Abstract
Age-related hearing loss (ARHL) is among the most prevalent and complex disorders in older adults. However, the pathogenesis of ARHL remains poorly understood. Using a single-cell transcriptomic landscape of mouse cochlea at five time points (1, 2, 5, 12 and 15 months), we found that the levels of human antigen R (HuR)—a classical RNA-binding protein—increase with age. Here we show that HuR is specifically transported from the nucleus to the cytoplasm in hair cells in both aging mice and nonhuman primates. HuR overexpression in cochlea could successfully alleviate ARHL in aged mice. Meanwhile, HuR deficiency led to premature hearing dysfunction characterized by degeneration of stereocilia and the subsequent loss of hair cells. RNA immunoprecipitation sequencing analysis revealed that HuR can bind to messenger RNAs that enable stereocilia maintenance, including Gnai3. Adeno-associated virus-mediated Gnai3 overexpression partially rescues the hearing defects in HuR-deficient mice. Taken together, these findings indicate that HuR is a potential therapeutic target for ARHL.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The raw data of the RNA Immunoprecipitation sequencing analysis have been deposited in the Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra), under accession code PRJNA1221523. Raw data of RNA sequencing have been deposited in the Sequence Read Archive under accession codes PRJNA1221644. Source data are provided with this study. All other data are available from the corresponding authors upon request.
Change history
08 July 2025
A Correction to this paper has been published: https://doi.org/10.1038/s43587-025-00936-9
References
Bowl, M. R. & Dawson, S. J. Age-related hearing loss. Cold Spring Harb. Perspect. Med. 9, a033217 (2019).
Chadha, S., Kamenov, K. & Cieza, A. The world report on hearing, 2021. Bull. World Health Organ. 99, 242–242A (2021).
Blackwell, D., Lucas, J. & Clarke, T. Summary health statistics for U.S. adults: national health interview survey, 2012. Vital Health Stat. 10, 1–161 (2014).
Feng, L. et al. Associations between age-related hearing loss, cognitive decline, and depression in Chinese centenarians and oldest-old adults. Ther. Adv. Chronic Dis. 13, 204062232210848 (2022).
Lin, F. & Albert, M. Hearing loss and dementia—who is listening? Aging Ment. Health 18, 671–673 (2014).
Schuknecht, H. & Gacek, M. Cochlear pathology in presbycusis. Ann. Otol. Rhinol. Laryngol. 102, 1–16 (1993).
KK, O. Age-related hearing loss: the status of Schuknecht’s typology. Curr. Opin. Otolaryngol. Head Neck Surg. 12, 439–443 (2004).
Hu, S. et al. Age-related hearing loss and its potential drug candidates: a systematic review. Chin. Med. 18, 121 (2023).
Jing, Z. et al. Prestin is the motor protein of cochlear outer hair cells. Nature 405, 149–155 (2000).
Burns, J. C. & Corwin, J. T. A historical to present-day account of efforts to answer the question: ‘What puts the brakes on mammalian hair cell regeneration?’. Hear. Res. 297, 52–67 (2013).
Du, W. et al. Rescue of auditory function by a single administration of AAV-TMPRSS3 gene therapy in aged mice of human recessive deafness DFNB8. Mol. Ther. 31, 2796–2810 (2023).
Fu, X. et al. Tuberous sclerosis complex-mediated mTORC1 overactivation promotes age-related hearing loss. J. Clin. Invest. 128, 4938–4955 (2018).
Fu, X. et al. Activation of Rictor/mTORC2 signaling acts as a pivotal strategy to protect against sensorineural hearing loss. Proc. Natl Acad. Sci. USA 119, e2107357119 (2022).
Zhang, Q. et al. Nuclear speckle specific hnRNP D-like prevents age- and AD-related cognitive decline by modulating RNA splicing. Mol. Neurodegener. 16, 66 (2021).
Kim, G. et al. ALS genetics: gains, losses, and implications for future therapies. Neuron 108, 822–842 (2020).
Rhine, K. et al. ALS/FTLD-linked mutations in FUS glycine residues cause accelerated gelation and reduced interactions with wild-type FUS. Mol. Cell 80, 666–681 (2020).
