Abstract
F-Actin cytoskeleton remodeling is vital for cell migration, organ development and immune responses. The small GTPase CDC42, a key regulator of F-actin dynamics, cycles between inactive GDP- and active GTP-bound states. However, mechanisms governing CDC42 turnover and their biological significance remain unclear. Here we show that KLHL23-mediated polyubiquitylation of CDC42•GTP and RhoGDI-mediated sequestration of CDC42•GDP spatiotemporally co-inactivate CDC42, preserving membrane dynamics and homeostasis during migration. KLHL23–Cul3 acts as the E3 ligase for CDC42 degradation, with KLHL23 and RhoGDI competing for CDC42’s switch II region, enhancing selectivity toward CDC42•GTP and CDC42•GDP, respectively. KLHL23 depletion disrupts membrane homeostasis, inducing excessive protrusions and promoting metastasis. Notably, the CDC42-Y64C germline variant in Takenouchi–Kosaki Syndrome escapes KLHL23-mediated degradation. Fluorescence resonance energy transfer assays reveal that KLHL23 and RhoGDI coordinately inactivate CDC42 in a spatiotemporal manner. These findings highlight the biological and clinical relevance of the KLHL23/RhoGDI–CDC42 axis, presenting new avenues for therapeutic exploration.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data supporting the findings of this study are included within the main article, Extended Data figures, Supplementary Information, Supplementary Table and Supplementary Videos. Source data are provided with this paper.
References
Garlick, E., Thomas, S. G. & Owen, D. M. Super-resolution imaging approaches for quantifying F-actin in immune cells. Front. Cell Dev. Biol. 9, 676066 (2021).
Mercaldo, V. et al. Altered striatal actin dynamics drives behavioral inflexibility in a mouse model of fragile X syndrome. Neuron 111, 1760–1775 (2023).
Yamada, S., Mizukoshi, T., Tokunaga, A. & Sakakibara, S. I. Inka2, a novel Pak4 inhibitor, regulates actin dynamics in neuronal development. PLoS Genet. 18, e1010438 (2022).
Nunes de Almeida, F., Walther, R. F., Pressé, M. T., Vlassaks, E. & Pichaud, F. CDC42 defines apical identity and regulates epithelial morphogenesis by promoting apical recruitment of PAR6–aPKC and Crumbs. Development 146, dev175497 (2019).
Haga, R. B. & Ridley, A. J. Rho GTPases: regulation and roles in cancer cell biology. Small GTPases 7, 207–221 (2016).
Endo, M. & Cerione, R. A. The brain-specific splice variant of the CDC42 GTPase works together with the kinase ACK to downregulate the EGF receptor in promoting neurogenesis. J. Biol. Chem. 298, 102564 (2022).
Zhang, Z., Liu, M. & Zheng, Y. Role of Rho GTPases in stem cell regulation. Biochem. Soc. Trans. 49, 2941–2955 (2021).
Amoah, A. et al. Aging of human hematopoietic stem cells is linked to changes in CDC42 activity. Haematologica 107, 393–402 (2022).
Melendez, J. et al. CDC42 coordinates proliferation, polarity, migration, and differentiation of small intestinal epithelial cells in mice. Gastroenterology 145, 808–819 (2013).
Murphy, N. P., Binti Ahmad Mokhtar, A. M., Mott, H. R. & Owen, D. Molecular subversion of CDC42 signalling in cancer. Biochem. Soc. Trans. 49, 1425–1442 (2021).
Coppola, S. et al. Mutations at the C-terminus of CDC42 cause distinct hematopoietic and autoinflammatory disorders. J. Allergy Clin. Immunol. 150, 223–228 (2022).
Gernez, Y. et al. Severe autoinflammation in 4 patients with C-terminal variants in cell division control protein 42 homolog (CDC42) successfully treated with IL-1β inhibition. J. Allergy Clin. Immunol. 144, 1122–1125 (2019).
Lam, M. T. et al. A novel disorder involving dyshematopoiesis, inflammation, and HLH due to aberrant CDC42 function. J. Exp. Med. 216, 2778–2799 (2019).
Martinelli, S. et al. Functional dysregulation of CDC42 causes diverse developmental phenotypes. Am. J. Hum. Genet. 102, 309–320 (2018).
