Abstract
Irisin is an exercise-induced myokine that confers multiple physiological benefits, including browning of subcutaneous adipose tissue in mice. However, the underlying cellular and molecular mechanisms of irisin’s effects on obesity are unclear. Here, we show that irisin modulates adipose tissue inflammation by increasing interleukin (IL)-33 production and preserving ST2+ regulatory T cells in white adipose tissues. Administered chronically to high-fat-diet-fed male mice, irisin preserves visceral adipose tissue (VAT) levels of ST2+ regulatory T cells, an important immunomodulatory population that usually contracts after long-term high-fat-diet feeding. This protection results from increases in IL-33-producing mesenchymal stromal cells and IL-33 levels in VAT. These effects are primary, as irisin directly induces IL-33 expression in cultured VAT mesenchymal stromal cells. Irisin-mediated changes in VAT IL-33 dynamics are accompanied by IL-33-dependent upregulation of thermogenic gene expression in subcutaneous adipose tissue. These irisin-driven cell–cell and inter-tissue interactions improve obesity and glucose intolerance, and increase energy expenditure, with no reduced food intake and muscle loss in obese mice.
Similar content being viewed by others
Main
Obesity affects more than 40% of adults in the United States, and its prevalence worldwide involves over a billion people1. Individuals with obesity are prone to develop metabolic comorbidities such as type 2 diabetes and cardiovascular diseases2. These metabolic syndromes derive, at least in part, from chronic, low-grade inflammation of the adipose tissues, which disrupts tissue and glucose homeostasis3,4. Adipose tissue inflammation further impedes the development and activation of thermogenic fat5, the specialized fat depot within which adipocytes transform fuel into heat via running futile chemical cycles6,7. There are two types of characterized thermogenic fat: brown and beige fat. The latter arises within white adipose tissue, and confers a variety of metabolic benefits, including increased energy expenditure, improved glucose homeostasis and insulin sensitivity. Restoring the activity of thermogenic fat is a promising strategy to combat the metabolic defects induced by obesity8.
Exercise improves several metabolic features associated with obesity, including promoting glucose and insulin sensitivity9,10, as well as lowering adipose tissue inflammation. Our lab identified a functional polypeptide termed irisin from peroxisome proliferator-activated receptor γ coactivator 1α-expressing and exercised muscle over 10 years ago; irisin recapitulates certain aspects of endurance physical training, including stimulating beiging of subcutaneous white adipose tissue11. Irisin is the cleaved product from a type-1 membrane protein FNDC5 (fibronectin domain III (FNIII) containing 5), whose levels are increased with exercise12,13,14,15. Its effects are mediated via αV integrins, and irisin binding and signalling requires activation of integrin receptors by a stress-induced co-factor heat-shock protein 90α (HSP90α)16. This molecular link between irisin action and stress motivated us to examine in detail the role of chronic irisin administration on the metabolic and immunological parameters of diet-induced obese mice.
VAT, compared with other fat depots, is most closely associated with the metabolic pathologies that develop in individuals with obesity17. VAT is more resistant to ‘beiging’ compared to subcutaneous adipose tissue (SAT); yet it is highly metabolically active and presents a unique repertoire of immunocytes and immunomodulatory cells that maintain adipose tissue homeostasis18. This repertoire includes anti-inflammatory macrophages, type 2 innate lymphoid cells (ILC2s), regulatory T (Treg) cells and specialized stromal cells that modulate the activity and dynamics of the immunocytes. VAT-Treg cells are a well-characterized Treg population that keeps immunocytes and adipocytes in check to maintain metabolic homeostasis19. While preserving the Treg lineage-defining transcription factor, FOXP3, and the classic Treg signature, VAT-Treg cells differ from their lymphoid-organ counterparts by a distinct transcriptome and T cell antigen receptor repertoire20,21. Furthermore, VAT-Treg cells have distinct growth and survival factor dependencies, most notably PPARγ22 and the cytokine IL-33 and its receptor ST2 (refs. 21,23,24). IL-33 is mainly produced by a subset of mesenchymal stromal cells (mSCs) in VAT that have poor adipogenic potential, and locally produced IL-33 is a key determinant of VAT-Treg accumulation and function25,26.
IL-33 is known for its role in inducing type 2 immunity in allergic diseases. It also modulates VAT inflammation and improves metabolic indices in obese mice through regulating ST2+ immunocytes, including ST2+ Treg cells and ILC2s27. In addition, several studies have demonstrated a role for IL-33 in promoting adipose tissue thermogenesis, although the detailed mechanism is under some debate. IL-33-mediated UCP1 expression primarily occurs during beiging of white adipose tissue in adult animals, and the involvement of ST2+ immunocytes, including SAT-ILC2s, requires further investigation26,27,28,29,30.
Here, we demonstrate that irisin modulates adipose tissue inflammation and promotes thermogenesis in obese mice through an IL-33–ST2 axis in white adipose tissues, largely operating through ST2+ Treg cells. These irisin-induced pathways collectively reduce fat content without loss of muscle mass and greatly improve glucose tolerance and insulin sensitivity in diet-induced obese mice.
Results
Chronic irisin treatment increases energy expenditure, and improves obesity and glucose homeostasis in HFD-fed mice
We previously showed that short-term irisin treatment stimulated beiging of white adipose tissue and improved glucose intolerance in mice. However, whether promoting thermogenesis is the only irisin action accounting for all irisin-mediated effects in glucose homeostasis and insulin insensitivity and what are the cellular pathways stimulated by irisin are not known. Here, we investigated the effects and underlying mechanisms of chronic irisin treatment in diet-induced obese mice. We used adeno-associated virus 8 (AAV8) to express FLAG-tagged irisin or GFP (control) in the mouse liver. Prior studies indicated that this route of administration stably elevates irisin concentrations to levels approximately 20-fold above basal levels in the plasma from 4 weeks until at least 6 months after AAV injection31. To observe any potential irisin-mediated effects at different stages of obesity progression, we assessed the effects of irisin in a prevention model with mice challenged by a high-fat-diet (HFD). Thus, we injected 4-week-old C57BL/6 male mice with the AAVs via the tail vein and then switched to a HFD 5 weeks later. The AAV injections resulted in increased plasma irisin levels, which were stable throughout the course (18 weeks) of HFD feeding (Extended Data Fig. 1a).
To study irisin’s actions in this model, we conducted metabolic and tissue analyses at the indicated time points (Fig. 1a). Both groups of animals were rendered obese by feeding a HFD; however, the weights of mice that received AAV-irisin were lower than those of control mice after 8 weeks of HFD feeding, without a significant change in food intake (Fig. 1b,c). Body composition analysis by magnetic resonance imaging (MRI) suggested that the lower body weight of the irisin group was largely due to a reduction in fat mass (Fig. 1d) with no loss in lean mass. Adipose tissue weights suggested that the subcutaneous depots were largely responsible for the irisin-mediated reduction in fat mass at this stage (Extended Data Fig. 1b,c). There was also improved glucose tolerance and insulin sensitivity in the irisin compared with the GFP group, as measured by intraperitoneal glucose tolerance tests (GTTs) and insulin tolerance tests (ITTs), respectively (Fig. 1e,f). These effects were not linked to changes in physical activity (Extended Data Fig. 1e,f). Interestingly, differences in body weight, glucose tolerance and insulin sensitivity between the irisin and GFP groups were not detected either at the early stage (8 weeks) of HFD feeding (Extended Data Fig. 1g,h) or in age-matched, normal chow-fed mice at the late stage (corresponding to 18 weeks of HFD feeding; Extended Data Fig. 2a–g). These data show that chronic elevation of circulating irisin reduces diet-induced obesity and its closely linked metabolic defects.
Four-week-old male C57BL/6J mice were intravenously (i.v.) injected with AAV-GFP (n = 15) or AAV-irisin (n = 15). Five weeks later, the mice were fed a HFD. After 8 weeks of HFD feeding, half of the mice were euthanized for tissue analysis, and the other half of the mice were kept until 18 weeks of HFD feeding. a, Schematic of the HFD experiment. b, Body weight measurements (n = 15, 15 until week 17; n = 10, 10 after week 17). Unpaired two-sided t-test was used for calculating P values. c, Food consumption measurements (n = 10, 10). Unpaired two-sided t-test was used for calculating P values. d, Body composition analysis by MRI (n = 8, 8). Unpaired two-sided t-test was used for calculating P values (**P = 0.0047). e, Intraperitoneal GTTs (n = 8, 5). Two-way analysis of variance (ANOVA) was used for calculating P values (**P = 0.0011, ****P < 0.0001 and ****P < 0.0001). f, Intraperitoneal ITTs (n = 5, 5). Two-way ANOVA was used for calculating P values (**P = 0.0018, ****P < 0.0001 and ***P = 0.0003). CLAMS, comprehensive lab animal monitoring system. Mean ± s.d. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, NS, not significant. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustration in a created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
The strong increase in energy expenditure in response to elevated irisin levels may be suggestive of activation of thermogenesis in the fat depot(s), given that short-term irisin treatments were shown to promote adipose tissue thermogenesis (Extended Data Fig. 1d). Therefore, we examined the gene expression profiles of brown adipose tissue (BAT), inguinal white adipose tissue (iWAT) and epididymal adipose tissue (eWAT) from the mice fed a HFD for 18 weeks using whole-tissue RNA sequencing (RNA-seq). Pathway analysis (fold change > 2; adjusted P value < 0.05) revealed that a few processes were upregulated by irisin, exclusively in iWAT, including thermogenesis (Extended Data Fig. 1k). Several years ago, we identified a UCP1-independent thermogenic pathway, the futile creatine cycle (FCC)32. This pathway involves the repeated phosphorylation and dephosphorylation of creatine and is largely catalysed by mitochondrial creatine kinase CKB and the creatine phosphatase TNAP32,33. To investigate if thermogenic programmes were stimulated by irisin in the adipose tissues, we investigated the expression of the key mediators of different futile cycles using quantitative PCR with reverse transcription (RT–qPCR). Irisin elevation significantly increased expression of both Ucp1 (~1.9-fold) and the key FCC genes Ckb (3.3-fold) and Alpl (gene encoding TNAP; 2.4-fold), specifically in iWAT (Extended Data Fig. 1l). These changes were not observed in eWAT and BAT, nor in age-matched, normal chow-fed mice at the late stage (corresponding to 18 weeks of HFD feeding; Extended Data Fig. 2h–j). No significant changes in lipolysis-related gene expression events were detected (Extended Data Fig. 2k).
Irisin inhibits inflammation and enhances the survival of ST2+ Treg cells in eWAT of obese mice
Obesity in both mice and humans is associated with chronic, low-grade inflammation, which impairs glucose homeostasis and insulin sensitivity as well as impeding thermogenesis3,4. To explore whether the metabolic benefits of irisin reflected immunomodulatory processes, we performed a comprehensive flow cytometric analysis of eWAT, iWAT and BAT from HFD-fed versus normal chow-fed mice administered either AAV-irisin or AAV-GFP. This analysis encompassed both the adaptive and innate arms of the immune system (see Extended Data Fig. 3a,b for gating).
We first profiled immunocyte populations at the 18-week time point, concurrent with the metabolic changes illustrated in Fig. 1e,f. There were no changes in the representation of either total immunocytes (CD45+ cells) or total macrophages (CD64+ cells) under any of the four experimental conditions (Fig. 2a,b). In contrast, CD11chi pro-inflammatory macrophages were more frequent in HFD-fed mice administered either control AAV or AAV-irisin (Fig. 2b). Irisin did not influence the representation of total T cells in normal chow-fed mice; T cells trended higher in HFD-fed mice injected with control AAV, which was largely normalized when they were administered AAV-irisin instead (Fig. 2c). Irisin treatment decreased the fraction of CD8+ (although not non-Treg CD4+) T cells under both feeding regimes (Fig. 2c). As was previously reported20, and was recapitulated in this experiment, the total eWAT-Treg (Foxp3+CD4+) population, in particular the ST2+ subtype, is typically strongly reduced in mice fed a HFD long term. Importantly, irisin treatment strongly attenuated these Treg cell reductions, almost totally in the case of ST2+ Treg cells (Fig. 2d), known to be important for controlling metabolic indices34,35.
Four-week-old male C57BL/6J mice were intravenously injected with AAV-GFP or AAV-irisin. Five weeks later, the mice were fed either a normal chow (NC) or HFD for 18 weeks (a–d, n = 3, 3, 8, 8). a–d, Flow cytometric analysis of the number of total CD45+ cells (a), and percentages of total and CD11chi macrophages (b), as well as total T, CD8+ and Foxp3−CD4+ T cells (c) in eWAT from mice fed a NC or HFD for 18 weeks. d, Representative gating strategy for analysing total (black) and ST2+ (red) eWAT-Treg cells (left); summary quantification of their percentages (right). One-way ANOVA was used for calculating P values (b: *P = 0.0459 and *P = 0.0464; c: **P = 0.0167, ***P = 0.0086 and *P = 0.0252; d: *P = 0.0299, *P = 0.0335, *P = 0.0207 and ***P = 0.0002). MFs, macrophages; CD11chi, CD11chi. Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
For a more detailed analysis, we converted the 13-parameter T cell data into a t-distributed stochastic neighbour embedding plot, which allowed us to quantify ten distinct clusters (Extended Data Fig. 3c). Eighteen weeks of HFD feeding increased the representation of activated CD4+ conventional T cells and CD8+ T cells, while decreasing that of Treg cells, specifically the ST2+ subtype; the fraction of activated γδ T cells was also reduced (Extended Data Fig. 3d). Irisin prevented the increase in CD4+ and CD8+ effector T cells, coincident with a strong enrichment in ST2+ Treg cells and a trending increase in the fraction of naive CD8+ T cells (Extended Data Fig. 3d). A differential density plot between the immunocyte landscapes of mice on a HFD treated with AAV-GFP versus AAV-irisin highlighted the irisin-induced decreases in CD4+ and CD8+ effector T cells and increases in ST2+ Treg cells and naive CD8+ T cells (Extended Data Fig. 3e). Thus, the two approaches to screening the effects of irisin on immunocyte profiles gave similar results.
