Abstract
This study integrates bioinformatics analysis and in vitro experiments to elucidate the role of Target of EGR1 (TOE1) in hepatocellular carcinoma (HCC) progression. TOE1, a deadenylase belonging to the DEDD exonuclease superfamily, is primarily localized in the Cajal bodies of the nucleus. While the biological functions of TOE1 have been partially characterized, its role in cancer remains unclear. We analyzed TOE1 expression, its correlation with clinicalpathological features, prognosis, and immune infiltration in HCC using databases and platforms such as The Cancer Genome Atlas (TCGA), COSMIC, cBioPortal, and TIMER. Additionally, we validated TOE1 expression levels in clinical samples. Through the knockdown of TOE1 in HCC cell lines and subsequent RNA-seq analysis, we explored its functional role and potential molecular mechanisms in HCC malignant progression. The research focused on uncovering the role of TOE1 in HCC development and its underlying mechanisms. Comprehensive bioinformatics interrogation revealed that TOE1 was significantly upregulated in HCC and was correlated with poor patient prognosis. It may influence tumor development by modulating the stemness of tumor cells and immune cell infiltration, thereby reshaping the tumor microenvironment. Furthermore, in vitro experiments suggested that TOE1 promotes HCC proliferation and metastatic potential via modulating the Hippo signaling pathway. This research highlights the pivotal role of TOE1 in HCC, indicating its promise as a novel target for early detection, therapeutic strategies, immunological intervention, and prognosis assessment in HCC. These findings provide fresh perspectives for precision medicine in the context of HCC.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is categorized among the most frequently diagnosed malignancies globally and serves as the principal cause of cancer mortality in many developing countries. In most regions, HCC exhibits higher incidence and mortality rates in males compared to females1. Multiple elements contribute to the evolution of primary liver cancer, including hepatitis B and C viral infections, aflatoxin exposure, heavy alcohol intake, obesity, tobacco use, and type 2 diabetes mellitus2. For early-stage patients, surgical resection can be performed in only 5–15% of cases, but the five-year survival rate is only 18%. In the case of advanced-stage patients, interventions like ablation therapy and chemotherapy demonstrate restricted therapeutic efficacy3,4. Additionally, HCC is characterized by high recurrence and metastasis rates, resulting in poor prognosis and high mortality5. Therefore, improving the survival outcomes of HCC patients requires urgent exploration of its pathogenesis and identification of specific therapeutic targets.
Early growth response factor 1 (EGR1), a member of the EGR family, functions as a critical transcription factor6. The protein encoded by this gene contains an activation domain, an inhibitory domain, and three Cys2-His zinc finger motifs, enabling it to specifically identify and attach to target genes for transcriptional regulation7,8. EGR1 is widely distributed in various cells and is involved in tissue repair, immune responses, and fibrosis9. In tumorigenesis, EGR1 exerts a substantial influence on cell proliferation, angiogenesis, invasion, and immune responses10,11,12. Target of EGR1 (TOE1) is a target of EGR1 and is regulated by it. EGR1 modulates TOE1 expression by binding to its promoter region. However, the functional studies of TOE1 in cancer remain limited.
TOE1 is primarily localized in the nucleolus and Cajal bodies of the cell nucleus13. It is pivotal in regulating the maturation of small nuclear RNAs. TOE1 directly interacts with viral trans-activation response elements, thereby inhibiting HIV-1 replication. It is also involved in early embryonic development and telomere maintenance14. The CCCH zinc finger domain in the middle region of TOE1 is a C3H-type zinc finger protein. Zinc finger proteins participate in multiple biological functions, including tumor cell proliferation, migration, replication, repair, translation, and programmed cell death, thereby influencing tumor invasion, apoptosis, and drug resistance. However, research on TOE1 in cancer is scarce, with studies suggesting that TOE1 regulates cervical cancer cell growth through p2115.
The current research was designed to explore the function of TOE1 in the progression of HCC. We knocked down TOE1 in the HepG2 cell line and Huh7 cell line derived from HCC tissues. Functional evaluations were undertaken to measure the impacts of TOE1 knockdown on HCC cells. Additionally, high-throughput RNA sequencing (RNA-Seq) was performed to compare the global gene expression landscapes between TOE1-silenced and control groups, thereby identifying all genes targeted by TOE1. The findings revealed that TOE1 knockdown led to substantial changes in the expression of numerous genes. These results offer valuable perspectives on the function of TOE1 during the progression of liver cancer.
Methods
Microarray data collection
We obtained RNA-seq datasets and corresponding clinical data (including survival status, clinical staging, and pathological staging) from The Cancer Genome Atlas (TCGA; https://portal.gdc.cancer.gov/), comprising 371 tumor specimens and 50 normal hepatic tissue samples. Data processing and integration were performed with R version 4.4.2, and the R packages “limma” and “ggplot2” were employed for comparing TOE1 expression. For single-cell level validation in LIHC, we further analyzed two LIHC-specific single-cell datasets (GSE140228 and GSE98638) through TISCH16.
Survival analysis
Samples with incomplete survival records or undefined survival status were excluded during data preprocessing. Univariate Cox regression models were constructed using the “survival” package in R, and Kaplan-Meier survival curves were generated for each cancer subtype. The prognostic significance of TOE1 across various cancer subtypes was assessed: Survival endpoints included Overall Survival (OS), Disease-Specific Survival (DSS), Disease-Free Survival (DFS), and Progression-Free Survival (PFS). Prognostic significance was evaluated using Hazard Ratios (HRs), 95% confidence intervals (CIs), and P-value17.
Tumor Immune Estimation Resource (TIMER) 2.0
TIMER 2.0 (http://timer.cistrome.org/) is a web-based platform for systematic analysis of tumor-infiltrating immune cells across 32 TCGA cancer types, integrating 10,897 tumor samples from TCGA program18. We used the Gene_DE module to evaluate differential expression of the TOE1 gene between tumor and adjacent normal tissues across all TCGA cancer types. The Wilcoxon rank-sum test was utilized to evaluate statistical significance, with P < 0.05 considered significant.
cBioPortal analysis
The cBioPortal for Cancer Genomics (http://cbioportal.org) stands as a comprehensive web-based platform integrating data from multiple platforms. We queried the cBioPortal database to analyze TOE1 gene expression in TCGA datasets and to explore cancer mutation types.
TOE1 analysis with genomic instability and stemness
Tumor genomic heterogeneity closely influences the effects of immune checkpoint inhibitors (ICIs) therapy and its association with patient prognosis, including tumor mutation burden (TMB), microsatellite instability (MSI), tumor purity, and other indices19. We integrated TCGA sample MSI scores and tumor purity data and performed Spearman correlation analysis. TMB scores were computed using the “maftools” package in R20. Tumor stemness scores were derived using the OCLR algorithm proposed by Tarthiane-M et al., which calculates DNA stemness score (DNAss) based on methylation features for various malignant tumors21. Finally, stemness scores were integrated with gene expression data, and visualizations were performed using Pearson correlation analysis22.
TOE1 expression and immune cell infiltration correlation analysis
Through the SangerBox website platform (http://www.sangerbox.com), we analyzed the relationship between TOE1 expression levels and tumor microenvironment (TME) immune cell profiles across 33 human cancer types using three computational algorithms: QUANTISEQ, XCELL, and TIMER. The QUANTISEQ algorithm was then used to evaluate the relative abundance of nine immune cell types in LIHC patients based on TOE1 expression.
