Abstract
Gestational Diabetes Mellitus (GDM) is an emerging maternal health problem with increasing incidences. The lack of complete understanding of its pathophysiological mechanisms and novel regulatory biomarkers makes early diagnosis difficult. High-throughput RNA sequencing and computational bioinformatics analyses were conducted to identify novel hub genes, and their regulatory mechanisms were validated through qRT-PCR, western blot, and siRNA-mediated knockdown studies. Intermediate metabolites and circulatory levels of amino acids in the serum of GDM patients and healthy controls were measured. Transcriptomic studies identified SNW1 as the most sensitive and specific biomarker, significantly up-regulated in GDM (fold change = 1.09; p < 0.001). Metabolomic studies indicated significantly elevated gluconeogenesis in GDM, evidenced by decreased levels of alanine and increased levels of pyruvate and glucose compared to controls. siRNA-mediated knockdown of SNW1 in PANC1 cells resulted in significant down-regulation of alanine aminotransferase (ALT/GPT) and insulin receptor substrate (IRS1), while glucose transporters (GLUT2/GLUT4) and insulin (INS) were significantly up-regulated at both mRNA and protein levels. This study identified SNW1 as a novel insulin-resistant gene that induces hyperglycemia by elevating gluconeogenesis and decreasing glucose uptake. SNW1 may be considered a potential therapeutic target with clinical utility for the management of GDM.
Similar content being viewed by others
Introduction
Gestational Diabetes mellitus (GDM) is often associated with maternal and fetal complications with increased risk for future metabolic diseases in both mother and child. In recent years, the incidence of GDM has increased dramatically worldwide, as approximately 15 to 20% of pregnancies are affected by GDM yearly1. In Southeast Asia, a higher prevalence of GDM with an average incidence of 10.1% has been reported2,3. Indian Diabetes Federation (IDF) has also estimated that around 4 million women with GDM are being reported yearly4. Despite being a significant cause of fetal morbidity and mortality, gestational diabetes mellitus (GDM) often goes undiagnosed until 24–28 weeks of pregnancy. The delay is due to the reliance on the oral glucose tolerance test (OGTT), the gold standard method for diagnosing GDM. This test measures blood glucose levels, which reflect the final consequence of GDM and appear only after the condition has significantly progressed5. Furthermore, OGTT is cumbersome and non-compliant to the patient as its invasive, require fasting for 8–12 h and is often associated with nausea and vomiting6. Hence, there is utmost urgency for sensitive and specific biomarkers that may offer early diagnosis so that timely interventions may be implemented to improve pregnancy outcomes.
GDM is a multifactorial disease in which genes and environment regulate the onset and progression of its pathophysiology. However, its complete mechanism remains obscure. Various studies have reported that the failure of pancreatic β cells to compensate for the increasing insulin demand followed by chronic insulin resistance are critical components of the pathophysiology of GDM7,8,9. Insulin resistance has long been known to play an important role in hyperglycemia10. Although, mechanism of insulin resistance is known, several insulin resistant genes and their possible mechanism are still behind curtain11. Hence, it is essential to identify novel insulin-resistant genes and understand their molecular mechanisms. This will provide a deeper insight into the pathophysiology of GDM. Such knowledge is crucial for developing strategies for early diagnosis, prevention, and improved management of the condition.
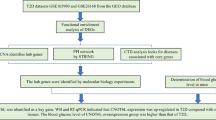
In this “era of omics”, the high throughput sequencing and metabolomics, combined with computational and statistical approach, has played a significant role in discovering novel regulators and biomarkers related to particular diseases12. In the present study, we employed RNA sequencing and NMR-based serum metabolomics analysis to identify novel regulatory and diagnostic biomarkers. These biomarkers aim to provide a detailed understanding of the pathophysiological mechanisms underlying GDM. This approach also holds potential for the early detection, timely prevention, and effective management of the condition. Networking, module extraction and functional analysis was applied to reveal the hub genes which were further validated in a large sample size and assessed for their sensitivity and specificity in predicting GDM. Further based on MCODE score, fold change, sensitivity and specificity and functional analysis potential diagnostic biomarker was selected. To verify whether this identified biomarker corresponds to the insulin-resistant gene, serum metabolomics and in-vitro validation were conducted. Recent studies have primarily focused on well-established insulin-resistant genes like PPARG, TCF7L2, and IRS1 in glucose metabolism and insulin signaling (Li et al., 2022). In contrast, our study uniquely identifies SNW1 as a novel biomarker involved in gestational diabetes mellitus (GDM), demonstrating its dual role in hyperglycemia regulation through transcriptomics and metabolomics analysis.
Results
Patient data
The present study is an integration of Transcriptomic and metabolomics where each study has 3 groups (NGT/NC, Pre-GDM and Post-GDM). The complete study design is shown in electronic supplementary material (ESM, Figure S1). The clinical and demographic details of patients and subjects involved in the Transcriptomic study are listed in Table 1. Clearly evident that the Clinical and biochemical parameters of NGT and GDM patients are significantly different, the GDM group has increased FPG and HbA1c levels when compared to NGT (Table 1).
The clinical and demographic details of patients and subjects involved in the metabolomics study are listed in Table 2.