Wurth, L. Versatility of RNA-Binding proteins in cancer. Comp. Funct. Genomics 2012, 178525 (2012).
Govindaraju, S. & Lee, B. S. Adaptive and maladaptive expression of the mRNA regulatory protein HuR. World J. Biol. Chem. 4, 111–118 (2013).
Schultz, C. W. et al. Understanding and targeting the disease-related RNAbinding protein human antigen R (HuR). Interdiscip. Rev. RNA 11, e1581 (2020).
Peng, W. et al. Elevated HuR in pancreas promotes a pancreatitis-like inflammatory microenvironment that facilitates tumor development. Mol. Cell. Biol. 38, e00427–17 (2018).
Atasoy, U. et al. Regulation of eotaxin gene expression by TNF-alpha and IL-4 through mRNA stabilization: involvement of the RNA-binding protein HuR. J. Immunol. 171, 4369–4378 (2003).
M, S. et al. Effects of infliximab therapy on gene expression levels of tumor necrosis factor alpha, tristetraprolin, T cell intracellular antigen 1, and Hu antigen R in patients with rheumatoid arthritis. Arthritis Rheum. 56, 2160–2169 (2007).
Fu, X., Zhai, S. & Yuan, J. Endothelial HuR deletion reduces the expression of proatherogenic molecules and attenuates atherosclerosis. Int. Immunopharmacol. 65, 248–255 (2018).
Green, L. C. et al. Human antigen R as a therapeutic target in pathological cardiac hypertrophy. JCI Insight 4, e121541 (2019).
Ding, F. et al. circHIPK3 prevents cardiac senescence by acting as a scaffold to recruit ubiquitin ligase to degrade HuR. Theranostics 12, 7550–7566 (2022).
Huai, Y. et al. HuR-positive stress granules: potential targets for age-related osteoporosis. Aging Cell 23, e14053 (2024).
Sun, G. et al. Single-cell transcriptomic atlas of mouse cochlear aging. Protein Cell 14, 180–201 (2022).
Brennan, C. M. & Steitz, J. A. HuR and mRNA stability. Cell. Mol. Life Sci. 58, 266–277 (2001).
Marie, A. et al. Senescence-accelerated mouse prone 8 (SAMP8) as a model of age-related hearing loss. Neurosci. Lett. 656, 138–143 (2017).
Menardo, J. et al. Oxidative stress, inflammation, and autophagic stress as the key mechanisms of premature age-related hearing loss in SAMP8 mouse Cochlea. Antioxid. Redox Signal 16, 263–274 (2012).
Pham, T. B. et al. Attenuation of age-related hearing impairment in senescence-accelerated mouse prone 8 (SAMP8) mice treated with fatty acid synthase inhibitor CMS121. J. Mol. Neurosci. 73, 307–315 (2023).
Ohlemiller, K. K., Dahl, A. R. & Gagnon, P. M. Divergent aging characteristics in CBA/J and CBA/CaJ mouse cochleae. J. Assoc. Res. Otolaryngol. 11, 605–623 (2010).
Sha, S. H. et al. Age-related auditory pathology in the CBA/J mouse. Hear. Res. 243, 87–94 (2008).
Dimri, G. P. et al. A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc. Natl Acad. Sci. USA 92, 9363–9367 (1995).
Lee, B. Y. et al. Senescence‐associated β‐galactosidase is lysosomal β‐galactosidase. Aging Cell 5, 187–195 (2006).
Chellappan, R. et al. SRI‐42127, a novel small molecule inhibitor of the RNA regulator HuR, potently attenuates glial activation in a model of lipopolysaccharide‐induced neuroinflammation. Glia 70, 155–172 (2022).
Sorge, R. E. et al. Inhibition of the RNA regulator HuR by SRI-42127 attenuates neuropathic pain after nerve injury through suppression of neuroinflammatory responses. Neurotherapeutics 19, 1649–1661 (2022).
Tan, F. et al. AAV-ie enables safe and efficient gene transfer to inner ear cells. Nat. Commun. 10, 3733 (2019).