Drug sensitivity analysis
Drug sensitivity of anticancer drugs was predicted for each tumor specimen in TCGA-LIHC using the R package “oncoPredict"23. Ridge regression was conducted via the “oncoPredict” algorithm, and all aforementioned predictions were performed using data from the Genomics of Drug Sensitivity in Cancer (GDSC) database24.
Cell culture and transfection
We obtained the human HCC cell lines HepG2 and Huh7 from Hunan Fenghui Biotechnology Co., Ltd. Cells were maintained in high-glucose DMEM supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin-amphotericin B, under standard culture conditions (37 °C, 5% CO2). TOE1 expression was knocked down using TOE1-targeted siRNA (TOE1, NM_025077.4; siRNA sequence: 5′-GGAAGAGUUUGCUGAACCA-3′, designated as siTOE1). For transfection, 200 nM siRNA was used per 60-mm dish. Transfection was performed using Lipo8000 (Beyotime, C0533) following the supplier’s standardized protocol. Cells were harvested 24 h post-transfection for TOE1 expression analysis via qPCR and western blot.
Cell counting Kit-8 (CCK-8) assay
Cell proliferation and viability were quantitatively analyzed using CCK-8 assays. Following siRNA transfection, cells were treated with 10 µL of CCK-8 reagent at three time points (24, 48, and 72 h) to monitor growth kinetics. Cells were incubated at standard culture conditions for 40 min, then absorbance at 450 nm was read. Viability was calculated from the absorbance data.
Transwell assay
Cell invasion ability was evaluated by Matrigel-coated chambers. Matrigel (diluted in serum-free high-glucose DMEM) was added into Transwell chambers incubated (37 °C, 5% CO2) for 30 min to form a gel matrix. The upper chamber received 5 × 10⁴ cells in suspension, while the lower chamber contained 750 µL containing 15% FBS high-glucose DMEM medium. Following 24-hour incubation, migrated cells were stained and microscopically quantified. The cell count penetrating the membrane served as an indicator of invasive potential.
Raw data cleaning and alignment of RNA-seq
Forty-eight hours post-TOE1 transfection, cellular samples were harvested and submitted for high-throughput sequencing (Illumina NovaSeq 6000 platform; Shanghai OE Biotech). Raw sequencing reads were cleaned and mapped to known reference gene sequences using similarity-based methods to determine gene expression levels in each sample. Read counts mapping to protein-coding genes were quantified per sample using HTSeq-count, followed by removal of universally unexpressed genes (zero counts across all samples).
Differential expression gene (DEG) was analyzed. Gene expression analysis was conducted with DESeq2, implementing BaseMean-based count normalization, Log2 fold change calculation, and negative Binomial model for differential expression testing. DEGs were selected using the criteria of |log2FC| > 0.26 and P < 0.01. Gene ontology (GO) enrichment analysis was performed to describe the functional roles of DEGs based on GO annotations. For each GO category, we quantified the DEG representation and assessed enrichment significance through hypergeometric testing, with statistical significance defined as a false discovery rate (FDR) < 0.05. Statistical significance was assessed via Fisher’s exact test for each GO category (biological processes/cellular components/molecular functions), with P-values indicating enrichment confidence. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis employed hypergeometric testing to identify significantly enriched pathways (FDR < 0.05).
Western blotting
After 24-hour transfection of cells with siTOE1, total cellular proteins were isolated using RIPA lysis buffer (Beyotime, P0013B), strictly in accordance with the manufacturer’s instructions. Protein samples (30 µg per lane) were electrophoresed on 4–12% gradient gels (ACE, 11 wells), and subsequently transferred to 0.45 μm PVDF membranes (Millipore). The PVDF membranes containing protein were blocked with 8% skim milk for 1 h at 25 °C. Following blocking, membranes were probed with primary antibodies (overnight at 4 °C). The antibodies used in this study are anti-GAPDH (1:5000, ABcanol), anti-Yap (1:1000, ABcanol), anti-p-Yap (1:1000, ABcanol), anti-TOE1 (1:1000, Proteintech), anti-ULK1 (1:1000, ABcanol), anti-BECN1 (1:1000, ABcanol), anti-P21 (1:500, ZEN BIO), anti-P27 (1:1000, CST), anti-CCND1 (1:1000, CST) and anti-CCNE1 (1:1000, CST). After each primary antibody incubation, the PVDF membranes were incubated with species-matched secondary antibody (ABclonal, 1:10000) for 1 h at 25 °C. Protein bands were developed using ultra-sensitive ECL detection system (Beyotime, P0018S). ImageJ software was used to quantify the densities of the protein bands, and the values were normalized to the expression of GAPDH.
RNA isolation and quantitative reverse transcription polymerase chain reaction (qRT‑PCR)
Total RNA was isolated from Huh7 and HepG2 cells 48 h post-TOE1 transfection using RNA extraction kit (Fastagen, 220011) according to the manufacturer’s protocols. cDNA was reverse transcribed from total RNA and preserved at −80 °C. Quantitative PCR analysis was conducted with SYBR Green reagents (Yeasen, 11201ES08). Relative gene expression levels were calculated by normalizing to GAPDH endogenous control and applying the 2 − ΔΔCT quantification method. All primer sequences are detailed in Supplementary Table S1.
Tissue specimen acquisition
A single pair of human HCC tissue and adjacent non-tumorous liver tissue was acquired from a 58-year-old male patient who underwent surgical resection at The Seventh Affiliated Hospital of Sun Yat-sen University in 2023. All participants provided informed consent, with ethics approval obtained (Approval Number: KY-2025-366-01).
Immunohistochemical (IHC) staining
For IHC staining, 5 μm thick sections underwent xylene deparaffinization and ethanol gradient rehydration for IHC. Antigen retrieval was conducted with 10 mM citrate buffer (pH 6.0) via microwave irradiation (100 °C, 20 min). Endogenous peroxidase activity was quenched, followed by 5% goat serum blocking. Tissue sections were immunostained with the anti-TOE1 antibody at 4 °C overnight (1:50; 16203-1-AP, Proteintech). After being washed with PBS, sections received HRP-conjugated secondary antibody (37 °C, 40 min), followed by 3,3’-diaminobenzidine (DAB) staining (37 °C, 5 min) and hematoxylin counterstaining for 30 s.
Statistical analysis
Results are displayed as mean ± standard deviation (SD), based on three or more biological replicates. Statistical significance was determined by one-way analysis of variance (ANOVA) or two-tailed unpaired t-tests, with *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001, indicating significance levels. Results were displayed as histograms created using GraphPad Prism 8.0.2.
Results
TOE1 expression is upregulated in HCC
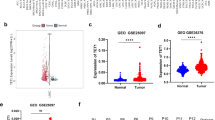
To investigate the abnormal expression of the TOE1 gene in HCC tissues, we conducted an expression analysis. Using the TIMER 2.0 platform, TOE1 expression levels were systematically compared between malignant tumors and paired adjacent normal tissues in 32 cancer types (Fig. 1A). Results showed significantly higher TOE1 expression in LIHC, cholangiocarcinoma (CHOL), esophageal cancer (ESCA), head and neck squamous cell carcinoma (HNSC), and stomach adenocarcinoma (STAD) (Wilcoxon rank-sum test, P < 0.05). Conversely, TOE1 was downregulated in kidney chromophobe carcinoma (KICH), kidney clear cell carcinoma (KIRC), lung squamous cell carcinoma (LUSC), prostate cancer (PRAD), and thyroid cancer (THCA) (P < 0.05). To investigate TOE1’s role in HCC, we assessed the association between TOE1 expression and clinical-pathological variables in LIHC patients from the TCGA database, including tumor type, patient gender, age, race, T-stage, N-stage, alcohol consumption, and body mass index (BMI). The results showed that in LIHC, TOE1 expression was overexpressed in primary cancer (Fig. 1B), significantly increased in advanced disease (Fig. 1G), and both increased with alcohol-free and alcohol consumption (Fig. 1H). Patients with low BMI exhibited significantly higher TOE1 expression compared to others (Fig. 1I). TOE1 expression levels demonstrated no significant associations with gender, age, or race (Fig. 1C–F). Overall, HCC tissues exhibited significantly elevated TOE1 expression relative to normal counterparts.
