Abstract
Cellular lipid metabolism is subject to strong homeostatic regulation, but the players involved in and mechanisms underlying these pathways remain largely uncharacterized. Here we develop a ‘feeding–fishing’ approach coupling membrane editing using optogenetic lipid-modifying enzymes (feeding) with organelle membrane proteomics through proximity labeling (fishing) to elucidate molecular players and pathways involved in the homeostasis of phosphatidic acid (PA), a multifunctional lipid central to glycerolipid metabolism. This approach identified several PA-metabolizing enzymes and lipid transfer proteins enriched in and depleted from PA-fed membranes. Mechanistic analysis revealed that PA homeostasis in the cytosolic leaflets of the plasma membrane and lysosomes is mediated by both local PA metabolism and the action of lipid transfer proteins that carry out interorganelle lipid transport before subsequent metabolism. More broadly, the interfacing of membrane editing to controllably modify membrane lipid composition with organelle membrane proteomics using proximity labeling represents a strategy for revealing mechanisms governing lipid homeostasis.

Similar content being viewed by others
Main
Membranes are complex and dynamic assemblies that regulate myriad cellular processes. The lipid composition of membranes, crucial to membrane biophysical properties and the functions of the membrane-associated proteome, is determined by a balance of activities of lipid-metabolizing enzymes and lipid transfer proteins (LTPs). Efforts to decode these lipid–protein networks are complicated by the rapid diffusion, trafficking and, in many cases, short metabolic half-lives of lipids.
One type of lipid emblematic of these challenges is phosphatidic acid (PA)1. A central player in lipid metabolism, PA is produced by at least four different biosynthetic routes and metabolized into phospholipids, triglycerides or fatty acids by three pathways2,3. Despite representing only 1% of the lipidome, PA is present in most organelle membranes, and its levels transiently rise downstream of signaling from numerous cell-surface receptors and soluble signaling enzymes4,5. Because such PA bursts induce potent signaling events, PA is subject to strong homeostatic regulation, with its levels tightly controlled in space and time by modulation of both biosynthetic flux and regulated interorganelle transport by LTPs4,5,6,7. Although several PA-metabolizing enzymes and PA-specific LTPs have been characterized, it is not well understood how cells sense changes to levels of PA to regulate the activities of these enzymes and transporters to ultimately restore PA levels and achieve homeostasis.
Here, we develop and exploit a ‘feeding–fishing’ strategy to map proteins involved in such homeostatic processes and identify mechanisms of how cells respond to changes in the lipid composition of their organelle membranes. In this approach, we combined membrane editing using an optogenetic superactive phospholipase D (superPLD) with proximity-dependent biotinylation using a membrane-tethered TurboID, followed by tandem mass tagging (TMT)-based mass spectrometry (MS) to enable protein identification and quantification. In this manner, feeding PA to a membrane and fishing out proteins associated with that membrane would identify proteins that accumulate on or are depleted from a target membrane in response to changes in PA levels within that membrane.
Our studies reveal changes to the membrane recruitment of several PA-metabolizing enzymes and LTPs in PA-fed membranes that collectively facilitate removal of excess PA. Analysis of lipid-metabolizing enzymes points to roles for certain PA metabolic pathways in clearing PA pools from the plasma membrane and lysosomes. Mechanistic investigations using confocal microscopy to visualize PA pools and lipidomics to track changes to the phospholipidome revealed divergent roles for several LTPs recruited to PA-fed membranes in affecting PA metabolism. In particular, we found that Nir2, an LTP known to transport PA between the plasma membrane and ER, impacted PA homeostasis not only at the plasma membrane but also at other organelles, and SCP2, a broad-spectrum LTP implicated in sterol and fatty acid transport, can also mediate PA clearance. We additionally point to unexpected roles for members of the SMP and ORD domain-containing LTP families in PA homeostasis, including PDZD8, TEX2 and ORP1L. Collectively, membrane editing with superPLD coupled to proximity labeling enables identification of regulators of PA homeostasis, revealing cellular mechanisms underlying rapid PA metabolism and interorganelle transport.
Results
Design of the feeding–fishing proteomics strategy
Membrane editing is an emerging strategy to precisely manipulate the lipid composition of organelle membranes within living cells to study the physiological functions of individual lipids8,9. We previously devoted substantial efforts toward development of membrane editors for PA. First, we designed an optogenetic PLD (optoPLD) to acutely produce PA on target organelle membranes10. PLDs produce PA through the hydrolysis of abundant phospholipids, and optoPLD uses a blue-light-inducible heterodimerization system to recruit PLD onto desired membranes. Moreover, the ability of PLDs to additionally catalyze transphosphatidylation with exogenously supplied primary alcohols to produce a variety of natural and unnatural phospholipids underscores the versatility of optoPLD for editing the phospholipidome11.
To overcome the relatively modest activity of the first-generation optoPLD, we greatly improved its performance by using directed evolution in mammalian cells12. The resultant superPLDs, which acquired several mutations that optimized performance of this disulfide-containing, secreted protein from Streptomyces sp. PMF in the mammalian intracellular environment, exhibited up to 100-fold higher activity in cells compared to the original editor (wild-type optoPLD). SuperPLDs were demonstrated as potent membrane editors for spatiotemporally defined editing of phospholipids with organelle-level precision in live cells.
Our feeding–fishing strategy to identify proteins associated with PA-fed membranes involves superPLD-enabled membrane editing coupled to proximity proteomics (Fig. 1a). Currently, there are two widely used types of proximity labeling enzymes with distinct labeling mechanisms: promiscuous biotin ligase (for example, TurboID13) and engineered peroxidase (for example, APEX2 (ref. 14)). Promiscuous biotin ligase releases a biotin-AMP intermediate that reacts with lysine residues on proximal proteins, whereas proximity labeling with engineered ascorbate peroxidase is mediated by biotin-phenoxyl radicals that react with tyrosine residues15. These enzymes are generally expressed in intracellular compartments, where the reactive intermediates are highly confined, and their cross-membrane diffusion is rare.
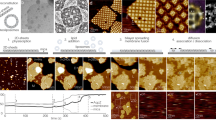
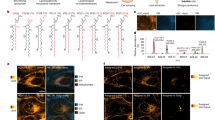
a, Feeding–fishing strategy couples membrane editing and proximity labeling for elucidation of regulators of PA homeostasis on different organelle membranes. An optogenetic superPLD is targeted to a membrane of interest to feed it with PA, and a proximity labeling enzyme is anchored to the same membrane to enable biotinylation to fish out proteins recruited to the lipid-edited membrane. PL, proximity labeling. b, Comparison of APEX2 and TurboID for proximal organelle membrane proteomics, evaluating their ability to selectively label a model protein (optoPLD) associated with the same versus different membranes. APEX2 labeling was performed using a 30-min incubation with 500 μM biotin-phenol followed by 1 min with H2O2. TurboID labeling was performed using a 10-min incubation with 500 μM biotin. PM, plasma membrane. c, Quantification of biotinylated optoPLD in b (n = 3 for APEX2 and n = 2 for TurboID, where n indicates independent biological replicates). d,e, Immunofluorescence images of cells expressing APEX2 (d) or TurboID (e) targeted to plasma membrane or lysosomes. Shown are representative images from two independent experiments. Scale bars, 10 μm. f, Western blots of cells expressing TurboID targeted to either the plasma membrane (KRAS CAAX domain19), lysosomes (p18/LAMTOR1 lysosomal-targeting sequence20 or TMEM192 (ref. 65)) or the ER (P450 2C1 (residues 1–29)66). Labeling was performed by 10-min incubation with 500 μM biotin. The rightmost two lanes indicate cells without expressing TurboID treated with or without biotin. Shown are representative blots from two independent experiments. g, One-shot transduction strategy enables efficient and transient coexpression of the feeding–fishing components. h,i, Immunofluorescence images of cells coexpressing TurboID and superPLD targeted to the plasma membrane (h) or lysosomes (i; Lyso). Shown are representative images from three independent experiments. hv, blue light (470 nm). Scale bars, 20 μm.
Because there are limited examples of tethering these enzymes to membranes to map membrane-proximal proteomes, we first explored the feasibility of using these two types of proximity labeling systems in this application, that is, to selectively label proteins that reside on the same membrane they are tagged to. We expressed APEX2 or TurboID fused to a plasma membrane targeting sequence, Lyn10, to localize the enzyme to the cytoplasmic leaflet of the plasma membrane. In the same cells, we coexpressed the wild-type optoPLD (from a bicistronic expression vector, CRY2–mCherry–PLD–P2A–CIBN–Tag where Tag is an organelle-targeting sequence) targeted to the cytosolic leaflet of either the plasma membrane or endoplasmic reticulum (ER) membrane. Surprisingly, plasma membrane-tagged APEX2 showed no difference in the labeling of optoPLD on the same (plasma membrane) versus different (ER) membrane, whereas TurboID preferentially labeled optoPLD on the same membrane (Fig. 1b,c). Consistent with this result, TurboID, when targeted to specific membranes, exhibited better colocalization with biotinylated proteins than APEX2 (Fig. 1d,e). From these results, we concluded that TurboID would be an optimal enzyme for mapping proteins associated with membranes of interest.
Among various membranes that could be targeted by superPLD and TurboID, we selected the cytoplasmic leaflets of the plasma membrane and lysosomal membrane because of the proposed localization and physiological functions of mammalian PLD1/2, which can initiate PA-based signaling on these membranes16,17,18. We previously showed that CIBN fused to the CAAX domain of KRAS19 (CIBN–CAAX) and the lysosome-targeting sequence of p18/LAMTOR1 (ref. 20) (p18–CIBN) can mediate effective superPLD recruitment and consequent PA enrichment on plasma membrane and lysosomes, respectively12. Using the same sequences for targeting TurboID to these membranes, we confirmed that these membrane-tethered TurboIDs showed equally high and promiscuous labeling activity to the originally reported ER membrane-tethered TurboID13 (Fig. 1f).
The feeding–fishing strategy requires coexpression of both TurboID and superPLD in cells at high efficiency. To avoid potential long-term effects of chronic expression, we performed transient expression by one-shot, lentivirus-based transduction. Although optoPLD or superPLD is typically expressed using a P2A self-cleavable peptide to ensure equimolar expression of CRY2–mCherry–PLD and CIBN–Tag, we found that for this application, separating these two components enabled more efficient transduction because of the limited size of DNA inserts that each lentivirus can encapsulate21. Therefore, our optimized protocol involved a triple lentivirus transduction system with one lentivirus each harboring TurboID–Tag, CRY2–mCherry–PLD or CIBN–Tag (Fig. 1g). To facilitate the incorporation of lentivirus into cells, we used spinfection, which is a method to apply mild centrifugation force during transduction22. The ratio of three lentivirus strains was optimized, on the basis of their titers, to achieve the highest coexpression efficiency while keeping cell viability unaffected. With these modifications, we achieved high efficiency (>75%) for transient cotransduction of HEK 293T cells with TurboID and superPLD targeted to either the plasma membrane or lysosomes (Fig. 1h,i).