Tao, Y. et al. AAV-ie-K558R mediated cochlear gene therapy and hair cell regeneration. Signal Transduct. Target Ther. 7, 109 (2022).
Zhao, C. et al. AAV-ie-mediated UCP2 overexpression accelerates inner hair cell loss during aging in vivo. Mol Med 28, 124 (2022).
Lu, Y. C. et al. Gene therapy with a synthetic adeno-associated viral vector improves audiovestibular phenotypes in Pjvk-mutant mice. JCI Insight 7, e152941 (2022).
Gu, X. et al. Transduction of adeno-associated virus vectors targeting hair cells and supporting cells in the neonatal mouse cochlea. Front. Cell Neurosci. 13, 8 (2019).
He, Z. X. et al. HuR in the medial prefrontal cortex is critical for stress-induced synaptic dysfunction and depressive-like symptoms in mice. Cereb. Cortex 29, 2737–2747 (2019).
Suhl, J. A. et al. A 3′ untranslated region variant in FMR1 eliminates neuronal activity-dependent translation of FMRP by disrupting binding of the RNA-binding protein HuR. Proc. Natl Acad. Sci. USA 112, E6553–E6561 (2015).
Han, X. et al. Loss of RNA-binding protein HuR leads to defective ependymal cells and hydrocephalus. J. Neurosci. 42, 202–219 (2022).
Liu, L. et al. HuR enhances early restitution of the intestinal epithelium by increasing Cdc42 translation. Mol. Cell. Biol. 37, e00574–00516 (2017).
Nishikawa, S. & Sasaki, F. Internalization of styryl dye FM1-43 in the hair cells of lateral line organs in Xenopus larvae. J. Histochem. Cytochem. 44, 733–741 (1996).
Gale, J. E. et al. FM1-43 dye behaves as a permeant blocker of the hair-cell mechanotransducer channel. J. Neurosci. 21, 7013–7025 (2001).
Makabe, A. et al. Systemic fluorescent gentamicin enters neonatal mouse hair cells predominantly through sensory mechanoelectrical transduction channels. J. Assoc. Res. Otolaryngol. 21, 137–149 (2020).
Du, T. T. et al. LMO7 deficiency reveals the significance of the cuticular plate for hearing function. Nat. Commun. 10, 1117 (2019).
Liu, Y. et al. Critical role of spectrin in hearing development and deafness. Sci. Adv. 5, eaav7803 (2019).
Gao, Y. et al. Impaired surface expression and conductance of the KCNQ4 channel lead to sensorineural hearing loss. J. Cell. Mol. Med. 17, 889–900 (2013).
Pyott, S. J. & Duncan, R. K. BK channels in the vertebrate inner ear. Int. Rev. Neurobiol. 128, 369–399 (2016).
Tilney, L. G., Tilney, M. S. & DeRosier, D. J. Actin filaments, stereocilia, and hair cells: how cells count and measure. Annu. Rev. Cell Biol. 8, 257–274 (1992).
Bartles, J. R. Parallel actin bundles and their multiple actin-bundling proteins. Curr. Opin. Cell Biol. 12, 72–78 (2000).
Katsuno, T. et al. TRIOBP-5 sculpts stereocilia rootlets and stiffens supporting cells enabling hearing. JCI Insight 4, e128561 (2019).
Krey, J. F. et al. ANKRD24 organizes TRIOBP to reinforce stereocilia insertion points. J. Cell Biol. 221, e202109134 (2022).
Prasad, S. et al. Radixin modulates the function of outer hair cell stereocilia. Commun. Biol. 3, 792 (2020).
Carlton, A. J. et al. Loss of Baiap2l2 destabilizes the transducing stereocilia of cochlear hair cells and leads to deafness. J. Physiol. 599, 1173–1198 (2021).
Bullen, A. et al. Ultrastructural defects in stereocilia and tectorial membrane in aging mouse and human cochleae. J. Neurosci. Res. 98, 1745–1763 (2020).
Jeng, J. Y. et al. Age-related changes in the biophysical and morphological characteristics of mouse cochlear outer hair cells. J. Physiol. 598, 3891–3910 (2020).
