Abstract
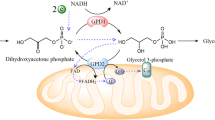
Dihydroxyacetone phosphate (DHAP), glycerol-3-phosphate (Gro3P) and reduced/oxidized nicotinamide adenine dinucleotide (NADH/NAD⁺) are key metabolites of the Gro3P shuttle, which transfers reducing equivalents between the cytosol and mitochondria. Targeted activation of Gro3P biosynthesis has recently emerged as a promising strategy to alleviate reductive stress. However, because Gro3P constitutes the backbone of triglycerides, its accumulation can promote extensive lipogenesis. Here we show that a genetically encoded tool based on a di-domain glycerol-3-phosphate dehydrogenase from the alga Chlamydomonas reinhardtii (CrGPDH) effectively operates both the alternative Gro3P shunt, which regenerates NAD⁺ while converting DHAP to Gro3P, and the glycerol shunt, which converts Gro3P to glycerol and inorganic phosphate, across transformed and primary mammalian cell cultures as well as mouse liver. CrGPDH expression supported proliferation of cancer cells under respiratory chain inhibition or hypoxia, as well as patient-derived fibroblasts with mitochondrial dysfunction. Moreover, CrGPDH decreased triglyceride levels in kidney cancer cell lines and reversed ethanol-induced triglyceride accumulation in mouse liver. Thus, CrGPDH represents a promising xenotopic tool to alleviate redox imbalance and associated impaired lipogenesis in conditions ranging from primary mitochondrial diseases to steatosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq data presented in this work are available at the Gene Expression Omnibus database under accession number GEO: GSE312066. The H. sapiens GRCh38 reference genome was used for mapping RNA-seq reads. Source data are provided with this paper.
Code availability
No new code was generated during this study.
References
Xiao, W., Wang, R. S., Handy, D. E. & Loscalzo, J. NAD(H) and NADP(H) redox couples and cellular energy metabolism. Antioxid. Redox Signal 28, 251–272 (2018).
Alonso-Lavin, A. J., Bajic, D. & Poyatos, J. F. Tolerance to NADH/NAD+ imbalance anticipates aging and anti-aging interventions. iScience 24, 102697 (2021).
Titov, D. V. et al. Complementation of mitochondrial electron transport chain by manipulation of the NAD+/NADH ratio. Science 352, 231–235 (2016).
Nelson, D. & Michael, M. Lehninger Principles of Biochemistry (W. H. Freeman, 2017).
Liu, S. et al. Glycerol-3-phosphate biosynthesis regenerates cytosolic NAD+ to alleviate mitochondrial disease. Cell Metab. 33, 1974–1987 (2021).
Xiao, W. & Loscalzo, J. Metabolic responses to reductive stress. Antioxid. Redox Signal 32, 1330–1347 (2020).
Pan, X. et al. A genetically encoded tool to increase cellular NADH/NAD+ ratio in living cells. Nat. Chem. Biol. 20, 594–604 (2024).
Yang, R., Guo, Z. & Li, B. NADH reductive stress drives metabolic reprogramming. Trends Cell Biol https://doi.org/10.1016/j.tcb.2025.07.005 (2025).
Yang, R. et al. Identification of purine biosynthesis as an NADH-sensing pathway to mediate energy stress. Nat. Commun. 13, 7031 (2022).
Weiss-Sadan, T. et al. NRF2 activation induces NADH-reductive stress, providing a metabolic vulnerability in lung cancer. Cell Metab. 35, 487–503 (2023).
Gu, L. et al. Cullin 3 RING E3 ligase inactivation causes NRF2-dependent NADH reductive stress, hepatic lipodystrophy, and systemic insulin resistance. Proc. Natl Acad. Sci. USA 121, e2320934121 (2024).
Goodman, R. P. et al. Hepatic NADH reductive stress underlies common variation in metabolic traits. Nature 583, 122–126 (2020).
Sharma, R. et al. Circulating markers of NADH-reductive stress correlate with mitochondrial disease severity. J. Clin. Invest. 131, e136055 (2021).
Wang, B. et al. Aldehyde dehydrogenase 1A1 increases NADH levels and promotes tumor growth via glutathione/dihydrolipoic acid-dependent NAD+ reduction. Oncotarget 8, 67043–67055 (2017).
