Abstract
In aging, skeletal muscle regeneration declines due to alterations in both myogenic and non-myogenic cells and their interactions. This regenerative dysfunction is not understood comprehensively or with high spatiotemporal resolution. We collected an integrated atlas of 273,923 single-cell transcriptomes and high-resolution spatial transcriptomic maps from muscles of young, old and geriatric mice (~5, 20 and 26 months old) at multiple time points following myotoxin injury. We identified eight immune cell types that displayed accelerated or delayed dynamics by age. We observed muscle stem cell states and trajectories specific to old and geriatric muscles and evaluated their association with senescence by scoring experimentally derived and curated gene signatures in both single-cell and spatial transcriptomic data. This revealed an elevation of senescent-like muscle stem cell subsets within injury zones uniquely in aged muscles. This Resource provides a holistic portrait of the altered cellular states underlying muscle regenerative decline across mouse lifespan.
Similar content being viewed by others
Main
Skeletal muscle is heterogeneously composed of interacting immune, stromal and myogenic cells that contribute to the maintenance and regeneration of muscle by regulating muscle stem cell (MuSC) quiescence, proliferation and differentiation1. MuSCs are found between the basal lamina and the plasma membrane of myofibers and are essential for the initial development of muscle and in muscle regeneration2,3. The MuSC population is maintained across multiple cycles of growth and regeneration by asymmetrical division, which generates additional MuSCs and Myod1+ myoblasts2,3. Myoblasts further expand, differentiate and fuse to form new Myog+ myocytes2,3. The paired box protein 7 (Pax7) transcription factor, predominantly expressed in MuSCs, regulates the expression of myogenic regulatory factors such as Myf5 and Myod1 (ref. 3).
There are extrinsic changes, such as an increase in fibro-adipogenic progenitors (FAPs), and intrinsic factors, such as a reduction in asymmetrical self-renewal and an increase in senescent MuSCs, that disrupt skeletal muscle homeostasis and regeneration in aging2. Aged MuSCs exhibit a decline in self-renewal and ability to differentiate, thus reducing the MuSC pool2,4,5,6. Compared to young MuSCs, fewer aged MuSCs are found in quiescence, due to either elevated activation or entering a pre-senescent state5,6. It remains unclear how these extrinsic and intrinsic changes are integrated systematically and how heterogeneities related to cellular senescence within both myogenic and non-myogenic cell populations contribute, in part due to a paucity of holistic analyses of these alterations. Further, it has been posited that temporal and spatial discoordination between the dynamics in key cell types during the repair process leads to inefficient outcomes2. We refer to Supplementary Note 1 for more details on the cell-type dynamics and senescence in skeletal muscle.
Single-cell methods have been used previously to understand skeletal muscle homeostasis and regeneration at various ages7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23. Recent reports have provided insights into MuSC dysfunction with aging. A recent human skeletal muscle study observed a decline in the proportion of MuSCs with age and that IGFN1, which is needed for myoblast fusion and differentiation, is decreased in old (~75 years old) muscle19. This report found no difference in Cdkn2a expression by age, but did identify a senescent myonuclei population that expressed Cdkn1a that was more frequent in old than young human samples19. A recent study on mouse skeletal muscle aging based on mass cytometry observed that CD47hi MuSCs occur with a higher frequency in aged mice and exhibit poor regenerative capacity24. Another report demonstrated that quiescent MuSCs in aged mice have reduced expression of cyclin D1, which is needed for proper MuSC activation and muscle regeneration25.
To evaluate the factors that contribute to the age-related decline in skeletal muscle regeneration in a more comprehensive manner, we have generated single-cell RNA-sequencing (scRNA-seq) datasets of uninjured (day 0) and myotoxin-injured (days 1, 2, 3.5, 5 and 7) tibialis anterior (TA) muscles from young, old and geriatric mice. We identified a total of 29 cell types, 8 of which had a significant difference in their abundances throughout regeneration by age. We confirmed changes in age-specific T cell abundance by immunohistochemistry and flow cytometry. Given the role that cellular senescence plays in limiting cell contributions in aged tissues, we tested a series of tools to bioinformatically identify senescence in these single-cell data and found a transfer-learning-based scoring approach accurately classified senescent-like myogenic cells across ages and cell cycling states. This scoring approach revealed that senescent-like subsets exist at key transitional self-renewal states within the myogenic stem/progenitor cell pseudotime trajectory across all ages but are elevated in aged muscles. We confirmed this age-specific difference in senescent-like MuSCs with spatial transcriptomics. This resource of mouse skeletal muscle aging provides a more comprehensive portrait of the changing cellular states underlying skeletal muscle regeneration across mouse lifespan.
Results
scRNA-seq of skeletal muscle regeneration
To comprehensively evaluate skeletal muscle homeostasis and regeneration throughout aging, we performed scRNA-seq on 65 mouse skeletal muscle samples with the 10x Chromium v2 and v3 platforms. Muscle damage was induced in young (4–7 months old (mo)), old (20 mo) and geriatric (26 mo) C57BL/6J mice by injecting the TA muscles with notexin. The injured and uninjured muscles were collected at six time points (days 0, 1, 2, 3.5, 5 and 7; Fig. 1a,b). Together, these 65 scRNA-seq samples, newly reported here and from our two prior reports11,26, contained a total of 365,710 cell barcodes before quality control and filtering (Extended Data Fig. 1a and Supplementary Table 1). All samples were processed by aligning sequencing reads to the mm10 mouse reference genome, removing ambient RNA signatures with SoupX27, removing low-quality cells and identifying and removing doublets with DoubletFinder28 (Fig. 1c and Extended Data Fig. 1). The samples were integrated with Harmony29 to correct for batch effects (Fig. 1c). The final dataset contained 273,923 cells. We observed that the cell number was relatively evenly distributed across the three age groups (Fig. 1d,g), and within each age group, the number of cells from each time point was relatively consistent (Fig. 1e).
a,b, Overview of experimental design (a) and summary of replicates (b). 3′ scRNA-seq (10× Chromium v2 and v3) was performed on dissociated TA muscles from young (4–7 mo), old (20 mo) and geriatric (26 mo) mice (both sexes) 0–7 days post injury (dpi) with notexin. n = 3–4 replicates per age and dpi. c, Processing workflow. Each scRNA-seq sample was aligned to the mm10 mouse reference genome, ambient RNA was removed by SoupX, low-quality cells were identified and removed, and doublets were identified and removed. All samples were then integrated with Harmony, resulting in a final dataset containing 273,923 cells from 65 samples. See Extended Data Fig. 1 and Supplementary Table 1 for additional detail. QC, quality control; UMI, unique molecular identifier. d, Fraction of cells from each age group. e, Fraction of cells from each dpi within each age group. f,g, Uniform manifold approximation and projection (UMAP) representations of the final dataset. Cells colored by manually assigned cell-type IDs based on the expression of hallmark skeletal muscle genes (see Extended Data Figs. 2 and 3) (f). Cells are colored by age group, with all other cells in gray (g). DC, dendritic cell.
Multistep identification of cell types by clustering
After data integration, shared nearest-neighbor (SNN) clustering was performed, and canonical marker genes were used to manually identify cell types. The initial clustering resulted in 24 clusters that each received a unique cell-type annotation. Of these, nine were myeloid cell clusters (monocytes, macrophages, dendritic cells and neutrophils) that exhibited similar expression profiles. To further clarify the myeloid cells found in the final dataset, the nine myeloid cell clusters were subset out, re-clustered and re-embedded (Extended Data Fig. 2a–c). This resulted in 15 clusters that each received a unique cell-type annotation based on known myeloid markers (Extended Data Fig. 2c,d). Although this re-clustering did not further clarify the monocyte and macrophage annotations, it did help to identify more specific dendritic cell and T cell subtypes (Extended Data Fig. 2c,d). The dendritic cell and T cell-subtype annotations were transferred from the myeloid subset back to the final dataset based on the cell barcode. The monocyte and macrophage annotations were not changed based on the myeloid subset. With the additional myeloid subset annotations, we identified 29 distinct cell types in the final dataset (Fig. 1f and Extended Data Fig. 3). Compared to prior scRNA-seq and single-nucleus RNA-sequencing skeletal muscle studies, we identified similar broad cell types and we identified more specific endothelial, FAP and immune cell subsets7,9,10,11,12,13,14,15,16,17,18,19,20,21,30.
Lymphoid cell types were 5 of the 17 immune cell clusters. We identified a rare B cell cluster (0.47% of the final dataset) that expressed Cd19, Cd22 and Ms4a1 (Extended Data Fig. 3a). We identified a natural killer (NK) cell cluster that expressed Nkg7, Gzma, Klra4 and Klre1 but, importantly, did not express T cell markers (Extended Data Fig. 3a). We identified three T cell clusters, all of which expressed Cd8a and Cd8b1. One of the T cell populations more strongly expressed Cd4, while the other two T cell populations expressed Cd3e. One of the Cd3e+ clusters also strongly expressed the cycling markers Cdk1 and Hmgb2 and were thus identified as cycling Cd3e+ T cells (Extended Data Fig. 3a). The other Cd3e+ T cell population appears to be non-cycling. The B cells, NK cells and the non-cycling Cd3e+ T cells were identified as unique clusters in the initial clustering of the final dataset. The Cd4+ and cycling Cd3e+ T cells were identified when we subset and re-clustered the myeloid populations (Extended Data Figs. 2c,d and 3a).
Although we incubated the single-cell suspension in erythrocyte lysis buffer, we did see a small (0.42% of the final dataset) erythrocyte cluster that uniquely and strongly expressed a variety of hemoglobin genes, including Hba-a1 and Hbb-bs (Extended Data Fig. 3a). Erythrocytes are not a native cell type in skeletal muscle, so we have excluded them from the cell-type dynamics analysis.
We identified three FAP populations (adipogenic, pro-remodeling and stem), a tenocytes population and a Schwann and neural/glial cell population. All three of the FAP populations expressed Pdgfra and Col3a1 (Extended Data Fig. 3b). The adipogenic FAPs also expressed Adam12, Bmp5, Myoc, Col1a1, Dcn, Mmp2 and Apod. The pro-remodeling FAPs uniquely expressed cycling genes like Cdk1 and Tyms in addition to other FAP markers like Tnfaip6, Il33, Adam12, Bgn and Hdlbp. The stem FAPs also expressed Igfbp5, Dpp4, Cd34, Gsn and Mmp2. The tenocyte population expressed some FAP markers like Col1a1, Dcn and Apod, and they expressed tenocyte-specific markers Tnmd and Scx. The Schwann and neural/glial cell cluster expressed Ptn and Mpz (Extended Data Fig. 3b).
We identified a pericyte and smooth muscle cell cluster that expressed the pericyte-specific genes Rgs5 and Acta2, Myl9 and Myh11 (Extended Data Fig. 3c)31. Four endothelial clusters were identified, and they shared strong expression of Cdh5 and Pecam1 (Extended Data Fig. 3c). The arterial endothelial cells uniquely expressed Alpl and Hey1, the capillary endothelial cells strongly expressed Lpl, and the venous endothelial cells expressed Vwf, Hif1a, Icam1, Lrg1 and Aplnr (Extended Data Fig. 3c). The fourth endothelial cluster expressed endothelial markers like Cdh5 and Pecam1 and myeloid markers like S100a8/S100a9, Csf1 and Itgam (Extended Data Fig. 3c). This cluster was small (0.073% of the final dataset) and was made up of cells from multiple replicates from the three ages at 0–2 dpi (Extended Data Fig. 5e). Because this cluster was not unique to a single replicate, or single age, or single time point, we have maintained it in our analysis.
Comparison of cell-type dynamics across mouse lifespan
For each time point independent of age, we calculated the percentage of cells from each cell type. This calculation represents the cell-type abundance within the scRNA-seq dataset, not within skeletal muscle. Multinucleated myofibers are not well captured in the single-cell isolation protocol, so the abundance of myonuclei is underrepresented within the scRNA-seq dataset. Additionally, scRNA-seq libraries are limited by cell capture number, so the cell-type abundances reported are conditional on the prevalence of other cell types. In agreement with previous scRNA-seq findings, uninjured skeletal muscle (day 0) was mainly composed of endothelial cells (49.2%), FAPs (20.4%) and myonuclei (15.1%), and there were small populations of immune cells (2.7%) and MuSCs and progenitors (2.0%; Extended Data Fig. 4a and Supplementary Table 1a)7,11. Of the endothelial cells at day 0, the most prominent subtype was the capillary endothelial cell (42.4%; Fig. 2k, Extended Data Fig. 4b and Supplementary Table 1b). The FAPs present at day 0 were mainly adipogenic FAPs (15.6%; Fig. 2k, Extended Data Fig. 4b and Supplementary Table 1b). Of the immune cells present at day 0, there were a variety of monocytes and macrophages (47.2%), B cells (14.6%), non-cycling Cd3e+ T cells (11.6%), Cd209a+ dendritic cells (10.6%) and neutrophils (9.3%; Extended Data Fig. 4d and Supplementary Table 1d)7.