Next, we assessed whether the irisin effects were confined to long-term HFD feeding. At an 8-week time point, mice treated with the control AAV showed a twofold increase in total immunocytes (CD45+ cells) when fed a HFD compared with normal chow; this increase did not occur in AAV-irisin-treated mice (Extended Data Fig. 3f). Unlike at the 18-week time point, irisin reduced the representation of total macrophages and T cells that accumulated with HFD feeding but did not preferentially affect the pro-inflammatory macrophages, CD8+ T cells or non-Treg CD4+ T cells (Extended Data Fig. 3g,h). At 8 weeks, representations of total Treg cells were similar under the four conditions, although there was a slight decrease in the dominant ST2+ subset in irisin-treated, normal chow-fed mice (Extended Data Fig. 3i). The different immunocyte profiles at the 8-week and 18-week time points align with previous observations on the dynamics of eWAT-Treg cells in response to HFD feeding, that is, an initial expansion phase followed by a sharp contraction36.
For an in situ view, we quantified crown-like structures (CLSs), a classical marker of inflammation and adipocyte death7, in eWAT or iWAT of HFD-fed mice administered AAV-irisin or control AAV-GFP. Irisin-treated mice on a HFD for 8 weeks showed a significant reduction in eWAT CLS counts compared with those of control-treated littermates (Supplementary Fig. 1a). As anticipated, CLS counts were higher at 18 weeks than at 8 weeks of HFD feeding in control-treated mice; irisin treatment reduced this number by 40% (Supplementary Fig. 1a). In contrast, no significant irisin-mediated reduction of CLS counts was observed in iWAT at either 8 or 18 weeks of HFD (Supplementary Fig. 1b).
Interestingly, the immunomodulatory effects of irisin were not observed in iWAT or BAT at either 8 or 18 weeks of HFD feeding (Extended Data Fig. 4a–d).
Irisin directly induces IL-33 expression in eWAT stromal cells
To ascertain whether irisin affected ST2+ Treg cells in eWAT directly or indirectly, we cultured cytofluorimetrically sorted total Treg cells from eWAT with either HSP90α plus irisin or control buffer (phosphate-buffered saline or PBS) and compared their ability to expand the ST2+ Treg subset (Fig. 3a). HSP90α, an important co-factor that is required for irisin receptor activation, was supplemented here to allow irisin actions. No significant difference was observed between the two groups, suggesting that irisin’s effect on eWAT-Treg cells may not be direct.
a, Freshly isolated eWAT-Treg cells from 12-week-old male Foxp3.Thy1.1 mice were cultured with or without irisin/HSP90α for 3 days, followed by flow cytometric analysis. Left, experimental scheme; right, summary data for total Treg numbers and fraction of ST2+ Treg cells. n = 3. b, Summary data for numbers of IL-33+ mSCs from the mice in Fig. 2a (n = 3, 3, 6 and 7). One-way ANOVA was used to calculate P values (*P = 0.0127). c, ELISA measurements of IL-33 concentration in eWAT lysates from mice in Fig. 1e (n = 3, 3, 6, 6). One-way ANOVA was used for calculating P values (**P = 0.0019, ****P < 0.0001 and ****P < 0.0001). d, ELISA measurements of IL-33 concentration in plasma from mice in Fig. 1e (n = 6, 9). Unpaired two-sided t-test was used for calculating P values (*P = 0.0342). e, Freshly isolated IL-33 (EGFP)+ or IL-33 (EGFP)− mSCs from 12-week-old male Il33.Egfp mice were cultured with PBS or the indicated concentrations of irisin/HSP90α for 3 days, followed by flow cytometric analysis. Left, experimental scheme; right, summary data for numbers and mean fluorescence intensity of IL-33 (EGFP)+ mSCs. n = 4. One-way ANOVA was used for calculating P values (**P = 0.0019, ***P = 0.0006, ****P < 0.0001). f, Freshly isolated IL-33 (EGFP)− mSCs from 12-week-old male Il33.Egfp mice were cultured with PBS or 10 nM irisin/HSP90α for 3 days, and the conditioned medium treated with either control IgG or anti-IL-33 antibody (2 μg ml−1) was subsequently used for culturing freshly isolated eWAT-Treg cells for another 3 days followed by flow cytometric analysis. Top, experimental scheme; bottom, summary data for fractions (left) and numbers (right) of ST2+ Treg cells (n = 2, 3, 5). One-way ANOVA was used for calculating P values (left: ****P < 0.0001, **P = 0.00156 and ****P < 0.0001; right: ****P < 0.0001, *P = 0.0122 and ****P < 0.0001). g, Volcano plot showing transcripts differentially expressed (fold change > 2, P < 0.05) by IL-33 (EGFP)− mSCs cultured with PBS or 10 nM irisin/HSP90α for 3 days (left). Transcripts from an adipogenesis up-signature are highlighted in magenta (right). Triplicate samples. Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustrations in a, e and f created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
Previous studies have shown that IL-33 secreted by eWAT mSCs is a crucial driver of ST2+ Treg accumulation in that fat depot25. Therefore, we assessed IL-33 expression by mSCs (CD45−CD31−PDGFRα+Sca-1+ cells) in eWAT, iWAT and BAT of mice treated as per Fig. 1a (see Extended Data Fig. 5a for gating). Irisin treatment resulted in an ~3-fold increase in the numbers of IL-33-producing mSCs in eWAT of irisin-treated, compared with control-treated, mice after 18 weeks of HFD feeding (Fig. 3b). This effect was not evident in mice on a HFD for only 8 weeks, nor in those on normal chow (Extended Data Fig. 5b). We also measured IL-33 concentrations in whole-tissue lysates of the three fat depots under the four conditions. Consistent with previous findings37,38, 18 weeks of HFD induced a twofold increase in the IL-33 concentration of eWAT, while irisin treatment boosted this increase by another twofold compared with control AAV treatment (Fig. 3c). Irisin did not significantly influence the numbers of IL-33-producing mSCs in iWAT or BAT at either time point (Extended Data Fig. 5c). This finding aligns with the comparable ST2+ Treg cell numbers in these two fat depots from irisin-treated and control AAV-treated obese mice (Extended Data Fig. 3d–g). However, we did detect about twofold more IL-33 in iWAT (but not BAT) of irisin-treated versus control-treated obese mice (Extended Data Fig. 5d). Plasma concentrations of IL-33 were also significantly higher (~2-fold increase) in obese mice treated with AAV-irisin (Fig. 3d). Importantly, the irisin–IL-33 axis appeared to selectively influence ST2+ eWAT-Treg cells, as there was no significant impact on ILC2s, another ST2-expressing cell type, in any adipose depot at either time point (Extended Data Fig. 5e).
These findings raised two important questions: does irisin act directly on eWAT mSCs to induce IL-33 expression? And, if so, does it induce immunocyte-promoting IL-33+ mSCs to proliferate, or does it direct adipocyte-generating IL-33− mSCs to produce IL-33 de novo (and perhaps change fate)25? To address these questions, we cytofluorimetrically sorted EGFP+ or EGFP− mSCs from eWAT of Il33-Ires-Egfp reporter mice, incubated them with irisin plus HSP90α or PBS for 3 days and quantified numbers of IL-33-expressing cells and their levels of IL-33 expression (Fig. 3e). Concerning the cultures of sorted IL-33+ mSCs, irisin did not increase their numbers but did enhance Il33 transcription, evidenced by the higher mean fluorescence intensity of their Il33 transcript reporter, EGFP (Fig. 3e). With respect to the cultures of sorted IL-33− mSCs, irisin induced an increase in both their numbers and Il33 transcript (EGFP) expression levels in a dose-dependent manner (Fig. 3e). In the absence of HSP90α, irisin failed to induce Il33 in eWAT mSCs (Extended Data Fig. 5f), consistent with our previous findings that HSP90α is necessary for irisin to bind to its receptor16.
The above findings raised the question of whether irisin/HSP90α-treated mSCs from eWAT acted directly on the ST2+ Treg population to induce its expansion. We co-cultured eWAT-Treg cells with control IgG-treated conditioned medium from PBS-treated IL-33 (EGFP)− mSCs from eWAT, or with either control IgG-treated or anti-IL-33-treated conditioned medium from irisin/HSP90α-treated counterparts (Fig. 3f). Conditioned medium from irisin/HSP90α-treated mSCs induced a >2-fold increase in the numbers and fractional representation of ST2+ eWAT-Treg cells (Fig. 3f), demonstrating that irisin-treated mSCs can directly affect ST2+ Treg cells.
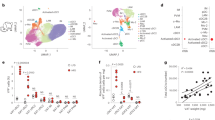
Our previous studies discriminated two major lineages of eWAT mSCs: IL-33+PPARγ− immunocyte-promoting cells and IL-33−PPARγ+ adipocyte-generating cells25. To determine whether irisin simply induced IL-33 expression in IL-33− cells or instead modified their adipogenic potential, we performed populational-level RNA-seq on irisin/HSP90α-treated versus PBS-treated IL-33 (EGFP)− eWAT mSC cultures. Consistent with the flow cytometric analysis, irisin induced a tenfold increase in Il33 transcripts, along with those encoding other immunomodulatory cytokines and chemokines such as Il6, Fgf23, Ccl2, Ccl7 and Cxcl5. Conversely, expression of 75% of an adipogenesis up genetic signature (147 of 195 genes), including Fabp4, Adipoq, Cd36 and Pparg, was attenuated by irisin addition (Fig. 3g). This finding suggests a potential shift in cell fate or at the least a transcriptional impact much greater than simply turning on IL-33 expression.
Our group identified integrin αVβ5 as the major irisin receptor in bone14. To investigate the role of this integrin in irisin-mediated IL-33 production, we first checked expression of transcripts encoding αVβ5 in visceral adipose tissue mesenchymal stromal cells (VmSCs) from existing RNA-seq data39. VmSCs expressed transcripts encoding both the αV and β5 subunits, with two IL-33-producing VmSC subtypes (VmSC1 and VmSC2) expressing significantly higher levels of the β5 subunit (Extended Data Fig. 6a). We next performed RT–qPCR and fluorescence imaging with cytofluorimetrically sorted EGFP− VmSCs that had been treated with irisin plus HSP90α or PBS in the presence of an anti-αVβ5 antibody40 or isotype-control IgG for 3 days. The irisin-induced increase in levels of IL-33 mRNA and the numbers of IL-33-expressing cells were largely blocked by αVβ5 antibody but not control IgG treatment (Extended Data Fig. 6b,c). Moreover, irisin-stimulated integrin signalling (pFAK/FAK ratio) in these cells was ablated (Extended Data Fig. 6d).
Irisin controls the accumulation of ST2+ eWAT-Treg cells through IL-33
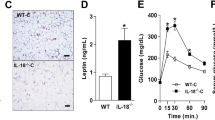
We then performed loss-of-function and complementation experiments with whole-body FNDC5-deficient (Fndc5−/−) mice. Flow cytometric analysis revealed that 16-week-old mutant mice had a normal number of eWAT mSCs but fewer of them expressed IL-33 in comparison with their wild-type littermates (Fig. 4a). Not surprisingly, then, these mutant mice, while hosting a CD4+ T cell compartment of normal size, had a deficit in both total and ST2+ Treg cells in eWAT, but not in spleen (Fig. 4b and Extended Data Fig. 7a).
a, Flow cytometric analysis of IL-33+ mSCs from 16-week-old male Fndc5+/+ (n = 6) or Fndc5−/− (n = 5) littermates. Left, representative gating strategy for analysing IL-33+ mSCs; right, summary data for numbers of total and fractions of IL-33+ mSCs. Unpaired two-sided t-test was used to calculate P values (****P < 0.0001). b, Flow cytometric analysis of eWAT-Treg cells from 16-week-old male Fndc5+/+ (n = 6) or Fndc5−/− (n = 5) littermates. Left, representative gating strategy for analysing total (black) and ST2+ (red) eWAT-Treg cells; right, summary data for numbers of CD4+ cells, as well as fractions of total and ST2+ Treg cells. Unpaired two-sided t-test was used to calculate P values (**P = 0.0031 and **P = 0.0083). c,d, Eight-week-old male Fndc5+/+ or Fndc5−/− littermates were intraperitoneally (i.p.) injected with PBS or IL-33 (40 ng per gram body weight) twice a week and were euthanized 3 days after the last injection. n = 6–9. c, Experimental scheme. d, Upper, representative gating strategy for analysing total (black) and ST2+ (red) eWAT-Treg cells; Lower, summary data for numbers of CD4+ cells, as well as fractions of total and ST2+ Treg cells. n = 9, 9, 6, 8. One-way ANOVA was used to calculate P values (left: **P = 0.0027 and **P = 0.0037, middle: ****P < 0.0001 and ****P < 0.0001, right: ****P < 0.0001, *P = 0.0102 and ****P < 0.0001). Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustrations in c created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
To determine whether the reduction in ST2+ eWAT-Treg cells in Fndc5−/− mice was primarily due to their deficit in IL-33 expression, we conducted an in vivo complementation experiment. We administered recombinant IL-33 or PBS vehicle to 8-week-old Fndc5−/− and Fndc5+/+ littermates and performed comprehensive cytofluorometric immunophenotyping (Fig. 4c). We chose this young age because it precedes the characteristic expansion of the ST2+ eWAT-Treg compartment beginning at around 12 weeks, thus leaving an open niche for IL-33 to manifest its growth/survival effects. Administering IL-33 to both wild-type and mutant mice led to increases in total CD4+, total Treg and ST2+ Treg cells in eWAT (Fig. 4d). The reduction in ST2+ eWAT-Treg cells observed in PBS-treated Fndc5−/− mice compared with their Fndc5+/+ littermates was completely reversed by IL-33 treatment, suggesting that this cytokine is a key mediator through which irisin regulates the ST2+ eWAT-Treg population. IL-33 failed to promote the accumulation of total Treg cells in iWAT of Fndc5−/− mice, as it did in Fndc5+/+ littermates, although there was an increase in the representation of the ST2+ subtype in both cases (Extended Data Fig. 7b).