Quintana, A., Kruse, S. E., Kapur, R. P., Sanz, E. & Palmiter, R. D. Complex I deficiency due to loss of Ndufs4 in the brain results in progressive encephalopathy resembling Leigh syndrome. Proc. Natl Acad. Sci. USA 107, 10996–11001 (2010).
Singh, C. et al. ChREBP is activated by reductive stress and mediates GCKR-associated metabolic traits. Cell Metab. 36, 144–158 (2024).
Li, H. et al. Lactate dehydrogenase and glycerol-3-phosphate dehydrogenase cooperatively regulate growth and carbohydrate metabolism during Drosophila melanogaster larval development. Development 146, dev175315 (2019).
Boudreau, A. et al. Metabolic plasticity underpins innate and acquired resistance to LDHA inhibition. Nat. Chem. Biol. 12, 779–786 (2016).
Zhai, X. et al. AMPK-regulated glycerol excretion maintains metabolic crosstalk between reductive and energetic stress. Nat. Cell Biol. 27, 141–153 (2025).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337 (2016).
Tighanimine, K. et al. A homoeostatic switch causing glycerol-3-phosphate and phosphoethanolamine accumulation triggers senescence by rewiring lipid metabolism. Nat. Metab. 6, 323–342 (2024).
Tiwari, V. et al. Glycerol-3-phosphate activates ChREBP, FGF21 transcription and lipogenesis in citrin deficiency. Nat. Metab. 7, 2284–2299 (2025).
Mugabo, Y. et al. Identification of a mammalian glycerol-3-phosphate phosphatase: role in metabolism and signaling in pancreatic beta-cells and hepatocytes. Proc. Natl Acad. Sci. USA 113, E430–E439 (2016).
Al-Mass, A. et al. Hepatic glycerol shunt and glycerol-3-phosphate phosphatase control liver metabolism and glucodetoxification under hyperglycemia. Mol. Metab. 66, 101609 (2022).
Al-Mass, A. et al. Glycerol-3-phosphate phosphatase operates a glycerol shunt in pancreatic beta-cells that controls insulin secretion and metabolic stress. Mol. Metab. 60, 101471 (2022).
Possik, E. et al. Glycerol 3-phosphate phosphatase/PGPH-2 counters metabolic stress and promotes healthy aging via a glycogen sensing-AMPK-HLH-30-autophagy axis in C. elegans. Nat. Commun. 14, 5214 (2023).
He, Q. et al. The unusual di-domain structure of Dunaliella salina glycerol-3-phosphate dehydrogenase enables direct conversion of dihydroxyacetone phosphate to glycerol. Plant J. 102, 153–164 (2020).
He, Y. et al. Cloning and characterization of two novel chloroplastic glycerol-3-phosphate dehydrogenases from Dunaliella viridis. Plant Mol. Biol. 71, 193–205 (2009).
Morales-Sanchez, D., Kim, Y., Terng, E. L., Peterson, L. & Cerutti, H. A multidomain enzyme, with glycerol-3-phosphate dehydrogenase and phosphatase activities, is involved in a chloroplastic pathway for glycerol synthesis in Chlamydomonas reinhardtii. Plant J. 90, 1079–1092 (2017).
Petrovic, D. et al. Ergothioneine improves healthspan of aged animals by enhancing cGPDH activity through CSE-dependent persulfidation. Cell Metab. 37, 542–556 (2025).
Parkhitko, A. A. & Cracan, V. Xenotopic synthetic biology: prospective tools for delaying aging and age-related diseases. Sci. Adv. 11, eadu1710 (2025).
Zhao, Y. et al. SoNar, a highly responsive NAD+/NADH sensor, allows high-throughput metabolic screening of anti-tumor agents. Cell Metab. 21, 777–789 (2015).
Tao, R. et al. Genetically encoded fluorescent sensors reveal dynamic regulation of NADPH metabolism. Nat. Methods 14, 720–728 (2017).
Cracan, V., Titov, D. V., Shen, H., Grabarek, Z. & Mootha, V. K. A genetically encoded tool for manipulation of NADP+/NADPH in living cells. Nat. Chem. Biol. 13, 1088–1095 (2017).
King, M. P. & Attardi, G. Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 246, 500–503 (1989).
Eales, K. L., Hollinshead, K. E. & Tennant, D. A. Hypoxia and metabolic adaptation of cancer cells. Oncogenesis 5, e190 (2016).