Endo, M., Druso, J. E. & Cerione, R. A. The two splice variant forms of Cdc42 exert distinct and essential functions in neurogenesis. J. Biol. Chem. 295, 4498–4512 (2020).
Takenouchi, T., Okamoto, N., Ida, S., Uehara, T. & Kosaki, K. Further evidence of a mutation in CDC42 as a cause of a recognizable syndromic form of thrombocytopenia. Am. J. Med. Genet. A 170a, 852–855 (2016).
Hamada, N. et al. Neuropathophysiological significance of the c.1449T > C/p.(Tyr64Cys) mutation in the CDC42 gene responsible for Takenouchi–Kosaki syndrome. Biochem. Biophys. Res. Commun. 529, 1033–1037 (2020).
Motokawa, M. et al. A hot-spot mutation in CDC42 (p.Tyr64Cys) and novel phenotypes in the third patient with Takenouchi–Kosaki syndrome. J. Hum. Genet. 63, 387–390 (2018).
Uehara, T. et al. Pathogenetic basis of Takenouchi–Kosaki syndrome: electron microscopy study using platelets in patients and functional studies in a Caenorhabditis elegans model. Sci. Rep. 9, 4418 (2019).
Ishikawa, K., Uchiyama, T., Kaname, T., Kawai, T. & Ishiguro, A. Autoimmune hemolytic anemia associated with Takenouchi–Kosaki syndrome. Pediatr. Int. 63, 1528–1530 (2021).
Hart, M. J., Eva, A., Evans, T., Aaronson, S. A. & Cerione, R. A. Catalysis of guanine nucleotide exchange on the CDC42Hs protein by the dbl oncogene product. Nature 354, 311–314 (1991).
Hoffman, G. R., Nassar, N. & Cerione, R. A. Structure of the Rho family GTP-binding protein CDC42 in complex with the multifunctional regulator RhoGDI. Cell 100, 345–356 (2000).
Johnson, J. L., Erickson, J. W. & Cerione, R. A. New insights into how the Rho guanine nucleotide dissociation inhibitor regulates the interaction of CDC42 with membranes. J. Biol. Chem. 284, 23860–23871 (2009).
Squair, D. R. & Virdee, S. A new dawn beyond lysine ubiquitination. Nat. Chem. Biol. 18, 802–811 (2022).
Hodge, R. G. & Ridley, A. J. Regulating Rho GTPases and their regulators. Nat. Rev. Mol. Cell Biol. 17, 496–510 (2016).
Murali, A. et al. Ubiquitin-dependent regulation of CDC42 by XIAP. Cell Death Dis. 8, e2900 (2017).
Oberoi, T. K. et al. IAPs regulate the plasticity of cell migration by directly targeting RAC1 for degradation. EMBO J. 31, 14–28 (2012).
Asmar, A. J. et al. A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development. Nat. Commun. 14, 4499 (2023).
Peng, J. M. et al. Actin cytoskeleton remodeling drives epithelial-mesenchymal transition for hepatoma invasion and metastasis in mice. Hepatology 67, 2226–2243 (2018).
Julio, A. R. et al. Delineating cysteine-reactive compound modulation of cellular proteostasis processes. Nat. Chem. Biol. 21, 693–705 (2024).
Békés, M., Langley, D. R. & Crews, C. M. PROTAC targeted protein degraders: the past is prologue. Nat. Rev. Drug Discov. 21, 181–200 (2022).
Martin, K. et al. Spatio-temporal co-ordination of RhoA, RAC1 and CDC42 activation during prototypical edge protrusion and retraction dynamics. Sci. Rep. 6, 21901 (2016).
Xu, L. et al. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature 425, 316–321 (2003).
Furukawa, M., He, Y. J., Borchers, C. & Xiong, Y. Targeting of protein ubiquitination by BTB–Cullin 3–Roc1 ubiquitin ligases. Nat. Cell Biol. 5, 1001–1007 (2003).
Harper, J. W. & Schulman, B. A. Cullin-RING ubiquitin ligase regulatory circuits: a quarter century beyond the F-box hypothesis. Annu. Rev. Biochem. 90, 403–429 (2021).