Identification of proteins enriched on PA-fed membranes
We next performed proximity proteomics to identify the effects of local PA production on the proteomes of different membranes. We treated HEK 293T cells coexpressing TurboID and superPLD, targeted to either the plasma membrane or lysosomes, for 30 min with intermittent blue light (470 nm, 5 s per 1 min) to activate superPLD, followed by a 3-min labeling with 150 μM biotin (Fig. 1g). As a negative control, we used cells transduced with TurboID and a catalytically dead superPLD (H167A;H440A, deadPLD) that does not produce PA. Catalytically dead control samples were subjected to the same 470-nm blue-light irradiation protocol to account for any potential membrane environmental changes because of foreign protein recruitment or light-induced side effects23. Biotinylated proteins from two conditions (superPLD versus deadPLD) and three replicates were combined for TMT 16-plex proteomics analysis to identify proteins that were selectively enriched on or depleted from PA-fed membranes (Fig. 2a).
a, Schematic of feeding–fishing proximity proteomics workflow. Biotinylated proteins from HEK 293T cells coexpressing TurboID and either superPLD or deadPLD targeted to the same membrane (either plasma membrane or lysosomes) were enriched by streptavidin–agarose pulldown, eluted from resin and subjected to TMT labeling and multiplexed proteomics. b,c, Volcano plots showing differential enrichment of proteins on PA-fed versus unfed plasma membranes (b) or lysosomes (c). Proteins with known functions in lipid transport and lipid metabolism are colored in magenta and green, respectively, and proteins that were further functionally characterized in this study are indicated in italic. d, Table summarizing proteins that showed significance (abundance ratio P < 0.05) in enrichment or depletion on PA-fed versus unfed plasma membranes or lysosomes. Protein hits are ordered by fold enrichment and the hits with particularly high significance (abundance ratio P < 0.001) are shown in bold. Two-sided Student’s t-tests were used for P-value calculation of the reported ratios. GO, Gene Ontology.
In these feeding–fishing proteomics experiments, we detected 4,401 and 4,260 biotinylated proteins from the plasma membrane and lysosomes, respectively, with high correlations between replicates (Fig. 2). To identify proteins that were enriched on or depleted from PA-fed membranes, we calculated protein enrichment in the superPLD samples relative to deadPLD controls and plotted abundance ratios and associated P values (Fig. 2b,c). Among the hits that showed a significant enrichment on PA-fed membranes (based on the abundance ratio P value) were two known PA-metabolizing enzymes, Lipins 1 and 2 (LPIN1/2), which degrade PA into diacylglycerol (DAG). Correspondingly, DAG kinase δ (DGKD), which produces PA from DAG, was found to be depleted from PA-fed membranes. Furthermore, the best-characterized mammalian PA transfer protein, Nir2 (PITPNM1), was also enriched on both plasma membranes and lysosomes upon PA production, supporting the validity of the feeding–fishing proteomics. Western blot and immunofluorescence analysis of representative target proteins revealed that the enrichment was because of differences in their labeling and localization patterns rather than their global expression levels (Extended Data Fig. 1).
We used the Gene Ontology biological process annotation to comprehensively highlight proteins associated with lipid biology: lipid localization (which includes lipid transport), lipid metabolic process, regulation of lipid localization, regulation of lipid metabolism and response to lipids. For the plasma membrane feeding–fishing studies, 42 proteins associated with these terms were enriched on this membrane and 18 were depleted; similarly, for lysosomal feeding–fishing experiments, 40 such proteins were enriched and 25 were depleted from lysosomal membranes (Fig. 2d and Supplementary Table 1).
Beyond proteins related to lipid transport and metabolism, interactome analysis using the STRING network revealed additional insights, including notable changes in mitochondrial proteins, which were enriched on PA-fed plasma membranes and depleted from lysosome-depleted membranes (Extended Data Fig. 2). An examination of mitochondrial morphology found alterations in cells expressing superPLD, where mitochondria appeared as round, fragmented structures (Extended Data Fig. 3a). Interestingly, a similar phenotype was recently reported upon induction of acute DAG synthesis at the outer mitochondrial membrane, whose lipid composition was found to regulate membrane-shaping proteins that alter mitochondrial morphology24. Consistent with this finding, our data showed that the tendency and kinetics of mitochondrial fragmentation were strongly influenced by the site of PA production, with PA generated on membranes more closely associated with mitochondria (for example, ER) being more effective than PA produced on more distal membranes (for example, plasma membrane) (Extended Data Fig. 3b,c).
Lipid transporters Nir2 and SCP2 reduce local PA enrichment
Nir2 is a well-characterized PA transfer protein that translocates to ER–plasma membrane contact sites upon PA production (for example, following PLC activation) and facilitates PA transfer from the plasma membrane to the ER6. Because our feeding–fishing proteomics detected Nir2 to be enriched not only at the plasma membrane but also on lysosomes upon PA feeding, we overexpressed an miRFP-tagged Nir2 construct to monitor its localization in mammalian cells. miRFP–Nir2 exhibited mostly cytosolic but weakly ER-associated localization under basal conditions, in agreement with a previous study using GFP–Nir2 (ref. 6). Upon light-induced superPLD recruitment to the plasma membrane or to lysosomes, miRFP–Nir2 relocalized to these membranes, strongly colocalizing with superPLD (Fig. 3a,b and Extended Data Fig. 4a,b).
a,b, Confocal images of cells coexpressing miRFP–Nir2 and either superPLD or deadPLD targeted to the plasma membrane (a) or lysosomes (b), whose recruitment was induced by intermittent blue-light illumination (470 nm, 5 s per 1 min). c, Schematic of approach to visualize PA localization with GFP–Spo20 upon PA feeding by superPLD in the presence or absence of forced expression of LTPs. d,e, Confocal images of cells coexpressing a PA-binding probe GFP–Spo20 and either PM-superPLD (d) or lyso-superPLD (e), without or with stable expression of V5–Nir2. f,g, Quantification of d and e, measuring colocalization between GFP–Spo20 and superPLD. Each graph shows the colocalization time course in the cells with empty vector (black), V5–Nir2 (red) or a mutant form of V5–Nir2 (T59A) deficient in lipid transfer (blue). Solid lines indicate the mean and shaded areas indicate the s.d. Statistical analysis was performed using a two-sided repeated-measures ANOVA (n = 34 cells, **P = 0.0032 for Nir2 and P = 0.1949 for T59A (f); n = 20 cells, **P = 0.0315 for Nir2 and P = 0.2021 for T59A (g)). Three independent experiments were performed with similar results. h,i, Confocal images of cells coexpressing the PA-binding probe GFP–Spo20 and either PM-superPLD (h) or lyso-superPLD (i), without or with expression of proSCP2. j,k, Similar plots to f and g using data from h and i. Each graph shows the colocalization time course in control cells (black) or cells stably expressing V5–Nir2 (red), SCP2 (blue) or proSCP2 (yellow). Note that the V5 tag was omitted from SCP2 and proSCP2 to avoid potential interference with their post-translational cleavage. Statistical analysis was performed using a two-sided repeated-measures ANOVA (n = 20 cells, *P = 0.0334 for Nir2, P = 0.3646 for SCP2 and P = 0.0691 for proSCP2 (j); n = 20 cells, **P = 0.0008 for Nir2, P = 0.0315 for SCP2 and **P = 0.0002 for proSCP2 (k)). Three independent experiments were performed with similar results. Scale bars, 10 μm. NS, not significant.
We then tested whether Nir2 could exert its PA transfer function on these PA-fed membranes. To minimize any potential interference on its function caused by the fluorescent protein fusion, we replaced miRFP with a V5 tag (GKPIPNPLLGLDST) and used lentiviral transduction to generate HEK 293T cells stably expressing V5–Nir2 (Extended Data Fig. 4c). To monitor PA localization in these cells, we coexpressed superPLD with the PA reporter GFP–Spo20 (ref. 25). GFP–Spo20 was chosen over GFP–PASS, a version designed with a nuclear export signal for improved sensitivity24, because the predominantly nuclear localization and lack of cytosolic signal of GFP–Spo20 simplified the quantitative colocalization analysis (Fig. 3c). In control wild-type cells, superPLD targeted either to the plasma membrane or to lysosomes triggered PA enrichment on these membranes, as illustrated by accumulation of GFP–Spo20 at the membranes where superPLD was localized (Fig. 3d,e). By contrast, in cells stably expressing V5–Nir2, this PA enrichment did not occur. Quantification of the overlap between GFP–Spo20 and superPLD fluorescence upon superPLD activation revealed little colocalization in V5–Nir2-expressing cells but increasing colocalization over time in cells expressing either the T59A lipid transfer mutant of Nir2 or empty vector, as continuous superPLD activation would be expected to elevate PA levels (Fig. 3f,g).
We then set out to discover new PA transport proteins from among the feeding–fishing proteomics hits. We selected three additional LTPs—SCP2, PDZD8 and OSBPL1A/ORP1L—that were significantly enriched on PA-fed membranes. As with Nir2, we produced HEK 293T cells stably expressing V5-tagged versions of each LTP (Extended Data Fig. 4c). Consistent with previous reports, PDZD8–V5 localized to the ER and lysosomes26, and V5–ORP1L localized to lysosomes27. V5–SCP2 localized exclusively to punctate structures that partially colocalized with lysosomes (Extended Data Fig. 4d). Expression of the SCP2 gene leads to two protein isoforms, a full-length 58-kDa SCP2 (also known as SCP-x) and a precursor 15-kDa polypeptide, proSCP2; both of these proteins undergo post-translational cleavage to produce a 13-kDa C-terminal fragment known as mature SCP2 (mSCP2)28. The N-terminal remnant of full-length SCP2/SCP-x, after proteolytic cleavage in the peroxisomes, becomes a functional peroxisomal thiolase29, which explains the punctate localization of our V5–SCP2 construct. In line with previous studies30, a V5 fusion to mSCP2 showed increased cytosolic localization with much weaker association with peroxisomes (Extended Data Fig. 4e).
We then assessed the effects of these LTPs on local PA enrichment. Whereas stable cell lines expressing the respective LTP did not show any noticeable difference in superPLD expression or activity levels (Extended Data Fig. 4f–h), HEK 293T cells stably expressing SCP2/SCP-x or proSCP2 showed a significant delay in the colocalization between GFP–Spo20 and superPLD to either membrane, indicating reduced PA enrichment on these membranes (Fig. 3h–k). The PA reduction effect with proSCP2 was stronger than with SCP2/SCP-x, which aligns with differences in their maturation efficiencies, as cleavage of SCP-x is partial, whereas proSCP2 undergoes complete cleavage28,31. To directly assess the capacity of SCP2 in mediating PA clearance through its lipid transfer activity, we reconstituted its activity in vitro using a fluorescence resonance energy transfer dequenching assay with tethered donor and acceptor liposomes32. In these studies, purified mSCP2 was anchored to donor liposomes to increase its effective local concentration, and fluorescence dequenching was used to assess lipid transport from donor to acceptor liposomes. We found that mSCP2 exhibited lipid transfer activity in a manner dependent on both PA and mSCP2 concentrations (Extended Data Fig. 5). These studies suggest that SCP2, characterized as a transporter of fatty acids and several other types of phospholipids, may also function as a PA transfer protein in cells.
