Abstract
Focal adhesion kinase (FAK) is overexpressed and hyperactivated in triple-negative breast cancer, driving tumor aggressiveness and cancer stem cell–mediated therapy resistance. Therefore, targeting FAK signalling represents a promising therapeutic strategy. In this study, a series of indole and bis-indole-1,2,4-triazoles were synthesized and evaluated as anti-TNBC agents targeting FAK. Compounds 3c, 4c, and 5c displayed potent cytotoxicity (IC₅₀ = 41–77 µg/mL) with minimal toxicity to normal cells, outperforming precursor compound 2. Wound-healing assay revealed significant inhibition of cell migration, particularly by 4c. Cell cycle analysis revealed that 4c induced S-phase arrest in MCF-7 cells and G1-phase arrest in MDA-MB-231 cells, accompanied by significant apoptosis. In MDA-MB-231 cells, 4c triggered extensive total apoptosis (90.84%) with minimal necrosis. Gene expression studies demonstrated that 4c markedly downregulated PTK2 (FAK), CCL5, and BCL2, while upregulating CASP3, highlighting its dual role as FAK inhibitor and apoptosis inducer. Importantly, 4c efficiently suppressed FAK protein expression (61.3%) in TNBC, compared to the FAK inhibitor GSK-2256098 (70.7%). In vivo toxicity assessment confirmed good tolerability in mice without profound hepatic or renal impairments, while docking and ADMET analyses confirmed strong FAK binding affinity, and favourable pharmacokinetics of 4c. Collectively, 4c emerges as a promising FAK-targeted candidate for TNBC therapy.
Similar content being viewed by others
Introduction
Breast cancer (BC) is the second leading cause of cancer-related mortality among women, and about 1–3 million cases are diagnosed worldwide every year1,2. Triple-negative breast cancer (TNBC) is the most aggressive and therapy-resistant breast cancer form, represents 20% of total BC cases and mostly affects young women under 40 years old3,4. TNBC is characterized by the low/lack of expression of three key receptors, including HER2 (human epidermal growth factor receptor 2), PR (progesterone receptors) and ER (estrogen receptors) and therefore not sensitive to available hormonal targeted therapies5. Clinically, TNBC is often associated with high histological grade, poor prognosis, increased risk of recurrence, and metastatic spread at an early stage. TNBC metastasizing is highly aggressive and frequently metastasizes to the lungs and central nervous system. TNBC is a highly heterogeneous disease, and in vitro cell line models are commonly classified into distinct molecular subtypes based on their genetic, transcriptomic, and morphological characteristics. The most widely classification describes six TNBC subtypes, including Basal-like 1 (BL1), Basal-like 2 (BL2), Immunomodulatory (IM), Mesenchymal (M), Mesenchymal stem-like (MSL), and Luminal androgen receptor (LAR) subtypes6,7,8. The BL1 subtype is characterized by elevated expression of genes involved in cell-cycle regulation and DNA damage response, with representative models including MDA-MB-468, HCC1937, HCC2157, HCC1599, HCC1143, HCC3153, and HCC389. BL2 tumors are enriched in growth-factor signaling pathways, such as EGFR and MET, and include cell lines such as SUM149PT, HCC1806, HCC70, CAL-851, and HDQ-P19. Immunomodulatory (IM) subtype is associated with immune-related pathways and cytokine signaling, includes cell lines such as HCC1187 and DU44759. The mesenchymal-like (M) subtype exhibits epithelial-to-mesenchymal transition (EMT) and enhanced cell motility, commonly represented by BT-549, CAL-51, and CAL-1209. The mesenchymal stem-like (MSL) subtype shares mesenchymal features but displays lower proliferative activity and higher angiogenic potential, with MDA-MB-231, HS578T, MDA-MB-157, SUM159PT, andMDA-MB-436 as typical models. The LAR subtype is driven by androgen receptor signaling and displays a luminal-like gene expression profile, with MDA-MB-453 considered a gold-standard model, in addition to SUM185PE, HCC2185, CAL-148, and MFM-223 models9. Collectively, these aggressive biological features contribute to limited therapeutic options, poor treatment outcomes, and reduced overall survival in patients with TNBC10. Currently, no targeted therapies are available for TNBC, and the most popular therapeutic options are the standard cytotoxic chemotherapies with significant side effects, lack of long-term efficacy, and tendency for rapid resistance development5,11. Hence, there is an urgent need for novel, effective, selective and safe treatment alternatives with recognized mechanism(s) of action.
Focal adhesion kinase (FAK) is a cytoplasmic tyrosine kinase that plays a vital role in regulating tumor development processes such as invasion, metastasis, and angiogenesis through both kinase-dependent and kinase-independent pathways12,13. Notably, FAK is frequently overexpressed in various human cancers, including TNBC cells, where its elevated levels are associated with enhanced invasion and metastasis14. Consequently, FAK inhibitors are emerging as one of the most promising targeted therapeutic approaches for TNBC treatment12. Several five-membered heterocyclic compounds, including oxadiazoles, triazoles, and thiadiazoles, have been identified as potent FAK inhibitors with significant anti-proliferative activity15,16,17.
1,2,4-Triazole derivatives have attracted renewed attention from organic and medicinal chemists, as several new hybrids with broader biological activity have recently been developed18,19,20. These compounds serve as key pharmacophores, interacting strongly with biological receptors due to their dipole moment, ability to form hydrogen bonds, good solubility, and resistance to both chemical and metabolic degradation21. Recently, four 1,2,4-triazole derivatives were synthesized, demonstrating potent anti-proliferative effects and significantly inhibiting FAK kinase at nanomolar concentrations22.
Moreover, indoles have gained significant interest due to their synthetic versatility and notable biological relevance, particularly in cancer research23. Interestingly, numerous studies indicate that conjugating indole with 1,2,4-triazole could significantly boost their anticancer activity24,25,26. In this context and as part of our continuing efforts to identify novel bioactive anticancer agents20,27,28,29,30, the present study aimed to design, synthesize, and characterize a new series of indole and bis-indole-linked 1,2,4-triazole derivatives, evaluating their in vitro anticancer potential against breast cancer cell lines, with special emphasis on triple-negative breast cancer (MDA-MB-231). The research investigated the possible mechanism underlying their cytotoxic action through cell-cycle arrest, apoptosis induction, and FAK inhibition. In addition, the selectivity and preliminary safety profile of the most active compounds, through in vitro and in vivo assays alongside in silico studies, was investigated.
Result and discussion
Chemistry
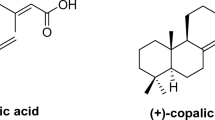
The synthesis of new indole-linked 1,2,4-triazole derivatives 3–8 was accomplished according to the synthetic routes illustrated in Fig. 1. Initially, the key precursor 5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazole-3-thiol (2) was synthesized through a straightforward one-pot fusion reaction between indole-3-propionic acid (1) and thiocarbohydrazide. Subsequently, the thiol (SH) functionality of compound 2 was subjected to S-alkylation reaction with various chloroacetamide derivatives, namely 2-chloro-N-phenylacetamide, 2-chloro-N-(4-chlorophenyl)acetamide, 2-chloro-N-(4-fluorophenyl)acetamide and 2-chloro-N-(4-(N-(pyridin-2-yl)sulfamoyl)phenyl)acetamide, which afforded the corresponding S-acetamide derivatives 3a-d (Fig. 1).
Synthesis of 1,2,4-triazole derivatives 2, 3a-d, 4a-c and 5a-d, reagent and condition: (i) oil bath, fusion at 130–140 °C for 4h; (ii) chloro acetamide derivatives, acetone, K2CO3, reflux; (iii) N-substituted indole-3-carboxaldehydes, EtOH/AcOH, refluxß (iv) chloro acetamide derivatives, acetone, K2CO3, reflux.
The 1HNMR spectra of 3a-d confirmed successful S-alkylation by the disappearance of the SH proton signal and the appearance of new singlet signals at δ 10.59–10.05 ppm, and δ 4.17–4.05 ppm (integrating for two protons), corresponding to NH and SCH₂-protons, respectively. For instance, the 1HNMR spectrum of 3a in DMSO-d₆ displayed four singlet signals at δ 10.77, 10.31, 5.93, and 4.06 assigned to 2NH, NH₂, and CH₂ protons, respectively. Additionally, multiplet signals observed at δ 3.05 ppm were assigned to the ethylene (CH₂–CH₂) protons linking the indole and triazole rings, along with characteristic aromatic proton resonances. The 13C NMR spectrum of 3a (DMSO-d6) further supported the proposed structure, showing three distinct CH₂ carbon signals at δ 39.91, 25.48, and 22.81 ppm, in addition to the expected aromatic carbon resonances (see Experimental Section).
Furthermore, a series of bis-indolyl-triazole Schiff bases 4a-c was synthesized through acid-catalyzed condensation reaction of compound 2 with indole-3-carboxaldehyde, N-ethyl and/or N-benzyl indole-3-carboxaldehydes under reflux in absolute ethanol (Fig. 1).
The 1HNMR spectra of compounds 4a–c confirmed Schiff base formation by the disappearance of the NH₂ proton signals and the appearance of new singlet resonances in the range δ 13.62–9.46 ppm, corresponding to SH, NH, and azomethine (CH=N) protons. For example, the 1HNMR spectrum of compound 4b (DMSO-d₆) displayed three singlet characteristic singlet signals at δ 13.62, 10.78 and 9.50 ppm which were attributed to SH, NH and CH=N protons, respectively. Additionally, quartet and triplet signals at δ 4.23 and 1.38 ppm were assigned to the NCH₂CH₃ protons, respectively, while signals observed at δ 3.07–3.06 ppm corresponded to the two methylene (CH₂) groups. The aromatic proton resonances were observed in the expected region. The 13C NMR spectrum (DMSO-d6) of 4b further supported the proposed structure, displaying CH₂ and CH₃ carbon signals at δ 50.69, 26.65, 22.43, and 15.69 ppm, alongside signals from aromatic carbons (Ar–C) (see Experimental Section).
In a subsequent synthetic route, S-alkylation of the bis-indolyl Schiff bases 4a–c with the different chloroacetamide derivatives, namely 2-chloro-N-phenylacetamide, 2-chloro-N-(4-chlorophenyl)acetamide, 2-chloro-N-(4-fluorophenyl) acetamide and 2-chloro-N-(4-(N-(pyridin-2-yl)sulfamoyl)phenyl)acetamide] yielded the corresponding bis-indolyl S-acetamide derivatives 5a-d (Fig. 1).
The 1HNMR spectra confirmed successful derivatization by the disappearance of SH signals and the appearance of new singlets for SCH₂ protons at δ 4.04–4.95 ppm. For instance, the 1HNMR spectra of compound 5a displayed characteristic singlet signals at δ 11.15, 10.81, 10.44, 8.66, and 4.95 ppm, corresponding to NH, azomethine (CH=N), and SCH₂ protons, respectively, in addition to multiplet signals at δ 3.07–2.99 assigned to the ethylene (CH₂–CH₂) linker and the expected aromatic proton signals. The 13CNMR spectrum of 5a further supported the proposed structure, displaying CH₂ carbon signals at δ 53.85, 25.04, and 22.91 ppm, consistent with the proposed structures (see Experimental Section).
