Abstract
Epithelial-mesenchymal transition (EMT) is a critical process in cancer cell motility and metastasis. Monoterpene indole alkaloids (MIAs) have been widely investigated for biological activities, but rarely been explored for cell motility inhibition. This study aimed to discover natural products, especially focusing on MIAs, that inhibit EMT in cancer cells, based on our screening using multiple plant extracts. We found that an extract of the aerial parts of Uncaria scandens (Sm.) Wall. (Rubiaceae) decreased cancer cell motility. Targeted isolation exhibited eight MIAs. Among them, strictosamide and mitraphylline suppressed the invasion and migration of the cells. The alteration of the mRNA expression of the EMT effectors and transcription factors suggested that the EMT signaling pathway is related to the suppression of cancer cell motility by both compounds. RT-PCR gene array suggested inhibition of integrin α4 signaling as a potential mechanism of the EMT inhibition, which was supported by the quantitative analysis of the mRNA expressions of the related genes. Together, present study is the first report highlighting the cell motility-suppressive effects of strictosamide and mitraphylline. Our results suggested the potential applicability of strictosamide and mitraphylline for the prevention of metastasis.
Similar content being viewed by others
Introduction
Cancer is more difficult to treat when metastasis occurs. Despite advances in treatment, metastasis is the most common cause of cancer-related deaths1. Epithelial-mesenchymal transition (EMT), a developmental process in which epithelial cells are transformed into spindle-shaped mesenchymal cells, enhances mobility, invasion, and resistance to apoptotic stimuli in cancer cells, which advances their metastatic properties2. A fundamental event in EMT includes downregulation of the epithelial marker E-cadherin, whereas the expressions of mesenchymal markers, such as N-cadherin, vimentin, and fibronectin, are upregulated3,4. Several experimental studies have suggested that EMT inhibition can help prevent metastasis5,6,7. Hence, small molecules that target EMT may be valuable candidates as chemotherapeutic agents that reduce malignancy.
Monoterpene indole alkaloids (MIAs) are phytochemicals derived from strictosidine which is biosynthesized by condensation of tryptamine and secologanin8. MIAs exhibit structural complexity and diversity during subsequent biosynthetic pathway, and this diversity makes them a promising source of drug candidates9. The most well-known clinical applications of MIAs are vinblastine and vincristine [chemotherapeutic agents from Catharanthus roseus (L.) G. Don (Apocynaceae)], reserpine [an antihypertensive drug from Rauvolfia serpentina (L.) Benth. ex Kurz (Apocynaceae)], and quinine [an antimalarial drug from Cinchona spp. (Rubiaceae)]. Despite extensive pharmacological investigations on MIAs, their inhibitory activities on cell motility remain unexplored.
As a part of our research program aiming to discover natural products that suppress cancer cell motility10,11,12, we investigated the phytochemistry and pharmacology of aerial parts of Uncaria scandens (Sm.) Wall. (Rubiaceae). The genus Uncaria is a rich source of alkaloids, especially MIAs13. The species that have been well-characterized for their bioactive chemical constituents include Uncaria rhynchophylla (Miq.) Miq. and Uncaria tomentosa (Willd. ex Schult.) DC14,15,16,17. However, MIA constituents of U. scandens have rarely been investigated. Yang et al.18 performed a chemical investigation of U. scandens; however, only triterpenes and flavonoids were reported. A study using liquid chromatography-tandem mass spectrometry (LC-MS/MS) profiling suggested the possible occurrence of MIAs in U. scandens; however, these were not fully confirmed by further experiments19.
Here, we describe the bioactivity- and chemical profile-guided discovery of strictosamide (1) and mitraphylline (2), two MIAs isolated from U. scandens, as bioactive compounds against the motility of H1975, AGS, and CaCo2 cells, which are human lung, gastric, and colorectal cancer cells, respectively, via EMT suppression. The three cancer cells used in this study harbored different states of mutations in cancer-critical genes: AGS has mutant KRAS, PIK3CA, TP53, and CTNNB1; CaCo2 has wild-type KRAS, BRAF, and PIK3CA; and H1975 has mutant EGFR, PIK3CA, and TP5320.
Materials and methods
Plant materials
The aerial parts of Uncaria scandens (Sm.) Wall. were collected on March 9, 2011, from the Dong community in K’Bang district, Gia Lai province, Vietnam. The plant was authenticated by T. T. Bach (Vietnam Academy of Science and Technology, Hanoi, Vietnam). A voucher specimen (VK 4455) was deposited in the herbarium of the Korea Research Institute of Bioscience and Biotechnology (Daejeon, Republic of Korea).
Isolation and structural Elucidation of compounds
The dried aerial parts of U. scandens (550 g) were extracted with MeOH (3 × 5.5 L, for 2 h each) by ultrasonication at room temperature. The crude extract (47.6 g) was obtained by removing the extraction solvent in vacuo. The extract was suspended in H2O and washed with n-hexane to remove fat, wax, and chlorophyll. The aqueous residue was acidified using HCl to attain a pH of 2–3, and the nonpolar compounds were extracted from the acidic suspension using EtOAc. Thereafter, NH4OH was added to aqueous residue to obtain a pH of 11–12 and alkaloid-rich fraction (0.8 g) was obtained by extracting it with CH2Cl2. Extra-pure grade solvents used for extraction and fractionation were purchased from Daejung Chemicals & Metals Co. Ltd. (Si-heung, Korea). The total extract and fractions were desiccated with a nitrogen blower and a centrifugal vacuum evaporator before further chemical and biological analysis.
Compound 2 (165.0 mg) was obtained as a MeOH-insoluble pellet of the alkaloid-rich fraction during sample preparation for the preparative HPLC. The remaining alkaloid-rich fraction was separated using preparative HPLC (Waters 600 system) equipped with a Hector C18 column (250 × 21.2 mm, 5 μm, RS Tech, Daejeon, Korea) and eluted with a linear gradient of H2O-MeCN (75:25 → 10:90 for 25 min, 12 mL/min) to obtain purified compound 1 (7.1 mg, tR 11 min) and four fractions D1–D4. D2 (77.3 mg) was further separated into seven subfractions (D2a–D2g) using silica gel flash chromatography and eluted with a gradient of CH2Cl2-MeOH. Compounds 3–6 (3, 1.7 mg, tR 6.5 min; 4, 1.9 mg, tR 7.0 min; 5, 0.8 mg, tR 14.0 min; 6, 1.8 mg, tR 8.0 min) were purified from D2b using preparative HPLC equipped with a Spursil C18–EP column (250 × 10 mm, 5 μm, Dikma Technologies, Foothill Ranch, CA, USA) and eluted using a linear gradient of H2O-MeCN (85:15 → 60:40 for 20 min, 4 mL/min). Compounds 7 and 8 (7, 1.5 mg, tR 9.5 min; 8, 2.1 mg, tR 11.0 min) were isolated from D2d by preparative HPLC equipped with a Hector C18 column (250 × 10 mm, 5 μm) and eluted with a linear gradient of H2O-MeCN (90:10 → 40:60 for 15 min, 4 mL/min). The solvents used for preparative HPLC were purchased from J. T. Baker (Avantor, Phillipsburg, NJ, USA; #4218-03, water; 9017-03, MeCN).