In contrast to SCP2, expression of two other LTPs tested, PDZD8 and ORP1L, did not induce a decrease in PA enrichment from PA-fed membranes (Fig. 4a–d). Because both PDZD8 and ORP1L can bind to PA in vitro33,34, we examined whether superPLD-induced PA enrichment caused any changes in their subcellular localization using GFP fusions to these proteins. PDZD8–GFP was found mostly at the ER, similar to PDZD8–V5 immunofluorescence, and GFP–ORP1L showed some lysosomal association along with a cytosolic pool (Extended Data Fig. 6a,b). It is noteworthy that the lysosomal pool of GFP–ORP1L exhibited limited colocalization with lysosome-targeted superPLD, suggesting that ORP1L and the p18/LAMTOR-targeted superPLD may reside in different microdomains. In addition to PDZD8, another SMP domain-containing LTP35, TEX2, was also enriched on PA-fed lysosomes in feeding–fishing proteomics. Both PDZD8 and TEX2 showed relatively high association with lysosome-targeted superPLD (Fig. 4e,f and Extended Data Fig. 6c,d). We found that acute PA production on lysosomes led to increased association between ER and lysosomes, and knockdown of PDZD8 mitigated this effect (Fig. 4g,h). Because both PDZD8 and TEX2 have been reported to localize to and mediate lipid transfer at ER–lysosome contact sites36, their detection in the feeding–fishing proteomics may reflect increased ER–lysosome association mediated by PA, rather than direct interactions with PA.
a,b, Confocal images of cells coexpressing the PA-binding probe GFP–Spo20 and mCherry-fused superPLD targeted to lysosomes, whose recruitment was induced by intermittent blue-light illumination (470 nm, 5 s per 1 min), without or with stable expression of PDZD8–V5 (a) or V5–ORP1L (b). c,d, Quantification of a and b, measuring colocalization between GFP–Spo20 and superPLD. Each graph shows the colocalization time course in control cells (black) or cells stably expressing V5–PDZD8 (blue) or V5–ORP1L (red). Solid lines indicate the mean and shaded areas indicate the s.d. Statistical analysis was performed using a two-sided repeated-measures ANOVA (n = 20 cells, P = 0.8562 for PDZD8 and P = 0.5025 for ORP1L (c); n = 20 cells, P = 0.3918 for PDZD8 and P = 0.5803 for ORP1L (d)). Two independent experiments were performed with similar results. e, Confocal images of cells expressing PDZD8–EGFP with coexpression of superPLD targeted to lysosomes. Representative images acquired 60 min after superPLD recruitment by intermittent blue-light illumination (470 nm, 5 s per 1 min) from 3 independent experiments are shown. Scale bars, 5 μm. f, Zoomed-in images of e for the areas marked with the dashed rectangles. Scale bar, 2 μm. g, Confocal images of cells expressing lysosome-anchored LOVPLD, costained with ER and lysosomal markers (calnexin and LAMP2, respectively), treated with or without light. Shown are representative images from three biological replicates. Scale bars, 20 µm. h, Quantification of the Pearson correlation coefficient between ER and lysosomes in LOVPLD-expressing cells treated with (red) or without (gray) light or treated with light in PDZD8-knockdown cells (blue). Black horizontal bars indicate the mean and vertical error bars indicate the s.d. Statistical significance was determined using a one-way ANOVA followed by post hoc honestly significant difference (HSD) Tukey test (n = 100 cells, **P = 0.002 and P = 0.4).
Cells maintain homeostasis to counteract PA overproduction
In our feeding–fishing proteomics studies, several enzymes that degrade or produce PA were, respectively, either enriched on or depleted from PA-fed membranes. Notably, these enzymes displayed differential abundances between PA-fed plasma membranes and lysosomes (Extended Data Fig. 7a). For example, all three isoforms of the lipin PA phosphatases (LPIN1/2/3) were enriched on PA-fed plasma membranes, whereas only LPIN1 was detected on PA-fed lysosomes and at lower abundance levels (Fig. 2a and Extended Data Fig. 1d). To understand the potential impact of these differences on PA metabolism and overall phospholipid homeostasis, we investigated the global changes to the phospholipidome by liquid chromatography (LC)–MS-based lipidomics after superPLD-mediated membrane editing to feed PA to either the plasma membrane or lysosomes (Fig. 5a,b).
a, Schematic depiction of PA feeding coupled to lipidomics to examine changes to cellular phospholipid levels. Lentiviral spinfection was used to express superPLD or deadPLD targeted to the plasma membrane or lysosomes. Then, 48 h after spinfection, cells were illuminated with or without intermittent blue light (470 nm, 5 s per 1 min) for 60 min to induce PLD recruitment to the plasma membrane or lysosomes. Cellular lipids were then extracted and analyzed by LC–MS. b, The CDP-DAG pathway and Kennedy pathway are two major pathways by which PA is converted to other phospholipids. PE, phosphatidylethanolamine; TAG, triacylglycerol. c, Heat maps depicting fold changes in individual (top) and total (bottom) levels of phospholipid species in cells after (+hv 470 nm) versus before (dark) 60-min illumination with 470-nm blue light to recruit deadPLD or superPLD to either the plasma membrane or lysosomes. d, Heat maps depicting fold changes in individual (top) and total (bottom) levels of phospholipid species, comparing cells expressing superPLD versus deadPLD, after (+hv 470 nm) their targeting to either the plasma membrane or lysosomes (n = 3 replicates per condition). e,f, Heat maps showing fold changes in individual and total levels of phospholipid species in cells stably expressing V5–Nir2, V5–SCP2, PDZD8–V5 or V5–ORP1L, 60 min after recruitment of superPLD to either the plasma membrane (e) or lysosomes (f). Fold changes were calculated on the basis of phospholipid levels in control HEK 293T cells expressing empty V5 vector and otherwise treated identically.
We found that HEK 293T cells expressing plasma membrane or lysosome-targeted superPLD exhibited light-dependent increases in PA levels after blue-light-induced superPLD recruitment to the target organelle membrane (Fig. 5c). Such increases in PA required superPLD activity, as they were not observed upon recruitment of deadPLD to these membranes (Fig. 5c). A more pronounced PA increase was observed at the plasma membrane compared to lysosomes upon PA feeding, indicating a slower net turnover of PA molecules at the plasma membrane. This finding is interesting considering the higher abundance of lipins at PA-fed plasma membranes (Extended Data Fig. 7a), which suggests an increased conversion of plasma membrane PA pools to DAG.
The conversion of PA to DAG is bidirectional in cells (where the reverse reaction is catalyzed by DAG kinases), and the rapid turnover of DAG molecules has been observed with an estimated half-life of one to several minutes37,38,39. By contrast, the conversion of PA to cytidine diphosphate (CDP)-DAG (and subsequently to phosphatidylglycerol (PG) and phosphatidylinositol (PtdIns)) is unidirectional. It is possible that the sustained buildup of PA at the plasma membrane results from increased PA flux within the bidirectional PA–DAG pathway, eventually leading to slower net PA turnover, and the increased abundance of lipins and DAG kinase η (DGKH) on PA-fed plasma membrane could be involved in this event (Extended Data Fig. 7). Moreover, despite superPLD consuming phosphatidylcholine (PC; with a preference for abundant species with 34:1 and 36:1 acyl chains10) for PA production, the levels of these lipid species did not change, suggesting that a substantial portion of PA produced on this timescale is routed to the Kennedy pathway as well to replenish these phospholipids (Fig. 5b). Concomitantly, the increased levels of PtdIns in PA-fed lysosomes may be accounted for by PA flux into the PA–CDP-DAG pathway.
SuperPLD expression results in moderate background PA production activity even in the dark12, which we addressed by using a next-generation, ultralow-background membrane editor for PA production termed LOVPLD40; thus, we hypothesized that superPLD-expressing cells would exhibit chronically elevated PA levels. However, in contrast to the robust PA increase observed during acute PA production (Fig. 5c), the overall difference resulting from chronic background PA production, analyzed by comparing deadPLD-expressing versus superPLD-expressing cells, was modest or even absent (Fig. 5d). Instead, a striking increase was observed in PG levels, suggesting that cells maintain PA homeostasis by removing excess PA molecules through the CDP-DAG pathway. Interestingly, small interfering RNA (siRNA)-mediated knockdown of a mitochondrial CDP-DAG synthase (CDS), TAMM41 (refs. 41,42), attenuated this PG increase in superPLD-expressing cells (Extended Data Fig. 8a,b). Because PG synthesis occurs in mitochondria and TAMM41 is responsible for the CDS activity localized there41, our lipidomics analysis and the observed changes in mitochondrial morphology upon acute PA production (Extended Data Fig. 3b,c) suggest that PA trafficking to and metabolism on mitochondria may serve as a mechanism to remove excess PA.
LTPs maintain PA homeostasis
Beyond these changes in recruitment of PA-metabolizing enzymes, the feeding–fishing experiments also revealed recruitment of several LTPs that could potentially be involved in restoring PA homeostasis (Extended Data Fig. 7b). Indeed, our confocal microscopy analysis in cells coexpressing superPLD and the PA-binding probe demonstrated that forced expression of two such LTPs, Nir2 and SCP2, can diminish local enrichment of PA at the plasma membrane and lysosomal membranes (Fig. 3). To directly investigate the roles of these lipid transporters in the global regulation of PA metabolism, we performed lipidomics analysis on cells subjected to PA feeding of either the plasma membrane or lysosomes that also overexpressed one of these LTPs. We found that forced expression of either Nir2 or SCP2 caused a significant attenuation in the global increase of PA levels that occurs upon PA feeding of either organelle membrane (Fig. 5e,f). These two LTPs had a stronger effect in decreasing excess PA at the plasma membrane compared to at lysosomes, a finding that aligns with our lipidomics analysis indicating slower baseline levels of PA turnover at the plasma membrane (Fig. 5c). Combined with the imaging analysis, these lipidomics results suggest that Nir2 and SCP2 can facilitate PA clearance at both local and global levels.
Overexpression of PDZD8 also led to a modest but significant decrease in cellular PA levels (Fig. 5e,f). Taken together with the increased ER–lysosome association observed upon PA production (Fig. 4g,h), it is plausible that PDZD8 and TEX2 contribute indirectly to PA clearance either by mediating lipid transfer at or facilitating formation of ER–lysosome contact sites, rather than by directly binding to PA or PA-fed membranes. Lastly, in cells overexpressing ORP1L, we observed an unexpected elevation in PA levels upon PA feeding, particularly at lysosomes (Fig. 5f). This PA enrichment persisted when full-length ORP1L was replaced with ORP1L variants bearing amino acid substitutions in either the PtdIns4P-binding site (H651A;H652A)27, in a well-conserved lipid-binding motif, EQVSHHPP, located within the ORD domain43, or its FFAT motif (F476A;Y477A)27, which mediates ER targeting through interactions with VAPA/B. However, the PA enrichment was antagonized by expression of an ORP1L mutant in its ORD domain that is deficient in cholesterol binding (Y583A)44 (Extended Data Fig. 8c,d).