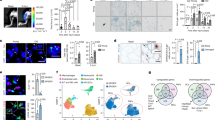
a–i, Line plots showing the relative abundance of Ccr2+ monocytes/macrophages (a), Mrc1+ monocytes/macrophages (b), Cx3cr1+ monocytes/macrophages (c), cycling monocytes/macrophages (d), patrolling monocytes/macrophages (e), NK cells (f), non-cycling T cells (g), B cells (h), and MuSCs and progenitors (i) as a fraction of total cells from 0 to 7 dpi. For each sample, the number of cells of the reported type was divided by the total number of cells (excluding erythrocytes). Points are each sample (n = 3–4). Ribbon is the standard deviation. Statistical significance of age-specific cell-type dynamics was evaluated using a one-way analysis of variance (ANOVA) and false discovery rate (FDR)-corrected P values are reported (Extended Data Fig. 6). j, Statistical significance of age-specific cell-type dynamics differences as reported by FDR-corrected P values from a one-way ANOVA. Red line denotes the FDR-corrected P = 0.05 threshold. k, Stacked bar plot of the fraction of cell types across all dpi values. All immune cells were grouped into ‘Immune cells (17 cell types)’. l, Scatterplots of the fraction of all T cells (three cell types) at 0 and 5 dpi from the scRNA-seq data. For each sample, the number of all T cells (three cell types) was divided by the total number of cells (excluding erythrocytes). Points are each sample (n = 3–4). Line is the mean for each age group. Significance was evaluated using a two-sided Welch’s t-test. m, Immunohistochemical analysis of CD3+ T cells at 0 and 5 dpi in TA muscles from young, old and geriatric mice. These are representative images from n = 2 samples per age and time point where four images were taken for each sample. Scale bar, 50 µm. Arrow denotes CD3+ T cell. n, Flow cytometric analysis gating strategy of TCRβ+ T cells. o, Scatterplots of the fraction of TCRβ+ cells of all CD45+ cells at 0 and 5 dpi. Points are each sample (n = 2–7). Line is the mean for each age group. Significance was evaluated using a two-sided Welch’s t-test. p, Violin plot of the T cell exhaustion score in all T cells by age at 5 dpi. Significance was evaluated using a two-sided Mann–Whitney U test. q, Dot plot showing the expression frequency and average expression level of select T cell, cycling, inhibitory and T cell exhaustion markers by age at 5 dpi. Prog TEX, progenitor exhausted T cell; Trans TEX, transitory exhausted T cell; Term TEX, terminal exhausted T cell.
Following injury (days 1, 2, 3.5 and 5), the most abundant general cell type was the immune cells (60.9%, 79.7%, 75.2% and 63.2%, respectively; Fig. 2k, Extended Data Fig. 4b and Supplementary Table 1b). As in previous studies7,11, non-immune cells like endothelial cells, FAPs and myonuclei were present following injury, but at transiently lower relative abundances. As expected, the most abundant immune cells immediately after injury (day 1) were neutrophils (32.0%), Ccr2+ monocytes/macrophages (19.7%), and Ctsa+ patrolling monocytes/macrophages (13.7%; Extended Data Fig. 4c and Supplementary Table 1c)7,11. Immediately following injury (days 1 and 2), there was a more pro-inflammatory environment, as evident by the abundance of Ccr2+ monocytes/macrophages. This was followed by a shift at days 3.5 and 5 to a more anti-inflammatory cell population, as evident by the peak in abundance of Cx3cr1+ monocytes/macrophages at day 5 (28.7%; Extended Data Fig. 4c and Supplementary Table 1c)7,11. By day 7, the cell-type abundances were returning to the abundances observed at day 0, but there was still a substantial immune cell population (32.1%; Extended Data Fig. 4a and Supplementary Table 1a). The immune population at day 7 mainly consists of Cx3cr1+ monocytes/macrophages (24.1%), Ctsa+ patrolling monocytes/macrophages (13.2%) and non-cycling Cd3e+ T cells (17.8%; Extended Data Fig. 4d and Supplementary Table 1d).
We next conducted an analysis of cell-type dynamics across age groups by comparing the abundance of a given cell type over the entire injury time course. Of the 28 cell types, 8 were identified as having significantly different cell-type dynamics across the three ages using a nonlinear modeling approach with multiple-hypotheses correction (Fig. 2j and Supplementary Table 2). In response to injury, we first observed an increase in neutrophils that peaked in abundance at day 1 and returned to day 0 levels by day 3.5 (Extended Data Fig. 5n). We also observed a peak in the abundance of the Ccr2+ monocytes/macrophages at days 1 and 2 (Fig. 2a), while the cycling monocytes/macrophages peaked in abundance at day 3.5 (Fig. 2d) and the Cx3cr1+ monocytes/macrophages peaked in abundance at days 3.5 and 5 (Fig. 2c). The Mrc1+ monocytes/macrophages responded early to injury (day 2) and remained high through day 5 (Fig. 2b). The geriatric Mrc1+ monocytes/macrophages maintained a higher abundance from day 2 to day 7 compared to the young and old cells (Fig. 2b). The young and old Ctsa+ patrolling monocytes/macrophages peaked in abundance at day 2, while the geriatric Ctsa+ patrolling monocytes/macrophages peaked in abundance at day 3.5 (Fig. 2e). Additionally, some lymphoid cell types like NK cells, Cd3e+ non-cycling T cells and B cells started to increase in abundance at day 2, day 3.5 and day 5, respectively (Fig. 2f–h). We observed a similar pattern when looking at all three T cell populations combined (Extended Data Fig. 5t). The geriatric NK cells did not increase in abundance within the 7-day time course, unlike the young and old cells (Fig. 2f).
When looking at all three T cell populations combined, we detected very few T cells and no age-specific differences in abundance at day 0. However, at day 5, we observed a higher abundance of T cells in old samples compared to both the young and the geriatric samples (Fig. 2l). To confirm this, we performed immunohistochemistry on sectioned TA muscles and observed that CD3+ T cells are detected more abundantly at day 5 compared to day 0 in the TAs of young, old and geriatric mice (Fig. 2m). We further used flow cytometry to quantify CD45+CD11c−CD11b−TCRβ+ T cells at days 0 and 5 from dissociated TA muscles of young, old and geriatric mice (Fig. 2n). We detected a low abundance of TCRβ+ T cells of all CD45+ hematopoietic cells at day 0, but still detected a significantly higher T cell abundance in old mice compared to young (Fig. 2o; Welch’s t-test, *P value = 0.001). Further, we observed an increase in T cell abundance from day 0 to day 5 and a significantly higher T cell abundance in old mice compared to young mice at 5 dpi (Fig. 2o; Welch’s t-test, *P value = 0.02). Together, these results suggest the abundance of the T cell pool is elevated specifically in older muscles (20 months of age), but not preserved in geriatric ages. To investigate the differences between T cell populations by age at 5 dpi, we calculated a T cell exhaustion score and looked at the expression of exhausted T cell markers (Fig. 2p,q). We found that the old and geriatric T cells have a higher T cell exhaustion score compared to the young (Fig. 2p; Mann–Whitney U test, *P = 2.2 × 10−16 and *P = 1.1 × 10−12, respectively). Genes that have been shown to mark progenitors of exhausted T cells were more highly expressed in old T cells (Bach2, Irf4, Batf, Slamf6, Tcf7). Additionally, genes that have been shown to mark transitory (Tbx21, Runx1/Runx2/Runx3) and terminally (Eomes, Tox, Nr4a1/Nr4a2/Nr4a3, Cd38, Entpd1) exhausted T cells are more highly expressed in both the old and geriatric T cells (Fig. 2q)32,33. Further, we observed that T cell inhibitory genes (Ctla4, Pdcd1, Lag3, Havcr2) were also elevated in the geriatric T cells. We reason that the increase in T cell abundance observed only in old muscles but not in geriatric muscles can be explained by T cells shifting to an inhibited, exhausted state in older muscles and, therefore, being unable to proliferate productively by geriatric ages.
Independent of age, we detected few MuSCs and progenitors immediately following injury (days 1 and 2; 2.1% and 1.1%, respectively) and the abundance of MuSCs and progenitors peaked at day 5 (9.7%; Extended Data Fig. 4a and Supplementary Table 1a). This was in agreement with previous studies7,11. The peak in abundance of MuSCs and progenitors did vary by age with the young cells peaking at day 5, the old cells peaking at day 3.5 and the geriatric cells peaking at day 7 (Fig. 2i). This difference in peak abundance did not result in a statistically significant difference in the MuSCs and progenitors dynamics by age, but it did demonstrate a delayed response by the geriatric MuSCs and progenitors. Independent of age, we detected the most myonuclei at days 0 and 7 (15.1% and 10.6%, respectively; Extended Data Fig. 4a and Supplementary Table 1a). The myonuclei dynamics were very similar between the three ages, but we detected more myonuclei in old and geriatric samples at both days 0 and 7 compared to young (Fig. 2j and Extended Data Fig. 5m).
Senescence scoring based on single-cell transcriptomics
Next, we sought to investigate age-specific differences in senescence within skeletal muscle regeneration. Hallmarks of mammalian aging include stem cell exhaustion, altered cellular communication and cellular senescence34. Identifying senescent cells in scRNA-seq data is challenging because the markers traditionally used to identify senescent cells are lowly expressed, expressed in select cell types in single-cell data and/or assayed in terms of cellular localization and enzymatic function35. For example, senescent cells are commonly identified by persistent expression of cell cycle regulators p16 (Cdkn2a), p21 (Cdkn1a), p53 and/or Rb36. Senescent cells are also marked by the senescence-associated secretory phenotype (SASP), which includes pro-inflammatory cytokines and chemokines, growth modulators, angiogenic factors and matrix metalloproteinases (for example, Mmp3)37. To examine individual gene signatures of senescence, we quantified the abundance of Cdkn2a (encodes p16), Cdkn1a (encodes p21), Mmp3 (a senescence-associated matrix metalloproteinase) and Glb1 (encodes senescence-associated β-galactosidase) across all cell types and ages (Fig. 3a).
a, Dot plot showing the expression frequency and average expression level of select senescence markers. b–d, Scatterplots of the fraction of Cdkn2a+ cells within young, old and geriatric MuSCs and progenitors (n = 20, 21 and 24, respectively) (b), young, old and geriatric T cells (cycling; Cd3e+) (n = 15, 16, 19, respectively) (c), and young, old and geriatric FAPs (adipogenic; n = 20, 21 and 24, respectively) (d). Points are the fraction for each sample. Horizontal line is the median for each age group. Significance was evaluated using a two-sided Mann–Whitney U test. e, Line plot of the fraction of MuSCs and progenitors that coexpress Cdkn2a and Cdkn1a from 0 to 7 dpi. Points are the fraction for each sample (n = 4). Ribbon is the standard deviation. Statistical significance of age-specific dynamics was evaluated using a one-way ANOVA, and the FDR-corrected P value is reported. f, Scatterplot of the fraction of MuSCs and progenitors at 3.5 dpi that coexpress Cdkn2a and Cdkn1a by age. Points are the fraction for each sample (n = 4). Horizontal line is the median for each age group. Significance was evaluated using a two-sided Student’s t-test. g, Scatterplot of the normalized expression level of Cdkn2a and Cdkn1a in MuSCs and progenitors at 3.5 dpi. The density is shown on the top and to the right. Blue line represents the linear trend. Ribbon is the confidence interval. The inset contains the Pearson correlation coefficient and its statistical significance, calculated with the R function cor.test(alternative = ‘two.sided’). h, Significantly upregulated or downregulated GO terms between Cdkn2a+ and Cdkn1a+ (double positive) and all other (other) MuSCs and progenitors at 3.5 dpi. The normalized enrichment score and the FDR-corrected q values were obtained from GSEA. i,j, Feature plots of the final dataset with cells colored by the rescaled two-way FBR Sen Score (i) and the rescaled one-way FBR Sen Score (j). Cells are randomly plotted. k,l, Violin plots of the two-way (k) and one-way (l) FBR Sen Scores in select cell types. m–p, ROC curves based on the coexpression of Cdkn2a and Cdkn1a for the six two-way in vitro Sen Scores (m), the four two-way in vivo Sen Scores (n), the six one-way Sen Scores (o) and select two-way Sen Scores split by age (p). The AUC is reported for each ROC curve.