Identification of differentially expressed genes (DEGs) and novel targets
Comparison of NGT (BC4) group with “Pre-term GDM” group (BC5 and BC7) yielded a total of 943 DEGs while comparison of NGT (BC4) group with “Post-term GDM” group (BC6) results in 470 DEGs. DEGs in both the GDM groups (i.e. Pre GDM group and Post GDM group) have been visualized through volcano plot and heat map (Fig. 1A and B) ; for high resolution image of Fig. 1A see ESM Fig. S2). After extracting the control (NGT/BC4) data from both GDM groups, we compared both groups and found 84 common elements in both, which was excluded from further analysis. After exclusion, 859 DEGs were found to be exclusively related to the “Pre-term GDM” group, while 386 DEGs were found unique to the “Post-term GDM” group (Fig. 1C). Further downstream processing has been performed for those 859 DEGs related to the Pre GDM group. The significant biological processes, molecular functions, cellular components and metabolic pathways identified for DEGs are shown in the ESM, Fig. S3 and S4. Among these DEGs, 311 were up-regulated while 548 were down-regulated. All the enriched DEGs have a false discovery rate of 0.1% with p value < 0.05 and logFC>+-1. We have not considered the post GDM group because blood sugar level was restored to normal level in those GDM women after the term.
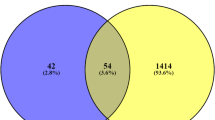
Next, we were interested to know the novel DEGs which have not been reported yet in the context of GDM. To achieve this, DEGs list were compared with the gene list obtained from the OMIM (https://www.omim.org/)13 and Gene Cards (https://www.genecards.org/)14. Comparison reveals total 258 novel up-regulated DEGs while 434 down-regulated DEGs (Fig. 1D).
DEGs Screening and Visualization: DEGs identification and screening among GDM pre-term (BC5, BC7), post-term (BC6), and NGT (BC4) group identified total of 859 DEGs which are being visualized by (A) volcano plot (B) Heat map and (C).ven diagram. (D). Further comparison of DEGs with gene list obtained from Gene cards and OMIM revealed a total of 692 DEGs out of 859 DEGs as shown in ven diagram. Of this 692 DEGs, 434 DEGs are down-regulated while 258 are up-regulated DEGs.
PPI network and module extraction analysis identified thirteen hub genes
The PPI network of DEGs with a combined score > 0.9 was constructed and visualized. The network consists of 215 nodes and 545 edges (Fig. 2, Zoomed image in ESM Fig. S5) The top four significant modules and networks of hub genes were identified using the MCODE plugin of Cytoscape software (Fig. 2A-D). We identified thirteen novel genes as hub genes with degrees ≥ 6. Among these genes, SNAPC5, GNG7, AP1M1, PPIL6, CSTF1, and SNW1 are up-regulated, while ATP6V1D, AP1G1, GTF2A2, POLR3K, NUP160, DHX38, and RAB5C are down-regulated in GDM samples.
Network Construction and module extraction: Protein-protein interaction network of identified DEGs were constructed using STRING and Cytoscape. Most significant modules were extracted from the main network using MCODE plugin of the cytoscape. Module extraction led to the identification of four most significant modules. Red indicates up-regulated DEG while blue indicate down-regulated DEG.
SNW1 as novel and most potent biomarker
We selected thirteen novel hub genes based on the networking and module creation (SNAPC5, GNG7, AP1M1, PPIL6, CSTF1, SNW1, ATP6V1D, AP1G1, GTF2A2, POLR3K, NUP160, DHX38, and RAB5C) for their validation through qRT-PCR analysis. The mRNA expression of novel hub genes was compared in the blood samples of NGT and GDM patients. The results revealed similar variation trends of DEGs expression profile to the RNA-Seq samples. Among the thirteen hub genes identified, SNW1 has highest fold change expression (fold change = 3; p = 0.01) (Fig. 3A). Furthermore, to evaluate the diagnostic potential of these biomarkers, we performed receiver operating characteristic curve (ROC) analysis. Out of six up-regulated hub genes, SNW1 have highest sensitivity and specificity (AUC = 0.988, sensitivity and specificity = 96.7) (Fig. 3B). MCODE analysis have revealed highest score for SNW1 further making it a potent target (Fig. 3C). Functional analysis has revealed their possible role in insulin signalling, amino acid metabolism and gluconeogenetic pathway (Fig. 3D). Hence, based on all these four parameters namely expression value, diagnostic potential, MCODE score and functional analysis SNW1 was identified as most potential biomarker.
Biomarker Selection: (A) Identified hub genes were validated using RT-qPCR. Fold change analysis revealed higher expression of SNW1. (B) ROC plot analysis shows higher sensitivity and specificity (96.7%) with higher area under curve (AUC 0.988). Genes highlighted in red colour are upregulated whereas genes highlighted in blue colour are downregulated. (C) MCODE analysis revealed the highest MCODE score (12.75) for SNW1 among all Hub genes. (D) Functional enrichment analysis revealed a possible role in insulin and glucose metabolism.
Gluconeogenesis is upregulated in GDM
The functional analysis has revealed the role of SNW1 in glucose metabolism and gluconeogenesis. In order to investigate the status of gluconeogenetic pathway in GDM patient, the concentration levels of metabolites involved in gluconeogenesis were evaluated. The NMR spectral measurement followed by multivariate ANOVA analysis (see ESM, Table S2 and Fig. S6) revealed the comparatively lower level of amino acids (glycine, alanine, valine etc.) in pre term-GDM (PrT) as compared to normal control (NC) and post-term GDM (PoT) while the level of glucose and pyruvate was found to be higher in GDM compared to the control (Fig. 4). Similarly, the level of other metabolites like lactate and succinate was found to be lower in GDM patient compared to normal control (Fig. 4). Ratiometric analysis revealed the lower level of alanine-to-glucose ratio (AGR), succinate-to-pyruvate ratio (SPR) and succinate-to -glucose ratio (SGR) in pre term-GDM (PrT) compared to normal control (NC) and post-term GDM (PoT) patients (Fig. 5).
Serum metabolite profiling: The representative graph compares the Circulatory level of metabolites in NC (n = 49), PrT (n = 32) and PoT (n = 18).
