Abstract
Neural cells in the adult human central nervous system (CNS) display extensive transcriptional heterogeneity. How different layers of epigenetic regulation underpin this heterogeneity is poorly understood. Here we profile, at the single-nuclei epigenomic level, distinct regions of the adult human CNS, for chromatin accessibility and simultaneously for the histone modifications H3K27me3 and H3K27ac. We unveil a putative SOX10 enhancer and primed chromatin signatures at HOX loci in spinal-cord-derived human oligodendroglia (OLG) and astrocytes, but not microglia. These signatures in adult OLG were reminiscent of developmental profiles but were decoupled from robust gene expression. Moreover, using high-resolution Micro-C, we show that induced pluripotent stem-cell-derived human OLGs exhibit a HOX chromatin architecture compatible with the primed chromatin in adult OLGs, bearing a strong resemblance not only to OLG developmental architecture but also to high-grade pontine gliomas. Thus, epigenetic memory from developmental states in adult OLG not only enables them to promptly transcribe Hox family genes during regeneration but also makes them susceptible to gliomagenesis.
Similar content being viewed by others
Main
The human central nervous system (CNS) contains diverse neuronal and glial populations that together support sensory processing, motor control and higher cognitive function. Mature oligodendrocytes (MOLs) are a glial cell type that wraps neuronal axons with myelin, enabling rapid saltatory conduction1,2,3. MOLs are primarily found within the white matter (WM) areas of the CNS, whereas their progenitor population, oligodendrocyte precursor cells (OPCs), are uniformly distributed throughout the CNS. Previous studies that investigate the oligodendroglial (OLG) lineage have identified region-specific transcriptomic differences in the human CNS4,5.
While the transcriptome of human neural populations has been well characterized, the underlying regulatory chromatin landscape remains largely unknown. Chromatin accessibility provides a snapshot of the regulatory blueprint underlying cell states6,7 and has been used to identify organ-specific regulatory elements8,9, developmental regulatory circuits9,10 and disease-associated effects11,12,13,14,15,16. However, chromatin accessibility represents one regulatory layer. Histone post-translational modifications (PTMs) and DNA modifications also have essential roles in transcriptional regulation17. Single-cell studies of DNA methylation18 and individual histone marks19,20,21,22,23,24,25,26,27,28,29 have started elucidating epigenetic states in the adult mouse CNS, but similar characterization of the adult human CNS remains limited. In humans, DNA methylation and chromatin architecture have been analyzed at single-cell resolution30. Single-cell H3K27me3 profiling has been performed in a glioblastoma sample23 alongside bulk analyses11, and several histone modifications in sorted CNS subpopulations31,32. Recent multimodal assays have enabled joint single-cell capture of H3K27ac/RNA in the mouse brain and human M1 cortex25,33. However, joint single-cell profiling of multiple histone modifications across the adult human CNS is still lacking.
Here we provide a single-cell chromatin accessibility dataset of the adult human CNS across three anterior–posterior axis regions, together with a joint multimodal single-cell histone PTM dataset in the adult human CNS. These resources, available at the UCSC Cell Browser and UCSC Genome Browser34 (https://cns-nanocuttag-atac.cells.ucsc.edu), define H3K27me3 and H3K27ac landscapes across major cell populations across regions, and enabled the identification of a putative SOX10 enhancer and lineage-specific transcription factor (TF) regulatory networks. We show that adult OLGs show chromatin accessibility, histone PTM patterns and three-dimensional (3D) architecture reminiscent of their developmental counterparts. This suggests that epigenetic memory of key developmental genes persists in adult OLGs, potentially priming them for rapid activation of HOX loci during regenerative responses, while also making them susceptible to hijacking in tumor transformation, such as gliomagenesis.
Results
Single-nucleus assay for transposase-accessible chromatin using sequencing of the adult human CNS reveals differential chromatin and TF motif accessibility in different CNS cell types
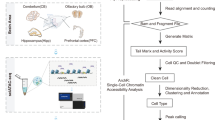
We collected a cohort of 60 tissue samples from 20 postmortem donors ranging in age from 34 to 74 years old, with equal representation of both sexes, and frozen tissue samples from three distinct regions of the CNS from each donor—primary motor cortex (Brodmann area 4 (BA4)), cerebellum (CB) and cervical spinal cord (CSC; Supplementary Table 1). Based on tissue quality metrics5 (Methods), we isolated WM-dominant areas from the tissue, dissociated them into single-nuclei suspensions and performed single-nucleus assay for transposase-accessible chromatin using sequencing (snATAC–seq) using the 10x Genomics Chromium platform (Fig. 1a). After sequencing and stringent quality control (QC) based on the number of unique reads and per-cell transcription start site (TSS) enrichment score (Extended Data Fig. 1a–c), we retained 108,626 nuclei representing all three regions (motor cortex = 55,037 cells, CB = 34,819 cells, CSC = 18,770 cells), with a median of 8,154 fragments per cell, and a median TSS enrichment (TSSe) score of 10.7 (Extended Data Fig. 1b). We built a binned genome count matrix35,36 followed by latent semantic indexing to obtain 16 distinct clusters across the three regions (Fig. 1b,c). To annotate cell types, we constructed a gene-activity matrix and assigned a metagene score to the broad cell types present in the CNS22 (Methods). Using the scores, we identified all major cell types, including cerebellar excitatory neurons (CBEX = 21,801 cells), cerebellar inhibitory neurons (CBINH = 1,301 cells), cortical excitatory neurons (CXEX = 9,671 cells) and cortical inhibitory neurons (CXINH = 4,353 cells). Conversely, the glial cell populations were relatively homogeneous, and we identified MOLs (MOL = 45,325 cells), OPCs (OPC = 5,130 cells), microglia (MIGL = 9,528 cells) and astrocytes (AST = 10,098 cells). We also found pericytes/endothelial cells (ENDO = 1,419 cells; Extended Data Fig. 2c,d). We identified marker genes for the different populations including SOX10 for the OLG lineage, PDGFRA for OPCs, PLP1 for MOLs, AQP4 for AST, AIF1 for MIGL, RBFOX3 for excitatory neurons and GAD1 for inhibitory neurons (Fig. 1c and Extended Data Fig. 2d). We confirmed the validity of our metagene-derived annotations by integrating the gene activity object with a paired single-nuclei transcriptomic dataset of the same cohort5, a previously published dataset37 and a prefrontal adult prefrontal cortex biopsy dataset that we generated for benchmarking (Extended Data Fig. 2a,b).
a, Schematic for snATAC–seq and nanoCUT&Tag experiments in adult human CNS tissue. b, Two-dimensional UMAP of the snATAC dataset colored by clusters and labeled by cell type. c, Gene activity scores for different genes in the identified cell types. d, Two-dimensional UMAP of the H3K27ac nanoCUT&Tag dataset with cell annotations based on integration with snATAC–seq. e, Heatmap showing snATAC–seq differentially accessible peaks across different clusters and cell types. f, nanoCUT&Tag genome browser snapshot showing H3K27ac (top) and H3K27me3 (bottom) pseudobulk signal distribution across different marker genes for each cell type. g, nanoCUT&Tag meta-signal enrichment plots for H3K27ac (green) and H3K27me3 (red) in the MOL population. Top: line plots showing signal enrichment for the two modalities at different peak sets. Middle and bottom: heatmap showing H3K27ac (middle) and H3K27me3 (bottom) signal enrichment across different peak sets. Peak sets (left to right)—H3K27me3 peaks, H3K27ac peaks, ATAC peaks and ATAC peaks from a previously published dataset32. h, Trimodal clustering of the genome highlights patterns of signal distribution across all cell types. i, Correlation matrix of ATAC, H3K27ac and H3K27me3 signals in each cell type shows strong correlation between active marks for individual cell types, and anticorrelation with the repressive H3K27me3. UMAP, uniform manifold approximation and projection; norm., normalized. Schematic in a created in BioRender; Castelo-Branco, G. https://biorender.com/orfbpyi (2025).
We then investigated TF motif accessibility differences in the different cell types in our dataset, using chromVAR38. Clustering on the motif deviations identified marker TFs for the different populations, for instance SOX TFs for OLG, or a specific RORA enrichment in the cerebellar neurons, which is required for cerebellar Purkinje cell maturation39 (Extended Data Fig. 2e,f). We observed lower correlation in endothelial cell annotations between our dataset and the integrated reference datasets (Extended Data Fig. 2a,b). This reduced correlation likely stems from a combination of biological and technical factors, including the inherent low representation of endothelial cells in the dataset and reduced chromatin coverage for this cell type in snATAC–seq data. Consequently, the downstream motif enrichment analysis (Extended Data Fig. 2e) did not reveal clear motif enrichments for endothelial cells, while it is clear for other cell types. Interestingly, while the neurons clustered distinctly according to region and broad electrophysiological profiles, we did not observe the same distinctions within the glial populations (Extended Data Fig. 2g), suggesting that the chromatin states of glial cells may be more region-agnostic and present plasticity to account for their varied functions.
To investigate whether chromatin accessibility changes with age, we analyzed differential accessibility across major cell types in donors younger than 50 (L50) and older than 50 (M50) years. Although age-associated changes were most evident in the neuronal populations, especially in CXEX population (Extended Data Fig. 3a and Supplementary Table 4), the number of nuclei per cell type, region and age group was too small to provide statistical power for additional comparisons and strong conclusions considering the age variable. Sex-based differences were primarily restricted to gonosomal genes (Extended Data Fig. 3b and Supplementary Table 4).
Single-nucleus nanoCUT&Tag H3K27ac and H3K27me3 profiling of major cell populations in the adult human CNS
Histone PTMs provide key functional information about local chromatin state17,40,41,42. We adapted our recently developed nanoCUT&Tag29 for archival human tissue (Methods) and simultaneously mapped H3K27ac (active mark) and H3K27me3 (repressive mark) in three CSCs and three cortical samples from four donors (Fig. 1a,d). Custom nano-Tn5 barcodes enabled demultiplexing of the two modalities, while the shared cell barcode linked each modality to the same cell, yielding 66,113 and 66,727 barcodes for H3K27ac and H3K27me3, respectively, and with 58,696 shared cells bearing both marks29 (Extended Data Fig. 4a,b and Methods). We captured a median of 3,038 and 1,315 unique fragments for H3K27ac and H3K27me3, respectively, comparable to metrics in our published mouse datasets and higher than in a H3K27ac droplet Paired-Tag dataset in the human M1 cortex29,33 (Extended Data Fig. 4c). The fraction of reads in peaks was 0.29 and 0.17 for H3K27ac and H3K27me3, respectively, consistent with the expectations for the nanoCUT&Tag chemistry and with the quality of archival tissue (Extended Data Fig. 4d and Methods). Comparison of snATAC and H3K27ac peaks with ENCODE brain-specific candidate cis-regulatory elements (cCREs) showed strong concordance, with 86.9% overlap between H3K27ac and ENCODE, 90.7% overlap between snATAC and ENCODE and with 78.8% overlap between snATAC and H3K27ac (Extended Data Fig. 4e). In all cell types, most peaks were intronic (median = 49%) and intergenic (median = 26%), consistent with the expected location of regulatory elements.
H3K27ac marks active enhancers and promoters40, and correlates strongly with chromatin accessibility. We therefore integrated the H3K27ac and snATAC datasets (Extended Data Fig. 4f,g and Methods), and, after label transfer, we identified MOL (40,268 cells), CXEX (8,821 cells), MIGL (4,297 cells), AST (2,169 cells), CXINH (1,774 cells), OPC (820 cells) and ENDO (547 cells) populations in the H3K27ac dataset (Extended Data Fig. 4h). Genome browser tracks showed the expected enrichment of the respective signal for each of the identified cell types (Fig. 1f). To ascertain the quality of cell-type-specific histone PTM landscapes, we clustered genes according to their joint H3K27ac and H3K27me3 profiles in each cell type. This revealed distinct groups of genes, with increased active or repressive marks. We then performed a cell-type enrichment analysis on the genes in the active-mark-enriched clusters and confirmed that they were most strongly associated with the same cell types from which they were derived (Extended Data Fig. 5).
To assess antibody specificity, we generated meta-signal plots for both modalities using H3K27ac and H3K27me3 peaks in MOLs. Each mark showed strong enrichment at its corresponding peak set with minimal cross-mapping, confirming signal specificity43. We also verified robust H3K27ac enrichment and the absence of H3K27me3 at snATAC peaks, and in a published bulk H3K27ac dataset32 (Fig. 1g). As no comparable human brain H3K27me3 dataset exists for benchmarking, our data represent a unique resource providing both H3K27ac and H3K27me3 at single-cell resolution in the adult brain and spinal cord (SC).