A Bruker Avance III HD 500 spectrometer (Bruker BioSpin, Billerica, MA, USA) at the Chronic and Metabolic Diseases Research Center of Sookmyung Women’s University was used for the structural identification of isolates. Pure forms of compounds 3–6 were unstable because isomerization occurs at room temperature14. Raw NMR data is provided as supplementary material (Figs. S1–S16).
LC–MS/MS analysis
The alkaloid-rich fraction of U. scandens was analyzed using a Waters Acquity UPLC system coupled with a Waters Vion IMS QTOF mass spectrometer (Waters Co., Milford, MA, USA). Chromatographic separation was performed using a mobile phase comprising 0.1% formic acid in water (A) and MeCN (B). Samples were injected into a Waters Acquity BEH C18 (100 × 2.1 mm, 1.7 μm; #186002353) column with a gradient method at a flow rate of 0.3 mL/min:10–60% of B (0–20 min), followed by 3 min washing and 3 min reconditioning. MS/MS analysis was performed in positive ion mode, and leucine enkephalin at m/z 556.2771 was selected as the lock mass. LC-MS grade solvents were acquired from Fisher Chemical (Thermo Fisher Scientific, Waltham, MA, USA; #W6, water; A995, MeCN).
Cell culture
Roswell Park Memorial Institute (RPMI; #CM058-050) or Dulbecco’s modified Eagle’s medium (DMEM; #CM001-050) (GenDepot, Katy, TX, USA) supplemented with 10% fetal bovine serum (FBS, #F0600-050, GenDepot) and 1% penicillin/streptomycin solution (#CA005-010, GenDepot) was used to maintain the human cancer cell lines H1975 (H. sapiens lung cancer; CVCL_1511; #CRL-5908), AGS (H. sapiens gastric cancer; CVCL_0139; #CRL-1739), and CaCo2 (H. sapiens colorectal cancer; CVCL_0025; #HTB-37). Cell lines were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were maintained at 37 °C in a humidified atmosphere containing 5% CO2. The α4β1 agonist (Integrin modulator 1, #HY-134130) was purchased from MedChemExpress (Monmouth Juntion, NJ, USA).
Cell viability
Cells (3 × 103/well) were seeded in 96-well plates, incubated overnight, and exposed to compounds at concentrations of 25, 12.5, 6.25, 3.13, and 1.56 µM for 48 h. MTT (#M2128, Sigma Aldrich, St. Louis, MO, USA) was added to the cultures, and cells were lysed after 4 h using 150 µL of DMSO (#D8418, Sigma Aldrich) at 37 °C. Absorbance was measured at 570 nm using a microplate reader (BioTek, Winooski, VT, USA).
Invasion assay
Invasion assay was conducted in Transwell chambers (Corning, NY, USA) containing polycarbonate membranes with 8 μm pores coated with 1% gelatin. Cells were plated at 5–7 × 104/well in DMEM or RPMI containing 0.2% bovine serum albumin (BSA) in the upper compartment of the chamber with or without the tested compounds. Subsequently, the chemoattractant fibronectin was added to the lower chamber with RPMI or DMEM (600 µM) containing 0.2% BSA. Following 24 h of incubation, the infiltrating cells were fixed using a Diff-Quik kit (Polysciences, Inc., Warrington, PA, USA). Cells were mechanically removed from the membrane in the chamber using a cotton swab, and cells clinging to the underside of the filter were stained and counted under a light microscope11.
Scratch wound healing assay
The scratch wound healing assay was performed using the IncuCyte system (Sartorius, Göttingen, Germany). H1975, AGS, and CaCo2 cells were seeded in Essen ImageLock 96-well plates (4 × 104/well). After an overnight incubation, a scratch wound was made on a monolayer of confluent cells using a wound marker (Sartorius). After wounding, cell debris was cleared with PBS, and the cells were incubated in DMEM or RPMI culture medium supplemented with 2% FBS, with or without the tested compounds.
Quantitative reverse-transcription PCR
The cells were plated at a density of 2 × 105/well on a 6-well plate and grown overnight. The cells were then treated with the compounds or DMSO. Total RNA was isolated from the cells using RNAiso Plus (#9109, TaKaRa, Otsu, Japan). The RNA (1 µg) was converted to cDNA using M-MLV reverse transcriptase (#28025021, Invitrogen, Carlsbad, CA, USA). qPCR was performed using SYBR Green qPCR PreMIX (#RT500, Enzynomics, Seoul, Korea). qRT-PCR was performed using a CFX instrument (Bio-Rad, Hercules, CA, USA). The list of the sequences of primers used in this study is shown in Table 1.
Western blot
Whole cell lysates were prepared as follows: cells were plated in 6-well plates, incubated overnight, treated with DMSO or strictosamide and mitraphylline, washed twice with ice-cold PBS, and lysed with a lysis buffer. Protein concentrations in the lysates were determined using a bicinchoninic acid (BCA) assay (#23225, Thermo Fisher Scientific, Waltham, MA, USA). Antibodies against N-cadherin, FAK, and p-FAK (Tyr397) were detected with a horseradish peroxidase-conjugated secondary antibody using an Immobilon Western Chemiluminescent HRP Substrate Kit (#WBKLS, Millipore, Billerica, MA, USA) and luminescence imaging. The relative densities of the bands were obtained by normalizing the density of β-actin (#4967, Cell Signaling Technology, Danvers, MA, USA) bands in each sample using Multi-Gauge 3.0 software (Fuji Photo Film Co., Ltd., Tokyo, Japan). Values are represented as arbitrary units of densitometry, corresponding to the signal intensity. The PageRuler Plus Prestained Proten Ladder (#26619, Thermo Fisher Scientific) served as a ladder. The primary and secondary antibodies used for Western blots are presented in Table 2. Full-length uncropped blots are provided as supplementary material (Figs. S18–S25).
Real-time PCR gene array
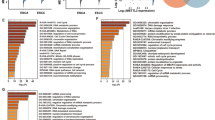
The Human Cell Motility RT2 Profiler™ PCR Array (Qiagen, SA Biosciences, Valencia, CA, USA) was used to assess the expression of motility-related genes in AGS cells that were treated with strictosamide and mitraphylline. Expression levels of 84 key genes associated with cell motility were quantified according to the manufacturer’s instructions. Fold-changes in mRNA expression were calculated based on the cycle threshold values obtained by RT-PCR. A scatter plot of the test versus control samples demonstrated the validity of the experiment. Downregulated markers were evaluated for their contribution to biological processes using Metascape (https://metascape.org)21.