Elevated Target of EGR1 (TOE1) expression in hepatocellular carcinoma (HCC) tissues. (A) TOE1 expression in different cancer types from TCGA datasets. (B) Expression of TOE1 in Liver Hepatocellular Carcinoma (LIHC) tumors and normal tissues. (C–I) Analysis of TOE1 mRNA expression in patients with different clinicopathological parameters, including gender, age, race, staged T, individual cancer stage, alcohol consumption, and body mass index. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant. BMI: Body Mass Index.
Gene mutation analysis of LIHC
To characterize TOE1 genetic alterations in LIHC, we analyzed mutation profiles using the cBioPortal platform (Fig. 2A). The primary mutation type was “amplification” in most cancers, followed by “shallow deletion”, with diverse mutations identified. Further investigation of the specific mutations and mutation sites of TOE1 in cancer revealed that most mutations occurred in the CAF1 domain and were nonsense mutations (Fig. 2B). Using the COSMIC online database25, we obtained an overview of the mutation types. Consistently, the major mutation types were nonsense substitutions (61.84%), with the most common substitution types being C > T (32.26%) and G > A (28.49%) (Fig. 2C and D).
Mutation analysis of TOE1 in pan-cancer. (A) TOE1 alterations in pan-cancers in the cBioPortal database. (B) Types, loci, and number of TOE1 gene mutations in pan-cancer. (C) Types of TOE1 alterations in pan-cancers. (D) Sites of TOE1 mutation in pan-cancers.
TOE1 high expression independently predicts poor prognosis in LIHC patients
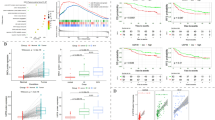
To evaluate the prognostic utility of TOE1 across cancers, univariate Cox proportional hazards regression analysis was performed for four survival endpoints: OS, DFS, DSS, and PFS. Cox proportional hazards model analysis revealed elevated TOE1 expression as a prognostic factor for OS across six types of cancers, including LIHC, GBM-LGG, LGG, LAML, ACC, and KICH (Fig. 3A). Further studies demonstrated that TOE1 expression showed significant DSS correlations with LIHC, GBM-LGG, LGG, ACC, and KICH tumor types (Fig. 3B). Additionally, high TOE1 expression was associated with lower DFS in KIRP tumors (Fig. 3C). Furthermore, TOE1 expression was found to correlate with poor prognosis across various cancers, including LIHC, GBM-LGG, LGG, and ACC, as assessed by PFS (Fig. 3D).
High TOE1 expression correlates with poor clinical outcomes of patients. (A–D) The forest plots shows the relationship of TOE1 expression with patient overall survival, disease-specific survival, disease-free interval, and progression-free interval. (E–H) Kaplan-Meier analysis of correlation between TOE1 expression and overall survival, disease specific survival, disease-free survival and progression free survival in LIHC patients. HR: Hazard ratios.
Given the significant association between LIHC and TOE1-mediated poor prognosis observed in the Cox regression analysis, we conducted survival analysis using the Kaplan-Meier plotter tool to assess the relationship of TOE1 expression with LIHC. A significant correlation was observed between high TOE1 expression and shorter OS (P = 6.3e-8), DSS (P = 7.1e-5), and PFS (P = 1.4e-3) (Fig. 3E, G) and it was also correlated with shorter DFS (P = 0.01) (Fig. 3F, H). These findings suggest that TOE1’s high expression may contribute to adverse prognosis in LIHC patients.
Role of TOE1 in the TME of HCC
To investigate TOE1 expression in cancerous and immune cells within the TME of HCC, we analyzed two single-cell RNA-seq datasets, GSE140228 and GSE98638. TOE1 expression was observed not just in tumor cells, but across various immune cell types, including T cells [CD4 T cells, CD8 T cells, Tprolif, and regulatory T cells (Tregs)], B cells, monocytes/macrophages, NK cells, and DCs (Fig. 4A). The extensive expression of TOE1 across diverse immune cell types underscores its potential functional role in the TME of LIHC (Fig. 4B).
Associations of the TOE1 expression level with immune cell infiltration. Single-cell sequencing analysis of TOE1 expression in (A) GSE140228 and (B) GSE98638. (C) The relationship between TOE1 expression and immune cell infiltration levels were analyzed in pan-cancer using QUANTISEQ Algorithm. (D–L) Individual TOE1 expression for LIHC patients and its correlation with infiltration levels of various immune cell types, including monocytes, M1 macrophages, NK cells, neutrophils, M2 macrophages, CD8 T cells, Tregs, B cells, and CD4 T cells. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
To further investigate the link between TOE1 expression and immune cell infiltration, we integrated three well-established cross-tumor immune scoring algorithms—QUANTISEQ, XCELL, and TIMER—based on immune infiltration scores. Findings revealed that TOE1 levels significantly influenced the infiltration levels of most immune cell types (Fig. 4C and Supplementary Fig. 1). Macrophages, natural killer (NK) cells, CD8 T cells, and Tregs were identified as the four immune cell types most strongly associated with TOE1 expression. Specifically, TOE1 showed significant positive correlations with B cells (r = 0.20), M1 macrophages (r = 0.22), M2 macrophages (r = 0.31), neutrophils (r = 0.10), CD8 T cells (r = 0.21), Tregs (r = 0.24), NK cells (r = 0.13), and CD4 T cells (r = 0.12) in LIHC (Figs. 4D–L). Although the correlation coefficients between TOE1 expression and immune cell infiltration were moderate, the associations remained statistically significant and consistent across multiple immune subsets. These results suggest that TOE1 may exert a broad but relatively subtle regulatory influence on immune cell infiltration within the HCC TME.
Interestingly, despite potential discrepancies among the three algorithms, TOE1 consistently showed robust associations with immune cell infiltration across various cancer types, including LIHC, KIRC, pan-kidney cohort (KIPAN), BRCA, and PAAD, exhibiting either robust positive or significant negative correlations. These results underscore the potential for conducting more in-depth investigations into the role of immune cell infiltration in the TME of various cancers.
Prediction of ICI treatment response by TOE1
Tumor stemness scores show strong associations with the development of drug resistance and sustained tumor cell proliferation during cancer treatment26. Typically, when the tumor stemness index is high, there is insufficient immune cell infiltration in the TME, and the expression level of PD-L1 is relatively low21. We evaluated the Pearson correlation between TOE1 gene expression and DNAss and RNA stemness scores (RNAss) across different tumor types. The analysis revealed that in tumors such as ovarian cancer (OV), thymoma (THYM), glioblastoma multiforme (GBM), lower grade glioma (LGG), and CHOL, TOE1 expression was significantly associated with DNAss (Fig. 5A). In contrast, in tumors like KIPAN, KIRC, PRAD, diffuse large B-cell lymphoma (DLBC), and testicular germ cell tumors (TGCT), TOE1 expression showed a stronger correlation with RNAss (Fig. 5B).