Garcia-Bermudez, J. et al. Aspartate is a limiting metabolite for cancer cell proliferation under hypoxia and in tumours. Nat. Cell Biol. 20, 775–781 (2018).
Kim, W. et al. Polyunsaturated fatty acid desaturation is a mechanism for glycolytic NAD+ recycling. Cell Metab. 29, 856–870 (2019).
Lister, A. et al. NRF2 regulates the glutamine transporter Slc38a3 (SNAT3) in kidney in response to metabolic acidosis. Sci. Rep. 8, 5629 (2018).
Nissen-Meyer, L. S. & Chaudhry, F. A. Protein kinase C phosphorylates the system N glutamine transporter SN1 (Slc38a3) and regulates its membrane trafficking and degradation. Front. Endocrinol. 4, 138 (2013).
Hagglund, M. G. et al. Identification of SLC38A7 (SNAT7) protein as a glutamine transporter expressed in neurons. J. Biol. Chem. 286, 20500–20511 (2011).
Chen, H. F., Chuang, H. C. & Tan, T. H. Regulation of dual-specificity phosphatase (DUSP) ubiquitination and protein stability. Int. J. Mol. Sci. 20, 2668 (2019).
Choe, M., Einav, T., Phillips, R. & Titov, D. V. Glycolysis model shows that allostery maintains high ATP and limits accumulation of intermediates. Biophys. J. 124, 1562–1586 (2025).
Zechner, C. & Rhee, E. P. Phosphate sensing in health and disease. Curr. Opin. Nephrol. Hypertens. 33, 361–367 (2024).
Zhou, W. et al. Kidney glycolysis serves as a mammalian phosphate sensor that maintains phosphate homeostasis. J. Clin. Invest. 133, e164610 (2023).
Simic, P. et al. Glycerol-3-phosphate contributes to the increase in FGF23 production in chronic kidney disease. Am. J. Physiol. Renal Physiol. 328, F165–F172 (2025).
Dambrova, M. et al. Acylcarnitines: nomenclature, biomarkers, therapeutic potential, drug targets, and clinical trials. Pharmacol. Rev. 74, 506–551 (2022).
Truby, L. K. et al. Circulating long chain acylcarnitines and outcomes in diabetic heart failure: an HF-ACTION clinical trial substudy. Cardiovasc. Diabetol. 20, 161 (2021).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 49, D1541–D1547 (2021).
Maiguel, D., Morita, M., Pei, Z., Jia, Z. & Watkins, P. A. ACSF2: a medium-chain acyl-coa synthetase with a potential role in neuronal differentiation. Preprint at bioRxiv https://doi.org/10.1101/2022.03.28.486105 (2022).
Chen, J. et al. ACSF2 and lysine lactylation contribute to renal tubule injury in diabetes. Diabetologia 67, 1429–1443 (2024).
Shen, H. et al. The human knockout gene CLYBL connects itaconate to vitamin B12. Cell 171, 771–782 (2017).
Liu, R. et al. Cellular pan-chain acyl-CoA profiling reveals SLC25A42/SLC25A16 in mitochondrial CoA import and metabolism. Nat Metab. 7, 1871–1888 (2025).
Sierra, A. Y. et al. CPT1c is localized in endoplasmic reticulum of neurons and has carnitine palmitoyltransferase activity. J. Biol. Chem. 283, 6878–6885 (2008).
Qi, X., Li, Q., Che, X., Wang, Q. & Wu, G. The uniqueness of clear cell renal cell carcinoma: summary of the process and abnormality of glucose metabolism and lipid metabolism in ccRCC. Front. Oncol. 11, 727778 (2021).
Pan, X. & Cracan, V. Translocation renal cell carcinoma says no to the Warburg effect. Nat. Metab. 7, 438–440 (2025).
Yao, C. H. et al. Uncoupled glycerol-3-phosphate shuttle in kidney cancer reveals that cytosolic GPD is essential to support lipid synthesis. Mol. Cell 83, 1340–1349 (2023).
Kayani, S. et al. SURF1 deficiency: expanding on disease phenotype and assessing disease burden by describing clinical and biochemical phenotype. Am. J. Med. Genet. A 197, e63947 (2025).
Zhu, Z. et al. SURF1, encoding a factor involved in the biogenesis of cytochrome c oxidase, is mutated in Leigh syndrome. Nat. Genet. 20, 337–343 (1998).