Immunofluorescence imaging
Cancer cells were cultured on cover slips at the density of 1 × 105 in a 12-well plate. After the overnight, cells were treated with DMSO or compounds for 24 h. Immunofluorescence (IF) staining was performed as previously described22, using α-tubulin monoclonal antibody (B-5-1-2), Alexa Fluor™ 488 (#78833, Thermo Fisher Scientific) and Alexa Fluor™ 568 Phalloidin (#A12380, Thermo Fisher Scientific). K1-Fluo Confocal Laser Scanning Microscope (Nanoscope Systems, Daejeon, Republic of Korea) was used to capture the images.
Statistical analysis
Data are expressed as means ± standard deviation. All statistical analyses were performed using Sigma Plot 12.5 software (Systat Software Inc., Chicago, IL, USA). The statistical significance between two groups was compared using the Student’s t-test and p values < 0.05 were considered statistically significant.
Results
Bioactivity- and chemical profile-based isolation of monoterpene indole alkaloids from Uncaria scandens
U. scandens was selected in this study because its crude extract showed inhibitory activity in primary screening of AGS cells using an invasion assay. AGS cell line is commonly used in drug screening studies due to its well-established characteristics and reproducible responses to various treatments23,24. Given the known production of alkaloids in this species, an acid–base partitioning method was employed for fractionation. Among the resulting fractions, both the EtOAc and alkaloid-rich fractions exhibited the most potent activity (Fig. 1A). The alkaloid-rich fraction was prioritized for further chemical investigation due to the reported abundance of alkaloids in the Uncaria genus. Furthermore, since liquid–liquid partitioning often distributes individual compounds across multiple fractions, it was hypothesized that alkaloids might also be present in the EtOAc fraction, potentially contributing to its observed bioactivity.
Bioactivity screening and chemical investigation of U. scandens. (A) Representative images showing each insert in the invasion assay and the relative number of invaded cells. AGS cells were treated with fractions (5 µg/mL; with 0.1% DMSO) for 24 h. Data represent mean ± S.D. * p < 0.05; ** p < 0.01. (B) Chemical structures of the eight isolated compounds 1–8. (C) The LC-MS base peak ion (BPI) chromatogram of the alkaloid-rich fraction of U. scandens. The isolated compounds are denoted on the corresponding chromatographic peaks. Notably, the intensities of chromatographic peaks do not represent absolute quantity of each compound because the MS signal intensity is heavily affected by ionization efficiency of each molecule.
LC-MS/MS profiling of the alkaloid-rich fraction suggested the abundance of MIAs, characterized by multiple MS ion features with predicted molecular formulae containing two nitrogen atoms and falling within the m/z 350–400 range. These results also supported the presence of shared alkaloids between the alkaloid-rich and EtOAc fractions (Fig. S17). Strictosamide and mitraphylline were annotated from the chemical profile by MS/MS spectral matching against the GNPS library25; however, many MS features remained unidentified. The alkaloid-rich fraction was subjected to chromatographic isolation and eight compounds were purified. We compared the NMR spectral data of the obtained compounds with that data available in literature and identified the isolates as strictosamide (1), mitraphylline (2), uncarine A (3), uncarine C (4), uncarine E (5), uncarine F (6), lyaloside (7), and 5(S)-5-carbomethoxystrictosidine (8) (Fig. 1B,C)26,27,28,29,30. Of the eight compounds, strictosamide (1) and mitraphylline (2) were further evaluated for their inhibitory activity against cancer cell motility as their isolated amounts were abundant and gap in literature for the investigation.
Strictosamide and mitraphylline suppress the cell motility in human cancer cells
We used three different cell lines in the following experimental designs, mainly because these cell lines were isolated from different types of cancers, but there are some background differences. The H1975 cell line, derived from human non-small cell lung cancer (NSCLC), carries EGFR mutations (L858R and T790M) and exhibits epithelial characteristics under basal conditions. However, H1975 cells are capable of undergoing epithelial-mesenchymal transition (EMT) upon stimulation with factors such as TGF-β, contributing to enhanced migratory ability and potential resistance to EGFR-targeted therapies31,32,33. The AGS cell line, derived from human gastric adenocarcinoma, harbors mutations in key oncogenes and tumor suppressors, including KRAS, PIK3CA, TP53, and CTNNB1, which are frequently altered in gastric cancer and implicated in EMT regulation. While AGS cells typically exhibit an epithelial phenotype and are not highly metastatic under basal conditions, they can undergo epithelial-mesenchymal transition (EMT) upon stimulation with cytokines such as TGF-β. These features make AGS a relevant and responsive model for investigating EMT-associated signaling pathways, stemness traits, and drug responses in the context of gastric cancer progression34,35,36,37,38,39,40. On the other hand, CaCo2 cell line, derived from human colorectal adenocarcinoma, exhibits a well-differentiated epithelial phenotype and forms tight junctions and brush-border-like structures, making it a classical model for intestinal barrier and absorption studies41,42. Mutations, including TP53 (a well-known tumor suppressor gene), KRAS-WT, and PIK3CA-WT, contribute to the cell line’s transformation and resistance to certain chemotherapeutic agents in CaCo2 cells43,44,45. Strictrosamide (1) and mitraphylline (2) were evaluated for their effects on the viability of H1975, AGS, and CaCo2 cells at concentrations from 1.56 to 25 µM. These compounds were cytotoxic at high concentrations, but their effects were not potent (Fig. 2A). Based on these results, the concentrations for the Transwell invasion assay were determined to be 1 and 2.5 µM. Strictosamide (1) inhibited the invasion of H1975, AGS, and CaCo2 cells by approximately 21, 27, and 11%, respectively, at 2.5 µM after 24 h, whereas mitraphylline (2) inhibited the invasion by approximately 24, 23, and 17%, respectively, under the same conditions (Fig. 2B). In the wound-healing assay using the IncuCyte system, strictosamide (1) and mitraphylline (2) decreased the number of migrating cancer cells in a dose-dependent manner (Fig. 2C). In particular, mitraphylline (2) exerted a strong inhibitory effect on AGS cell migration. Our results suggested that these two compounds suppressed the motility of H1975, AGS, and CaCo2 cells.
The effects of strictosamide (1) and mitraphylline (2) on cell viability and motility of H1975, AGS, and CaCo2 cells. (A) The cell viability of H1975, AGS, and CaCo2 cells treated with strictosamide (1) and mitraphylline (2) for 48 h. Data are represented as the mean ± S.D. of the differences between the cell viability of cells affected with strictosamide (1) and mitraphylline (2) compared with that of DMSO-treated controls. (B) Representative images of each insert and relative number of invaded cells in the Transwell invasion assay. Cells were treated with strictosamide (1) and mitraphylline (2) at concentrations of 1 and 2.5 µM for 24 h. (C) Representative images of each wound area and quantitative data of relative wound density in the scratch wounding migration assay using the IncuCyte system. Overlaid on phase contrast images, the green line indicates the initial wound line, while the red line indicates the migrated cell area at the specified time period. Data are represented as the mean ± S.D. *p < 0.05; ** p < 0.01; *** p < 0.001.