Biological activities
Cytotoxic activity
The cytotoxic activities of the newly synthesized 1,2,4-triazole derivatives were evaluated against two human breast cancer cell lines (MCF-7 and MDA-MB-231) and a normal human fibroblast cell line (hFB) using the MTT assay after 24 h and 48 h of exposure. The half-maximal inhibitory concentration (IC50) values were summarized in Table 1. Overall, several derivatives demonstrated enhanced cytotoxicity compared to the parent compound 2, which exhibited relatively weak activity with IC50 values above 140 µg/mL on MCF-7 cells and more than 230 µg/mL on MDA-MB-231 cells after both incubation periods. This observation indicates that structural modification significantly improved the anti-proliferative potential of the synthesized analogues. Among the tested compounds, 1,2,4-triazole derivatives 3c, 4c, and 5c displayed the most potent activities with IC₅₀ values in the range of 41–78 µg/mL against both MCF-7 and MDA-MB-231 cell lines after 48 h of exposure (Fig. 2). Notably, compounds 3c, 4c and 5c demonstrated lower toxicity toward normal fibroblasts (IC₅₀ ≈ 130–183 µg/mL), suggesting a favorable selectivity profile. While, derivatives 3a, 3b, 4a, 5a, 5b, and 5d exhibited moderate cytotoxicity with IC50 values in the range of 50–99 µg/mL against both MCF-7 and MDA-MB-231 cell lines after 48 h. On the other hand, compounds 3d, 4b, and 5e showed weak cytotoxicity with IC50 > 140 µg/mL. A time-dependent increase in activity was evident for most active derivatives, with lower IC50 values observed after 48 h compared to 24 h, indicating that prolonged exposure enhances cytotoxic efficacy. Overall, 3c and 4c emerged as the most promising derivatives due to their potent and selective anti-proliferative effects.
Evaluation of cytotoxic response of compounds 2, 3c, 4c, and 5c on hFB, MCF-7, and MDA-MB-231 cell lines after treatment with compounds 2, 3c, 4c, and 5c for 24h and 48h. Data are presented as mean ± SEM (n = 3).
Wound healing assay
Wound healing assay was conducted to evaluate the cell migration response to a scratched region (Fig. 3A). Migration areas were quantified in µm2 to compare the effects of compounds 2, 3c, 4c, and 5c against non-treated cells. Breast cancer cell lines (MCF-7 and MDA-MB-231) and human skin fibroblast (hFB) cells were exposed to half the IC50 concentration (IC25) of the tested compounds for 48 h to identify the most potent compounds to inhibit cell migration effectively (Fig. 3B).
Wound healing assay of hFB, MCF-7, and MDA-MB-231 cell lines after treatment with compounds 2, 3c, 4c, and 5c by half the inhibitory concentration 50 (IC25) regarding to non-treated cells (NC) for different time intervals (0, 24, and 48 h.). (A) Representative photo panel for the scratched areas of the treated cell lines at different time points with scale bar 100 µm. (B) Scoring the closure rate of gap area of migration (µm2) regarding to the time (h) in different treatments. Data are presented as mean ± SEM (n = 3 independent biological replicates), and the significance variances were calculated regarding to control or the parent compound; * p < 0.05, ** p < 0.01, and *** p < 0.001.
The results illustrated that the delay in cell migration increased with time line, and the derivative compounds (S3c, S4c, and S5c) inhibited cell migration significantly at the same time point compared to the non-treated cells, or the parental compound 2. For hFB cells, compounds 4c and 5c declined cell migration remarkably at p < 0.01, and compounds 2 and 3c inhibited cell migration highly significantly at P < 0.001 in comparison to non-treated cells after 48h. For MCF-7 and MDA-MB-231 cells, the potential of derivative compounds (3c, 4c, and 5c) to inhibit closure of the scratched areas was delayed highly significantly (p < 0.001) from the initial time points compared to non-treated cells or cells treated with the principal compound 2.
The most active 1,2,4-triazole derivatives 3c, 4c, and 5c were further investigated for their cytotoxic mechanisms of action at 37.5 µg/mL for 48 h, in comparison to the parent compound 2.
Cell cycle analysis
Both MCF-7 and MDA-MB-231 cells responded differently to compounds 2, 3c, 4c, and 5c (37.5 µg/mL, 48 h). In MCF-7 cells, untreated controls displayed typical G1 predominance with minimal sub-G1 apoptosis. Compound 2 significantly reduced S-phase cells while increasing G2/M fraction (G2/M arrest), without substantial sub-G1 induction. Compound 4c prominently arrested cells at S-phase with marked apoptosis, while 3c and 5c induced S-phase arrest with minimal 3c and moderate 5c apoptosis, respectively (Fig. 4A). In MDA-MB-231 cells, all four compounds significantly arrested cells at the G1 phase (limiting DNA synthesis and mitosis), with 3c showing the weakest effect. Compounds 4c and 5c markedly elevated sub-G1 populations (apoptosis), with 4c demonstrating superior potency (Fig. 4B, Supp. Table 2).
Cell Cycle analysis of MCF7 and MDA-MB-231 cells. (A) It represents MCF7 cell frequencies among the cell cycle phases (G0/G1, S, and G2/M) in the different treatment conditions. (B) It represents MDA-MB-231 cell frequencies among the cell cycle phases in the different treatment conditions. Data are presented as mean ± SEM (n = 3 independent biological replicates), * 0.05 ≥ P ≥ 0.01; **: 0.01 > P ≥ 0.001; *** P < 0.001, **** < 0.0001. One-way ANOVA followed by Bonferroni’s multiple comparisons test was used for statistical analysis.
Apoptosis induction (annexin V/PI)
Supporting the sub-G1 cell cycle data, treatment with compound 4c significantly reduced the percentage of viable MCF-7 cells compared with untreated controls or compound 2 (Fig. 5A). This reduction was accompanied by a significant increase in early apoptotic (Annexin V + /PI −) and late apoptosis/secondary necrosis (Annexin V + /PI +) cell populations, in addition to a modest elevation in necrotic cells (Annexin V − /PI +) (Fig. 5B,C, Supp. Table 3). Compound 5c moderately but significantly, increased early apoptotic and late apoptosis/secondary necrosis populations in MCF-7 cells, whereas compounds 2 and 3c exerted negligible effects (Fig. 5B).
Annexin V-FITC/PI flow cytometric analysis of apoptosis in MCF-7 and MDA-MB-231 cells. The figure shows the percentage of (A) viable cells (Annexin V − /PI −), (B) early apoptotic (Annexin V + /PI −) and late apoptosis/secondary necrosis (Annexin V + /PI +) populations, and (C) primary necrotic cells (Annexin V − /PI +) under the indicated treatment conditions in both cancer cell lines. Data are presented as mean ± SEM (n = 3 independent biological replicates). Statistical significance indicators above the bars represent comparisons between each treated group and the corresponding untreated control, * 0.05 ≥ P ≥ 0.01; ** 0.01 > P ≥ 0.001; *** P < 0.001; **** P < 0.0001. Statistical analysis was performed using one-way ANOVA followed by Bonferroni’s multiple comparisons test.
In MDA-MB-231 (TNBC) cells, compound 4c demonstrated markedly stronger pro-apoptotic activity. The percentage of viable cells decreased from 96.56% (control) to 8.54% (Fig. 5A). Early apoptotic cells (Annexin V + /PI −) increased dramatically from 0.4% to 62.04%, while the late apoptosis/secondary necrosis population (Annexin V + /PI +) increased from 0.58% to 28.8%, yielding a total apoptotic of 90.84% (Fig. 5B, Supp. Table 3). In contrast, primary necrotic cells (Annexin V − /PI +) remained minimal (0.62%) (Fig. 5C).
Compound 5c induced apoptosis in MDA-MB-231 cells, although to a lesser extent (total apoptosis: 17.85%), with early apoptosis increasing to 13.44% and late apoptosis/secondary necrosis to 3.86%, accompanied by a reduction in viable cells to 80.4%. Compounds 2 and 3c exhibited mild apoptotic effects (5.2% and 4.6% total apoptosis, respectively).
Thus, 4c most potently drives TNBC apoptosis, with 5c intermediate between mild 2/3c effects and 4c’s extreme potency, consistent with prior reports of 1,2,4-triazolo-indolyl conjugates in TNBC models. The synthetic derivatives, particularly 4c, effectively arrested cell cycle progression in the TNBC MDA-MB-231 cell line at G1 phase while potently inducing programmed cell death (90.84% total apoptosis), thereby inhibiting tumor cell proliferation and migration. These findings align with previous reports demonstrating the cell cycle arrest and ROS-mediated apoptosis induction by 1,2,4-triazolo-linked indolyl conjugates across multiple cancer cell lines, including TNBC MDA-MB-231 models31.
Gene expression dysregulation
FAK is widely implicated in cancer growth, survival, motility, and drug resistance across multiple malignancies32. In breast cancer, FAK expression is elevated in several tumor cell lines, including the TNBC model MDA-MB-231, acting as a key player in mesenchymal cell migration33,34.
Consequently, targeting FAK/PTK2 expression and connected genes has emerged as an effective strategy for inhibiting tumor survival, immunosuppression, and migration/metastasis-associated programs32,35. Accordingly, we evaluated the effect of the most active synthesized indole-linked 1,2,4-triazole derivatives on the expression of the FAK gene (PTK2) and selected downstream functional readouts. Using RT-qPCR, we quantified mRNA expression levels of PTK2, apoptosis-linked genes (BCL2 and CASP3), and migration/invasion-linked genes (CCL5, vimentin, and MMP14) across two breast cancer cell lines: MDA-MB-231 and MCF-7. Our results demonstrated that the parent compound 2 modulated the expression of apoptosis-related genes in the MDA-MB-231 cell line, markedly decreasing BCL2 (anti-apoptotic) and increasing CASP3 (pro-apoptotic) expression, while showing no significant effect in the MCF-7 cell line. Compound 3c significantly upregulated the levels of both BCL2 and CASP3 in MDA-MB-231 cells, without any observable impact on MCF-7 cells. Notably, compound 4c markedly downregulated the mRNA expression levels of PTK2 (direct on-target inhibition of FAK), together with strong downregulation of the pro-tumor chemokine CCL5, and the anti-apoptotic regulator BCL2, while significantly raising the levels of the proapoptotic mediator CASP3 in the MDA-MB-231 cells. Interestingly, compound 4c also induced a significant suppression of CCL5 expression level in MCF-7 cells. In contrast, compound 5c significantly elevated VIM and CASP3 in the MDA-MB-231 cells and enhanced CCL5 expression levels in the MCF-7 cell line (Fig. 6). Collectively, these findings suggest that compound 4c exhibits a strong inhibitory potential against mRNA expression of FAK/PTK2 and induces concomitant changes in apoptosis- and migration-related transcripts in MDA-MB-231 cells. This is consistent with the findings of Chen et al.33, who demonstrated that their newly developed FAK inhibitor activated caspase-3, promoting the formation of cleaved caspase-3. They also reported a dose-dependent inhibition of Bcl-xL (a BCL2 family protein) by their FAK inhibitor, further supporting its apoptotic activity. Furthermore, the downregulated expression of CCL5 by compound 4c in MDA-MB 231 can be correlated with the downregulation of FAK/PTK2, as it was previously reported that silenced FAK reduces CCL5 expression levels, indicating that FAK is a key regulator of CCL5 expression36. Hence, downregulation of CCL5 is in line with the FAK inhibitory action of our 4c compound. Importantly, in breast cancer models, CCL5 was found to increase the migration/invasion process37 downregulation of CCL5 is associated with decreased cell migration, as observed in our study. Also, it can be deduced that compound 4c suppressed FAK-induced tumour immune evasion, which is mediated by CCL538. However, mechanistic studies are required to establish causality.