Sue, C. M. et al. Differential features of patients with mutations in two COX assembly genes, SURF1 and SCO2. Ann. Neurol. 47, 589–595 (2000).
Wedatilake, Y. et al. SURF1 deficiency: a multi-centre natural history study. Orphanet J. Rare Dis. 8, 96 (2013).
Choe, M. & Titov, D. V. Genetically encoded tools for measuring and manipulating metabolism. Nat. Chem. Biol. 18, 451–460 (2022).
Gohil, V. M. et al. Meclizine inhibits mitochondrial respiration through direct targeting of cytosolic phosphoethanolamine metabolism. J. Biol. Chem. 288, 35387–35395 (2013).
Shimano, H. & Sato, R. SREBP-regulated lipid metabolism: convergent physiology—divergent pathophysiology. Nat. Rev. Endocrinol. 13, 710–730 (2017).
Li, Z. et al. Cancer cells depend on environmental lipids for proliferation when electron acceptors are limited. Nat. Metab. 4, 711–723 (2022).
Pettit, F. H., Pelley, J. W. & Reed, L. J. Regulation of pyruvate dehydrogenase kinase and phosphatase by acetyl-CoA/CoA and NADH/NAD ratios. Biochem. Biophys. Res. Commun. 65, 575–582 (1975).
Oliveira, A. P. et al. Regulation of yeast central metabolism by enzyme phosphorylation. Mol. Syst. Biol. 8, 623 (2012).
Ozbalci, C., Storck, E. M. & Eggert, U. S. RNAi transfection results in lipidome changes. Proteomics 19, e1800298 (2019).
Qu, H. et al. Mitochondrial glycerol 3-phosphate dehydrogenase deficiency exacerbates lipotoxic cardiomyopathy. iScience 27, 109796 (2024).
Rusu, P. et al. GPD1 specifically marks dormant glioma stem cells with a distinct metabolic profile. Cell Stem Cell 25, 241–257 (2019).
Liu, R. et al. A HIF1alpha-GPD1 feedforward loop inhibits the progression of renal clear cell carcinoma via mitochondrial function and lipid metabolism. J. Exp. Clin. Cancer Res. 40, 188 (2021).
Zhou, C. et al. Identification of glycerol-3-phosphate dehydrogenase 1 as a tumour suppressor in human breast cancer. Oncotarget 8, 101309–101324 (2017).
Zhang, W. et al. Allosteric activation of the metabolic enzyme GPD1 inhibits bladder cancer growth via the lysoPC–PAFR–TRPV2 axis. J. Hematol. Oncol. 15, 93 (2022).
Kawano, Y. & Cohen, D. E. Mechanisms of hepatic triglyceride accumulation in non-alcoholic fatty liver disease. J. Gastroenterol. 48, 434–441 (2013).
Nagle, C. A., Klett, E. L. & Coleman, R. A. Hepatic triacylglycerol accumulation and insulin resistance. J. Lipid Res. 50, S74–S79 (2009).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Scott, D. A. Analysis of melanoma cell glutamine metabolism by stable isotope tracing and gas chromatography-mass spectrometry. Methods Mol. Biol. 2265, 91–110 (2021).
Pino, L. K. et al. The Skyline ecosystem: informatics for quantitative mass spectrometry proteomics. Mass Spectrom. Rev. 39, 229–244 (2020).
Millard, P. et al. IsoCor: isotope correction for high-resolution MS labeling experiments. Bioinformatics 35, 4484–4487 (2019).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Acknowledgements
We thank David Scott (SBP Discovery Cancer Metabolism Core, La Jolla, CA) for technical support. This work was supported by grants from the National Institutes of Health (R35GM142495 and R03CA286706 to V.C. and R01DK134675 to R.P.G.). R.P.G. was supported by a Career Award for Medical Scientist (Burroughs Welcome Fund). The Seahorse XFe96 analyser in the Saez laboratory (TSRI) was supported by 1S10OD16357. The Metabolomics Platform (RRID: SCR_022932) was supported by the University of Chicago Comprehensive Cancer Center Support Grant (P30 CA014599). This is manuscript 1080 from the Scintillon Institute.