Strictosamide and mitraphylline regulates the markers of epithelial-mesenchymal transition
To determine whether the motility suppression by strictosamide (1) and mitraphylline (2) involved any regulation of EMT, the mRNA expression levels of three EMT effectors (E-cadherin, N-cadherin, and vimentin) were measured. The results showed that E-cadherin tended to be slightly upregulated, while the expression of N-cadherin mRNA was significantly downregulated by the two compounds in the three cell lines, whereas vimentin mRNA expression was not significantly downregulated in H1975 cells treated with strictosamide (1) or in CaCo2 cells treated with mitraphylline (2) (Fig. 3A). Examination of N-cadherin protein levels supported the downregulation of N-cadherin by strictosamide and mitraphylline (Fig. 3B). The mRNA expression of the EMT transcription factors Twist, Snail, Slug, Zeb1, and Zeb27 were examined to determine if they were affected by strictosamide and mitraphylline. Strictosamide (1) significantly suppressed all EMT transcriptional regulators, including Twist, in H1975 cells, whereas it was more weakly modulated in CaCo2 cells than in the H1975 and AGS cell lines. In contrast, mitraphylline (2) did not significantly affect Slug and Zeb1 mRNA expression in the H1975 cell line, whereas it significantly downregulated other markers. Although mitraphylline (2) significantly downregulated all markers in the AGS cell line, the markers Slug, Zeb1, and Zeb2 exhibited a more dramatic inhibitory effect on mitraphylline (2) than on the H1975 and CaCo2 cell lines. Our results showed that both compounds decreased the expression of EMT transcription factors, suggesting that the EMT signaling pathway is related to the suppression of cancer cell motility by strictosamide (1) and mitraphylline (2) (Fig. 3C).
Strictosamide (1) and mitraphylline (2) downregulated the expression of EMT effectors and transcription factors. (A) The mRNA expression of EMT effectors (N-cadherin, vimentin and E-cadherin) were analyzed by qRT-PCR. The mRNA expressions were normalized against β-actin. (B) The protein levels of N-cadherin were measured. β-actin served as a loading control. (C) The mRNA expressions of EMT transcription factors (Twist, Snail, Slug, Zeb1, and Zeb2) were analyzed by qRT-PCR. The mRNA expressions were normalized against β-actin. Data are represented as the mean ± S.D. * p < 0.05; ** p < 0.01; *** p < 0.001.
Strictosamide and mitraphylline downregulated several cell motility-related factors and suppressing ITGA4 signaling
To identify the mechanisms of action of strictosamide (1) and mitraphylline (2) against EMT, the mRNA expression profiles of strictosamide- and mitraphylline-treated cells were compared with those of controls using a Human Cell Motility RT2 Profiler™ PCR Array containing 84 cell motility-related genes. The array identified cell motility target genes, such as CAPN1, RHO, RHOA, RHOB, WASF1, WASL, CAV1, EGF, SRC, and ITGA4. Among them, ITGA4 was the most downregulated by strictosamide (1) (− 17.42, fold change) and mitraphylline (2) (− 11.42, fold change) (Fig. 4A). Based on the PCR array results, downregulated genes were screened for potential associations using the Search Tool for the Retrieval of Interacting Genes (STRING) database46. A protein-protein association network comprising CAV1, CAPN1, ITGA4, PTK2, RHO, RHOA, RHOB, SRC, WASF1, and WASL was acquired (Fig. 4B). The RT-PCR gene array results were further validated by quantifying the relative mRNA expression of genes listed in the protein network. Focal adhesion kinase (FAK) was also examined because it is associated with integrin signaling47,48. Strictosamide (1) downregulated all genes (Fig. 4C), confirming the PCR array profiling, and significantly downregulated ITGA4, FAK, and SRC mRNA expression in all the cell lines. However, mitraphylline (2) had different effects on these cell lines. RT2 PCR array analysis confirmed the downregulation of ITGA4, SRC, RHOB, WASF1, and CAV1 in AGS cells. Mitraphylline (2) significantly downregulated RHO, RHOA, and ITGA4 in H1975 cells but did not significantly alter the expression of other regulators. The expression of ITGA4, FAK, and SRC was not downregulated in CaCo2 cells after mitraphylline (2) treatment, whereas the expression of CAPN1, WASL1, and RHOB was significantly downregulated. This result suggests that mitraphylline (2) has different mechanical effects on CaCo2 cell line. These results suggested that strictosamide (1) and mitraphylline (2) inhibited cancer cell motility via ITGA4 signaling (Fig. 4C).
Strictosamide (1) and mitraphylline (2) downregulated several cell motility-related factors. (A) Pathway-focused gene expression analysis using the Human Cell Motility RT2 Profiler™ PCR Array was performed to examine the mechanisms by which strictosamide and mitraphylline modulate cell motility. The scatter plot compares the gene on the PCR Array between the DMSO-strictosamide and DMSO-mitraphylline. The number of fold changes for ITGA4 arranged downwards is shown in the figure. (B) A network of interrelated genes affected by strictosamide (1) and mitraphylline (2). (C) Quantitative analysis of the mRNA expressions of CAV1, CAPN1, WASF1, WASL, RHO, RHOA, RHOB, ITGA4, FAK, SRC in H1975, AGS, and CaCo2 cells treated with 2.5 µM of strictosamide (1) and mitraphylline (2). Data are represented as the mean ± S.D. * p < 0.05; ** p < 0.01; *** p < 0.001.
Effect of strictosamide and mitraphylline on cell morphology and reorganization of the actin cytoskeleton
We further analyzed the effect of downregulated markers on biological processes in Metascape. The results showed that the effect of the alkaloids is associated with regulating actin cytoskeleton regulation and regulate cell adhesion (Fig. 5A). Cell motility is linked to EMT, loss of the cell-cell adhesion protein E-cadherin, rupture of cell-cell junctions, and actin cytoskeleton reorganization49. Therefore, tubulin and F-actin staining were performed to examine the distribution of cell morphology and cytoskeletal components in strictosamide (1) and mitraphylline (2) treatment compared to control. The results reveal that the cell-cell adhesion of the treated cells was tighter and mesenchymal cell morphology was less observed compared to the control group (Fig. 5B). Interactions between the extracellular matrix, regulation of the actin cytoskeleton, mesenchymal cell adhesion, and cell motility are regulated through focal adhesions. High expression of FAK is related to the EMT process and increased cell migration50,51,52. For a further validation, we checked total-FAK and phosphorylated-FAK (Tyr397) protein levels to investigate the effect of the alkaloids against EMT and ITGA4-associated FAK activation. The result revealed that the level of phosphor-FAK (Tyr 397) and/or the level of total FAK were decreased compared to the control group (Fig. 5C).