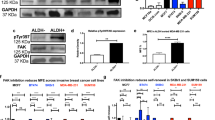
RT-qPCR results of FAK-related genes in breast cancer cell lines following treatment with the most active compounds. The figure shows the expression levels of PTK2 (FAK), PDL, VEGFR2, VIM, CCL5, CDH1, BCL2, and CASP3 in MDA-MB-231 and MCF-7 cells that were treated with the parent compound (2) and the selected active derivatives (3c, 4c, and 5c) alongside untreated controls. Relative expression is presented as fold change calculated by the comparative method (2−ΔΔCt) normalized to GAPDH. Log scale was used to visualize both up- and downregulation. Data are represented as mean ± SEM of three independent biological replicates (n = 3), *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001.
FAK protein levels reduction
FAK protein levels were quantified in MDA-MB-231 breast cancer cells following treatment with the IC50 concentration of the test compounds using the Human FAK SimpleStep ELISA® Kit (Abcam, ab187395). The highest FAK level (59.09 ± 2.29 ng/mL) was detected in the control untreated cells (Table 2, Fig. 7). Meanwhile, treatment with the compound 4c significantly reduced FAK levels to 22.88 ± 0.88 ng/mL, representing a 61.3% inhibition relative to the control. 4c-mediated reduction of FAK levels was comparable to that induced by the reference FAK inhibitor GSK-2256098, which lowered FAK expression to 17.28 ± 0.67 ng/mL, corresponding to 70.7% inhibition. As well, compound 3c treatment significantly decreased FAK levels to 29.33 ± 1.13 ng/mL (50.4% inhibition), while compound 2 caused a moderately significant reduction, resulting in a FAK concentration of 41.31 ± 1.60 ng/mL (30.1% inhibition) (Fig. 7, Table 2). Overall, all tested compounds suppressed FAK expression in MDA-MB-231 cells in a concentration-dependent manner, with compound 4c showing the most potent inhibitory activity, followed by compounds 3c and 2. These findings suggest that the synthesized compounds, especially 4c, may serve as effective modulators of the FAK pathway comparable to the standard inhibitor.
Effect of compounds 2, 3c, and 4c on FAK protein levels in MDA-MB-231 cells Welch’s t test was sued for statistical pairwise comparison. Abbreviations: FAK: Focal adhesion kinase, GSK-2256098: A reference FAK inhibitor.
From all the above-mentioned in vitro studies, we suggest that the decline of cell migration together with inhibition of gene and protein expressions of CCL5 and FAK markers might contribute, remarkably, to the anti-migratory potential of the investigated compounds on breast cancer cell lines and, in particular, the TNBC cells. These findings are in agreement with the previous reports that illustrated the direct correlation between cell migration and the protein expressions of chemotactic CCL539, or focal adhesion kinase (FAK) protein markers40, which play crucial roles in cell migration, invasion, and metastatic pathways. Furthermore, the new indole-linked 1,2,4-triazole derivatives activated the programmed cell death pathways on the tested cancer cell line (MCF-7 and MDA-MB-231 cells) through inhibition of oncogenic markers BCL2, CCL5 and FAK and activation of apoptotic markers CASP3 and VIM pathways remarkably. This apoptotic response could result from the release of ROS from the indole-linked 1,2,4-triazole derivatives, which enhanced cell cycle arrest remarkably in both breast cancer cell lines. These observations agree with previous reports on numerous cancer cell lines, including TNBC MDA-MB-231 cells, which exhibited apoptotic response, cell cycle arrest and potential release of ROS after treatment with 1,2,4-triazolo-linked-indolyl conjugates31.
Structure–activity relationship (SAR) analysis
A comparative SAR analysis of the most active compounds 3c, 4c and 5c revealed key structural features likely influencing their biological activity. Compound 4c exhibited the highest potency, which can be attributed to its bis-indole framework connected through a 1,2,4-triazole ring bearing a thiol group. The dual indole units enhance aromatic interactions, while the triazole and thiol functionalities facilitate hydrogen bonding and sulfur-mediated binding. A benzyl substituent further improved lipophilicity and molecular flexibility, supporting effective target engagement. Compound 3c, with a single indole and a para-fluorophenyl thioamide substituent, retained essential pharmacophores but shows reduced activity, likely due to lower aromatic density despite beneficial electronic effects from fluorine. Compound 5c displayed increased steric hindrance and structural rigidity arising from extended substitution around the triazole–thioamide region. Overall, these findings emphasize the critical role of enhanced aromaticity, balanced molecular flexibility, and accessible sulfur-containing functionalities in maximizing biological activity, with compound 4c representing the most favorable structural configuration within this series.
In vivo toxicity study
Administration of a single intraperitoneal dose of the synthesized chemical compound of 12.5, 25, and 50 mg/kg to mice showed no clinical toxicity signs, including behavioral alterations or mortality in the following 48 h.
The impact of the chemical compound 4c on the histopathological differences in
-
a.
Liver
Liver tissues of the negative control group: Demonstrated regular hepatic architecture with mild cloudy swelling of hepatocytes. Liver tissues of 12.5 mg-treated group: Displayed moderate hydropic degeneration and mild intra-lobular neutrophilic infiltration. Liver tissues of the 25 mg treated group: Revealed similar hepatocellular degeneration with more extensive neutrophilic infiltration involving both portal and intra-lobular areas. Liver tissues of the 50 mg treated group: Showed marked histopathological damage, including dense neutrophilic infiltration and patchy necrosis (Fig. 8, Table 3).
The histopathological findings of the hepatic tissues of different experimental groups receiving a single intraperitoneal dose of the synthesized compound 4c (12.5, 25, and 50 mg/kg), as compared to the negative control group. Photomicrographs (a and b): Sections in the liver of negative control mice showing regular hepatic lobular architecture with mild diffused cloudy swelling of hepatocytes (yellow arrows) (left H&E stain X100; right H&E stain, X400). Photomicrographs (c and d): Sections in the liver of 12.5 mg 4c-treated mice showing preserved hepatic lobular architecture with moderate diffuse hydropic degeneration of hepatocytes (yellow arrow) and focal intra-lobular aggregates of neutrophils (red arrows) (H&E stain, left X100; right, X400). Photomicrographs (e and f): Sections in the liver of 25 mg 4c-treated mice showing moderate diffuse hydropic degeneration of hepatocytes (yellow arrow) and focal portal and intra-lobular aggregates of neutrophils (red arrows) (H&E stain, left X100; right, X400). Photomicrographs (g and h) sections in the liver of 50 mg 4c-treated mice; showing dense focal intra-lobular aggregates of neutrophils (red arrows) with patchy hepatocellular necrosis (H&E stain, left X200; right, X400).
-
b.
Kidney
Renal tissues of the negative control group: Showed normal histological structure with intact glomeruli and renal tubules. Renal tissues of the 12.5 mg treated group: Demonstrated mild focal epithelial degeneration and slight irregularity in glomerular contours. Renal tissues of the 25 mg treated group: Exhibited changes similar to the 12.5 mg group, with mild degeneration and irregular contours. Renal tissues of the 50 mg treated group: Showed more severe alterations, including variable glomerular sizes, focal sclerosis, cortical inflammation, and medullary fibrosis (Fig. 9, Table 4).
The histopathological findings of the renal tissues of different experimental groups receiving a single intraperitoneal dose of the synthesized compound 4c (12.5, 25, and 50 mg/kg), as compared to the negative control group. Photomicrographs (a, b, c, and d) sections: in the kidney of negative control mice showing average glomeruli in number and size (yellow arrows), with intact proximal (blue arrow) and collecting tubules (red arrow) (H&E stain, upper photos X200, lower photos X400). Photomicrographs (e, f, g, and h): Sections in the kidney of 12.5 mg 4c-treated mice showing average number and size of glomeruli (yellow arrows), with mild irregularity in the contour as well as mild focal epithelial degeneration of proximal (blue arrow) and collecting tubules (red arrow). (H&E stain; upper left X200; upper right, lower left and lower right X400). Photomicrographs (i, j, k, and l): Sections in the kidney of 25 mg 4c-treated mice showing average number and size of glomeruli (yellow arrows), with mild irregularity in the contour as well as mild focal epithelial degeneration of proximal (blue arrow) and collecting tubules (red arrow). (H&E stain, upper left X200; upper right, lower left and lower right X400) Photomicrographs (m, n, o, and p): Sections in the kidney of 50 mg 4c-treated mice showing variable glomerular sizes with focal sclerosis (yellow arrows) with focal cortical inflammation (blue arrow) and medullary fibrosis (red arrow) (H&E stain, X400).
Impact of different concentrations of the compound 4c on serum levels of aspartate aminotransferase (AST) and Kidney-injury molecule 1 (KIM-1)
In order to find out which concentration of 4c could induce hepatotoxicity and to evaluate the proper performance of the liver, serum AST levels were measured. Concentrations of 12.5, 25, and 50 mg/kg caused a non-significant increase in AST levels by 2.7 for a concentration of 12.5 and a significant increase for the other two concentrations by 10.51 and 22.15%, respectively, as compared to negative control mice (Table 5). On the other hand, KIM-1 was employed to evaluate the degree of the inflammation in the renal tissues, the i.p. administration of different concentrations of 12.5, 25, and 50 mg/kg of the 4c resulted in a significant elevation of serum KIM-1 levels by 295.33, 653.27, and 1035.55%, respectively, as compared to negative control group (Table 5).
Impact of different concentrations of the compound 4c on oxidative stress status of lipid peroxidation and nitric oxide (NO) production in the hepatic and renal tissues
The extent of oxidative injury in the liver and kidney was evaluated by estimating levels of malondialdehyde (MDA), an indicator of lipid peroxidation, and nitric oxide (NO). A statistically significant increment in MDA levels was detected in mice that received an intraperitoneal dose of different concentrations of 25 and 50 mg/kg of 4c by 17.86, and 35.22% respectively, while the dose of 12.5 mg/kg (0.55%) did not cause a significant change, compared to the negative control mice (Table 8). On the other side, hepatic NO contents were significantly increased in mice receiving different concentrations of 12.5, 25, and 50 mg/kg of 4c by 80.74, 170.64, and 197.19%, respectively, as compared to the negative control mice (Table 6).
Regarding the stimulated oxidative stress status in the renal tissues, the renal MDA contents were increased significantly in the treated mice by 14.85, 25.85, and 28.50%, respectively, as compared to negative control mice, for concentrations of 12.5, 25, and 50 mg/kg of 4c. Similarly, renal NO was significantly increased for concentrations of 25 and 50 mg/kg (30.71, and 49.94% respectively), while a concentration of 12.5 mg/kg of 4c exhibited a non-significant change, compared to the negative control mice (Table 6).