LTPs affect PA localization and signaling
Our proteomics and lipidomics analyses suggest that cells maintain PA homeostasis by harnessing two pathways for PA consumption, the Kennedy pathway and CDP-DAG pathway, and multiple LTPs including Nir2 and SCP2 facilitate PA trafficking and ultimate clearance in cells. To further explore how the native forms of these LTPs may regulate PA distribution and signaling as a complement to overexpression studies, we depleted the endogenous pools of these LTPs by siRNA-mediated knockdown (Extended Data Fig. 9a,b). We found that, in SCP2-depleted cells, stable expression of superPLD resulted in substantial cell death, potentially because of exacerbation of lipid metabolic defects upon SCP2 depletion by chronic levels of excess PA production that occur upon superPLD expression, even in the dark45. Therefore, we switched from superPLD to LOVPLD to reduce the effect of chronic background PA production. Whereas SCP2-depleted cells showed drastically reduced the expression levels of all transfected plasmids (~25% and ~50% lower levels of LOVPLD and PA probe expression level, respectively, than in control cells; Extended Data Fig. 9c,d), the patterns of PA enrichment were largely consistent with those in control cells (Fig. 6a,b; quantification shown in Extended Data Fig. 9e,f). Interestingly, in cells depleted of Nir2, PA distribution was markedly different but nonoverlapping with the site of PA production. Instead, in Nir2-depleted cells, PA production by LOVPLD either at the plasma membrane or on lysosomes led to PA accumulation in a perinuclear compartment that corresponded to the Golgi apparatus (Fig. 6a–c).
a,b, Confocal microscopy images of HEK 293T cells, with or without depletion of Nir2 (siNir2) or SCP2 (siSCP2), coexpressing a PA-binding probe (GFP–PASS) and LOVPLD (containing mCherry) localized on either plasma membrane (a) or lysosomes (b). Images were acquired 0 and 30 min after incubation with intermittent blue-light illumination (470 nm, 500 ms per 5 s). c, Images of Nir2-depleted HEK 293T cells from b, followed by fixation, permeabilization and immunostaining with Golgi marker GRASP65. Scale bars, 20 μm. d, Relative mTOR signaling activity in control or LTP-depleted HEK 293T cells stably expressing LOVPLD, measured by quantification of phospho-S6K level using western blot. LOVPLD was localized to the plasma membrane, lysosomes or ER, and 30-min incubation with intermittent blue-light illumination (470 nm, 500 ms per 5 s) was used to activate PA production by LOVPLD. DsiRNA targeted to a respective LTP was used to deplete Nir2 (siNir2), SCP2 (siSCP2) or PDZD8 (siPDZD8) (n = 4 biological replicates, except for siPDZD8, where n = 2 biological replicates). Black horizontal bars indicate the mean and vertical error bars indicate the s.d. Statistical analysis was performed using a one-way ANOVA followed by post hoc Tukey HSD test (**P = 0.008, **P = 0.001 and P = 0.16 for plasma membrane LOVPLD control versus siRNA samples; *P = 0.01, **P = 0.009 and P = 0.9 for lysosome LOVPLD control versus siRNA samples; P = 0.21, 0.14 and 0.9 for ER LOVPLD control versus siRNA samples).
Lastly, we asked whether changes in PA trafficking and metabolism caused by knockdown of these LTPs led to functional consequences. We focused on mTOR signaling, which is known to be activated by PA produced by PLDs46. In cells stably expressing LOVPLDs at the plasma membrane, lysosome or ER, we observed light-dependent increases in the activity of mTOR signaling, measured by the phosphorylation of p70 S6 kinase 1 (S6K) (Fig. 6d and Extended Data Fig. 9g). The depletion of Nir2 or SCP2 but not PDZD8 further increased the mTOR activity, correlating with the abilities of these proteins to clear excess PA produced by membrane editing. Notably, a similar degree of increase was observed in LTP-depleted cells without PA overproduction (Extended Data Fig. 9h,i), potentially reflecting changes in their baseline PA metabolism. Collectively, these knockdown studies support that endogenous pools of Nir2 and SCP2 can affect PA localization and cellular signaling pathways.
Discussion
In this study, we introduced feeding–fishing as a general strategy to identify regulators of lipid homeostasis at the organelle level. In this approach, membrane editing is used to feed lipids of interest to designated organelle membranes and proximity labeling is used to fish out proteins recruited to those same membranes. We implemented this concept to reveal proteins and potential mechanisms associated with PA metabolism and transport on the cytosolic leaflets of two distinct organelle membranes: the plasma membrane and the lysosomal membrane. We subsequently analyzed the consequences of the observed enrichment of several protein hits identified in the feeding–fishing proteomics using a combination of confocal microscopy to visualize the subcellular localization of PA using a PA-binding probe and LC–MS-based lipidomics to assess changes to the phospholipidome. A constant challenge in these studies is that local elevations in PA induced by membrane editing with optogenetic superPLDs is opposed by homeostatic mechanisms, that is, local PA metabolism and PA transport to distal membranes followed by subsequent metabolism. Our ability to produce transient elevations in PA on target membranes relies on the high activities of superPLDs to achieve such acute increases to PA levels12.
Our studies revealed that, following membrane editing to locally elevate PA on the plasma membrane and lysosomes, certain metabolic enzymes are recruited to deplete the excess PA. Among the several metabolic fates of PA, the enzymes involved in PA–DAG interconversion were dynamically regulated on PA-fed membranes, whereas minor changes in recruitment were observed for enzymes that mediate metabolism to lysophosphatidic acid. The synthases that convert PA to CDP-DAG (CDS1/2) were not reliably detected in our proteomics experiments, likely because these are multipass transmembrane proteins localized in ER membranes. It would be interesting to expand the feeding–fishing approach to additional organelle membranes involved in PA metabolism (for example, ER, Golgi complex and mitochondria).
A limitation of our study design is the moderate background activity of superPLD that is present even without light stimulation. Light-independent PG accumulation was observed in cells expressing superPLD, indicating that this background superPLD activity altered lipid metabolism even in this relatively short-term period (that is, 48 h after lentiviral transduction). Thus, future implementations of feeding–fishing proteomics would benefit from using LOVPLD, a next-generation, ultralow-background optogenetic membrane editor, for PA feeding40.
In addition to insights related to the contributions of different PA-metabolizing enzymes to PA homeostasis on the plasma membrane and lysosomal membrane, the feeding–fishing studies revealed the translocation of several LTPs to PA-fed plasma membranes and lysosomes that could return excess PA and its metabolites to the ER and/or other organelles for further metabolism. Notably, both a PA-specific LTP, Nir2, and a more broad-spectrum LTP, SCP2, could reduce local PA enrichment and global PA levels under the conditions when PA was supplied to plasma membrane or lysosomes. The finding that SCP2 can reduce local PA enrichment suggests that this soluble LTP can mediate intracellular transport of PA, in addition to cholesterol and several other lipids that it has previously been found to transport28,47,48,49. SCP2 contains an N-terminal amphipathic helix that preferentially binds to membranes enriched in negatively charged lipids such as PtdIns(4,5)P2 (refs. 50,51) and its expression was reported to selectively alter the distribution of phospholipids at the plasma membrane52. Whereas our in vitro lipid transfer assay used SCP2 tethered to donor liposomes to approximate its physiological local concentration (870 nM to micromolar range53), it remains undetermined whether such tethering would occur or be required for SCP2 function in cells. Identifying additional factors, beyond PA itself, that could accelerate SCP2-mediated activity or localization would be an interesting direction for future study.
Interestingly, depletion of endogenous Nir2 resulted in striking redistribution of PA produced at either the plasma membrane or lysosomal membrane to the Golgi apparatus. A previous study reported that depletion of Nir2 causes a reduction in the DAG level in the Golgi apparatus and leads to a substantial inhibition of protein transport from the trans-Golgi network to the plasma membrane54. Consistent with those findings, our results indicate that Nir2 depletion can affect PA distribution and negatively affect the PA-to-DAG conversion. Notably, the same study suggested that Nir2 RNA interference increased the levels of PC at the Golgi apparatus by ~20% through upregulation of PC synthesis from DAG and CDP-choline (Fig. 5b), whereas total cellular PC levels remained the same54. In mammalian cells, synthesis of PC from DAG is mainly catalyzed by CEPT1 and CPT1, which localize on the ER and Golgi complex, respectively55,56. Because Nir2 supports PA transport to the ER6, where PA can be metabolized into DAG and, subsequently, PC, it is plausible that Nir2-depleted cells provide compensatory PC synthesis through pathways from PA to DAG and then to PC that at least in part involves Golgi-localized enzymes. Identification of alternative LTPs responsible for PA trafficking to the Golgi complex requires further investigation.
In contrast to Nir2, SCP2, PDZD8 and TEX2, whose overexpression antagonized the PA accumulation elicited by membrane editing, ORP1L forced expression led to an unexpected further increase in PA levels. ORP1L binds to PA34, but the functional relevance of such binding and whether it is accompanied by PA extraction and interorganelle transport remain unknown. Our structure–function studies revealed that the PA increase induced by ORP1L required the cholesterol-binding site in its ORD domain but not the EQVSHHPP signature motif found in all ORP-family proteins43,57, which is consistent with the lack of enrichment of other members of the ORP family in our feeding–fishing proteomics results. In other ORP-family proteins, the ORD domain mediates lipid transfer by extracting the lipid from one membrane and depositing it into another58,59,60. It can have differential affinities for two different lipids whose binding sites partially overlap, and such dual affinities are critical for vectorial transport when coupled to other metabolic steps on the origin and destination membranes. The implication of the ORD domain from ORP1L in the paradoxical increase in PA seen upon ORP1L forced expression on PA-fed membranes suggests that mechanisms more complex than simple transport to the ER and subsequent metabolism, as would be predicted by analogy with other ORP proteins, might be involved. Therefore, the roles of ORP1L in PA metabolism warrant further detailed study.
Intriguingly, several other proteins related to cholesterol biosynthesis and trafficking were identified in the feeding–fishing proteomics. For example, SOAT1, which catalyzes the formation of fatty acid–cholesterol esters on the ER, and LIPA, which catalyzes the reverse reaction, were both enriched on PA-fed membranes. By contrast, STARD3, an LTP that mediates ER-to-lysosome cholesterol transport61, was depleted from PA-fed membranes. Investigation of these proteins might shed light on new and unexpected regulatory mechanisms between PA and cholesterol metabolism and homeostasis. Lastly, the enrichment of select members of the bridge-like LTP family to PA-fed membranes (VPS13B to the plasma membrane and VPS13C to the lysosome) suggests that increased bulk phospholipid flow between these organelles may occur as a result of changes to PA metabolism induced by superPLD-mediated membrane editing.
Beyond elucidating mechanisms that implicate select PA-metabolizing enzymes and transporters in mediating PA homeostasis at the organelle and cellular levels, our study is notable for the introduction of the feeding–fishing strategy. This approach combining membrane editing with proximity labeling is geared to reveal how targeted modifications to the lipid composition of individual organelle membranes impacts the proteomes of such membranes, information that can point to new ways that cells can sense and correct imbalances in lipid metabolism. By exploiting organelle-specific PC hydrolysis by optogenetic PLDs, we focused here on regulators of PA metabolism at the plasma membrane and lysosomes. Yet, our optogenetic PLDs can use exogenously supplied primary alcohols in transphosphatidylation reactions to produce other phospholipids10,12,40,62 (Extended Data Fig. 10), and a growing collection of membrane editors targeting diverse types of lipids (for example, phosphoinositides, DAGs and sterols) is emerging19,62,63,64. The interfacing of such tools with organelle membrane proteomics using proximity labeling in feeding–fishing (or, conversely, fasting–fishing) experiments represents a powerful and generalizable strategy for elucidating mechanisms governing lipid homeostasis.
Methods
Plasmids and cloning
References and/or sequences of plasmids and primers used for this study are provided (Supplementary Tables 2 and 3). Membrane-targeted TurboID and optoPLD were cloned into pCDNA5/FRT/TO for transient transfection and pCDH-CMV-MCS-EF1α-Puro for lentiviral transduction. For plasma membrane targeting, the CAAX domain of KRAS19 (GKKKKKKSKTKCVIM) was fused to the C terminus of the constructs after a linker sequence (GGSGSLYK). For lysosomal membrane targeting, the p18 domain20 (MGCCYSSENEDSDQDREERKLLLDPSSPPTKALNGAEPNY) followed by a linker sequence (GGRGSGSGSGSGSGSGSGSGSG) was fused to the N terminus of the constructs. PM-iLID-LOVPLD40 was cloned into pSBtet-Pur (Addgene, 60507) for stable expression.