Effect of strictosamide (1) and mitraphylline (2) on cell morphology and FAK protein level of cancer cells. (A) Biological processes linked to the pathways by strictosamide (1) and mitraphylline (2) found in Metascape. (B) The microtubule organization in the H1975, CaCo2, and AGS cells was examined using immunofluorescence microscopy following a 24 h treatment with strictosamide (1) and mitraphylline (2). Microtubules were stained with α-tubulin antibodies (green), while actin was stained with Alexa Fluor 568 phalloidin (red). (C) The protein levels of p-FAK and total FAK were measured. Cells were treated with 1 and 2.5 µM of strictosamide (1) and mitraphylline (2) for 48 h. β-actin served as a loading control. Data are represented as the mean ± S.D. * p < 0.05; ** p < 0.01; *** p < 0.001.
The responses against strictosamide and mitraphylline in cells exposed to Α4β1 agonist
We checked the responses against strictosamide (1) and mitraphylline (2) in cells treated with an α4β1 agonist molecule developed by Baiula and co-authors53 (Fig. 6A). We first performed a cell viability assay to determine the appropriate concentration and used the α4β1 agonist at concentrations of 2.5 µM in qRT-PCR and Transwell invasion assays (Fig. 6B).
The responses to strictosamide (1) and mitraphylline (2) were evaluated in cells exposed α4β1 agonist. (A) Chemical structure of the α4β1 agonist molecule. (B) Cell viability in ITGA4 agonist treated in AGS cells. Cells were treated with α4β1 agonist for 48 h with indicated concentration. (C) Relative mRNA levels of N-cadherin, ITGA4 and FAK, after treatment with strictosamide (1) and mitraphylline (2) for 48 h in AGS cells. (D) Representative images of each insert and relative number of invaded cells in the Transwell invasion assay. AGS cells were treated with strictosamide (1) and mitraphylline (2) at concentrations of 1 and 2.5 µM for 24 h. The α4β1 agonist was treated at a concentration of 2.5 µM. Data are presented as mean ± S.D. Statistical significance was determined as follows: *p < 0.05, **p < 0.01, ***p < 0.001, NS: not significant (p > 0.05) compared to DMSO-treated cells; #p < 0.05, ##p < 0.01, ###p < 0.001, NS: not significant (p > 0.05) compared to α4β1 agonist-treated cells; @p < 0.05, @@p < 0.01, @@@p < 0.001, NS: not significant (p > 0.05) compared to strictosamide (1)- or mitraphylline (2)-treated cells.
In α4β1 agonist treated cells, the mRNA levels of N-cadherin, ITGA4 and FAK were upregulated. However, in groups of cells co-treated with the α4β1 agonist and the alkaloids, N-cadherin and ITGA4 levels retained their capacity to be downregulated by strictosamide (1) and mitraphylline (2), whereas FAK levels were not significantly downregulated (Fig. 6C). We then checked these results with a phenotypic assay. The results were consistent in that the number of invaded cells induced by the α4β1 agonist treatment was reduced when co-treated with the alkaloids (Fig. 6D). Overall, treatment of strictosamide (1) and mitraphylline (2) continued to significantly suppress cell motility, although the response was attenuated in the cells treated with the ITGA4 agonist. Comparing between strictosamide (1) and mitraphylline (2), mitraphylline (2) exhibited a stronger inhibitory effect in α4β1 agonist-treated cells.
Discussion
Integrins provide outside-in signals that control cell destiny and mediate physical adhesion to appropriate matrices/ligands54. Among the integrin family, ITGA4 is crucial for cell survival, proliferation, migration, invasion, and tumor growth and spread, and high ITGA4 expression is associated with metastasis in many cancer types55. It has several ligands including fibronectin, osteopontin, fibrillin, thrombospondin, ADAM, COMP, and L148. Fibronectin is an important mesenchymal ECM glycoprotein involved in cell motility, matrix adherence, and cell-to-cell contact56. Furthermore, integrin clustering activates focal adhesion kinase (FAK) to cause tyrosine phosphorylation57. The focal adhesion complex is composed of the structural proteins vinculin, paxillin, and talin, along with SRC and FAK48. Several signaling pathways are controlled by FAK activation, which also causes EMT in different cancer cells58. The activation of the SRC/FAK pathway in cancer cells can result in several forms of invasion, which require different Rho GTPases59. Rho, RhoA, and RhoB family GTPases are important regulators of cell adhesion, cytoskeleton, and cell migration. RhoA activation promotes the production of actin stress fibers and protein clusters associated with focal adhesions60,61. RHOA upregulation promotes angiogenesis and cell migration62. The results of the present study indicated that strictosamide and mitraphylline affect cell motility by decreasing the expression of ITGA4-related genes (FAK, SRC, and RhoA) and EMT markers (N-cadherin, Vimentin, Twist, Snail, Slug, Zeb1, and Zeb2).
EMT is also intricately linked to the acquisition of cancer stem cell (CSC)-like properties and the development of resistance to anticancer therapies63,64. Cells undergoing EMT often exhibit stemness markers and features such as self-renewal and tumor-initiating capacity, which contribute to tumor heterogeneity and recurrence. Furthermore, EMT-driven changes in gene expression can promote survival pathways and reduce drug uptake or increase drug efflux, collectively leading to chemoresistance65,66. Promoting migratory and invasive properties, epithelial-to-mesenchymal transition (EMT) is functionally and mechanistically linked to the acquisition of cancer stem cell (CSC) characteristics. EMT markers such as Snail, Slug, Twist, and Zeb1 have been directly implicated in the induction of stemness in epithelial cancer cells66,67,68,69,. Therefore, the modulation of EMT by the investigated compounds in our study may have broader implications beyond cell motility, potentially influencing CSC-related traits and therapeutic resistance.