We then used the ATAC, H3K27ac and H3K27me3 human CNS single-nuclei datasets to perform trimodal clustering of the entire genome and identify patterns and correlations among the three modalities in neural cell types (Fig. 1h and Methods). Although we identified the expected patterns of the three signals across the genome, the overall matrix looked scattered, highlighting not only the complexity of the regulatory genome but also the sparsity of the datasets. We identified different regions of the genome that were (1) accessible in all cell types, likely corresponding to housekeeping genes; (2) specifically accessible in each cell type with corresponding H3K27ac signal; and (3) presented shared H3K27me3 in all cell types (Fig. 1h). We then used the signal in all genomic bins to examine correlations among the different cell types, and observed, as expected, strong anticorrelation between the active ATAC and H3K27ac marks with the inactive H3K27me3 mark (Fig. 1i). Interestingly, chromatin accessibility exhibited a higher correlation across the different cell types overall, unlike H3K27ac, which presented higher correlation between glial populations or between neuronal populations (Fig. 1i). These findings suggest that H3K27ac may be a better discriminant of cell-type-specific regulatory activity compared to chromatin accessibility.
Identification of a new enhancer for the SOX10 gene in human OLGs
Promoters and enhancers exhibit open chromatin and H3K27ac enrichment, making our datasets well-suited to identify cCREs (Extended Data Fig. 6a,b, Supplementary Table 2 and Methods). To validate this, we focused on the SOX10 gene, which has several well-characterized enhancers44. Indeed, the SOX10 promoter was co-accessible with peaks corresponding to the enhancers in OLG, but not other lineages45 (Extended Data Fig. 6a,b). Notably, we found that the promoter was also co-accessible with two distal upstream peaks, previously not associated with SOX10 (Fig. 2a and Extended Data Fig. 6b). One peak corresponded to the ANKRD54 gene promoter, which we found accessible in all cell types. However, the second peak, located 91 kb upstream of SOX10, was accessible only in the OLG lineage, suggesting it may be an uncharacterized OLG-specific enhancer.
a, snATAC–seq genome browser snapshot showing pseudobulk chromatin accessibility signal in OPCs and MOLs at the identified SOX10-distal enhancer (gray) and SOX10 locus (pink) and the corresponding loops identified using Cicero. b, The V plot showing tagmentation pattern and density in a 3.5-kb locus around the SOX10-distal enhancer. Top: density plot showing density of the Tn5 insertion events, seen enriched at the site of the enhancer. Bottom right: fragment size distribution, with subnucleosomal and mononucleosomal bands visible. Bottom left: scatterplot of fragments. Dots represent midpoints of fragments in the OLG lineage and colored by density of fragments. A 330-bp locus zoom-in at the enhancer site. TF footprint is seen as dip in the Tn5 insertion frequency (top) and is marked by a red dotted line. The V plot shows density of fragments at the center of the footprint. A TFAP2A motif was found in the center of the footprint. c, Schematic showing plasmid and lentivirus setup for CRISPRa experiment. d, Gene expression (qPCR) of five genes from CRISPRa experiment with guides targeting either the SOX10 enhancer (turquoise) or an NTC (gray). n = 4 biological replicates; data shown as mean ± s.e.m.; statistics, two-sided t test performed; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. NS, nonsignificant; NTC, nontargeting control. Schematic in c created in BioRender; Castelo-Branco, G. https://biorender.com/3ekzpp9 (2025).
Interestingly, this distal enhancer was co-accessible with the CDC42EP1 promoter, located 300-kb away (Extended Data Fig. 6b). CDC42EP1 encodes the effector protein of CDC42, which is associated with myelin sheath compaction in MOLs46. Notably, we observed increased accessibility of the CDC42EP1 promoter specifically within MOLs, but not OPCs (Extended Data Fig. 6b–d). We checked the co-accessibility links within the OPCs and MOLs separately, and found SOX10 interactions with the canonical enhancers, and the new enhancers in both populations, but the CDC42EP1 promoter–enhancer connection only in MOLs (Supplementary Table 2), suggesting that the chromatin looping concerning this enhancer is altered upon OLG lineage progression. We also queried this locus in our nanoCUT&Tag dataset and observed an increase in the H3K27ac signal at the identified enhancer in MOLs and OPCs, with an increase in H3K27ac at the CDC42EP1 promoter in MOLs specifically (Extended Data Fig. 6e).
This enhancer was evolutionarily conserved across four species, including rhesus monkey and mouse (Extended Data Fig. 6c), and displayed positive PhyloP scores47, characteristic of enhancer evolution48. Using volcano (V) plots49,50 in a 3.5-kb locus spanning the enhancer, we found a strong density of fragments corresponding to the subnucleosomal and mononucleosomal bands, but also intermediate-length fragments, indicative of dynamic TF-bound open chromatin (Fig. 2b). A focused 300-bp window revealed a clear TF footprint in the tagmentation density, corroborated by the expected fragment distribution in the V plot (Fig. 2b). A motif analysis of the core footprint identified a TFAP2A motif coinciding well with the expected binding site from the V plot. The AP2-α TF has been shown to regulate SOX10 expression, but specifically through the U3 enhancer44, which is distinct from this distal enhancer. This suggests that a new distal enhancer may be regulating SOX10 expression in the OLG lineage.
To test enhancer function, we performed a CRISPR activation (CRISPRa) assay in induced pluripotent stem cell (iPSC)-derived human OPCs51,52, using dead-Cas9 fused to the transcriptional co-activator p300 (dCas9–p300 (ref. 53); Fig. 2c). Recruiting dCas9–p300 to the enhancer element increased SOX10 expression but not neighboring genes EIF3L, ANKRD54, and PICK1 or CDC42EP1 (Fig. 2d). Collectively, these findings propose that, in addition to U2 and U3 (Extended Data Fig. 6f), this new distal enhancer can regulate SOX10 in the human OPCs.
Core regulatory TF networks in adult human neural cell types
Using the two datasets, we constructed a core TF regulatory network for each identified cell type based on enhancer and TF motif accessibility (Fig. 3a and Methods). Alongside expected TFs such as OLIG2 in MOLs and OPCs, IRF2 in MIGL and CUX2 and SATB1 in cortical neurons, we identified new regulators, including FOXN2 in MIGL and ZBTB20 and HMX1 in AST (Fig. 3b and Supplementary Table 3). To assess FOXN2 specificity in microglia, we compared it with other forkhead box (FOX) TFs in the network—FOXK1, FOXO3, FOXI1, FOXJ1, FOXJ2 and FOXP3. Although FOXK1 and FOXO3 showed similar expression levels to FOXN2, both had lower regulatory strength. FOXK1 has been implicated in the regulation of STAT1 expression in microglia54, and FOXO3 has well-characterized roles in microglial responses to neuroinflammation and phagocytosis55,56, suggesting that FOXN2 may have a distinct regulatory role in this lineage. In OPCs, we identified PAX3 potentially reflecting a dorsal lineage contribution11, and PRRX1, a regulator of OPC quiescence57, which was absent in MOLs. Notably, with the exception of CAMTA1 in CXINH, TF expression level did not necessarily predict regulatory strength; ZBTB20, which is upregulated after cerebral ischemia in neural progenitor cells58, was the highest expressed TF in the AST, MOL and OPC populations but ranked only 9th, 20th and 29th in regulatory strength, respectively (Fig. 3c,d).
a, Schematic for constructing the regulatory TF circuit and assigning a regulatory strength based on the difference in the incoming and outgoing connections for each TF node in the network. b, Ranked scatterplot showing the strength of core TFs identified within the regulatory network for different cell types. Strength is measured by the difference in number of outgoing connections (out degree) and incoming connections (in degree). Top TFs are marked for each population. c, Same plot as in a, but size and color intensity of each dot represents the average (log-normalized) expression of that TF in the population. The TF with the highest expression in each network is shown in red. d, Network of the top 15 TFs (by expression) in each cell type. Density of edges in the network reflects the correlation between the TF strength and the expression of the TF. e, Scatterplot showing the strength of the shared TFs in the OPC and MOL core networks. Red line represents linear regression line, with 95% confidence interval shown as shaded region. f, Same as in e, but size of the dots represents the difference in the strength of the TF in the OPC and MOL populations. TFs stronger in OPC and MOL are shown in purple and orange, respectively. A subset of the HOX TFs is marked in red. g, Ranked scatterplot highlighting the position of the HOX TFs in the OPC population. h, Same as g, but in MOL population. CRC, core regulatory circuit.
We also investigated how TF networks change during OLG differentiation by comparing OPCs and MOLs. Over 90% TFs in the networks were shared among the two populations and showed comparable regulatory strength (Fig. 2e). However, each population also contained distinct high-strength regulators that may support lineage-specific functions, including ARX, MYC, SIX1 and FOXF2 in OPCs, and NFAT5, ARNT, ZNF566 and ZNF333 in MOLs (Supplementary Table 3).
We then examined TFs whose regulatory strength changed across OLG differentiation and found, among others, BHLHE22 as a strong TF in the MOLs (Fig. 3f). BHLHE22 expression has been shown to increase in OPCs upon T3 stimulation and has a role in differentiation and myelination59. In OPCs, ONECUT1, which regulates NKX6-2 expression, a key TF in OPCs, showed stronger activity. Interestingly, we found several HOX TFs, including HOXA3, HOXA5 and HOXB6, as strong regulators in OPCs, but weaker in MOLs (Fig. 3f and Extended Data Fig. 6g), suggesting higher regulatory potential in progenitors that diminishes with differentiation (Fig. 3g,h). The HOX TFs appeared only in the OLG lineage and not in other cell types, indicating a lineage-specific regulatory role (Supplementary Table 3).
SC adult OLGs exhibit increased accessibility at HOX family genes, decoupled from gene expression
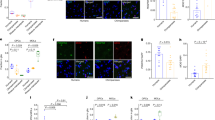
The HOX family of proteins comprises evolutionarily conserved TFs that have key roles in the developmental patterning of the embryonic SC60. As we observed potential regulatory activity in adult OLGs, we asked whether there was regional specificity. Differential accessibility analysis between motor cortex (BA4) and CSC-derived OLGs revealed regionally enriched genes previously reported by snRNA-seq, such as PAX3, SKAP2, SPARC, HCN2 in CSC OLGs and FOXG1, NELL1 in BA4-OLGs5 (Fig. 4a). Notably, 30 HOX cluster genes showed higher accessibility in CSC OLGs than in cortical OLGs (Fig. 4a,b and Extended Data Fig. 6g). Genome browser tracks showed that the OLG lineage and AST, but not MIGL, presented increased HOX accessibility in the SC (Fig. 4c), matching the developmental HOX expression pattern of the cervical spinal cord61.
a, Volcano plot showing differentially accessible peaks in SC and cortical OLGs. Previously characterized marker genes are shown in red and labeled. HOX cluster-associated peaks are shown in blue. Two-sided t test with Benjamini–Hochberg correction. Thresholds, adjusted P = 0.001, log(FC) = 1.5. b, Same as in a, but highlighting the specific HOX clusters identified as being differentially accessible. Two-sided t test with Benjamini–Hochberg correction. c, snATAC–seq genome browser snapshot showing pseudobulk chromatin accessibility signal in AST, MIGL, MOL and OPC populations from the CSC and motor cortex at the HOXA locus. MIGL signal is depleted in both regions, whereas AST, OPC and MOL exhibit accessibility in SC. d, Schematic showing workflow for the multiome experiment and UMAP embedding of the multiome ATAC with cell types annotated using the multiome-RNA. e, Stacked violin plots showing the expression (top) and promoter accessibility (bottom) in cortex and SC-derived MOLs and OPCs from a multiome experiment. Most differentially accessible HOX genes and OLG marker genes are shown. Schematic in d created in BioRender; Castelo-Branco, G. https://biorender.com/orfbpyi (2025).
As promoter accessibility can reflect transcriptional activity, we coprofiled chromatin accessibility and RNA using the 10x Genomics multiome platform on cervical spinal cord and cortical samples (Fig. 4d). Integration with the snATAC dataset confirmed strong concordance between accessibility and expression of core OLG genes, such as PTPRZ1 (OPCs), PLP1 (MOLs) and the TF-encoding SOX10, OLIG1, OLIG2 genes were highly correlated (Fig. 4e). In contrast, eight HOX genes showed elevated accessibility in CSC OLGs, but only residual RNA expression (Fig. 4e). To eliminate sequencing depth as a factor, we analyzed HOX expression in a high-depth single-cell transcriptomic atlas4 of the adult human brain, which revealed similar residual HOX gene expression (Extended Data Fig. 6j), indicating a transcriptional and epigenomic decoupling at HOX loci in adult OLGs, unlike during development.