Furness, D. N. et al. Progressive hearing loss and gradual deterioration of sensory hair bundles in the ears of mice lacking the actin-binding protein Eps8L2. Proc. Natl Acad. Sci. USA 110, 13898–13903 (2013).
Yasuda, S. P. et al. c.753A>G genome editing of a Cdh23ahl allele delays age-related hearing loss and degeneration of cochlear hair cells in C57BL/6J mice. Hear. Res. 389, 107926 (2020).
Noben-Trauth, K., Zheng, Q. Y. & Johnson, K. R. Association of cadherin 23 with polygenic inheritance and genetic modification of sensorineural hearing loss. Nat. Genet. 35, 21–23 (2003).
Mendia, C. et al. Clarin-2 gene supplementation durably preserves hearing in a model of progressive hearing loss. Mol. Ther. 32, 800–817 (2024).
Mesgarzadeh, J. S., Buxbaum, J. N. & Wiseman, R. L. Stress-responsive regulation of extracellular proteostasis. J. Cell Biol. 221, e202112104 (2022).
Di Micco, R. et al. Cellular senescence in ageing: from mechanisms to therapeutic opportunities. Nat. Rev. Mol. Cell Biol. 22, 75–95 (2021).
Mukherjee, N. et al. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol. Cell 43, 327–339 (2011).
Fallmann, J. et al. AREsite2: an enhanced database for the comprehensive investigation of AU/GU/U-rich elements. Nucleic Acids Res. 44, D90–D95 (2016).
Beer Hammer, S. et al. Galphai proteins are indispensable for hearing. Cell. Physiol. Biochem. 47, 1509–1532 (2018).
Si, R. et al. HuR/Cx40 downregulation causes coronary microvascular dysfunction in type 2 diabetes. JCI Insight 6, e147982 (2021).
Beaudoin-Chabot, C. et al. The unfolded protein response reverses the effects of glucose on lifespan in chemically-sterilized C. elegans. Nat. Commun. 13, 5889 (2022).
Zhang, Z. et al. Hepatic HuR modulates lipid homeostasis in response to high-fat diet. Nat. Commun. 11, 3067 (2020).
Ezan, J. et al. Primary cilium migration depends on G-protein signalling control of subapical cytoskeleton. Nat. Cell Biol. 15, 1107–1115 (2013).
Krey, J. F. et al. Control of stereocilia length during development of hair bundles. PLoS Biol. 21, e3001964 (2023).
Manor, U. & Kachar, B. Dynamic length regulation of sensory stereocilia. Semin. Cell Dev. Biol. 19, 502–510 (2008).
Vogl, C. et al. The BEACH protein LRBA is required for hair bundle maintenance in cochlear hair cells and for hearing. EMBO Rep. 18, 2015–2029 (2017).
Behlouli, A. et al. EPS8, encoding an actin-binding protein of cochlear hair cell stereocilia, is a new causal gene for autosomal recessive profound deafness. Orphanet. J. Rare Dis. 17, 55 (2014).
Re, A. et al. RNA-protein interactions: an overview. Methods Mol. Biol. 1097, 491–521 (2014).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant nos. 82322019 to F.X.L., 82192863 to G.J.G., 82271175 to F.X.L., 82001204 to C.R.J., 82301320 to L.P.P., 82201294 to B.X.L., 82201296 to L.W., 82301305 to L.Z.Y., 82330033 to C.R.J., 82030029 to C.R.J. and 92149304 to C.R.J.). National Key R&D Program of China (grant nos. 2021YFA1101300 to F.X.L., 2021YFA1101800 to C.R.J. and 2020YFA0112503 to C.R.J.). The Natural Science Foundation from Shandong Province (grant nos. ZR2022QH338 to L.Z.Y., ZR2021QH269 to L.W. and ZR2022QH205 to B.X.L.). Natural Science Foundation of Jiangsu Province (grant nos. BK20232007 to C.R.J.), Science and Technology Department of Sichuan Province (grant nos. 2021YFS0371 to C.R.J.), Shenzhen Science and Technology Program (grant nos. JCYJ20190814093401920 to C.R.J. and JCYJ20210324125608022 to C.R.J.), 2022 Open Project Fund of Guangdong Academy of Medica Sciences (grant no. YKY-KF202201 to C.R.J.). We acknowledge the support of the Medical Science and Technology Innovation Center at Shandong First Medical University for valuable expertise and advice on the fluorescent imaging assays by confocal microscope.