Strictosamide (1) and mitraphylline (2) downregulate the expression of other cell motility factors in combination with ITGA4/FAK/SRC/RHOs axis or independently. CAPN1 induces membrane detachment rate associated with peripheral focal adhesions, resulting in cell motility70. While Cav1 is associated with metastasis, it is also involved in cancer-related processes, such as tumor growth and drug resistance71. Cav1 phosphorylation promotes cell motility by promoting the association of stabilized FAK with focal adhesions through the RHO/ROCK signaling cascade72. In contrast, Wiskott-Aldrich syndrome proteins (WASPs) are involved in cellular processes through the transmission of GTPase activation signals73. WASF1/2/3 and WASL proteins promote the Arp2/3 (actin-related protein 2/3) complex in response to extracellular signaling, resulting in actin fiber recruitment and induction of actin polymerization74. Previous studies showed that WASF1 and WASL promote cancer cell motility75,76. With these knowledges, the results of the present study suggest that strictosamide and mitraphylline regulate multiple signaling pathways. Even through the pharmacological aspects of investigation of strictosamide (1) and mitraphylline (2) have been broadly studied, the present study is the first report— to the best of our knowledge—to highlight the cell motility-suppressive effects of the compounds. Strictosamide (1) has been previously reported to exhibit anti-inflammatory77,78 and anthelmintic79 activities, whereas mitraphylline (2) has shown anti-inflammatory80 and immunomodulatory81 activities. Notably, strictosamide (1) and mitraphylline (2) possess different skeleton types, although they are classified as MIAs. Strictosamide (1) is classified as having a rigid vallesiachotaman-type skeleton, whereas mitraphylline (2) is a spirocyclic oxindole alkaloid specifically distributed in the genus Uncaria82,83. However, future studies should comprehensively investigate the morphological changes of these alkaloids on actin and other cytoskeletal components that play a role in EMT processes and mechanical effects on the ITGA4/FAK axis. Differences in cellular processes because of different factors such as ITGA4 expression differences in cell lines and ECM substrates may cause differences in sensitivity and response to these alkaloids. Therefore, experimental designs such as ITGA4 overexpression and knockdown cell modeling or response tests with ITGA4 small molecule inhibitors can guide future studies. Another noteworthy point is that, mitraphylline (2) modulated EMT-related gene expression in a cell-type-specific manner. While ITGA4, FAK, and SRC were significantly downregulated in AGS and H1975 cells, CaCo2 cells showed limited changes in these targets, with downregulation observed instead in CAPN1, WASL1, and RHOB. This difference likely reflects the epithelial phenotype and lower EMT plasticity of CaCo2 cells, along with potential variations in basal signaling, mutational profile, or compound bioavailability84,85. These findings underscore the importance of cellular context in interpreting EMT modulation by natural compounds. Further investigations can help expand the potential of these two compounds and their analogs as anti-metastatic agents in different types of cancer therapy.
It should be noted that mitraphylline (2) undergoes spontaneous isomerization to isomitraphylline, a 7-epimer given via retro-Mannich interconversion of the spiro structure86. It was reported that the isomerization of mitraphylline (2) occurs in normal cell culture condition14. Thus, the biological activity shown in this study should be considered as a result from a mixture of mitraphylline (2) and isomitraphylline. This needs to be considered for further investigations performed in a molecular level.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Ganesh, K. & Massagué, J. Targeting metastatic cancer. Nat. Med. 27, 34–44 (2021).
Mittal, V. Epithelial mesenchymal transition in aggressive lung cancers. Adv. Exp. Med. Biol. 890, 37–56 (2016).
Said, N. A. B. M. & Williams, E. D. Growth factors in induction of epithelial-mesenchymal transition and metastasis. Cells Tissues Organs. 193, 85–97 (2011).
Lamouille, S., Xu, J. & Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell. Biol. 15, 178–196 (2014).
Santamaria, P. G., Moreno-Bueno, G., Portillo, F. & Cano, A. EMT: Present and future in clinical oncology. Mol. Oncol. 11, 718–738 (2017).
Ribatti, D., Tamma, R. & Annese, T. Epithelial-mesenchymal transition in cancer: A historical overview. Transl. Oncol. 13, 100773 (2020).
Huang, Y., Hong, W. & Wei, X. The molecular mechanisms and therapeutic strategies of EMT in tumor progression and metastasis. J. Hematol. Oncol. 15, 129 (2022).
Pan, Q., Mustafa, N. R., Tang, K., Choi, Y. H. & Verpoorte, R. Monoterpenoid Indole alkaloids biosynthesis and its regulation in Catharanthus roseus: A literature review from genes to metabolites. Phytochem. Rev. 15, 221–250 (2016).
O’Connor, S. E. & Maresh, J. J. Chemistry and biology of monoterpene Indole alkaloid biosynthesis. Nat. Prod. Rep. 23, 532–547 (2006).
Zhou, R. et al. The lichen secondary metabolite Atranorin suppresses lung cancer cell motility and tumorigenesis. Sci. Rep. 7, 8136 (2017).
Le, T. C. et al. Marine depsipeptide nobilamide I inhibits cancer cell motility and tumorigenicity via suppressing epithelial-mesenchymal transition and MMP2/9 expression. ACS Omega. 7, 1722–1732 (2022).
Pulat, S. et al. Marinobazzanan, a bazzanane-type sesquiterpenoid, suppresses the cell motility and tumorigenesis in cancer cells. Mar. Drugs. 21, 153 (2023).
Qin, N. et al. Recent research progress of Uncaria spp. based on alkaloids: Phytochemistry, Pharmacology and structural chemistry. Eur. J. Med. Chem. 210, 112960 (2021).
Kaiser, S. et al. Cat-s claw oxindole alkaloid isomerization induced by cell incubation and cytotoxic activity against T24 and RT4 human bladder Cancer cell lines. Planta Med. 79, 1413–1420 (2013).
Ndagijimana, A. et al. A review on Indole alkaloids isolated from Uncaria rhynchophylla and their Pharmacological studies. Fitoterapia 86, 35–47 (2013).
Azevedo, B. C. et al. Antioxidant activity of an aqueous leaf extract from Uncaria tomentosa and its major alkaloids Mitraphylline and Isomitraphylline in Caenorhabditis elegans. Molecules 24, 3299 (2019).
Chen, Z. et al. Monoterpene Indole alkaloids in Uncaria rhynchophlly (Miq.) jacks chinensis and their chemotaxonomic significance. Biochem. Syst. Ecol. 91, 104057 (2020).
Yang, A. et al. Chemical constituents of Uncaria scandens. Chem. Nat. Compd. 54, 793–794 (2018).
Zhang, J. G., Huang, X. Y., Ma, Y. B., Chen, J. J. & Geng, C. A. UFLC-PDA-MS/MS profiling of seven Uncaria species integrated with Melatonin/5-Hydroxytryptamine receptors agonistic assay. Nat. Prod. Bioprospect. 10, 23–36 (2020).
Bairoch, A. The Cellosaurus, a cell-line knowledge resource. J. Biomol. Tech. 29, 25–38 (2018).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Gamage, C. D. B. et al. Libertellenone T, a novel compound isolated from Endolichenic fungus, induces G2/M phase arrest, apoptosis, and autophagy by activating the ROS/JNK pathway in colorectal cancer cells. Cancers. 15, 489 (2023).
Flobak, Å. et al. A high-throughput drug combination screen of targeted small molecule inhibitors in cancer cell lines. Sci. Data 6, 237 (2019).
Mendiola, M. et al. Characterisation of new in vitro models and identification of potentially active drugs in angiosarcoma. Biomed. Pharmacother. 173, 116397 (2024).
Wang, M. et al. Sharing and community curation of mass spectrometry data with global natural products social molecular networking. Nat. Biotechnol. 34, 828–837 (2016).