Expression of these genes depended more on cell type than age, when considering all time points and samples together (Fig. 3a). Given that Cdkn1a was widely expressed across many cell types, we focused on the common senescence hallmark gene Cdkn2a. Cdkn2a transcripts were rarely detected, in agreement with previous observations in the Tabular Muris Senis project8, and primarily observed in the MuSCs and progenitors and cycling T cells. We observed age-associated changes in the relative abundance of Cdkn2a+ cells within any given cell type infrequently significant (Fig. 3b–d). There was no significant difference in the fraction of Cdkn2a+ MuSCs and progenitors by age (Fig. 3b). We did observe a significant difference in the fraction of Cdkn2a+ cycling T cells between the young and old ages (Fig. 3c; Mann–Whitney U test, *P value = 0.04) and in the fraction of Cdkn2a+ adipogenic FAPs between the young and geriatric ages (Fig. 3d; Mann–Whitney U test, *P value = 0.02*).
Within the MuSCs and progenitors, we observed an increase in the fraction of cells that coexpressed Cdkn2a and Cdkn1a from day 0 to day 3.5, after which it returned to near day 0 levels (Fig. 3e). Although there was no significant difference by age in the abundance of these double-positive cells across all time points, there was a significant difference at day 3.5 between the young and old ages (Student’s t-test, *P value = 0.03) and the young and geriatric ages (Student’s t-test, *P value = 0.01; Fig. 3e,f). Although Cdkn2a and Cdkn1a are both cell cycle inhibitors and senescence markers, their expression was not correlated in MuSCs and progenitors at day 3.5 on the individual cell level in this dataset, possibly due to transcript detection dropout (Fig. 3g). We considered the double-positive Cdkn2a+ and Cdkn1a+ cells as candidate senescent MuSCs and progenitors and performed gene-set enrichment analysis (GSEA) on day 3.5 at their peak abundance. GSEA found that double-positive MuSCs and progenitors are enriched for gene sets associated with muscle weakness and various mitosis-related processes but diminished in muscle contraction and cytoskeletal processes (Fig. 3h). Collectively, these GSEA results suggest that double-positive MuSCs and progenitors have signatures of dysregulated muscle function and stalled cell cycle-related gene expression.
We then used the Cdkn2a+ Cdkn1a+ MuSCs and progenitors as a candidate cell population to evaluate broader senescence signatures at the single-cell level. We tested two senescence scoring methods38,39 and 14 senescence-signature (SenSig) gene lists38,40,41,42,43,44,45 (Supplementary Table 2). We refer to the first method as the two-way senescence score (Sen Score) because it calculates a score based on a list of upregulated and downregulated genes. Within this method, we tested nine gene lists that were experimentally derived38,40,44. We refer to the second method as the one-way Sen Score because it calculates a score based on a list of upregulated genes. Within this method, we tested six gene lists, two of which were taken from Gene Ontology (GO) databases42,43, one was curated in another report41 and three were experimentally derived38,45. We refer to the Methods, Fig. 4 and Supplementary Table 2 for more details on these two methods and the gene lists. We note that two of these gene lists are derived from bulk RNA-seq differential expression analyses of p16+ and p16− cells selected based on transgenic reporter status. One gene list (foreign body response, or FBR) was generated by Cherry et al. from p16+ versus p16− CD29+ cells isolated from an FBR-driven skeletal muscle fibrosis model in adult p16-CreERT2;Ai14 reporter mice38. A second gene list (‘aged chondrocytes’) was generated from p16+ versus p16− Aggrecan+ chondrocytes isolated from 20-mo p16-tdTom;Aggrecan-CreERT2;Ai6 mice (B.O. Diekman, personal communication)46,47,48. To compare how the choice of gene list and method impacted senescence scoring across different cell types, we examined the two-way and one-way FBR Sen Scores in the final scRNA-seq dataset. The two-way FBR Sen Scores were more consistently low across most cell clusters and exhibited high scores most notably in the FAPs and MuSCs and progenitors (Fig. 3i,k). The one-way FBR Sen Scores had broader distribution, with many more cell types exhibiting high scores, including FAPs, MuSCs and progenitors, endothelial cells, pericytes and smooth muscle cells and monocytes/macrophages (Fig. 3j,l). Whereas the two-way FBR Sen Scores were mean-centered around zero for each cell-type cluster due to their z-scored counts (Fig. 3k), one-way FBR Sen Scores had varied cell-type averages (Fig. 3l). These differences complicate establishing a threshold for senescence between cell types in the one-way Sen Scores.
a–f, ROC curves based on the coexpression of Cdkn2a and Cdkn1a for the nine two-way Sen Scores (a–c) and the six one-way Sen Scores (d–f) split into young (a and d), old (b and e) and geriatric (c and f) MuSCs and progenitors. g–j, ROC curves based on the coexpression of Cdkn2a and Cdkn1a for the nine two-way Sen Scores (g and h) and the six one-way Sen Scores (i and j) split into G1 (g and i) and non-G1 (S, G2, and M) (h and j) MuSCs and progenitors. k, Table of the AUC for each ROC curve. l–n, Venn diagrams of the unique and shared upregulated genes found in the two-way in vitro FBR SenSig gene lists (l), the two-way in vivo FBR SenSig gene lists (m) and the one-way in vivo FBR SenSig gene lists (n).
We established a scoring approach calibrated for sensitivity and specificity in discriminating Cdkn2a+ Cdkn1a+ MuSCs and progenitors across all ages and time points and present the results in receiver operating characteristic (ROC) curves with performance reported using an area under the curve (AUC) metric (Fig. 3m,p). Between the two senescence scoring methods and the 14 SenSig gene sets, the one-way FBR method performed the best of any experimentally derived approach (AUC = 0.86) and was comparable to the ontology-curated one-way GO:SASP approach (AUC = 0.88; Fig. 3m–o). Notably, it performed far better than the recently described SenMayo list while using the same single-sample GSEA method (AUC = 0.63). Moreover, the one-way FBR Sen Score accurately discriminated double-positive senescent-like MuSCs and progenitors across all three ages (AUCs = 0.85–0.86), suggesting it captures common features of senescence irrespective of age (Fig. 4d–f,k). We concluded that the one-way FBR method was able to accurately identify senescent-like cells in a manner that is not biased by a highly curated gene list.
While evaluating the SenSig gene lists, we posited that gene lists derived from aged mice would perform better when scoring aged cells. However, this is not the case as the SPIDER+/− young MuSCs and the SPIDER+/− geriatric MuSCs gene lists did not perform well in young (AUC = 0.47) and geriatric (AUC = 0.30) cells, respectively (Fig. 3p).
Refined analysis of myogenic subsets
In the final dataset, we identified two broad myogenic clusters (Fig. 1f). We observed a cluster of MuSCs and progenitors that expressed the myogenic transcription factor Pax7 (ref. 3) and a cluster of myonuclei that expressed Acta1, Myh1 and Myh4, genes critical for the contractile function of mature skeletal muscle cells49 (Extended Data Fig. 3c). We subsetted out these myogenic clusters and re-clustered and re-embedded the cells, resulting in nine distinct subclusters for refined annotation (Fig. 5a,b). We identified four progenitor populations that expressed Pax7, Myf5, Myod1, Myog, Mymk and Mymx and three myonuclei subtypes (IIx, IIb, IIx/IIb) that expressed Acta1, Ckm and Tnnt3 (ref. 49). We observed two transcriptomically variant clusters, which expressed both myogenic markers and either endothelial cell markers like Cd34, Cdh5 and Pecam1 or monocyte and macrophage markers like Ccr2 and C1qa (Extended Data Fig. 7). We suspected these clusters were dominated by doublets. The first doublet subcluster contained cells that coexpressed Pax7 and Pecam1, suggesting that these were MuSCs and progenitors and endothelial cell doublets (Extended Data Fig. 7a). The second doublet subcluster contained cells that coexpressed Acta1 and C1qa, suggesting that these were myonuclei and monocyte/macrophage doublets (Extended Data Fig. 7b). We designated these subclusters as ‘doublets 1’ and ‘doublets 2’, respectively, and excluded them from subsequent analyses involving the myogenic subset.
a–c, Pseudotime organization of the myogenic cell subset after re-clustering. Myogenic cells were re-embedded with UMAP (a) and with PHATE (b) and are colored by general myogenic IDs. PHATE embeddings were used by Monocle 3 to organize the cells in pseudotime where the earliest pseudotime is in the upper-left corner and the latest pseudotime is in the upper-right corner. Myogenic cells were organized into 25 approximately equal bins of increasing pseudotime values (c). d, Each cell was assigned an S phase score and a G2M phase score using Seurat’s Cell Cycle Scoring method. Scatterplot of the two scores with the cells colored by the predicted cell cycle phase based on the two scores. e–g, Polar coordinates (d) were converted to Cartesian coordinates and rescaled to fit in a range from 0 to 1. Cells are colored by the predicted cell cycle phase based on the two scores (e) and by the log-normalized expression of Cdk1 (f) and Cdk4 (g). h–k, Dot plots of the average log-normalized expression of Pax7 (h), Myog (i) and Acta1 (j) and by the fraction of non-G1 (S, G2 and M) cells (k) in each dpi and pseudotime-based myogenic cell-state bin. The size of the circle is the percentage of cells in each pseudotime-based myogenic cell-state bin for each age and dpi combination. l, Top 40 differentially upregulated and downregulated genes between cells in MuSCs 1 and MuSCs 2–6 at day 3.5. All genes highlighted here have an FDR-corrected q value < 0.05.
The type IIx cluster expressed Myh1, the type IIb cluster expressed Myh4, and the type IIx/IIb cluster expressed both Myh1 and Myh4 (Fig. 5b and Extended Data Fig. 8a–g)50. A previous study also identified a myonuclei cluster that expressed both Myh1 and Myh4, but this study concluded that this cluster represented cells with high metabolic activity, not cells that represent a transitional state between types IIx and IIb9. We interpret that the type IIx/IIb cluster identified here represents a transitional state between types IIx and IIb because 39% of the cells in this cluster coexpress Myh1 and Myh4 and because this cluster does not differentially express markers of high metabolic activity (Tnnc2, Tnni2, Mb, Cox6a2, Cox6c, Atp5e, Atp5g1; Extended Data Fig. 8f–h)9,51,52. Additionally, this transitional fiber type is the most common transitional fiber type in rat and mouse muscle fibers53,54. Notably, the type IIx/IIb cluster had a lower percentage of mitochondrial reads than the type IIx and type IIb clusters, indicating that these cells were not clustering together due to being lower quality (Extended Data Fig. 8I). We did not identify a cluster of neuromuscular junction or myotendinous junction cells due to a lack of Chrne or Col22a1 expression (Extended Data Fig. 8h)9,20.
We then re-embedded the myogenic subset using Potential of Heat-diffusion for Affinity-based Trajectory Embedding (PHATE)55, and these embeddings were used by Monocle 3 (refs. 56,57,58) to organize the cells in pseudotime (Fig. 5b,c). The pseudotime values were grouped into 25 bins that contain approximately equal numbers of cells. As the cells progressed through pseudotime, the order in which myogenic markers were expressed followed a typical trajectory of myogenesis. Early pseudotime bins had predominant expression of Pax7 and Myf5, but no strong expression of activation markers. Further in the pseudotime progression, cells still expressed Pax7 and Myf5, but they also expressed cycling markers such as Cdk1/Cdk4. In later pseudotime bins, cells expressed Myod1, Myog, Mymx and Mymk, markers of committed and fusing progenitors. In the latest pseudotime bins, cells expressed Acta1, Ckm, Myh1 and Myh4, markers of myogenic maturation (Extended Data Fig. 9a–h).