Dhanoa, B. S., Cogliati, T., Satish, A. G., Bruford, E. A. & Friedman, J. S. Update on the Kelch-like (KLHL) gene family. Hum. Genomics 7, 13 (2013).
Kobayashi, A. et al. Oxidative stress sensor KEAP1 functions as an adaptor for CUL3-based E3 ligase to regulate proteasomal degradation of NRF2. Mol. Cell Biol. 24, 7130–7139 (2004).
Liu, C. C. et al. Cul3-KLHL20 ubiquitin ligase governs the turnover of ULK1 and VPS34 complexes to control autophagy termination. Mol. Cell 61, 84–97 (2016).
Leonard, D. et al. The identification and characterization of a GDP-dissociation inhibitor (GDI) for the CDC42Hs protein. J. Biol. Chem. 267, 22860–22868 (1992).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Hodgson, L. et al. FRET binding antenna reports spatiotemporal dynamics of GDI–CDC42 GTPase interactions. Nat. Chem. Biol. 12, 802–809 (2016).
Garcia-Mata, R., Boulter, E. & Burridge, K. The ‘invisible hand’: regulation of Rho GTPases by RhoGDIs. Nat. Rev. Mol. Cell Biol. 12, 493–504 (2011).
Santoro, C. et al. Intermittent macrothrombocytopenia in a novel patient with Takenouchi–Kosaki syndrome and review of literature. Eur. J. Med. Genet. 64, 104358 (2021).
Nakashima, M. et al. De novo variants in CUL3 are associated with global developmental delays with or without infantile spasms. J. Hum. Genet. 65, 727–734 (2020).
Kato, K., Miya, F., Oka, Y., Mizuno, S. & Saitoh, S. A novel missense variant in CUL3 shows altered binding ability to BTB-adaptor proteins leading to diverse phenotypes of CUL3-related disorders. J. Hum. Genet. 66, 491–498 (2021).
Castel, P. Defective protein degradation in genetic disorders. Biochim. Biophys. Acta Mol. Basis Dis. 1868, 166366 (2022).
Daimon, E., Shibukawa, Y., Thanasegaran, S., Yamazaki, N. & Okamoto, N. Macrothrombocytopenia of Takenouchi–Kosaki syndrome is ameliorated by CDC42 specific- and lipidation inhibitors in MEG-01 cells. Sci. Rep. 11, 17990 (2021).
Chirnomas, D., Hornberger, K. R. & Crews, C. M. Protein degraders enter the clinic—a new approach to cancer therapy. Nat. Rev. Clin. Oncol. 20, 265–278 (2023).
Clayton, N. S. & Ridley, A. J. Targeting Rho GTPase signaling networks in cancer. Front. Cell Dev. Biol. 8, 222 (2020).
Peng, J. M., Tseng, R. H., Shih, T. C. & Hsieh, S. Y. CAMK2N1 suppresses hepatoma growth through inhibiting E2F1-mediated cell-cycle signaling. Cancer Lett. 497, 66–76 (2021).
Bera, R. et al. Functional genomics identified a novel protein tyrosine phosphatase receptor type F-mediated growth inhibition in hepatocarcinogenesis. Hepatology 59, 2238–2250 (2014).
Peng, J. M., Lin, S. H., Yu, M. C. & Hsieh, S. Y. CLIC1 recruits PIP5K1A/C to induce cell-matrix adhesions for tumor metastasis. J. Clin. Investig. 131, e133525 (2021).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Laskowski, R. A., Jabłońska, J., Pravda, L., Vařeková, R. S. & Thornton, J. M. PDBsum: structural summaries of PDB entries. Protein Sci. 27, 129–134 (2018).
Kaplan, W. & Littlejohn, T. G. Swiss-PDB Viewer (Deep View). Brief. Bioinform. 2, 195–197 (2001).
Bowie, J. U., Lüthy, R. & Eisenberg, D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 253, 164–170 (1991).