Ratio-metric analysis: The representative shows alteration in circulatory metabolic ratio (A) Alanine-to-Glucose (AGR), (B) Succinate-to-Pyruvate (SPR), (C) Succinate-to-Glucose are significantly decreased in PrT and PoT when compared to NC.
The major precursors for gluconeogenesis are amino acids and lactate while glucose is the final end product. Pyruvate, being the key metabolite in gluconeogenetic as well as kerb’s cycle, may be utilised either for glucose production or consumed in Kreb’s cycle for ATP production (Fig. 6A and B) (Supplementary Fig. 1). NMR analysis revealed the lower level of different amino acids and lactate (precursor of gluconeogenesis) while the higher level of glucose (final product of gluconeogenesis) indicating the elevated gluconeogenesis in GDM patients when compared to normal control. Further, the lower alanine-to-glucose ratio (AGR) in GDM patient confirms the elevated gluconeogenesis. On the other hand, lower level of succinate while higher level of pyruvate and glucose indicate the lower consumption of pyruvate for ATP production. Overall, the result indicates that the gluconeogenesis is elevated in GDM patient compared to normal control.
Summary and integration of omics study: (A) From metabolomics study, we found lower level of amino acids like alanine, lactate and succinate while higher level of glucose and pyruvate in GDM patients compared to normal pregnant women. The higher level of glucose and pyruvate while lower level of alanine, lactate and succinate indicates elevated glucose production and diminished glucose utilization. (B) Integration of transcriptome and metabolome revealed SNW1, a novel diagnostic biomarker identified from transcriptome, as a regulator of gluconeogenesis. Protein-protein interaction network revealed link between Key regulatory enzyme alanine aminotransferase (ALT/GPT) and SNW1. Alanine aminotransferase (ALT/GPT) converts alanine into pyruvate precursor of gluconeogenesis. (C) Pearson correlation analysis performed for identifying the serum metabolic profiles correlated with alanine-to-glucose ratio and (D) Spearman rank correlation analysis performed for identifying the serum metabolic profiles correlated with transcriptomic marker feature SNW1. The vertical bar graphs illustrate positive correlations (highlighted in pale pink) and negative correlations (highlighted in blue). Statistically significant correlations are marked with asterisks: ****, ***, ** and * for p-value < 0.0001, p-value < 0.001, p-value < 0.01, and p-value < 0.05.
The biological relevance of the transcriptomic and metabolic alterations and their association was evaluated using statistical correlation analysis methods. First, we performed the pearson correlation analysis for identifying the serum metabolic profiles correlated with alanine-to-glucose ratio (AGR, Fig. 11A). Circulatory AGR demonstrated a strong positive association with metabolites linked to energy metabolism (e.g., Lactate, Pyruvate) and amino acid metabolism (e.g., Alanine, Glycine, Isoleucine). Further, we used spearman rank correlation analysis for identifying the serum metabolic profiles correlated with transcriptomic marker feature SNW1 (Fig. 11B). The SNW1 appears positively correlated with metabolites involved in methylation and energy balance (e.g., Betaine, Choline) and negatively correlated with certain TCA cycle intermediates (e.g., Citrate) and energy substrates like Glucose (mM). The correlation analysis suggests that alterations in amino acid and energy metabolism are key features associated with both Alanine/Glucose and SNW1, potentially reflecting metabolic reprogramming in GDM patients.
SNW1 promotes hyperglycemia in GDM
During gluconeogenesis, alanine is converted into pyruvate by an enzyme called alanine-amino transferase or glutamine-pyruvate aminotransferase (ALT/GPT). Protein-protein interaction network analysis has revealed the direct link of SNW1 and ALT. Hence, in order to investigate the possible role of SNW1 in glucose metabolism, in-vitro validation assay was performed. Transfection efficiency was assured by qRT-PCR, SNW1 was down-regulated in SNW1 siRNA group compared to control siRNA and control group (Fig. 7A). To assess the effect of SNW1 knock-down on insulin and glucose metabolism pathway, the expression level of some biomarkers related to these pathways were evaluated at RNA as well as protein level. Downregulation of SNW1 via siRNA, leads to the downregulation of ALT and IRS1 (Fig. 7B, F) revealing the possible regulatory action of SNW1 in glucose and insulin metabolism. Further, SNW1 knockdown leads to the upregulation of insulin and GLUT2/4 (Fig. 7C, D, E). Hormone assay also revealed the lower level of glucose and GPT while higher level of GLUT4 and insulin upon SNW1 knockdown (Fig. 7G, H, I, J). Similarly, at protein level, expression of GLUT4 was up while that of IRS1 was down when SNW1 was knockdown (Fig. 8) (Supplementary Fig. 2). Thus, SNW1 is promoting hyperglycemic condition in GDM patients by enhancing glucose production via gluconeogenesis and inhibit peripheral glucose uptake by inhibiting the insulin production as well as glucose transporter.
In vitro validation assays: PANC1 cells were transfected with SNW1 siRNA and quantified for key genes related to glucose and insulin metabolism at RNA and protein level. (A-F): SNW1 knockdown causes downregulation of IRS1 and GPT while upregulation of INS and GLUT2/4. SNW1, being a transcription factor, promotes the expression of IRS1 and GPT while inhibit the expression of INS and GLUT2/4. (G-J) Hormone assays revealed low glucose uptake while high insulin level in SNW1 knockdown PANC1 cells compared to the control PANC1 cells, similarly, serum level of GPT was low while GLUT4 level was high in SNW1 knockdown PANC1 cells compared to the control PANC1 cells.
Western blot analysis: Downregulation of SNW1 inhibits the IRS1 while promote GLUT4 expression: Expression of GLUT4 was found to be upregulated while that of IRS1 downregulated when SNW1 was knock-down with SiRNA. Thus, SNW1 is inhibiting the glucose uptake by downregulating the expression of glucose transporter like GLUT4 while up-regulating the expression of IRS1.