Author information
Authors and Affiliations
Contributions
X.P. performed all experiments with assistance from A.L.Z., S.M. and A.P. S.M. performed protein purification and enzyme kinetics experiments. H.S., N.B.T. and R.M.P. performed all the lipid profiling, metabolomic profiling and stable-isotope-tracing experiments. R.P.G. and N.S. performed mouse experiments. S.V. and J.R.C. performed LC–MS experiments to screen GPDHs. X.P., V.C. and A.L.Z. wrote the paper, with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
V.C., X.P. and A.L.Z. are listed as inventors on a patent application (no. 63/812,699: ‘Enzyme-based compositions and methods for treating mitochondrial dysfunction’) based on sequences and activities of proteins described in this paper. V.C. is listed as an inventor on a patent application on the therapeutic uses of LbNOX and TPNOX (US patent application US20190017034A1). The other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Melanie McReynolds, Yi Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Gimenez-Cassina and Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effect of CrGPDH expression in HeLa cells on metabolic features, bioenergetics and GSH/GSSG ratio.
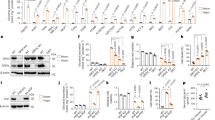
Lactate (a) and pyruvate (b) levels in media incubated for 24 hours (spent media) with HeLa cells expressing Luciferase or CrGPDH. Glucose consumption (c), lactate production (d) and glutamine consumption (e) by HeLa cells expressing CrGPDH. Oxygen consumption rate (OCR) (f) and extracellular acidification rate (ECAR) (g) of HeLa cells expressing CrGPDH before and after separate additions of 1 µM piericidin A (Pier) or 1 µM antimycin A (ANT) measured in pyruvate-free HEPES/DMEM+dFBS media containing 5 mM glucose. GSH (h), GSSG (i) and the GSH/GSSG ratios (j) in HeLa cells expressing CrGPDH. Two hundred μM menadione (vitamin K3) was used as a positive control in (h-j). Experiments in (a-e, h-j) were performed with pyruvate-free DMEM supplemented with dialysed FBS (DMEM+dFBS) containing 25 mM glucose. Values are mean ± s.d.; n = 5, 6 in (a), n = 6, 6 in (b), n = 11, 6 in (c), n = 11, 11 in (d), n = 12, 10 in (e), n = 45, 15, 30, 55, 25, 30 in (f), n = 50, 20, 30, 60, 30, 30 in (g), n = 3 in (h-j) biologically independent samples. The statistical significance indicated for (a-e) represents a two-tailed unpaired t test; for (f-g) represents a One-Way ANOVA followed by Šídák multiple comparison test; for (h-j) represents a Two-Way ANOVA followed by uncorrected Fisher’s least significant difference test.
Extended Data Fig. 2 SoNar and iNAP1 imaging of HeLa cells expressing CrGPDH.
(a) Widefield images of HeLa cells with lentivirus mediated LUC and CrGPDH expression under Dox control transiently expressing SoNar. Experiments were performed in the imaging medium (DMEM without pyruvate, fluorescent vitamins and phenol red with 5 mM glucose, 25 mM HEPES, pH 7.4 and 1% dialyzed FBS), or after addition of 1 μM antimycin A. Scale bars: 20 µm. (b) Quantification of the time course measurements of the fluorescence ratio (F400/488) for HeLa cells with lentivirus mediated LUC and CrGPDH expression under Dox control transiently expressing SoNar for conditions shown in (a). (c) Widefield images of HeLa cells with lentivirus mediated LUC and CrGPDH expression under Dox control transiently expressing iNAP1. Experiments were performed in the imaging medium (DMEM without pyruvate, fluorescent vitamins and phenol red with 5 mM glucose, 25 mM HEPES, pH 7.4 and 1% dialyzed FBS), or when cells were switched to the imaging medium without glucose but with 10 mM pyruvate. Scale bars: 20 µm. Quantification of the time course measurements of the fluorescence ratio (F400/488) for HeLa cells with lentivirus mediated LUC and CrGPDH expression under Dox control transiently expressing iNAP1 (d) or inactive variant which does not bind NADPH (iNAPC) (e) in the imaging medium (DMEM without pyruvate, fluorescent vitamins and phenol red with 5 mM glucose, 25 mM HEPES, pH 7.4 and 1% dialyzed FBS), or when cells were switched to the basal medium without glucose but with 10 mM pyruvate. Values are mean ± s.d.; n = 27, 30 in (b), n = 29, 27 in (d), n = 20, 20 in (e) biologically independent samples.
Extended Data Fig. 3 Metabolic flux features of CrGPDH expression HeLa cells.