Author information
Authors and Affiliations
Contributions
X.F., R.C. and J.G. designed and supervised the research. S.G. and X.F. wrote the paper. S.G., J.C., G.H., Y.S. and P.L. performed the experiments. M.X., W.Y., G.S., J.Q., Y.X., S.L. and S.C. contributed to the materials and analyzed the data. X.B., Z.L., Y.W., W.L. and X.Z. provided technical support for this research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Shintaro Iwasaki and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
https://www.ncbi.nlm.nih.gov/sra/PRJNA1221523
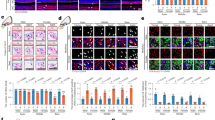
a, Representative fluorescence images of HuR in the SV of 1-month-old and 12-month-old mice. Scale bar, 10 μm. b, Representative fluorescence images of HuR in the SGN of 1-month-old and 12-month-old mice. Scale bars, 20 μm. c, Representative fluorescence images of HuR in the SCs of 1-month-old and 12-month-old mice. Scale bars, 10 μm. d, The fluorescence ratio of HuR signals in the cytoplasm and nucleus in Fig. 1k. The fluorescence intensity was quantified using Image J (NIH). n = 5 mice. e, Corresponding line intensity measurements for HuR protein and DAPI expression indicated that HuR is expressed only in the nucleus of HCs in young mice (1-month-old), while a portion of the HuR protein is shuttled to the cytoplasm of the HCs in aged mice (12-month-old). The fluorescence intensity was quantified using ImageJ (NIH). f, HuR immunostaining in the cochlea of young and old Macaca fascicularis. Scale bars, 20 μm. g, The fluorescence ratio of HuR signals in the cytoplasm and nucleus in (f). The fluorescence intensity was quantified using ImageJ (NIH). n = 5 Macaca fascicularis. h, Corresponding line intensity measurements for HuR protein and DAPI expression indicated that HuR is expressed only in the nucleus of OHCs in young Macaca fascicularis, while a portion of the HuR protein is shuttled to the cytoplasm of the OHCs in aged Macaca fascicularis. The fluorescence intensity was quantified using ImageJ (NIH). The data are presented as the mean ± standard error of the mean. The P-values were determined by a two-tailed t-test.
Extended Data Fig. 2 Inhibition of HuR shuttling to the cytoplasm in mouse models and successful overexpression of HuR in hair cells.
a, Western blot analysis and quantification of p53 and p21 expression (relative to GAPDH) in the cochlea of 1-month-old (1M) and 5-month-old (5M) SAMP8 mice (n = 3 mice). b, Real-time PCR analysis of p16, p21, and p53 mRNA expression (relative to β-actin) in the cochlea of 1-month-old (1M) and 5-month-old (5M) SAMP8 mice (n = 3 mice). c, The corresponding line intensity measurements show HuR protein and DAPI expression in the 1-month-old (1M), 2-month-old (2M), and 5-month-old (5M) SAMP8 mice. d, The fluorescence ratio of HuR signal between the cytoplasm and nucleus in 1-month-old (1M), 2-month-old (2M), and 5-month-old (5M) SAMP8 mice. The fluorescence intensity was quantified using ImageJ (NIH). n = 5 mice. e, HuR immunostaining in the cochlea of SAMP8 mice and SAMP8 mice injected at P50 with HuR translocation inhibitor SRI-42127. Scale bars, 10 μm. f, The corresponding line intensity measurements show HuR protein and DAPI expression in the control (CON) and SRI-42127-treated (SRI-42127) groups. g–h, The fluorescence ratio of HuR signal between the cytoplasm and nucleus in control and SRI-42127-treated groups. The fluorescence intensity was quantified using ImageJ (NIH). n = 5 mice. i, Comparison of transduction efficiency between AAV-ie-control and AAV-ie-HuR based on HA (green) and Phalloidin (red) immunostaining. Both AAV-ie-control and AAV-ie-HuR were found to efficiently transduce SCs and HCs throughout the cochlea. Scale bar, 50 μm. j,k, Histograms showing the percentage of HA-labeled SCs and HCs. n = 5 mice. The data are presented as the mean ± standard error of the mean. The P-values were determined by a two-tailed t-test in a, b, g, h, i and j, and a one-way ANOVA followed by Tukey’s multiple comparison test in d.