Wiederstein, M. & Sippl, M. J. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, W407–W410 (2007).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Acknowledgements
We sincerely thank J.-M. Egly from the Department of Functional Genomics and Cancer, IGBMC, CNRS/INSERM/University of Strasbourg, for critical review of this paper. We are grateful to S.-P. Chai for technical assistance with protein ubiquitylation assays and Y.-H. Yi for support with live-cell imaging studies. We extend our appreciation to H.-L. Huang for administrative support, as well as the Microscopy Core and Clinical Proteomics Core of Chang Gung Memorial Hospital for their technical expertise. We thank the National RNAi Core Facility of Taiwan for providing lentivirus-based shRNA clones. This work was supported by research grants from the Chang Gung Medical Foundation (CORPG3L0011~2, CORPG3L0441~5, CMRPG3N0071~3 and OMRPG3M0011 to S.-Y.H.), National Health Research Institute (NHRI-EX114-11217BI to S.-Y.H.) and the Ministry of Science and Technology (MOST 109-2314-B-182A-061-MY3 to S.-Y.H.), Taiwan.
Author information
Authors and Affiliations
Contributions
S.-Y.H. conceived the project and designed the experiments. P.-C.L., H.-C.C., Y.-H.H., C.-C.L. and C.-J.Y. performed the immunoblotting, immunoprecipitation and protein ubiquitylation assays and the related data analyses. P.-C.L. and H.-C.C. conducted the imaging studies and data analyses. Y.-C.L. and P.-C.L. performed the molecular modeling. H.-C.C. wrote the Methods and figure legends, and S.-Y.H. wrote the paper and edited the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
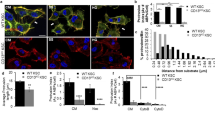
Extended Data Fig. 1 Knockdown of KLHL23 using different shRNA clones consistently enhances CDC42 bioactivities.
Three shRNA clones targeting distinct regions of KLHL23 mRNA were used to transfect cells, and silencing efficiency was validated by quantitative RT-PCR. a,b, Quantification of filopodia number (a) and F-actin intensity (b) per cell, shown as dot plots (mean ± s.d.). Data were pooled from three independent experiments. (a) shLuc (n = 17), shKLHL23-1 (n = 19), shKLHL23-2 (n = 17), and shKLHL23-3 (n = 16). (b) F-actin intensity was measured in three hepatoma cell lines: HepG2 — shLuc (n = 11), shKLHL23-1 (n = 12), shKLHL23-2 (n = 14); Hep3B — shLuc (n = 10), shKLHL23-1 (n = 17), shKLHL23-2 (n = 14); Huh7 — shLuc (n = 9), shKLHL23-1 (n = 19), shKLHL23-2 (n = 15). Statistical significance was assessed using two-tailed t-tests comparing each KLHL23 knockdown group to the corresponding control. c, Dot plots (mean ± SD) present the relative intensity of CDC42-FLARE/FRET signals in the actin cortex of the plasma membrane. shLuc (controls): n = 33 cells; shKLHL23 (silenced KLHL23 expression): n = 45 cells. Statistical significance was assessed using two-tailed t-tests.
Extended Data Fig. 2 The KLHL23-Cul3 complex is an E3 ligase targeting CDC42 for polyubiquitylation and degradation.
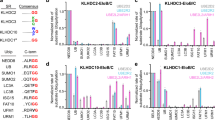
a,b, Immunoblots for CDC42 in HepG2 and HEK293 cells with or without KLHL23 depletion (shLuc or shK23) or ectopic KLHL23 expression (K23). MG132: a 26S proteasomal inhibitor. c, MLN4924 (MLN; a neddylation inhibitor) exhibits a dose-dependent suppression of KLHL23-mediated CDC42 degradation. d, Relative amounts of CDC42 harboring different mutations affected by ectopic expression of KLHL23 wild-type, T17N (constitutive inactivated mutant), and Q61L (constitutively active mutant) variants of CDC42, respectively. K107R, K133R, K135R, K144R, K153R, K163R, K166R, K183R, and K184R were CDC42-Q61L variants with a mutation at the indicated lysine residue. e, Tandem mass spectra of Ub-conjugated lysine residues of CDC42. f, Tandem mass spectrum of a peptide derived from ubiquitinated CDC42 exhibited Ub conjugation at the K11 residue of Ub, as indicated by a red circle.