Plasmids encoding PITPNM1 (Nir2), SCP2, PDZD8, TEX2 or OSBPL1A (ORP1L) were purchased from the DNASU plasmid repository, and their open reading frames were cloned into pCDH-CMV-MCS-EF1α-Puro with an optional V5 tag to generate stable cell lines overexpressing these proteins. V5–Nir2, SCP2, proSCP2 (residues 405–547 of SCP2), V5–ORP1L, PDZD8–V5 and TEX2–V5 were used for analyzing their functions in PA trafficking and metabolism in live cells (note that the V5 tag was omitted from SCP2 and proSCP2 to avoid potential interference with their post-translational cleavage). PITPNM1, PDZD8, TEX2 and OSBPL1A were also cloned into pCDNA3.1 along with EGFP or miRFP fluorescent proteins for localization studies in live cells.
For visualization of PA localization in live cells, a Spo20 PA-binding domain25 (MDNCSGSRRRDRLHVKLKSLRNKIHKQLHPNCRFDDATKTS) or PASS domain67 (Addgene, 193970) fused to EGFP was cloned into pCDH-CMV-MCS-EF1α-Puro.
Mammalian cell culture and transient transfection
Cells were grown in DMEM (Corning) supplemented with 10% FBS (Corning), 1% penicillin–streptomycin (Corning) and 1 mM sodium pyruvate (Thermo Fisher) at 37 °C in a 5% CO2 atmosphere. For poly(l-lysine) pretreatment, cell plates were treated with 0.1 mg ml−1 poly(l-lysine) (Sigma Aldrich, P2636) in PBS for 1 h at 37 °C, followed by triple rinses with autoclaved deionized water. For fibronectin coating, cell plates were treated with 20 μg ml−1 human plasma fibronectin (Millipore Sigma, FC010) in PBS for 5–15 min at 37 °C.
For transient transfection, HEK 293T cells were transfected using Lipofectamine 2000 (Invitrogen, 11668019) or PEI MAX (Polysciences, 24765). Cells were incubated in regular DMEM containing plasmids premixed with Lipofectamine 2000 (1–1.5 µg of total plasmids and 3 µl of Lipofectamine 2000 or PEI MAX for cells in 35-mm dish) and the cells were incubated for 20–24 h before experiments.
For transient gene knockdown, HEK 293T cells were transfected using Lipofectamine RNAiMAX (Invitrogen, 13778075). Cells were incubated in regular medium containing DsiRNA (Integrated DNA Technologies) premixed with Lipofectamine RNAiMAX (5 pmol of DsiRNA and 1.5 µl of Lipofectamine RNAiMAX for cells in a 24-well plate) and the cells were incubated for 40–48 h before experiments. A list of DsiRNA sequences is provided in Supplementary Table 3.
Highly efficient multigene expression by virus semipurification and spinfection
HEK 293T cells were transfected using Lipofectamine 2000 or PEI MAX for lentivirus production. HEK 293T cells seeded on a six-well plate were incubated in Transfectagro (Corning) or Opti-MEM (Gibco) supplemented with 10% FBS containing plasmids premixed with Lipofectamine 2000 or PEI MAX (0.5 µg of envelope plasmid, 1 µg of packaging plasmid, 1.5 µg of transfer plasmid and 6 µl of Lipofectamine 2000 or PEI MAX per well for a six-well plate). Then, 12–16 h after transfection, the transfection medium was replaced with regular DMEM and media were collected 40–48 h and 60–72 h after transfection to obtain virus-containing media. For multivirus transduction (for example, to introduce all feeding–fishing components), the collected virus media were concentrated by centrifugation at 100,000g (24,000 rpm in an SW 41 Ti swinging-bucket rotor) for 90 min at 4 °C. After ultracentrifugation, the supernatant was carefully decanted and the virus concentrate was resuspended in fresh DMEM. The ratio of lentivirus packaging cells and transduced cells used for this study was 1.5:1 for pCDH-CRY2-mCherry-PLD constructs and 0.16:1 for all the other constructs, optimized on the basis of the lentivirus titer of each construct.
For spinfection, HEK 293T cells seeded on a six-well plate (pretreated with poly(l-lysine) or fibronectin) were incubated in virus-resuspended medium supplemented with 0.4 µg ml−1 polybrene (Millipore Sigma). The plate was centrifuged at 931 or 1,000g for 2 h at 37 °C, followed by the replacement of virus-containing medium with fresh DMEM (as well as DsiRNA precomplexed with Lipofectamine RNAiMAX for knockdown study). The six-well plate was covered with aluminum foil to keep cells in the dark and the cells were incubated for 40–48 h before experiments.
Setup for optogenetics experiments
A homemade light box was built by attaching four strips of dimmable, 12-V blue light-emitting diode (LED) tape light (1000Bulbs.com, 2835-60-IP65-B1203) on the inside of a Styrofoam box. For optogenetics experiments, the light box was placed inside the CO2 incubator using an AC outlet power bank (Omars, 24,000 mAh, 80 W) as a power supply. An outlet timer (BN-LINK) was used to switch the light on and off automatically to enable 3-s intervals of 470-nm light per 1 min.
For imaging and mTOR signaling experiments using LOVPLD (Figs. 4g,h and 6, and Extended Data Figs. 3 and 4a,b), an AMUZA system consisting of a blue LED array (470 nm), an LED array driver and a pulse generator with 10% duty cycle (500 ms on, 5 s off) was used. Light power was typically around 10 mW cm−2 (measured by Thorlabs PM100D).
Membrane TurboID and PA TurboID labeling and proteomics
Cells expressing optoPLD and TurboID were illuminated for 30 min with intermittent blue light (470 nm, 5-s pulses every 1 min), followed by 3-min treatment with 500 µM biotin. The cells were then washed five times with PBS and lysed in RIPA lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS, 1 mM EDTA and 1× cOmplete protease inhibitor). For proteomics, the cells from all six wells in a six-well plate (~2 × 107 cells total) were combined to prepare each sample. After sonication and centrifugation, the lysate supernatant was incubated with Pierce high-capacity streptavidin–agarose beads (Thermo Fisher Scientific, 20359) overnight at 4 °C on a rotator. A 10-µl volume of bead solution was used for cells per well in 6-well plate.
After overnight incubation, the streptavidin beads were washed twice with RIPA buffer, once with 1 M KCl, once with 0.1 M Na2CO3, once with 2 M urea in 10 mM Tris-HCl and twice with RIPA buffer. For western blot analysis, the washed beads were boiled for 10 min in 3× Laemmli sample buffer supplemented with 25 mM biotin to elute biotinylated proteins off the beads. For proteomics, the beads were further washed twice with HEPES buffer (50 mM HEPES and 150 mM NaCl) and once with sample elution buffer (50 mM HEPES, 150 mM NaCl, 1% SDS, 0.1% Triton X-100 and 25 mM biotin, pH 8). The washed beads were boiled for 5 min in sample elution buffer for protein elution. The elution process was repeated for three times, each time replacing the elution buffer, to increase the protein yield. The samples were then quantified by BCA assay and submitted to Proteomics and Metabolomics Facility at Cornell University for TMT labeling and proteomics analysis. Four samples in triplicate were grouped for the labeling and analysis. Detailed workflows of the proteomics analyses are described in Supplementary Information.
Antibody and dilutions used for validation experiments (Extended Data Fig. 1c–e) were the following: anti-LPIN1 polyclonal antibody (Proteintech, 27026-1-AP; 1:1,000), anti-LPIN2 monoclonal antibody (Santa Cruz Biotechnology, sc-514353; 1:100), anti-Nir2 polyclonal antibody (Proteintech, 26983-1-AP; 1:1,000), anti-PDZD8 polyclonal antibody (Proteintech, 25512-1-AP; 1:1,000), anti-SCP-x polyclonal antibody (Proteintech, 14397-1-AP; 1:1,000), anti-SCP-2 polyclonal antibody (Proteintech, 23006-1-AP; 1:1,000), anti-DGKD polyclonal antibody (Abcepta, AP8126b; 1:1,000), anti-DGKH polyclonal antibody (Proteintech, 13873-1-AP; 1:1,000) and anti-β-tubulin monoclonal antibody (Cell Signaling Technology, 86298). Detection was conducted using chemiluminescence with the SuperSignal West Pico PLUS chemiluminescent substrate (Thermo Fisher Scientific, 34580) or SuperSignal West Atto ultimate-sensitivity substrate (Thermo Fisher Scientific, A38556) and acquisition on a Bio-Rad ChemiDoc MP system.
Time-course imaging by confocal microscopy
Cells were seeded on 35-mm glass-bottom imaging dishes (Matsunami Glass) coated with poly(l-lysine) or fibronectin. Unless otherwise noted, images were acquired every 1 min for 1 h at 37 °C using Zeiss Zen Blue 2.3 on a Zeiss LSM 800 confocal laser-scanning microscope equipped with Plan Apochromat objectives (×40, numerical aperture: 1.4) and two GaAsP photomultiplier tube detectors. Solid-state lasers (488, 561 and 640 nm) were used to excite GFP, mCherry and miRFP, respectively, and the 488-nm laser irradiation also served as a stimulus for activating optoPLD recruitment to the plasma membrane, ER or lysosomes.
Imaging experiments with LOVPLD (Figs. 4g,h and 6, and Extended Data Figs. 3 and 4a,b) were acquired using a Zeiss AxioObserver inverted microscope equipped with a Yokogawa spinning-disk confocal head, Cascade II:512 camera and four-color laser launch (405-nm diode, 491-nm diode-pumped solid-state laser (DPSS), 561-nm DPSS and 640-nm diode, all at 50 mW). Images were acquired using Slidebook software (Intelligent Imaging Innovations) through a ×100 oil-immersion objective. Emission filters were 528/38 nm for GFP and MitoTracker, 617/73 nm for mCherry and 700/75 nm for AlexaFluor 647. Mitochondrial morphology was observed by staining cells with 100 nM MitoTracker Green (Thermo Fisher Scientific, M7514) for 5 min before experiments.
Quantification of PA enrichment on the membranes
HEK 293T cells coexpressing a PA-binding probe (EGFP–Spo20), superPLD (CRY2–mCherry–superPLDmed, derived from superPLD clone 1-12 (ref. 12)) targeted to different organelle membranes and the indicated LTP were imaged as described above. Colocalization analysis between PA-binding probe and superPLD was carried out on ImageJ/FIJI as follows. Firstly, a region of interest (ROI) was drawn around each cell expressing both constructs. Secondly, for each ROI, the mCherry signal of the superPLD was used to generate a binary mask. Thirdly, the ratio of the EGFP signal from the PA-binding probe found in the mask, as compared to the total signal, was calculated to obtain the colocalization ratio. Lastly, the fold change in this colocalization ratio was tracked over the course of the time series and plotted with the use of matplotlib in Python. Statistical analysis was performed using a repeated-measures analysis of variance (ANOVA).