In line with this restoration of ST2+ Treg cells in eWAT by IL-33, we observed comparable fold changes of ST2+ conventional (Foxp3−) CD4+ T cells, ILC2s and eosinophils in IL-33-treated versus PBS-treated mice regardless of their FNDC5 status (Extended Data Fig. 7c–e). However, IL-33 had no effect on the accumulation of these IL-33-responsive cell types in iWAT of Fndc5−/− mice (Extended Data Fig. 7f).
Loss of FNDC5 decreases IL-33 levels and exacerbates diabetic phenotypes, while chronic irisin treatment can rescue these effects
Lean FNDC5-deficient mice did not show obvious metabolic defects in our previous studies14. We further investigated the role of irisin in regulating IL-33 and its downstream effects in Fndc5−/− mice challenged by HFD feeding. The same experiment as described in Fig. 1 was performed with diet-induced obese mice that did or did not express FNDC5 (Fig. 5a and irisin levels in Extended Data Fig. 7g). Fndc5−/− mice gained more fat mass after 17 weeks of HFD feeding than control mice, and replenishing irisin via AAV administration prevented further fat deposition due to the absence of FNDC5 (Fig. 5b). Furthermore, Fndc5−/− mice displayed decreased glucose sensitivity and increased resistance to insulin after 16 weeks of HFD feeding, which was almost completely rescued by AAV-mediated chronic irisin treatment (Fig. 5c). Pearson analysis (r = −0.6370, P = 0.0080) revealed an inverse correlation between the area under the ITT curves and body weights (Extended Data Fig. 7h). Of note, in the absence of FNDC5, tissue levels of IL-33 in eWAT and plasma were significantly lower, while ectopic irisin expression successfully restored the disrupted IL-33 levels (Fig. 5d). These data collectively demonstrate that irisin, the functional moiety cleaved from FNDC5, is sufficient to complement ablation of the Fndc5 gene by stimulating IL-33 production and promoting a healthy metabolic tenor in obese mice.
a, Schematic of HFD feeding experiment with male FNDC5 null mice. Four-week-old Fndc5+/+ or Fndc5−/−littermates were injected with AAV-irisin or AAV-GFP via tail vein and then started HFD feeding after 5 weeks. Experiments were performed at the indicated time points. b, Body composition analysis by MRI after 17 weeks of HFD feeding (n = 6, 5, 6, 5). Two-way ANOVA was used for calculating P values (*P = 0.0200, ***P = 0.0005 and *P = 0.0357). c, GTT and ITT analysis of the indicated mice fed a HFD for 16 weeks (n = 4, 5, 7, 8). Two-way ANOVA was used to calculate P values (GTT: ***P = 0.0001, ****P < 0.0001 and ****P < 0.0001; ITT: *P = 0.0163, ****P < 0.0001 and ****P < 0.0001). d, ELISA measurements of IL-33 concentration in eWAT lysates (left, n = 7, 6, 7, 4) and in plasma (right, n = 4, 4, 4, 4) from indicated mice fed a HFD for 18 weeks. Brown–Forsythe and Welch ANOVA was used for calculating P values (left: **P = 0.0072, *P = 0.0147 and ***P = 0.0004; right: *P = 0.0498, *P = 0.0399 and **P = 0.0097). Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P ≤ 0.0001. Significant but irrelevant P values are not indicated. Experiments were repeated three times with similar results. Mouse illustration in a created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
Inhibition of IL-33 abolishes irisin-mediated metabolic benefits in HFD-treated mice
To critically examine the role of this irisin–IL-33 axis in improving metabolic defects in the adipose tissues, we inhibited IL-33 function with an IL-33trap protein (Fig. 6a); this is a recombinant soluble ST2-based construct that was previously shown to bind IL-33 and inhibit IL-33-induced inflammatory responses both in vitro and in vivo21,39. The IL-33trap was produced and purified under endotoxin-free conditions from HEK293 cells (Extended Data Fig. 8a,b). We then performed a HFD experiment similar to Fig. 1a along with long-term IL-33trap treatment. As illustrated in Fig. 6a (see irisin levels in Extended Data Fig. 8c), we initiated twice-a-week administration of IL-33trap (100 μg per mouse) immediately after the diet was switched.
a, Schematic of HFD feeding experiment with long-term IL-33trap treatment. IL-33trap treatments (twice a week, 100 μg per mice per time point) started in the same week of diet switch, and the flow analysis was performed with male mice fed a HFD for 18 weeks. b–d, Flow cytometric analysis of the number of total CD4+ T cells (b, left), the fractions of total (b, middle) and ST2+ (b, right) Treg cells, the number of total mSCs (d, left) and the fraction of IL-33+ mSCs (d, right). n = 6, 6, 6, 6. One-way ANOVA was used for calculating P values (b: left: ****P < 0.0001 and ****P < 0.0001; middle: ****P < 0.0001, ****P < 0.0001 and ****P < 0.0001; right: ****P < 0.0001, ****P < 0.0001 and ****P < 0.0001; c: **P = 0.0026 and ****P < 0.0001; d: ****P < 0.0001 and ****P < 0.0001). e, Body composition analysis of the indicated mice by MRI after 18 weeks of HFD feeding (n = 6, 6, 6, 6). One-way ANOVA was used for calculating P values (*P = 0.0108 and *P = 0.0202). f, GTT and ITT analysis of the indicated mice fed a HFD for 17 weeks (n = 6, 6, 6, 6). Two-way ANOVA was used for calculating P values (GTT: ****P < 0.0001 and ****P < 0.0001; ITT: ****P < 0.0001 and ****P < 0.0001). g, Energy expenditure measured in CLAMS cages over one week. n = 4, 4, 4, 4. The last 4 days of measurement are shown on the left and the averaged values in the light and dark periods are summarized on the right. Two-way ANOVA was used for calculating P values (P values between AAV-irisin PBS and AAV-irisin IL-33trap: left: **P < 0.0046, *P < 0.0162, **P < 0.0036 nd **P < 0.0011; right: **P = 0.0045 and *P = 0.0285). h, RT–qPCR measuring mRNAs associated with thermogenesis and adipogenesis in iWAT of the mice that received AAV-irisin or AAV-GFP, treated with IL-33trap and fed a HFD for 18 weeks (n = 3, 3, 3, 3). Two-way ANOVA was used for calculating P values (P values between AAV-GFP PBS and AAV-irisin PBS, as well as AAV-irisin PBS and AAV-irisin IL-33trap: Fabp4: *P = 0.0222 and *P = 0.0488; Ucp1: ****P < 0.0001 and ****P < 0.0001; Alpl: **P = 0.0064 and *P = 0.0127; Dio2: ****P < 0.0001 and ****P < 0.0001; Elovl3: ****P < 0.0001 and ****P < 0.0001; Cox8b: **P = 0.0027 and **P = 0.0063; Cidea: ***P = 0.0003 and ***P = 0.0008). Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustration in a created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
The action of IL-33trap was first evaluated by quantifying immunocyte populations that respond to IL-33 in eWAT. Flow cytometric analysis revealed a complete reversal of the characteristic irisin-induced expansion of the ST2+ Treg eWAT compartment (Fig. 6b). Irisin-mediated contraction of other immunocyte populations was also completely reversed by the IL-33trap (Fig. 6c). Notably, while no significant changes in the number of total mSCs were found, the irisin-mediated increase of IL-33+ mSCs was fully suppressed by IL-33trap, strongly suggesting that the expansion of IL-33+ mSCs involves a positive feedback loop requiring locally active IL-33 (Fig. 6d). Taken together, these results show that long-term IL-33trap treatment inhibits the accumulation of IL-33+ stromal cells and ST2+ Treg cells in eWAT.
The influence of IL-33 inhibition on irisin-mediated improvements in systemic metabolism was studied next. Indeed, without changing lean mass, IL-33trap treatment reversed the irisin-mediated reduction of fat mass and whole-body mass (Fig. 6e and Extended Data Fig. 8d). Similarly, the improvements in glucose homeostasis and insulin sensitivity in response to irisin treatment, as determined by GTTs and ITTs, were abolished completely by IL-33trap in mice fed a HFD for 17 weeks (Fig. 6f). Control PBS-treated mice that received AAV-irisin again exhibited increased energy expenditure and elevated expression of thermogenic signature genes in the iWAT, both of which were ablated by IL-33trap administration (Fig. 6g,h). The food intake and activity of the animals were not significantly altered by IL-33trap (Extended Data Fig. 8e–g). Interestingly, IL-33trap did not alter the fraction of IL-33+ mSCs in the control mice that received AAV-GFP, yet it resulted in a decreased accumulation of eWAT-ST2+ Treg cells (Fig. 6b,d). However, the changes in the metabolic effects mediated by IL-33trap aligned with the changes in IL-33+ mSCs as the metabolic defects did not worsen with IL-33trap treatment in the control GFP group (Fig. 6e–h). The reduction of body weight and iWAT mass in the irisin-treated group was largely gone with IL-33trap treatment (Extended Data Fig. 8h–j).
Irisin improves metabolic health largely through ST2+ Treg cells
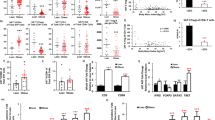
Prior studies show that global ablation of IL-33 results in a significant reduction in the eWAT-Treg compartment, while lymphoid-organ Treg cells are largely unaffected21,23,41. Neutralization of IL-33 selectively inhibited the irisin-induced accumulation of ST2+ Treg cells in eWAT of obese mice without affecting other ST2+ cell types, suggesting that Treg cells might be a key conduit of the irisin–IL-33 axis. To test this hypothesis, we ablated Il1rl1, the gene encoding ST2, specifically in Treg cells by generating Foxp3-Cre.Il1rl1fl/fl mice (referred to as Il1rl1Δtreg mice) and examined their immunological and metabolic properties after HFD feeding (Fig. 7a and irisin levels in Extended Data Fig. 8k). Flow cytometric analysis confirmed a nearly complete absence of ST2+ eWAT-Treg cells in Il1rl1Δtreg mice (Fig. 7b). Importantly, irisin’s ability to suppress the accumulation of immunocytes, particularly CD11Chi macrophages and CD8+ T cells, in eWAT was attenuated in the mutant mice (Fig. 7c). Moreover, the irisin-induced increase of IL-33-producing eWAT mSCs in the wild-type mice was comparable with that in the mutant mice (1.5-fold increase for the wild-type and 1.7-fold increase for the mutant; Fig. 7d).
a, Schematic of HFD feeding experiment with male ST2 Treg null mice (Il1rl1Δtreg) and their littermates (Il1rl1wt). Mice received AAV-irisin or AAV-GFP at 4 weeks old and were then fed a HFD for 17 weeks. GTTs and ITTs were performed with mice fed a HFD for 12 weeks. Flow cytometric analysis was performed with mice fed a HFD for 17 weeks. b–d, Flow cytometric analysis of the number of total Treg cells and the fraction of ST2+ Treg cells (n = 11, 10, 4, 7) (b), the number of total CD45+ cells, the percentages of total and CD11chi macrophages (c, top), the percentages of total T cells, CD8+ T cells and Foxp3−CD4+ cells (c, bottom) and the number of total mSCs and the fraction of IL-33+mSCs (d) One-way ANOVA was used for calculating P values (b: ****P < 0.0001, ****P < 0.0001 and ****P < 0.0001; c, top: ***P = 0.00011, ***P = 0.00074, *P = 0.038; ****P < 0.0001, *P = 0.031 and ****P < 0.0001; c, bottom: ****P < 0.0001, *P = 0.036, ****P < 0.0001, ****P < 0.0001 and **P = 0.0015; d: ****P < 0.0001, *P < 0.027 and *P < 0.018). e, GTT and ITT analyses of the indicated mice fed a HFD for 12 weeks (n = 11, 10, 4, 7). Two-way ANOVA was used for calculating P values (GTT: ****P < 0.0001, and **P = 0.0093; ITT: ****P < 0.0001, and **P = 0.0015). Note, normalized values are presented for this plot because multiple independent cohorts were examined at diverse times. f, RT–qPCR measuring mRNAs associated with thermogenesis and adipogenesis in iWAT of the indicated male mice received AAV-irisin or AAV-GFP, and fed a HFD for 17 weeks (n = 11, 10, 4, 7). Two-way ANOVA was used for calculating P values (P values between Il1rl1wt AAV-GFP and Il1rl1wt AAV-irisin, as well as Il1rl1wt AAV-irisin and Il1rl1Δtreg AAV-irisin: Ppargc1a: ***P = 0.0004 and ***P = 0.0006; Ucp1: ****P < 0.0001 and **P = 0.0043; Ckb: ***P = 0.0002 and NS P = 0.6285; Alpl: **P = 0.0050 and NS P = 0.9976; Dio2: **P = 0.0048 and *P = 0.0123; Elovl3: ****P < 0.0001 and ****P < 0.0001). Mean ± s.d.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Significant but irrelevant P values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustration in a created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
In addition, the absence of ST2+ Treg cells largely reversed the metabolic benefits mediated by irisin. The irisin-induced improvements in insulin sensitivity and glucose tolerance typical of wild-type mice were strongly reduced in littermates lacking ST2+ Treg cells, evidenced by a >50% reduction in both ITTs and GTTs, respectively (Fig. 7e). Expression of Ucp1 and FCC genes were further induced by irisin in the absence of ST2+ Treg (2.8-fold versus 3.3-fold for Ucp1, 2.6-fold versus 6.7-fold for Ckb, and 2.3-fold versus 2.8-fold for Alpl; Fig. 7f). Consistent with Fig. 1i, irisin had no impact on the thermogenic gene expression in eWAT of either Il1rl1Δtreg or Il1rl1wt mice, although several adipogenic genes (Fabp4, Adipoq and Lep) were upregulated in Il1rl1Δtreg mice as expected42 (Extended Data Fig. 8l).