In silico study
Molecular Docking study
To elucidate the molecular basis of FAK inhibition, docking simulations were conducted for the most active synthesized compounds 3c and 4c within the FAK active site (PDB ID: 2JKK). The docking results are summarized in Table 9. The co-crystallized ligand BI9 displayed the best docking score (-10.0 kcal/mol), forming conventional hydrogen bonds with Asp564 and Cys502, in addition to multiple hydrophobic interactions (Table 7, Fig. 10).
(A) The 3D conformations of the co-crystalline ligand, BI9 (yellow), and the re-docked ligand (green), compound 3c (brown), compound 4c (cyan) within the binding pocket of FAK (PDB ID: 2JKK); indicating that they were superimposed in the same position. (B) 2D conformations of the re-docked BI9 within the FAK binding pocket (PDB ID: 2JKK). (C) 2D conformations of compound 3c within the FAK binding pocket (PDB ID: 2JKK). (D) The 2D conformations of the re-docked compound 4c within the FAK binding pocket (PDB ID: 2JKK).
Among the synthesized derivatives, compound 4c exhibited a high binding affinity (− 9.5 kcal/mol), closely approximating that of BI9. It established several hydrogen bonds with Glu430, Asn551, Arg550, and Ile428, along with attractive charge, π-donor H-bond, π-anion and π-alkyl interactions mediated by its bis-indole and phenyl moieties with key residues, including Glu506, Glu430, Leu567, Leu553, Ala452, and Ile428. These interactions, along with extensive van der Waals contacts, support the formation of a highly stable complex within the ATP-binding cleft of FAK (Fig. 10).
Compound 3c also showed strong binding affinity (-8.7 kcal/mol), forming three conventional hydrogen bonds with Glu506, Gly430, and Asp564, as well as carbon-hydrogen and halogen bonds involving its fluorine atom and residues Gly563 and Asn551. Its aromatic systems contributed to further stabilization via π-alkyl interactions with Leu567, Leu553, Val484, Ile428 and Ala452 (Fig. 10). Collectively, the docking outcomes correlate well with the ELISA findings, confirming that compound 4c exhibits the strongest affinity for FAK and achieves the greatest inhibition of FAK protein expression in MDA-MB-231 cells. The presence of bis-indole and amino-triazole-thione functionalities appears to enhance binding through multiple hydrogen bonding and π-π stacking interactions, thereby reinforcing its potential as a promising FAK-targeted anticancer agent.
Drug likeliness properties and ADMET prediction
The drug-likeness properties of compounds 3c and 4c were evaluated using the SwissADME online tool (http://www.swissadme.ch/index.php), and their profiles were compared to the reference drug with doxorubicin. The calculated physicochemical parameters were interpreted according to Lipinski’s Rule of Five and Veber’s criteria, as presented in Table 841,42.
The results indicated that both compounds generally satisfied Lipinski’s Rule of Five, with compound 4c displaying a single violation related to its lipophilicity (MLOGP > 4.15). In addition, both compounds complied with Veber’s criteria (nRB ≤ 10 and TPSA ≤ 140 Å2), supporting their potential for oral bioavailability. In comparison, the reference drug doxorubicin exhibited multiple violations of both Lipinski’s and Veber’s criteria due to its high molecular weight, elevated TPSA, and excessive numbers of hydrogen bond donors/acceptors.
On the other hand, the ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) properties of compounds 3c and 4c were predicted using the pkCSM online server (http://biosig.unimelb.edu.au/pkcsm/), and the results were compared with doxorubicin (Table 9).
Both compounds showed moderate solubility and high intestinal absorption (87.39% and 90.99%, respectively) suggest good gastrointestinal uptake, markedly higher than doxorubicin (62.63%). Both derivatives exhibited low skin permeability, consistent with limited transdermal diffusion. Notably, 3c was identified as a P-glycoprotein substrate, while 4c was not, implying that 4c may have a lower tendency for efflux-mediated resistance. In addition, both compounds were predicted to inhibit P-gp I and II, unlike doxorubicin. The distribution parameters revealed low tissue distribution (log VDss < 0), with 4c exhibiting moderate blood–brain barrier penetration (log BB = 0.238) and higher CNS permeability than 3c or doxorubicin. Both derivatives were substrates of CYP3A4, indicating hepatic metabolism via this isoform, and acted as inhibitors of several CYP enzymes, suggesting possible drug–drug interactions during co-administration. The predicted total clearance values revealed that 4c may be more efficiently cleared than 3c, although both were lower than doxorubicin.
Toxicity predictions showed 3c mutagenic (Ames test) while 4c non-mutagenic (like doxorubicin); both hepatotoxic but non-sensitizing, hERG II positive (like doxorubicin), indicating cardiotoxicity risk, with 4c having a higher LD₅₀ than 3c.
Compound 4c potent FAK suppression-rivalling GSK-2256098, follows a temporal cascade as predicted by molecular docking: compound 4c binds FAK’s ATP-binding pocket, forming hydrogen bonds with hinge residues Met499/Glu500/Glu506, confirming immediate kinase inhibition that prevents Y397 autophosphorylation. Inactive FAK cannot recruit Src, disrupting the FAK-Src complex and downstream PI3K/Akt and MAPK/ERK survival pathways43. Kinase-dead FAK loses conformational stability and undergoes ubiquitin-proteasomal degradation after 12h, consistent with ATP-competitive inhibitors like GSK225609844. This PI3K/Akt inhibition triggers FoxO3a dephosphorylation and nuclear translocation45, contributing to secondary transcriptional repression of PTK2 mRNA. This temporal cascade—kinase inhibition → signaling disruption → protein degradation → mRNA reduction—explain 4c’s dual suppression of FAK expression at both protein and gene levels.
Conclusion
In summary, a new series of indole and bis-indole-linked 1,2,4-triazole derivatives was successfully synthesized and characterized, leading to the identification of promising anticancer candidates. Among the synthesized compounds, 4c exhibited the most potent cytotoxic activity against MCF-7 and MDA-MB-231 breast cancer cells, with significant selectivity toward cancer cells over normal fibroblasts. Mechanistic investigations revealed that 4c induced S-phase arrest in MCF-7 cells and G1-phase arrest in MDA-MB-231 cells. The striking 4c-mediated effect was apoptosis induction, specifically in MDA-MB-231 cells, approaching almost 90% of the treated population. Furthermore, gene expression studies demonstrated that 4c markedly downregulated PTK2 (FAK), CCL5, and BCL2, while upregulating CASP3, highlighting its role as a dual FAK inhibitor and apoptosis inducer. Moreover, 4c significantly suppressed FAK expression, indicating a FAK-targeted mode of action consistent with its molecular docking profile.
In vivo toxicity study demonstrated that 4c is well-tolerated at doses up to 12.5 mg/kg, without detectable hepatotoxicity or nephrotoxicity, this was based on the biochemical and histopathological findings that indicated that the lowest concentration of the compound 4c (12.5 mg/kg/on day) exhibited the safest toxological profile and could be regarded as a promising candidate as a chemotherapeutic agent. Histopathologically, the study evaluated structural changes in liver and kidney tissues across the three experimental groups, as compared to the negative control group, and the results demonstrated a dose-dependent toxic pattern of tissue alteration in both organs. Mild changes were noted at lower doses (12.5 and 25 mg), while severe tissue damage, including necrosis and fibrosis, was observed at the highest dose (50 mg). These findings suggest potential toxicity risks associated with increasing dosage, underlining the need for cautious dose selection in therapeutic or experimental applications.
Furthermore, the current study demonstrated that the lowest concentration (12.5 mg/kg/on day) was capable of inducing the lowest extent of oxidative stress in the liver and the kidney, in addition, the lowest percentage of change as compared to negative control was recorded for this concentration for both serum levels of AST and KIM-1, as biomarkers related to hepatic dysfunction and renal inflammation. Finally, the histopathological findings of the hepatic and renal tissues run in agreement with the biochemical findings, indicating the lowest toxicity potential of the lowest dose of the synthesized compound 4c.
Therefore, further experimental studies and safety testing using different doses up to 12.5 mg/kg are required. In addition, further validation in cancer-induced animal models and comparing its therapeutic activities to standard chemodrugs such as doxorubicin or cisplatin is essential to confirm the therapeutic anti-cancer potential and safety of the synthesized compound 4c.
On the other hand, in silico ADMET predictions supported its favorable pharmacokinetic and safety characteristics. Collectively, these findings highlight compound 4c as a promising FAK-modulating bis-indole 1,2,4-triazole scaffold with potent anticancer activity, mechanistic selectivity, and an acceptable safety profile, warranting further preclinical optimization and mechanistic exploration as a lead candidate for TNBC therapy.
Experimental
Chemistry
Materials and reagent
All reagents and solvents were of commercial grade. Indole-3-propionic acid was purchased from Sigma-Aldrich ChemieGmeH, Taufkirchen, Germany. Melting points were determined on the digital melting point apparatus (Electrothermal 9100, Electrothermal Engineering Ltd., serial No. 8694, Rochford, United Kingdom) and are uncorrected. The reaction progress was monitored by thin-layer chromatography (TLC) using silica gel plates (POLYGRAM SILG/UV254, 0.20 mm), which were visualized under UV light 254 and 365 nm. The Fourier Transform Infrared (ATR-FTIR) spectra were obtained employing an FTIR spectrometer (Vertex 70, Bruker); the spectra are determined within a spectral range of 4000–400 cm−1 with a spectral resolution of 4 cm−1. The 1H and 13C NMR spectra were recorded using a JEOL-ECA-50 NMR instrument at 500 and 125 MHz, respectively, using TMS as the internal standard, National Research Center, Egypt. Hydrogen coupling patterns are described as (s) singlet, (d) doublet, (t) triplet, (q) quartered, (m) multiple. Chemical shifts were defined as parts per million (ppm) relative to the solvent peak. El-MSElectron ionization mass spectrometry 70 electron volt. Isq thermo scientific Italy 2009. 5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4- triazole-3-thiol (2) was prepared according to the reported methods46.
General procedure for the preparation of 5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazole-3-thiol (2)
A mixture of 3-(1H-indol-3-yl)propanoic acid (1, 0.01mol) and thiocarbohydrazide (0.01 mol) was ground and mixed in a mortar and then heated in an oil bath at 130 °C for 5h. After cooling, a dilute aqueous solution of Na2CO3 (10 mL, 20%) was added, and the mixture was heated at 70 °C for 30 min. The precipitate formed upon cooling was collected by filtration, washed with water, air-dried, and crystallized from methanol.
General procedure for the preparation of 2-((5-(2-(1H-Indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazol-3-yl)thio)-N- acetamide derivatives (3)
A mixture of 5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazole-3-thiol (2, 0.01mol) and 2-chloro acetamide derivatives (0.01mol) in dry acetone (20 mL) containing anhydrous potassium carbonate (0.02 mol), was heated under reflux for 14–16 h (monitored by TLC). The reaction mixture was cooled to room temperature and poured into ice water. The Precipitate formed was filtered off, washed with water, air-dried, and re-crystallized from n-hexane/ethyl acetate.