Discussion
Gestational diabetes mellitus (GDM) has become a global women’s health problem with increasing incidences worldwide15. Moreover, GDM poses a threat to both mother and foetal health16. However, its complete pathophysiological mechanism is still unknown. Several molecular players related to insulin resistance are yet to be deciphered17,18,19. Hence, there is a great need to decipher the novel regulatory molecule and understand their pathophysiological mechanism which will help enhance foetal and maternal health.
In the present study, we performed the high-throughput RNA sequencing and metabolome profiling. RNA - sequencing of peripheral maternal blood from GDM and NGT women, lead to the identification of statistically significant DEGs (Log FC>+-1; p value < 0.05). Based on the MCODE score (12.7), log fold change value (1.5), and sensitivity and specificity (96.7%) score, SNW1 was identified to be the most potent novel biomarker. The functional analysis revealed its possible role in insulin metabolism. Further, literature search revealed SNW1 as a transcriptional regulator suspecting it as an insulin resistant gene20,21,22,23. For being an insulin resistant gene, it must inhibit glucose uptake and promote endogenous glucose production via gluconeogenesis10,11,24,25,26. The elevated gluconeogenesis in GDM patients was confirmed through metabolomics. Metabolomics study identified higher level of glucose (end product of gluconeogenesis) and pyruvate (intermediate of gluconeogenesis) whiles the lower level of amino acids and lactate (precursors of gluconeogenesis) in GDM patients. Protein-protein interaction network identified the possible link between SNW1 and GPT/ALT (enzyme which converts alanine into pyruvate). Finally, in-vitro assays were carried to confirm the regulatory action of SNW1. The siRNA mediated knockdown of SNW1, revealed that SNW1 was inhibiting glucose uptake by downregulating GLUT2/4 and insulin while promoting gluconeogenesis by upregulating the ALT/GPT enzyme at mRNA as well as protein level.
GDM is characterised by insulin resistance, however, several of the insulin resistant genes are not deciphered yet27. From our RNA-seq study we identified SNW1 as most potent biomarker involved in insulin metabolism, however, studies on the role of SNW1 in hyperglycemic condition, especially GDM, is lacking. SNW domain-containing protein 1 (SNW1), a member of the SNW gene family, is involved in splicing and transcription. SNW1 is a transcriptional co-activator as well as co-repressor. It has been shown to be the transcriptional regulator of the NF-κB pathway, which has been implicated in various biological processes including insulin resistance28,29,30. Besides, SNW1 has also been shown to regulate the Notch and BMP signaling pathway31,32,33,34, which is associated with hyperglycemic condition. However, direct evidence indicating the SNW1 as an insulin resistant gene is lacking. Our results identified the SNW1 as novel insulin resistant genes and the regulator of hyperglycemia in GDM patients. SNW1 with higher sensitivity and specificity was found to be 3 fold higher in GDM patients compared to normal mothers. Knockdown of SNW1 via siRNA in PANC1 cells, leads to the upregulation of insulin and glucose transporters like GLUT2/4 at mRNA as well as protein level. Similarly, siRNA knockdown of SNW1 causes lower glucose and higher insulin level indicating that higher expression of SNW1 inhibits glucose uptake. Further, the SNW1 causes the up regulation of ALT/GPT at mRNA as well as protein level. ALT enzyme is involved in the gluconeogenesis. Elevated ALT level in response to higher expression of SNW1 leads to the elevated endogenous glucose production. Thus, higher expression of SNW1 inhibits glucose uptake as well as elevates the endogenous glucose production, thereby causing hyperglycemic condition (Fig. 9). These both features i.e. inhibition of glucose uptake and endogenous glucose production, makes SNW1, a novel insulin resistant gene.
Diagrammatic representation of SNW1 mediated hyperglycemia in GDM Patient. SNW1 promotes gluconeogenesis by up-regulating the expression of ALT while inhibit glucose uptake by down-regulating the expression of glucose transporter like GLUT2/4.
Previous studies have primarily focused on well-known insulin-resistant genes such as PPARG, TCF7L2, and IRS1, which are implicated in glucose metabolism and insulin signalling (Li et al., 2022; Su et al., 2023)25,35. Unlike these studies, our research uniquely identifies SNW1 as a novel biomarker involved in the pathophysiology of gestational diabetes mellitus (GDM). Through the integration of transcriptomics and metabolomics data, we demonstrate SNW1’s dual role in hyperglycemia: inhibiting glucose uptake via downregulation of GLUT2/4 and promoting gluconeogenesis through upregulation of ALT/GPT. This combined functional mechanism, supported by in-vitro knockdown assays, distinguishes SNW1 from previously studied insulin-resistant genes. Moreover, while earlier research linked SNW1 to transcriptional regulation in other biological pathways, such as NF-κB and Notch signaling (Verma et al., 2019; Hong et al., 2019)20,21, our study is the first to highlight its direct role in hyperglycemia and GDM, paving the way for novel therapeutic targets.
Concluding remarks
In conclusion, the present study explores the transcriptomics analysis in combination with serum metabolomics analysis for identifying novel hub genes and regulatory biomarkers for early diagnosis and management of gestational diabetes mellitus (GDM). The study’s methodology involved collection of blood plasma and serum samples, RNA-sequencing, and serum metabolomics analysis, with a focus on the identification of differentially expressed genes (DEGs) and their impact on metabolic pathway alterations. The study revealed the regulatory role of the insulin-resistant SNW1 gene in glucose metabolism and gluconeogenesis. SNW1 is novel insulin resistant genes which inhibits the glucose uptake by downregulating the insulin and glucose transporter while increases the endogenous glucose production via elevating the alanine aminotransferase enzyme, a key enzyme in the gluconeogenesis. In vitro validation assays confirmed the regulatory action of SNW1, demonstrating its role in inhibiting glucose uptake and promoting endogenous glucose production. Further, the integration of transcriptomic and metabolomic data revealed a link between SNW1 and the altered levels of metabolites involved in gluconeogenesis, providing insights into the molecular mechanisms underlying hyperglycemia in GDM and also proposes SNW1 as a potential therapeutic target for the management of GDM. Taken together, the study identified SNW1 as a sensitive and specific biomarker significantly up-regulated in GDM, with a potential role in inducing hyperglycemia by elevating gluconeogenesis and decreasing glucose uptake.