Irisin’s effects on obesity in a therapeutic setting
To assess irisin’s effects in a therapeutic setting, we fed 6-week-old male C57BL/6 mice on a HFD. We then administered AAV-irisin or AAV-GFP to them after 14 weeks of HFD feeding, and analysed them 10 weeks after AAV administration (Extended Data Fig. 9a and irisin levels in Extended Data Fig. 9b). We did not observe significant differences between the two groups in body and fat mass, GTT, ITT, energy expenditure, thermogenic gene expression in iWAT and immunocyte profiles in eWAT (Extended Data Fig. 9c–o). Given that this protocol did not result in irisin-induced expansion of total or ST2+ Treg populations, the absence of irisin-induced metabolic benefits, at least in eWAT, is to be expected. The lack of Treg expansion likely reflects the very few ST2+ Treg cells available for expansion at the time of irisin administration, both because few eWAT-Treg cells existed at the earlier onset of HFD feeding20 and because of the prolonged HFD exposure in the absence of irisin.
The role of the IL-33–ST2 axis in irisin-mediated effects in iWAT
As we observed increased lL-33 levels in iWAT without changes in the representation of total or ST2+ Treg populations (Extended Data Fig. 3e), we examined the role of IL-33 in irisin-mediated thermogenesis using primary iWAT cultures. The stromal vascular fraction (SVF) was isolated from iWAT of 4-week-old male C57BL/6 mice, and the culture was treated with irisin/HSP90α either before induction of adipocyte differentiation or throughout the culture at the indicated time points (Extended Data Fig. 10a). Total RNAs were extracted 4 h after norepinephrine (a hormone molecule that potently induces thermogenesis) treatment on the last day. IL-33 mRNA levels were significantly upregulated by irisin treatment compared with PBS treatment for both methods. However, continuous irisin treatment showed a much more significant increase, which cannot be further induced by noradrenaline treatment. ST2 is expressed at the cell surface during adipogenesis of 3T3-L1 cells, with mRNA level peaking on day 2 after induction of differentiation and the protein level accumulating in the mature adipocytes41. We then tested levels of Il1rl1 and Il33 transcripts and the signature genes of different thermogenic pathways (Ucp1 and Alpl) in different batches of our iWAT cultures. Il1rl1 transcripts levels seemed to be positively correlated with transcripts encoding IL-33 and TNAP, and negatively correlated with transcripts encoding UCP1 (Extended Data Fig. 10b). Furthermore, a low dose of recombinant IL-33 (1 ng ml−1) significantly induced Alpl expression in differentiated iWAT cultures (Extended Data Fig. 10c). Therefore, we proposed a mechanistic model underpinning irisin action through the IL-33–ST2 axis in iWAT based on these ex vivo data (Fig. 8).
Adipose tissues in male obese mice secrete irisin co-factor under stress, which activates irisin integrin receptor on the mSCs. Irisin binds and signals through the activated receptor for IL-33 production. In visceral fat tissue, the secreted soluble form of IL-33 acts on ST2+ Treg cells to protect the visceral fat Treg population and keep the visceral fat inflammation in check; in SAT, IL-33 targets the ST2+ adipocytes (and/or other ST2+ cell types) to stimulate thermogenesis and control the fat mass. None of the data exclude the possibility of the role of elevated circulating IL-33 in mediating effects in SAT. APCs, adipose precursor cells. Cell illustrations created in BioRender: top row, A, M. https://biorender.com/syavjpa (2026); bottom row, A, M. https://biorender.com/ywnwq02 (2026).
Discussion
Previous studies have shown beneficial effects of irisin on obese mice and ex vivo slices of human adipose tissues11,43,44. However, no clear mechanism of action was elucidated, except that the expression of genes of the classical thermogenic pathway, like UCP1, was elevated. Here, we demonstrate that chronic irisin treatment restrains weight gain in mice exposed to a HFD, as it markedly improves glucose intolerance and insulin resistance. An interesting and unusual feature of this response is that a divergence in both body weight and metabolic dysfunction does not take place for the first 8 weeks of exposure to HFD, because this time course correlates roughly with the development of substantial adipose tissue inflammation. At a later stage of HFD treatment (18 weeks), body weights diverged without changes in lean mass. Although body weight and fat mass correlate with improved metabolic indices, changes in adiposity under control of irisin may not reflect the full extent of the irisin-mediated increase of glucose tolerance and insulin sensitivity, as irisin treatment still increases these metabolic effects compared with the weight-matched GFP-treated mice (Extended Data Fig. 1I). So far, there is no acceptable quantitative way to address the direct relationship of body weight and fat mass to the metabolic changes. We then asked whether irisin might affect these critical metabolic parameters through actions on immunocytes in the adipose tissue itself. Numerous studies have shown that modulation of the immunocyte complement can have profound effects on metabolic disease in experimental models36,40.
We also assessed the effects of chronic irisin treatment in healthy lean mice. Our prior mechanistic study on irisin–integrin interaction demonstrated that irisin action requires stress-induced co-factor HSP90α, suggesting that irisin would function only when there is a substantial challenge, such as in the diseased state16. In this study, the observation that irisin did not further improve metabolic health of the lean mice supports these previous findings.
Evidence presented here indicates that irisin-stimulated improvements in obesity and diabetes require processed soluble IL-33, most of which appears to come from stromal cells in the eWAT45. Moreover, irisin can act directly on these stromal cells in vitro (Fig. 3e), activating their IL-33 production and secretion, providing a direct link between irisin and a critical immunomodulatory pathway. Such immunological changes are not associated with body weight or tissue mass changes (Extended Data Figs. 1c,i,j and 8i). IL-33 itself has been previously shown to play a role in adipose tissue biology; for instance, global and mSC-specific ablation of IL-33 resulted in a significant decrease of Treg cells in eWAT, and administration of external IL-33 to obese mice reduced VAT inflammation and improved metabolic indices21,23,25,46. As shown in this study, IL-33 clearly functions on adipose tissues in at least two distinct ways. First, IL-33 acts on ST2+ Treg cells to promote their survival. Treg cells are critical to suppress inflammation in adipose tissues, especially in VAT, and are also known to have direct impacts on adipocytes and their precursors (for example, on lipolysis and terminal maturation), and thereby systemic metabolism42,47,48,49,50. Secondly, IL-33 neutralization appears to have metabolic effects that are greater than those that can be explained by its actions on Treg cells. This observation suggests that IL-33 might also act directly on mature adipocytes, especially in the iWAT.
In iWAT, irisin induces thermogenesis through IL-33 (Fig. 6h), and irisin upregulates IL-33 protein levels in iWAT (Extended Data Fig. 5d). It is very likely that iWAT IL-33 is produced by stromal adipocyte precursors analogous to those in eWAT (IL-33− VmSCs). However, we did not observe a significant increase in the total or ST2+ Treg populations in iWAT with irisin treatment. This may reflect a deficiency in ST2 co-receptors or their downstream signalling, or inhibition of ST2 and its wired signalling in iWAT given different cytokine milieux at the different fat depots. Of note, HFD feeding did not significantly reduce these Treg populations (Fig. 2i and Extended Data Fig. 3e), making irisin-mediated Treg changes difficult to be detected. Extended Data Fig. 10 and refs. 51,52 suggest that IL-33 might directly act on ST2+ adipocytes to induce thermogenesis in iWAT.
In this study, we primarily investigated the effects of irisin in obesity and diabetes in male mice. It has been previously established that female mice are protected against HFD-induced metabolic syndrome53,54. This is unlike humans, where both males and females show adipose tissue inflammation in obesity55.
The mechanisms uncovered here have important implications for other tissues and other diseases. Chronic irisin treatments prevent neurodegeneration in multiple models of Alzheimer’s disease and Parkinson’s disease. Some of these effects appear to be through direct actions on neurons and glial cells31,56,57,58. However, IL-33, expressed mainly in endothelial cells, astrocytes and oligodendrocytes in the brain, can regulate cognition59. Moderate increases in IL-33 levels reversed memory deficits in APP/PS1 mice (an Alzheimer’s disease mouse model), whereas administration of IL-33 in wild-type mice induced neuroinflammation that further led to impaired memory60,61. Whether similar mechanisms demonstrated in this study can contribute to some of the irisin-mediated benefits in neurodegeneration will be important to investigate.
It will also be important to understand how irisin increases IL-33 expression in stromal cells. While it has been reported that the adipocyte precursor cells that respond to irisin abundantly express integrin αV, β1 and β5 (ref. 62), the subclass of integrins that function as irisin receptors in this context remain to be determined.
Lastly, the ability of irisin to affect body weight and metabolic dysfunction without decreasing food intake, suggests an approach different from the current glucagon-like peptide-1-based drugs, which have primary effects on food intake. Whether irisin can be used as a complement to these other therapies remains to be determined.
Methods
Mouse strains and husbandry
Mouse husbandry and experimentation were performed according to protocols approved by the Institutional Animal Care and Use Committee of the Beth Israel Deaconess Medical Center under protocol 072-2020-23 and Harvard Medical School under protocol 1257. Male mice were used for this study. Unless otherwise noted, mice were housed at 22 °C with 40–50% humidity under a 12-h light–dark cycle, with free access to food and water. Mice were fed with either normal chow (Formulab Diet, 5008) or a HFD (Research Diets, D12492i). All strains were on a C57BL/6J background unless otherwise stated. Fndc5 floxed mice were developed with the Texas A&M Institute for Genomic Medicine and crossed with EIIa-cre mice to generate germline deletion of Fndc5 (exons 2 and 3). Experiments were performed with sex- and age-matched global FNDC5-deficient and littermate wild-type control mice. B6.Foxp3-Cre (016959) mice were purchased from The Jackson Laboratory. Il1rl1fl mice were donated by V. Kuchroo. B6.Foxp3-Thy1.1 mice were obtained from A. Rudensky. B6.Il33-Egfp mice were donated by P. Bryce (now commercially available at The Jackson Laboratory with stock no. 30619). Il1rl1Δtreg mice were generated by crossing Il1rl1fl mice with B6.Foxp3-Cre mice, and this mouse line was bred and utilized in the animal facility of the New Research Building at Harvard Medical School. The age, strain and number of mice used in the individual experiments are noted in the corresponding figure legends.
Production of AAV
Mouse irisin (open reading frame 1–140, containing the N-terminal signal peptide) plus a five-amino-acid linker and a C-terminal FLAG-tag was cloned into the pENN.AAV.CB7.CI.pm20d1flag.WPRE.rBG vector (Addgene plasmid no. 132682) to replace pm20d1flag (pENN.AAV.CB7.CI.PM20D1flag.WPRE.rBG was a gift from J. Long; 132682). The final plasmid was sequenced to confirm the correct insertion of irisin open reading frame. AAV (serotype 8) was packaged at the Boston Children’s Hospital Viral Core with a titre of 3.15 × 1013 genome copies (GCs) per ml. AAV8-GFP (pENN.AAV.CB7.CI.eGFP.WPRE.rBG) was used as control, obtained from Addgene (105542), and packaged at the Boston Children’s Hospital Viral Core with a titre of 2.10 × 1013 GCs per ml.
Tail vein injections
Mice were injected into the tail vein with AAV8-GFP or AAV8-irisin-FLAG (1 × 1010 GCs per mouse) diluted in PBS to a final volume of 100 μl.
IL-33 treatment
Eight-week-old Fndc5−/− male mice and their wild-type littermates were administered recombinant mouse IL-33 (BioLegend, 580506; 200 ng per mouse in a total volume of 100 μl) or PBS (100 μl) via intraperitoneal injection on day 0 and day 4. Tissues were analysed on day 7.
IL-33trap treatment
Nine-week-old male C57BL/6 mice were injected with AAV8-irisin or AAV8-GFP and switched to a HFD (day 0). The IL-33trap (100 µg per mouse with a total volume of 100 µl) or PBS (100 µl) were injected intraperitoneally on day 2 and day 5. Injections were continued twice a week for a total of 18 weeks until the animals were euthanized.