HOXA/HOXD genes are primed for expression in subsets of SC-derived adult human OLG
Because HOX genes present open chromatin but attenuated expression in adult human OLGs, we examined H3K27ac and H3K27me3 deposition in SC OLGs and cortical OLGs at these loci. Cortical cells displayed a uniform H3K27me3 across the clusters, reflecting the canonical pattern of Polycomb Repressive Complex 2 (PRC2) mediated repression62 (Fig. 5a,b and Extended Data Fig. 7a,b). In contrast, H3K27me3 and H3K27ac exhibited anticorrelated gradients across the HOXA–HOXD clusters in SC OLGs (Fig. 5a,b), reminiscent of HOX gene collinearity during development61,62,63. The 3′ ends of the clusters (for example, HOXA1 to HOXA7) showed elevated ATAC, H3K27ac with reduced H3K27me3, while the 5′ ends (for example, HOXA10 to HOXA13) showed the opposite, with a sharp transition in the middle. This pattern matches the developmental CSC HOX activation profile. Chromatin accessibility was also lower in differentiated postmitotic MOLs relative to OPCs (Fig. 4c), supporting the idea that OPCs retain epigenetic memory of developmental chromatin states, potentially through mitotic inheritance.
a, nanoCUT&Tag and snATAC–seq genome browser tracks showing H3K27ac, H3K27me3 and ATAC pseudobulk signals in OLGs at the HOXA in CSC (upright track, darker shade) and motor cortex (inverted track, lighter shade). b, Same as in a, but for the HOXD locus. c, Gaussian smoothed normalized signal from ATAC (blue), H3K27ac (green) and H3K27me3 (red) across the HOXA cluster with a 50-kb flanking region upstream and downstream. Gray bars show the locations of the cumulative ‘signal boundaries’ identified in each modality. Color intensity reflects the cumulative strength of the signal boundary. d, Same as in c, but with each modality separated out. HOXA directionality is shown at the top, and arrows beneath show the medium (two modalities) and strong (three modalities) signal boundaries. e, nanoCUT&Tag and snATAC–seq genome browser track of the HOXA cluster showing the location of the strong signal boundaries and the corresponding inactive, primed and silenced chromatin domains. f, nanoCUT&Tag and snATAC–seq genome browser tracks showing the ATAC (blue), H3K27ac (green) and H3K27me3 (red) pseudobulk signal in the microglial and astrocyte populations at HOXD in both SC (CSC) and cortex (BA4). g, nanoCUT&Tag genome browser track around the HOXD locus (marked with dotted lines) with H3K27me3 (red) and H3K27ac (green) pseudobulk signal in SC OLGs. Single-cell tracks are shown below and sorted by decreasing H3K27me3 signal. Group 1 cells exhibit moderate H3K27me3 at the 3′ end, while group 2 cells show H3K27me3 depletion, and the amount of H3K27ac remains the same in both groups, suggesting that group 2 cells may be expressing low levels of HOX genes.
To assess domain organization across the HOX clusters, we compared signal transitions across chromatin accessibility, H3K27ac and H3K27me3, which revealed multiple discrete domains with coordinated changes (Methods). Within the HOXA cluster, we identified three strong, one moderate and one weak border (Fig. 5c,d). Two strong borders flanked HOXA1 to HOXA4 and were present in all modalities (Fig. 5c–e). A weak border around HOXA7 marked a region with increased H3K27me3 and increased accessibility, while a strong border at HOXA10 corresponded to the shift from predominantly accessible to predominantly H3K27me3-repressed chromatin. A moderate border marked the decrease of the heavily inactive signature at the 5′ end of the cluster (after HOXA13). The strong border identified at the 3′ end of the cluster suggested further nuance. Although the levels of H3K27me3 at the 3′ end (HOXA1 to HOXA7) were far lower than the 5′ end, it was distinctly greater than the flanking chromatin immediately upstream of the cluster and to the left of the identified border. Thus, we could demarcate the following three regulatory domains around the HOXA cluster: (1) inactive chromatin upstream of the cluster, (2) primed chromatin at the 3′ end and (3) silenced chromatin at the 5′ end (Fig. 5e). While the HOXB and HOXC clusters displayed moderate borders at the 3′ end of the respective clusters, the HOXD cluster displayed strong borders at the 3′ end and in the middle of the cluster, as in the HOXA cluster (Extended Data Fig. 7c). Thus, this indicates three distinct levels of the H3K27me3 repressive mark at HOX loci in adult OLGs, suggesting that this increased level of H3K727me3 at the 3′ end might be sufficient to prevent gene expression and maintain the genes in a primed state. In line with the accessibility pattern seen in SC astrocytes earlier, we also found a similar H3K27ac and H3K27me3 pattern within this population that was absent in cortical astrocytes and in both SC and cortical microglia (Fig. 5f and Extended Data Fig. 7d), suggesting that these priming mechanisms could also be operational in astrocytes.
The multimodal nanoCUT&Tag data enabled joint analysis of H3K27ac and H3K27me3 within the same cell. In the HOXD cluster, all cells showed high H3K27me3 at the 5′ end (HOXD8–HOXD13), but the 3′ end displayed two groups with either medium (group 1) or low (group 2) H3K27me3. In contrast, H3K27ac at the 3′ end was uniform across all cells (Fig. 5g). This suggests that, while most adult OLGs retain a primed repressive state at the HOX loci, a subset of OLGs lacking H3K27me3 but with enriched H3K27ac may be capable of low-level HOX expression and could explain the low RNA expression seen in single-cell transcriptomic studies. Definitive confirmation would require simultaneous single-cell coprofiling of H3K27ac, H3K27me3 and RNA.
Developmental architecture of HOX genes in iPSC-derived human OLGs
HOX gene expression is shaped by chromatin architecture during development62,63. Given the observed epigenetic memory of the developmental state of chromatin at the HOX genes in adult OLGs, we questioned whether the 3D chromatin conformation has a role in the epigenetic state of HOX genes in OLGs. During embryogenesis, HOX activation involves dissolution of a single topologically associated domain (TAD) and formation of two domains (c-Dom and t-Dom) that connect HOX genes to flanking enhancers61,63. To examine the TAD structure in OLGs, we generated high-resolution Micro-C maps64 in iPSC-derived human OPCs51,52 and in human primary B cells to compare them with those of another developmental lineage (Fig. 6a and Extended Data Fig. 8a–e). We were able to capture broad compartment-level information65 and TAD structures66 at a resolution of 5 kb, which corresponded with the well-characterized SOX9–KCNJ2 locus (Extended Data Fig. 8b). The active A compartments65 identified in the OPCs also corresponded to regions of high accessibility in OPCs, further strengthening the validity of the data (Extended Data Fig. 8f). Finally, we could also identify cell-type-specific loops and interactions in both B cells and the human OPCs (Extended Data Fig. 8g).
a, Normalized Micro-C contact matrix at 5-kb resolution at HOXA (left) and HOXD (right) loci and corresponding ATAC, H3K27ac and H3K27me3 tracks in human adult SC OLGs showing the c-Dom and t-Dom TAD structures, including the sub-TAD contacts. Contacts with distal enhancers is shown by the gray bars. b, Micro-C contact matrix showing contacts between HOX genes and flanking enhancers in hOPCs, in contrast to B cells. c, Correlation matrix of ATAC, H3K27ac and H3K27me3 signals in cortex and SC-derived OLGs across all four HOX clusters. d, Contact matrix showing long-range interaction between miR10b and LINC01116, virtual 4c (anchored on LINC01116) H3K27me3, H3K27ac, ATAC and inferred loops are shown. e, scATAC, H3K27ac and H3K27me3 tracks showing signal distribution at HOXD and distal LINC01116 in SC OLGs and cortex OLGs. f, Schematic showing plasmid and lentivirus setup for CRISPRi/a experiment. g, Gene expression changes (qPCR) of five genes from the CRISPRi/a experiment. Left, data from CRISPRa (left, dCas9–p300) and CRISPRi (right, dCas9–KRAB) targeting either miR10b (turquoise), LINC01116 (red) or an NTC (gray). n = 4 biological replicates; data shown as mean ± s.e.m.; statistics, one-way ANOVA with Tukey’s post hoc test; two-sided t test performed; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001. ANOVA, analysis of variance. Schematic in f created in BioRender; Castelo-Branco, G. https://biorender.com/orfbpyi (2025).
Around the HOXA and HOXD clusters, we observed distinct differences between hOPCs and B cells (Fig. 6b,c). In B cells, each cluster was sequestered into a single insulated TAD. In OPCs, however, each cluster was split into two larger TADs, with the 3′ and 5′ ends of the cluster engaging in interactions with distinct upstream and downstream regions (Fig. 6b,c), reminiscent of the developmental c-Dom and t-Dom configuration during activation in the development63. Overlaying these data with accessibility and histone-mark profiles from adult SC OLGs revealed that the boundary among these domains aligned with the strong chromatin border separating primed from repressed HOX regions suggesting the 3′ and 5′ genes of each cluster might be associated with an active and silent TAD, respectively (Fig. 6a). Within the TADs, we also observed subcontacts between regions outside the cluster. Interestingly, within the HOXA active TAD, we observed contact with a region in the SKAP2 locus, which contains a well-known enhancer regulating the expression of 3′ HOXA genes during development67 (Fig. 6a).
We then investigated whether the expression of these HOX genes might be regulated by putative distal cCREs within the TAD domains in hOPCs. We observed a long-range contact between the HOXD-embedded microRNA MIR10B and a distal enhancer within the long noncoding RNA LINC01116 (Fig. 6d), a regulatory contact previously seen in astrocytes68. This enhancer displayed increased accessibility and H3K27ac in SC OLG (Fig. 6e), suggesting that it may be active in these cells. To functionally assess its role in HOXD gene regulation, we performed CRISPRi/a assays (Fig. 6f and Extended Data Fig. 8h). Targeting the enhancer with dCas9–KRAB repressed MIR10B, HOXD1 and HOXD4 gene expression, while activation with dCas9–p300 led to an increase in their expression (Fig. 6g). Notably, HOXD8, situated close to MIR10B but in the adjacent c-Dom, remained insulated from the effects of KRAB and p300 (Fig. 6g). These results suggest that both the histone modification landscape and 3D architecture at the HOX loci regulate their transcriptional state in OLGs.
HOX genes with primed chromatin in SC OLG are activated in high-grade gliomas
Ectopic activation of HOX genes is a feature of many cancers61,69. Midline H3K27M high-grade gliomas (HGGs) have been shown to have OPC origins11,70. These gliomas exhibit strong spatiotemporal specificity and disrupted PRC2 function70,71 (Extended Data Fig. 9a). Additionally, their HOX chromatin architecture faithfully reflects the developmental identity of their cell of origin11.
Given the primed accessibility, histone marks and 3D architecture we observed around HOX loci in SC OLGs, we asked whether this state relates to HOX activation in H3K27M gliomas. Several HOX genes activated in pontine HGGs11, including HOXA1, HOXA3, HOXA5, HOXB4 and HOXD, exhibited primed promoters in SC OLGs but not in cortical OLGs, where they remain H3K27me3-repressed (Fig. 7a). These genes are expressed in posterior fossa group A ependymomas (PFA-EP) tumors and H3.1/H3.3K27M pontine HGG, but not more anterior H3.3K27M thalamic HGGs (Fig. 7b and Extended Data Fig. 9c,d). We also analyzed other SC patterning genes (NKX2-1, NKX6-1, NOTCH1, HES5, HES6, PAX3, PAX6, PAX7, DBX1, IRX3) and cortical patterning genes (IRX3, LX1, DLX2, OTX1, ZIC1, ZIC4) but found that most retained strong H3K27me3, minimal H3K27ac and ATAC signal across promoters in both regions (Extended Data Fig. 9b).
a, nanoCUT&Tag and snATAC–seq normalized promoter accessibility (ATAC), H3K27ac and H3K27me3 signal in SC OLGs (top) and cortical OLGs (bottom) at all HOX genes. Asterisk indicates the genes previously identified to be expressed in pontine HGGs11. b, Normalized Hi–C contact matrix in H3.3K27M pontine HGG11 at the HOXA locus (marked by dotted lines) and corresponding ATAC, H3K27ac and H3K27me3 signals in SC OLGs and H3K27ac and H3K27me3 in H3.3K27M pontine HGG, showing similarity in mark distribution in nondiseased conditions and gliomas. c, Aggregate pileup analysis of hOPC loops (Micro-C) at HOXA (top) and HOXD (bottom) in pontine HGG, PFA-EP and thalamic HGG (left to right)11. d, Insulation score from the Micro-C matrix across a 5-Mb window spanning HOXA (left) and HOXD (right) in the three HGGs (green, pink, yellow, from ref. 11) and B cells (blue) overlaid on the hOPC insulation profile (black).
We then overlaid the chromatin architecture and chromatin immunoprecipitation followed by sequencing (ChIP–seq) data from thalamic and pontine H3.3K27M HGG11. Strikingly, pontine HGG presented H3K27ac/H3K27me3 patterns and sub-TAD structures closely resembling those in human SC OLG and hOPCs (Fig. 6c and Extended Data Fig. 9c,d). Interestingly, aggregate loop analysis around the HOX-associated interactions in hOPCs revealed stronger signals in pontine tumors, but could also be observed in the thalamic and posterior fossa tumors (Fig. 7c). Insulation profiles across HOX loci were comparable between hOPCs and all three tumor types and distinct from B cells (Fig. 7d), although the H3K27me3/H3K27ac distributions varied across tumors (Extended Data Fig. 9c,d). Collectively, these findings suggest that the primed state of these genes in the nondiseased context in OLG in the posterior CNS may drive their expression upon PRC2 disruption in distinct brain tumors, being in line with the regional identity of the cell of origin in HGGs being a key determinant in their arisal11.