Pseudotemporal analysis of myogenesis progression
To directly compare myogenesis in regeneration responses, we assembled an annotated ‘cartography’ of myogenic progression arrayed across both day after injury and myogenic pseudotime. We first examined the percentage of cells that fell within each of the 25 initial pseudotime bins by time point and age, and then used expression frequency of myogenic marker genes to inform myogenic cell-state annotations (Extended Data Fig. 9a–h). Pseudotime bins 1–6 exhibited age-specific cell abundances, with bins 1–2 predominantly containing cells from young mice, bin 3 from young and old mice, bin 4 contained cells from old mice only, bin 5 from old and geriatric mice, and bin 6 from geriatric mice only. Cells in bins 1–6 expressed Pax7 and Myf5, which we annotated as ‘MuSCs’ (with sub-stages 1–6 preserved). Cells in bins 7–13 at 0 dpi expressed Pax7, Myf5 and Myod1 and lowly expressed cycling markers Cdk1 and Cdk4, which we annotated as ‘non-cycling MPCs’ (MPC, myogenic progenitor cell). Cells in bin 7 and dpi 1–3.5 and cells in bins 8–11 and dpi 1–7 expressed Myod1 and cycling markers Cdk1 and Cdk4, which we annotated as ‘cycling MPCs’. Cells in bin 12 at dpi 0–7 and cells in bin 13 at dpi 1–2 have diminishing expression of Myf5, Cdk1 and Cdk4, and increasing expression of Myog and Mymk, which we annotated as ‘committing MPCs’. Cells in pseudotime bin 13 and dpi 3.5–7 highly expressed Myog and Mymk and lowly expressed Cdk1 and Cdk4, which we annotated as ‘fusing myocytes’. Cells in pseudotime bins 14–25 and dpi 0–7 expressed Acta1, which we annotated as ‘myonuclei’. These pseudotime-informed myogenic cell-state aggregates (summarized in Extended Data Fig. 9i) were used in subsequent analyses.
To infer the cell cycle phases, we first assigned each myogenic cell S phase and G2M phase scores using Seurat’s standard Cell Cycle Scoring method59 (Fig. 5d). We treated these scores as polar coordinates, which were converted to Cartesian coordinates and normalized to be within a range from 0 to 1 (‘normalized theta’; Fig. 5e–g and Extended Data Fig. 10a,b). We assessed all myogenic cells within this cell cycle progression from a normalized theta of 0 to 1, corresponding to the continuum of G1–S–G2M stages (Fig. 5e and Extended Data Fig. 10a,b). When considering all MuSCs and progenitors, the distribution of normalized theta values increased from day 0 to day 3.5 and nearly returned to day 0 levels by day 7 in all age groups, suggesting a return to quiescence as expected (Extended Data Fig. 10e). We observed a shift to higher normalized theta values at 1 dpi in the geriatric samples compared to the young and old myogenic cells, suggesting an age-skewed cell cycle induction in early injury response that may represent a precocious activation phenotype. Differences by age group were minimal after 3.5 dpi. When examining all myogenic cells by the 25 pseudotime bins, we observed a shift in normalized theta values at bin 7 persisting through bin 13 (Extended Data Fig. 10f). Notably, pseudotime bins 7–13 also highly expressed the cycling markers Cdk1 and Cdk4 (Extended Data Fig. 9g,h).
We then found that Cdk1 and Cdk4 are more highly expressed in cells predicted to be in S or G2M than in cells predicted to be in G1 (Fig. 5f,g). Seurat’s standard G1 cutoff is at a normalized theta value of 0.25. Based on the expression of cycling markers Cdk1 and Cdk4 and the distribution of cells across the normalized theta values, we extended the G1 cutoff to 0.375 for this analysis (Fig. 5f,g and Extended Data Fig. 10c,d). For simplicity, cells with a normalized theta below and above 0.375 were classified as ‘G1’ and ‘non-G1’ (S/G2/M), respectively (Extended Data Fig. 10c,d).
We then calculated the percentage of cells within each pseudotime-informed myogenic cell state and time point by age group. Within MuSCs 1–6 at 0–1 and 5–7 dpi, we observed high levels of Pax7 (Fig. 5h). We also detected a high fraction of non-G1 cells in MuSCs 1–6, especially at 1–3.5 dpi in the old and geriatric mice (Fig. 5k). This suggested that more of the old and geriatric MuSCs were actively cycling after injury compared to the young MuSCs. We compared the 3.5 dpi cells in MuSCs 1 and MuSCs 2–6 by differential gene expression and found that the quiescence-associated genes Cdkn1c (encoding p57Kip2) and Socs3 were upregulated in MuSCs 1 and numerous translation-associated genes such as Rps29 were upregulated in MuSCs 2–6 (Fig. 5l). These expression profiles suggest that MuSCs 1 cells are in a less activated state MuSCs 2–6 cells.
We observed an inverse relationship between the average Myog expression and the fraction of non-G1 cells in the cycling MPCs, committing MPCs and fusing myocytes, as expected for differentiating myogenic cells (Fig. 5i,k). In all ages, we detected the highest average expression of Myog in the fusing myocytes population (Fig. 5i). We detected myonuclei in all ages and at every dpi, but there were fewer myonuclei with lower average expression of Acta1 at dpi 1–3.5 in the geriatric mice compared to the young and old mice (Fig. 5j). These results present an integrated cellular cartography of myogenic trajectories through regeneration, which exhibits age-associated cellular trajectories, particularly in the MuSC pool.
Senescence scoring across the myogenic cell cycle
To explore how cellular senescence manifests within this organized cartography of myogenesis, we focused on the one-way FBR Sen Score, which performed well across ages (Fig. 3o). To identify cells with senescent-like identities, we defined a threshold within the one-way FBR Sen Score based on its relationship with Cdkn2a and Cdkn1a expression (Extended Data Fig. 10g,h). We set a one-way FBR Sen Score threshold at 2,412, where 50% of cells above this value coexpressed Cdkn2a and Cdkn1a, and classified cells above this threshold as ‘Sen Score-high’ and senescent-like (Fig. 6a). We further observed through ROC analysis that the one-way FBR Sen Score could accurately identify double-positive Cdkn2a and Cdkn1a cells from both the G1 and non-G1 fractions of MuSCs and progenitors (Fig. 6b).
a, Scatterplot of the normalized Cartesian coordinate cell cycle scores and the one-way FBR senescence score (Sen Score) in all MuSCs and progenitors. Cells colored in red coexpress Cdkn2a and Cdkn1a. All other cells are colored in gray. The vertical line is the G1 cutoff, and the horizontal line is where 50% of the cells above this line coexpress Cdkn2a and Cdkn1a. b, ROC curves based on the coexpression of Cdkn2a and Cdkn1a in the MuSCs and progenitors (all ages and dpi) in G1 and non-G1 (S, G2 and M) when using the one-way FBR Sen Score. AUC is reported for each ROC curve. c,d, Dot plots of the fraction of cells that coexpress Cdkn2a and Cdkn1a (c) and the fraction of one-way FBR Sen Score-high cells (d) in each dpi and pseudotime-based myogenic cell-state bin. The size of the circle is the percentage of cells in each pseudotime-based myogenic cell-state bin for each age and dpi combination. e–g, Scatterplots of the fraction of MuSCs 1 (e), MuSCs 2–6 (f) and all MPCs (g) at 3.5 dpi that coexpress Cdkn2a and Cdkn1a by age. Points are the fraction for each sample (n = 2–4) and the horizontal line is the median for each age group. Significance was evaluated using a two-sided Welch’s t-test. h–j, Scatterplots of the fraction of MuSCs 1 (h), MuSCs 2–6 (i) and all MPCs (j) at 3.5 dpi that have a high one-way FBR Sen Score by age. Points are the fraction for each sample (n = 2–4), and the horizontal line is the median for each age group. Significance was evaluated using a two-sided Welch’s t-test.
We aimed to quantify the prevalence of cellular senescence within the cartography of myogenesis by age using both double-positive expression status and Sen Score, reasoning that the scores might capture a more expansive set of senescent-like cells. We observed a correspondence between the fraction of cells that coexpress Cdkn2a and Cdkn1a and the fraction of cells that have a high Sen Score in most ages, cell states and time points (Fig. 6c,d). Focusing on 3.5 dpi, we found a significantly higher fraction of Cdkn2a+ and Cdkn1a+ cells in geriatric MuSCs 1 compared to young MuSCs 1 (Fig. 6e; Welch’s t-test, *P value = 0.007). We did not observe age-specific differences in the fraction of Cdkn2a+ and Cdkn1a+ in MuSCs 2–6 (Fig. 6f). We found a significantly higher fraction of Cdkn2a+ and Cdkn1a+ in both old and geriatric MPCs (from both cycling and committing states) compared to young MPCs (Fig. 6g; Welch’s t-tests, *P values = 0.04 and 0.04, respectively). We observed a similar pattern in the fraction of cells that have a high one-way FBR Sen Score, with significantly higher frequencies in the old compared to young MuSCs 1 (Fig. 6h; Welch’s t-test, *P value = 0.006). We did not observe age-specific differences in the fraction of Sen Score high MuSCs 2–6 or MPCs (Fig. 6i,j). Together, these observations point to a transitory senescent-like cell population that is abundant at the self-renewing MuSC 1 stage across all ages, but increases in older mice, potentially underlying a stalled stem cell self-renewal in mouse muscle aging.
Evaluating muscle regeneration with spatial transcriptomics
To further investigate the aging-associated accumulation of senescent-like myogenic cells, we generated Curio Seeker spatial transcriptomic datasets from young and geriatric TAs at 5 dpi. Deconvolution was performed using the aging skeletal muscle regeneration scRNA-seq dataset presented here, and 15 cell types were identified in the spatial transcriptomic datasets (Fig. 7a,c). We delineated the spatial transcriptomic maps into an injury zone and a noninjury zone (Fig. 7b,d) based on immune cell density (Methods). We predominantly detected myonuclei, especially within the noninjury zone, in both the young and geriatric samples, as expected (Fig. 7e,f). Within the injury zone, we detected a greater diversity of cell types, including MuSCs and progenitors, fusing myocytes, monocytes/macrophages, B cells, T cells, dendritic cells, FAPs and endothelial cells (Fig. 7e,f). We observed some age-specific cell-type composition differences including a higher fraction of fusing myocytes and FAPs in the young injury zone and a higher fraction of monocytes/macrophages, T cells and myonuclei in the geriatric injury zone (Fig. 7e,f). Also, as expected, we detected a higher fraction of MuSCs and progenitors and fusing myocytes in the injury zone than the noninjury zone (Fig. 7e–l). We calculated a one-way FBR Sen Score based on a refined FBR SenSig gene list in the spatial datasets (Fig. 7m,n). There were significantly more FBR score-high senescent-like geriatric MuSCs and progenitors in the whole spatial dataset and specifically in the injury zone (Fig. 7o; Mann–Whitney U test, *P = 0.02 and *P = 0.03, respectively). When considering all other cell types, we found that there were significantly more geriatric senescent-like non-MuSC and progenitor cells in the whole spatial dataset and specifically within the noninjury zone (Fig. 7o; Mann–Whitney U test, *P = 7.53 × 10−38 and *P = 1.35 × 10−36, respectively). These high-resolution spatial transcriptomics maps provide further support for the conclusions from the scRNA-seq results that a senescent-like MuSC and progenitor cell population is more abundant in the geriatric mice, suggesting stalled stem cell self-renewal in aging.
a–d, Spatial transcriptomic maps of young (a and b) and geriatric (c and d) TA muscles 5 days after injury. Beads are colored by cell type (a and c) and injury zone (b and d). Scale bars, 500 µm. e,f, Percentage of cells of select cell types in the injury zone (e) and the noninjury zone (f) in the young and geriatric samples. g–n, Zoomed-in section of the young (g, i, k and m) and geriatric (h, j, l and n) spatial maps (as outlined in a–d) with beads colored by all cell types (g and h), MuSCs and progenitors (i and j), fusing myocytes (k and l) and one-way FBR Sen Score (m and n). o, Box plots of the one-way FBR Sen Score in the MuSCs and progenitors and all other cell types within the full transcriptomic map and within the injury and noninjury zones. The center line is the median, box limits are the upper and lower quartiles, whiskers are the 1.5 times the interquartile range and diamonds are the outliers. From left to right, n = 168, 99, 120, 88, 48, 11, 31,936, 2,2821, 11,361, 9,609, 20,575 and 13,212. Significance was evaluated using a two-sided Mann–Whitney U test.
Discussion
We utilized the profiling depth and complexity of scRNA-seq and associated computational analyses to generate a comprehensive compendium of 273,923 single-cell transcriptomes from regenerating TA muscles throughout mouse lifespan. To our knowledge, our dataset is the most comprehensive portrait of muscle repair at the single-cell level to date, as it includes three ages (young, old and geriatric), six time points (days 0, 1, 2, 3.5, 5 and 7) and 29 different cell-type clusters (Fig. 1 and Extended Data Figs. 1 and 3). Additionally, compared to previous scRNA-seq and single-nucleus RNA-sequencing skeletal muscle studies, we have identified more specific endothelial, FAP and immune cell subtypes7,9,10,11,12,13,14,15,16,17,18,19,20,21,30.