Prediction of treatment response to immune checkpoint inhibitors by TOE1. (A)Correlation between tumor stemness and TOE1 levels using DNAss. (B)Correlation between tumor stemness and TOE1 level using RNAss. (C) Correlation analysis between TOE1 expression and microsatellite instability, (D) tumor mutation burden, (E) tumor ploidy, and (F) tumor purity.
Given that TMB and MSI are critical determinants of the response to immune checkpoint inhibitor therapy, we conducted a further investigation into the associations between TOE1 expression and TMB as well as MSI. In most tumor types, TOE1 expression levels were positively correlated with both TMB and MSI. Notably, the correlation between TOE1 and MSI was particularly significant in patients with acute myeloid leukemia (LAML), THYM, adrenocortical carcinoma (ACC), GBM, and colorectal adenocarcinoma (COAD) (Fig. 5C). Additionally, the correlation between TOE1 and TMB scores was strongest in DLBC, GBM, KIPAN, PRAD, and TGCT (Fig. 5D).
Assessing tumor purity and ploidy is crucial for understanding cancer genome evolution and tumor heterogeneity, which are also considered key factors influencing the efficacy of ICIs. TOE1 expression was significantly correlated with tumor ploidy, particularly in medullary THYM, uterine sarcoma (UCS), GBM, KIRC, and pheochromocytoma and paraganglioma (PCPG) (Fig. 5E). Furthermore, in uveal melanoma (UVM), PRAD, CHOL, KIPAN, sarcoma (SARC), and ACC, TOE1 expression was also significantly associated with tumor purity (Fig. 5F).
In this context, the positive correlation between DNAss, RNAss, and TOE1 expression may suggest that PD-L1-based therapies are less effective in patients with high TOE1 expression. Similarly, in LIHC patients, both TMB and MSI showed significant positive correlations with TOE1 expression. Additionally, tumor purity and ploidy were significantly positively correlated with TOE1 expression. Therefore, further studies are needed to identify targeted therapies for LIHC patients with high TOE1 expression.
Correlation between TOE1 and LIHC drug sensitivity
Improving tumor cell sensitivity to different drugs remains a critical challenge in LIHC treatment. Based on the GDSC database, we investigated the sensitivity of LIHC patients with different TOE1 expression levels to targeted therapy and chemotherapy. We identified the top 10 drugs (Bortezomib, Daporinad, Dinaciclib, Docetaxel, Eg5_9814, Paclitaxel, Sepantronium bromide, Vincristine, Vinblastine, and Podophyllotoxin bromide) that showed differential sensitivity based on TOE1 expression (Fig. 6A–J). Interestingly, for all 10 drugs, the high-TOE1 expression group exhibited higher sensitivity compared to the low-expression group. Among these, Bortezomib, Daporinad, Docetaxel, and Sepantronium bromide showed the highest sensitivity in the high-TOE1 expression group. These findings provide valuable insights for identifying targeted therapies for patients with high TOE1 expression in LIHC.
Relationship of TOE1 expression with Genomics of Drug Sensitivity in Cancer (GDSC) drug response. (A–J) Bortezomib, Daporinad, Dinaciclib, Docetaxel, Eg5_9814, Paclitaxel, Sepantronium bromide, Vincristine, Vinblastine and Podophyllotoxin bromide, respectively. **P < 0.01, ****P < 0.0001.
Validation of the expression of TOE1 in HCC
To explore the functional involvement of TOE1 in HCC development, we performed immunohistochemical analysis on paired tumor and adjacent non-tumorous liver tissues from HCC patients. The results showed significantly more intense TOE1 protein staining in HCC specimens compared to matched adjacent non-tumorous tissues, demonstrating elevated TOE1 expression in malignant hepatocytes (Fig. 7).
Validation of TOE1 expression in HCC using immunohistochemistry analysis. (A) Representative images of adjacent non-tumor tissues and HCC tumor tissues. (B) Quantification of TOE1 expression levels based on the average optical density (AOD) from six randomly selected fields per group. ****P < 0.0001.
Validation of TOE1 knockdown in HCC cell lines
To investigate the biological roles of TOE1 in HCC, we knocked down TOE1 in HCC cell lines HepG2 and Huh7. Three siRNAs targeting the TOE1 transcript (NM_025077.4)—siTOE1-1, siTOE1-2, and siTOE1-3—were designed and transfected into HepG2 cells. qPCR analysis showed that siTOE1-2 exhibited the most efficient transcriptional interference compared to siTOE1-1 and siTOE1-3 (Fig. 8A), prompting its selection for subsequent experiments.
Verification of the silencing effect of TOE1 knockdown by transfection with non-specific control siRNA (NC) or TOE1 siRNA in HCC cells. (A) TOE1 mRNA levels in HepG2 cells by qRT-PCR. (B) TOE1 mRNA levels in HepG2 cells by qRT-PCR. (C) TOE1 mRNA levels in Huh7 cells by qRT-PCR. (D)TOE1 protein levels in HepG2 cells by western blot analysis. (E)TOE1 protein levels in Huh7 cells by western blot analysis. *P < 0.05, ***P < 0.001, ****P < 0.0001; NC, non-specific control; siTOE1, TOE1 knockdown.
TOE1 knockdown inhibits the malignant proliferation of HCC cells. (A, C, D) In Transwell assay, detection of migration ability of HepG2 and Huh7 cells. (B, E, and F) In Transwell assay, detection of invasion ability of HepG2 and Huh7 cells. (G) Cell viability of different groups of cells in HepG2. (H) Cell viability of different groups of cells in Huh7 cells. **P < 0.01, ***P < 0.001, ****P < 0.0001; NC, non-specific control; siTOE1, TOE1 knockdown.
We transfected siTOE1-2 (siTOE1) and non-targeting control siRNA (NC) into HepG2 and Huh7 cells. Total RNA and protein samples were isolated from the transfected cells for subsequent qPCR and western blot analyses. qPCR data revealed significantly reduced TOE1 mRNA levels in both HepG2 and Huh7 cell lines compared to the control group (Fig. 8B–C). Western blot confirmed a corresponding decrease in TOE1 protein expression following siTOE1 treatment (Fig. 8D–E). These results demonstrated successful knockdown of TOE1 in HCC cells.
TOE1 knockdown inhibits the malignant proliferation of HCC cells
To investigate the role of TOE1 in HCC development, we evaluated the effects of TOE1 knockdown on the migration and invasion capabilities of HCC cells. Our experimental data indicated that TOE1 knockdown significantly inhibited the migration and invasion of HCC cells, as observed in both HepG2 and Huh7 cell lines (Fig. 9A–F). Furthermore, CCK-8 assays showed that TOE1 silencing led to a significant reduction in the proliferation of HCC cells (Fig. 9G and H). These findings suggest that TOE1 serves as a key regulator of HCC cell migration, invasion, and proliferation progression.
TOE1 knockdown alters gene expression profiles in HepG2 cells
To elucidate the molecular pathways mediating TOE1 expression in HCC development, we performed RNA-seq analysis comparing gene expression patterns between TOE1 siRNA knockdown and scramble siRNA-treated controls in HepG2 cells. The fragments per kilobase million (FPKM) method was used to normalize gene expression levels. We obtained expression results for 22,373 genes and generated a stacked column chart to visualize the distribution of gene expression levels (Fig. 10A).