Extended Data Fig. 3 Lack of HuR leads to profound hering loss due to HC defects.
a, Representative H&E staining images of HCs, SGNs, and the SV in the middle turns of cochleae from WT and HuR-deficient (Atoh1-HuR−/− and Pax2-HuR−/−) mice. Scale bar, 50 μm. b,c, Graphs illustrating the average numbers of SGNs (b) and average thickness (c) of the SV in WT and HuR-deficient mice (Atoh1-HuR−/− and Pax2-HuR−/−). n = 5 mice. d, Whole-mount IF staining of the OC from P60 and P90 WT and HuR-deficient (Atoh1-HuR−/− and Pax2-HuR−/−) mice. Phalloidin (F-actin, red); Myo7A (HCs, green). Scale bar, 50 μm. n = 7 mice. e,f, Percentage of OHCs loss in P60 (e) and P90 (f) WT and HuR-deficient (Atoh1-HuR−/− and Pax2-HuR−/−) mice. n = 7 mice. g, TUNEL analysis of the OHCs in WT, Atoh1-HuR−/− and Pax2-HuR−/− cochlea at P60. Scale bar, 50 μm. h, The mean number of TUNEL-positive OHCs. n = 10 mice. The data are presented as the mean ± standard error of the mean. The P-values were determined by a one-way ANOVA followed by Tukey’s multiple comparison test.
Extended Data Fig. 4 HuR knockout in supporting cochlear cells does not affect hearing.
a, Schematic diagram of the transgenic HuR conditional knockout mice (Sox2-HuR−/−). b, IF staining for HuR expression in the SCs in P40 Sox2-HuR−/− mice. The white asterisks indicate supporting cells not labeled by the HuR antibody. Scale bar, 50 μm. c,d, Western blot analysis and quantification of HuR expression in the cochlea of Sox2-HuR−/− and WT mice. n = 3 mice. e, ABR thresholds of the WT and Sox2-HuR−/− mice at P60. n = 5 mice. The data are presented as the mean ± standard error of the mean. The P-values were determined by a two-tailed t-test.
Extended Data Fig. 5 Successful overexpression of HuR in HuR-deficient mouse models.
a, Comparison of Anc80L65-control and Anc80L65-HuR transduction efficiency based on HA (green) and Myo7 (red) immunostaining. Both Anc80L65-control and Anc80L65-HuR constructs were found to efficiently transduce HCs throughout the cochlea. Scale bar, 50 μm. b,c, Histograms showing the percentage of HA-labeled IHCs and OHCs (n = 8 mice for CON; n = 10 mice for Atoh1-HuR−/− and Pax2-HuR−/−). The data are presented as the mean ± standard error of the mean. The P-values were determined by a one-way ANOVA followed by Tukey’s multiple comparison test.
Extended Data Fig. 6 MET activity is affected in HuR mutant mice.