The immune, stromal and myogenic cells found in skeletal muscle contribute to muscle maintenance and regeneration by regulating MuSC quiescence, proliferation and differentiation1. It has been shown that an imbalance in immune cell populations during injury response can disrupt proper muscle repair1,2. To investigate this, we compared the change in cell-type abundances over our regeneration time course between young, old and geriatric muscles. As expected, neutrophils are one of the first immune cell types to peak in abundance (Extended Data Fig. 5l)4. We also observe monocyte and macrophage populations that express pro-inflammatory markers like Ccr2 and patrolling markers like Ctsa responding soon after injury (days 1–2) when we expect the muscle environment to be enriched with pro-inflammatory cytokines (Fig. 2a,e)1,4. Monocytes and macrophages that express pro-inflammatory markers clear cellular debris and promote myogenic cell proliferation1,60. There should be a shift to monocytes and macrophages that express anti-inflammatory marker C1qa at 4–7 dpi (Fig. 2b,c and Extended Data Fig. 3a)4. We do broadly observe a shift from monocytes and macrophages that express pro-inflammatory markers to anti-inflammatory markers, but there are significant differences by age (Fig. 2a–e). This difference in monocyte and macrophage dynamics could explain the age-related decline in muscle repair because if macrophages do not clear cellular debris or promote myogenic cell proliferation and differentiation, the muscle remains inflamed and there are repeated cycles of necrosis and regeneration60. The damaged myofibers are then replaced with adipose tissue, fibrotic tissue or bone, instead of new myofibers60.
In addition to age-specific differences in the dynamics of the monocyte and macrophage populations, we observe age-specific differences in the T cell dynamics (Fig. 2l–o and Extended Data Fig. 5t). It has previously been shown that regulatory T cells, marked by Cd4 and Foxp3, accumulate in injured muscle and peak in abundance at day 4 (ref. 4). We detected Foxp3 expression in a few T cells, specifically in the T cell cycling and T cell non-cycling populations, which both highly express Cd3e (Extended Data Fig. 3a). Although we cannot confidently identify any of our three T cell populations as regulatory T cells, we do observe a peak in T cell abundances at days 5 and 7 (age-specific). There is miscoordination of the T cell response, which in turn could impact the ability of aged muscle to repair itself.
One factor that has been shown to contribute to the reduced functionality of MuSCs in aged tissues is the establishment of senescent MuSCs2,6. Prior studies have used the cell cycle proteins p16, p21, p53 and Rb to differentiate between dividing and non-dividing cells, but the non-dividing cells can include senescent cells and quiescent cells36. Senescence-associated beta-galactosidase is also commonly used to identify senescent cells, but it is also detected in quiescent cells and in stressed cells36,61. Because these markers are not unique to senescent cells and because senescent cells are heterogeneous, it has been challenging to identify biomarkers that can accurately and consistently identify senescent cells across species, tissues and conditions62. Indeed, recent large consortia have been established to develop new tools to detect and bioinformatically identify senescent cells with robustness and precision throughout mammalian tissues and lifespans63. Additional studies have aimed to understand the role of cellular senescence and its mechanisms in muscle aging44,64,65,66.
Given the role that cellular senescence plays in limiting cell contributions in aged tissues, we tested a series of tools to bioinformatically identify senescence in these single-cell data and found a transfer-learning-based scoring approach accurately classified senescent-like myogenic cells across ages and cell cycling states. The approach described here to quantitatively assess various senescence scoring approaches and reference gene lists in discriminating senescent or senescent-like cells (Figs. 3 and 5) may provide a template for future studies using single-cell data. Notably, here we concluded that a skeletal muscle FBR gene list more accurately and robustly discriminated senescent-like Cdkn2a+ and Cdkn1a+ MuSCs and progenitors in these muscle regeneration datasets than did a variety of experimental and curated gene lists, including the recently described SenMayo list41. In particular, at 3.5 dpi, we observed a higher fraction of Cdkn2a+ and Cdkn1a+ cells in the old and geriatric MuSCs associated with a self-renewing cell stage (MuSCs 1; Fig. 6e). Likewise, we observed a similar pattern in the fraction of one-way FBR Sen Score-high cells, with significantly higher frequencies in the old compared to young MuSCs 1 (Fig. 6h).
We identified similar broad cell types within Curio Seeker spatial transcriptomic maps from young and geriatric TAs at 5 dpi (Fig. 7a,c). As expected, the injury zone contains a greater diversity of cell types, including a higher fraction of MuSCs and progenitors and fusing myocytes (Fig. 7e–l). In agreement with the scRNA-seq data, we observed significantly more senescent-like geriatric MuSCs and progenitors in the whole spatial dataset and specifically in the injury zone (Fig. 7o). The spatial datasets support the conclusion discussed with the scRNA-seq dataset where we observed a senescent-like MuSC and progenitor population that is more abundant in the geriatric mice, suggesting stalled stem cell self-renewal in mouse muscle aging. Together, these observations point to a transitory senescent-like cell population that is abundant at the self-renewing MuSC stage during regeneration across all ages of mice. This population of senescent-like MuSCs increases within the injury zone and in older mice, suggesting that a stalled stem cell self-renewal state underlies the regenerative dysfunction in mouse aging.
Methods
Mouse muscle injury and single-cell isolation
The Cornell University Institutional Animal Care and Use Committee approved all animal protocols (approval no. 2014-0085), and experiments were performed in compliance with its institutional guidelines. Mice were maintained at 70–73 °F on a 14-h light/10-h dark cycle with humidity mainly at 40%. Muscle injury was induced in young (4–7 mo), old (20 mo) and geriatric (26 mo) C57BL/6J mice (Jackson Laboratory, 000664; NIA Aged Rodent Colonies) by injecting both TA muscles with 10 µl of notexin (10 µg ml−1; Latoxan). The mice were euthanized, and TA muscles were collected at 0, 1, 2, 3.5, 5 and 7 dpi with n = 3–4 biological replicates per sample. Each TA was processed independently to generate single-cell suspensions. At each time point, the young and old samples are biological replicates of TA muscles from distinct mice, and the geriatric samples are biological replicates of two TA muscles from each of two mice. A mixture of male and female mice was used. See Supplementary Table 1 for additional detail. Muscles were digested with 8 mg ml−1 collagenase D (Roche) and 10 U ml−1 Dispase II (Roche) and then manually dissociated to generate cell suspensions. Myofiber debris was removed by filtering the cell suspensions through a 100-µm and then a 40-µm filter (Corning Cellgro, 431752 and 431750). After filtration, erythrocytes were removed by incubating the cell suspension in erythrocyte lysis buffer (IBI Scientific, 89135-030).
scRNA-seq library preparation
After digestion, the single-cell suspensions were washed and resuspended in 0.04% BSA in PBS at a concentration of 106 cells per ml. A hemocytometer was used to manually count the cells to determine the concentration of the suspension. scRNA-seq libraries were prepared using the Chromium Single Cell 3′ reagent kit v3 (10x Genomics) following the manufacturer’s protocol67. Cells were diluted into the Chromium Single Cell A Chip to yield a recovery of 6,000 single-cell transcriptomes with <5% doublet rate. Libraries were sequenced on the NextSeq 500 (Illumina)68. The sequencing data were aligned to the mouse reference genome (mm10) using Cell Ranger v5.0.0 (10x Genomics)67.
Preprocessing scRNA-seq data
From the gene expression matrix, the downstream analysis was carried out in R (v3.6.1). First, ambient RNA signal was removed using the default SoupX (v1.4.5) workflow (autoEstCounts and adjustCounts; https://github.com/constantAmateur/SoupX/)27. Samples were then preprocessed using the standard Seurat (v3.2.3) workflow (NormalizeData, ScaleData, FindVariableFeatures, RunPCA, FindNeighbors, FindClusters and RunUMAP; https://github.com/satijalab/seurat/)59. Cells with fewer than 200 genes, fewer than 750 unique molecular identifiers and more than 25% of unique transcripts derived from mitochondrial genes were removed. After preprocessing, DoubletFinder (v2.0.3) was used to identify putative doublets in each dataset28. The estimated doublet rate was 5% according to the 10x Chromium handbook. The putative doublets were removed from each dataset. Next, the datasets were merged and then subjected to batch correction with Harmony (https://github.com/immunogenomics/harmony/; v1.0)29. Seurat was then used to process the integrated data. Dimensions accounting for 95% of the total variance were used to generate SNN graphs using FindNeighbors, and SNN clustering was performed using FindClusters. A clustering resolution of 0.8 was used resulting in 24 initial clusters.
Cell-type annotation in scRNA-seq data
Cell types were determined by expression of canonical genes. Each of the 24 initial clusters received a unique cell-type annotation. The nine myeloid clusters were challenging to differentiate between, so these clusters were subset out (Subset) and re-clustered using a resolution of 0.5 (FindNeighbors, FindClusters) resulting in 15 initial clusters. More specific myeloid cell-type annotations were assigned based on expression of canonical myeloid genes. This did not help to clarify the monocyte and macrophage annotations, but it did help to identify more specific dendritic cell and T cell subtypes. These more specific annotations were transferred from the myeloid subset back to the complete integrated object based on the cell barcode.
Analysis of cell-type dynamics
We generated a table with the number of cells from each sample (n = 65) in each cell-type annotation (n = 29). We removed the erythrocytes from this analysis because they are not a native cell type in skeletal muscle. Next, for each sample, we calculated the percentage of cells in each cell-type annotation. The mean and standard deviation were calculated from each age and time point for every cell type. The solid line is the mean percentage of the given cell type, the ribbon is the standard deviation around the mean and the points are the values from individual replicates. We evaluated whether there was a significant difference in the cell-type dynamics over all six time points using nonlinear modeling. The dynamics for each cell type were fit to some nonlinear equation (for example, quadratic, cubic, quartic) independent and dependent on age. The type of equation used for each cell type was selected based on the confidence interval and significance (P < 0.05) of the leading coefficient. If the leading coefficient was significantly different from zero, it was concluded that the leading coefficient was needed. If the leading coefficient was not significantly different than zero, it was concluded that the leading coefficient was not needed, and the degree of the equation went down one. No modeling equation went below the second degree. The null hypothesis predicted that the coefficients of the nonlinear equation were the same across the age groups, while the alternative hypothesis predicted that the coefficients of the nonlinear equation were different across the age groups. We conducted a one-way ANOVA to see if the alternative hypothesis fits the data significantly better than the null hypothesis and we used FDR as the multiple-comparison test correction (using the ANOVA and p.adjust (method = fdr) functions in R, respectively).
Muscle immunohistochemical analysis
Muscle injury was induced in young, old and geriatric C57BL/6J female mice by injecting both TA muscles with 10 µl of notexin (10 µg ml−1; Latoxan). The mice were euthanized, and TA muscles were collected at 0 and 5 dpi. The TA muscles were coated in Tissue-Tek O.C.T. Compound (Sakura Finetek, 4583), snap frozen in liquid nitrogen-cooled isopentane (Thermo Scientific Chemicals, AA19387AP) and then stored at −80 °C. Frozen TA muscles were sectioned with a cryostat transversely at 5-µm thickness, and section slides were stored at −20 °C until stained. Sections were fixed with 4% PFA (Electron Microscopy Sciences, 15710) for 10 min, washed with 1× PBS, and blocked with 3% BSA (Rockland Immunochemicals, RLBSA50) at room temperature for 1 h. Sections were washed with 1× PBS and then stained with rat anti-mouse CD3 (eBioscience, 14-0032-82) at 1:100 dilution in blocking buffer overnight at 4 C. Sections were then washed and stained with Alexa Fluor Plus 750 Phalloidin (Life Technologies, A30105) at a 1:500 dilution and goat anti-rat 488 (Invitrogen, A-11006) at a 1:250 dilution in blocking buffer for 1 h at room temperature protected from light. Sections were then washed with 1× PBS and stained with 5 mg ml−1 DAPI (Life Technologies, D3571) at a 1:1,000 dilution. Slides were mounted with glycergel mounting medium (Agilent, C056330-2) and stored at 4 °C before imaging. Images were acquired using a Nikon Eclipse Ti-E microscope (Micro-Video Instruments), and were analyzed using NIS Elements 5.11.03 software and ImageJ 2.1.0.