TOE1 knockdown widely affects the gene expression profile in HepG2 cells. (A) Distribution of gene expression in NC and TOE1 knockdown samples. (B) The Pearson correlation analysis results revealed high transcriptional concordance between NC and siTOE1-treated samples. (C) TOE1 expression quantified by RNA sequencing data. (D) Volcano plot of TOE1-Knockdown Transcriptome. Differentially expressed genes (DEGs) were identified in TOE1-depleted cells compared to scramble controls (|log2FC|>0.26, adjusted P < 0.01). (E) A heatmap was created to display the expression levels of each DEG. **P < 0.01; NC, non-specific control; siTOE1, TOE1 knockdown.
A Pearson correlation matrix constructed from FPKM values demonstrated high consistency between biological replicates in both groups, with the diagonal heatmap reflecting strong correlation within the TOE1-knockdown and control samples (Fig. 10B). RNA-seq analysis further confirmed the effective knockdown of TOE1 (Fig. 10C). Differential expression analysis (|log₂FC| > 0.26, P < 0.01) identified 3,275 DEGs, including 1,492 upregulated and 1,783 downregulated genes (Fig. 10D). Additionally, the consistent expression patterns of these DEGs across three biological replicates in both siTOE1 and control samples were confirmed (Fig. 10E). These data indicate that TOE1 broadly modulates transcriptional networks in HepG2 cells.
Functional analysis of DEGs following TOE1 knockdown in HepG2 cells
To characterize the molecular consequences of TOE1 knockdown, GO enrichment analysis was performed on both up- and down regulated DEGs. GO analysis was categorized into three main classifications: biological process, cellular component, and molecular function. Functional enrichment analysis revealed that the down regulated DEGs were enriched in processes such as cholesterol biosynthetic process, positive regulation of endothelial cell migration, and angiogenesis, as well as molecular functions such as extracellular matrix structural constituent and clathrin adapter activity (Fig. 11A). Up regulated DEGs were enriched in processes such as rRNA processing, regulation of transcription from DNA templates, and nuclear lumen, as well as molecular functions such as protein binding and RNA binding (Fig. 11C).
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses were performed to identify functional alterations associated with TOE1 silencing in HepG2 cells. (A) Top 30 enriched GO terms for downregulated genes after TOE1 knockdown. (B) Top 20 enriched KEGG pathways with downregulated after TOE1 knockdown. (C) Top 30 GO terms for upregulated genes after TOE1 knockdown. (D) Top 20 enriched KEGG pathways with upregulated pathways after TOE1 knockdown. NC, non-specific control; siTOE1, TOE1 knockdown.
To further elucidate the potential functional pathways involved in TOE1-mediated HCC development, we performed KEGG pathway analysis (Fig. 11B–D). The analysis demonstrated that the downregulated DEGs were significantly enriched in key pathways such as cellular senescence, focal adhesion, and the Hippo signaling pathway (Fig. 11B). The upregulated DEGs showed significant enrichment in pathways such as Autophagy, Apoptosis − Multiple Species, and Apoptosis (Fig. 11D). These results indicate that TOE1 regulates multiple signaling pathways involved in HCC development, including tumor autophagy and apoptosis.
Validation of TOE1-regulated genes in HCC cells
To validate key DEGs linked to cancer progression, we selected ten DEGs for functional annotation. Specifically, Cyclin-Dependent Kinase 2 (CDK2), Cyclin E1 (CCNE1), Cyclin D1 (CCND1), P21, and P27kip are genes involved in cell cycle regulation; Ferritin Heavy Chain 1 (FTH1) is associated with ferroptosis; Unc-51-like kinase 1 (ULK1), Beclin 1 (BECN1), and Autophagy-Related 3 (ATG3) are associated with autophagy; Ubiquitin C (UBC) is associated with ubiquitination. Transcriptome data showed that among the ten DEGs, four genes (CDK2, CCNE1, CCND1, and ULK1) were downregulated (Fig. 12A–D), while six genes (UBC, FTH1, P27kip, P21, ATG3, and BECN1) were upregulated (Fig. 12E–J). To validate the RNA-seq findings, we performed qPCR on selected DEGs following TOE1 knockdown in HCC cells, confirming significant alterations in mRNA levels. The findings were in accordance with the results derived from the RNA-seq analysis (Fig. 13). Furthermore, western blot analysis provided further validation for the alterations in the protein-level expression of CCNE1, ULK1, BECN1, CCND1, p27, and p21, showing significant decreases in CCNE1 and CCND1 expression, as well as significant increases in ULK1, BECN1, p27, and p21 expression after interference (Fig. 14). These findings indicate that TOE1 exerts a positive regulatory effect on the expression of P21, FTH1, P27 kip, BECN1, ATG3, and UBC, while negatively regulating CDK2, CCNE1, and CCND1.
Transcriptome results of expression levels of genes regulated by TOE1. (A-C) Transcriptome expression levels of three DEGs downregulated upon siTOE1 compared to NC cells. (D-I) Transcriptome expression levels of six DEGs upregulated upon siTOE1 compared to NC cells. *P < 0.05, **P < 0.01, ***P < 0.001; NC, non-specific control; siTOE1, TOE1 knockdown.
Validation of genes regulated by TOE1 in HepG2 and Huh7 cells by qPCR. (A-C) qPCR validation of three DEGs downregulated upon siTOE1 compared to NC cells. (D-I) qPCR validation of six DEGs upregulated upon siTOE1 compared to NC cells. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; NC, non-specific control; siTOE1, TOE1 knockdown.
Protein levels of CCNE1, ULK1, BECN1, CCND1, P27, and P21 in HCC cells analyzed by Western blot. (A) Western blot analysis of CCNE1, ULK1, BECN1, CCND1, P27, and P21 in HepG2 cells transfected with NC or siTOE1 (GAPDH as loading control). (B) Western blot analysis of CCNE1, ULK1, BECN1, CCND1, P27, and P21 in Huh7 cells transfected with NC or siTOE1 (GAPDH as loading control). (C-H) Quantitative analysis of the relative protein levels normalized to GAPDH for each target protein in HepG2 and Huh7 cells. *P < 0.05, **P < 0.01, ***P < 0.001, ns: not significant; NC, non-specific control; siTOE1, TOE1 knockdown.
TOE1 regulates HCC development via the Hippo signaling pathway
KEGG pathway enrichment analysis of the downregulated genes in the siTOE1 group revealed their enrichment in the Hippo signaling pathway. Members of the Hippo pathway, including Yes1 - Associated Transcriptional Regulator (YAP1), TEA Domain Transcription Factor 1 (TEAD1), and TEA Domain Transcription Factor 2 (TEAD2), exhibited downregulation in the siTOE1 group (Fig. 15A-B). qRT-PCR validation confirmed these findings, and Western blot analysis revealed a marked reduction in YAP1 protein abundance after TOE1 knockdown (Fig. 15D-E). Taken together, these outcomes indicate that TOE1 knockdown leads to the downregulation of members of the Hippo signaling pathway. Therefore, TOE1 may regulate HCC development through the Hippo signaling pathway.
TOE1 promotes HCC progression through the Hippo pathway. (A-C) Fragments Per Kilobase Million (FPKM) levels and mRNA Expression levels of YAP1, TEAD1, and TEAD2 in HCC cells with siTOE1 compared to NC. (D) Western blot analysis of YAP1 protein levels in HepG2 cells after siTOE1 and NC (GAPDH loading control). Quantification of relative YAP1 protein levels is shown in the lower panel. (E) Western blot analysis of YAP1 expression in Huh7 cells after siTOE1 and NC (GAPDH as loading control). *P < 0.05, **P < 0.01, ***P < 0.001; NC, non-specific control; siTOE1, TOE1 knockdown.