2-((5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazol-3-yl)thio)-N-phenylacetamide (3a)
Gray ppt, m.p.112-4 °C; yield: 56% for C20H20N6OS (M.wt 392); IR (cm−1) 3414, 3322 (NH2), 3233 (NH), 1679 (C=O), 1612 (C=N), 1583 (C=C); 1H NMR (500 MHz, DMSO-d6) δ 10.77, 10.31 (s, 2H, 2NH), 7.54–6.95 (m, 10H, Ar–H), 5.93 (s, 2H, NH2), 4.06 (s, 2H, S-CH2), 3.05 (m, 4H, 2CH2); 13C NMR (125 MHz, DMSO-d6) δ 166.71 (C=O), 156.78 (C=N of triazole), 151.43 (C=N of triazole), 139.35, 151.43, 139.35, 136.81, 129.33, 127.50, 124.06, 122.97, 121.50, 119.70, 118.79, 111.93, 39.9 (S-CH2), 25.48 (CH2), 22.81 (CH2).
2-((5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazol-3-yl)thio)-N-(4-chlorophenyl) acetamide (3b)
Off white ppt, m.p. 111–3 °C; yield: 56%; IR (cm−1) 3296 (NH2), 3055 (NH), 1668 (C=O), 1630 (C=N), 1592 (C=C);1H NMR (500 MHz, DMSO-d6) δ 10.77, 10.05 (s, 2H, 2NH), 7.92–6.94 (m, 9H, Ar–H), 5.93 (s, 2H, NH2), 4.17 (s, 2H, S-CH2), 3.06, 3.02 (m, 4H, 2CH2); 13C NMR (125 MHz, DMSO-D6) δ 167.88 (C=O), 161.60 (C=N of triazole), 159.83 (C=N of triazole), 156.18, 154.52, 151.38, 136.70, 134.36, 128.08, 127.45, 126.14, 123.16, 121.34, 118.64, 113.75, 111.74, 35.20 (S-CH2), 25.45 (CH2), 22.88 (CH2); m/z: 426/428 (M+/M+ + 2) for C20H19ClN6OS (M.wt 426.92).
2-((5-(2-(1H-Indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazol-3-yl)thio)-N-(4-fluorophenyl) acetamide (3c)
Brown ppt, m.p. 104-6 °C; yield: 56% for C20H19FN6OS (M.wt 410.47); IR (cm−1) 3282 (NH2), 3202 (NH), 1656 (C=O), 1605 (C=N), 1546 (C=C) ;1H NMR (500 MHz, DMSO-d6) δ 10.82, 10.51 (2s, 2H, 2NH), 7.58–7.03 (m, 9H, Ar–H), 5.97 (s, 2H, NH2), 4.05 (s, 2H, S-CH2), 3.05, 3.02 (m, 4H, 2CH2); 13C NMR (125 MHz, DMSO-d6) δ 184.93 (C=O), 169.23, 166.73, 166.11, 159.63, 157.70, 156.87, 152.30, 151.19, 144.74, 136.85, 135.83, 135.15, 127.57, 122.98, 121.49, 118.77, 115.90, 115.73, 111.94, 56.60 (S-CH2), 25.58 (CH2), 24.49 (CH2).
2-((5-(2-(1H-Indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazol-3-yl)thio)-N-(4-(N-(pyridin-2-yl) sulfamoyl)phenyl)acetamide (3d)
Pale brown ppt, m.p. 79–81 °C; yield: %; IR (cm−1) 3353 (NH2), 3186, 3052 (NH), 1659 (C=O), 1630 (C=N), 1591 (C=C) ;1H NMR (500 MHz, DMSO-d6) δ 10.78, 10.59, 10.37 (3s, 3H, 3NH), 7.94–6.67 (m, 13H, Ar–H), 5.91 (s, 2H, NH2), 4.06 (s, 2H, S-CH2), 3.04–2.95 (m, 4H, 2CH2); 13C NMR (125 MHz, DMSO-d6) δ 175.36 (C=O), 164.72, 163.09, 162.77, 160.94, 151.83, 146.04, 145.30, 134.90, 134.65, 127.68, 123.30, 122.91, 122.83, 116.60, 111.54, 111.38, 101.01, 54.12 (S-CH2), 26.46 (CH2), 22.79 (CH2); m/z: 548 (M+) for C25H24N8O3S2 (M.wt 548.64).
General procedure for the preparation of 5-(2-(N-substituted-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazole-3-thiol (4)
A mixture of 5-(2-(1H-indol-3-yl)ethyl)-4-amino-4H-1,2,4-triazole-3-thiol (2, 0.01mol) and N-substituted indole-3-carboxaldehydes (0.01mol) in absolute ethanol (10 mL) containing a few drops of glacial acetic acid was refluxed for 10-12h. After all, starting compounds was consumed as indicated by TLC (n-hexane/ethyl acetate 3:1), the reaction mixture was cooled to room temperature and poured onto ice water. The Precipitate formed was filtered off, washed with water, air-dried, and re-crystallized from n-hexane–ethyl acetate.
5-(2-(1H-Indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazole-3-thiol (4a)
Canary ppt, m.p. 214-6 °C; yield: 95%; IR (cm−1) 3278, 3193 (NH), 1631 (C=N), 1591 (C=C), 1241 (C=S); 1H NMR (500 MHz, DMSO-d6) δ 13.61 (s, 1H, SH), 12.03, 10.78 (2s, 2H, 2NH), 9.50 (s, 1H, CH=N), 8.17 (d, 1H, indolyl H-4`), 8.10 (s, 1H, indolyl H-2`), 7.49, 7.43 (2d, 2H, indolyl H-4 & H-7`), 7.26–7.11 (m, 6H, Ar–H), 3.07 (m, 4H, 2CH2); 13C NMR(125 MHz, DMSO-d6) δ 162.55 (S-C=N of triazole), 161.90 (C=N), 151.21 (C=N of triazole), 137.97, 136.78, 135.90, 123.82, 123.10, 122.68, 122.57, 122.12, 121.50, 121.50, 118.80, 111.96, 26.69 (CH2), 22.46 (CH2); m/z: 386 (M+) for C21H18N6S (M.wt 386.48).
5-(2-(1H-indol-3-yl)ethyl)-4-(((1-ethyl-1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazole-3-thiol (4b)
Green ppt, m.p. 234–6 °C; yield: 81%; IR (cm−1) 3266 (NH), 1630 (C=N), 1593 (C=C), 1240 (C=S); 1H NMR (500 MHz, DMSO-d6) δ 13.62 (s, 1H, SH), 10.78 (s, 1H, NH), 9.50 (s, 1H, CH=N), 8.88 (m, 2H, indolyl H-2` & H-4`), 8.42–7.11 (m, 8H, Ar–H), 4.23 (q, 2H, CH2), 3.07, 3.06 (m, 4H, 2CH2), 1.38 (t, 3H, CH3); 13C NMR(125 MHz, DMSO-d6) δ 162.55 (S-C=N of triazole), 161.85 (C=N), 151.16 (C=N of triazole), 137.69, 137.57, 136.91, 123.08, 122.78, 121.47, 118.75, 118.64, 113.31, 111.92, 111.38, 50.69 (N-CH2), 26.65 (CH2), 22.43 (CH2), 15.69 (CH3); m/z: 414 (M+) for C23H22N6S (M.wt 414.53).
5-(2-(1H-indol-3-yl)ethyl)-4-(((1-benzyl-1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazole-3-thiol (4c)
Brown ppt, m.p. 176–8 °C; yield: 85%; IR (cm−1) 3216 (NH), 2767 (SH), 1617 (C=N), 1593 (C=C);1H NMR (500 MHz, DMSO-d6) δ 13.63 (s, 1H, SH), 10.78 (s, 1H, NH), 9.53 (s, 1H, CH=N), 8.26, 8.19 (m, 2H, indolyl H-2` & H-4`), 7.30–7.11 (m, 13H, Ar–H), 5.50 (s, 2H, N-CH2), 3.08, 3.07 (m, 4H, 2CH2); 13C NMR(125 MHz, DMSO-d6) δ 185.25 (C=S), 171.38 (C=N), 161.91 (C=N), 151.23 (C=N), 137.99, 137.53, 136.79, 135.90, 129.26, 127.84, 123.11, 121.52, 118.82, 113.37, 112.97, 111.97, 60.32 (N-CH2), 26.96 (CH2), 22.47 (CH2); m/z: 476 (M+) for C28H24N6S (M.wt 476.60).
General procedure for the preparation of 2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-acetamide derivatives (5a-d)
A mixture of Schiff’s base (4a, 0.01mol) and 2-chloro acetamide derivatives (0.01mol) in dry acetone (20 mL) containing anhydrous potassium carbonate (0.02 mol), was heated under reflux for 14–16 h (monitored by TLC). The reaction mixture was cooled to room temperature and poured into ice water. The Precipitate formed was filtered off, washed with water, air-dried, and re-crystallized from n-hexane/ethyl acetate.
2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-phenyl-acetamide (5a)
Brown ppt, m.p. 125–7 °C; yield: %; IR (cm−1) 3328, 3227, 3148 (NH), 1675 (C=O), 1612 (C=N), 1580 (C=C); 1H NMR (500 MHz, DMSO-d6) δ 11.15, 10.81, 10.44 (s, 3H, 3NH), 8.66 (s, 1H, CH=N), 8.20–6.96 (m, 15H, Ar–H), 4.95 (s, 2H, CH2), 3.07–2.99 (m, 4H. CH2-CH2); 13C NMR (125 MHz, DMSO-d6) δ 176.51 (C=O), 155.77 (S-C=N of triazole), 154.40 (C=N), 142.15 (C=N of triazole), 138.94, 136.81, 132.55, 129.64, 127.75, 126.68, 124.54, 124.30, 122.70, 122.18, 119.74, 118.86, 113.58, 112.50, 11.97, 110.91, 110.15, 53.85 (S-CH2), 25.04 (CH2), 22.91 (CH2); m/z: 519 (M+) for C29H25N7OS (M.wt 519.63).
2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-(4-chloro-phenyl)acetamide (5b)
Green ppt, m.p. 89-91 °C; yield: %; IR (cm−1) 3260, 3225, 3165 (NH), 1685 (C=O), 1630 (C=N), 1591 (C=C); ;1H NMR (500 MHz, DMSO-d6) δ 12.08, 11.51, 10.75 (s, 3H, 3NH), 8.64 (s, 1H, CH=N), 8.15 (d, 1H, indolyl H-4`), 8.00 (s, 1H, indolyl H-2`), 7.81–6.83 (m, 12H, Ar–H), 4.15 (s, 2H, CH2), 3.08–2.99 (m, 4H, CH2-CH2); 13C NMR (125 MHz, DMSO-d6) δ 168.75 (C=O), 154.39 (C=N of triazole), 141.46 (C=N), 129.54, 128.09, 122.81, 121.47, 119.62, 118.81, 118.74, 113.85, 112.16, 111.87, 54.20 (S-CH2), 25.45 (CH2), 23.14 (CH2); m/z: 588/590 (M+/M+ + 2) for C29H23Cl2N7OS (M.wt 588.51).