Discussion
In this study, we profiled single-cell chromatin accessibility and histone modifications across adult human motor cortex, CB and CSC, generating large-scale snATAC–seq and nanoCUT&Tag datasets and high-resolution Micro-C maps in human OPCs. These integrated epigenomic resources enabled the identification of a previously uncharacterized SOX10 enhancer active in the OLG lineage and provided a framework to investigate lineage-specific and region-specific regulatory states.
We identified elevated chromatin accessibility at HOXA and HOXD clusters in SC OLGs, despite low transcript levels. Accessibility and H3K27ac were concentrated at the 3′ ends, whereas a sharp transition to H3K27me3-enriched chromatin defined the 5′ regions. This pattern suggests that subsets of HOX genes remain in a primed, developmentally derived state. Priming was evident in OLGs and astrocytes but absent in microglia, consistent with their distinct developmental origins. OLGs and astrocytes arise from neuroectoderm that undergoes HOX-dependent patterning, whereas microglia originate from the yolk sac after this patterning is complete, likely explaining the lack of priming in microglia. Epigenetic priming is commonly associated with pluripotent cells72. However, we have recently shown that mouse OPCs can activate immune genes in response to an inflammatory insult and that these immune genes are also maintained in a primed state, potentially to enable rapid activation73,74,75.
HOX genes are well studied in development and remain expressed in various adult lineages76,77. In the oligodendrocyte lineage, however, their patterning identity is largely lost postnatally, with Hox transcripts actively downregulated78, and expressed at lower levels than in neurons79, suggesting different mechanisms of regulation of Hox genes in different neural lineages. In contrast, Hox gene activation is strongly correlated with various cancers61. We observed that chromatin architecture around HOX clusters in SC OLGs closely resembled that of H3.3K27M pontine pediatric HGG cells, arising from OPCs. Promoter priming and long-range enhancer contacts at genes activated in pons HGG suggest these loci are poised for activation in posterior OLGs but not in anterior regions such as cortex, where H3K27me3 likely enforces repression. Thalamic HGGs may instead rely on anterior developmental regulators such as OTX1, ZIC1 and ZIC4 (ref. 11). While some pons-expressed HOX genes were primed in SC OPCs, others were not, likely reflecting regional differences and additional effects of the H3K27M mutation that are not seen in the nonmutant SC OPCs80,81. Nonetheless, the reduced H3K27me3 levels around HOX genes activated in glioma support the idea that primed chromatin states may facilitate reactivation of a subset of these genes in pontine HGGs. However, other regulatory mechanisms beyond those studied here may also be involved in HOX activation and control in adult OLGs.
Our data also suggest that HOX priming may be deleterious, as aberrant HOX activation contributes to tumorigenesis. Within the active HOXD TAD, we observed contact between HOXD3 and LINC01116, whose enhancer can drive astrocyte hyperproliferation when activated, a glioma-like phenotype68. We show that this enhancer functionally regulates 3′ HOXD genes in hOPCs, with both activation and repression altering expression within the same t-Dom but not the adjacent c-Dom, indicating a stable, domain-restricted regulatory architecture. This suggests that the pathway may become activated only in specific cellular contexts.
While our data outline a mechanism for HOX activation in disease, they do not explain why epigenetic memory is maintained; it could simply reflect retained signal from the putative cells of origin, as suggested earlier11. Our findings build upon previous work demonstrating that H3K27me3 is crucial for keeping HOX genes inactive and preserving positional identity76,82,83. We previously found that its depletion derepresses HOX genes in neonatal mouse OPCs74. The simultaneous accessibility and H3K27me3 we observe likely represent a poised, primed state in SC OPCs, although promoter-proximal paused RNA polymerase II may also contribute to maintaining accessibility at these developmental loci84. HOX genes are re-expressed in regenerative contexts, including limb injury, where they are strongly upregulated in regenerative stem cells84. Regeneration is closely linked to high proliferative capacity, and OPCs are both progenitor-like and highly proliferative, particularly after demyelinating injury85. This may underlie the pronounced HOX priming we observe in SC OPCs, which are more proliferative than those in the brain86. Consistent with this, the predicted HOX TF activity from our TF network is higher in OPCs than in MOLs, suggesting that priming may support rapid gene activation during regeneration or remyelination and diminishes with differentiation.
In conclusion, this study provides a comprehensive view of chromatin regulation in adult human neural cells and shows that developmental epigenetic states can persist in adult glia. These latent programs may support regenerative competence but may also predispose cells to aberrant activation in tumorigenic contexts.
Limitations of the study
Our multiome data provide insights into the single-cell histone PTM landscapes of human neural cells in the adult CNS, particularly in the motor cortex and the CSC, revealing region-specific regulation of developmental genes. We identify TFs that act as new putative regulators of specific CNS lineages, which nevertheless will need to be validated with functional experiments in future studies. While our CRISPRa assays support SOX10 enhancer activity, full functional characterization may require a targeted deletion of the enhancer. Our findings highlight that mapping of histone PTMs in different regions is relevant, and it would be of interest to investigate differences between further posterior CNS regions, such as thoracic and lumbar SC, and other anterior brain regions, such as the thalamus and hippocampus, among others. As our findings reveal developmental epigenetic memory relevant for HGG, probing other CNS regions might reveal additional disease susceptibilities.
Our nanoCUT&Tag dataset exhibits increased sparsity compared to our previously published mouse dataset, although we attribute this disparity to the inherent challenges of working with frozen archival tissue, as opposed to fresh mouse tissue. While canonical bivalency refers to the co-occurrence of H3K27me3 and H3K4me3, we find H3K27ac and H3K27me3 at the same genomic locus. However, higher-resolution methods would be needed to ascertain whether the modifications were found on separate H3 tails of the same nucleosome.
Our chromatin architecture data suggest that the HOXA and HOXD clusters may also be divided into two separate domains, which are reflective of the active chromatin architecture seen in development when these genes are being expressed. Interestingly, the strong recapitulation of the spatiotemporal context of HOX gene expression in our chromatin data suggests that there might be a role for epigenetic memory, wherein the cells retain some memory of where they came from. Nevertheless, our Micro-C data have high resolution but are still acquired at the bulk level. Moreover, the human iPSC-derived human OPCs are patterned for forebrain identity. Inducing a more posterior identity in iPSC-derived human OPCs or probing the chromatin architecture of human OLGs across different regions of the adult CNS could further elucidate the extent to which the mechanisms described here are broad.
Methods
Human tissue collection and processing strategy
Adult postmortem fresh-frozen tissue was obtained from the MRC Sudden Death Brain Bank in Edinburgh with full ethical approval (16/ES/0084) and consent. Work in Sweden was performed under the ethical permit 2016/589-31, with amendment 2019-01503, granted by the Swedish Ethical Review Authority (EPM). Tissue was collected from 20 different donors (n = 10 male, n = 10 female) within the ages of 34–74 years old (Supplementary Table 1). No statistical method was used to predetermine sample size. Data collection and analysis were not performed blind to the experimental conditions. Each donor donated fresh-frozen WM from the following three tissue regions: primary motor cortex (BA4), arbor vitae cerebelli and fasciculi cuneatus and gracilis from the CSC. Tissue was processed semirandomly, ensuring that each batch of experiments included both sexes and all three tissue regions. Completed libraries were again randomly multiplexed during sequencing to minimize batch effects.
Tissue dissociation and nuclei isolation
A total of 50–100 mg of frozen tissue were placed in a 1.5 ml tube and chilled in a mortar using liquid nitrogen. A chilled pestle was used to crush the tissue, followed by resuspension in 500 µl of nuclei permeabilization buffer (5% BSA, 0.2% IGEPAL, 1 mM DTT, 1× ethylenediaminetetraacetic acid (EDTA)-free protease inhibitor in PBS). Resuspended tissue was kept on ice for 15 min, with gentle pipetting every 5 min. The homogenized suspension was filtered through a 30 µm filter, followed by a 10 µm filter. An equal volume of 50% iodixanol solution was added and mixed thoroughly. Five-hundred microliter of 29% iodixanol were gently underlaid using a syringe and needle, forming two phases. Samples were centrifuged at 13,500g for 20 min at 4 °C. Supernatant was removed and the nuclear pellet was resuspended in wash buffer (2% BSA in PBS). Samples were spun at 1,000g for 5 min at 4 °C. Supernatant was discarded and the pellet was resuspended in 30 µl 1× diluted nuclei buffer (DNB; snATAC–seq; 10x Genomics) or 30 µl 1× antibody buffer (nanoCUT&Tag; recipe shown in corresponding section) or 30 µl 1× DNB with 1 U µl−1 RNase inhibitor (multiome; 10x Genomics).
Nano-Tn5 purification and loading
Nanobody-Tn5 fusion proteins were purified as described earlier29. Purified enzyme was loaded using barcoded oligonucleotides. First, an equimolar mixture of 100 µM Tn5_P5_MeA_BcdX_0N, Tn5_P5_MeA_BcdX_1N, Tn5_P5_MeA_BcdX_2N and Tn5_P5_MeA_BcdX_3N was mixed with an equimolar amount of 100 µM Tn5_Rev oligonucleotide. The oligonucleotide mixture was denatured by incubating at 95 °C for 5 min in a thermocycler and allowed to anneal slowly by ramping down the temperature by 0.1 °C s−1. Furthermore, 8 µl annealed oligonucleotide, 42 µl glycerol, 44.1 µl 2× dialysis buffer (100 mM HEPES-KOH, pH 7.2, 200 mM NaCl, 0.2 mM EDTA, 2 mM DTT (freshly added), 0.2% Triton X-100, 20% glycerol) and 5.9 µl antimouse nano-Tn5 (5 mg ml−1, 67.7 µM) were combined to obtain a final volume of 100 µl. Alternatively, 8 μl annealed oligonucleotides, 42 μl glycerol, 45.7 μl 2× dialysis buffer and 4.3 µl antirabbit nano-Tn5 (6.8 mg ml−1, 93 µM) were mixed to the same final volume.
snATAC–seq library preparation and sequencing
Dissociated nuclei were counted and incubated at 37 °C for 60 min in the tagmentation mix. Tagmented nuclei were loaded onto the Chromium Chip H (10x Genomics) according to the manufacturer’s instructions. The Chromium Single Cell ATAC Library and Gel Bead Kit (v1.1; 10x Genomics) was used to generate single-nuclei libraries. All libraries were sequenced on the Illumina NovaSeq 6000 with either the S Prime, S1 or S2 flow cell and a 50-8-16-49 read setup.
nanoCUT&Tag
Multinano-CUT&Tag libraries were prepared as described earlier. Briefly, dissociated nuclei in antibody buffer (20 mM HEPES, pH 7.5, 150 mM NaCl, 0.05 mM spermidine, 1× protease inhibitor, 0.05% digitonin, 0.01% IGEPAL, 2% BSA, 2 mM EDTA in dH2O) were counted and 80,000–120,000 nuclei transferred to 0.5-ml microfuge Eppendorf tubes. Nuclei were topped up to 96 µl with antibody buffer. Mouse H3K27me3 antibody (1:100; Abcam, ab6002), rabbit H3K27ac antibody (1:100; Abcam, ab177178), barcoded antirabbit nano-Tn5 (1:100) and barcoded antimouse nano-Tn5 (1:100) were added to the nuclear suspension (final volume of 100 µl). Samples were then incubated overnight at 4 °C on a rotator. After overnight incubation, cells were centrifuged at 600g for 3 min and washed twice with Dig-300 buffer (20 mM HEPES, pH 7.5, 300 mM NaCl, 0.5 mM spermidine, 1× protease inhibitor, 0.05% digitonin, 0.01% IGEPAL, 2% BSA in dH2O). After the second wash, nuclei were resuspended in 100-µl tagmentation buffer (20 mM HEPES, pH 7.5, 300 mM NaCl, 0.5 mM spermidine, 1× protease inhibitor, 10 mM MgCl2, 0.05% digitonin, 0.01% IGEPAL, 2% BSA in dH2O) and incubated at 37 °C for 60 min. Tagmentation was stopped by adding 100-µl STOP buffer (12.5 mM EDTA in 1× DNB (10x Genomics) supplemented with 2% BSA). Nuclei were centrifuged at 600g for 3 min and washed twice with 1× DNB/BSA to remove traces of EDTA. After the second wash, 185 µl of supernatant was removed and nuclei were resuspended in the remaining 15 µl. A total of 2 µl were used for counting (1:5 diluted in trypan blue).
nanoCUT&Tag library preparation and sequencing
Single-cell indexing was performed according to Chromium Next GEM Single Cell ATAC Library and Gel Bead Kit (v1.1; 10x Genomics) instructions. Nuclei of 8 µl were added to 7 µl ATAC buffer B (10x Genomics) and loaded onto the Chromium Chip H. GEM incubation and post-GEM incubation clean-up were performed according to Chromium Next GEM Single Cell ATAC Reagent Kits (v1.1) instructions (step 2.0–3.2). Of the 40 µl of eluted sample, 2 µl was used to measure the concentration using the Qubit dsDNA HS Assay kit. The remaining sample was used for P7 tagmentation by mixing with a tagmentation reaction containing 2× TD buffer (20 mM Tris, pH 7.5, 20% dimethylformamide, 10 mM MgCl2), 1 µl of 10 ng µl−1 MeB-loaded standard Tn5 and dH2O to a final volume of 100 µl, followed by incubation at 37 °C for 30 min in a thermocycler. After tagmentation, samples were purified using DNA Clean and Concentrator-5 (Zymo) according to the manufacturer’s instructions and eluted in 40 µl Zymo elution buffer. Purified DNA was used as input for the Sample Index PCR in the Chromium Next GEM Single Cell ATAC Reagent Kits (v1.1; step 4.1) and samples were amplified for 11–15 cycles. After the Sample Index Double-Sided Size Selection was performed according to the manufacturer’s instructions. Library quality was checked on the Agilent bioanalyzer and sequenced on the Illumina NovaSeq 6000 S Prime flow cell (100c kit) with a custom read1 (R1_seq, 5′-GCGATCGAGGACGGCAGATGTGTATAAGAGACAG-3′) primer, custom index2 (I2_seq, 5′-CTGTCTCTTATACACATCTGCCGTCCTCGATCGC-3′) primer and a 36-8-48-36 read setup.