Immune cell flow cytometric analysis
Single-cell suspensions of uninjured day 0 and injured day 5 TAs and gastrocnemius muscles from young, old and geriatric C57BL/6J female mice were collected in the same way as the single-cell suspensions for scRNA-seq library preparation. However, the single cells were suspended in 90% FBS and 10% dimethyl sulfoxide and frozen at −80 °C. When ready to use, the single-cell suspensions were thawed in a 37 °C water bath and then transferred to 15-ml conical tubes. The cells were washed with staining buffer (1× PBS + 0.5% BSA + 2 mM EDTA) before being spun at 500g for 3 min and transferred to a 96-well round-bottom plate. Cells were incubated with FC block (TruStain FcX PLUS (anti-mouse CD16/32), BioLegend, 156604) at 4 °C for 5 min. Cells were then washed with staining buffer, spun at 500 g for 3 min and incubated with viability dye (Fixable viability dye APCe780, eBioscience, 65-0865-14) and surface antibody (see ‘Flow cytometry antibodies’) in Brilliant Stain Buffer Plus (BD Biosciences, 566385) and staining buffer for 30 min at 4 °C in the dark. Aliquots of cell samples were counted on a MoxiZ Mini Automated Cell Counter. After incubation, we followed the manufacturer’s protocol (FOXP3 transcription factor fixation/permeabilization kit, eBioscience, 00-5521-00) and cells were washed with staining buffer, spun and resuspended in FoxP3 1× perm solution (10× Permeabilization buffer, Invitrogen, 00-8333-56) and incubated for 30 min at 4 °C in the dark. Cells were washed with 1× perm solution, spun twice and resuspended in an intracellular antibody stain. Cells were incubated in intracellular antibody stain for 30 min at 4 °C in the dark. Cells were washed with 1× perm and spun twice before being resuspended in staining buffer and transferred to 40 µM blue-capped flow tubes. Fluorescence-minus-one controls were prepared using a mixture of young, old and geriatric uninjured and injured TA and gastrocnemius muscles. Flow cytometry was performed using a FACSymphony A3 (BD), and data were analyzed in FlowJo 10.5.3. Gates were determined using fluorescence-minus-one controls.
Flow cytometry antibodies
Antibody | Clone | Fluorophore | Manufacturer catalog no. | Dilution strategy |
|---|---|---|---|---|
CD45 | 30-F11 | PECy7 | Invitrogen, 25-0451-82 | 1:250 |
CD11b | M1/70 | BUV395 | BD Biosciences, 563553 | 1:250 |
CD11c | N418 | PerCPCy5.5 | Invitrogen, 45-0114-82 | 1:250 |
TCRβ | H57-597 | APC | eBioscience, 17-5961-82 | 1:250 |
CD19 | 1D3 | BV650 | BD Biosciences, 563235 | 1:250 |
T cell exhaustion scoring
We grouped the three T cell populations (including Cd3e+ cycling and non-cycling T cells and Cd4+ T cells) and z-scored all genes. The T cell exhaustion score was calculated using a transfer-learning method developed by Cherry et al. and a T cell exhaustion gene list from Bengsch et al.38,69. The Mann–Whitney U test was performed on the T cell exhaustion score between ages.
Senescence scoring
We tested two senescence scoring methods along with 14 senescence gene lists (Supplementary Table 2) to identify senescent-like cells within the scRNA-seq dataset. The two-way senescence score (Sen Score) was calculated using a transfer-learning method developed by Cherry et al.38. With this method, we tested 14 gene lists: stimulation independent, replicative, oncogene, ion-radiation induced, senescence-associated core, aged-chondrocyte, two-way FBR, SPiDER young MuSCs and SPiDER geriatric MuSCs. The one-way Sen Score was calculated using single-sample GSEA with the R package escape (v1.4.2)39 as per Saul et al.41. With this method, we tested six gene lists: one-way FBR, SenMayo, GO:Senescence, GO:SASP, lung uninjured and lung injured.
We evaluated the ability of the two methods and the 14 gene lists to accurately identify senescent-like MuSCs and progenitors by calculating a ROC curve. For each MuSC and progenitor cell, we evaluated whether it expressed both Cdkn2a and Cdkn1a. This analysis was done for all MuSCs and progenitors (Fig. 3m–p), MuSCs and progenitors split by age (Figs. 3p and 4a–f), and MuSCs and progenitors split by G1 status (Figs. 4g–j and 6b). The AUC was calculated for each ROC curve (Figs. 3m–p, 4k and 5b).
Given that the one-way Sen Score with the FBR gene list performed the best (AUC = 0.86), we focused on that for further analyses. We next set a threshold of senescence based on the one-way FBR Sen Score where 50% of the MuSCs and progenitors with at least that score coexpress Cdkn2a and Cdkn1a. For the one-way FBR Sen Score, MuSCs and progenitors with a score ≥ 2412.562 were called ‘senescent-like’, while all other cells were called ‘not senescent-like’ (Fig. 6a and Extended Data Fig. 10g,h). We refer to Supplementary Note 2 for more details on how the senescence scoring was performed and evaluated.
Cell cycle scoring
To each cell in the final dataset, we assigned an S phase score, a G2M phase score and a discrete phase classification (G1/S/G2M) using Seurat’s standard Cell Cycle Scoring method59. We have treated the S phase and G2M phase score as polar coordinates to help us visualize how cells are progressing through the cell cycle (Fig. 5d). We converted the polar coordinates to Cartesian coordinates and normalized the theta values so that they range from 0 to 1 to ensure that cells in G1 have the lowest theta values followed by cells in S and G2M (Fig. 5e and Extended Data Fig. 10a,b). This enables us to see how cells are progressing linearly through the cell cycle. Seurat’s standard cutoff between cells classified as G1 versus cells classified as S is at the normalized theta value of 0.25. Looking at the distribution of cells across the normalized theta values as well as the expression of cell cycle markers Cdk1 and Cdk4, we decided to extend the G1 to S cutoff to 0.375 (Fig. 5e–g and Extended Data Fig. 10c,d). Cells with a normalized theta value ≥ 0.375 are considered non-G1 (S/G2/M).
Myogenic cell subsets
From the final dataset, the cells with the cell-type IDs ‘MuSCs and progenitors’ and ‘myonuclei’ were subset out and the Seurat workflow was partially rerun (ScaleData, FindVariableFeatures, RunPCA, FindNeighbors, FindClusters and RunUMAP). Dimensions accounting for 95% of the total variance were used to generate SNN graphs (FindNeighbors) and SNN clustering was performed (FindClusters)59. A clustering resolution of 0.7 was used resulting in nine clusters. These nine clusters were assigned general cell-type IDs based on canonical myogenic markers. Of the nine clusters, four were progenitor subtypes, three were myonuclei subtypes, and two were doublets (Fig. 5a).
To more specifically identify the doublet clusters, we looked at the coexpression of myogenic and non-myogenic markers (Extended Data Fig. 7). Using GetAssayData, we extracted the log-normalized expression values of Pax7, Pecam1, Acta1 and C1qa in each cell in the nine myogenic clusters. For each of the nine clusters, we plotted cells by their expression values of Pax7 and Pecam1 and by their expression values of Acta1 and C1qa. A density plot was plotted along the x and y axes using ggmarginal(type = ‘density’; Extended Data Fig. 7). The two clusters identified as doublets were excluded from the remaining myogenic subset analyses.
To identify the myonuclei clusters more specifically, we looked at the expression of myonuclei markers, markers of high metabolic activity, and the percentage of unique transcripts derived from mitochondrial genes (Extended Data Fig. 8). Using GetAssayData, we extracted the log-normalized expression values of Myh1 and Myh4 in each cell in the three myonuclei clusters. For every cell, as defined by the cell barcode, we determined whether the expression value equaled zero (no expression) or exceeded zero (expression) for both Myh1 and Myh4 independently. For each of the three myonuclei clusters, the fraction of cells that expressed Myh1 and Myh4, only Myh1, only Myh4, and neither Myh1 nor Myh4 were calculated by dividing the number of cells that expressed Myh1 and Myh4, only Myh1, only Myh4, and neither Myh1 nor Myh4 by the total number of cells within each myonuclei cluster (Extended Data Fig. 8d–g).
Harmony embedding values from the dimensions accounting for 95% of the total variance were used for further dimensional reduction with PHATE, using phateR (v1.0.7; Fig. 5b,c)55. The PHATE embedding values were used by Monocle 3 (v1.0.0)56,57,58. The normal Monocle 3 workflow was used (cluster_cells, estimate_size_factors, learngraph, order_cells) where L1.sigma = 0.4 and the root cell was in the progenitor 1 cluster. The pseudotime values for each cell as defined by Monocle 3 were transferred from the Monocle 3 CDS object to the myogenic cells only Seurat object by cell barcode. The pseudotime values were divided into 25 bins with approximately equal numbers of cells (1,089–1,090 cells per bin; Fig. 5c). We assigned myogenic cell-type IDs based on known myogenic marker expression in each pseudotime bin and dpi (Extended Data Fig. 9). We refer to the Supplementary Note 2 for more details on how the myogenic cell-type IDs were assigned.
We focused on dpi 3.5 and compared the cells in MuSCs 1, MuSCs 2–6 and all MPCs (this includes cells classified as cycling MPCs and cells classified as committing MPCs). For each sample, we calculated the fraction of cells that had log-normalized Cdkn2a and Cdkn1a counts greater than 0 (Fig. 6e–g). Within these same groupings, we also calculated the fraction of cells that had a one-Way FBR Sen Score greater than 2412.562 (Fig. 7h–j). We conducted two-sided Welch’s t-tests to evaluate whether there was a significant difference between ages.
Differential expression and GSEA
For select comparisons, we used Seurat’s FindAllMarkers() function to identify genes that were differentially expressed between groups. In the myogenic subset with all ages at day 3.5, we conducted this analysis between the cells in ‘MuSCs 1’ and the cells in ‘MuSCs 2–6’ (Fig. 5l). In the MuSCs and progenitors with all ages at day 3.5, we performed this analysis between cells that coexpressed Cdkn2a and Cdkn1a (we refer to these cells as ‘double positive’) and all other cells (we refer to these cells as ‘other’). Genes that had an FDR-corrected P value ≤ 0.05 were ranked by average log2 fold change and used in a GSEA (v4.1.0). The gene-set databases used included h.all.v2023.1.Hs.symbols.gmt, c2.all.v2023.1.Hs.symbols.gmt, 5.all.v2023.1.Hs.symbols.gmt and c8.all.v2023.1.Hs.symbols.gmt. Significant GO terms (FDR q value ≤ 0.25) were ranked by the normalized enrichment score (enrichment scores normalized by the size of the gene set; Fig. 3h).
Tissue collection for spatial transcriptomics
Muscle injury was induced in young and geriatric C57BL/6J female mice by injecting both TA muscles with 10 μl of notexin (10 μg ml−1; Latoxan). The mice were euthanized, and TA muscles were collected at 5 dpi. The TA muscles were coated in Tissue-Tek O.C.T. Compound (Sakura Finetek, 4583), snap frozen in liquid nitrogen-cooled isopentane (Thermo Scientific Chemicals, AA19387AP), and then stored at −80 °C. Frozen TA muscles were sectioned with a cryostat transversely at 10-µm-thick slices at −20 °C. Slide-seq spatial transcriptomics70 experiments were performed using the Curio Seeker Kit (Curio Bioscience). The sections were mounted on Curio Seeker tiles (Curio Bioscience) and further processed to generate spatial transcriptomics libraries.
Library preparation for spatial transcriptomics
Sequencing libraries were generated for the mounted young (Curio Tile ID: A0018_011) and geriatric (Curio Tile ID: A0018_012) TA muscle sections with the Curio Seeker Spatial Transcriptomics kit. After RNA hybridization and reverse transcription, the tissue sections underwent enzymatic digestion, and the beads were detached from the glass tile and suspended in solution. This was followed by second-strand cDNA synthesis and amplification. Sequencing libraries were prepared using the Nextera XT DNA sample preparation kit, pooled and sequenced on an Illumina NextSeq 2,000 system using a P2 flow cell and a 100-cycle kit (read 1 = 50 bp, read 2 = 72 bp, index 1 = 8 bp, index 2 = 8 bp). The sequencing data were processed with the slidesnake pipeline (https://github.com/mckellardw/slide_snake/). The ‘seeker_v3.1_internalTrim’ recipe was used to trim adaptors (Cutadapt v4.6)71 and low-quality reads before mapping to the mouse genome (GRCm39) using the STAR Solo (STAR v2.7.10b) pipeline that generates a feature-by-bead barcode expression matrix.
Preprocessing Slide-seq data for quality control
AnnData (v0.10.3)72 objects were generated from the count matrices of each sample, and Scanpy (v1.9.3)73 was used to load the spatial coordinates of each bead as provided by Curio Bioscience. Beads with fewer than 20 transcripts and genes detected in less than ten beads were removed. The expression data were then scaled and log-normalized. Additionally, the spatial distances between all pairs of beads were calculated and beads with less than ten neighboring beads within 100 μm were removed74.