2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-(4-fluoro-phenyl)acetamide (5c)
Ppt, m.p. °C; yield: %; for C29H24FN7OS (537.62); IR (cm−1) 3387, 3226 (NH), 1664 (C=O), 1617 (C=N), 1595 (C=C); 1H NMR (500 MHz, DMSO-d6) δ 12.02, 10.76 (s, 3H, 3NH), 9.52 (s, 1H, CH=N), 8.18 (d, 1H, indolyl H-4`), 8.08 (s, 1H, indolyl H-2`), 7.50–711 (m, 12H, Ar–H), 4.09 (s, 2H, CH2), 3.08–3.03 (m, 4H. CH2-CH2); 13C NMR (125 MHz, DMSO-d6) 168.65 (C=O), 154.36 (S-C=N of triazole), 141.61 (C=N), 136.51(C=N), 133.95, 130.90, 129.58, 127.83, 126.54, 123.98, 122.22, 119.67, 118.37, 113.47, 112.47, 110.71, 54.08 (S-CH2), 25.22 (CH2), 22.89 (CH2).
2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-(4-methoxyphenyl)acetamide (5d)
Pale green ppt, m.p. 75–7 °C; yield: %; for C30H27N7O2S (549.65) IR (cm−1) 3267, 3187, 3115 (NH), 1680 (C=O), 1631 (C=N), 1591 (C=C); 1H NMR (500 MHz, DMSO-d6) δ 12.02, 10.76 (s, 3H, 3NH), 9.52 (s, 1H, CH=N), 8.18 (d, 1H, indolyl H-4`), 8.08 (s, 1H, indolyl H-2`), 7.50–6.79 (m, 12H, Ar–H), 4.13 (s, 2H, CH2), 3.35 (s, 3H, OCH3), 3.08–3.03 (m, 4H. CH2-CH2); 13C NMR (125 MHz, DMSO-d6) δ 167.18 (C=O), 156.36 (C=N of triazole), 141.69 (C=N), 137.06, 136.03, 132.21, 127.81, 126.78, 124.21, 122.96, 122.17, 121.42, 119.58, 118.86, 114.49, 113.93, 112.11, 111.89, 110.85, 110.34, 55.84 (OCH3), 54.02 (CH2), 25.24 (CH2), 23.21 (CH2).
2-((5-(2-(1H-indol-3-yl)ethyl)-4-(((1H-indol-3-yl)methylene)amino)-4H-1,2,4-triazol-3-yl)thio)-N-(4-(N-(pyridin-2-yl)sulfamoyl)phenyl)acetamide (5e)
Brown ppt, m.p. 180–2 °C; yield: %; IR (cm−1) 3260, 3225, 3202, 3182 (NH), 1664 (C=O), 1630 (C=N), 1591 (C=C); 1H NMR (500 MHz, DMSO-d6) δ 12.78, 10.69 (s, 4H, 4NH), 8.64 (s, 1H, CH=N), 8.12–6.71 (m, 18H, Ar–H), 4.14 (s, 2H, CH2), 3.08–3.01 (m, 4H. CH2-CH2); 13CNMR (125 MHz, DMSO-d6) δ 166.86 (C=O), 163.82 (S-C=N of triazole), 155.55 (C=N), 153.10 (C=N of pyridine), 145.13 (C=N), 139.63, 137.94, 136.74, 128.20, 122.35, 121.42, 119.12, 118.71, 114.18, 111.91, 56.97 (S-CH2), 26.55 (CH2), 23.06 (CH2); m/z: 675 (M+) for C34H29N9O3S2 (675.79).
Biological activities
Ethical approval
All care and procedures for cell culture experiments and the in vivo studies used in this study were performed in accordance with relevant guidelines and regulations of the Declaration of Helsinki, and in accordance with ARRIVE guidelines. Approval was granted by the Medical Research Ethics Committee (MREC) of the NRC, Egypt (Approval no. 13060130).
Cytotoxicity (MTT) assay
The adenocarcinoma breast cancer cell lines MDA-MB-231 and MCF-7 were purchased from the American Type Culture Collection (ATCC, Rockville, MD, USA), and the normal human skin fibroblast (hFB) cell line was obtained from the Banco de Células de Rio de Janeiro (BCRJ, Brazil). The tested cell lines were cultured in RPMI-1640 medium (MCF-7 and hFB cells) or DMEM (MDA-MB-231 cells) containing 10% fetal bovine serum (heat-inactivated), supplemented with 5% antibiotics (penicillin and streptomycin), and incubated in a humidified air chamber with 5% CO2. The cell lines were propagated in cell culture flasks until the monolayer reached 85% confluency. Subsequently, the cells were sub-cultured in 96-well plates at a density of 5,000 cells/ well and treated with the synthesized compounds at variable concentrations for different time intervals (24 h and 48 h). The cytotoxic activity was estimated using the 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay as described by Hansen et al.47.
Briefly, after the treatment period, 20 µL of MTT solution (5 mg/mL in PBS) was added to each well and incubated for 4 h at 37 °C. The medium was then carefully removed, and the resulting insoluble formazan crystals were dissolved in 100 µL of DMSO, and the absorbance was measured at 570 nm using a microplate reader. The data were expressed as the mean percentage of viable cells as compared to the respective control cultures treated with the solvent from three independent experiments (n = 3). The half maximal growth inhibitory concentration IC50 values were calculated from the line equation of the dose–dependent curve of each compound.
Evaluation of cell migration, chemotaxis, and invasion capacity using the cell migration assay
In this protocol, we used the standard methodology for wound healing assay using live-cell microscopy as described by48.
Sample preparation: creating the gap
A cell-free gap was created in a cell monolayer by physical exclusion, as a “cell exclusion” collective cell migration assay, to simulate wounding and to create a gap by scratching a confluent monolayer with a pipette tip. The cultured plates were washed twice with phosphate buffer saline to remove debris from damaged or dead cells, particularly after physical scratching. Then they were incubated with complete media containing the tested chemicals for different time intervals 24, and 48h. The cultured plates were rinsed twice with phosphate-buffered saline, and image acquisition is started at each related time point.
Image processing and analysis
Once the digital images were recorded, the gap size was measured as a function of time using Olympus microscopy with a digital camera (PowerShot A20, Canon, USA).
Having measured the gap area for each frame in the wound healing experiment, we plotted the gap area as a function of time to derive the cell migration rate v migration and also the t1/2gap value according to48.
In addition, by plotting the gap area versus time, the cell migration rate was readily extrapolated from the data well before the gap had fully closed in, and the actual cell migration rate was calculated according to real-world units of µm2/hr.
Cell cycle analysis
Cell cycle analysis was performed according to Nunez 49. MDA-MB-231 (TNBC) and MCF-7 (non-TNBC) cells were cultured in DMEM HG or RPMI, respectively, supplemented with 100 U/mL penicillin, 100 mg/mL streptomycin, and 10% heat-inactivated FBS at 37 °C in a humidified atmosphere containing 5% CO₂. After 48h treatment with test compounds, 1 × 105 cells were trypsinized, washed twice with ice-cold PBS (pH 7.4), fixed in 60% ice-cold ethanol (4 °C, 1h), then stained with RNase A (50 µg/mL) and propidium iodide (10 µg/mL) for 20 min at 37 °C in the dark. DNA content was analyzed by flow cytometry (FL2 channel, λex/em 535/617 nm; ACEA Novocyte™, ACEA Biosciences), acquiring 12,000 events/sample, with distribution calculated using NovoExpress™ software. All experiments were performed in three independent biological replicates (n = 3), each derived from separate cell cultures and conducted on different days. One sample per treatment condition was analyzed within each biological replicate.
Apoptosis and necrosis analysis
After treatment with the TNBC and non-TNBC with the test compounds for 48h, apoptosis and necrosis cell populations are determined using the Annexin V-FITC apoptosis detection kit (Abcam Inc., Cambridge Science Park, Cambridge, UK) according to the manufacturer’s instructions. Briefly, 1 × 105 cells were collected by trypsinization and washed twice with ice-cold PBS (pH 7.4). Then, cells were incubated in the dark with 0.5 ml of Annexin V-FITC/PI solution for 30 min at room temperature. After staining, cells are injected via ACEA Novocyte™ flowcytometer (ACEA Biosciences Inc., San Diego, CA, USA) and analyzed for FITC and PI fluorescent signals using FL1 and FL2 signal detectors, respectively (λex/em 488/530 nm for FITC and λex/em 535/617 nm for PI). A minimum of 12,000 events were acquired per sample.
Initial gating was performed on forward scatter (FSC) versus side scatter (SSC) plots to exclude cellular debris and aggregates. Doublet discrimination was applied using FSC-area versus FSC-height parameters to ensure analysis of single-cell populations only. Quadrant gates were subsequently established on Annexin V-FITC versus PI dot plots using unstained, single-stained, and compensation controls. Cell populations were defined as follows:
-
Annexin V − /PI − : viable cells
-
Annexin V + /PI − : early apoptotic cells
-
Annexin V + /PI + : late apoptosis/secondary necrosis
-
Annexin V − /PI + : primary necrotic cells
All apoptosis experiments were conducted in three independent biological replicates (n = 3) performed on separate days from independently cultured cells.
Real-time quantitative PCR for gene expression
Total RNA was isolated from treated and untreated MDA-MB-231 and MCF-7 cell lines using a total RNA purification kit (NORGEN, Canada) following the manufacturer’s instructions. RNA concentration and purity were checked using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, USA). cDNA was reverse-transcribed from 1 μg of extracted RNA using Revert Aid First Strand cDNA synthesis Kit (Thermo Fisher Scientific, Lithuania). Quantitative real time-PCR reaction was performed in an ABI 7500 fast PCR system (Applied Biosystems, USA) using Hera plus SYBR Green qPCR kit (Willow fort, Birmingham, UK). The primers of the endogenous control gene (GAPDH) and the target genes were designed using net primer software (https://www.premierbiosoft.com/netprimer/), NCBI primer blast software (https://www.ncbi.nlm.nih.gov/tools/primer-blast), and UCSC software (https://genome.ucsc.edu/cgi-bin/hgPcr). Designed primers were purchased from willowfort.co.uk. and listed in Table 10. Melting curve analysis was done to ensure the specificity of the amplification product. The ΔCT method was used for gene expression calculations35,50. Experiments were performed with three independent biological replicates (n = 3) per condition for each cell line. Each biological replicate corresponded to an independently seeded and treated cell culture which was processed independently through RNA extraction, cDNA synthesis, and qPCR measurement51. Results are presented as mean value ± SEM.
ELISA analysis of FAK expression
Focal adhesion kinase (FAK) expression, in the MDA-MB-231 cell line without and after treatment with compounds 2, 3c, or 4c, was quantified using the Human SimpleStep ELISA® Kit (Abcam, Cambridge, UK; catalog no. ab187395) following the manufacturer’s protocol22. Briefly, Samples or standard serials were incubated for 1 h with the supplied antibody cocktail at room temperature on a shaker set to 400 rpm. The ELISA plates were then washed extensively to remove unconjugated antibodies. Then, 3, 3’, 5, 5’-Tetramethylbenzidine (TMB) substrate was added for 10 min in the dark, then the reaction was stopped, and absorbance was measured at 450 nm. The average zero standard was subtracted from all readings. Duplicate readings of the positive control dilutions were averaged and plotted against their concentrations. A smooth curve was fitted through these points to construct the standard curve. Protein concentrations of unknown samples were then calculated based on this standard curve.