To be mentioned here is that the present study suffers from a limitation of low sample size taken for RNA-sequencing, but rigorous bioinformatics analysis combined with a large sample size in validation cohort minimizes the variability in datasets. Nevertheless, further validation studies are required to elucidate the role of SNW1 in causing hyperglycaemia.
Materials and methods
Sample collection and grouping
The study was approved by the Institutional Ethics Committee of the Banaras Hindu University (F.Sc./2018-19/human ethics/0008). This study was carried out following the required local ethical recommendations (Banaras Hindu University). All procedures followed were following the ethical standards of the responsible committee on human experimentation. All methods were conducted in compliance with relevant guidelines and regulations. Informed consent was obtained from all patients for being included in the study.
GDM and healthy pregnant women (NGT) were attended at Gynaecological out-patient ward of Sir Sundar Lal hospital, Institute of Medical Science, Banaras Hindu University. Single-step OGTT was performed, and blood sugar level ≥ 140 mg/dl was taken as the cutoff value (As per WHO) for GDM diagnosis. The 5 ml of peripheral blood of age-matched GDM and NGT subjects were collected in an EDTA vial (for transcriptomics) and plain vial (for serum metabolomics) with the help of a sterile syringe. Informed consent was obtained from participating subjects.
For transcriptomics analysis, two separate cohorts of patients namely the sequencing cohort (N = 15) and the validation cohort (N = 60) were formed. In sequencing cohort, blood from healthy pregnant women (NGT, n = 3), GDM patients before term [Pre-term GDM (24–36 gestational weeks), N = 6], and GDM patients after term [Post-term GDM (GDM mother after delivery), N = 6] was collected. For NMR based clinical metabolomics analysis, the serum was collected from NGT (N = 49), pre-term GDM (N = 32) and post-term GDM (N = 18) patients. For validation, 30 cases and 30 controls were taken.
According to the manufacturer’s instructions, total RNA was extracted using TRIzol® Reagent Kit (Thermo Fisher Scientific, USA). RNA was quantified by measuring absorbance at 260 and 280 nm using NanoDrop™ One/OneCMicrovolume UV-Vis Spectrophotometer. In addition, the integrity of 28s and 18s ribosomal RNA was assessed on 1% agarose gel. For serum isolation, blood collected in red top collections tubes were left undisturbed for 60 min (at Room Temperature) for proper clotting. Clot was removed by centrifugation at 1000–1500 g for 10 min in cooling centrifuge. Subsequently, serum was isolated and stored at -80 °C with appropriate labelling until the acquisition of NMR data.
Transcriptomics analysis
RNA-Sequencing and DEG screening
RNA-sequencing was performed on portable nanopore sequencing platform (Oxford Nanopore-MinION). Single strand cDNA synthesis was performed using Agilent Sure select kit. Raw data quality was assessed, and gene expression was measured in units of fragments per kilobase of exon model per million mapped reads (FPKM). The library’s identification of DEGs was performed using the Bioconductor package edgeR (https://bioconductor.org/packages/release/bioc/html/edgeR.html). The p-value and false discovery rate (FDR) was corrected using the Benjamini-Hochberg (BH) procedure36,37. Fold Change > 2 or FoldChange < 0.5 and FDR value < 0.05 were set as the threshold for selecting statistically significant regulated differentially expressed genes.
Gene network construction, module extraction and functional analysis
STRING v.11.538 and Cytoscape v.3.8.239 was used to get protein-protein interaction network with combined score (CS) > 0.9 as set criteria. Molecular Complex Detection (MCODE) plugin of Cytoscape software was applied to analyze the different modules of the PPI network40,41. The criteria for the MCODE analysis comprised: degree cut-off = 2, MCODE scores > 5, max depth = 100, k-score = 2, and node score cut-off = 0.2. The hub genes were excavated for degrees ≥ 13.
In order to predict the function and pathways, the DEGs enriched in modules were subjected to gene ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) metabolic pathway mapping through DAVID v 6.842. The statistically significant (p < 0.05) GO terms were plotted against –log10 of p-value.
Validation of hub genes
According to the manufacturer’s instructions, RNA was isolated from the validation cohort using TRIzol® Reagent Kit (Thermo Fisher Scientific, USA) and reverse transcribed into single-stranded cDNA using Revert Aid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, USA). A 20ul PCR reaction was prepared according to GCC Biotech qPCR SYBR Green Master Mix and the qPCR reaction was performed using Quant Studio 5 system (Applied biosystem, Thermo Fisher Scientific, USA). The 18s rRNA was used as a housekeeping gene to normalize expression value. All the reactions were run in duplicate using primer sequences listed in electronic supplementary material (ESM, Table S1). The dCt and 2^-ddCt method was used to analyze the expression level. All primer details are listed in Table S1(in supplementary). Receiver operating characteristic (ROC) curves were plotted by MedCalc® 20.0 (MedCalc Software Ltd, Ostend, Belgium); the area under the curve (AUC) values, as well as sensitivity and specificity values, were also calculated respectively.