Cell-type deconvolution in Slide-seq data
The Curio Seeker technology utilizes a dense array of beads, 10 μm in diameter, that capture polyadenylated mRNA from a tissue section placed on the array70. The beads collect transcriptomes predominantly from cells in direct contact with them. Although the size of the beads allows for close to single-cell spatial profiling, multiple cells contribute to the transcriptome captured on each bead75. Deconvolution approaches, where the gene expression signature from a reference single-cell dataset is used to predict the cell-type contribution of each bead transcriptome, are used to construct cell-type maps. Here, we applied the cell2location model (v0.1.3)75 to deconvolve the spatial transcriptomics data against the aged scRNA-seq dataset presented in this paper. The scRNA-seq reference was filtered to retain only genes that are highly expressed and informative for identifying rare cell types (cell_count_cutoff = 5, cell_percent_cutoff = 0.01, and nonz_mean_cutoff = 1.12). Using these refined gene lists, cell-type-specific expression signatures were derived using negative binomial regression. The resulting signatures were applied to the Slide-seq data to assign cell-type identities based on the highest prediction scores, with hyperparameters set to N_cells_per_location = 1 and detection_alpha = 20.
Defining the injury zone in Slide-seq data
To define the injury zone, we used a k-nearest-neighbors algorithm to calculate immune cell density around each tissue spot. Each spot was assigned an Immune_neighbors score based on the number of immune cells (T cells, B cells, NK cells, dendritic cells, neutrophils and monocytes/macrophages) within its 300 nearest neighbors. We established injury zones by identifying spots with an Immune_neighbors score exceeding 20. Additionally, to focus on the high-density injury zone, additional specific spatial criteria were applied: spots with y coordinates less than 1,500 μm for the young sample, and spots with x coordinates greater than 1,945 μm for the geriatric sample were classified as within the injury zone. Spots failing to meet these criteria were designated as a noninjury zone. To refine the injury zone identification and remove isolated spots, we applied a k-nearest-neighbors algorithm with k = 20, focusing on neighbors classified as ‘injury’ spots. We reclassified spots with more than ten ‘injury’ neighbors as part of the injury zone and spots with ten or fewer ‘injury’ neighbors as ‘noninjury’, effectively discarding isolated spots and small clusters. This step ensured that the defined injury zones represented coherent clusters of high-density injury spots.
Senescence score calculation in Slide-seq data
The one-way FBR Sen Score was calculated on a refined FBR gene list. This refined list contained 44 genes that were captured in more than two beads assigned with the MuSCs and progenitors label in both samples. To calculate the Sen Score, we used the Scanpy tool score_genes function with the parameters set to 1,000 bins and a control size of 50. The score was calculated on the normalized matrix of each sample. Age-specific significance was evaluated using a two-sided Mann–Whitney U test.
Statistics and reproducibility
No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications7,9,10,11,12,13,14,15,16,17,18,19,20,21,30. For the new scRNA-seq data presented here, replicates for the same age and time point were not all processed on the same day. Data collection and analysis were not performed blind to the conditions of the experiments. When statistical tests were used, the assumptions of the test were met before conducting the statistical test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
A complete list of metadata and Gene Expression Omnibus (GEO) accessions for the scRNA-seq data can be found in Supplementary Table 1. Previously published scRNA-seq data are deposited in the GEO under accessions GSE143437, GSE159500 and GSE162172. Newly collected scRNA-seq data from young (4.7 mo; days 1 and 3.5) and geriatric (26 mo) mice are deposited in the GEO under accession GSE232106. Newly collected Curio Seeker spatial transcriptomics data from young (4 mo) and geriatric (26 mo) mice are deposited in GEO under accession GSE266933. Fully processed Seurat objects for the final dataset (Fig. 1) and the myogenic subset (Figs. 4 and 5) and the fully processed Scanpy objects for the young and geriatric spatial transcriptomics data (Fig. 7) are available for download on Dryad (https://doi.org/10.5061/dryad.kkwh70sbv). Gene lists used in the senescence scoring analysis are compiled in Supplementary Table 2. Source data are provided with this paper.
Code availability
All newly developed code for the scRNA-seq analysis is available on GitHub (https://github.com/ldwalter/). Code related to the spatial transcriptomics analysis is also available on GitHub (https://github.com/ntekasi/ST_MuSCs/).
References
Bentzinger, C. F., Wang, Y. X., Dumont, N. A. & Rudnicki, M. A. Cellular dynamics in the muscle satellite cell niche. EMBO Rep. 14, 1062–1072 (2013).
Blau, H. M., Cosgrove, B. D. & Ho, A. T. V. The central role of muscle stem cells in regenerative failure with aging. Nat. Med. 21, 854–862 (2015).
Wang, Y. X. & Rudnicki, M. A. Satellite cells, the engines of muscle repair. Nat. Rev. Mol. Cell Biol. 13, 127–133 (2011).
Tidball, J. G. Regulation of muscle growth and regeneration by the immune system. Nat. Rev. Immunol. 17, 165–178 (2017).
Chakkalakal, J. V., Jones, K. M., Basson, M. A. & Brack, A. S. The aged niche disrupts muscle stem cell quiescence. Nature 490, 355–360 (2012).
Sousa-Victor, P. et al. Geriatric muscle stem cells switch reversible quiescence into senescence. Nature 506, 316–321 (2014).
Oprescu, S. N., Yue, F., Qiu, J., Brito, L. F. & Kuang, S. Temporal dynamics and heterogeneity of cell populations during skeletal muscle regeneration. iScience 23, 100993 (2020).
Almanzar, N. et al. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Chemello, F. et al. Degenerative and regenerative pathways underlying Duchenne muscular dystrophy revealed by single-nucleus RNA sequencing. Proc. Natl Acad. Sci. USA 117, 29691–29701 (2020).
Cutler, A. A. et al. The regenerating skeletal muscle niche drives satellite cell return to quiescence. iScience 25, 104444 (2022).
De Micheli, A. J. et al. Single-cell analysis of the muscle stem cell hierarchy identifies heterotypic communication signals involved in skeletal muscle regeneration. Cell Rep. 30, 3583–3595 (2020).
Dell’Orso, S. et al. Single cell analysis of adult mouse skeletal muscle stem cells in homeostatic and regenerative conditions. Dev. Camb. Engl. 146, dev174177 (2019).
Dos Santos, M. et al. Single-nucleus RNA-seq and FISH identify coordinated transcriptional activity in mammalian myofibers. Nat. Commun. 11, 5102 (2020).
Giordani, L. et al. High-dimensional single-cell cartography reveals novel skeletal muscle-resident cell populations. Mol. Cell 74, 609–621 (2019).
Jin, R. M., Warunek, J. & Wohlfert, E. A. Chronic infection stunts macrophage heterogeneity and disrupts immune-mediated myogenesis. JCI Insight https://doi.org/10.1172/jci.insight.121549 (2018).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779 (2020).
Kimmel, J. C., Hwang, A. B., Scaramozza, A., Marshall, W. F. & Brack, A. S. Aging induces aberrant state transition kinetics in murine muscle stem cells. Development 147, dev183855 (2020).
Li, H. et al. Muscle-secreted granulocyte colony-stimulating factor functions as metabolic niche factor ameliorating loss of muscle stem cells in aged mice. EMBO J. 38, e102154 (2019).
Perez, K. et al. Single nuclei profiling identifies cell specific markers of skeletal muscle aging, frailty and senescence. Aging 14, 9393–9422 (2022).
Petrany, M. J. et al. Single-nucleus RNA-seq identifies transcriptional heterogeneity in multinucleated skeletal myofibers. Nat. Commun. 11, 6374 (2020).
Stepien, D. M. et al. Tuning macrophage phenotype to mitigate skeletal muscle fibrosis. J. Immunol. 204, 2203–2215 (2020).
Tabula Muris Consortium et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Kimmel, J. C. et al. Murine single-cell RNA-seq reveals cell-identity- and tissue-specific trajectories of aging. Genome Res. 29, 2088–2103 (2019).
Porpiglia, E. et al. Elevated CD47 is a hallmark of dysfunctional aged muscle stem cells that can be targeted to augment regeneration. Cell Stem Cell 29, 1653–1668 (2022).
Brett, J. O. et al. Exercise rejuvenates quiescent skeletal muscle stem cells in old mice through restoration of Cyclin D1. Nat. Metab. 2, 307–317 (2020).
McKellar, D. W. et al. Large-scale integration of single-cell transcriptomic data captures transitional progenitor states in mouse skeletal muscle regeneration. Commun. Biol. 4, 1280 (2021).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Dos Santos, M. et al. Opposing gene regulatory programs governing myofiber development and maturation revealed at single nucleus resolution. Nat. Commun. 14, 4333 (2023).
Mitchell, T. S., Bradley, J., Robinson, G. S., Shima, D. T. & Ng, Y. -S. RGS5 expression is a quantitative measure of pericyte coverage of blood vessels. Angiogenesis 11, 141–151 (2008).
Belk, J. A., Daniel, B. & Satpathy, A. T. Epigenetic regulation of T cell exhaustion. Nat. Immunol. 23, 848–860 (2022).
Hudson, W. H. et al. Proliferating transitory T cells with an effector-like transcriptional signature emerge from PD-1+ stem-like CD8+ T cells during chronic infection. Immunity 51, 1043–1058 (2019).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Lähnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Matjusaitis, M., Chin, G., Sarnoski, E. A. & Stolzing, A. Biomarkers to identify and isolate senescent cells. Ageing Res. Rev. 29, 1–12 (2016).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
Cherry, C. et al. Transfer learning in a biomaterial fibrosis model identifies in vivo senescence heterogeneity and contributions to vascularization and matrix production across species and diverse pathologies. Geroscience 45, 2559–2587 (2023).
Borcherding, N. et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun. Biol. 4, 122 (2021).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660 (2017).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Fridman, A. L. & Tainsky, M. A. Critical pathways in cellular senescence and immortalization revealed by gene expression profiling. Oncogene 27, 5975–5987 (2008).
Reactome. Senescence-associated secretory phenotype (SASP). https://www.reactome.org/content/detail/R-HSA-2559582
Moiseeva, V. et al. Senescence atlas reveals an aged-like inflamed niche that blunts muscle regeneration. Nature 613, 169–178 (2023).
Reyes, N. S. et al. Sentinel p16INK4a+ cells in the basement membrane form a reparative niche in the lung. Science 378, 192–201 (2022).
Henry, S. P. et al. Generation of aggrecan-CreERT2 knockin mice for inducible Cre activity in adult cartilage. Genesis 47, 805–814 (2009).
Liu, J.-Y. et al. Cells exhibiting strong p16INK4a promoter activation in vivo display features of senescence. Proc. Natl Acad. Sci. USA 116, 2603–2611 (2019).
Sessions, G. A. et al. Controlled induction and targeted elimination of p16INK4a-expressing chondrocytes in cartilage explant culture. FASEB J. 33, 12364–12373 (2019).
Lyons, G. E., Ontell, M., Cox, R., Sassoon, D. & Buckingham, M. The expression of myosin genes in developing skeletal muscle in the mouse embryo. J. Cell Biol. 111, 1465–1476 (1990).
Schiaffino, S. & Reggiani, C. Fiber types in mammalian skeletal muscles. Physiol. Rev. 91, 1447–1531 (2011).
Pette, D. & Staron, R. The molecular diversity of mammalian muscle fibers. Physiology 8, 153–157 (1993).
Pette, D. & Staron, R. S. Myosin isoforms, muscle fiber types and transitions. Microsc. Res. Tech. 50, 500–509 (2000).
Caiozzo, V. J. et al. Single-fiber myosin heavy chain polymorphism: how many patterns and what proportions? Am. J. Physiol. Regul. Integr. Comp. Physiol. 285, R570–R580 (2003).
Glaser, B. W., You, G., Zhang, M. & Medler, S. Relative proportions of hybrid fibres are unaffected by 6 weeks of running exercise in mouse skeletal muscles. Exp. Physiol. 95, 211–221 (2010).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Sciorati, C., Rigamonti, E., Manfredi, A. A. & Rovere-Querini, P. Cell death, clearance and immunity in the skeletal muscle. Cell Death Differ. 23, 927–937 (2016).
Cosgrove, B. D. et al. Rejuvenation of the muscle stem cell population restores strength to injured aged muscles. Nat. Med. 20, 255–264 (2014).
Roy, A. L. et al. A blueprint for characterizing senescence. Cell 183, 1143–1146 (2020).
Lee, P. J. et al. NIH SenNet Consortium to map senescent cells throughout the human lifespan to understand physiological health. Nat. Aging 2, 1090–1100 (2022).
Wang, Y. X. et al. A single cell spatial temporal atlas of skeletal muscle reveals cellular neighborhoods that orchestrate regeneration and become disrupted in aging. Preprint at bioRxiv https://doi.org/10.1101/2022.06.10.494732 (2022).
