n, n = 2 biological replicates, each with 3 technical replicates. h–n, Data are displayed as mean ± s.d. Statistical comparisons were performed using a one-way ANOVA on replicates followed by a Tukey’s post-hoc test; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; NS, not significant. Exact P values are provided in the Source Data.
Extended Data Fig. 11 Androgens have minimal effects on cell growth in vitro in ST-EPN and DIPG compared with PFA-EPN.
a–c, ST-EPN cell lines (ST1 and ST4) and DIPG cell lines (DIPG004 and DIPG007) were cultured with testosterone (a), oestradiol (b) or progesterone (c) for 7 days at concentrations ranging from 10 nM to 1 uM. Cell counts were normalized to vehicle control for each line. d,e, ST-EPN cell lines (ST1 and ST4) and DIPG cell lines (DIPG004 and DIPG007) were cultured with the AR inhibitor enzalutamide (d) or MTX-23 (e) at concentrations ranging from 2.5 µM to 20 µM for 7 days. Cell counts were normalized to the vehicle control for each line. f,g, ST-EPN cell lines (ST1 and ST4) and DIPG cell lines (DIPG004 and DIPG007) were pre-treated with 50 nM testosterone for 7 days, followed by treatment with the AR inhibitor enzalutamide (f) or MTX-23 (g) at concentrations ranging from 2.5 µM to 20 µM for 7 days. Cell counts were normalized to the vehicle control for each line. a–g, n = 2 biological replicates, each with 3 technical replicates. Data are displayed as mean ± s.d. Statistical comparisons were performed using a one-way ANOVA on replicates followed by a Tukey’s post-hoc test; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; NS, not significant. Exact P values are provided in the Source Data. h, Schematic of proposed cellular hierarchy for PFA-EPNs. Created in BioRender; Pomada Villalbi, A. https://BioRender.com/y8qtbvf (2025).
Supplementary information
Supplementary Figure 1 (download PDF )
Uncropped images of western blots for Fig. 5a–d
Supplementary Tables (download ZIP )
Supplementary Tables 1–8
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, J., Ong, W., Rasnitsyn, A. et al. Androgen activity in the male embryonic hindbrain drives lethal PFA ependymoma. Nature (2026). https://doi.org/10.1038/s41586-026-10264-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10264-6