Multiome library prep
Tissue dissociation and nuclei extraction were performed as described above. A total of 10,000 nuclei were counted and used for bulk tagmentation followed by loading on the Chromium Next GEM Single Cell Chip J. Single-cell indexing and library preparation were performed using the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression kit, according to the manufacturer’s instructions. Libraries were sequenced on the Illumina NovaSeq 6000 S Prime flow cell (100c kit), with a 50-8-24-49 read setup.
iPS-derived hOPC cell cultures
hOPCs were derived from the human iPS line C27 (ref. 87) in Steven Goldman’s lab, with the protocol described in ref. 51. Work in Sweden was performed under the ethical permit 2020-00398, with amendment 2023-04598-02, granted by the Swedish EPM. Corning six-well cell culture plates were precoated with poly-L-ornithine (PLO; Sigma-Aldrich, P4957-50ML) and incubated for 1 h at 37 °C. PLO was removed and wells were rinsed thrice using sterile 1× DPBS−/− (Thermo Fisher Scientific, 14190144) followed by overnight incubation with 5 µg ml−1 laminin (Corning, 354232) in HBSS+/+ (Thermo Fisher Scientific, 24020117). After removing the laminin, 1 million iPS-derived hOPC cells (C27 line) were directly seeded into the plate and expanded for 3 weeks before splitting. Cells were cultured in proliferation media (DMEM/F12 (Invitrogen, 11330-057) containing 1× B27 (Invitrogen, 12587-010), 1× N1 (Sigma-Aldrich, N6530), 1× NEAA (Invitrogen, 11140-050), 60 ng ml−1 T3 (Sigma-Aldrich, T5516-1MG), 1 µM dcAMP (Sigma-Aldrich, D0260), 100 ng ml−1 biotin (Sigma-Aldrich, B4639), 10 ng ml−1 PDGF-AA (R&D, 221-AA-50), 10 ng ml−1 IGF-1 (R&D, 291-G1-050) and 10 ng ml−1 NT3 (R&D, 267-N3-025)), which was refreshed every 2 days.
ATAC–seq in hOPCs
ATAC–seq was performed as described previously7. A total of 60,000 cultured hOPCs were collected, washed with 1× PBS, and incubated in lysis buffer (0.1% IGEPAL, 10 mM Tris–HCl, pH 7.4, 10 mM NaCl, 3 mM MgCl2) on ice for 5 min. Lysed cells were centrifuged at 500g for 20 min at 4 °C. The nuclei pellet was resuspended in tagmentation mix (2× TD buffer, Tn5 enzyme, in dH2O) and incubated at 37 °C for 30 min. Tagmented DNA was purified using the Qiagen minElute Purification kit and PAGE purified to remove adaptor dimers. Libraries were sequenced on an Illumina NovaSeq 6000 with a 50-8-8-50 read setup.
B cell collection
Peripheral mononuclear cells were freshly isolated by Ficoll (GE Healthcare, 17-1440-03) gradient centrifugation from buffy coats obtained through Karolinska University Hospital of three healthy female donors (age = 28, 29 and 39 years). Study procedures were conducted under ethical permit 2009/2107-31-2 approved by the Swedish EPM. B cells were then enriched by negative selection using the EasySep Human B Cell Enrichment Kit II (STEMCELL Technologies, 17963) without CD43 depletion, according to the manufacturer’s instructions. B cells were then stained for 30 min on ice with anti-CD3 (clone SK7; 560176), anti-CD14 (clone MφP9; 560180), anti-CD16 APC-Cy7 (clone 3G8; 560195), anti-CD19 APC (clone HIB19; 561742), anti-IgG BV510 (clone G18-145; BD Biosciences, 563247), anti-CD27 PE-Cy5.5 (clone 0323; Novus Biologicals, NBP1-43426) or BV711 (clone O323; 302833), anti-IgD pacific blue (clone IA6-2; 348224), anti-IgM BV570 (clone MMH-88; 314517) and Zombie NIR fixable viability dye (BioLegend, 423106). Cells were then washed in PBS (Sigma-Aldrich, D8537) and filtered. Between 106 and 2 × 106 CD27+CD19+ live B cells were sorted using a SH800 Cell Sorter (Sony) into an RPMI medium (R8758) with 10% heat-inactivated fetal bovine serum (F7524), 100 U ml−1 penicillin and 100 μg ml−1 streptomycin (Sigma-Aldrich, P4458). Cells were washed in PBS, centrifuged and stored at −80 °C as dry pellet before proceeding with Micro-C.
Processing of human prefrontal cortex biopsy for scRNA-seq
The scRNA-seq prefrontal cortex sample was selected from a cohort of patients with hydrocephalus, a 76-year-old female, planning to undergo cerebrospinal fluid (CSF) diversion surgery either with a ventriculoperitoneal shunt placement or ventriculocisternostomy and without diagnosed CNS malignancy, hematoma, infection or inflammation. Patients received oral and written information and were required to sign an informed consent. The biopsy procedure was performed by a standard operative procedure under general anesthesia. The placement of burr hole on the frontal lobe followed by an opening of dura and cortex was performed in a routine fashion for ventriculoperitoneal shunt or ventriculocisternostomy surgery. The biopsy was sharply dissected with knife through the leptomeninges, forming a tissue block consisting of cortex and subcortex, and placed in a sterile tube containing CSF. Biopsies were immediately placed on ice and transported to the nearby laboratory. After biopsy was taken, hemostasis was achieved using diathermy and the CSF diversion surgery continued. The wound was closed routinely. No surgical insults were detected in the study population. Patients and relatives received oral and written information about the study before inclusion and provided signed informed consent on hospital admission. Patients who did not wish to participate or were unable to understand the information or provide signed informed consent were excluded. Patient data and samples were anonymized for the research group. The study was approved by the Stockholm Region’s ethical committee (2016/1062-31/2 and 2018/843-32). Fresh tissue sample processing was performed within the hours before surgery. The tissue was dissociated, followed by red cell and debris removal, and cell counting for sequencing (scRNA-seq; 10x Genomics). After dissociation, only non-neuronal cells were retrieved.
FASTQ files of prefrontal cortex scRNA-seq were processed using Cellranger count (v6.12) and aligned to the GRCh38 genome assembly. Gene count matrix was processed with Seurat (v4). For QC filtering and downstream dimension reduction and clustering, cells with filters nFeature_RNA > 200 & nFeature_RNA < 10,000 & percent_mito ≤ 5 were retained, which led to a final dataset of 2,798 cells. Cells were annotated using canonical markers and compared to the full scATAC–seq dataset and multiome datasets from this study, using the gene activities from the scATAC–seq as a proxy of the RNA expression. The final dataset included different non-neuronal cell types, 186 astrocytes (ASTRO), 714 microglia (MIGL), 1,826 oligodendrocytes (OL) and 72 OPCs. Integration with scATAC–seq cortex data was performed with Seurat (v4) FindTransferAnchors on the intersecting variable features (from the top 3,000 most variable features). Anchors were identified between the scRNA-seq prefrontal cortex genes and the gene activities from the corresponding genes on the scATAC–seq dataset. Predicted labels were transferred using Labeltransfer with dims = 1:20 and weight.reduction = ‘cca’.
Micro-C
Micro-C was performed by the National Genomics Infrastructure, Stockholm, Sweden, using the commercially available Micro-C kit (Dovetail Genomics, 21006), with 100,000–200,000 iPS-derived hOPCs as input. Briefly, cells were crosslinked and enzymatically digested using MNase to allow for nucleosome-resolution fragmentation. Free ends were ligated with biotin-containing adaptors. Ligated fragments were reverse-crosslinked and amplified to introduce sequencing handles. Libraries were generated in three separate biological replicates and were sequenced as a pilot on the Illumina NovaSeq 6000 S Prime flow cell with a 2 × 150 bp (300c) kit with a 151-19-10-151 read setup. After QC of the pilot and checking for library complexity (with the ‘preseq’ tool), two of three hOPC replicates and all three B cell replicates were resequenced on a large S4 flow cell to a depth of 6 billion reads.
gRNA cloning strategy
Four guides per locus were cloned by annealing 1 µl of each complementary oligo (Supplementary Table 5) to a final volume of 25 µl annealing buffer (combine 500 µl 1 M Tris–HCl, pH 8.0, with 500 µl 5 M NaCl to final volume of 50 ml with ddH2O) and incubated at 95 °C for 3 min, cooled to 22 °C at 0.1 °C s−1 and diluted in 75 µl ddH2O to yield 1 µM. Lentiviral U6–GG–acceptor plasmid was prelinearized using BsmBI (20 U µl−1; NEB, R0580) and BsiWI-HF (20 U µl−1; NEB, R3553) and isolated on gel and aliquoted at 30 ng µl−1. T4 ligation was carried out mixing 1 µl of 1 µM annealed oligos, 30 ng linearized GG–acceptor plasmid, 0.2 µl BsmBI (20 U µl−1), 0.2 µl BsiWI-HF (20 U µl−1), 0.5 µl T4 DNA ligase (40 U µl−1; NEB, M0202), 1 µl 10× T4 DNA ligase buffer and 6.1 µl ddH2O and incubating for 10 min at 20 °C, followed by 5 min at 37 °C and 5 min at 80 °C. One microliter of the ligation reaction was added to 10 µl of competent cells and incubated on ice for 10 min. Cells were heat-shocked at 42 °C for 30 s, returned to ice for 5 min, and then supplemented with 100 µl SOC medium. The cells were incubated at 37 °C for 60 min with shaking at 250 rpm and plated on agar plates containing ampicillin for overnight selection. Colonies were isolated and expanded in liquid LB medium prior to plasmid isolation using the ZymoPURE Plasmid Miniprep Kit (Zymo Research, D4210) according to the manufacturer’s instructions. Purified plasmids were verified by Sanger sequencing using the primer 5′-CGATACAAGGCTGTTAGAGAG-3′.
Lentiviral packaging
dCas9–KRAB–BFP, dCas9–p300–BFP and U6–GG–acceptor lentiviral plasmids were a gift from CRISPR Functional Genomics, Karolinska Institute. Lentivirus was packaged in HEK293 cells cultured in DMEM (Gibco, 31966021) supplemented with 10% FBS (Gibco, 10500064) on precoated poly-l-lysine (Sigma-Aldrich, P4707) culture ware seeded to be 70–80% confluent after >24 h. A total of 16 µg of transfer vector, pCMV-VSV-G, pRSV-REV and pCgpV (Takara, 631278) were mixed at a ratio of 3:1:1:1 with 32 µl lipofectamine-2000 (Thermo Fisher Scientific, 11668030) and Opti-MEM (Gibco, 31985070) according to the manufacture’s protocol and added to the cells. Viral particles were collected from the media after 48 h, centrifuged at 500g for 10 min and filtered through 0.45 µm before incubating with Lenti-X concentrator (Takara, 631231) at a ratio of 3:1 and incubated for >30 min at 4 °C, before centrifugation at 1,500g for 45 min at 4 °C and resuspended in DMEM at a 20× or 100× concentration.
Lentiviral transduction of hOPCs
Concentrated lentivirus was added for <24 h after seeding to the media at a ratio of 1:4:1:3, and growth factors and supplements were adjusted accordingly. dCas9–BFP and gRNA-mCherry transduced hOPC at a rate of 25–20% and >90%, respectively. After 24h, the media was replaced, and cells were kept for 3 days before collecting using miRNeasy Micro Kit (Qiagen, 217084) according to the manufacturer’s protocol. cDNA library was prepared using High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific, 4368814). RT-qPCR was performed on a StepOnePlus System (Applied Biosystems) in triplicate and with reverse transcriptase-negative reactions to control for genomic DNA. Fast SYBR Green Master Mix (Applied Biosystems, 4385616) was used according to the manufacturer’s protocol, each PCR reaction had a final volume of 10 µl and 2 µl of diluted cDNA and reverse transcriptase, respectively, and incubated at 20 s at 95 °C, 40× cycles of 3 s of 95 °C and 30 s of 60 °C, followed by 15 s at 95 °C, 60 s at 60 °C and 15 s at 95 °C. Melting curve was obtained for each PCR product after each run, to control for primer dimers and gene-specific peaks. Expression levels were calculated by dividing the quantity by the geometric mean of the housekeeping genes.