Discussion
HCC ranks among the most prevalent malignant tumors worldwide, with a persistently low long-term survival rate despite advancements in early-stage diagnosis and treatment. Given the unique nature of liver tissue, HCC is characterized by high aggressiveness, metastatic potential, and extreme heterogeneity, consequently causing a five-year survival rate of only 5% for HCC patients27. Therefore, identifying new influencing factors for HCC holds significant importance for its diagnosis and treatment. However, comprehensive research exploring the association between TOE1 expression and the development of HCC remains limited. The objective of this investigation is to thoroughly explore the function of TOE1 in HCC and its therapeutic potential by the analysis of public databases, transcriptomic profiling, and in vitro experiments.
TOE1, as a member of the zinc finger protein family, may play a role similar to other zinc finger proteins, which have been implicated in tumor progression and prognosis. For instance, elevated ZNF689 levels can influence HCC growth and contribute to poor patient outcomes28,29. ZNF746 promotes HCC progression by suppressing PGC-1α transcription30, while ZNF687 enhances the formation of tumor spheres, invasion, and chemotherapy resistance in HCC cells by upregulating pluripotency-related factors such as BMI1, OCT4, and NANOG31. Additionally, ZNFs like ZNF233, ZNF143, and ZNF384 have been shown to accelerate the cell cycle and drive tumor growth32,33,34. Based on bioinformatics analysis across multiple databases, significant overexpression of TOE1 was observed in HCC patients, which correlates with advanced tumor stages and unfavorable prognostic factors. Our findings suggest that TOE1 may drive HCC progression by promoting tumor cell proliferation, migration, and invasion. Furthermore, immunohistochemical analysis confirmed the significantly high expression of TOE1 in HCC tissues, underscoring its potential as a valuable biomarker for HCC.
The Hippo signaling pathway, initially identified in Drosophila melanogaster, represents a highly conserved kinase cascade regulating pivotal biological processes, including proliferation, differentiation, and survival35. In HCC, our findings reveal that TOE1 negatively regulates key Hippo pathway components—YAP1, TEAD1, and TEAD2—which are frequently dysregulated in human malignancies. Aberrant activation of Hippo effectors, particularly nuclear accumulation of YAP/TAZ and elevated TEAD expression, correlates with poor prognosis across cancers36, a phenomenon prominently observed in HCC37. Mechanistically, TOE1 knockdown suppresses HCC proliferation and invasion by inhibiting Hippo signaling effectors YAP1 and TEAD1/2, as well as cyclin-dependent kinases CCND1, CCNE1, and CDK2. This inhibition of CDK activity disrupts cell cycle progression, a critical driver of tumor proliferation.
The cyclin-dependent kinase inhibitor p21 (CDKN1A) serves as a critical cell cycle regulator by inhibiting CDK activity, leading to G2/M arrest38. Our findings indicate that TOE1 knockdown elevates p21 expression, thereby inducing G2/M arrest and effectively halting cancer cell division. This aligns with Hippo signaling’s canonical tumor-suppressive role in restraining YAP/TAZ-TEAD transcriptional programs. Notably, TOE1’s dual regulation of Hippo effectors and cyclins suggests a feedforward loop that amplifies proliferative signals, a mechanism that may be conserved across malignancies.
Autophagy exhibits tumor suppressive effects by degrading damaged cellular components and the elimination of aberrant cells, thereby impeding the initiation and advancement of malignant tumors39. BECN1 is considered a tumor suppressor that activates autophagy and prevents tumorigenesis40. ATG3, as a key enzyme of autophagy, plays a crucial role in autophagosome membrane formation. FTH1 is a core component of ferritin responsible for the storage and release of iron. The high expression of FTH1 may inhibit tumor cells by promoting iron-dependent cell death proliferation and invasion. In addition to its role in cell cycle regulation, TOE1 modulates tumor metabolism through autophagy and ferroptosis. Our data show that TOE1 depletion upregulates autophagy regulators BECN1 and ATG3, as well as ferroptosis-associated FTH1. This highlights TOE1’s role in metabolic reprogramming and underscores its tumor-suppressive function during early carcinogenesis. The coordinated modulation of autophagy and cell cycle checkpoints by TOE1 suggests a delicate balance between pro-survival and pro-death pathways, fine-tuning tumor fitness in HCC.
Furthermore, TOE1’s association with immunosuppressive microenvironments, characterized by increased M2 macrophages, Tregs, and reduced CD8+ T cell activity, highlights its potential to dampen antitumor immunity. It is noteworthy that some correlations between TOE1 expression and immune infiltration levels exhibited modest r values. Such moderate correlations are frequently observed in complex tumor immune microenvironments, where multiple factors jointly determine immune cell recruitment and functional heterogeneity41,42. Nevertheless, the consistent positive trends across immune subsets imply that TOE1 may indirectly modulate immune responses, possibly through post-transcriptional regulation or Hippo–YAP/TAZ–related signaling. This aligns with emerging evidence linking Hippo signaling to immune evasion, suggesting that TOE1 inhibition might synergize with immune checkpoint therapies43,44. Additionally, TOE1-high tumors exhibit heightened sensitivity to chemotherapeutics, such as Bortezomib, offering a rationale for stratified treatment strategies. TOE1’s expression may influence tumor progression by indirectly affecting immune cell infiltration in the tumor microenvironment. Its significant positive correlation with tumor stemness scores further supports its critical role in tumor microenvironment remodeling.
Our single-cell transcriptomic analysis revealed that TOE1 is expressed in multiple immune cell populations within the HCC microenvironment, including macrophages and T cells. Although its immune-specific role has not yet been experimentally defined, TOE1 functions as a nuclear RNA-processing factor45,46 and has been reported to interact with components of the Hippo signaling pathway. Recent evidence indicates that aberrant Hippo-YAP activation promotes tumor immune escape by regulating macrophage-mediated phagocytosis, while YAP inhibition reshapes macrophage distribution and function44. Moreover, YAP/TAZ activation enhances the expression of Arg1, an M2 macrophage marker47, and Wnt5a-mediated YAP/TAZ upregulation facilitates TGF-β1–induced M2 polarization48. In HCC, hepatocyte YAP activation promotes CCL2-dependent macrophage recruitment and drives M2 polarization49. Taken together, these findings suggest that TOE1 may influence macrophage function in HCC by modulating the Hippo-YAP/TAZ pathway. Further functional studies are warranted to elucidate the mechanistic role of TOE1 in immune cells, particularly macrophages, and its potential significance in regulating the HCC immune microenvironment.
Despite the comprehensive exploration of TOE1’s function in HCC in this research, there are still some limitations. First, this study mainly focused on HepG2 and Huh7 cell lines and lacked validation on other HCC cell lines and primary cells. Second, the specific molecular mechanism of TOE1 regulating the Hippo signaling pathway still needs to be further explored. In addition, the drug sensitivity analysis in this study revealed that elevated TOE1 expression could correlate with the response to some chemotherapeutic drugs, but its potential as a therapeutic target still needs further validation.
In summary, the current investigation revealed the critical function of TOE1 in HCC and its potential mechanism to influence tumor progression through multiple molecular mechanisms. TOE1 emerges as a promising target for diagnosis, therapy, and prognostic assessment of HCC. Further studies on the mechanism of TOE1 in different cancer types and its clinical application value as a therapeutic target are needed in the future.
Data availability
The datasets generated and analysed during the current study are available in the GEO repository, under accession number GSE304375. Publicly available datasets were obtained from the Gene Expression Omnibus under accession numbers GSE140228 and GSE98638.