In vivo acute toxicity study
Chemicals and reagents
Biochemical analyses of aspartate aminotransferase (AST), malondialdehyde (MDA), and Nitric oxide (NO) were conducted using commercially available colorimetric kits from Biodignostic Chemical Company, Egypt. The ELISA kit for Kidney Injury Molecule-1 (KIM-1) was purchased from CUSABIO (catalog No: CCSB-E08808r).
Animal source and care
Albino white mice weighing between 25 and 30 g were sourced from the animal house of the National Research Centre (NRC), Egypt. The animals were housed in the animal house of NRC. The mice were kept in cages and maintained under standard conditions (12:12 h light/dark cycle, controlled room temperature (23 ± 2 °C), stress-free, ad libitum water, standard diets, and sanitary conditions). Before commencing the experiment, the mice were allowed to acclimatize for a period of one week to reduce stress.
Acute toxicity study
To evaluate the toxicity of the synthesized compound 4c, an in vivo toxicity assay was conducted by treating mice with a single intraperitoneal (i.p.) injection of three different concentrations of this compound, and then biochemical and histopathological analyses following treatment were performed. The control group was administered solely with the vehicle, DMSO, while the other three groups of mice received a single i.p. dose of increasing concentrations of 4c (12.5, 25, and 50 mg/kg/day) dissolved in the vehicle, DMSO.
Experimental design
Sixteen albino mice (n = 4) were used and allocated randomly into four different groups:
Group (1): Negative control mice receiving a single intraperitoneal (i.p.) dose of the vehicle, DMSO.
Group (2): (12.5 mg/kg 4c received group) Mice received a single i.p. dose of 12.5 mg/kg of 4c.
Group (3): (25 mg/kg 4c received group) Mice received a single i.p. dose of 25 mg/kg of compound 4c.
Group (4): (50 mg/kg 4c received group) Mice received a single i.p. dose of 50 mg/kg of compound 4c.
The mice were observed for the next 48 h, for morphological features, behavioral alterations, or mortality and other signs of toxicity.
Collection and preparation of blood and tissue samples
At the end of the experiment, Thiopental sodium (50 mg/kg, i.p., Thiopental®, Biochemie GmbH, Vienna, Austria) was used to anesthetize mice that were fasted overnight, with free access to water; then blood was collected from the orbital plexus, and the anesthetized mice were sacrificed by cervical dislocation. Blood samples were centrifuged at 3000 rpm and stored at -80 °C for further biochemical examinations. Under deep anaesthesia, the mice were slaughtered and sacrificed by cervical dislocation. Livers and kidneys were rapidly excised, cleaned, dissected, and washed with normal saline. Tissue homogenates were prepared by homogenization of the liver and kidney samples in cold Phosphate-buffered saline (PBS; pH 7.4). Subsequently, homogenates were centrifuged (10,000×g for 20 min at 4 °C), and the clear supernatants were quickly collected and stored at -80 °C until use. Liver and kidney samples (n = 3) were specified for biochemical analysis of oxidative stress.
Biochemical investigations
Biochemical assays in mice serum and tissue homogenates were carried out on a UV/Vis Spectrophotometer (Aglient Technologies, India). To evaluate hepatic function, serum levels of aminotransferase (AST) was estimated52. To evaluate renal function, serum levels of kidney-injury molecule-1 (Kim-1) was measured colorimetrically using the ELISA technique.
Evaluation of oxidative stress status in the hepatic and renal tissues
Lipid peroxidation (LPO) Lipid peroxidation in the hepatic and renal tissues was estimated colorimetrically, according to Ohkawa et al.53. Lipid peroxidation levels were estimated by measuring thiobarbituric acid reactive substances (TBARS) as malondialdehyde (MDA), a product of lipid peroxidation. The reaction between thiobarbituric acid and MDA, in acidic medium, resulted in pink colored end-product that was measured spectrophotometrically at 534 nm. The results were expressed as nmol/g of wet tissue.
Determination of NO content Nitric oxide (NO) production in the hepatic and renal tissues was determined by estimating the total nitrite level, which is determined by reaction with Griess reagent using sodium nitrite as a standard according to Montgomery and Dymock54, spectrophotometrically at 540 nm. The data were expressed as µmol/g of wet tissue.
Histopathological investigation
Samples of liver and kidneys were collected from all groups, immediately fixed in 10% neutral buffered formalin, and routinely processed according to Suvarna et al.55. Sections of 4-5μm thickness were prepared and stained with Hematoxylin and Eosin (H&E) for histopathological investigation by a light microscope (Olympus BX50, Tokyo, Japan). Signs of organ tissue damage are recorded56. The histology of each organ was compared with that of the negative control organ.
Statistical analysis
Data are presented mean ± standard error of the mean (SEM) from three independent experiments. Statistical analyses were conducted using GraphPad Prism version 8.0.2 or SPSS version 20. The normality of data distribution was evaluated with the Shapiro–Wilk test. For pairwise comparisons, Welch’s t-test was applied to parametric data, while the Mann–Whitney test was utilized for nonparametric data. Multiple group comparisons were performed using one-way analysis of variance (ANOVA) followed by Tukey’s, or Bonferroni post hoc tests. A p-value less than 0.05 was considered statistically significant.
In-silico studies
Molecular Docking
Molecular docking was done using PyRx version 8 AutoDock Vina57. Focal Adhesion Kinase (PDB ID: 2JKK) three-dimensional structures were obtained from the RCSB protein data bank (https://www.rcsb.org/access on 25 September 2025)58. The unwanted co-crystallized ligands and water molecules were removed, and the enzymes were prepared using VEGA ZZ, saved as pdb and converted to PDBQT format by Autodock vina tools. The chemical structure of co-crystalline ligand, 2-({5-chloro-2-[(2-methoxy-4-morpholin-4-ylphenyl)amino]pyrimidin-4-yl}amino)-n-methylbenzamide (BI9) was obtained from the PubChem database as an SDF file. While the structure of the specified molecules was constructed with ChemDraw ultra-10.0 and saved as an SDF files. After that, all the SDF files were minimized utilizing the MMFF94 force field, then converted to a pdbqt file by OpenBable tools in pyrx software. AutoDock Tools establish the grid box size and center. The FAK active site was centered at x = 8.28, y = 0.137, and z = 26.55, with dimensions of 19.01 × 18.56 × 24.12 Å. The co-crystalline ligand was re-docked into the enzyme active sites to verify our docking protocol. PyRx software presents the 8 most suitable docking poses of the ligand–protein complex after the docking is completed and subsequently ranked according to the binding energy. We have selected the first docking pose, which is the most suitable pose where the ligands have the lowest binding energy, zero root-mean-square deviation (RMSD) and strongly interact with the protein’s catalytic cavity and visualized them using BIOVIA Discovery Studio Visualizer to have a great insight into ligand binding position in the protein cavity.
Drug likeness and ADMET prediction
Drug likeness and some ADMET endpoints of the most active compounds, 3c, and 4c, were predicted utilizing the SwissADME tool, accessible at http://www.swissadme.ch/index.php and pkCSM website (http://biosig.unimelb.edu.au/pkcsm/)59,60. Drug likeness analyses were performed on the computed physicochemical descriptors, such as molecular MW, LogP, HBA, HBD, nRB, MR, and TPSA, using Lipinski’s rule of five and the Veber filter rule in consideration41,42.
Data availability
The IR, NMR and Mass spectra are available in the supporting information file.
References
Zhang, J. et al. Natural products and derivatives for breast cancer treatment: From drug discovery to molecular mechanism. Phytomedicine 129, 155600. https://doi.org/10.1016/j.phymed.2024.155600 (2024).
Arnold, M. et al. Current and future burden of breast cancer: Global statistics for 2020 and 2040. Breast 66, 15–23. https://doi.org/10.1016/j.breast.2022.08.010 (2022).
Morris, G. J. et al. Differences in breast carcinoma characteristics in newly diagnosed African-American and Caucasian patients. Cancer 110, 876–884. https://doi.org/10.1002/cncr.22836 (2007).
Zagami, P. & Carey, L. A. Triple negative breast cancer: Pitfalls and progress. NPJ Breast Cancer 8, 95. https://doi.org/10.1038/s41523-022-00468-0 (2022).
Almansour, N. M. Triple-Negative breast cancer: A brief review about epidemiology, risk factors, signaling pathways, treatment and role of artificial intelligence. Front. Mol. Biosci. https://doi.org/10.3389/fmolb.2022.836417 (2022).
Yin, L., Duan, J.-J., Bian, X.-W. & Yu, S.-C. Triple-negative breast cancer molecular subtyping and treatment progress.. Breast Cancer Res. 22, 61. https://doi.org/10.1186/s13058-020-01296-5 (2020).
Lehmann, B. D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767. https://doi.org/10.1172/JCI45014 (2011).
Espinosa Fernandez, J. R. et al. Identification of triple-negative breast cancer cell lines classified under the same molecular subtype using different molecular characterization techniques: Implications for translational research. PLoS ONE 15, e0231953. https://doi.org/10.1371/journal.pone.0231953 (2020).
Yin, L., Duan, J.-J., Bian, X.-W. & Yu, S. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 22, 61. https://doi.org/10.1186/s13058-020-01296-5 (2020).
Ge, W. et al. Synthesis and structure-activity relationship studies of parthenolide derivatives as potential anti-triple negative breast cancer agents. Eur. J. Med. Chem. 166, 445–469. https://doi.org/10.1016/j.ejmech.2019.01.058 (2019).
Dass, S. A. et al. Triple negative breast cancer: A review of present and future diagnostic modalities. Medicina 57, 62. https://doi.org/10.3390/medicina57010062 (2021).
Yang, F. et al. Discovery of novel chloropyramine-cinnamic acid hybrids as potential FAK inhibitors for intervention of metastatic triple-negative breast cancer. Bioorg. Med. Chem. 66, 116809. https://doi.org/10.1016/j.bmc.2022.116809 (2022).
Lu, Y. & Sun, H. Progress in the development of small molecular inhibitors of focal adhesion kinase (FAK). J. Med. Chem. 63, 14382–14403. https://doi.org/10.1021/acs.jmedchem.0c01248 (2020).
Zhou, J., Yi, Q. & Tang, L. The roles of nuclear focal adhesion kinase (FAK) on cancer: A focused review. J. Exp. Clin. Cancer Res. 38, 250. https://doi.org/10.1186/s13046-019-1265-1 (2019).
Jia, Y. et al. Thiophenol-formaldehyde triazole causes apoptosis induction in ovary cancer cells and prevents tumor growth formation in mice model. Eur. J. Med. Chem. 172, 62–70. https://doi.org/10.1016/j.ejmech.2019.03.033 (2019).
Altıntop, M. D. et al. Design, synthesis, in vitro and in silico evaluation of a new series of oxadiazole-based anticancer agents as potential Akt and FAK inhibitors. Eur. J. Med. Chem. 155, 905–924. https://doi.org/10.1016/j.ejmech.2018.06.049 (2018).
Yang, X.-H. et al. Synthesis, biological evaluation, and molecular docking studies of 1,3,4-thiadiazol-2-amide derivatives as novel anticancer agents. Bioorg. Med. Chem. 20, 2789–2795. https://doi.org/10.1016/j.bmc.2012.03.040 (2012).
Mustafa, M. et al. Synthesis, antiproliferative, anti-tubulin activity, and docking study of new 1,2,4-triazoles as potential combretastatin analogues. Eur. J. Med. Chem. 141, 293–305. https://doi.org/10.1016/j.ejmech.2017.09.063 (2017).