ELISA
For irisin-FLAG ELISA, blood was collected via the submandibular vein in heparin-coated tubes (365985, BD Microtainer) and then centrifuged. The plasma fraction was collected and stored at −80 °C until analysis. Ninety-six-well plates (R&D, DY990) were coated with an anti-irisin capture antibody (R&D, MAB8880) in PBS overnight at 4 °C. The next day, plates were washed four times with 0.1% PBST and blocked with 1% BSA for 1 h at room temperature (RT). After four washes with 0.1%, PBST, standards (0–1,000 ng ml−1) and plasma samples were added to the plate and incubated for 2 h at RT. Plates were incubated with an anti-FLAG detection antibody for 2 h in RT after washing 4× with 0.1% PBST. Next, the samples were incubated in horseradish peroxidase-linked secondary antibody for 30 min following 4× PBST wash. 3,3’,5,5’-Tetramethylbenzidine chromogen (ab171522, Abcam) was used as a detection system and, finally, the absorbance was measured at 450 nm after adding stop solution (ab171529, Abcam). Measurement was done using a plate reader (FLUOstar Omega, BMG Labtech). Irisin-FLAG concentration was quantified from a standard curve of recombinant irisin-FLAG (AG-40B-0136-C010, Adipogen).
For the IL-33 ELISA, blood was collected via the submandibular vein in heparin-coated tubes (365985, BD Microtainer) and then centrifuged. The plasma fraction was collected and stored at −80 °C until analysis; adipose tissues were homogenized in RIPA buffer (Sigma) and centrifuged at 45,000g for 30 min at 4 °C. Supernatants were taken and measured using the pre-coated 96-well microplate as outlined in the manufacturer’s protocol of the mIL-33 kit (Abcam, ab213475) and was read on a plate reader (FLUOstar Omega, BMG Labtech).
Recombinant protein production
Mouse IL-33trap protein was produced in suspension HEK293 cells (Expi293F cells; Life Technologies, A14527). The cell line was authenticated by maintaining cells below 5 × 106 cells per ml with viability > 95%. Mouse IgG1 control mammalian expression plasmid was used for transient transfection, and the produced IgG1 in the cell culture medium with glycosylation was probed using western blot and SDS–PAGE followed by silver staining. Mycoplasma contamination was tested on a monthly basis to ensure the cell line is mycoplasma free. Expi293F (1 l) cells grown in expression medium (Life Technologies, A1435101) at a density of 2.8 × 106 were transfected with 1 mg pEF-mIL-33-trap DNA plasmid (gift from Beyaert laboratory in Belgium)39, and 3 mg sterile 25-kDa linear PEI mix in Opti-Plex Complexation Buffer (Life Technologies A4096801). Proteins were expressed at 37 °C, 8% CO2, >80% humidity with shaking at 125 rpm for 4 days. Enhancers (Life Technologies, A14524) were added 22 h after transfection to boost protein expression. For IL-33trap protein purification from the Expi293F culture, all the following steps were performed at 4 °C or on ice. Cells were pelleted at 600g for 20 min, and the medium was subjected to an additional 2 h of centrifugation at 1,000g. The supernatant was filtered through a 0.22-μm filter unit and was concentrated tenfold using the Tangential Filtration System with a 50,000-Da molecular weight cut-off (Paul) before being applied to a Ni-Excel affinity column (Cytiva, 17371201) equilibrated in phosphate buffer (10 mM HEPES pH 7.4, 150 mM NaCl, 20 mM imidazole). After thorough washes with wash buffer (10 mM HEPES pH 7.4, 150 mM NaCl, 30 mM imidazole), the column was eluted with elution buffer (10 mM HEPES pH 7.4, 150 mM NaCl, 500 mM imidazole). The eluent was concentrated with a using an Amicon Ultra-15 filter Unit (Millipore) with a 50,000-Da molecular weight cut-off before further purification through a Superdex 200 10/300 GL gel-filtration column (Cytiva, 17517501) equilibrated with PBS buffer. The level of endotoxin was measured using the Toxin Sensor Chromogenic LAL Endotoxin Assay Kit (GeneScript, L00350). Protein was further concentrated to >1 mg ml−1 before being aliquoted, frozen in liquid nitrogen and stored at −80 °C. In all cases, the HEPES pH given is at 23 °C.
Irisin was purified from Expi293 cells, and HSP90α was purified from Escherichia coli as previously described16.
mRNA expression analysis
Total RNA was isolated from inguinal fat or epididymal fat tissues using TRIzol reagent (Invitrogen, 15596018) and RNeasy Mini purification kit (Qiagen, 74104) according to the manufacturer’s protocol. Tissues were homogenized in TRIzol reagent using a bead homogenizer for 20 min at maximum speed (Qiagen, TissueLyser II). DNA was digested on a column using the RNase-Free DNase Set (Qiagen, 79254). RNA was reverse transcribed using the High-Capacity cDNA Reverse Transcription kit with RNase Inhibitor (Applied Biosystems, 4374966). The resulting cDNA was analysed by RT–qPCR using SYBR green fluorescent dye 2× qPCR master mix (Promega) in a QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems). Briefly, cDNA was mixed with 250–500 nmol primers and the GoTaq qPCR System (Promega, A6002). Relative mRNA levels of the gene of interest were normalized to the mRNA level of Rplp0, and fold change was calculated using the ΔΔCt method. Unless stated otherwise, primer sequences (Supplementary Fig. 2) were chosen from PrimerBank63,64,65,66 and have been validated in ref. 16.
Whole-tissue and populational RNA-seq library preparation and data analysis
For adipose tissue RNA-seq, RNA integrity was assessed using the Bioanalyzer 2100 system (Agilent Technologies). Messenger RNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first-strand cDNA was synthesized using random hexamer primers, and then the second-strand cDNA synthesis was either the same way as the first strand (for non-strand-specific library) or synthesized using dUTP, instead of dTTP to generate the directional library (for strand-specific library). The library was ready after end repair, A-tailing adaptor ligation, size selection, amplification and purification. The library was checked with Qubit and real-time PCR for quantification and bioanalyzer for size distribution detection. After library quality control, different libraries were pooled based on the effective concentration and targeted data amount, then subjected to Illumina sequencing (Illumina NovaSeq 6000). The basic principle of sequencing is ‘Sequencing by Synthesis’, where fluorescently labelled dNTPs, DNA polymerase and adaptor primers are added to the sequencing flow cell for amplification. As each sequencing cluster extends its complementary strand, the addition of each fluorescently labelled dNTP releases a corresponding fluorescence signal. The sequencer captures these fluorescence signals and converts them into sequencing peaks through computer software, thereby obtaining the sequence information of the target fragment.
Raw data (raw reads) of FASTQ format were firstly processed through in-house Perl scripts. In this step, clean data (clean reads) were obtained by removing reads containing adaptors, reads containing ploy-N and low-quality reads from raw data. At the same time, Q20, Q30 and GC content of the clean data were calculated. All the downstream analyses were based on the clean data with high quality. Reference genome and gene model annotation files were downloaded from the genome website directly. An index of the reference genome was built using Hisat2 v2.0.5, and paired-end clean 1 reads were aligned to the reference genome (GRCm38/mm10) using Hisat2 v2.0.5. We selected Hisat2 as the mapping tool because Hisat2 can generate a database of splice junctions based on the gene model annotation file and thus a better mapping result than other non-splice mapping tools. FeatureCounts v1.5.0-p3 was used to count the read numbers mapped to each gene. And then fragments per kilobase of transcript per million mapped reads of each gene was calculated based on the length of the gene and reads count mapped to this gene. The expected number of fragments per kilobase of transcript sequence per millions base pairs sequenced considers the effect of sequencing depth and gene length for the reads count at the same time, and is currently the most commonly used method for estimating gene expression levels.
Differential gene expression testing was performed using the DESeq2 R package (1.20.0). Differential expression analysis for two conditions/groups was performed. DESeq2 provides statistical programmes for determining differential expression in digital gene expression data using models based on negative binomial distribution. The resulting P value is adjusted using the Benjamini and Hochberg method to control the error discovery rate. The corrected P value ≤ 0.05 and |log2(fold change)| ≥ 1 was set as the threshold of significant differential expression. KEGG is a database resource for understanding high-level functions and utilities of the biological system, such as the cell, the organism and the ecosystem, from molecular-level information, especially large-scale molecular datasets generated by genome sequencing and other high-throughput experimental technologies (https://www.genome.jp/kegg/). We used cluster Profiler R package to test the statistical enrichment of differentially expressed genes in KEGG pathways.
For mSCs, 1,000 cells were lysed in 5 μl Buffer TCL (QIAGEN) with 1% 2-mercaptoethanol (Sigma). Library construction and sequencing as well as data processing were performed according to standard Immunological Genome Project protocols (https://www.immgen.org/img/Protocols/ImmGenULI_RNAseq_methods.pdf). Smart-seq2 libraries were prepared and sequenced as previously described. Briefly, RNA was captured and purified using RNAClean XP beads (Beckman Coulter), and polyadenylated mRNA was selected using an anchored oligo(dT) primer (5´-AAGCAGTGGTATCAACGCAGAGTACT30VN-3´). Polyadenylated mRNA was converted to cDNA by the reverse-transcription reaction followed by limited PCR amplification of first-strand cDNA. The Nextera XT DNA Library Preparation Kit (Illumina) was used for Tn5 transposon-based fragmentation followed by PCR amplification for an additional 12 cycles using barcoded primers such that each sample carried a specific combination of Illumina P5 and P7 barcodes. Paired-end sequencing was performed on an Illumina NextSeq 500 (two full NextSeq runs per plate for an average of 10 M raw reads per sample) using two 38-base-pair reads with no further trimming. Reads were aligned to the mouse genome (GENCODE GRCm38/mm10 primary assembly and gene annotations vM16) using STAR 2.7.3a. Transcripts annotated as ribosomal RNA were removed, and gene-level quantification was calculated using the Subread 2.0 command featureCounts. The DESeq2 package from Bioconductor was used to normalize raw read counts according to the median of ratios method (samples with <1 M uniquely mapped reads or with fewer than 8,000 genes with over 10 reads were excluded from normalization). Normalized data were converted to GCT and CLS files, which were used in downstream analyses. Additional quality control after normalization included removal of biological replicates with a poor Pearson’s correlation (<0.9) and/or poor congregation by principal component analysis of the top 1,000 variable genes. Normalized reads were further filtered by minimal expression over 10. Data were analysed by Multiplot Studio in the GenePattern software package. The adipogenesis up-signature came from gene-set enrichment analysis.
Cell isolation and flow cytometry
Adipose tissue was dissected, minced and digested for 20 min with 1.5 mg ml−1 collagenase type II (C6885, Sigma) in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 2% FCS in a 37 °C water bath with shaking. The digested material was filtered through a 100-μm nylon cell strainer, digested with ammonium–chloride–potassium lysing buffer, and then filtered through a 40-μm nylon cell strainer. The SVF was collected after centrifugation at 650g for 5 min. For immunocyte analysis, cells were stained with anti-CD45 (30-F11), anti-TCRβ (H57-597), anti-CD4 (RM4-5), anti-CD8a (53-6.7), SiglecF (S17007L) and anti-Thy1.1 (OX-7) monoclonal antibodies (all from BioLegend); anti-ST2 (RMST2-2) (eBioscience); and LIVE/DEAD Fixable Violet Dead Cell Stain Kit (Invitrogen) or DAPI. For mSC analysis, cells were stained with anti-CD45 (30-F11), anti-CD31 (390), anti-PDGFRα (APA5), anti-Sca-1 (D7) monoclonal antibodies (all from BioLegend); and LIVE/DEAD Fixable Violet Dead Cell Stain Kit (Invitrogen) or DAPI. For intracellular staining, cells were fixed, permeabilized and incubated with anti-Foxp3 (FJK-16s, eBioscience) monoclonal antibody at RT for 30 min, or anti-IL-33 (AFS626, R&D) monoclonal antibody at 4 °C overnight followed by donkey anti-goat IgG secondary antibody (Jackson ImmunoResearch) at RT for 1 h according to the manufacturer’s instructions (eBioscience). Cells were acquired using FACSymphony A5 flow cytometers (BD Biosciences) and were sorted using a FACSAria (BD) cell sorter. Data were analysed using FlowJo 10 software. Detailed antibody information is listed in Supplementary Fig. 3.
Cell culture
Sorted mSCs from eWAT of 12-week-old Il33-Egfp mice were cultured in growth medium (DMEM/F12 supplemented with 10% FBS and 1% penicillin–streptomycin) with PBS or HSP90α for 30 min followed by irisin for 3 days. Supernatant from the cultured mSCs was collected as mSC conditioned medium. For integrin agonist antibody treatment, 1 h before irisin/HSP90α or PBS treatment, the cultures were pretreated with 1 ng ml−1 of a control IgG (Abcam, 206200) or anti-αVβ5 antibody (Abcam, 177004).
eWAT-Treg cells were sorted from 12-week-old Foxp3-Thy1.1 mice, cultured in RPMI 1640 supplemented with 10% FBS, 50 μM β-mercaptoethanol, 1% penicillin–streptomycin, 2,000 U ml−1 human recombinant IL-2 (200-02, PeproTech) and were stimulated with anti-CD3/anti-CD28 dynabeads (11452D, Thermo Fisher) in the presence of conditioned medium treated with either control IgG or anti-IL-33 antibody (2 μg ml−1, AG-27B-0013PF-C100, AdipoGen Life Sciences) from mSCs for 3 days at 37 °C in a 5% CO2 atmosphere.