RNA extraction, cDNA synthesis and qRT–PCR
Cells were collected with QIAzol (Qiagen) and stored at −80 °C until further processing. RNA was extracted with the miRNeasy mini kit (Qiagen) according to the manufacturer’s instructions. Contaminating DNA was degraded by the treatment of the samples with RNase-free DNase (Qiagen) in column. Thus, 350 ng RNA was used to synthesize cDNA with the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems), including RNase inhibitor (Applied Biosystems), with annealing for 10 min at 25 °C and extending for 2 h at 37 °C and inactivation for 5 min at 85 °C. The cDNA was diluted 1:5 in H2O and 2.5 μl was used in the qRT–PCR reactions with 2 μl of Fast SYBR Green Master Mix (Applied Biosystems) and 5 pmol of each primer in a final volume of 10 μl. The reactions were run on a StepOnePlus System (Applied Biosystems) in duplicate and with reverse transcriptase-negative reactions to control for genomic DNA. The running conditions were 20 s at 95 °C, followed by 40 cycles of 3 s of 95 °C and 30 s of 60 °C, then 15 s at 95 °C, 1 min at 60 °C and 15 s at 95 °C. Melt curves were generated to control primer dimers and gene-specific peaks. Relative standard curves for each gene were generated to obtain relative expression values. ‘GAPDH’ and ‘b-Act’ were run as housekeeping genes. Expression levels were then calculated by dividing the relative expression value by the geometric mean of the housekeeping genes. Samples were normalized per experiment. Data distribution was assumed to be normal, but this was not formally tested.
snATAC–seq data preprocessing and QC
FASTQ files generated from sequencing were processed using ‘cellranger-atac count’ with the default parameters. Samples were aggregated using the ‘cellranger-atac aggr’ with default parameters, but with the normalization omitted using the flag ‘—normalize=none’.
Thus, 2-kb count matrices were built using a custom script ‘build_large_mtx.py’ that is a modified version of episcanpy’s88,89 build_atac_mtx.py script and allows for reading in files in batches. TSSe scores were generated using the ArchR90 package and only cells with TSSe >7 and number of unique fragments of >3,000 were retained.
snATAC–seq peak calling
The Fragments file (fragments.tsv.gz) was split according to the cell-type annotation and peaks were called using the ‘callpeak’ function from MACS2 (ref. 91) with the following parameters: ‘-f BED -g hs -q 0.05 –shift -100 –extsize 200 –nomodel –call-summits –keep-dup = 1’. Peak annotation was performed using the HOMER92 annotatePeaks.pl function and a custom GTF file with miRNA and snoRNA removed.
snATAC–seq downstream analysis
After cell filtering, the top 100,000 features were retained and normalization was performed using term frequency-inverse document frequency, followed by singular vector decomposition. A nearest-neighbor graph was built in the lower-dimensional space, followed by Leiden clustering. The highly variable features from each cluster were retained and used to repeat the term frequency-inverse document frequency, singular vector decomposition, graph building and clustering for a total of three iterations. Batch correction was performed using Harmony93. Differentially accessible regions were identified using both the ‘rank_features’ function in episcanpy (v0.3.2) with Benjamini–Hochberg correction for multiple testing as well as the diffxpy (https://github.com/theislab/diffxpy/) package with sex, age and tissue added as covariates.
Gene activity matrix and cell-type annotation
A gene activity matrix was built using a 5 kb promoter region flanking the TSS of genes. The count matrix was smoothened using MAGIC94 with default parameters to improve the signal. The top 50 distinct differentially accessible genes for all cell types found in our published snRNA-seq dataset were used as cell-type metagenes and aggregate gene activity was calculated for all genes within each cell-type metagene, generating a metagene score for each cell. Metagene scores for the different cell types and individual marker genes were used to assign the clusters to broad cell types. We could not identify SC-derived neurons, although it is known that these neurons are particularly sensitive and susceptible to hypoxia, possibly leading to difficulty in isolating them. An overwhelming proportion of all cerebellar cells in the dataset was composed of the CBEX cells (also known as cerebellar granular cells). These cells are tiny and densely packed within the granular layer of the CB, close to the gray matter–WM border. We suspect that the skewed distribution may have arisen from imprecise dissection during the collection of WM from the tissue.
Integration with snRNA-seq data
The annotation of the cell types based on transcriptome data from external datasets was performed as discussed in refs. 5,37. The expression matrix mentioned in the ref. 37 was downloaded from Gene Expression Omnibus repository with the accession GSE118257, and then converted to Seurat95 (v4.3.0.1) object. The dataset mentioned in ref. 5 was downloaded from https://cellxgene.cziscience.com/collections/9d63fcf1-5ca0-4006-8d8f-872f3327dbe9 as a Seurat object including all the cell types. scATAC h5ad file was converted to Seurat using SeuratDisk R library with the Convert function with assay = ‘peaks’ to h5Seurat format. Gene activities were calculated based on the chromatin accessibility signal using ENSEMBL gene annotations (EnsDb.Hsapiens.v86). For each annotated gene, the region of the promoter (500 bp upstream of the annotated TSS) and the gene body were considered (promoter + genebody). Gene activities were calculated using FeatureMatrix function from Signac96 (v1.10.0) with the promoter + genebody in GRanges format, from GenomicRanges97 library, and the cellranger-atac 2.0 fragments.tsv for all the samples. Then, they were added to the Seurat/Signac object as a new assay. Label transfer was performed using Seurat. The FindIntegrationAnchors function was applied to find anchors between scRNA reference dataset and the query scATAC gene activities. The FindTransferAnchors function was used to find transfer anchors from each reference to the query, and then the resulting anchors were used to perform label transfer with the TransferData function, using canonical correlation analysis with 20 dimensions.
Trimodal genomic clustering
A total of 10-kb genomic bins spanning the hg38 genome were used as input peaks to the deepTools98 compute-matrix function. ATAC, H3K27ac and H3K27me3 bigwig files from each population were used to generate a matrix of normalized signal in each genomic bin. The mean signal across each bin across each modality and each cell type was used to hierarchically cluster the bins (rows) and celltype + modality (columns). Pearson correlation was used to identify the correlation among each column (identifying similar and dissimilar celltype + modalities based on whole-genome patterns).
Co-accessibility analysis
Co-accessible regions were identified using Cicero99. Cell-type-specific pseudobulk BAM files and single-cell count matrices were provided as input to identify pairs of genomic bins with increased co-accessibility across different cells of a population. Co-accessibility score cutoffs of 0.25 and 0.5 were used to identify significant interactions and high-confidence interactions, respectively.
TF regulatory network
The Core Regulatory Circuit package100 was used to identify the core TF network. Briefly, H3K27ac BAM and bigwig files and Super Enhancers (using ROSE algorithm) for each cell population were used as input to FIMO. FIMO then evaluates TF motif enrichment through a P-value cutoff based on a log-likelihood ratio test. For the OPC and CXINH populations, the ATAC–seq data (BAM, bigwig, Super Enhancers) were used instead due to the sparsity of the H3K27ac data101. A network was built by inferring the number of interacting TF motifs in the proximal super enhancer of a TF. The TF strength was assigned based on the difference between the number of outbound edges (regulated TFs) and inbound edges (targeting TFs). A higher score suggests that a TF has a stronger regulatory influence over other TFs.
TF footprinting analysis
Footprinting analysis was performed using the Regulatory Genomics Toolbox102. Briefly, peaks were called for the different cell types, and the HMM-based Identification of TF footprints (HINT) framework was applied to identify active TF-binding sites using default parameters.
nanoCUT&Tag signal enrichment
The k-means algorithm (k = 10) implemented in deepTools (v3.5.1) was used to cluster all genes (1-kb-padded TSS) based on the H3K27ac and H3K27me3 signals for each cell type. We then queried the genes identified in the clusters that displayed high H3K27ac and low H3K27me3 and used gget enrichr103, predicting the enriched cell type based on the genes (database = ‘celltypes’).
Micro-C balancing and transformation
Raw contact matrices were normalized and balanced using iterative correction and eigen decomposition as implemented in the ‘cooler’104 package. The ‘hicTransform’ package was used to generate observed/expected counts with the ‘–method obs_exp’ flag.
Compartment analysis and aggregate pileup analysis
Micro-C compartments were identified by computing the principal component analysis eigenvectors of the hic matrix using the ‘hicPCA’ command from the hicExplorer package105,106,107. Aggregate pileup was performed using the ‘coolpuppy’ package108. Briefly, signal aggregation was done in all hic matrices (hOPCs, B cells, PFA-EP, pontine HGG, thalamic HGG) at genomic coordinates corresponding to the identified loops.
Insulation and boundary strength analysis
The ‘cooltools’109 Python API was used to process the contact matrices and identify the insulation scores within the normalized contact frequency data. Briefly, a diamond-shaped window is used to slide along the genome, with one of the corners on the main diagonal of the contact matrix, and contacts within the window at each position are summed up. Windows with low sums are marked as putative boundaries and as insulating regions immediately upstream and downstream. Boundaries were identified in the 10-kb contact matrix with a sliding window size of 100 kb.
Loop calling and virtual 4C analysis
Virtual 4C identifies loci that exhibit increased contact frequency with a reference locus of interest (viewpoint analysis) and was performed using the ‘hicPlotViewpoint’ function in the HiCExplorer106 package. Loops were called on the 5-kb contact matrix using the ‘mustache’110 package and looking within a maximum distance of 100 Mb.
Additional computational analysis
Description of the following computational analysis is given in Supplementary Methods: motif analysis, integration of multiome ATAC with snATAC–seq data, nanoCUT&Tag data preprocessing and cell calling, nanoCUT&Tag peak calling and bigwig track generation, integration of H3K27ac nanoCUT&Tag with snATAC–seq data, regulatory domain border identification, micro-C data preprocessing and Hi–C to Cool matrix conversion.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw Human data have been deposited in the European Genome-Phenome Archive (EGA) under EGA accession EGAD50000000410 for the scATAC–seq, nanoCT-seq, SC multiome and hOPCs Micro-C data (EGAD50000001542) for the scRNA-seq from the human prefrontal cortex biopsy and EGAD50000001535 for B cells Micro-C data. Browsable UMAPS and tracks are available at UCSC Cell Browser and UCSC Genome Browser34 (https://cns-nanocuttag-atac.cells.ucsc.edu) and https://ki.se/en/mbb/oligointernode. Source data are provided with this paper.
Code availability
All code is available at https://github.com/mkabbe/snATACnanoCT_AdultHumanCNS.
References
Jäkel, S. & Dimou, L. Glial cells and their function in the adult brain: a journey through the history of their ablation. Front. Cell. Neurosci. 11, 24–24 (2017).
Fünfschilling, U. et al. Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature 485, 517–521 (2012).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023).
Seeker, L. A. et al. Brain matters: unveiling the distinct contributions of region, age, and sex to glia diversity and CNS function. Acta Neuropathol. Commun. 11, 84 (2023).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Zhang, K. et al. A single-cell atlas of chromatin accessibility in the human genome. Cell 184, 5985–6001 (2021).
Domcke, S. et al. A human cell atlas of fetal chromatin accessibility. Science 370, eaba7612 (2020).
Mannens, C. C. A. et al. Chromatin accessibility during human first-trimester neurodevelopment. Nature 647, 179–186 (2025).
Jessa, S. et al. K27M in canonical and noncanonical H3 variants occurs in distinct oligodendroglial cell lineages in brain midline gliomas. Nat. Genet. 54, 1865–1880 (2022).
Liu, I. et al. The landscape of tumor cell states and spatial organization in H3-K27M mutant diffuse midline glioma across age and location. Nat. Genet. 54, 1881–1894 (2022).
Li, Y. E. et al. A comparative atlas of single-cell chromatin accessibility in the human brain. Science 382, eadf7044 (2023).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Thornton, C. A. et al. Spatially mapped single-cell chromatin accessibility. Nat. Commun. 12, 1274 (2021).
Corces, M. R. et al. Single-cell epigenomic analyses implicate candidate causal variants at inherited risk loci for Alzheimer’s and Parkinson’s diseases. Nat. Genet. 52, 1158–1168 (2020).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Liu, H. et al. DNA methylation atlas of the mouse brain at single-cell resolution. Nature 598, 120–128 (2021).
Grosselin, K. et al. High-throughput single-cell ChIP–seq identifies heterogeneity of chromatin states in breast cancer. Nat. Genet. 51, 1060–1066 (2019).