NMR based serum metabolomics analysis
Preserved serum samples were thawed and prepared by mixing 300 µL serum with equal volume of sodium phosphate buffer (0.9% saline, buffer strength 50 mM prepared in 100% D2O, pH 7.4) followed by centrifugation at 16,278 g for 5 min. The NMR based serum metabolic profiling is performed as described previously from our lab43,44,45. Briefly, the prepared serum samples was transferred to a 5 mm NMR tube (Wilmad Glass, USA) along with a coaxial NMR tube containing 1.0 mM TSP (sodium salt of 3-trimethylsilyl-(2,2,3,3)-propionic acid-d4; Sigma–Aldrich (St. Louis, MO, USA)) dissolved in D2O (Sigma–Aldrich, St. Louis, MO, USA) was inserted separately that served as an external reference (offering a final apparent concentration approximately equal to 0.1 mM). Serum metabolic profiles were measured using the one dimensional (1D)1H NMR spectra recorded on high-field 800 MHz NMR spectrometer (Bruker Avance III) equipped with a TCI cryogenic probe. The broad signals of protein and other macromolecules was suppressed using Bruker standard CPMG (Carr–Purcell–Maiboom–Gill) spin–echo pulse sequence with water pre-saturation and a T2 filter time of ~ 60 ms. The NMR spectra were manually phased and corrected for baseline using TopSpin 3.6.1 and PROCESSOR module of CHENOMX NMR Suite 8.6 software spectrum and finally calibrated for chemical shift with respect to the formate peak at δ8.43 ppm. Formate signal was also considered as an internal reference for concentration profiling (concentration was set to 10 µM). The NMR spectral after data processing, the spectrum was imported to the PROFILER Module of CHENOMX and the concentrations of metabolites were estimated in all the serum samples of GDM patients and NGT subjects.
Association between transcriptomics and metabolomics data
Literature search and protein-protein interaction network was used to establish link between identified novel biomarkers and the concentration of identified metabolites. PPI network showing link between transcriptomics and metabolomics has been provided in supplementary Fig. 1. Further, Statistical correlation analysis was performed to evaluate the relationships between the metabolic features and specific variables of interest, namely Alanine/Glucose and SNW1. Pearson Correlation method quantified linear relationships between the metabolic features and Alanine/Glucose. Whereas the non-parametric Spearman Rank Correlation approach was used to assess relationships between the metabolic features and SNW1.
Transfection and hormone assays
PANC1 cell line were cultured in DMEM (Sigma-Aldrich) supplemented with 10% fetal bovine serum (Sigma-Aldrich), 100 µg/mL streptomycin and 100 IU/mL penicillin 227 (Gibco, Grand Island, NY, USA) at 37 °C with 5% CO2 humidified atmosphere. Cells were cultured in 24-well format for 24 h in serum and antibiotic free medium prior to transfection. Biomarker specific siRNA and control siRNA were transfected in 24-well culture plate using Lipofectamine 3000 transfection reagent (Thermo Fisher). Cells and consumed media were harvested post 48 h of transfection. Retrieved media were used for hormone assays. RNA was isolated followed with cDNA synthesis and quantitative PCR for genes like insulin (INS), insulin receptor (INSR), glucose transporter (GLUT2/4, SGLT1), Insulin receptor substrate (IRS1/2), alanine amino-transferase (ALT/GPT) involved in insulin and glucose metabolism pathway.
Western blot analysis
Cells were cultured in 6 well plate, transfected with SNW1 siRNA and then collected by scraping. The cell pellets were washed once with PBS, and then lysed in RIPA buffer, 1 µl of lysate was used for determination of protein concentration using a colorimetric bicinchoninic acid assay (Pierce, Rockford, IL). The remaining sample was added an equal volume of 2× SDS-polyacrylamide gel electrophoresis sample loading buffer (62.5 mM Tris-HCl, pH 6.8, 4% SDS, 5% β-mercaptoethanol, 20% glycerol, 0.1% bromophenol blue), followed by incubation in boiling water for 5 min. Equivalent amounts of proteins were resolved in SDS-polyacrylamide gels, and then transferred to nitrocellulose membranes using a semidry transfer cell. The transblotted membrane was washed twice with Tris-buffered saline containing 0.1% Tween 20 (TBST). After blocking with TBST containing 5% nonfat milk for 40 min, the membrane was incubated with the appropriate primary antibody (1:1000) in TBST-1% nonfat milk at 4 °C overnight. The membrane was then washed three times with TBST for a total of 15 min, followed by incubation with goat anti-rabbit or anti-mouse IgG-horseradish peroxidase conjugates (1:2000) for 1 h at room temperature and four washes with TBST for a total of 1 h. The immunoblots were visualized by enhanced chemiluminescence.
Statistical analysis
All statistical analysis was performed using Graph Pad Prism 9 (Graph Pad Software, Inc., La Jolla, CA, USA). Unpaired t-test (parametric) and one-way ANOVA (analysis of variance) was applied to compare between two and more than two groups respectively. A p-value < 0.05 was considered statistically significant.
Data availability
All the RNA-seq data were deposited in the SRA database of the National Center for Biotechnology Information (NCBI) database under the BioProject accession number PRJNA735083. The NMR spectral Data has been deposited to ZENODO (ID: 6997180 and DOI:10.5281/zenodo.6997180). The datasets used and /or analyzed in this study are available on request as well. Dr. Pawan K. Dubey can be contacted for RNA-seq data while Dr. Dinesh Kumar can be contacted for spectral data.