a, Representative images of cochlear ribbon synapses in WT and HuR-deficient mice (Atoh1-HuR−/− and Pax2-HuR−/−). The pre-synaptic (CtBP2, red) and post-synaptic (GluR2, green) markers are both immunolabeled. Scale bar, 20 μm. b, The number of ribbon synapses (CtBP2+ and GluR2+) per IHC. n = 5 mice. c, FM1-43 uptake in P60 cochlea derived from WT, Atoh1-HuR−/−, and Pax2-HuR−/− mice. HCs from WT mice displayed robust FM1-43 uptake, whereas those from both Atoh1-HuR−/− and Pax2-HuR−/− mice displayed weak FM1-43 uptake. Scale bar, 20 μm. d, FM1-43 signal intensity was measured using ImageJ. n = 5 mice, 2 images per mouse. Each value represents the average of all cells within each image. The shaded bands are presented as the mean ± standard error of the mean. e, Phalloidin-labeled OHC stereocilia in P60 WT, Atoh1-HuR−/−, and Pax2-HuR−/− mice. Scale bar, 20 μm. f, The number of HCs with abnormal stereocilia in P60 WT, Atoh1-HuR−/−, and Pax2-HuR−/− mice. n = 6 mice. g, Whole-mount IF staining for KCNQ4, BK, Prestin, and Spectrin in the OC at P60 in WT and HuR conditional knockout mice (Atoh1-HuR−/− and Pax2-HuR−/−). Scale bars, 20 μm. The data are presented as the mean ± standard error of the mean. The P-values were determined by a one-way ANOVA followed by Tukey’s multiple comparison test.
Extended Data Fig. 7 The degeneration of stereocilia in HuR-deficient mice accelerates the aging of OHCs.
a, Representative pseudo-colored scanning electron micrographs of OHC bundles from 1-month-old and 12-month-old mice. Close-up views illustrate the three full rows (tallest [magenta], middle [blue], and short [yellow]) of stereocilia in the OHC bundles. The orange arrow points to the fused stereocilia, the green arrow points to the broken stereocilia, and the magenta arrow points to internalized stereocilia. Scale bar, 2 μm. b, TEM micrographs reveal the normal morphology of P90 WT OHCs and severe degenerative changes in Atoh1-HuR−/− and Pax2-HuR−/− OHCs. In HuR-deficient mice, the number of morphologically abnormal mitochondria and lysosomes increased. The red arrows indicate abnormal mitochondria, the yellow arrows point to lysosomes. Scale bar, 10 μm. c, Representative SA-β-gal staining of the OC of the cochlea in WT and HuR-deficient mice at P80. Yellow arrows represent the SA-β-gal-positive cells. The dashed lines indicate the region of the OC where the HCs are located. Scale bar, 50 μm. d,e, Representative immunofluorescence images of p21 in the OC of WT and HuR-deficient mice at P80 and quantitative analysis (e). Scale bar, 20 μm, n = 6 mice. The white arrows represent the p21-positive cells. f,g, Representative immunofluorescence images of γ-H2A.X in the OC of WT and HuR-deficient mice at P80 and quantitative analysis (g). Scale bar, 20 μm, n = 6 mice. The white arrows represent the γ-H2A.X-positive cells. The data are presented as the mean ± standard error of the mean. The P-values were determined by a one-way ANOVA followed by Tukey’s multiple comparison test.
Extended Data Fig. 8 Successful overexpression of Gnai3 in HuR-deficient mouse models.
a, Comparison of Anc80L65-control and Anc80L65-Gnai3 transduction efficiency based on HA (green) and Phalloidin (red) staining. Both Anc80L65-control and Anc80L65-Gnai3 were found to efficiently transduce HCs throughout the cochlea. Scale bar, 50 μm. b,c, Histograms showing the percentage of HA-labeled OHCs and IHCs. n = 8 mice. The data are presented as the mean ± standard error of the mean. The P-values were determined by a one-way ANOVA followed by Tukey’s multiple comparison test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2 and Tables 3 and 4.
Supplementary Data 1 (download XLS )
Source data for Supplementary Fig. 1.
Source data
Source Data Fig. 1 (download XLS )
Statistical source data.
Source Data Fig. 2 (download XLS )
Statistical source data.
Source Data Fig. 3 (download XLS )
Statistical source data.
Source Data Fig. 4 (download XLS )
Statistical source data.
Source Data Fig. 5 (download XLS )
Statistical source data.
Source Data Fig. 6 (download XLS )
Statistical source data.
Source Data Fig. 7 (download XLS )
Statistical source data.
Source Data Fig. 8 (download XLS )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Fig. 7 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 1 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLS )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, S., Cao, J., Hong, G. et al. mRNA metabolism regulator human antigen R (HuR) regulates age-related hearing loss in aged mice. Nat Aging 5, 848–867 (2025). https://doi.org/10.1038/s43587-025-00860-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-00860-y