Patty, B. J. & Hainer, S. J. Transcription factor chromatin profiling genome-wide using uliCUT&RUN in single cells and individual blastocysts. Nat. Protoc. 16, 2633–2666 (2021).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Bartosovic, M., Kabbe, M. & Castelo-Branco, G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat. Biotechnol. 39, 825–835 (2021).
Wu, S. J. et al. Single-cell CUT&Tag analysis of chromatin modifications in differentiation and tumor progression. Nat. Biotechnol. 39, 819–824 (2021).
Stuart, T. et al. Nanobody-tethered transposition enables multifactorial chromatin profiling at single-cell resolution. Nat. Biotechnol. 41, 806–812 (2023).
Zhu, C. et al. Joint profiling of histone modifications and transcriptome in single cells from mouse brain. Nat. Methods 18, 283–292 (2021).
Xie, Y. et al. Droplet-based single-cell joint profiling of histone modifications and transcriptomes. Nat. Struct. Mol. Biol. 30, 1428–1433 (2023).
Lochs, S. J. A. et al. Combinatorial single-cell profiling of major chromatin types with MAbID. Nat. Methods 21, 72–82 (2024).
Xiong, H., Wang, Q., Li, C. C. & He, A. Single-cell joint profiling of multiple epigenetic proteins and gene transcription. Sci. Adv. 10, eadi3664 (2024).
Bartosovic, M. & Castelo-Branco, G. Multimodal chromatin profiling using nanobody-based single-cell CUT&Tag. Nat. Biotechnol. 41, 794–805 (2023).
Tian, W. et al. Single-cell DNA methylation and 3D genome architecture in the human brain. Science 382, eadf5357 (2023).
Ziegler, K. C. et al. The brain neurovascular epigenome and its association with dementia. Neuron 114, 268–286 (2026).
Nott, A. et al. Brain cell type-specific enhancer-promoter interactome maps and disease-risk association. Science 366, 1134–1139 (2019).
Zemke, N. R. et al. Conserved and divergent gene regulatory programs of the mammalian neocortex. Nature 624, 390–402 (2023).
Speir, M. L. et al. UCSC Cell Browser: visualize your single-cell data. Bioinformatics 37, 4578–4580 (2021).
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Chen, X. R. et al. Mature Purkinje cells require the retinoic acid-related orphan receptor-α (RORα) to maintain climbing fiber mono-innervation and other adult characteristics. J. Neurosci. 33, 9546–9562 (2013).
Gräff, J. & Tsai, L. H. Histone acetylation: molecular mnemonics on the chromatin. Nat. Rev. Neurosci. 14, 97–111 (2013).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Ferrari, K. J. et al. Polycomb-dependent H3K27me1 and H3K27me2 regulate active transcription and enhancer fidelity. Mol. Cell 53, 49–62 (2014).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem. Cell 125, 315–326 (2006).
Wahlbuhl, M., Reiprich, S., Vogl, M. R., Bösl, M. R. & Wegner, M. Transcription factor Sox10 orchestrates activity of a neural crest-specific enhancer in the vicinity of its gene. Nucleic Acids Res. 40, 88–101 (2012).
Werner, T., Hammer, A., Wahlbuhl, M., Bösl, M. R. & Wegner, M. Multiple conserved regulatory elements with overlapping functions determine Sox10 expression in mouse embryogenesis. Nucleic Acids Res. 35, 6526–6538 (2007).
Thurnherr, T. et al. Cdc42 and Rac1 signaling are both required for and act synergistically in the correct formation of myelin sheaths in the CNS. J. Neurosci. 26, 10110–10119 (2006).
Ion Reporter. phyloP score. https://ionreporter.thermofisher.com/ionreporter/help/GUID-03D1F68A-E646-4B49-AD59-AF2F51874BD2.html (2024).
Rebeiz, M. & Tsiantis, M. Enhancer evolution and the origins of morphological novelty. Curr. Opin. Genet. Dev. 45, 115–123 (2017).
Henikoff, J. G., Belsky, J. A., Krassovsky, K., MacAlpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011).
Schep, A. N. et al. Structured nucleosome fingerprints enable high-resolution mapping of chromatin architecture within regulatory regions. Genome Res. 25, 1757–1770 (2015).
Wang, S. et al. Human iPSC-derived oligodendrocyte progenitor cells can myelinate and rescue a mouse model of congenital hypomyelination. Cell Stem Cell 12, 252–264 (2013).
Windrem, M. S. et al. Neonatal chimerization with human glial progenitor cells can both remyelinate and rescue the otherwise lethally hypomyelinated shiverer mouse. Cell Stem Cell 2, 553–565 (2008).
Hilton, I. B. et al. Epigenome editing by a CRISPR–Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Meleady, L. et al. Histone deacetylase 3 regulates microglial function through histone deacetylation. Epigenetics 18, 2241008 (2023).
Lee, J.-W. et al. TLR4 (Toll-like receptor 4) activation suppresses autophagy through inhibition of FOXO3 and impairs phagocytic capacity of microglia. Autophagy 15, 753–770 (2019).
Yan, D. et al. SIRT1/FOXO3-mediated autophagy signaling involved in manganese-induced neuroinflammation in microglia. Ecotoxicol. Environ. Saf. 256, 114872 (2023).
Wang, J. et al. Paired related homeobox protein 1 regulates quiescence in human oligodendrocyte progenitors. Cell Rep. 25, 3435 (2018).
Doeppner, T. R., Herz, J., Bähr, M., Tonchev, A. B. & Stoykova, A. Zbtb20 regulates developmental neurogenesis in the olfactory bulb and gliogenesis after adult brain injury. Mol. Neurobiol. 56, 567–582 (2019).
Dugas, J. C., Ibrahim, A. & Barres, B. A. The T3-induced gene KLF9 regulates oligodendrocyte differentiation and myelin regeneration. Mol. Cell. Neurosci. 50, 45–57 (2012).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Gonçalves, C. S., le Boiteux, E., Arnaud, P. & Costa, B. M. HOX gene cluster (de)regulation in brain: from neurodevelopment to malignant glial tumours. Cell. Mol. Life Sci. 77, 3797–3821 (2020).
Noordermeer, D. et al. The dynamic architecture of Hox gene clusters. Science 334, 222–225 (2011).
Rekaik, H. et al. Sequential and directional insulation by conserved CTCF sites underlies the Hox timer in stembryos. Nat. Genet. 55, 1164–1175 (2023).
Hsieh, T.-H. S. et al. Mapping nucleosome resolution chromosome folding in yeast by Micro-C. Cell 162, 108–119 (2015).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Cao, K. et al. SET1A/COMPASS and shadow enhancers in the regulation of homeotic gene expression. Genes Dev. 31, 787–801 (2017).
Deforzh, E. et al. Promoter and enhancer RNAs regulate chromatin reorganization and activation of miR-10b/HOXD locus, and neoplastic transformation in glioma. Mol. Cell 82, 1894–1908 (2022).
Bhatlekar, S., Fields, J. Z. & Boman, B. M. HOX genes and their role in the development of human cancers. J. Mol. Med. 92, 811–823 (2014).
Nagaraja, S. et al. Histone variant and cell context determine H3K27M reprogramming of the enhancer landscape and oncogenic state. Mol. Cell 76, 965–980 (2019).
Li, X.-N. Defining the cell of origin for diffuse midline gliomas. Nat. Genet. 54, 1770–1771 (2022).
Madrigal, P. et al. Epigenetic and transcriptional regulations prime cell fate before division during human pluripotent stem cell differentiation. Nat. Commun. 14, 405 (2023).
Falcão, A. M. et al. Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat. Med. 24, 1837–1844 (2018).
Meijer, M. et al. Epigenomic priming of immune genes implicates oligodendroglia in multiple sclerosis susceptibility. Neuron 110, 1193–1210 (2022).
Zheng, C. et al. Distinct transcriptomic and epigenomic responses of mature oligodendrocytes during disease progression in a mouse model of multiple sclerosis. Nat. Neurosci. 28, 2612–2627 (2025).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Morgan, R. Hox genes: a continuation of embryonic patterning? Trends Genet. 22, 67–69 (2006).
Marques, S. et al. Transcriptional convergence of oligodendrocyte lineage progenitors during development. Dev. Cell 46, 504–517 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
Hassan, U. et al. Morphological spectrum and survival analysis of diffuse midline glioma with H3K27M mutation. Cureus 13, e17267 (2021).
Eytan, K. et al. Pediatric glioblastoma cells are sensitive to drugs that inhibit eIF2α dephosphorylation and its phosphomimetic S51D variant. Front. Oncol. 12, 959133 (2022).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, eaai8236 (2017).
Wang, K. C., Helms, J. A. & Chang, H. Y. Regeneration, repair and remembering identity: the three Rs of Hox gene expression. Trends Cell Biol. 19, 268–275 (2009).
Levine, M. Paused RNA polymerase II as a developmental checkpoint. Cell 145, 502–511 (2011).
Keirstead, H. S. & Blakemore, W. F. The role of oligodendrocytes and oligodendrocyte progenitors in CNS remyelination. Adv. Exp. Med. Biol. 468, 183–197 (1999).
Li, N. & Leung, G. K. K. Oligodendrocyte precursor cells in spinal cord injury: a review and update. Biomed. Res. Int. 2015, 235195 (2015).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Danese, A. et al. EpiScanpy: integrated single-cell epigenomic analysis. Nat. Commun. 12, 5228 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871 (2018).
Saint-André, V. et al. Models of human core transcriptional regulatory circuitries. Genome Res. 26, 385–396 (2016).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Li, Z. et al. Identification of transcription factor binding sites using ATAC–seq. Genome Biol. 20, 45 (2019).
Luebbert, L. & Pachter, L. Efficient querying of genomic reference databases with gget. Bioinformatics 39, btac836 (2023).
Abdennur, A. & Mirny, L. A. Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 36, 311–316 (2020).
Ramírez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Wolff, J. et al. Galaxy HiCExplorer 3: a web server for reproducible Hi-C, capture Hi-C and single-cell Hi-C data analysis, quality control and visualization. Nucleic Acids Res. 48, W177–W184 (2020).
Wolff, J. et al. Galaxy HiCExplorer: a web server for reproducible Hi-C data analysis, quality control and visualization. Nucleic Acids Res. 46, W11–W16 (2018).
Flyamer, I. M., Illingworth, R. S. & Bickmore, W. A. Coolpup.py: versatile pile-up analysis of Hi-C data. Bioinformatics 36, 2980–2985 (2020).
Open2C et al. Cooltools: enabling high-resolution Hi-C analysis in Python. PLoS Comput. Biol. 20, e1012067 (2024).
Roayaei Ardakany, A., Gezer, H. T., Lonardi, S. & Ay, F. Mustache: multi-scale detection of chromatin loops from Hi-C and Micro-C maps using scale-space representation. Genome Biol. 21, 256 (2020).
Acknowledgements
We thank T. Jimenez-Beristain for writing human ethical permits, A. Pombo and G. La Manno for discussions, and N. Jabado and C. Kleinman for critically evaluating the paper. We thank the donors, their families and the Edinburgh Brain Bank for the archival tissue used in this study. We acknowledge support from the National Genomics Infrastructure in Stockholm, supported by Science for Life Laboratory, the Knut and Alice Wallenberg Foundation and the Swedish Research Council. Part of the computation/data handling was enabled by resources provided by the National Academic Infrastructure for Supercomputing in Sweden and Swedish National Infrastructure for Computing at the Uppsala Multidisciplinary Center for Advanced Computational Science, partially supported by the Swedish Research Council (grant agreements 2022-06725 and 2018-05973). Part of the computing was also performed in the Linnarsson group Monod Linux cluster at MBB-KI, and we thank P. Lönnerberg for maintenance and support. This project has been made possible in part by grants 2019-002427 and 2021-239069 from the Chan Zuckerberg Initiative DAF, an advised fund (project 200412) as part of the Human Cell Atlas Project (to A.W.), Swedish Research Council (grant 2022-00650 to M.J.) and the Swedish Brain Foundation (FO2021-0150 to M.J.). Work in G.C.-B.’s research group was supported by the Swedish Research Council (grant 2019-01360 and Distinguished Professor grant 2023-00324), the European Union (Horizon 2020 Research and Innovation Programme/European Research Council Advanced Grant SingleMS, grant agreement 101096064), the Swedish Brain Foundation (FO2023-0032), the Swedish Cancer Society (Cancerfonden grant 23 2945 Pj 01 H), Knut and Alice Wallenberg Foundation (grant 2019-0089 and Wallenberg Scholar grant 2023-0280), the Göran Gustafsson Foundation for Research in Natural Sciences and Medicine, the Swedish Society for Medical Research (grant JUB2019 and postdoctoral fellowship to K.E.C.), Olav Thon Foundation (to G.C.-B. and S.A.G.), Ming Wai Lau Centre for Reparative Medicine, and Strategic Research Area Stem Cells and Regenerative Medicine (Karolinska Institutet). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
M.K. and G.C.-B. conceptualized the project and designed the experiments. L.A.S. performed macrodissection of archival tissue blocks. M.K. and M.M. optimized the snATAC–seq protocol for archival tissue and M.K. optimized the nanoCUT&Tag protocol. M.K. performed single-cell experiments and analyzed the data, with the help of F.B.P. (chromVAR), E.A. (dataset integrations, annotations, comparisons and scRNA-seq analysis), and J.Z. and V.A. (data QC). K.E.C., Ö.D. and Y.K.L. performed the CRISPRi/a experiments. L.A., E.E. and T.H.M. collected biopsy tissue. K.E.C., N.R., M.J. and M.B. collected cells for and coordinated the Micro-C experiments with the National Genomics Infrastructure. M.K. analyzed the Micro-C data. D.v.B., N.B.-C. and A.R.L. provided the input for the computational analysis. S.A.G. provided iPSC-derived hOPCs. S.A.G. and A.W. secured funding and coordinated work in their labs, contributing to the paper. M.K. and G.C.-B. wrote the paper. All co-authors read and approved the paper.