Ferrari, F., Messana, I., Botta, B. & de Mello, J. F. Constituents of Guettarda platypoda. J. Nat. Prod. 49, 1150–1151 (1986).
Seki, H., Takayama, H., Aimi, N., Sakai, S-I. & Ponglux, D. A nuclear magnetic resonance study on the eleven stereoisomers of Heteroyohimbine-Type oxindole alkaloids. Chem. Pharm. Bull. 41, 2077–2086 (1993).
Aimi, N. et al. Isolation of alkaloids from cultured hybrid cells of Rauwolfia serpentina × Rhazya stricta. Chem. Pharm. Bull. 44, 1637–1639 (1996).
Achenbach, H. & Benirschke, M. Confirmation of the absolute configuration of Dolichantoside and isodolichantoside by synthesis from (–)-secologanin. Phytochemistry 44, 1387–1390 (1997).
Berger, A. et al. Loganin and Secologanin derived tryptamine-iridoid alkaloids from Palicourea crocea and Palicourea Padifolia (Rubiaceae). Phytochemistry 116, 162–169 (2015).
Li, L. T., Jiang, G., Chen, Q. & Zheng, J. N. Ki67 is a promising molecular target in the diagnosis of cancer (review). Mol. Med. Rep. 11, 1566–1572 (2015).
Yang, C. H. et al. EGFR over-expression in non-small cell lung cancers harboring EGFR mutations is associated with marked down-regulation of CD82. Biochim. Biophys. Acta. 1852, 1540–1549 (2015).
Lin, S. H. et al. Chidamide alleviates TGF-β-induced epithelial-mesenchymal transition in lung cancer cell lines. Mol. Biol. Rep. 43, 687–695 (2016).
Zhang, Z. et al. Advantages of restoring miR-205-3p expression for better prognosis of gastric cancer via prevention of epithelial-mesenchymal transition. J. Gastric Cancer 20, 212–224 (2020).
Kim, S. H. et al. Arsenic trioxide attenuates STAT-3 activity and epithelial-mesenchymal transition through induction of SHP-1 in gastric cancer cells. BMC Cancer. 18, 150 (2018).
Ghandi, M. et al. Next-generation characterization of the Cancer cell line encyclopedia. Nature 569, 503–508 (2019).
Forbes, S. A. et al. COSMIC: Exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–811 (2015).
He, Z., Dong, W., Li, Q., Qin, C. & Li, Y. Sauchinone prevents TGF-β-induced EMT and metastasis in gastric cancer cells. Biomed. Pharmacother. 101, 355–361 (2018).
Albano, F. et al. Representing ECM composition and EMT pathways in gastric cancer using a new metastatic gene signature. Front. Cell. Dev. Biol. 12, 1481818 (2024).
Schmuck, R. et al. Genotypic and phenotypic characterization of side population of gastric cancer cell lines. Am. J. Pathol. 178, 1792–1804 (2011).
Hidalgo, I. J., Raub, T. J. & Borchardt, R. T. Characterization of the human colon carcinoma cell line (Caco-2) as a model system for intestinal epithelial permeability. Gastroenterology 96, 736–749 (1989).
Keemink, J. & Bergström, C. A. S. Caco-2 cell conditions enabling studies of drug absorption from digestible lipid-based formulations. Pharm. Res. 35, 74 (2018).
Patankar, M., Eskelinen, S., Tuomisto, A., Mäkinen, M. J. & Karttunen, T. J. KRAS and BRAF mutations induce Anoikis resistance and characteristic 3D phenotypes in Caco–2 cells. Mol. Med. Rep. 20, 4634–4644 (2019).
Ikonomou, G. et al. Interplay between oncogenic K-Ras and wild-type H-Ras in Caco2 cell transformation. J. Proteomics 75, 5356–5369 (2012).
Liu, Y. & F Bodmer, W. Analysis of P53 mutations and their expression in 56 colorectal cancer cell lines. Proc. Natl. Acad. Sci. U. S. A. 103, 976–981 (2006).
Szklarczyk, D. et al. The STRING database in 2023: Protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Guan, J. L. Focal adhesion kinase in integrin signaling. Matrix Biol. 16, 195–200 (1997).
Mushtaq, U., Bashir, M., Nabi, S. & Khanday, F. A. Epidermal growth factor receptor and integrins Meet redox signaling through P66shc and Rac1. Cytokine 146, 155625 (2021).
Shankar, J. & Nabi, I. R. Actin cytoskeleton regulation of epithelial mesenchymal transition in metastatic cancer cells. PLoS ONE 10, e0119954 (2015).
Yoon, H., Dehart, J. P., Murphy, J. M. & Lim, S. T. Understanding the roles of FAK in cancer: Inhibitors, genetic models, and new insights. J. Histochem. Cytochem. 63, 114–128 (2015).
Zheng, D. et al. FAK regulates epithelial–mesenchymal transition in adenomyosis. Mol. Med. Rep. 18, 5461–5472 (2018).
Nurmagambetova, A., Mustyatsa, V., Saidova, A. & Vorobjev, I. Morphological and cytoskeleton changes in cells after EMT. Sci. Rep. 13, 22164 (2023).
Baiula, M. et al. New β-lactam derivatives modulate cell adhesion and signaling mediated by RGD-binding and leukocyte integrins. J. Med. Chem. 59, 9721–9742 (2016).
Shishido, S., Bönig, H. & Kim, Y. M. Role of integrin alpha4 in drug resistance of leukemia. Front. Oncol. 4, 99 (2014).
Xie, J. et al. Integrin Α4 up-regulation activates the Hedgehog pathway to promote arsenic and benzo[α]pyrene co-exposure-induced cancer stem cell-like property and tumorigenesis. Cancer Lett. 493, 143–155 (2020).
Gulubova, M. & Vlaykova, T. Immunohistochemical assessment of fibronectin and Tenascin and their integrin receptors Α5β1 and Α9β1 in gastric and colorectal cancers with lymph node and liver metastases. Acta Histochem. 108, 25–35 (2006).
Kornberg, L. J., Earp, H. S., Turner, C. E., Prockop, C. & Juliano, R. L. Signal transduction by integrins: Increased protein tyrosine phosphorylation caused by clustering of beta 1 integrins. Proc. Natl. Acad. Sci. U. S. A. 88, 8392–8396 (1991).
Huang, K. et al. Correlation between FAK and EGF-induced EMT in colorectal cancer cells. J. Oncol. 5428920 (2020).
Tornin, J. et al. FUS-CHOP promotes invasion in myxoid liposarcoma through a SRC/FAK/RHO/ROCK-dependent pathway. Neoplasia 20, 44–56 (2018).
Amano, M. et al. Phosphorylation and activation of myosin by Rho-associated kinase (Rho-kinase). J. Biol. Chem. 271, 20246–20249 (1996).