For the primary iWAT culture, the SVF of iWAT was isolated from 4-week-old male mice. iWAT was dissected and washed in Hank’s Buffered Saline Solution (HBSS) without calcium or magnesium (Corning). It was then minced and digested in HBSS containing 10 mg ml−1 Collagenase D (Roche), 3 U ml−1 Dispase II (Roche) and 10 mM CaCl2 for 45 min at 37 °C. After digestion, the cell suspension was combined with adipocyte culture medium (DMEM/F12 supplemented with 2.5 mM L-alanyl-L-glutamine, 10% FBS (GeminiBio), 100 U ml−1 penicillin (Gibco), 100 ug ml−1 streptomycin (Gibco) and 0.1 mg ml−1 Primocin; Invivogen) and filtered through a 100-mm cell strainer. Cells of the SVF were pelleted by centrifugation at 600g for 5 min, after which they were resuspended in adipocyte culture medium, filtered through a 40-mm cell strainer, pelleted as above and resuspended in adipocyte culture medium. The cells were plated on tissue culture plates and cultured at 37 °C with 10% CO2. Primocin was removed from the culture medium after 4 days and withheld during subsequent differentiation and experiments. Preadipocytes were cultured until 2 days after reaching confluency, then differentiated by addition of fresh adipocyte culture medium supplemented with 1 mM rosiglitazone (Cayman Chemical), 0.5 mM isobutylmethylxanthine (Sigma), 1 mM dexamethasone (Sigma) and 870 nM insulin (Sigma). Two days later (‘day 2’), and every 2 days thereafter, medium was replaced with fresh adipocyte culture medium supplemented with 1 mM rosiglitazone and 870 nM insulin. Beginning on day 8 (day 4 of differentiation), rosiglitazone was omitted from the medium. Depending on assay, experiments were performed at day 10 (day 6 of differentiation) as noted in the individual experiments. Cells were treated with 1 ng ml−1 mouse IL-33 (BioLegend, 580506) as indicated and analysed 4 h after 500 nM norepinephrine (Sigma, A9512) treatment.
Immunofluorescence imaging and analysis
Cells were plated on 35-mm Matek dishes (Thermo Fisher) and were fixed by 4% paraformaldehyde (1 volume of 16% paraformaldehyde (Electron Microscopy Sciences) diluted with three volumes of PBS) for 15 min at RT, washed three times with PBS, permeabilized by permeabilization solution (PBS supplemented with 0.25% Triton X-100) for 10 min at RT and blocked by blocking solution (PBS supplemented with 0.1% Triton X-100 and 5% FBS) for 1 h at RT. For immunofluorescence staining, anti-pFAK (rabbit, Invitrogen 700255) and anti-FAK (mouse) antibodies (mouse, Invitrogen 39-6500) at a dilution of 1:500 in blocking solution were used for primary labelling of phosphorylate FAK and total FAK, respectively; and anti-rabbit Alexa Fluor 488 (Invitrogen A-11008) and anti-mouse Alexa Fluor 568 (Invitrogen A-11004) at a dilution of 1:1,000 were used for secondary labelling. DAPI (3 μM; BioLegend) was used to stain nuclei. All fluorescence images for whole-dish scans were acquired at the DFCI Molecular Imaging Core using LASX Office 1.4.5 27713 on a Leica THUNDER Imager wide-field microscope. A Zeiss 980 confocal/spectral/Airyscan microscope was used to acquire high-magnification images. Nikon Elements 4.2 and Fiji67 were used for image processing and profile quantification. For IL-33-GFP-positive cell quantification, three dishes for each condition were scanned, and total GFP intensity was normalized against total DAPI intensity in each large image, and different conditions were normalized against PBS control. For pFAK/FAK ratio quantification, cells in different regions of interest were analysed, and ratiometric analysis was performed using total pFAK and FAK intensity from each cell.
Body composition measurement
Fat and lean mass composition of living mice was assessed using a 3-in-1 Echo MRI Composition analyzer (Echo Medical Systems).
Diet-induced obesity
Age-matched littermates and group housing were used for high-fat feeding experiments. At 8 weeks of age, mice were given a 60 kcal% fat rodent diet (D12492, Research Diets) ad libitum. Mouse weight and consumed food weight were assessed once per week.
Intraperitoneal GTT
Mice were fasted overnight, after which their weight and fasting blood glucose values were measured using tail nick blood and a OneTouch UltraMini glucose meter. Glucose was administered by intraperitoneal injection at a dose of 1 g per kilogram body weight, followed by blood glucose measurements over the next 2 h (0, 30, 60, 90 and 120 min).
Intraperitoneal ITT
Mice were fasted for 4 h, after which their weight and fasting blood glucose values were measured using tail nick blood and a OneTouch UltraMini glucose meter. Human insulin (Humulin R U-100 Insulin; HI-210) was administered via intraperitoneal injection at a dose of 1 U per kilogram body weight, followed by blood glucose measurements over the next 2 h (0, 30, 60, 90 and 120 min).
Indirect calorimetry
Mice were individually housed in Promethion metabolic cages (Sable Systems International) at 23 °C with a 12-h light–dark cycle with ad libitum food and water consumption. Mice were fed a 60 kcal% fat rodent diet for the duration of the experiment. After at least 2 days of acclimatization to the cages, energy expenditure, VO2, VCO2, food intake and movement were assessed using the Sable Systems’ Promethion apparatus, comprising a rack of 16 metabolic cages and another of 8 metabolic cages. This apparatus was enclosed in a temperature-controlled chamber, and during the experiment the temperature was adjusted as depicted in the graphs at RT (23 °C). For the IL-33trap experiment, mice were injected intraperitoneally with IL-33trap (100 μg per mouse) or the same volume of PBS (100 μl) as depicted. Macro 13 (data were binned every 30 min), from the ExpeData software system, was used to export metabolic variables of interest at each reading for each cage. Data collected by Promethion metabolic cages were analysed using CalR (version 1)68 and normalized by body weight.
Histology
Tissues were fixed in formalin fixative and embedded in paraffin and then passed over to Harvard Medical School Rodent Histopathology Core for tissue sectioning, staining and quantification. Fifty-micron-thick sections were cut, baked at 60 °C for 1 h, deparaffinized in xylene, rehydrated in a graded ethanol series and then stained. H&E staining was performed with Harris’ haematoxylin for 30 s and eosin for 2 min. Images were acquired at DFCI Molecular Imaging Core using LASX Office 1.4.5 27713 on a Leica THUNDER Imager wide-field microscope. Zen 3.7 microscopy software was used for image processing and profile quantification. Images were processed in the same way and made into masks using CellProfiler.33 for automated quantification. Cropped images for presentation were prepared using Fiji67. CLSs were quantified automatically from blinded images, and their density was expressed as cells or CLS per mm2 of tissue section.
Statistical analysis
Replicate numbers are described in the figure legends. For cellular assays, n corresponds to the number of experimental replicates (for example, independent treatments). For animal assays or tissue extracted from animals, n corresponds to the number of mice used per genotype or condition. Sample sizes were determined on the basis of previous experiments using similar methodologies. Unless otherwise stated, data are presented as the mean, and error bars indicate the standard error. Graphing and statistical analyses, including two-tailed Student’s t-test, one-way ANOVA and Fisher’s least significant difference test, were performed using Prism (GraphPad). To assess the significant enrichment of gene signatures in RNA-seq datasets, we used a chi-squared test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The whole-tissue and populational RNA-seq data have been deposited to the Gene Expression Omnibus database under GSE283234 for the 27-week-old HFD-fed (18 weeks of HFD feeding) male C57BL/6 mouse adipose tissue dataset and GSE282942 for the eWAT mSC (isolated from 10-week-old male C57BL/6 mice) dataset. Raw microscopy images are deposited on Figshare (https://doi.org/10.6084/m9.figshare.31151524)69. Source data are provided with this paper.
References
Ward, Z. J. et al. Projected U.S. state-level prevalence of adult obesity and severe obesity. N. Engl. J. Med. 381, 2440–2450 (2019).
Kong, Y. et al. Obesity: pathophysiology and therapeutic interventions. Mol. Biomed. 6, 25 (2025).
Hotamisligil, G. S. Inflammation, metaflammation and immunometabolic disorders. Nature 542, 177–185 (2017).
Zatterale, F. et al. Chronic adipose tissue inflammation linking obesity to insulin resistance and type 2 diabetes. Front. Physiol. 10, 1607 (2020).
Gómez-Ambrosi, J. Adipose tissue inflammation. Cells 12, 1484 (2023).
Villarroya, F., Cereijo, R., Gavaldà-Navarro, A., Villarroya, J. & Giralt, M. Inflammation of brown/beige adipose tissues in obesity and metabolic disease. J. Intern. Med. 284, 492–504 (2018).
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44 (2014).
Kajimura, S., Spiegelman, B. M. & Seale, P. Brown and beige fat: physiological roles beyond heat generation. Cell Metab. 22, 546–559 (2015).
Celik, O. & Yildiz, B. O. Obesity and physical exercise. Minerva Endocrinol. 46, 131–144 (2021).
Thyfault, J. P. & Bergouignan, A. Exercise and metabolic health: beyond skeletal muscle. Diabetologia. 63, 1464–1474 (2020).
Boström, P. et al. A PGC1-α-dependent myokine that drives brown-fat-like development of white fat and thermogenesis. Nature 481, 463–468 (2012).
Lee, P. et al. Irisin and FGF21 are cold-induced endocrine activators of brown fat function in humans. Cell Metab. 19, 302–309 (2014).
Jedrychowski, M. P. et al. Detection and quantitation of circulating human irisin by tandem mass spectrometry. Cell Metab. 22, 734–740 (2015).
Kim, H. et al. Irisin mediates effects on bone and fat via αV integrin receptors. Cell 175, 1756–1768 (2018).
Zhang, X., Hu, C., Wu, H. M., Ma, Z. G. & Tang, Q. Z. Fibronectin type III domain-containing 5 in cardiovascular and metabolic diseases: a promising biomarker and therapeutic target. Acta Pharmacol. Sin. 42, 1390–1400 (2021).
A, M. et al. Irisin acts through its integrin receptor in a two-step process involving extracellular Hsp90α. Mol Cell. 83, 1903–1920 (2023).
Neeland, I. J. et al. Visceral and ectopic fat, atherosclerosis, and cardiometabolic disease: a position statement. Lancet Diabetes Endocrinol. 7, 715–725 (2019).
Mittal, B. Subcutaneous adipose tissue & visceral adipose tissue. Indian J. Med. Res. 149, 571–573 (2019).
Li, C., Spallanzani, R. G. & Mathis, D. Visceral adipose tissue Tregs and the cells that nurture them. Immunol. Rev. 295, 114–125 (2020).
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nat. Med. 15, 930–939 (2009).
Kolodin, D. et al. Antigen- and cytokine-driven accumulation of regulatory T cells in visceral adipose tissue of lean mice. Cell Metab. 21, 543–557 (2015.
Cipolletta, D. et al. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature 486, 549–553 (2012).
Vasanthakumar, A. et al. The transcriptional regulators IRF4, BATF and IL-33 orchestrate development and maintenance of adipose tissue-resident regulatory T cells. Nat. Immunol. 16, 276–285 (2015).
Han, J. M. et al. IL-33 reverses an obesity-induced deficit in visceral adipose tissue ST2+ T regulatory cells and ameliorates adipose tissue inflammation and insulin resistance. J. Immunol. 194, 4777–4783 (2015).
Spallanzani, R. G. et al. Distinct immunocyte-promoting and adipocyte-generating stromal components coordinate adipose tissue immune and metabolic tenors. Sci. Immunol. 4, eaaw3658 (2019).
Ding, X. et al. IL-33-driven ILC2/eosinophil axis in fat is induced by sympathetic tone and suppressed by obesity. J. Endocrinol. 231, 35–48 (2016).
Goldberg, E. L. et al. IL-33 causes thermogenic failure in aging by expanding dysfunctional adipose ILC2. Cell Metab. 33, 2277–2287 (2021).
Brestoff, J. R. et al. Group 2 innate lymphoid cells promote beiging of white adipose tissue and limit obesity. Nature 519, 242–246 (2015).
Lee, M. W. et al. Activated type 2 innate lymphoid cells regulate beige fat biogenesis. Cell 160, 74–87 (2015).
Odegaard, J. I. et al. Perinatal licensing of thermogenesis by IL-33 and ST2. Cell 166, 841–854 (2016).
Islam MR,Valaris, S. et al. Exercise hormone irisin is a critical regulator of cognitive function. Nat Metab. 3, 1058–1070 (2021).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Sun, Y. et al. Mitochondrial TNAP controls thermogenesis by hydrolysis of phosphocreatine. Nature 593, 580–585 (2021).
Li, C. et al. Interferon-α-producing plasmacytoid dendritic cells drive the loss of adipose tissue regulatory T cells during obesity. Cell Metab. 33, 1610–1623 (2021).
Holgado, A. et al. IL-33trap is a novel IL-33-neutralizing biologic that inhibits allergic airway inflammation. J. Allergy Clin. Immunol. 144, 204–215 (2019).
Li, C. et al. TCR Transgenic mice reveal stepwise, multi-site acquisition of the distinctive fat-Treg phenotype. Cell 174, 285–299 (2018).
Mahlakõiv, T. et al. Stromal cells maintain immune cell homeostasis in adipose tissue via production of interleukin-33. Sci. Immunol. 4, eaax0416 (2019).
Schmitz, J. et al. IL-33, an interleukin-1-like cytokine that signals via the IL-1 receptor-related protein ST2 and induces T helper type 2-associated cytokines. Immunity 23, 479–490 (2005).
Holgado, A. et al. Single-chain soluble receptor fusion proteins as versatile cytokine inhibitors. Front. Immunol. 11, 1422 (2020).
Hu, B. et al. γδ T cells and adipocyte IL-17RC control fat innervation and thermogenesis. Nature 578, 610–614 (2020).
Molofsky, A. B. et al. Interleukin-33 and interferon-γ counter-regulate group 2 innate lymphoid cell activation during immune perturbation. Immunity 43, 161–174 (2015).