(a) Schematic showing carbons flux from glucose and glutamate. Brown circles represent 13C-labelled carbons from glucose, purple circles represent 13C-labelled carbons from glutamine. Heatmaps of the glucose carbon flux into glycolysis (b), TCA cycle (c), pentose phosphate pathway (PPP) and purine (d), and pyrimidine (e) metabolites, as well as TCA cycle intermediates from glutamine carbon transitions (f) in HeLa cells expressing CrGPDH and LUC after 8hr-culture with 10 mM U-¹³C₆ D-Glucose or 2 mM ¹³C₅, 99% L-Glutamine. Note that unless specified, the values in (b-f) are the abundances of the most labelled isotopologue. (g) Fractional labeling of citrate M + 4 (oxidative) and M + 5 (reductive) from ¹³C₅ glutamine. G6P: glucose-6-phosphate; F6P: fructose-6-phosphate; F1,6BP: fructose-1,6-bisphosphate; DHAP: dihydroxyacetone phosphate; Gro3P, glycerol-3-phosphate; AKG, α-ketoglutarate; 6PG: 6-phosphogluconate; R5P: ribose-5-phosphate. In heatmaps in (b-f), each column represents a biologically independent sample. Values are mean ± s.d.; n = 5 in (g) biologically independent samples. The statistical significance indicated for (g) represents a two-way ANOVA followed by Fisher’s LSD test without correction.
Extended Data Fig. 4 Analysis of transcriptomic features of CrGPDH expression in HeLa cells.
(a) Volcano plots that represent the log2 fold change (x axis) and adjusted p value for significance (y axis) of CrGPDH vs LUC expressing HeLa cells. Genes significantly different in expression at false discovery rate (FDR) of 5% are indicated in red (upregulated genes, log2 fold change above 1) or blue (downregulated genes, log2 fold change below -1). Grey dots represent genes without significant changes. (b) Gene ontology (GO) terms analysis of significant genes that are differentially changed between CrGPDH and LUC. BP: biological process; CC, cellular component; MF: molecular function. The Cnet plots for the subset of genes that correlate with GO terms BP in (c) or MF in (d). Analysis shown in (a-d) is based on n = 4 biologically independent samples per group.
Extended Data Fig. 5 Metabolic features of CrGPDH expression in 786-O cells.
Glycerol (a), lactate (b), pyruvate (c) and the lactate/pyruvate ratio (d) measured by GC-MS in media which was incubated for 24 hours (spent media) with 786-O cells expressing Luciferase (LUC) or CrGPDH. Glucose consumption (e), lactate production (f) and glutamine consumption (g) measured by the YSI 2900 Biochemistry Analyzer in media which was incubated for 24 hours (spent media) with 786-O cells expressing Luciferase (LUC) or CrGPDH. Targeted metabolomics of 786-O cells expressing CrGPDH relative to LUC control (h). Significantly increased metabolites (p value cutoff < 0.05, fold change >1) are highlighted in red, and significantly decreased metabolites (p value cutoff < 0.05, fold change < -1) are highlighted in dark blue, while gray dots represent metabolites without significant changes. Experiments in (a-g) were performed in pyruvate-free RPMI+dFBS containing 5 mM glucose; in (h) were performed in pyruvate-free RPMI+dFBS containing 25 mM glucose. UDP-GlcA: UDP-glucuronic acid. Values are mean ± s.d.; n = 6, 6 in (a-g) biologically independent samples. The statistical significance indicated for (a-g) represents a two-tailed unpaired t test. NS, no significant difference.
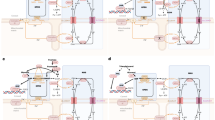
Extended Data Fig. 6 Diagram summarizing metabolic rewiring in clear cell renal cell carcinoma (ccRCC) cells expressing CrGPDH.
(a) In ccRCC cells, the Gro3P shuttle is functionally truncated, as cytosolic cGPDH is uncoupled from mitochondrial mGPDH, thereby supporting a robust Gro3P pool for TG synthesis57. (b) Expression of CrGPDH in these cells facilitates efficient Gro3P clearance, leading to suppression of TG synthesis. FFA: free fatty acid; TG: triglycerides; GL/FFA cycle: glycerolipid/free fatty acid cycle; GK: glycerol kinase.
Extended Data Fig. 7 Metabolic features of CrGPDH expression in IMR-90 primary human fibroblasts.