Abbreviations
- TOE1:
-
Target of EGR1
- HCC:
-
Hepatocellular carcinoma
- TCGA:
-
The Cancer Genome Atlas
- EGR1:
-
Early growth response factor 1
- RNA-Seq:
-
RNA sequencing
- LIHC:
-
Liver hepatocellular carcinoma
- TISCH:
-
Tumor Immune Single Cell Hub
- OS:
-
Overall survival
- DSS:
-
disease-specific survival
- DFS:
-
Disease-free Survival
- PFI:
-
Progression-free interval
- HR:
-
Hazard Ratio
- TIMER:
-
Tumor Immune Estimation Resource
- ICIs:
-
Immune checkpoint inhibitors
- TMB:
-
Tumor mutation burden
- MSI:
-
Microsatellite instability
- DNAss:
-
DNA stemness score
- RNAss:
-
RNA stemness score
- GDSC:
-
Genomics of Drug Sensitivity in Cancer
- FBS:
-
Fetal bovine serum
- IHC:
-
Immunohistochemical
- CHOL:
-
Cholangiocarcinoma
- PRAD:
-
Prostatic cancer
- KIPAN:
-
Pan-kidney carcinoma
- DLBC:
-
Diffuse large B-cell lymphoma
- TGCT:
-
Testicular germ cell tumors
- ESCA:
-
Esophageal carcinoma
- HNSC:
-
Head and neck squamous cell carcinoma
- STAD:
-
Stomach adenocarcinoma
- KICH:
-
Kidney Chromophobe
- KIRC:
-
Kidney renal clear cell carcinoma
- LUSC:
-
Lung squamous cell carcinoma
- THCA:
-
Thyroid cancer
- LGG:
-
Lower Grade Glioma
- UCS:
-
Uterine Carcinosarcoma
- PCPG:
-
Pheochromocytoma and Paraganglioma
- UVM:
-
Uveal Melanoma
- SARC:
-
Sarcoma
- BMI:
-
Body Mass Index
- NK cells:
-
Natural killer cells
- THYM:
-
Thymoma
- ACC:
-
Adrenocortical carcinoma
- GBM:
-
Glioblastoma multiforme
- DEG:
-
Differential Expression Gene
- FPKM:
-
Fragments per kilobase million
- CDK2:
-
Cyclin-Dependent Kinase 2
- CCNE1:
-
Cyclin E1
- CCND1:
-
Cyclin D1
- BECN1:
-
Beclin 1
- ATG3:
-
Autophagy-Related 3
- UBC:
-
Ubiquitin C
- YAP1:
-
Yes1-Associated Transcriptional Regulator
- TEAD1:
-
TEA Domain Transcription Factor 1
- TEAD2:
-
TEA Domain Transcription Factor 2
- LAML:
-
Acute Myeloid Leukemia
- PFS:
-
Progression Free Survival
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263. https://doi.org/10.3322/caac.21834 (2024).
Vogel, A., Meyer, T., Sapisochin, G., Salem, R. & Saborowski, A. Hepatocellular carcinoma. Lancet 400, 1345–1362. https://doi.org/10.1016/s0140-6736(22)01200-4 (2022).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 70, 7–30. https://doi.org/10.3322/caac.21590 (2020).
Anwanwan, D., Singh, S. K., Singh, S., Saikam, V. & Singh, R. Challenges in liver cancer and possible treatment approaches. Biochim. Et Biophys. Acta (BBA)-Reviews Cancer. 1873, 188314 (2020).
Kim, E. & Viatour, P. Hepatocellular carcinoma: old friends and new tricks. Exp. Mol. Med. 52, 1898–1907. https://doi.org/10.1038/s12276-020-00527-1 (2020).
Sukhatme, V. P. et al. A zinc finger-encoding gene coregulated with c-fos during growth and differentiation, and after cellular depolarization. Cell 53, 37–43. https://doi.org/10.1016/0092-8674(88)90485-0 (1988).
Christy, B. & Nathans, D. DNA binding site of the growth factor-inducible protein Zif268. Proc. Natl. Acad. Sci. U S A. 86, 8737–8741. https://doi.org/10.1073/pnas.86.22.8737 (1989).
Lemaire, P. et al. The serum-inducible mouse gene Krox-24 encodes a sequence-specific transcriptional activator. Mol. Cell. Biol. 10, 3456–3467. https://doi.org/10.1128/mcb.10.7.3456-3467.1990 (1990).
Wang, B. et al. The role of the transcription factor EGR1 in cancer. Front. Oncol. 11, 642547. https://doi.org/10.3389/fonc.2021.642547 (2021).
Chen, J., Zhan, Y., Xu, J., Wang, Y. & Gao, Q. EGR1 overexpression inhibits the occurrence of preeclampsia by binding to MicroRNA-574 promoter and upregulating GAB1. Reprod. Sci. 28, 1112–1121. https://doi.org/10.1007/s43032-020-00392-3 (2021).
Du, K., Wu, X., Ji, X., Liang, N. & Li, Z. Early growth response 1 promoted the invasion of glioblastoma multiforme by elevating HMGB1. J. Neurosurg. Sci. 67, 422–430. https://doi.org/10.23736/S0390-5616.20.05107-3 (2023).
Wu, Y. et al. Beta-Defensin 2 and 3 promote bacterial clearance of Pseudomonas aeruginosa by inhibiting macrophage autophagy through downregulation of early growth response Gene-1 and c-FOS. Front. Immunol. 9, 211. https://doi.org/10.3389/fimmu.2018.00211 (2018).
Fong, K. W. et al. Whole-genome screening identifies proteins localized to distinct nuclear bodies. J. Cell. Biol. 203, 149–164. https://doi.org/10.1083/jcb.201303145 (2013).
Deng, T. et al. TOE1 acts as a 3′ exonuclease for telomerase RNA and regulates telomere maintenance. Nucleic Acids Res. 47, 391–405 (2019).
Sperandio, S., Tardito, S., Surzycki, A., Latterich, M. & de Belle I. TOE1 interacts with p53 to modulate its transactivation potential. FEBS Lett. 583, 2165–2170. https://doi.org/10.1016/j.febslet.2009.06.004 (2009).
Sun, D. et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 49, D1420–D1430. https://doi.org/10.1093/nar/gkaa1020 (2021).
Zhang, Y. et al. SUV39H1 is a novel biomarker targeting oxidative phosphorylation in hepatitis B virus-associated hepatocellular carcinoma. BMC Cancer. 23 https://doi.org/10.1186/s12885-023-11633-4 (2023).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48, W509–W514. https://doi.org/10.1093/nar/gkaa407 (2020).
Ballhausen, A. et al. The shared frameshift mutation landscape of microsatellite-unstable cancers suggests immunoediting during tumor evolution. Nat. Commun. 11, 4740. https://doi.org/10.1038/s41467-020-18514-5 (2020).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905. https://doi.org/10.1038/nature08822 (2010).
Malta, T. M. et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173, 338–354e315. https://doi.org/10.1016/j.cell.2018.03.034 (2018).
Wang, Q. et al. NCAPG2 could be an immunological and prognostic biomarker: From pan-cancer analysis to pancreatic cancer validation. Front. Immunol. 14, https://doi.org/10.3389/fimmu.2023.1097403 (2023).
Maeser, D., Gruener, R. F. & Huang, R. S. oncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief. Bioinform 22, https://doi.org/10.1093/bib/bbab260 (2021).
Yang, W. et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955-961. https://doi.org/10.1093/nar/gks1111 (2013).