Xiao, T. et al. Tregs in visceral adipose tissue up-regulate circadian-clock expression to promote fitness and enforce a diurnal rhythm of lipolysis. Sci. Immunol. 7, eabl7641 (2022).
Zhang, Y. et al. Irisin exerts dual effects on browning and adipogenesis of human white adipocytes. Am. J. Physiol. Endocrinol. Metab. 311, E530–E541 (2016).
Li, H. et al. Effects of irisin on the differentiation and browning of human visceral white adipocytes. Am. J. Transl. Res. 11, 7410–7421 (2019).
Li, H. et al. The effect of irisin as a metabolic regulator and its therapeutic potential for obesity. Int. J. Endocrinol. 2021, 6572342 (2021).
Miller, A. M. et al. Interleukin-33 induces protective effects in adipose tissue inflammation during obesity in mice. Circ. Res. 107, 650–658 (2010).
Wang, G. et al. Adipose-tissue Treg cells restrain differentiation of stromal adipocyte precursors to promote insulin sensitivity and metabolic homeostasis. Immunity 57, 1345–1359 (2024).
Torres, S. V. et al. Two regulatory T cell populations in the visceral adipose tissue shape systemic metabolism. Nat. Immunol. 25, 496–511 (2024).
Revelo, X. S., Luck, H., Winer, S. & Winer, D. A. Morphological and inflammatory changes in visceral adipose tissue during obesity. Endocr. Pathol. 25, 93–101 (2014).
Lin, Y. et al. The chemerin-CMKLR1 axis limits thermogenesis by controlling a beige adipocyte/IL-33/type 2 innate immunity circuit. Sci. Immunol. 6, eabg9698 (2021).
Cao, S. et al. The IL-33/ST2 axis affects adipogenesis through regulating the TRAF6/RelA pathway. Int. J. Mol. Sci. 25, 12005 (2024).
Estell, E. G. et al. Irisin directly stimulates osteoclastogenesis and bone resorption in vitro and in vivo. Elife 9, e58172 (2020).
Pettersson, U. S., Waldén, T. B., Carlsson, P. O., Jansson, L. & Phillipson, M. Female mice are protected against high-fat diet induced metabolic syndrome and increase the regulatory T cell population in adipose tissue. PLoS ONE 7, e46057 (2012).
Vasanthakumar, A. et al. Sex-specific adipose tissue imprinting of regulatory T cells. Nature 579, 581–585 (2020).
Stępień, M. et al. Obesity indices and inflammatory markers in obese non-diabetic normo- and hypertensive patients: a comparative pilot study. Lipids Health Dis. 13, 29 (2014).
Wrann, C. D. et al. Exercise induces hippocampal BDNF through a PGC-1α/FNDC5 pathway. Cell Metab. 18, 649–659 (2013).
Kam, T. I. et al. Amelioration of pathologic α-synuclein-induced Parkinson’s disease by irisin. Proc. Natl Acad. Sci. USA 119, e2204835119 (2022).
Kim, E. et al. Irisin reduces amyloid-β by inducing the release of neprilysin from astrocytes following downregulation of ERK-STAT3 signaling. Neuron 111, 3619–3633 (2023).
Rao, X. et al. Dual roles of interleukin-33 in cognitive function by regulating central nervous system inflammation. J. Transl. Med. 20, 369 (2022).
Fu, A. K. et al. IL-33 ameliorates Alzheimer’s disease-like pathology and cognitive decline. Proc. Natl Acad. Sci. USA 113, E2705–E2713 (2016).
Reverchon, F. et al. Hippocampal interleukin-33 mediates neuroinflammation-induced cognitive impairments. J. Neuroinflammation 17, 268 (2020).
Oguri, Y. et al. CD81 controls beige fat progenitor cell growth and energy balance via FAK Signaling. Cell 182, 563–577 (2020).
Wang, X., Spandidos, A., Wang, H. & Seed, B. PrimerBank: a PCR primer database for quantitative gene expression analysis, 2012 update. Nucleic Acids Res. 40, D1144–D1149 (2012).
Spandidos, A., Wang, X., Wang, H. & Seed, B. PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res. 38, D792–D799 (2010).
Spandidos, A. et al. A comprehensive collection of experimentally validated primers for Polymerase Chain Reaction quantitation of murine transcript abundance. BMC Genomics 9, 633 (2008).
Wang, X. & Seed, B. A PCR primer bank for quantitative gene expression analysis. Nucleic Acids Res. 31, e154 (2003).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Mina, A. I. et al. CalR: A web-based analysis tool for indirect calorimetry experiments. Cell Metab. 28, 656–666 (2018).
A, Mu. Irisin ameliorates obesity and insulin-resistance via adipose-tissue IL-33 and regulatory T cells. Figshare https://doi.org/10.6084/m9.figshare.31151524.v1 (2026).
Acknowledgements
We gratefully acknowledge funding from the Freedom Together Foundation (FTF) to B.M.S. and D.M. and from the National Institutes of Health (NIH) to D.M. (DK092541). M.A. is supported by NIH F32 (1F32DK132864-01) and K01 (1K01DK141969) awards, L.R. by the Deutsche Forschungsgemeinschaft (DFG, German research foundation) – Projektnummer 507975958, and P.K.L. by NIH F32 (AG072874). We thank all members of the labs of D.M. and B.M.S. for helpful discussions. Work in the lab of R.B. is funded by Ghent University (bof/baf/4y/2024/01/511) and the Fund for Scientific Research Flanders (BRIDGEExcellence of Science Programme grant, EOS ID: 40007491). I.S.A. holds a fundamental mandate of the Foundation against Cancer (365C06721). We acknowledge H. Braun for help in IL-33trap development and the VIB Protein Core for help in optimizing the IL-33trap production protocol. We acknowledge the Immunology Flow Cytometry Facility at Harvard Medical School, Rodent Histopathology Core at Harvard Medical School, Energy Balance Core at BIDMC and the Molecular Imaging Core at Dana-Farber Cancer Institute. We also thank the E. Rosen lab for sharing their animal space with us for mouse breeding.
Author information
Authors and Affiliations
Contributions
M.A. and G.W. performed and analysed most of the cellular and animal experiments under the supervision of D.M. and B.M.S. N.Z. performed certain flow cytometry experiments. L.R. helped with the design of the animal experiments and assisted with the MRI and CLAMS experiments. D.B., Y.M. and Q.Z. assisted with some of the animal experiments, breeding and maintenance. C.C. performed the primary iWAT ex vivo experiments. P.K.L. assisted with VmSC sorting. I.A. and R.B. developed and initially characterized the IL-33trap and provided the IL-33trap expression plasmid and information related to IL-33trap expression and purification. R.B. instructed on the IL-33trap administration in mice. M.A., G.W., B.M.S. and D.M. co-wrote the paper, which all the other authors reviewed.
Corresponding authors
Ethics declarations
Competing interests
B.M.S. holds several patents on irisin: compositions and Methods for Brown Fat Inductions and Activity using Fndc5 (patent nos. 2848368, 11098092, 8969519 and 10093705), Methods for the Identification, Assessment, Prevention and Treatment of Neurological Disorders and Diseases Using FNDC5 (patent nos. 2014329606, 2020202676, 10,286,042 and 11,129,879). B.M.S. is an academic co-founder of Aevum Therapeutics, which is attempting to develop irisin as a therapeutic. M.A. is a consultant to Aevum Therapeutics. R.B. holds a patent on IL-33trap (patent no. US10703799B2). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Pei Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chronic irisin treatment increases energy expenditure, and improves glucose homeostasis of high-fat-diet-fed mice.
(A) ELISA measurements of irisin plasma concentrations in mice fed a HFD for 8 weeks (13 weeks after AAV injection; n = 12, 12, 5, 5, 5, 5) and 18 weeks (23 weeks after AAV injection; n = 12, 12, 5, 5, 5, 5). One-way ANOVA was used for calculating p values (**** p < 0.0001, **** p < 0.0001, **** p < 0.0001, and **** p < 0.0001). (B, C) Measurements of tissues from male mice fed a HFD or normal chow for 8 weeks (B) (n = 8, 8, 4, 4) or 18 weeks (C) (n = 8, 8, 3, 3) after AAV-Irisin or AAV-GFP administration. n = 8, 8, 4, 4. Two-way ANOVA was used for calculating p values (B: **** p < 0.0001, * p = 0.0299, and * p = 0.0110; C: **** p < 0.0001, and ** p = 0.0012). (D) Energy expenditure measured in CLAMS cages over one week. n = 5, 5. The last 4.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Two-way ANOVA was used for calculating p values (left: * p = 0.0161, * p = 0.0285, * p = 0.0277, and * p = 0.0308; right: ** p = 0.0064). (E) Total distance that animal traveled was measured in CLAMS cages over the last 4.5 days of the monitoring week. n = 5, 5, 5, 5. The last 4.5 days of tracking were shown on the left and the average values in the light and dark periods were summarized on the right. n = 5, 5. Two-way ANOVA was used for calculating p values. (F) Locomotive activity measured in CLAMS cages over one week. n = 5, 5. The last 4.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Two-way ANOVA was used for calculating p values. (G) Intraperitoneal glucose tolerance tests (GTTs) of mice fed a HFD for 8 weeks (n = 15, 15). Two-way ANOVA was used for calculating p values. (H) Intraperitoneal insulin tolerance tests (ITTs) of mice fed a HFD for 8 weeks (n = 15, 15). Two-way ANOVA was used for calculating p values. (I) Body weight (left), intraperitoneal glucose tolerance tests (GTTs, middle) and insulin tolerance tests (ITTs, right) of mice with similar weights from a re-run of Fig. 1a cohort fed a HFD for 18 weeks (n = 4, 4). Unpaired two-sided T test (body weight) or two-way ANOVA (GTTs and ITTs) was used for calculating p values (GTT: * p = 0.0486, ** p = 0.0094, and * p = 0.0123; ITT: * p = 0.0120, ** p = 0.0085, and *** p = 0.0008). (J) Fraction of ST2+ Treg cells in total Treg cells in eWAT from mice in (I). Unpaired two-sided T-test was used for calculating p values (*** p = 0.0001). (K) KEGG pathway enrichment analysis from the transcripts prepared from iWATs. The significant pathways (FDR < 0.05) for differentially expressed genes (FC > 2; FDR < 0.05) between mice received AAV-irisin and mice received the control AAV-GFP were indicated (n = 3, 3). (L) RT-qPCR measuring mRNAs associated with thermogenesis and adipogenesis in BAT (n = 7, 8), iWAT (n = 6, 6), and eWAT (n = 5, 8) of the mice received AAV-irisin or -GFP and fed a HFD for 18 weeks. Two-way ANOVA was used for calculating p values (BAT: * p = 0.0134, *** p = 0.0004, * p = 0.0170, ** p = 0.0099, * p = 0.0159; iWAT: ** p = 0.0016, *** p = 0.0007, ** p = 0.0024, **** p < 0.0001, *** p = 0.0002, ** p < 0.0017, *** p = 0.0003; eWAT: * p = 0.0336). Abbreviations as per Fig. 1. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 2 Chronic irisin treatment has no effects on energy expenditure and glucose homeostasis in normal-chow-fed mice.
4-week-old male C57BL6/J mice were intravenously injected with AAV-GFP or AAV-Irisin. The mice were fed a normal chow continuously without switching diet. Mice were analyzed in CLAMS cages after another 12 weeks of continuous normal chow feeding, and gene expression analysis was performed after another 18 weeks of continuous normal chow feeding. (A) Body weight measurements in CLAMS cages over one week (n = 8, 8). The last 3.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Unpaired two-sided T-test was used for calculating p values. (B) Food consumption measurements in CLAMS cages over one week (n = 8, 7). The last 3.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Unpaired two-sided T-test was used for calculating p values. (C) Energy expenditure measured in CLAMS cages over one week (n = 8, 8). The last 3.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Two-way ANOVA (left) or unpaired two-sided T-test (right) was used for calculating p values. (D) Locomotive activity measured in CLAMS cages over one week (n = 8, 8). The last 3.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Two-way ANOVA (left) or unpaired two-sided T-test (right) was used for calculating p values. (E) Intraperitoneal glucose tolerance tests (GTTs) of mice fed a normal chow for 18 weeks (n = 15, 15). Two-way ANOVA was used for calculating p values. (F) Intraperitoneal insulin tolerance tests (ITTs) of mice fed a normal chow for 18 weeks (n = 15, 15). Two-way ANOVA was used for calculating p values. (G) Distance traveled measured in CLAMS cages over one week. n = 8, 8. The last 3.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Unpaired two-sided T-test was used for calculating p values. (H–J) RT-qPCR measuring mRNAs associated with thermogenesis and adipogenesis in BAT (H) (n = 15, 15), iWAT (I) (n = 15, 15), and eWAT (J) (n = 9, 12) of the mice received AAV-irisin or -GFP and fed a normal chow for 5 weeks followed by another 18 weeks of feeding without diet switch. Two-way ANOVA was used for calculating p values (BAT: * p = 0.0211, **** p < 0.0001, * p = 0.0276, * p = 0.0294, **** p < 0.0001, *** p = 0.0006, and ** p = 0.0046; iWAT: * p = 0.0420; eWAT: *** p = 0.0002). (K) RT-qPCR measuring mRNAs associated with lipolysis in iWAT of the indicated mice received AAV-irisin or -GFP, and fed a HFD for 18 weeks (n = 7, 8). Two-way ANOVA was used for calculating p values. Abbreviations as per Fig. 1. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 3 Irisin selectively protects ST2+ Treg cells in the T cell compartment of eWAT in obese mice.