Immunofluorescence imaging
Cells were fixed in 4% formaldehyde for 10 min at room temperature, rinsed three times with PBS, permeabilized with 0.5% Triton X-100 in PBS for 15 min at room temperature and blocked with 1% BSA and 0.1% Tween-20 in PBS (PBS-T; blocking buffer) for 30 min. The cells were then treated with primary antibody solution in blocking buffer for 1 h at room temperature and rinsed three times with PBS-T. Afterward, the cells were treated with secondary antibody solution in blocking buffer for 1 h at room temperature and rinsed three times with PBS-T. After the final rinse, the cells were incubated in PBS-T for 5 min and stored in PBS supplemented with 1 µg ml−1 DAPI. Image acquisition by laser-scanning confocal microscopy was performed as described above using solid-state lasers (405, 488, 561 and 640 nm) to excite DAPI, AlexaFluor 488, mCherry and AlexaFluor 647, respectively. Pearson colocalization analysis was performed using the Coloc 2 plugin in ImageJ/FIJI.
Antibodies, affinity reagents and dilutions used for immunofluorescence staining were as follows: streptavidin–AlexaFluor 488 (Invitrogen, S11223; 1:50), anti-V5 antibody (Bio-Rad, MCA1360GA; 1:100), anti-calnexin antibody as an ER marker (Thermo Fisher Scientific, PA534754; 1:100), anti-LAMP2 antibody as a lysosomal marker (Santa Cruz Biotechnology, sc-18822; 1:100), anti-GRASP65 antibody as a Golgi marker (Santa Cruz Biotechnology, sc-374423; 1:100), anti-LPIN1 antibody (Cell Signaling Technology, 14906; 1:100), anti-mouse–AlexaFluor 488 antibody conjugate (Invitrogen, A21202; 1:500) and anti-mouse–AlexaFluor 647 (Invitrogen, A31571; 1:500).
In vitro liposome assay to measure lipid transfer activity
mSCP2 fused with an N-terminal 6×His tag was cloned into a pET28a vector and the construct was transformed into BL21 Rosetta2 Escherichia coli cells for purification. Cells were cultured in Terrific Broth medium at 37 °C to an optical density at 600 nm of 0.5 (approximately 2 h), and then expression was induced by addition of 0.2 mM IPTG followed by overnight incubation at 18 °C. Cells were harvested and lysed by sonication in buffer containing 150 mM NaCl, 10% glycerol, 5 mM β-mercaptoethanol, 1 mM PMSF and 50 mM sodium phosphate at pH 7.4. Lysate was clarified by centrifugation at 10,000g for 30 min, followed by incubation with TALON beads for 2 h at 4 °C. The resin was washed four times with lysis buffer, followed by elution with lysis buffer supplemented with 150 mM imidazole. The tether-only construct was produced as previously described68 and was purified similarly to mSCP2.
Liposomes were prepared as previously described32. Briefly, lipids were dissolved in chloroform and mixed in glass tubes in the indicated ratio for donor and acceptor liposomes separately. The mixture was dried under a stream of nitrogen to a thin film, followed by further drying in vacuo for 2 h. The lipid film was hydrated with buffer containing 25 mM Tris-HCl, 150 mM NaCl and 0.5 mM TCEP at pH 8.0 to a total lipid concentration of 2.5 mM. Liposomes were formed by ten freeze–thaw cycles in liquid N2 and 37 °C water bath, followed by extrusion 21 times through polycarbonate filters with a pore size of 100 nm.
Lipid transfer assays were set up at 25 °C in black flat-bottom 96-well plates, with 100-µl total volume containing 50 µM donor liposomes, 150 µM acceptor liposomes, 62.5 nM 6×His–PH domain tether (used to link donor and acceptor liposomes and, therefore, minimize spurious lipid exchange between donor–donor and acceptor–acceptor liposomes) and 187.5 nM 6×His–mSCP2 (His tag used to anchor mSCP2 to donor liposomes to increase local concentration), for a 1:800 protein-to-lipid ratio and a 1:3 tether-to-LTP ratio. For specific reactions, different compositions of liposomes or concentrations of LTP were used as indicated. The reaction was initiated by addition of proteins and was then monitored for 2 h, using an excitation wavelength of 465 nm and emission wavelength of 540 nm every 10 s, using a BioTek Synergy H1 microplate reader.
The following lipids were purchased: DOPC (Avanti Polar Lipids, 850375), PtdIns(4,5)P2 (Avanti Polar Lipids, 840046), DGS-NTA(Ni) (Avanti Polar Lipids, 790404), POPA (Avanti Polar Lipids, 840857), NBD-PA (Avanti Polar Lipids, 810176), NBD-PE (Avanti Polar Lipids, 810144), Rhod-PE (Avanti Polar Lipids, 810150), DOPE (Cayman, 15091) and DOPC (Echelon, L-1182).
Lipidomics analysis by LC–MS
HEK 293T cells expressing the indicated constructs were illuminated for 30 min with intermittent blue light (470 nm, 5-s pulses every min) and rinsed once in PBS; the cellular lipids were extracted using the Bligh–Dyer method10. Then, 250 μl of methanol was added to cells in a 35-mm dish on ice. Followed by the addition of 125 µl of 20 mM acetic acid and 100 µl of PBS, the cells were scraped off and transferred into a 1.5-ml centrifuge tube. After addition of 500 µl of chloroform, the tube was shaken vigorously for 3 min and centrifuged for 1 min at 10,000g. The bottom organic layer was transferred into a new tube and dried under a stream of N2 gas. The resulting lipid film was dissolved in 150 µl of chloroform and subjected to high-resolution LC–MS analysis.
LC–MS measurement was performed on an Agilent 6230 electrospray ionization time-of-flight MS instrument coupled to an Agilent 1260 high-performance LC instrument equipped with a Luna 3-µm silica LC column (Phenomenex; 50 × 2 mm) using a binary gradient elution system where solvent A was chloroform, methanol and ammonium hydroxide (85:15:0.5) and solvent B was chloroform, methanol, water and ammonium hydroxide (60:34:5:0.5). Separation was achieved using a linear gradient from 100% A to 100% B over 10 min. Phospholipid species were detected using an Agilent Jet Stream source operating in positive or negative mode, acquiring in an extended dynamic range of m/z 100–1,700 at one spectrum per second (gas temperature, 325 °C; drying gas, 12 L min−1; nebulizer, 35 psi; fragmentor, 300 V (for positive mode) and 250 V (for negative mode); sheath gas flow, 12 L min−1; Vcap, 3,000 V; nozzle voltage, 500 V.
The LC–MS data were analyzed on MassHunter quantitative analysis software using the ‘find compounds by formula’ tool. The search parameters were set as follows: source of formulas to confirm, database/library provided in Supplementary Table 4; matches per formula, 1 (automatically increase for isomeric compounds); Values to match, mass and retention time (retention time required); match tolerance, masses ± 20 ppm and retention times ± 0.200 min; expansion of values for chromatogram extraction, m/z ± 20 ppm and retention time 0.500 + 1.00 min; positive ion charge carriers, +H; negative ion charge carriers, −H.
Quantification of phosphatidyl alcohol production by LOVPLD
HEK 293T cells stably expressing doxycycline-inducible PM-iLID-LOVPLD were seeded in 12-well plates with 2.5 µg ml−1 doxycycline and incubated in the dark for 48 h. Each alcohol was first prepared as a 100× concentrated solution in PBS and then added to cells stably expressing PM-iLID-LOVPLD. The cells were immediately transferred to the light chamber and illuminated for 15 min with intermittent blue light (470 nm, 5-s pulses every 1 min). Bligh–Dyer extraction followed by LC–MS analysis was performed as described above.
Measurement of mTOR activity by quantification of phospho-S6K
HEK 293T cells were transduced with LOVPLD using lentivirus and spinfection as described above. The cells were incubated with 10 μM dorsomorphin (AMPK inhibitor) for 1 h at 37 °C, followed by a 30-min stimulation with intermittent blue-light illumination (470 nm, 500-ms pulses every 5 s). The cells were then lysed with RIPA lysis buffer supplemented with protease and phosphatase inhibitor cocktails (Thermo Fisher Scientific, 78439 and 78420). After sonication and centrifugation, the lysate supernatants were mixed with 6× Laemmli sample buffer to prepare the sample for western blot. The membrane was blotted with 1:1,000 dilutions of antibodies for phospho-S6K (T389; Cell Signaling Technology, #9205) or β-actin (Cell Signaling Technology, #5125), with detection by chemiluminescence using the SuperSignal West Pico PLUS chemiluminescent substrate (Thermo Fisher Scientific, 34580) and acquisition on a Bio-Rad ChemiDoc MP system.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Proteomics data are available from the PRIDE database under accession codes PXD070270 and PXD070378. Plasmids generated during the current study are listed along with their source information in Supplementary Information. Source data are provided with this paper.
Code availability
Custom Python scripts developed for proteomics and lipidomics analyses are available from GitHub (https://github.com/teipanda/Membrane-Turbo)69, along with the corresponding example datasets. Other scripts used for data processing and plotting were developed using a combination of ImageJ macros and Python and are available from the corresponding author upon request.
References
Liu, Y., Su, Y. & Wang, X. in Lipid-Mediated Protein Signaling (ed. Capelluto, D. G. S.) 159–176 (Springer, 2013).
Vance, J. E. Phospholipid synthesis and transport in mammalian cells. Traffic 16, 1–18 (2015).
Vance, J. E. & Vance, D. E. Phospholipid biosynthesis in mammalian cells. Biochem. Cell Biol. 82, 113–128 (2004).
Selvy, P. E., Lavieri, R. R., Lindsley, C. W. & Brown, H. A. Phospholipase D: enzymology, functionality, and chemical modulation. Chem. Rev. 111, 6064–6119 (2011).
Shulga, Y. V., Topham, M. K. & Epand, R. M. Regulation and functions of diacylglycerol kinases. Chem. Rev. 111, 6186–6208 (2011).
Kim, Y. J., Guzman-Hernandez, M.-L., Wisniewski, E. & Balla, T. Phosphatidylinositol–phosphatidic acid exchange by Nir2 at ER–PM contact sites maintains phosphoinositide signaling competence. Dev. Cell 33, 549–561 (2015).
Hussain, Z., Uyama, T., Tsuboi, K. & Ueda, N. Mammalian enzymes responsible for the biosynthesis of N-acylethanolamines. Biochim. Biophys. Acta 1862, 1546–1561 (2017).
Chen, P.-H. B., Li, X.-L. & Baskin, J. M. Synthetic lipid biology. Chem. Rev. 125, 2502–2560 (2025).
Luan, L., Frederick, N. P. & Baskin, J. M. Emerging approaches for studying lipid dynamics, metabolism, and interactions in cells. Annu. Rev. Biochem. 94, 417–446 (2025).
Tei, R. & Baskin, J. M. Spatiotemporal control of phosphatidic acid signaling with optogenetic, engineered phospholipase Ds. J. Cell Biol. 219, e201907013 (2020).
Chiu, D.-C. & Baskin, J. M. Imaging and editing the phospholipidome. Acc. Chem. Res. 55, 3088–3098 (2022).
Tei, R., Bagde, S. R., Fromme, J. C. & Baskin, J. M. Activity-based directed evolution of a membrane editor in mammalian cells. Nat. Chem. 15, 1030–1039 (2023).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Qin, W., Cho, K. F., Cavanagh, P. E. & Ting, A. Y. Deciphering molecular interactions by proximity labeling. Nat. Methods 18, 133–143 (2021).
Bowling, F. Z., Frohman, M. A. & Airola, M. V. Structure and regulation of human phospholipase D. Adv. Biol. Regul. 79, 100783 (2021).