Kaibuchi, K., Kuroda, S. & Amano, M. Regulation of the cytoskeleton and cell adhesion by the Rho family GTPases in mammalian cells. Annu. Rev. Biochem. 68, 459–486 (1999).
Zhao, L. et al. The effect of RhoA on human umbilical vein endothelial cell migration and angiogenesis in vitro. Oncol. Rep. 15, 1147–1152 (2006).
Chu, X. et al. Cancer stem cells: Advances in knowledge and implications for cancer therapy. Signal. Transduct. Target. Ther. 9, 170 (2024).
Dart, A. EMT in chemoresistance. Nat. Rev. Cancer 23, 349 (2023).
Pradella, D., Naro, C., Sette, C. & Ghigna, C. EMT and stemness: Flexible processes tuned by alternative splicing in development and cancer progression. Mol. Cancer 16, 8 (2017).
Roy, S., Sunkara, R. R., Parmar, M. Y., Shaikh, S. & Waghmare, S. K. EMT imparts cancer stemness and plasticity: New perspectives and therapeutic potential. Front. Biosci. 26, 238–265 (2021).
Zhou, C. et al. ZEB1 confers stem cell-like properties in breast cancer by targeting neurogenin-3. Oncotarget 8, 54388–54401 (2017).
Khales, S. A, Mozaffari-Jovin, S., Geerts, D. & Abbaszadegan, M. R. TWIST1 activates cancer stem cell marker genes to promote epithelial-mesenchymal transition and tumorigenesis in esophageal squamous cell carcinoma. BMC Cancer 22, 1272 (2022).
Luanpitpong, S. et al. SLUG is required for SOX9 stabilization and functions to promote cancer stem cells and metastasis in human lung carcinoma. Oncogene 35, 2824–2833 (2016).
Storr, S. J., Carragher, N. O., Frame, M. C., Parr, T. & Martin, S. G. The Calpain system and cancer. Nat. Rev. Cancer 11, 364–374 (2011).
Xiong, N. et al. Involvement of caveolin-1 in low shear stress-induced breast cancer cell motility and adhesion: Roles of FAK/Src and ROCK/p-MLC pathways. Biochim. Biophys. Acta Mol. Cell. Res. 1864, 12–22 (2017).
Joshi, B. et al. Phosphorylated caveolin-1 regulates Rho/ROCK-dependent focal adhesion dynamics and tumor cell migration and invasion. Cancer Res. 68, 8210–8220 (2008).
Stradal, T. E. B. et al. Regulation of actin dynamics by WASP and WAVE family proteins. Trends Cell. Biol. 14, 303–311 (2004).
Ghoshal, P., Teng, Y., Lesoon, L. A. & Cowell, J. K. HIF1A induces expression of the WASF3 metastasis-associated gene under hypoxic conditions. Int. J. Cancer 131, E905–915 (2012).
Zhong, X. P. et al. NCKAP1 improves patient outcome and inhibits cell growth by enhancing Rb1/p53 activation in hepatocellular carcinoma. Cell. Death Dis. 10, 369 (2019).
Hidalgo-Sastre, A. et al. Loss of Wasl improves pancreatic cancer outcome. JCI Insight 5, e127275 (2020).
Li, D. et al. Anti-inflammatory effect of the six compounds isolated from Nauclea officinalis Pierrc ex Pitard, and molecular mechanism of strictosamide via suppressing the NF-κB and MAPK signaling pathway in LPS-induced RAW 264.7 macrophages. J. Ethnopharmacol. 196, 66–74 (2017).
Jia, Q. et al. Strictosamide alleviates the inflammation in an acute ulcerative colitis (UC) model. J. Physiol. Biochem. 77, 283–294 (2021).
Aderibigbe, S. A., Idowu, S. O., Olaniyi, A. A., Wright, C. W. & Fatokun, A. A. Bioactivity and cytotoxicity profiling of Vincosamide and Strictosamide, anthelmintic epimers from Sarcocephalus latifolius (Smith) Bruce leaf. J. Ethnopharmacol. 265, 113142 (2021).
Rojas-Duran, R. et al. Anti-inflammatory activity of mitraphylline isolated from Uncaria tomentosa bark. J. Ethnopharmacol. 143, 801–804 (2012).
la Montserrat-de, S. et al. Pharmacological effects of mitraphylline from Uncaria tomentosa in primary human monocytes: Skew toward M2 macrophages. J. Ethnopharmacol. 170, 128–135 (2015).
Ahmad, R. & Salim, F. Oxindole Alkaloids of Uncaria (Rubiaceae, Subfamily Cinchonoideae). In Studies In Natural Products Chemistry 485–525 (Elsevier, 2015).
Lopes, A. A. et al. Unnatural spirocyclic oxindole alkaloids biosynthesis in Uncaria Guianensis. Sci. Rep. 9, 11349 (2019).
Sambuy, Y. et al. The Caco-2 cell line as a model of the intestinal barrier: Influence of cell and culture-related factors on Caco-2 cell functional characteristics. Cell. Biol. Toxicol. 21, 1–26 (2005).
Hilgendorf, C. et al. Caco-2 versus Caco-2/HT29-MTX co-cultured cell lines: Permeabilities via diffusion, inside- and outside-directed carrier-mediated transport. J. Pharm. Sci. 89, 63–75 (2000).
Laus, G., Brössner, D., Senn, G. & Wurst, K. Analysis of the kinetics of isomerization of Spiro oxindole alkaloids. J. Chem. Soc. Perkin Trans. 2, 1931–1936 (1996).
Acknowledgements
We thank Dr. Mehdi Beniddir (Université Paris-Saclay, France) and Dr. Pierre Le Pogam (Université Paris-Saclay, France) for constructive discussion on the distribution of monoterpene indole alkaloids in the family Rubiaceae.
Funding
This work was supported by the National Research Foundation of Korea Grants (NRF-2021K1A3A1A21038059, 2022R1A5A2021216, and RS-2024-00413760) funded by the Korean government (MSIT), the KRIBB Initiative Program of Korea, and project ĐTĐL.CN-72/22 in Vietnam.
Author information
Authors and Affiliations
Contributions
Mücahit Varlı: Data Curation, Investigation, Methodology, Visualization, Writing – original draft. Kyungha Lee: Data Curation, Investigation, Writing – original draft. Tran The Bach: Investigation, Resources, Writing – review & editing. Tran Duc Binh: Investigation, Resources, Writing – review & editing. Hangun Kim: Conceptualization, Project administration, Supervision, Writing – review & editing. Kyo Bin Kang: Conceptualization, Project administration, Supervision, Writing – review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Varlı, M., Lee, K., Bach, T.T. et al. Strictosamide and mitraphylline inhibit cancer cell motility by suppressing epithelial-mesenchymal transition via integrin α4-mediated signaling. Sci Rep 15, 22807 (2025). https://doi.org/10.1038/s41598-025-04064-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-04064-7