Extended Data Fig. 3 RhoGDI facilitates KLHL23 selectively targeting active CDC42·GTP.
a, In vitro polyubiquitylation assays reveal the effects of RhoGDI on KLHL23-mediated CDC42 polyubiquitylation. b, Immunoblots assessing the impact of RhoGDI on the levels of CDC42 variants in HEK293 cells with or without KLHL23 depletion. WT, Q61L, and T17N represent wild-type, constitutively active, and inactive CDC42 variants, respectively. c, The silencing of endogenous RhoGDI sensitizes T17N-CDC42 variant to KLHL23-mediated degradation.
Extended Data Fig. 4 Ribbon representation of binding of the KLHL domain of KLHL23 and RhoGDI to CDC42.
a, The binding model for CDC42 interaction with the KLHL domain of KLHL23 (left) or RhoGDI (right). The middle panel is the merged image for binding the Keltch domain and RhoGDI. The C-terminus of CDC42 interacts only with RhoGDI but not with KLHL. b, A close view of the superimposition of interaction sites on CDC42 shared by KLHL23 and RhoGDI. c, Y64 of CDC42 forms two hydrogen bonds with D42 and K52 of RhoGDI1. Notably, Y64 also forms two hydrogen bonds with H424 and S430 of KLHL23 (Extended Data Figs. 6b and 9a, d).
Extended Data Fig. 5 Validation of the predicted model using different diagnostic methods.
a, Ramachandran analysis indicates that over 99.2% of the backbone dihedral angles in the model structure are located in energetically allowed regions. b, Assessment of the 3D model of the Kelch domain using the Verify 3D software. c, ProSA is widely used to validate protein structures obtained from experimental methods and theoretical calculations. The Z-score indicates the overall quality of the modeled KLHL domain structure. As shown by the black dots in this figure, the general model quality (Z-score) of the modeled structure of the KLHL domain is within the range of scores characteristic of native proteins of a similar size determined through X-ray (light blue) or nuclear magnetic resonance imaging (dark blue) and deposited in the RCSB Protein Data Bank (https://www.rcsb.org/).
Extended Data Fig. 6 GDP or GTP promotes CDC42 binding preference to RhoGDI or KLHL23.
a,b, Ribbon representation of the shift of switch I and II of CDC42 by GDP or GTP binding to interact with RhoGDI or KLHL23. White-dashed circles: intramolecular H bonds formed between the third phosphate of GTP with the switch I and II of CDC42. Red dashed circles: inter-molecular H bonds formed between the third phosphate of GTP with RhoGDI (a) or KLHL23 (b). c, Protein-protein interaction by using immunoprecipitation. HEK 293 T cells were co-transfected with KLHL23 (upper panel) or RhoGDI (lower panel) and myc-tagged vector and CDC42 variants. Wt: wild-type CDC42. Immunoprecipitation was performed by using anti-myc antibodies.
Extended Data Fig. 7 Colocalization of KLHL23 and RhoGDI in the plasma membrane.
a, GFP-CDC42, GFP-KLHL23, and GFP-RhoGDI co-localize at the leading edge of the plasma membrane in migrating cells. Arrowheads indicate the plasma membrane edge at the leading front of migrating cells, as shown in the differential interference contrast (DIC) image (left panel). K23: KLHL23; C42: CDC42; GDI: RhoGDI. Scale bar = 10 μm. b, Time-lapse imaging for GFP-KLHL23 colocalized with the plasma membrane during filopodia formation of a HeLa cell 15 min after being reseeded on the fibronectin-coated plates. Left panel: A representative merged image shows colocalization of KLHL23 with PM at the plasma membrane and filopodia.
Extended Data Fig. 8 Optimization of CDC42 FLARE/FRET biosensor variants interacting with KLHL23 and RhoGDI in live cells.