(A, B) Representative gating strategy for analyzing lymphocytes (A), as well as CD11chigh macrophages (B). (C–E) t-SNE analysis of the T cell compartment from the flow cytometry data in Fig. 2a. (C) t-SNE plot depicting T cell clusters across all samples. (D) Summary data showing the fractions of each T cell cluster. n = 3, 3, 8, 8. Two-way ANOVA was used for calculating p values (* p = 0.0479, * p = 0.0314, and * p = 0.0058). (E) Individual (top, middle) and differential density plots (bottom) comparing AAV-Irisin with AAV-GFP under HFD conditions. (F–I) 4-week-old male C57BL6/J mice were intravenously injected with AAV-GFP or -Irisin. 5 weeks later, the mice were fed either a normal chow (NC) or HFD for 8 (n = 4, 4, 8, 8). Flow-cytometric analysis of the number of total CD45+ cells (F), percentages of total and CD11chigh macrophages (G), percentage of T, CD8+ and CD4+ T cells (H), and percentage of Foxp3− CD4+ T cells and ST2+ Treg cells (I) in eWATs from mice fed a NC or HFD for 8 weeks. One-way ANOVA was used for calculating p values (F: * p = 0.0107, and ** p = 0.0051; G: *** p = 0.0009, * p = 0.033, *** p = 0.0007 and *** p = 0.0002; H: * p = 0.0255; I: * p = 0.0116, and * p = 0.0313). Tconv, conventional CD4+ T cells; other abbreviations as per Fig. 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 4 Irisin has no effects on inflammation and ST2+ Treg cells in iWAT and BAT of obese mice.
(A–D) Flow-cytometric analysis of the number of total CD45+ cells, and percentages of total and CD11chigh macrophages, as well as total T, CD8+, Foxp3−CD4+ T cells, total and ST2+ Treg cells in iWATs and BATs from mice fed a HFD for either 8 weeks or 18 weeks as Extended Data Fig. 3F (n = 4, 4, 8, 8) and Fig. 2 (n = 3, 3, 8, 8). One-way ANOVA was used for calculating p values (A: * p = 0.0365, **** p < 0.0001, * p = 0.0436, and * p = 0.0445; B: * p = 0.0386; C: * p = 0.0134, * p = 0.0210, and * p = 0.0129; D: * p = 0.0429, ** p = 0.0019, and * p = 0.0166). Abbreviations as per Fig. 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 5 Irisin directly induces IL-33 expression in stromal cells from eWAT but not in those from iWAT or BAT.
(A) Representative gating strategy for analyzing IL-33+ mSCs. (B) Summary data for numbers of IL-33+ mSCs in eWATs from the male mice in Extended Data Fig. 3F (n = 4, 4, 8, 8). (C) Summary data for numbers of IL-33+ mSCs in iWATs and BATs from the mice in Extended Data Fig. 3F and Fig. 2 (left, n = 4, 4, 8, 8; right, n = 3, 3, 8, 8). (D) ELISA measurements of IL-33 concentration in iWAT and BAT lysates from a subset of mice in Fig. 3c (n = 3, 3, 3, 3). One-way ANOVA was used for calculating p values (* p = 0.0159, *** p = 0.0002, and ** p = 0.0050). (E) Summary data for numbers of ILC2s in eWAT, iWAT and BAT from the mice in Extended Data Fig. 4 (top, n = 4, 4, 8, 8; bottom, n = 3, 3, 8, 8). (F) Freshly isolated IL-33 (EGFP)− mSCs from 12 wk-old male Il33.Egfp mice were treated with PBS or 10 nM/Hsp90α irisin for 3 days, followed by flow-cytometric analysis. Left: experimental scheme; right: summary data for numbers and fractions of IL-33 (EGFP)+ mSCs. n = 3. Abbreviation: mSCs, mesenchymal stromal cells; ILC2s, type 2 innate lymphoid cells; other abbreviations as per Figs. 1 and 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustration in f created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
Extended Data Fig. 6 Irisin induces IL-33 expression in adipose tissues via integrin αVβ5 receptor.
(A) Normalized read counts of integrin αV and β5 subunits from prior RNA-seq data47. One-way ANOVA was used for calculating p values (ns p = 0.7013, * p = 0.0234, and ** p = 0.0081). (B) RT-qPCR measuring IL-33 mRNAs in flow cytometrically sorted IL-33 (EGFP)− mSCs from 12 wk-old male Il33.Egfp mice treated with PBS or 10 nM irisin/Hsp90α for 3 days in the presence of anti-αVβ5 or control IgG. One-way ANOVA was used for calculating p values (**** p < 0.0001, ns p = 0.9930, and **** p < 0.0001). (C) Fluorescence imaging and quantification of IL-33-GFP expression of flow cytometrically isolated IL-33 (EGFP)− mSCs from 12 wk-old male Il33.Egfp mice treated with PBS or 10 nM irisin/Hsp90α for 3 days in the presence of anti-αVβ5 or control IgG. Left: representative images of the treated mSCs, Scale bar = 100 μm; Right: normalized GFP signals from each wells. One-way ANOVA was used for calculating p values (**** p < 0.0001, ns p = 0.9948, and *** p = 0.0002). (D) Fluorescence imaging and quantification of integrin signaling of flow cytometrically isolated IL-33 (EGFP)− mSCs from 12 wk-old male Il33.Egfp mice treated with PBS or 10 nM irisin/Hsp90α for 3 days in the presence of anti-αVβ5 or control IgG. Left: representative images of the treated mSCs, Scale bar = 100 μm; Right: quantification of pFAK/FAK ratios from each wells. One-way ANOVA was used for calculating p values (**** p < 0.0001, ns p = 0.8338, and **** p < 0.0001). Abbreviations as per Figs. 1 and 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 7 Irisin promotes IL-33 response in iWAT.
(A) Summary data of the number of total CD4+ T cells, as well as the percentages of Treg cells and ST2+ Treg cells in the spleen of the indicated mice from the flow-cytometric analysis. (B–E) 8 wk-old male Fndc5+/+ or Fndc5−/− littermates were treated as per Fig. 4c. (B) Summary data for the number of total Treg cells and the fraction of ST2+ Treg cells in iWAT. n = 3, 4, 3, 3. One-way ANOVA was used for calculating p values (* p = 0.0269). (C, D) Representative gating strategy for analyzing ILC2s (C) and Eosinophils (D). (E, F) Summary data for numbers of ST2+Foxp3−CD4+ cells, ILC2s, and Eosinophils in eWAT (n = 9, 9, 6, 8) (E) and iWAT (n = 3, 4, 3, 3) (F). Brown-Forsythe and Welch ANOVA (E) or one-way ANOVA (F) was used for calculating p values (E: * p = 0.0238, ** p = 0.0011, * p = 0.0426, **** p < 0.0001, and ** p = 0.0011; F: * p = 0.0333, * p = 0.0293, and * p = 0.0161). (G) ELISA measurements of irisin plasma concentrations in male Fndc5+/+ and Fndc5−/− mice fed a HFD for 17 weeks after AAV injection; n = 10, 11, 13, 13. One-way ANOVA was used for calculating p values (*** p = 0.0001, and *** p = 0.0006). (H) Pearson correlation analysis of area under ITT curves and body weight of male Fndc5−/− mice fed a HFD for 17 weeks after AAV injection; n = 16. Abbreviations as per Figs. 4 and 5. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Experiments in B-F were repeated three times with similar results, and the rest of the experiments were repeated twice with similar results.
Extended Data Fig. 8 IL-33/ST2 axis is not required for regulating food intake, physical activities or thermogenic gene expression in eWAT of HFD-treated mice.
(A, B) Purification of recombinant IL-33trap (A) and measurement of endotoxin content (B). (C) ELISA measurements of irisin plasma concentrations in male mice fed a HFD for 17 weeks after AAV injection w and w/o chronic IL-33trap treatment (n = 13, 13, 13, 13). One-way ANOVA was used for calculating p values (**** p < 0.0001, * p = 0.0255, and **** p < 0.0001). (D) Body weight measurements of male mice fed a HFD for 17 weeks after AAV injection w and w/o chronic IL-33trap treatment (mice used for CLAMS); n = 6, 6, 6, 6. One-way ANOVA was used for calculating p values (** p = 0.0049, ** p = 0.0064, and ** p < 0.0033). (E–G) Accumulated food intake (E), locomotive activity (F) and total distance the animal traveled (G) measured in CLAMS cages over one week. n = 4, 4, 4, 4. The last 4 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Two-way ANOVA was used for calculating p values. (H–J) Body weight (H), eWAT weight (I) and iWAT weight (J) measurements (n = 11, 10, 4, 7). One-way ANOVA was used for generating p values (H: **** p < 0.0001, ns p = 0.9928, and * p = 0.0330; J: **** p < 0.0001, *** p = 0.0002, and ns p = 0.4604). (K) ELISA measurements of irisin plasma concentrations in mice; n = 12, 7. Unpaired two-sided T-test was used for calculating p values (ns p = 0.2468). (L) Related to Fig. 7f. RT-qPCR measuring mRNAs associated with thermogenesis and adipogenesis in eWAT of ST2 Treg null mice (Il1rl1Δtreg) and their littermates (Il1rl1wt) received AAV-irisin or -GFP and fed a HFD for 17 weeks (n = 11, 10, 4 7). Two-way ANOVA was used for calculating p values (**** p < 0.0001, **** p < 0.0001, and **** p < 0.0001). Abbreviations as per Fig. 1. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Extended Data Fig. 9 Irisin’s effects in obese mice in a therapeutic setting.
6-week-old male C57BL6/J mice were switched to HFD feeding. After 14 weeks, mice were intravenously injected with AAV-GFP (n = 15) or AAV-Irisin (n = 15). After another 10 weeks of HFD feeding, mice were subjected to metabolic analysis and finally sacrificed for tissue analysis. (A) Schematic of the HFD experiment. (B) ELISA measurements of irisin plasma concentrations in mice; n = 15, 15. Unpaired two-sided T-test was used for calculating p values (*** p = 0.0004). (C) Body weight measurements (n = 15, 15). (D) Body composition analysis by MRI (n = 15, 15). Unpaired two-sided T-test was used for calculating p values. (E) Intraperitoneal glucose tolerance tests (GTTs) (n = 15, 15). Two-way ANOVA was used for calculating p values. (F) Intraperitoneal insulin tolerance tests (ITTs) (n = 15, 15). Two-way ANOVA was used for calculating p values. (G–I) Energy expenditure measured (G), accumulated food intake (H) and distance traveled (I) in CLAMS cages over one week. n = 8, 8. The last 4.5 days of measurement were shown on the left and the averaged values in the light and dark periods were summarized on the right. Unpaired two-sided T-test was used for calculating p values. (J) RT-qPCR measuring mRNAs associated with thermogenesis and adipogenesis in iWAT (left) (n = 8, 8) and eWAT (right) (n = 7, 7) of HFD-induced obese mice received AAV-irisin or -GFP for 10 weeks. Two-way ANOVA was used for calculating p values (* p = 0.0170, ** p = 0.0061, and ** p = 0.0036). (K–O) Flow-cytometric analysis of the number of total CD45+ cells (K), percentages of total (L) and CD11chigh (M) macrophages, and total T, CD8+ and Foxp3− CD4+ T cells (N), as well as Treg cells and ST2+ Treg cells in eWATs from obese mice administered AAV for 10 weeks (n = 8, 8). Unpaired two-sided T-test was used for calculating p values (* p = 0.0173). Abbreviations as per Figs. 1 and 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results. Mouse illustration in a created in BioRender; A, M. https://biorender.com/5v6fzeu (2026).
Extended Data Fig. 10 Irisin’s action in iWAT through IL-33.
(A) RT-qPCR measuring mRNAs of ST2, IL-33 and genes associated with thermogenesis in different batches of differentiated primary iWAT cultures prepared from stromal vascular fraction (SVF) of 4-week-old male C57BL6 mice; n = 3, 3, 3, 3. Two-way ANOVA was used for calculating p values (left: **** p < 0.0001, *** p = 0.0001, and **** p < 0.0001; right: ns p = 0.9632, ** p = 0.0019, and ** p = 0.0021). (B) SVF from 4-week old male mice iWAT was differentiated in vitro with indicated Irisin/HSP90α and NE treatments (left). IL-33 mRNAs were measured using RT-qPCR (right) (n = 4, 4, 4, 4). Two-way ANOVA was used for calculating p values (Il1rl1: * p = 0.0160, ** p = 0.0016, and ** p < 0.0022; Il33: * p = 0.0436, * p = 0.0309, ** p = 0.0042, and ** p = 0.0032; Alpl: ** p = 0.0080, and ** p = 0.0059; Ucp1: ** p = 0.0010, ** p = 0.0043, * p = 0.0276, and *** p = 0.0003). (C) RT-qPCR measuring ALPL mRNA in differentiated primary iWAT cultures in the presence or absence of NE treatment; n = 4, 4, 4, 4. Two-way ANOVA was used for calculating p values (** p = 0.0028, **** p < 0.0001, and **** p < 0.0001). Abbreviations: NE: norepinephrine; as well as per Figs. 1 and 2. Mean ± SD.* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns: not significant. Significant but irrelevant p values are not indicated. Unless specifically mentioned, experiments were repeated twice with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3.
Source data
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed gel.
Source Data Figs. 1–7 and Extended Data Figs. 1–10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
A, M., Wang, G., Zammit, N.W. et al. Irisin ameliorates obesity and insulin resistance via adipose tissue IL-33 and regulatory T cells. Nat Metab 8, 885–901 (2026). https://doi.org/10.1038/s42255-026-01491-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-026-01491-2