Abbreviations
- GDM:
-
Gestational diabetes mellitus
- SNW1:
-
SNW domain-containing protein 1
- ALT/GPT:
-
Alanine aminotransferase
- IRS1:
-
Insulin receptor substrate
- GLUT2/ GLUT4:
-
Glucose transporters
- INS:
-
Insulin
- IDF:
-
Indian diabetes federation
- OGTT:
-
Oral glucose tolerance test
- NGT:
-
Healthy pregnant women
- EDTA:
-
Ethylenediaminetetraacetic acid
- DEGs:
-
Differentially expressed genes
- RT:
-
Room temperature
- FPKM:
-
Fragments per kilobase of Exon model per million mapped reads
- BH:
-
Benjamini-Hochberg
- MCODE:
-
Molecular complex detection
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- CPMG:
-
Carr–Purcell–Meiboom–Gill
- ROC :
-
Receiver operating characteristic curve
- AUROC:
-
Area under ROC curve
- FDR:
-
False discovery rate
- CI:
-
Confidence interval
- CS:
-
Combined score
- TSP:
-
Tri-methyl silyl propionic acid-d4
- D2O:
-
Deuterium oxide
- TBST:
-
Tris-buffered saline containing 0.1% Tween 20
- PrT:
-
Pre term-GDM
- NC:
-
Normal control
- PoT:
-
Post-term GDM
- AGR:
-
Alanine-to-glucose ratio
- SPR:
-
Succinate-to-pyruvate ratio
- SGR:
-
Succinate-to-glucose ratio
- ESM:
-
Electronic supplementary material
References
Alduayji, M. M. & Selim, M. Risk factors of gestational diabetes Mellitus among women attending an Antenatal Care Clinic in Prince Sultan Military Medical City (PSMMC), Riyadh, Kingdom of Saudi Arabia: a case-control study. Cureus 15, e44200.
Kunasegaran, T., Balasubramaniam, V. R. M. T., Arasoo, V. J. T., Palanisamy, U. D. & Ramadas, A. Gestational diabetes Mellitus in Southeast Asia: a scoping review. Int. J. Environ. Res. Public. Health. 18, 1272 (2021).
Lee, K. W. et al. Prevalence and risk factors of gestational diabetes mellitus in Asia: a systematic review and meta-analysis. BMC Pregnancy Childbirth. 18, 494 (2018).
Mantri, N. et al. National and regional prevalence of gestational diabetes mellitus in India: a systematic review and Meta-analysis. BMC Public. Health. 24, 527 (2024).
Moon, J. H. & Jang, H. C. Gestational diabetes Mellitus: diagnostic approaches and maternal-offspring complications. Diabetes Metab. J. 46, 3–14 (2022).
Jagannathan, R. et al. The oral glucose tolerance test: 100 years later. Diabetes Metab. Syndr. Obes. 13, 3787–3805 (2020).
Li, L. M., Jiang, B. G. & Sun, L. L. Frontiers | HNF1A: from monogenic diabetes to type 2 diabetes and gestational diabetes Mellitus. https://doi.org/10.3389/fendo.2022.829565
Alejandro, E. U. et al. Gestational diabetes Mellitus: a harbinger of the vicious cycle of diabetes. Int. J. Mol. Sci. 21, 5003 (2020).
Lin, J. Y. & Yin, R. X. Exposure to endocrine-disrupting chemicals and type 2 diabetes Mellitus in later life. Exposure Health. 15, 199–229 (2022).
Zhao, X. et al. Frontiers | the crucial role and mechanism of insulin resistance in metabolic disease. https://doi.org/10.3389/fendo.2023.1149239
Petersen, M. C. & Shulman, G. I. Mechanisms of insulin action and insulin resistance. Physiol. Rev. 98, 2133–2223 (2018).
Hertzog, A. et al. A narrative review of metabolomics in the era of -omics: integration into clinical practice for inborn errors of metabolism. Transl Pediatr. 11, 1704–1716 (2022).
Hamosh, A., Amberger, J. S., Bocchini, C., Scott, A. F. & Rasmussen, S. A. Online mendelian inheritance in man (OMIM®): Victor McKusick’s magnum opus. Am. J. Med. Genet. A. 185, 3259–3265 (2021).
Stelzer, G. et al. The GeneCards suite: from Gene Data Mining to Disease Genome sequence analyses. Curr. Protocols Bioinf. 54, 1301–13033 (2016).
Mazumder, T., Akter, E., Rahman, S. M., Islam, M. T. & Talukder, M. R. Prevalence and risk factors of gestational diabetes Mellitus in Bangladesh: findings from Demographic Health Survey 2017–2018. Int. J. Environ. Res. Public. Health. 19, 2583 (2022).
Thakur, A., Agrawal, S., Chakole, S. & Wandile, B. A Critical Review of Diagnostic Strategies and Maternal Offspring Complications in Gestational Diabetes Mellitus. Cureus 15, e51016.
Sharma, A. K. et al. Deep insight of the pathophysiology of gestational diabetes Mellitus. Cells 11, 2672 (2022).
Xie, Y., Lin, S., Xie, B. & Zhao, H. Frontiers | recent progress in metabolic reprogramming in gestational diabetes mellitus: a review. https://doi.org/10.3389/fendo.2023.1284160
Filardi, T. et al. Role in gestational diabetes pathophysiology and complications. Int. J. Mol. Sci. 21, 4020 (2020). Non-Coding RNA.
Hong, M., He, J. & Li, S. SNW1 regulates notch signaling in neuroblastoma through interacting with RBPJ. Biochem. Biophys. Res. Commun. 509, 869–876 (2019).
Verma, S., De Jesus, P., Chanda, S. K. & Verma, I. M. SNW1, a Novel Transcriptional Regulator of the NF-κB pathway. Mol. Cell. Biol. 39, e00415–e00418 (2019).
Pérez-García, A. et al. Posttranscriptional regulation of insulin resistance: implications for metabolic diseases. Biomolecules 12, 208 (2022).