Corresponding author
Ethics declarations
Competing interests
G.C.-B. and M.B. filed a patent application on nanoCUT&Tag (European patent application EP22160860.7), which was not pursued. G.C.-B., M.B. and M.K. are shareholders of Nexus Epigenomics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Bing Ren and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 QC and preprocessing of the snATAC-seq dataset.
a, Schematic showing downstream data analysis workflow. b, Density scatter QC plots for all 48 samples in the snATAC-seq dataset. Number of unique fragments (log scale) on x axis and TSS enrichment score on y axis. c, Violin plots showing number of unique fragments (log) from each donor (top) and each sample (bottom). Data in each violin plot composed of all cells within that donor (top) or sample (bottom). Black box within each violin plot represents the interquartile range (IQR), white dot is the median, and whiskers extend to 1.5× IQR.
Extended Data Fig. 2 Validation of cell-type annotations through integration with external datasets.
a, Correlation plot showing quality of cell type annotations using the metagene scores (y axis) and annotations from integration with previously published datasets (ref. 37—right, ref. 5—left). b, Correlation plot showing quality of cell type annotations from the snATAC-seq dataset when integrated with a human cortical biopsy dataset (this study), and the cortical biopsy clusters in a 2D-UMAP embedding (bottom). c, Bar chart and violin plots showing cell numbers and feature counts (log scale) for each cell type (left) and contribution of each sex to the metrics (right). Data from each plot are composed of total cell counts for individual cell types (left) and the sex distribution for those cells (right). See Supplementary Table 1 for detailed information on donor metadata. Black box within each violin plot represents the interquartile range (IQR), white dot is the median, and whiskers extend to 1.5× IQR. d, snATAC-seq genome coverage tracks for different marker genes in each identified cell type. All columns are group normalized (range is shown in the first panel for each column). Gene location and TSS orientation are shown below. e, TF motif enrichment matrix showing the top four TF motifs identified as being differentially accessible in each cell type. f, 2D UMAP showing cell clustering based on the chromVAR calculated TF motif deviations. Marker TF enrichment for each cell type is shown on the right. g, 2D UMAP (left) and bar chart (right) showing tissue composition of cell types in the snATAC-seq dataset.
Extended Data Fig. 3 Differential accessibility across age and sex in major CNS cell types.
a, Volcano plot showing differentially accessible genes in snATAC-seq based on age (less than or greater than 50 years old) in the different cell types. Two-sided t-test with Benjamini–Hochberg correction. Thresholds: adjusted P-value: 0.001, logFC = 1.5. b, Volcano plots showing differentially accessible genes in snATAC-seq based on sex in different cell types in the cortex (left) and spinal cord (right). Two-sided t-test with Benjamini–Hochberg correction. Thresholds: adjusted P-value: 0.001, logFC = 1.5.
Extended Data Fig. 4 Quality assessment of H3K27ac/H3K27me3 nanoCUT&Tag.
a, Workflow for demultiplexing and custom cell-calling for each nanoCUT&Tag antibody (H3K27ac, H3K27me3). b, Scatterplots used for custom cell calling in the H3K27ac (top) and H3K27me3 (bottom) modality. Number of unique fragments (log scale) shown on x axis and the fraction of reads in peaks (FRiP) shown on y axis. Selected cells shown in blue. c, Quality metric violin plots showing the number of unique features (left) and fraction of reads in peaks (FRiP) for H3K27ac (green) and H3K27me3 (red) in this dataset compared to those in a previously published dataset in mouse29. Black box within each violin plot represents the interquartile range (IQR), white dot is the median, and whiskers extend to 1.5× IQR. d, Violin plot showing the number of unique features (log scale) of the H3K27ac, H3K27me3 (this study) compared to the H3K27ac of a published dataset33. Black box within each violin plot represents the interquartile range (IQR), white dot is the median, and whiskers extend to 1.5× IQR. e, Venn diagrams showing the overlap between peaks called in our snATAC-seq dataset (blue), H3K27ac dataset (green) and a set of brain cCREs from the ENCODE database (pink). f, 2D UMAP co-embedding of the snATAC and H3K27ac nanoCUT&Tag datasets. g, 2D UMAP of the nanoCUT&Tag dataset from both modifications. Colored lines connect the same cell in both modalities. h, Bar chart and violin plot showing the cell number in each population (left) and distribution of feature counts (log scale) for each modality (middle, right). Data from each plot are composed of total cell counts for individual cell types. Black box within each violin plot represents the interquartile range (IQR), white dot is the median, and whiskers extend to 1.5× IQR.
Extended Data Fig. 5 Cell-type-specific enrichment of H3K27ac/H3K27me3 across gene TSS.
The k-means clustering of H3K27me3 and H3K27ac signal distribution at the TSS of all genes for each cell type. Genes in the clusters showing strong H3K27ac and weak H3K27me3 were used as input for gget enrichr analysis to identify enriched cell types. Top identified cell type is the cell type itself highlighting the strong cell-type-specific signal captured in our nanoCUT&Tag dataset.
Extended Data Fig. 6 Co-accessibility analysis identifies a distal SOX10 enhancer in OLGs.
a, Cumulative distribution of the co-accessibility score for all loops identified by Cicero. Red line shows the score cutoff (0.5) used for assessing high-quality interactions and captures the top 5% of all loops. b, snATAC-seq genome browser snapshot showing chromatin accessibility signal in all cell types at CDC42EP1 locus (left), identified SOX10-distal enhancer (middle) and SOX10 locus (right) and the corresponding loops identified using Cicero. Red columns highlight the CDC42EP1 and SOX10 genes, and the gray column highlights the new enhancer. c, UCSC Genome Browser snapshot showing the identified enhancer locus (light blue column) as well as overlap with previously identified ENCODE cCREs, PhyloP base conservation score and evolutionary conservation with different species. d, snATAC-seq genome browser track showing chromatin accessibility in OPCs and MOLs at the newly identified enhancer, SOX10 locus, and previously characterized U1-U5, D6 and D7 enhancers (purple) and overlap with thyroid hormone receptor motifs (T3R, green) and TFAP2A motifs (orange). e, snATAC-seq and nanoCUT&Tag genome browser tracks the CDC42EP1-enhancer-SOX10 locus showing ATAC, H3K27ac and H3K27me3 pseudobulk signal in MOLs and OPCs and Cicero links. f, FACS plots showing transduction efficiency (left) and gene expression changes (qPCR) of SOX10 when directing dCas9–p300 to the U2 enhancer, U3 enhancer, and a nontargeting control (NTC) (right). N = 4 biological replicates; data shown as mean ± s.e.m. Statistics: one-way ANOVA with Tukey’s post hoc test. Two-sided t-test performed. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, NS = nonsignificant. g, Scatterplot showing TF strength of shared core TFs in MOLs (x axis) and OPCs (y axis) and highlighting the identified HOX genes. h, List of identified HOX genes being differentially accessible in spinal cord OPCs and MOLs. i, Scatterplot showing correlation between the multiome-RNA and multiome-ATAC. j, Stacked violin plot showing gene expression levels of all HOX genes in OPCs in all regions from an adult human brain transcriptomic atlas4.
Extended Data Fig. 7 Chromatin accessibility and histone PTM patterns at HOX loci in OLGs.
a, snATAC-seq and nanoCUT&Tag genome browser tracks showing H3K27ac, H3K27me3 and ATAC pseudobulk signal in OLGs at the HOX-B and HOX-C clusters in cervical spinal cord (upright track, darker shade) and motor cortex (inverted track, lighter shade). Directionality of the clusters is shown by the arrow. b, Gaussian smoothed normalized signal from ATAC (blue), H3K27ac (green) and H3K27me3 (red) in spinal cord OLGs (solid line) and cortical OLGs (dotted line) across each HOX cluster with a 50 kb flanking region upstream and downstream. c, Gaussian smoothed normalized signal from ATAC (blue), H3K27ac (green) and H3K27me3 (red) across the HOXB, HOXC and HOXD clusters with a 50 kb flanking region upstream and downstream, separated by each modality. Gray bars show the location of ‘signal boundaries’ identified in each modality. Dotted lines mark the boundaries of each HOX cluster. Arrowheads underneath the plots mark intermediate and strong signal boundaries. HOX cluster directionality shown by arrow on top. d, snATAC-seq and nanoCUT&Tag genome browser tracks showing the ATAC (blue), H3K27ac (green), and H3K27me3 (red) pseudobulk signal in the microglial and astrocyte populations at HOXA in both spinal cord (CSC) and cortex (BA4).
Extended Data Fig. 8 High-resolution Micro-C in human iPS-derived OPCs and primary B-cells shows cell-type-specific architecture.
a, Experimental schematic showing collection of iPS-derived hOPCs and patient-derived B cells for Micro-C. Schematic of chromatin looping to bring enhancer and promoter in contact for transcription is shown below. Schematic in a created in BioRender. Castelo-branco, G. https://BioRender.com/orfbpyi (2025). b, Micro-C contact matrix in hOPCs at 5 kb resolution showing the TAD formed at the SOX9–KCNJ2 locus. c, Bar chart showing the sequencing reads (in millions) obtained for each replicate in the B cells and hOPCs. Libraries were first shallow sequenced to assess library quality (light gray bars) followed by deep sequencing (dark gray). B cell replicates correspond to 3 separate patients. hOPC replicates correspond to separate biological replicates. d, Cumulative distribution of compartment size identified in the hOPC and B-cell Micro-C data. Asterisk marks 1.5 Mb size as the upper size limit for 95% of all compartments. e, Contact matrix showing the normalized observed counts (lower triangle) and normalized obs/exp counts (upper triangle) at chromosome 7 in hOPCs. A and B compartments are shown along the sides of the matrix and exhibit strong correlation with ‘pockets’ of increased contact frequency. f, A/B compartments in hOPCs on chromosome 15 overlaid on chromatin accessibility data from hOPCs (unpublished), showing correlation between active A compartments and increased accessibility. g, Aggregate pileup analysis of differentially accessible loops in B cells and hOPCs showing cell type specificity of identified loops. h, Schematic of the 3’ TAD (c-Dom) and 5’ TAD (t-Dom) that flank the HOXD cluster (yellow) and the locations of genes tested in qPCR (HOXD1, HOXD4, HOXD8) and the miR10b and LINC01116 genes.
Extended Data Fig. 9 HOX-associated chromatin states in adult OLGs mirror regulatory programs in HGG.
a, Schematic of the adult human brain showing the location of pontine and thalamic gliomas along the A-P axis. Schematic in a created in BioRender. Castelo-branco, G. https://BioRender.com/orfbpyi (2025). b, snATAC-seq and nanoCUT&Tag normalized promoter accessibility (ATAC), H3K27ac and H3K27me3 signal in spinal cord OLGs (top) and cortical OLGs (bottom) at different developmental genes associated with brain and spinal cord patterning. c, Genome browser tracks showing nanoCUT&Tag H3K27ac and H3K27me3 pseudobulk signal coverage at the HOXA cluster in spinal cord (SC) derived adult human OLG (hOLG) and PFA-EP tumors, H3.3K27M pontine tumors, and H3.3K27M thalamic tumors11. d, Same as c, but at the HOXD locus.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods.
Supplementary Tables (download XLSX )
Supplementary Table 1: Human tissue metadata. Supplementary Table 2: Co-accessible loci (Cicero) for all cell types. Supplementary Table 3: TF networks for all cell types. Supplementary Table 4: Differentially accessible regions in glia across sex, age and tissue. Supplementary Table 5: gRNA sequences and qPCR primers. Supplementary Table 6: Plasmid sequences for lentiviral constructs.
Source data
Source Data Figs. 2 and 6 and Extended Data Fig. 6 (download XLSX )
Statistical source data for Figs. 2 and 6 and Extended Data Fig. 6.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kabbe, M., Agirre, E., Carlström, K.E. et al. Single-nucleus epigenomic profiling of the adult human central nervous system unveils epigenetic memory of developmental programs. Nat Neurosci (2026). https://doi.org/10.1038/s41593-026-02208-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41593-026-02208-0