Young, L. V. et al. Muscle injury induces a transient senescence-like state that is required for myofiber growth during muscle regeneration. FASEB J. 36, e22587 (2022).
Kurland, J. V. et al. Aging disrupts gene expression timing during muscle regeneration. Stem Cell Rep. 18, 1325–1339 (2023).
10x Genomics: resolving biology to advance human health. https://www.10xgenomics.com/ (2020).
Illumina. Sequencing and array-based solutions for genetic research. https://www.illumina.com/ (2020).
Bengsch, B. et al. Epigenomic-guided mass cytometry profiling reveals disease-specific features of exhausted CD8 T cells. Immunity 48, 1029–1045 (2018).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 39, 313–319 (2021).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Virshup, I., Rybakov, S., Theis, F. J., Angerer, P. & Wolf, F. A. anndata: Access and store annotated data matrices. J. Open Source Softw. 9, 4371 (2024).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Mantri, M., Zhang, H. H., Spanos, E., Ren, Y. A. & De Vlaminck, I. A spatiotemporal molecular atlas of the ovulating mouse ovary. Proc. Natl Acad. Sci. USA 121, e2317418121 (2024).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. 40, 661–671 (2022).
Acknowledgements
We thank P. Schweitzer and colleagues in the Genomics Facility (Research Resource Identifier RRID: SCR_021727) of the Cornell Biotechnology Resource Center in the Cornell Institute for Biotechnology for their help in performing scRNA-seq and spatial transcriptomic experiments. We thank L. Tesfa and colleagues in the Flow Cytometry Facility (RRID: SCR_021740) of the Cornell Biotechnology Resource Center for their help in performing flow cytometry experiments. We thank the Cornell Center for Animal Resources and Education for assisting in animal housing and care. We thank A. De Micheli for helping with mouse procedures and in generating some scRNA-seq data. We thank A. Grimson and D. Iu for the advice on evaluating T cell exhaustion. We thank C. Cherry in the Elisseeff research group and other members of the JHU-Mayo-National Institutes of Health (NIH) SenNet team for advice on senescence scoring methods. We thank B. Diekman and his research group for generating and providing the chondrocyte gene signature. We thank D. McKellar for providing code via a GitHub repository and helpful advice. This work was supported by the US NIH grants R01AG058630 (to B.D.C. and I.D.V.), R01AR081449 (to B.D.C. and I.D.V.), R01AI176681 (to I.D.V.), U54AG079779 (to B.D.C. and J.H.E.), T32HD057854 (to L.D.W.), F30OD032097 (to V.I.M.), R01AI105265 (to B.D.R.) and DP1AR076959 (to J.H.E.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. We also acknowledge funding support from the Bloomberg~Kimmel Institute (to J.H.E.) and a Morton Goldberg Professorship (to J.H.E.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper.
Author information
Authors and Affiliations
Contributions
L.D.W., V.I.M., B.D.R., J.H.E. and B.D.C. designed the study. L.D.W., J.L.O., I.N., E.H.H.F. and V.I.M. carried out the experiments. L.D.W. and I.N. performed computational analyses. L.D.W., J.L.O. and I.N. analyzed experimental data and prepared figures. B.D.R., I.D.V., J.H.E. and B.D.C. supervised the data analyses. L.D.W., I.N. and B.D.C. wrote the paper. All authors provided feedback and comments.
Corresponding author
Ethics declarations
Competing interests
J.H.E. was previously a consultant and holds equity in Unity Biotechnology and Aegeria Soft Tissue and is an advisor for Tessera Therapeutics, HapInScience, Regenity and Font Bio. The other authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Evaluating sample quality.
(A-C) Quality control workflow and resulting UMAPs with cells colored by manually assigned cell type IDs based on the expression of hallmark skeletal muscle genes. Prior to any quality-control there were 365,710 cells and 24 cell types were identified, including 6 low-quality (LQ) clusters (A). After ambient RNA removal with SoupX, 23 cell types were identified, including 5 LQ clusters (B). After ambient RNA removal with SoupX and removal of LQ cells based on the number of genes and UMIs and the percent of mitochondrial reads, there were 286,273 cells and 24 cell types were identified, including two doublet clusters (C). (D) This is the same UMAP as in (B), but the cells are colored by quality status. Cells that had <200 genes, <750 UMIs, and >25% mitochondrial reads are considered LQ. All other cells are considered high-quality (HQ). (E) This is the same UMAP as in (C), but the cells are colored by doublet status as determined by DoubletFinder using an estimated doublet rate of 5%. (F) For every cell type cluster in (C), the fraction of singlets and doublets was calculated. (G-I) Violin plots of the number of genes (G), the number of UMIs (H), and the percent of mitochondrial reads (I) in each sample.
Extended Data Fig. 2 Identifying additional immune cell types when focusing on just the myeloid cells.
(A) Workflow to clarify the myeloid cell types. The 9 myeloid clusters based on broad cell type IDs were subset from the final dataset, re-clustered, and re-embedded. More specific cell type IDs were manually assigned using genes known to mark myeloid cell types. Some of these more specific cell type IDs from the myeloid subset were transferred back to the final dataset (Fig. 1f) based on the cell barcodes. (B-C) UMAPs of the myeloid subset after re-embedding, the cells are colored by the broad cell type IDs originally identified in the final dataset (B). After re-clustering, re-embedding, and re-annotating based on known immune cell markers, the cells are colored by the more specific cell type IDs (C). If the same cell type is in (B) and (C), it has the same color designation in both UMAPs. (D) Dot plot showing the expression frequency and magnitude of genes used to manually assign the more specific cell type IDs as shown in (C).
Extended Data Fig. 3 Identifying cell types in skeletal muscle during homeostasis and regeneration.
(A-C) Dot plots showing the expression frequency and magnitude of genes used to manually assign cell type IDs, including the more specific immune cell type IDs. We identified 17 immune cell clusters (A), 5 FAPs, Tenocytes, and Neural cell clusters (B), and 7 myogenic, Pericyte/Smooth muscle, and endothelial cell clusters (C).
Extended Data Fig. 4 Determining the cell type composition in each day post injury.
(A-B) Stacked bar plots of the percent of each cell type found at a given day post injury (dpi). The three endothelial populations were combined into “Endo (3 CTs)”, the three FAPs populations were combined into “FAPs (3 CTs)”, and the 17 immune populations were combined into “Immune cells (17 CTs)” (A). Only the 17 immune populations are combined into “Immune cells (17 CTs)” (this plot is the same as in Fig. 2k) (B). (C-D) Stacked bar plots of the percent of each immune cell type out of all immune cells at a given dpi. The four dendritic cell populations were combined into “Dendritic cells (4 CTs)” and the three T cell populations were combined into “T cells (3 CTs)” (C). All individual immune cell type IDs (D).
Extended Data Fig. 5 Cell type dynamics (those that are not in main Fig. 2).
(A-T) Line plots for the remaining cell types not shown in Fig. 2 showing cell type relative abundance as a fraction of total cells from 0-7 days post injury (dpi). For each sample, the number of cells of the reported type was divided by the total number of cells (excluding erythrocytes). Points are each sample (n = 3-4). Ribbon is the standard deviation. Statistical significance of age-specific cell type dynamics was evaluated using a One-Way ANOVA and the p-values were corrected with FDR (see Extended Data Fig. 6).
Extended Data Fig. 6 Cell type dynamics non-linear modeling.
(A-B) Plots of the fraction of MuSCs and progenitors from 0-7 days post injury (dpi) by age. For each sample, the number of cells was divided by the total number of cells (excluding erythrocytes). The points are the fraction for each sample (n = 3-4) and the ribbon is the confidence interval. The black line is the non-linear model independent of age (A) and the three colored lines are the non-linear model dependent on age (B). Whether there was a significant difference in the non-linear models independent of and dependent on age was determined using a One-Way ANOVA and the p-values were corrected with FDR. (C-D) Same as in (A-B) but with NK cells.
Extended Data Fig. 7 Identified two doublet clusters in the myogenic subset that were not identified by DoubletFinder.
(A-B) Scatter plots of the log-normalized expression levels of Pax7 and Pecam1 (A) and Acta1 and C1qa (B) by general myogenic cell type IDs as defined in Fig. 5a. A density curve is plotted on each axis.
Extended Data Fig. 8 Identifying a transitional myonuclei state.
(A-C) Same UMAP of the myogenic subset as in Fig. 5a colored by general myogenic cell type IDs (A) and colored by the log-normalized expression of Myh1 (B) and Myh4 (C). (D-F) Scatter plots of the log-normalized expression levels of Myh1 and Myh4 in the myonuclei clusters Type IIx (D), Type IIb (E), and Type IIx/IIb (F). The dotted-line boxes highlight the cells that express the marker(s) that should be expressed in each myonuclei cluster. (G) Stacked bar plot of the fraction of cells within each myonuclei cluster that express a combination of Myh1 and Myh4. (H) Violin plots of the log-normalized expression levels of markers of high metabolic rate (Tnnc2, Tnni2, Mb, Cox6a2, Cox6c, Atp5e, Atp5g1) and myonuclei (Ckm, Myh2, Myh1, Myh4, Chrne, Col22a1) in each myonuclei cluster. (I) Violin plot of the percent of mitochondrial reads in each myonuclei cluster.
Extended Data Fig. 9 Myogenic subset pseudotime analysis.
(A-H) Dot Plots of the average log-normalized expression of Pax7 (A), Myf5 (B), Myod1 (C), Myog (D), Mymk (E), Acta1 (F), Cdk1 (G), and Cdk4 (H) in each day post injury (dpi) and pseudotime bin. The size of the circle is the percent of cells in each pseudotime bin for each age and dpi combination. (I) Schematic of newly assigned myogenic IDs (referred to as ‘pseudotime-based myogenic cell state bin’) based on the expression of known myogenic markers in each dpi and pseudotime bin.
Extended Data Fig. 10 Cell cycle workflow.
(A-C) The polar coordinates in Fig. 5d were converted to cartesian coordinates (A) and rescaled to fit in a range from 0 to 1 (this plot is the same as in Fig. 5e) (B). The cells are colored by the predicted cell cycle phase based on the two scores (A-B) and G1 status (C). (D) Distribution of MuSCs and progenitors across the normalized theta values. The black line represents the standard Seurat G1 cutoff (normalized theta = 0.25), and the red line represents our extended G1 cutoff (normalized theta = 0.375). (E) Violin plot of the distribution of the normalized theta values in MuSCs and progenitors split by age and day post injury (dpi). (F) Violin plot of the distribution of the normalized theta values in MuSCs and progenitors split by pseudotime bin. (G-H) Scatter plot of the normalized theta values and the One-way FBR senescence score in MuSCs and progenitors. The cells are colored by log-normalized Cdkn2a expression (G) and by Cdkn1a expression (H). The vertical line is the extended G1 cutoff, and the horizontal line is where 50% of the cells above this line co-express Cdkn2a and Cdkn1a.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1 and 2 and Tables 1 and 2.
Supplementary Table 1 (download CSV )
Summary of scRNA-seq samples in this study.
Supplementary Table 2 (download XLSX )
Lists of senescence gene signatures used in this study.
Source data
Source Data Fig. 2 (download XLSX )
Data table for 2a–i,l,o.
Source Data Fig. 3 (download XLSX )
Data table for 3b–f.
Source Data Fig. 5 (download XLSX )
Data table for 5l.
Source Data Fig. 6 (download XLSX )
Data table for 6e–j.
Source Data Fig. 7 (download XLSX )
Data table for 7o.
Source Data Extended Data Fig. 5 (download XLSX )
Data table for 5a–t.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Walter, L.D., Orton, J.L., Ntekas, I. et al. Transcriptomic analysis of skeletal muscle regeneration across mouse lifespan identifies altered stem cell states. Nat Aging 4, 1862–1881 (2024). https://doi.org/10.1038/s43587-024-00756-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-024-00756-3
This article is cited by
-
VISTA uncovers missing gene expression and spatial-induced information for spatial transcriptomic data analysis
Communications Biology (2026)
-
Muscle stem cells in Duchenne muscular dystrophy exhibit molecular impairments and altered cell fate trajectories impacting regenerative capacity
Cell Death & Disease (2025)
-
Tenascin-C from the tissue microenvironment promotes muscle stem cell maintenance and function through Annexin A2
Communications Biology (2025)
-
Regulation and function of Pax3 in muscle stem cell heterogeneity and stress response
Journal of Muscle Research and Cell Motility (2025)
-
Advancing biological understanding of cellular senescence with computational multiomics
Nature Genetics (2025)