Payankaulam, S., Raicu, A. M. & Arnosti, D. N. Transcriptional regulation of INSR, the insulin receptor Gene. Genes 10, 984 (2019).
Chandrasekaran, P. & Weiskirchen, R. Cellular and Molecular mechanisms of insulin resistance. Curr. Tissue Microenvironment Rep. 1–12 https://doi.org/10.1007/s43152-024-00056-3 (2024).
Li, M. et al. Trends in insulin resistance: insights into mechanisms and therapeutic strategy. Signal. Transduct. Target. Therapy. 7, 1–25 (2022).
Hatting, M., Tavares, C. D. J., Sharabi, K., Rines, A. K. & Puigserver, P. Insulin regulation of gluconeogenesis. Ann. N Y Acad. Sci. 1411, 21–35 (2018).
Plows, J. F., Stanley, J. L., Baker, P. N., Reynolds, C. M. & Vickers, M. H. The pathophysiology of gestational diabetes Mellitus. Int. J. Mol. Sci. 19, 3342 (2018).
Thiel, G., Guethlein, L. A. & Rössler, O. G. Insulin-responsive transcription factors. Biomolecules 11, 1886 (2021).
Zhou, S., Tang, X. & Chen, H. Z. Frontiers | sirtuins and insulin resistance. https://doi.org/10.3389/fendo.2018.00748
Feng, J. et al. The Role of JNk Signaling Pathway in Obesity-Driven Insulin Resistance | DMSO.
Wang, W. et al. BMP and Notch Signaling pathways differentially regulate Cardiomyocyte Proliferation during ventricle regeneration. Int. J. Biol. Sci. 17, 2157–2166 (2021).
Manzari-Tavakoli, A. et al. Frontiers | the Cross-talks among Bone morphogenetic protein (BMP) signaling and other prominent pathways involved in neural differentiation. https://doi.org/10.3389/fnmol.2022.827275
Wang, N. et al. Notch signaling negatively regulates BMP9-induced osteogenic differentiation of mesenchymal progenitor cells by inhibiting JunB expression. Oncotarget 8, 109661–109674 (2017).
Shih, H. Y., Chang, C. W., Chen, Y. C. & Cheng, Y. C. Identification of the Time Period during which BMP signaling regulates proliferation of neural progenitor cells in zebrafish. Int. J. Mol. Sci. 24, 1733 (2023).
Su, J., Tang, L., Luo, Y., Xu, J. & Ouyang, S. Research progress on drugs for diabetes based on insulin receptor/insulin receptor substrate. Biochem. Pharmacol. 217, 115830 (2023).
Jokelainen, O. et al. Differential expression analysis identifies a prognostically significant extracellular matrix–enriched gene signature in hyaluronan-positive clear cell renal cell carcinoma. Sci. Rep. 14, 10626 (2024).
Terra Machado, D., Brustolini, B., Côrtes, O. J., Martins, Y. & Grivet Mattoso Maia, M. A. Ribeiro De Vasconcelos, A. T. Inference of differentially expressed genes using generalized linear mixed models in a pairwise fashion. PeerJ 11, e15145 (2023).
Szklarczyk, D. et al. The STRING Database in 2023: protein–protein Association Networks and Functional Enrichment analyses for any sequenced genome of interest.
Shannon, P. et al. Cytoscape: a Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 13, 2498–2504 (2003).
Tian, L. et al. Integrated protein–protein Interaction and weighted gene co-expression network analysis Uncover Three Key genes in Hepatoblastoma. Front. Cell. Dev. Biol. 9, 631982 (2021).
Cao, L. et al. Identification of hub genes and potential molecular mechanisms in gastric cancer by integrated bioinformatics analysis. PeerJ 6, e5180 (2018).
Cao, F., Cheng, Y. S., Yu, L., Xu, Y. Y. & Wang, Y. Bioinformatics Analysis of differentially expressed genes and protein–protein Interaction Networks Associated with Functional pathways in Ulcerative Colitis. Med. Sci. Monit. 27, (2021). e927917-1-e927917-11.
Yadav, S. et al. NMR based serum metabolomics revealed metabolic signatures associated with oxidative stress and mitochondrial damage in brain stroke. Metab. Brain Dis. 39, 283–294 (2023).
Kusum, K. et al. Elevated circulatory proline to glutamine ratio (PQR) in endometriosis and its potential as a diagnostic biomarker. ACS Omega. 7, 14856–14866 (2022).
U, K. et al. Circulatory Glutamine/Glucose ratio for evaluating disease activity in Takayasu arteritis: a NMR based serum metabolomics study. PubMed (2020).
Acknowledgements
The present work was supported by Indian Council of Medical Research (ICMR) and by the Science and Engineering Research Board (SERB), Ministry of Science and Technology, Government of India (Grant No. CRG/2019/002237) and Incentive grant for faculty under IOE-BHU. SR acknowledges the financial support from UGC-BSR start-up grant (File No.: 30593/2021(BSR)). RT acknowledges the receipt of the JRF fellowship from the UGC, New Delhi, India (201610145908).
Author information
Authors and Affiliations
Contributions
PKD, RB and DK were responsible for study, concept, design and writing—review and editing AR, RT, RR, DK, AT, AK, SGR and SR, were responsible for acquisition and interpretationof data, statistical analysis. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement consent to participate
This study was carried out following the required local ethical recommendations (Banaras Hindu University). All procedures followed were following the ethical standards of the responsible committee on human experimentation. Informed consent was obtained from all patients for being included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bhushan, R., Trivedi, R., Raj, R. et al. Integration of transcriptomics and metabolomics data revealed role of insulin resistant SNW1 gene in the pathophysiology of gestational diabetes. Sci Rep 15, 4159 (2025). https://doi.org/10.1038/s41598-025-88485-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88485-4