Mustafa, M., Anwar, S., Elgamal, F., Ahmed, E. R. & Aly, O. M. Potent combretastatin A-4 analogs containing 1,2,4-triazole: Synthesis, antiproliferative, anti-tubulin activity, and docking study. Eur. J. Med. Chem. 183, 111697. https://doi.org/10.1016/j.ejmech.2019.111697 (2019).
Elrashedy, A., Ibrahim, N. E., Abo-Salem, H., Elaasser, M. M. & El-Sawy, E. R. Design, synthesis, and molecular modeling of new 1,2,4-triazole-containing indole compounds as aromatase antagonists for the treatment of breast cancer. Bioorg. Chem. 163, 108677. https://doi.org/10.1016/j.bioorg.2025.108677 (2025).
Malik, M. S., Ahmed, S. A., Althagafi, I. I., Ansari, M. A. & Kamal, A. Application of triazoles as bioisosteres and linkers in the development of microtubule targeting agents. RSC Med. Chem. 11, 327–348. https://doi.org/10.1039/C9MD00458K (2020).
Mustafa, M. et al. Discovery of antiproliferative and anti-FAK inhibitory activity of 1,2,4-triazole derivatives containing acetamido carboxylic acid skeleton. Bioorg. Med. Chem. Lett. 40, 127965. https://doi.org/10.1016/j.bmcl.2021.127965 (2021).
Kumar, S. & Ritika,. A brief review of the biological potential of indole derivatives. Futur. J. Pharm. Sci. 6, 121. https://doi.org/10.1186/s43094-020-00141-y (2020).
Yang, F. et al. Discovery of new indole-based 1,2,4-triazole derivatives as potent tubulin polymerization inhibitors with anticancer activity. New J. Chem. 45, 21869–21880. https://doi.org/10.1039/D1NJ03892C (2021).
Yele, V. et al. Synthesis and preclinical evaluation of indole triazole conjugates as microtubule targeting agents that are effective against MCF-7 breast cancer cell lines. Anticancer Agents Med. Chem. 21, 1047–1055. https://doi.org/10.2174/1871520620666200925102940 (2021).
Zahoor, A. F. et al. Synthesis, cytotoxic, and computational screening of some novel indole–1,2,4-triazole-based S-alkylated N-aryl acetamides. Biomedicines 11, 3078. https://doi.org/10.3390/biomedicines11113078 (2023).
Abo-Salem, H. M., Gibriel, A. A., El Awady, M. E. & Mandour, A. H. Synthesis, molecular docking and biological evaluation of novel flavone derivatives as potential anticancer agents targeting Akt. Med. Chem. 17, 158–170. https://doi.org/10.2174/1573406416666200306115035 (2020).
Fawzy, N. M., Ahmed, K. M., Abo-Salem, H. M. & Aly, M. S. Novel furochromone derivatives of potential anticancer activity targeting EGFR tyrosine kinase. Synthesis and molecular docking study. Russ. J. Bioorg. Chem. 48, 749–767. https://doi.org/10.1134/S1068162022040082 (2022).
Abo-Salem, H. M. et al. Synthesis, bioactivity assessment, molecular docking and ADMET studies of new chromone congeners exhibiting potent anticancer activity. Sci. Rep. 14, 9636. https://doi.org/10.1038/s41598-024-59606-2 (2024).
Abdelmegeed, H., Abo-Salem, H. M., Zayed, E. M. & El-Sawy, E. R. Anti colorectal cancer activity and in silico studies of novel pyridine nortopsentin analog as cyclin dependent kinase 6 inhibitor. Sci. Rep. 14, 26327. https://doi.org/10.1038/s41598-024-75411-3 (2024).
Yakkala, P. A. et al. Synthesis and cytotoxic activity of 1,2,4-triazolo-linked bis-indolyl conjugates as dual inhibitors of tankyrase and PI3K. Molecules 27, 7642. https://doi.org/10.3390/molecules27217642 (2022).
Hu, H.-H. et al. Roles and inhibitors of FAK in cancer: Current advances and future directions. Front. Pharmacol. https://doi.org/10.3389/fphar.2024.1274209 (2024).
Chen, T. et al. Design, synthesis and biological evaluation of novel FAK inhibitors with better selectivity over IR than TAE226. Bioorg. Chem. 124, 105790. https://doi.org/10.1016/j.bioorg.2022.105790 (2022).
Rachmi, E., Purnomo, B. B., Endharti, A. T. & Fitri, L. E. Afzelin inhibits migration of MDA-MB-231 cells by suppressing FAK expression and Rac1 activation. J. Appl. Pharm. Sci. 10, 77–82. https://doi.org/10.7324/JAPS.2020.101010 (2020).
Schlaepfer, D. D., Ojalill, M. & Stupack, D. G. Focal adhesion kinase signaling – tumor vulnerabilities and clinical opportunities. J. Cell Sci. https://doi.org/10.1242/jcs.261723 (2024).
Vaniotis, G. et al. Collagen IV-conveyed signals can regulate chemokine production and promote liver metastasis. Oncogene 37, 3790–3805. https://doi.org/10.1038/s41388-018-0242-z (2018).
An, G. et al. Effects of CCL5 on the biological behavior of breast cancer and the mechanisms of its interaction with tumor-associated macrophages. Oncol. Rep. https://doi.org/10.3892/or.2019.7344 (2019).
Jin, L. et al. FAK-induced time dependent cell response profiling: Prediction, identification, and analysis of anti-tumor immune natural products. Drug Des. Devel. Ther. 19, 9079–9097. https://doi.org/10.2147/DDDT.S533551 (2025).
Singh, S. K. et al. CCR5/CCL5 axis interaction promotes migratory and invasiveness of pancreatic cancer cells. Sci. Rep. 8, 1323. https://doi.org/10.1038/s41598-018-19643-0 (2018).
Masuyama, M. et al. Targeting focal adhesion kinase inhibits cell migration and non-angiogenic vascularization in malignant breast cancer. Breast Cancer 33, 188–199. https://doi.org/10.1007/s12282-025-01792-6 (2026).
Lipinski, C. A. Lead- and drug-like compounds: The rule-of-five revolution. Drug Discov. Today Technol. 1, 337–341. https://doi.org/10.1016/j.ddtec.2004.11.007 (2004).
Veber, D. F. et al. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 45, 2615–2623. https://doi.org/10.1021/jm020017n (2002).
Sulzmaier, F. J., Jean, C. & Schlaepfer, D. D. FAK in cancer: Mechanistic findings and clinical applications. Nat. Rev. Cancer 14, 598–610. https://doi.org/10.1038/nrc3792 (2014).
Zhang, J., He, D.-H., Zajac-Kaye, M. & Hochwald, S. N. A small molecule FAK kinase inhibitor, GSK2256098, inhibits growth and survival of pancreatic ductal adenocarcinoma cells. Cell Cycle 13, 3143–3149. https://doi.org/10.4161/15384101.2014.949550 (2014).
Link, W. et al. Chemical interrogation of FOXO3a nuclear translocation identifies potent and selective inhibitors of phosphoinositide 3-kinases. J. Biol. Chem. 284, 28392–28400. https://doi.org/10.1074/jbc.M109.038984 (2009).
Sevaille, L. et al. 1,2,4‐triazole‐3‐thione compounds as inhibitors of dizinc metallo‐β‐lactamases. ChemMedChem 12, 972–985. https://doi.org/10.1002/cmdc.201700186 (2017).
Hansen, M. B., Nielsen, S. E. & Berg, K. Re-examination and further development of a precise and rapid dye method for measuring cell growth/cell kill. J. Immunol. Methods 119, 203–210. https://doi.org/10.1016/0022-1759(89)90397-9 (1989).
Jonkman, J. E. N. et al. An introduction to the wound healing assay using live-cell microscopy. Cell Adh. Migr. 8, 440–451. https://doi.org/10.4161/cam.36224 (2014).
Nunez, R. DNA measurement and cell cycle analysis by flow cytometry. Curr. Issues Mol. Biol. 3, 67–70 (2001).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108. https://doi.org/10.1038/nprot.2008.73 (2008).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Reitman, S. & Frankel, S. A colorimetric method for the determination of serum glutamic oxalacetic and glutamic pyruvic transaminases. Am. J. Clin. Pathol. 28, 56–63. https://doi.org/10.1093/ajcp/28.1.56 (1957).
Ohkawa, H., Ohishi, N. & Yagi, K. Assay for lipid peroxides in animal tissues by thiobarbituric acid reaction. Anal. Biochem. 95, 351–358. https://doi.org/10.1016/0003-2697(79)90738-3 (1979).
Montgomery, H. A. C. & Dymock, J. F. Nitric oxide assay. Analyst 86, 414 (1961).
Suvarna, S. K., Layton, C. & Bancroft, J. D. Bancroft’s Theory and Practice of Histological Techniques (Churchill Livingstone Elsevier, 2019).
Bancroft, J. & Stevens, A. Theory and Practice of Histological Techniques 44th edn. (Churchil Livingstone, 1996).
Trott, O. & Olson, A. J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461. https://doi.org/10.1002/jcc.21334 (2010).
Lietha, D. & Eck, M. J. Crystal structures of the FAK kinase in complex with TAE226 and related bis-anilino pyrimidine inhibitors reveal a helical DFG conformation. PLoS ONE 3, e3800. https://doi.org/10.1371/journal.pone.0003800 (2008).
Daina, A., Michielin, O. & Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7, 42717. https://doi.org/10.1038/srep42717 (2017).
Pires, D. E. V., Blundell, T. L. & Ascher, D. B. pkCSM: Predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J. Med. Chem. 58, 4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104 (2015).
Acknowledgements
The authors express their gratitude to Dr. Esam R. Ahmed, VACSERA, Cairo, Egypt, for performing the Focal Adhesion Kinase (FAK) assay.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was supported by the National Research Centre (NRC), Egypt, under research project No. 13060130.
Author information
Authors and Affiliations
Contributions
H.A.A-S.: conceptualization, Synthesis, data analyses; N.A-S.: contributed to cell culture (MDA, MCF-7, and human fibroblast cells), primer design, RNA extraction, cDNA synthesis, real time gene expression analysis, Statistical analysis of gene expression, cell cycle and apoptosis results, revision; G.I.F. performing in vivo toxicity assay experiment, writing, revision; M.M. contributed to biological activities analyses and data interpretation, mainly cell cycle, annexin V/PI and gene expression analyses, writing and revision; E.A.M.: Synthesis, data analyses; M.A.M.A.: preforming cytotoxicity and wound healing assay, writing, review; H.M. A-S.: supervision, conceptualization, synthesis, Data curation, in-silico study, writing of the original draft, review and editing; R.A.M.A.: conceptualization, Methodology, generation of data, interpreting and analysis the results. All authors have read and agreed to publish the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abd El Salam, H.A., Abu-Shahba, N., Ibrahim Fouad, G. et al. New indole-linked 1,2,4-triazole derivatives as dual FAK inhibitors and apoptosis inducers targeting survival and migration in triple-negative breast cancer in-vitro. Sci Rep 16, 13134 (2026). https://doi.org/10.1038/s41598-026-41032-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-41032-1