a, Schematic representation of the GDI·CDC42 FLARE biosensor. FRET occurs when KLHL23 (left panel) or RhoGDI (right panel) binds to the antenna-CDC42 chimera. b, Representative fluorescence images of HT1080 cells co-overexpressing GDI·CDC42 FLARE derivatives and either KLHL23 or RhoGDI, reseeded on fibronectin-coated substrates. Scale bar, 20 μm. c, Quantification of normalized FRET/CFP emission ratios for GDI·CDC42 FLARE derivatives in the presence of excess KLHL23 (left panel) or RhoGDI (right panel), shown as mean ± s.d. UR denotes the CDC42-K163R/K166R variant. Sample sizes: for KLHL23 interaction—CDC42.WT (n = 49), UR (n = 50), T17N (n = 41), T17N-UR (n = 45), Q61L (n = 48), and Q61L-UR (n = 44); for RhoGDI interaction—CDC42.WT (n = 52) and UR (n = 52). Statistical analysis was performed using two-tailed t-tests. Data represent three independent experiments. d, Representative time-lapse images illustrating the analysis of the spatiotemporal relationship between FLARE interactions and membrane dynamics. e, Spatiotemporal profiles of FLARE/FRET intensities and membrane dynamics. Blue and red curves represent FLARE/FRET intensities for the interaction of KLHL23 with CDC42.T17N.UR and CDC42.Q61L.UR, respectively. Light blue and light red vertical lines indicate the distances of membrane movement corresponding to the same conditions. The plots show raw data prior to normalization and synchronization with membrane dynamics. f, Scheme of the revised model regulating CDC42 activity.
Extended Data Fig. 9 Computer modeling for interaction of CDC42 with the KLHL domain of KLHL23.
a, Molecular docking model showing hydrogen bond formation between CDC42-Y64 and KLHL23 residues: H424 and S430 (right panel), which are lost when Y64 is replaced with Y64C of CDC42 (left panel). b, Aerial view of the interaction between the KLHL domain (brown) and CDC42 (grey). c, d, The potentially critical role of Y64 in stabilizing the interaction by forming two hydrogen bonds, Y64-CDC42—H424-KLHL23 and Y64—S430-KLHL23. e, Replacement of Y64 with cysteine (Y64C) abolishes these hydrogen bonds, disrupting the interaction.
Extended Data Fig. 10 The Y64C-CDC42 variant demonstrates a comparable rescue effect to that of ub-resistant K163R/K166R-CDC42.
a, Immunoblot analysis confirming transfection efficiency.b, Representative images of wound-healing assays demonstrating cell migration efficiency of cells transfected with the indicated plasmids. Relative migration was measured 48 hours after wound creation. c, Dot plots summarize wound closure efficiency, expressed as the percentage of the wounded area remaining (mean ± s.d., n = 9), from three independent experiments. Two-sided t test. Mock: transfection with an empty vector; shKLHL23: shRNAs targeting KLHL23. d, Immunoblotting confirms transfection efficiency. Endogenous KLHL23 was minimally detectable without MG132 treatment. e, RNA sequencing analysis of total RNA from 20 human tissues. The data were derived from the NCBI Gene database (https://www.ncbi.nlm.nih.gov/gene/151230; PRJNA280600)2. f, Expression levels of Klhl23 across different mouse tissues.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1.
Supplementary Video 1 (download AVI )
CDC42 2G FRET mock (related to Fig. 1e,h): time-lapse FRET assays for CDC42 activation using a CDC42 2G biosensor in HT1080 cells transfected with an empty vector (mock).
Supplementary Video 2 (download AVI )
CDC42 2G FRET as KLHL23 overexpression (related to Fig. 1e,h): time-lapse FRET assays for CDC42 activation using a CDC42 2G biosensor in HT1080 cells with KLHL23 (OX) ectopic expression.
Supplementary Video 3 (download MP4 )
CDC42-Q61L-UR FLARE/FRET as interacting with KLHL23 (related to Fig. 5c, top): time-lapse FRET assays for the KLHL23–CDC42-Q61L-UR interaction in HT1080 cells during migration.
Supplementary Video 4
CDC42-UR FLARE/FRET as interacting with RhoGDI (related to Fig. 5c, bottom): time-lapse FRET assays for the RhoGDI–CDC42-UR interaction in HT1080 cells during migration.
Supplementary Video 5 (download MP4 )
CDC42-Q61L-UR FLARE/FRET as interacting with KLHL23 in filopodia (related to Fig. 5f): time-lapse FRET assays for CDC42 activation using a CDC42 2G biosensor in HT1080 cells with KLHL23 (OX) ectopic expression.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, PC., Chang, HC., Liu, YC. et al. KLHL23 and RhoGDI coordinate CDC42 inactivation ensuring membrane homeostasis. Nat Chem Biol (2025). https://doi.org/10.1038/s41589-025-02010-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-025-02010-2