Frias, M. A. et al. Phosphatidic acid drives mTORC1 lysosomal translocation in the absence of amino acids. J. Biol. Chem. 295, 263–274 (2020).
Hussain, S. S. et al. RalA and PLD1 promote lipid droplet growth in response to nutrient withdrawal. Cell Rep. 36, 109451 (2021).
Idevall-Hagren, O., Dickson, E. J., Hille, B., Toomre, D. K. & Camilli, P. D. Optogenetic control of phosphoinositide metabolism. Proc. Natl Acad. Sci. USA 109, E2316–E2323 (2012).
Nada, S., Mori, S., Takahashi, Y. & Okada, M. p18/LAMTOR1: a late endosome/lysosome-specific anchor protein for the mTORC1/MAPK signaling pathway. Methods Enzymol. 535, 249–263 (2014).
Kumar, M., Keller, B., Makalou, N. & Sutton, R. E. Systematic determination of the packaging limit of lentiviral vectors. Hum. Gene Ther. 12, 1893–1905 (2001).
Berggren, W. T., Lutz, M. & Modesto, V. General spinfection protocol. In StemBook (Harvard Stem Cell Institute, 2008).
Krassovka, J. M. et al. The impact of non-toxic blue light (453 nm) on cellular antioxidative capacity, TGF-β1 signaling, and myofibrogenesis of human skin fibroblasts. J. Photochem. Photobiol. B 209, 111952 (2020).
Pemberton, J. G. et al. Acute diacylglycerol production activates critical membrane-shaping proteins leading to mitochondrial tubulation and fission. Nat. Commun. 16, 2685 (2025).
Horchani, H., de Saint-Jean, M., Barelli, H. & Antonny, B. Interaction of the Spo20 membrane-sensor motif with phosphatidic acid and other anionic lipids, and influence of the membrane environment. PLoS ONE 9, e113484 (2014).
Elbaz-Alon, Y. et al. PDZD8 interacts with Protrudin and Rab7 at ER–late endosome membrane contact sites associated with mitochondria. Nat. Commun. 11, 3645 (2020).
Zhao, K. & Ridgway, N. D. Oxysterol-binding protein-related protein 1L regulates cholesterol egress from the endo-lysosomal system. Cell Rep. 19, 1807–1818 (2017).
Gallegos, A. M. et al. Gene structure, intracellular localization, and functional roles of sterol carrier protein-2. Prog. Lipid Res. 40, 498–563 (2001).
Wirtz, K. W. Phospholipid transfer proteins revisited. Biochem. J. 324, 353–360 (1997).
Schroeder, F. et al. Pro-sterol carrier protein-2: role of the N-terminal presequence in structure, function, and peroxisomal targeting. J. Biol. Chem. 275, 25547–25555 (2000).
Ossendorp, B. C. et al. Tissue-specific distribution of a peroxisomal 46-kDa protein related to the 58-kDa protein (sterol carrier protein X; sterol carrier protein 2/3-oxoacyl-CoA thiolase). Arch. Biochem. Biophys. 334, 251–260 (1996).
Valverde, D. P. et al. ATG2 transports lipids to promote autophagosome biogenesis. J. Cell Biol. 218, 1787–1798 (2019).
Gao, Y., Xiong, J., Chu, Q.-Z. & Ji, W.-K. PDZD8-mediated lipid transfer at contacts between the ER and late endosomes/lysosomes is required for neurite outgrowth. J. Cell Sci. 135, jcs255026 (2021).
Xu, Y., Liu, Y., Ridgway, N. D. & McMaster, C. R. Novel members of the human oxysterol-binding protein family bind phospholipids and regulate vesicle transport. J. Biol. Chem. 276, 18407–18414 (2001).
Reinisch, K. M. & De Camilli, P. SMP-domain proteins at membrane contact sites: structure and function. Biochim. Biophys. Acta 1861, 924–927 (2016).
Jeyasimman, D. et al. PDZD-8 and TEX-2 regulate endosomal PI(4,5)P2 homeostasis via lipid transport to promote embryogenesis in C. elegans. Nat. Commun. 12, 6065 (2021).
Schuhmacher, M. et al. Live-cell lipid biochemistry reveals a role of diacylglycerol side-chain composition for cellular lipid dynamics and protein affinities. Proc. Natl Acad. Sci. USA 117, 7729–7738 (2020).
Höglinger, D. et al. Trifunctional lipid probes for comprehensive studies of single lipid species in living cells. Proc. Natl Acad. Sci. USA 114, 1566–1571 (2017).
Jensen, J. B. et al. Biophysical physiology of phosphoinositide rapid dynamics and regulation in living cells. J. Gen. Physiol. 154, e202113074 (2022).
Li, X.-L., Tei, R., Uematsu, M. & Baskin, J. M. Ultralow background membrane editors for spatiotemporal control of phosphatidic acid metabolism and signaling. ACS Cent. Sci. 10, 543–554 (2024).
Blunsom, N. J., Gomez-Espinosa, E., Ashlin, T. G. & Cockcroft, S. Mitochondrial CDP-diacylglycerol synthase activity is due to the peripheral protein, TAMM41 and not due to the integral membrane protein, CDP-diacylglycerol synthase 1. Biochim. Biophys. Acta 1863, 284–298 (2018).
Jiao, H., Yin, Y. & Liu, Z. Structures of the mitochondrial CDP-DAG synthase Tam41 suggest a potential lipid substrate pathway from membrane to the active site. Structure 27, 1258–1269.e4 (2019).
Nakatsu, F. & Kawasaki, A. Functions of oxysterol-binding proteins at membrane contact sites and their control by phosphoinositide metabolism. Front. Cell Dev. Biol. 9, 664788 (2021).
Dong, J. et al. Allosteric enhancement of ORP1-mediated cholesterol transport by PI(4,5)P2/PI(3,4)P2. Nat. Commun. 10, 829 (2019).
Atshaves, B. P. et al. Effect of SCP-x gene ablation on branched-chain fatty acid metabolism. Am. J. Physiol. Gastrointest. Liver Physiol. 292, G939–G951 (2007).
Frias, M. A., Hatipoglu, A. & Foster, D. A. Regulation of mTOR by phosphatidic acid. Trends Endocrinol. Metab. 34, 170–180 (2023).
Galano, M., Venugopal, S. & Papadopoulos, V. Role of STAR and SCP2/SCPx in the transport of cholesterol and other lipids. Int. J. Mol. Sci. 23, 12115 (2022).
Puglielli, L., Rigotti, A., Greco, A. V., Santos, M. J. & Nervi, F. Sterol carrier protein-2 is involved in cholesterol transfer from the endoplasmic reticulum to the plasma membrane in human fibroblasts. J. Biol. Chem. 270, 18723–18726 (1995).
Schroeder, F. et al. Sterol carrier protein-2 functions in phosphatidylinositol transfer and signaling. Biochemistry 42, 3189–3202 (2003).
Huang, H., Ball, J. M., Billheimer, J. T. & Schroeder, F. The sterol carrier protein-2 amino terminus: a membrane interaction domain. Biochemistry 38, 13231–13243 (1999).
Schroeder, F. et al. Sterol carrier protein-2: new roles in regulating lipid rafts and signaling. Biochim. Biophys. Acta 1771, 700–718 (2007).
Gallegos, A. M. et al. Sterol carrier protein-2 expression alters plasma membrane lipid distribution and cholesterol dynamics. Biochemistry 40, 6493–6506 (2001).
Gianotti, A. R., Ferreyra, R. G. & Ermácora, M. R. Binding properties of sterol carrier protein 2 (SCP2) characterized using Laurdan. Biochim. Biophys. Acta 1866, 1143–1152 (2018).
Litvak, V., Dahan, N., Ramachandran, S., Sabanay, H. & Lev, S. Maintenance of the diacylglycerol level in the Golgi apparatus by the Nir2 protein is critical for Golgi secretory function. Nat. Cell Biol. 7, 225–234 (2005).
Horibata, Y. & Sugimoto, H. Differential contributions of choline phosphotransferases CPT1 and CEPT1 to the biosynthesis of choline phospholipids. J. Lipid Res. 62, 100100 (2021).
Dorighello, G., McPhee, M., Halliday, K., Dellaire, G. & Ridgway, N. D. Differential contributions of phosphotransferases CEPT1 and CHPT1 to phosphatidylcholine homeostasis and lipid droplet biogenesis. J. Biol. Chem. 299, 104578 (2023).
Depta, L., Whitmarsh-Everiss, T. & Laraia, L. Structure, function and small molecule modulation of intracellular sterol transport proteins. Bioorg. Med. Chem. 68, 116856 (2022).
Mesmin, B. et al. A four-step cycle driven by PI4P hydrolysis directs sterol/PI4P exchange by the ER–Golgi tether OSBP. Cell 155, 830–843 (2013).
Antonny, B., Bigay, J. & Mesmin, B. The oxysterol-binding protein cycle: burning off PI4P to transport cholesterol. Annu. Rev. Biochem. 87, 809–837 (2018).
Kovács, D. et al. Lipid exchange at ER–trans-Golgi contact sites governs polarized cargo sorting. J. Cell Biol. 223, e202307051 (2023).
Wilhelm, L. P. et al. STARD3 mediates endoplasmic reticulum-to-endosome cholesterol transport at membrane contact sites. EMBO J. 36, 1412–1433 (2017).
Tei, R. & Baskin, J. M. Induced proximity tools for precise manipulation of lipid signaling. Curr. Opin. Chem. Biol. 65, 93–100 (2021).
Zewe, J. P. et al. Probing the subcellular distribution of phosphatidylinositol reveals a surprising lack at the plasma membrane. J. Cell Biol. 219, e201906127 (2020).
Pham, H. et al. Development of a novel spatiotemporal depletion system for cellular cholesterol. J. Lipid Res. 63, 100178 (2022).
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Ahn, K., Szczesna-Skorupa, E. & Kemper, B. The amino-terminal 29 amino acids of cytochrome P450 2C1 are sufficient for retention in the endoplasmic reticulum. J. Biol. Chem. 268, 18726–18733 (1993).
Zhang, F. et al. Temporal production of the signaling lipid phosphatidic acid by phospholipase D2 determines the output of extracellular signal-regulated kinase signaling in cancer cells. Mol. Cell. Biol. 34, 84–95 (2014).
Bian, X., Saheki, Y. & De Camilli, P. Ca2+ releases E-Syt1 autoinhibition to couple ER–plasma membrane tethering with lipid transport. EMBO J. 37, 219–234 (2018).
Tei, R. teipanda/Membrane-Turbo: v2. Zenodo https://doi.org/10.5281/zenodo.17468627 (2025).
Acknowledgements
J.M.B. acknowledges support from the National Institutes of Health (R01GM151682). R.T. was supported by Honjo International, Funai Overseas, Cornell and Life Sciences Research Foundation fellowships. X.-L.L. was supported by a Natural Sciences and Engineering Research Council of Canada postgraduate fellowship. We acknowledge Cornell University Proteomics and Metabolomics Facility for their support in designing and analyzing proteomics study; specifically, we thank E. Anderson for TMT labeling of proteomics samples, Q. Fu for data analysis and S. Zhang for guidance. We thank A. Y. Ting and her lab at Stanford University for providing resources and instrumentation used in parts of this study.