Lactate (a) and pyruvate (b) measured by GC-MS in media which was incubated for 24 hours (spent media) with IMR-90 cells expressing Luciferase (LUC) and CrGPDH. Glucose consumption (c), lactate production (d) and glutamine consumption (e) measured by the YSI 2900 Biochemistry Analyzer in media which was incubated for 24 hours (spent media) with IMR-90 cells expressing CrGPDH or Luciferase (LUC). Targeted metabolomics of IMR-90 cells expressing CrGPDH relative to LUC control (f). Significantly increased metabolites (p value cutoff < 0.05, fold change > 0.7) are highlighted in red, and significantly decreased metabolites (p value cutoff < 0.05, fold change < -0.7) are highlighted in dark blue, while gray dots represent metabolites without significant changes. Experiments were performed in pyruvate-free DMEM+dFBS supplemented with 5 mM glucose in (a-e) or 25 mM glucose in (f). Values are mean ± s.d.; n = 6, 6 in (a-e) biologically independent samples. The statistical significance indicated for (a-e) represents a two-tailed unpaired t test. NS, no significant difference.
Extended Data Fig. 8 Hepatic metabolic features of CrGPDH expression in mice.
KEGG pathway enrichment analysis of altered metabolites in (a) CrGPDH versus GFP under water gavage, (b) ethanol versus water in GFP-expressing mice, and (c) CrGPDH versus GFP under ethanol gavage. X axis represents KEGG terms, Y axis represents log10 p value. Pathway analysis was performed in MetaboAnalyst 6.0 using metabolites with p < 0.05 in (a, b) and p < 0.2 in (c). (d-f) The schematics highlighting related metabolomic pathways and selected metabolites for indicated comparisons in (a-c). Heatmaps of the most impacted pyrimidines (g), beta-Alanine (h), arginine (i), TCA cycle (j) and glycerophospholipid (k) metabolites in indicated group. AKG, α-ketoglutarate; Gro3P, glycerol-3-phosphate; Gro3P-Cho, glycerol-3-phosphate Choline; P-Cho, Phosphocholine; PEA, Phosphoethanolamine; CDP-Etn, CDP-Ethanolamine; CDP-Cho, CDP-Choline. In heatmaps in (g-k), each column represents a biologically independent sample.
Extended Data Fig. 9 Hepatic lipid profiles of CrGPDH expression in mice.
Volcano plots of summed lipid classes. Comparisons are: (a) CrGPDH versus GFP under water gavage, (b) ethanol versus water in GFP-expressing mice, and (c) CrGPDH versus GFP under ethanol gavage. (d) Heatmaps of all detected lipids, clustered by each lipid class across the indicated groups. Significantly increased summed lipids (log2FC > 0.1, p < 0.05) are shown in red, decreased summed lipids (log2FC < -0.1, p < 0.05) in dark blue, and nonsignificant ones in gray. ChE: Cholesterol ester, CoQ: Coenzyme Q-like molecules, DG: Diglycerides, HexCer: Ceramide with hexose, LBPA: Glycerophosphoglycerols, LPC: Lysophosphatidylcholine, LPE: Lysophosphatidylethanolamine, LPI: Lysophosphatidylinositol, MePC: Methyl phosphatidylcholine, PA: Phosphatidic acid, PC: Phosphatidylcholine, PE: Phosphatidylethanolamine, PG: Phosphatidylglycerol, PI: Phosphatidylinositol, PS: Phosphatidylserine, SM: Sphingomyelin, TG: triglycerides. In heatmaps in (d), each column represents a biologically independent sample.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–10.
Supplementary Data 1 (download XLSX )
Source data for Supplementary Figures.
Supplementary Tables 1–4 (download XLSX )
Relevant to Methods.
Supplementary Tables 5–9 (download XLSX )
Relevant to raw metabolomics peak areas.
Supplementary Tables 10–14 (download XLSX )
Relevant to raw lipidomics peak areas.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download TIF )
Unprocessed western blots.
Source Data Fig. 2 (download TIF )
Unprocessed gel.
Source Data Fig. 6 (download TIF )
Unprocessed western blots.
Source Data Fig. 7 (download TIF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pan, X., Munan, S., Zuckerman, A.L. et al. A genetically encoded bifunctional enzyme mitigates redox imbalance and lipotoxicity. Nat Metab 8, 350–370 (2026). https://doi.org/10.1038/s42255-025-01450-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01450-3
This article is cited by
-
A metabolic safety valve for reductive stress
Nature Metabolism (2026)