JG, T. et al. COSMIC: the catalogue of somatic mutations in cancer. 47, D941–D947, (2019). https://doi.org/10.1093/nar/gky1015
Zhang, Y. et al. Tumor stemness score to estimate epithelial-to-mesenchymal transition (EMT) and cancer stem cells (CSCs) characterization and to predict the prognosis and immunotherapy response in bladder urothelial carcinoma. Stem Cell Res. Ther. 14, 15. https://doi.org/10.1186/s13287-023-03239-1 (2023).
Allemani, C. et al. Global surveillance of trends in cancer survival 2000-14 (CONCORD-3): analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries. Lancet 391, 1023–1075. https://doi.org/10.1016/s0140-6736(17)33326-3 (2018).
Shigematsu, S. et al. ZNF689 suppresses apoptosis of hepatocellular carcinoma cells through the down-regulation of Bcl-2 family members. Exp. Cell. Res. 317, 1851–1859. https://doi.org/10.1016/j.yexcr.2011.05.012 (2011).
Yi, P. S., Wu, B., Deng, D. W., Zhang, G. N. & Li, J. S. Positive expression of ZNF689 indicates poor prognosis of hepatocellular carcinoma. Oncol. Lett. 16, 5122–5130. https://doi.org/10.3892/ol.2018.9295 (2018).
Kim, H. et al. ZNF746/PARIS promotes the occurrence of hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 563, 98–104. https://doi.org/10.1016/j.bbrc.2021.05.051 (2021).
Zhang, T. et al. Overexpression of zinc finger protein 687 enhances tumorigenic capability and promotes recurrence of hepatocellular carcinoma. Oncogenesis 6, e363. https://doi.org/10.1038/oncsis.2017.63 (2017).
Xie, W. et al. Knockdown of ZNF233 suppresses hepatocellular carcinoma cell proliferation and tumorigenesis. Gene 679, 179–185. https://doi.org/10.1016/j.gene.2018.08.070 (2018).
Feng, Z. et al. ZNF143 expression is associated with COPD and tumor microenvironment in Non-Small cell lung cancer. Int. J. Chron. Obstruct Pulmon Dis. 17, 685–700. https://doi.org/10.2147/copd.S352392 (2022).
Yang, Y. et al. The pivotal role of ZNF384: driving the malignant behavior of serous ovarian cancer cells via the LIN28B/UBD axis. Cell. Biol. Toxicol. 40, 100. https://doi.org/10.1007/s10565-024-09938-6 (2024).
Ma, S., Meng, Z., Chen, R. & Guan, K. L. The Hippo pathway: biology and pathophysiology. Annu. Rev. Biochem. 88, 577–604. https://doi.org/10.1146/annurev-biochem-013118-111829 (2019).
Zanconato, F., Cordenonsi, M. & Piccolo, S. YAP/TAZ at the roots of cancer. Cancer Cell. 29, 783–803. https://doi.org/10.1016/j.ccell.2016.05.005 (2016).
Zhang, S. & Zhou, D. Role of the transcriptional coactivators YAP/TAZ in liver cancer. Curr. Opin. Cell. Biol. 61, 64–71. https://doi.org/10.1016/j.ceb.2019.07.006 (2019).
Niculescu, A. B. et al. Effects of p21(Cip1/Waf1) at both the G1/S and the G2/M cell cycle transitions: pRb is a critical determinant in blocking DNA replication and in preventing endoreduplication. Mol. Cell. Biol. 18, 629–643. https://doi.org/10.1128/mcb.18.1.629 (1998).
Bhutia, S. K., Mukhopadhyay, S., Sinha, N., Das, D. N. & Fisher, P. B. Autophagy: cancer’s friend or foe? Adv. Cancer Res. 118, 61–95 (2013).
Ye, J. et al. Targeting autophagy and beyond: deconvoluting the complexity of Beclin-1 from biological function to cancer therapy. Acta Pharm. Sin B. 13, 4688–4714. https://doi.org/10.1016/j.apsb.2023.08.008 (2023).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147. https://doi.org/10.1038/s41586-022-05400-x (2022).
Wang, Y. F. et al. Spatial maps of hepatocellular carcinoma transcriptomes reveal Spatial expression patterns in tumor immune microenvironment. Theranostics 12, 4163–4180. https://doi.org/10.7150/thno.71873 (2022).
Xu, C. et al. SIRPγ-expressing cancer stem-like cells promote immune escape of lung cancer via Hippo signaling. J. Clin. Invest. 132 https://doi.org/10.1172/jci141797 (2022).
Zhou, X. et al. The Hippo-YAP signaling pathway drives CD24-mediated immune evasion in esophageal squamous cell carcinoma via macrophage phagocytosis. Oncogene 43, 495–510. https://doi.org/10.1038/s41388-023-02923-z (2024).
Ma, T., Xiong, E. S., Lardelli, R. M. & Lykke-Andersen, J. Sm complex assembly and 5’ cap trimethylation promote selective processing of SnRNAs by the 3’ exonuclease TOE1. Proc. Natl. Acad. Sci. U S A. 121, e2315259121. https://doi.org/10.1073/pnas.2315259121 (2024).
Deng, T. et al. TOE1 acts as a 3’ exonuclease for telomerase RNA and regulates telomere maintenance. Nucleic Acids Res. 47, 391–405. https://doi.org/10.1093/nar/gky1019 (2019).
Rehrauer, H. et al. How asbestos drives the tissue towards tumors: YAP activation, macrophage and mesothelial precursor recruitment, RNA editing, and somatic mutations. Oncogene 37, 2645–2659. https://doi.org/10.1038/s41388-018-0153-z (2018).
Feng, Y. et al. The signaling protein Wnt5a promotes TGFβ1-mediated macrophage polarization and kidney fibrosis by inducing the transcriptional regulators Yap/Taz. J. Biol. Chem. 293, 19290–19302. https://doi.org/10.1074/jbc.RA118.005457 (2018).
Guo, X. et al. Single tumor-initiating cells evade immune clearance by recruiting type II macrophages. Genes Dev. 31, 247–259. https://doi.org/10.1101/gad.294348.116 (2017).
Funding
This research was funded by the National Natural Science Foundation of China, grant nos. 82100321, 82370353, the Shenzhen Natural Science Foundation Basic Research Surface Project (JCYJ20220530-144811027), and the Shenzhen Science and Technology Program (JCYJ2024081315-0437049).
Author information
Authors and Affiliations
Contributions
Jie Yu Ao: Conceived the study, designed experiments, performed functional assays (cell culture, qPCR, Western blot), analyzed data, and wrote the initial draft. Yi Cai: Conducted bioinformatics analyses, performed functional assays, validated gene expression data, and contributed to the manuscript revision. Si Ting Xu: Performed immunohistochemistry and tissue microarray analysis, interpreted IHC results, and assisted in manuscript preparation. Bo Yu Yang: Carried out RNA-seq data processing, functional enrichment analysis, and validation of differentially expressed genes. Xian Sun: Provided clinical samples, and supervised the entire study. Yan Shi: Supervised the entire study, designed the research framework, secured funding, and critically revised the manuscript for intellectual content.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethnic statement
The clinical trial was sanctioned by the Ethics Committee of School of Medicine of Sun Yat-sen University (Approval Number: KY-2025-366-01). This work is guided by the Declaration of Helsinki. Participants gave their written consent after being fully informed.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ao, J.Y., Cai, Y., Xu, S.T. et al. Transcriptomic analysis reveals the potential role of TOE1 in hepatocellular carcinoma. Sci Rep 16, 69 (2026). https://doi.org/10.1038/s41598-025-29408-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-29408-1