Author information
Authors and Affiliations
Contributions
R.T. and J.M.B. conceptualized the project, designed the experiments, interpreted the results and wrote the paper, with input from other authors. X.-L.L. carried out the lipidomics experiments using LOVPLD, L.L. carried out the in vitro liposome assays, and R.T. carried out all other experiments and data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Itay Budin, Carsten Schultz and the other, anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Validation of proteomics hits.
a, b, Schematic depiction of optogenetic superPLD (a) for light-dependent recruitment of PLDPMF on the selected membranes, used in the Feeding–Fishing proteomics, and LOVPLD (b), which is constitutively tethered onto the membranes and activated upon 470 nm blue light, used for secondary validation with high temporal resolution. c–e, Quantification of key proteomics hits in supernatant (c) and streptavidin pulldown (d), with quantification shown in (e). SCP2, DGKD, DGKH, and Tubulin were not detected by Western blot in the pulldown samples. n = 2 independent biological replicates, and similar results were obtained in three independent experiments. Statistical analysis was performed using one-way ANOVA (p = 0.0008, 0.05, 0.01, 0.18, 0.11, 0.69, 0.0001, 0.05, respectively). f, g, Confocal microscopy analysis of endogenous Lipin-1, stained using an α-Lipin-1 antibody, with or without LOVPLD activation by 470 nm blue light, revealing changes in Lipin-1 localization pattern upon PA production at the plasma membrane. Shown are representative images from three biological replicates.
Extended Data Fig. 2 Interactome analysis of proteins identified in Feeding–Fishing proteomics.
STRING network analysis of proteins found to be either enriched on (a, plasma membrane; b, lysosomes) or depleted from (c, plasma membrane; d, lysosomes) PA-fed membranes. Proteins that showed p-value < 0.01 were analyzed (interaction score > 0.400), and clustering was performed using k-means. Two-sided Student’s t-tests were used for p value calculation.
Extended Data Fig. 3 Mitochondrial dysmorphology in cells with hyperactivated PA production.
a, Confocal microscopy of HEK 293T cells expressing plasma membrane or lysosome-targeted superPLD or deadPLD. Mitochondria were stained with MitoTracker Green. b, c, Quantification (b) of mitochondrial morphology (Normal, Mix, or Round) in cells expressing LOVPLD anchored on endoplasmic reticulum (ER), lysosome (Lyso), or plasma membrane (PM) over the indicated time periods (0–30 min for ER and 0–180 min for Lyso and PM). 50–100 cells (exact numbers are provided in the Source Data) from each condition were analyzed. Mitochondrial dysmorphology was more rapidly observed in the order of: ER > Lyso > PM. Representative cells showing Normal, Mix, and Round mitochondria are indicated by gray, green, and magenta arrows in (c), respectively. Similar results have been obtained in three independent experiments. Scale bars, 20 μm.
Extended Data Fig. 4 Stable expression of lipid transfer proteins in HEK 293T cells.
a, b, Quantification of miRFP-Nir2 (T59A mutant with lipid transfer activity disabled) co-localization with PA-fed membranes. Black horizontal bars indicate means and vertical error bars indicate standard deviations. Statistical significance was determined using two-sided Student’s t-test (n = 10–16 cells with exact numbers provided in the Source Data, p = 8.6E-06 and 8.9E-07). c, Confocal images of HEK 293T cells stably expressing V5-tagged Nir2 (PITPNM1), PDZD8, ORP1L (OSBPL1A), full-length SCP2 (SCP-x), or the mature form of SCP2 (mSCP2). Cells were co-stained with α-LAMP1 and α-Calnexin antibodies as lysosome and ER markers. Shown are representative images from two independent experiments. d, e, Zoomed-in images of cells expressing V5-SCP-x (d) or V5-mSCP2 (e). V5-mSCP2 shows more cytosolic localization. Scale bars, 20 μm. f, Scheme of IMPACT labeling to quantify superPLD activity in live cells. g, h, Flow cytometry results of cells co-expressing superPLD targeted to the plasma membrane (PM; g) or lysosomes (Lyso; h) and the indicated lipid-transfer protein. mCherry signal (readout of superPLD expression level) and BODIPY signal (readout of PLD activity) are plotted. Wild-type cells expressing deadPLD were included as a negative control. Shown are representative plots from two independent experiments.
Extended Data Fig. 5 Mature SCP2 (mSCP2) exhibits lipid transfer activity in a PA concentration-dependent manner in an in vitro liposome assay.
a, Schematic design of lipid transfer assay. Donor liposomes include NBD conjugated to phosphatidic acid (NBD-PA), whose fluorescence signal is quenched by FRET by rhodamine conjugated to phosphatidylethanolamine (Rhod-PE). Lipid transfer from donor liposomes to acceptor liposomes results in dequenching and increased fluorescence signal. b, SDS-PAGE gel of purified mSCP2. c–e, Fluorescence signal readout using donor liposomes with different POPA percentages (c), negative control where mSCP2 or acceptor liposomes were omitted (d), and with different mSCP2 concentrations (e). Raw fluorescence signal is shown except for e, where fluorescence signal was normalized to the No Acceptor control. The concentration of mSCP2 was 187.5 nM unless otherwise noted. Shown are representative plots from two independent experiments.
Extended Data Fig. 6 PDZD8, ORP1L and TEX2 do not exhibit dynamic recruitment to PA-enriched membranes.
a, b, Confocal images of cells co-expressing PDZD8-EGFP (a) or EGFP-ORP1L (b) and superPLD or deadPLD targeted to lysosomes (Lyso). Images acquired 0, 30, and 60 min following recruitment of superPLD or deadPLD induced by intermittent blue light illumination (470 nm, 5 s per 1 min), representative of three biological replicates, are shown. Scale bars, 10 μm. c, Confocal images of cells expressing TEX2-EGFP, with co-expression of superPLD or deadPLD targeted to lysosomes (Lyso). Images acquired 60 min following superPLD/deadPLD recruitment, representative of three biological replicates, are shown. Scale bars, 5 μm. d, Zoomed-in images of (c) for the areas marked with the dashed rectangles. Scale bars, 2 μm.
Extended Data Fig. 7 Lipid-modifying enzymes and lipid transfer proteins may directly or indirectly regulate PA metabolism.
a, Differential enrichment and depletion of enzymes that mediate PA synthesis and degradation on PA-fed plasma membranes (left) and lysosomes (right). Shown in the table are fold changes in protein abundance between the PA-fed (superPLD-recruited) and the negative control (deadPLD-recruited) membranes and the mean abundance values for each condition (n = 3). The fold changes determined to be statistically significant by two-sided Student’s t-tests (abundance ratio p-value < 0.05) are shown in bold. Enzymes that mediate conversion of PA to LPA (for example, PLA2s) were not detected. b, Similar list as in (a), showing lipid transfer proteins that exhibit differential enrichment on or depletion from PA-fed plasma membranes (left) and lysosomes (right).
Extended Data Fig. 8 Lipidomics analysis reveals the effects of TAMM41 and ORP1L on the phospholipidome in superPLD-expressing cells.
a, Quantification of PG levels by lipidomics upon expression of either deadPLD in control (gray) or TAMM41 knockdown (cyan) cells, or superPLD in control (magenta) or TAMM41 knockdown (green) cells, showing that TAMM41 knockdown attenuates the increase of PG levels in superPLD-expressing cells. Total cellular lipids were extracted and analyzed after a 30 min incubation with intermittent blue light illumination (470 nm, 5 s per 1 min). PG species with average normalized ion counts > 0.05 are shown. Vertical error bars indicate standard deviations. Statistical analysis was performed using one-way ANOVA followed by post-hoc HSD Tukey test (n = 3, p-values are shown in the plots). b, Similar quantification as in (a), showing PA levels. Note that TAMM41 knockdown did not lead to a further increase in PA levels, despite the abolished PG increase, suggesting that cells may compensate for the loss by removing excess PA through an alternative mechanism. c, Domain maps summarizing ORP1L mutants used in the lipidomics study. d, Heat map analysis similar to that shown in Fig. 5f but instead depicting fold changes in individual and total levels of phospholipid species in cells stably expressing V5-ORP1L, V5-ORP1LH651A/H652A (HH/AA: ORD domain mutant lacking PI4P-binding motif), V5-ORP1LY583A (Y583A: ORD domain mutant lacking cholesterol-binding motif), or V5-ORP1LF476A/Y477A (FY/AA: an FFAT mutant deficient in binding to VAP). n = 3 replicates per condition.
Extended Data Fig. 9 Altered cell physiology in LTP-depleted cells.
a, Western blot of LOVPLD-expressing HEK 293T cells treated with DsiRNA to deplete Nir2, SCP-x/SCP2, or PDZD8. DsiRNA #1 (colored red) was used in the subsequent experiments. b, Quantification of (a), with relative levels compared to the control shown in parentheses. Plots are representative of two independent experiments. c, d, Expression levels of LOVPLD (c) and the PA probe GFP-PASS (d) in HEK 293T cells depleted with Nir2 (siNir2), SCP2 (siSCP2), or PDZD8 (siPDZD8), co-expressing PA probe and LOVPLD targeted to either plasma membrane (PM) or lysosomes (Lyso). Plots are representative of two independent experiments. e, f, Quantification of colocalization between LOVPLD and GFP-PASS shown in Fig. 6a, b. Black horizontal bars indicate means and vertical error bars indicate standard deviations. Statistical analysis was performed using one-way ANOVA followed by post-hoc Tukey-HSD test (n = 14–19 cells with exact numbers provided in the Source Data, p = 0.01 and 0.6 for (e) and 0.03 and 0.9 for (f) for Control vs. siRNA samples). g, Western blot of p-S6K to measure mTOR activity in LTP-depleted cells expressing LOVPLD, with quantification presented in Fig. 6d. LOVPLD was localized to plasma membrane (PM), lysosomes (Lyso), or endoplasmic reticulum (ER), and 30-min incubation with intermittent blue light illumination (470 nm, 500 ms per 5 s) was used to activate PA production by LOVPLD. β-actin was used as a loading control. h, Similar experiment as (g) but in LTP-depleted HEK 293T cells without LOVPLD activation. i, Quantification of (h). n = 4 (except for siPDZD8, where n = 2) biological replicates. Statistical analysis was performed using one-way ANOVA followed by post-hoc Tukey-HSD test (p = 0.004, 0.01, and 0.07 for Control vs. siRNA samples). Black horizontal bars indicate means and vertical error bars indicate standard deviations.
Extended Data Fig. 10 Membrane editors can produce a variety of unnatural phospholipids in cells.
a–d, LC–MS quantification of phosphatidyl alcohol lipid species extracted from HEK 293T cells expressing PM-targeted iLID-LOVPLD treated with butanol (a), ethanol (b), butynol (c), or azidopropanol (d), demonstrating that LOVPLD can mediate in situ production of phospholipids with customizable alcohols in light-dependent manner. Total cellular lipids were extracted and analyzed after a 15 min incubation with intermittent blue light illumination (470 nm, 5 s per 1 min). Black horizontal bars indicate means and vertical error bars indicate standard deviations. e–h, Similar quantification of PA species extracted from cells treated with butanol (e), ethanol (f), butynol (g), or azidopropanol (h), showing the lack of noticeable increase in PA levels.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–4 and Methods.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tei, R., Li, XL., Luan, L. et al. Membrane editing with proximity labeling reveals regulators of lipid homeostasis. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-025-02104-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-025-02104-x
This article is cited by
-
How to catch a lipid transporter
Nature Chemical Biology (2026)