Abstract
Gestational diabetes mellitus (GDM), a common pregnancy disease, has long-term negative effects on offspring health. Epigenetic changes may have important contributions to that, but the underlying mechanisms are not well understood. Here, we report the influence of GDM on DNA methylation of offspring (GDF1) oocytes and the possible mechanisms. Our results show that GDM induces genomic hyper-methylation of offspring oocytes, and at least a part of the altered methylation is inherited by F2 oocytes, which may be a reason for the inheritance of metabolic disorders. We further find that GDM exposure increases the expression of Ezh2 in oocytes. Ezh2 regulates DNA methylation via DNMT1, and Ezh2 knockdown reduces the genomic methylation level of GDF1 oocytes. These results suggest that GDM may induce oocyte genomic hyper-methylation of offspring via enhancing the Ezh2 expression recruiting more DNMT1 into nucleus.
Similar content being viewed by others
Introduction
Gestational diabetes mellitus (GDM), characterized as hyperglycemia during pregnancy, is one of the most common diseases of pregnancy and its prevalence has increased worldwide. The International Diabetes Federation (IDF) reported that 1 in 6 live births were complicated by GDM in 20191. The incidence of GDM widely varies from 2% to 50% depending on the diagnostic criteria and regions2. GDM is not only associated with pregnancy complications, including neurodevelopmental delay, increased perinatal mortality, and perinatal complications, but also causes congenital malformations3. The probability of fetal macrosomia for maternal GDM is 3-fold higher than normal women4. Although maternal GDM can be treated with insulin, the incidence of low-birth-weight infants is still higher compared with normal females5. Offspring of women with GDM have a higher risk of cardiac malformations6. Long-term outcomes of offspring of GDM mothers are also reported7. Offspring of mothers with diabetes during pregnancy have a higher risk of obesity8,9, impaired glucose tolerance, insulin resistance, and diabetes10,11. Similar results are also received in animal models12. Additionally, the deleterious influence of GDM on offspring would be transmitted to the subsequent generations13. Therefore, it’s crucial to elucidate how GDM affects offspring health.
Previous studies have shown that altered DNA methylation induced by GDM is observed in umbilical cord blood, placenta, fetoplacental endothelial cells, and offspring blood in humans14. GDM exposure also has negative effects on histone modification and miRNA expression in fetus and placentas15,16,17. Exposure to maternal GDM during pregnancy alters the expression of miR-15 and miR-15b, and DNA methylation of Txnip (thioredoxin-interacting protein gene) in skeletal muscles of adults18. Similar results are also reported in animal models19. Moreover, intrauterine hyperglycemia has negative effects on DNA methylation of F1 primordial germ cells (PGCs)20. These suggest that epigenetic changes may have vital contributions to offspring health of GDM mothers. However, the underlying mechanism are not well elucidated. Therefore, we investigated the effects of gestational hyperglycemia on genomic DNA methylation in offspring oocytes and the possible mechanisms using a mouse model.
Here, we show that GDM disrupts the DNA methylome in oocytes and the metabolism of offspring, which can be partly transmitted to the subsequent generations via females. We further find that GDM increases the expression of EZH2 in oocytes, which interacts with DNMT1 but not DNMT3A to increase the whole genome DNA methylation of oocytes. Our results suggest that the increased EZH2 induced by GDM recruits DNMT1 into nucleus leading to the hyper-methylation of oocytes.
Results
GDM has negative contributions to the metabolism of F1 and F2 offspring
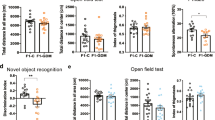
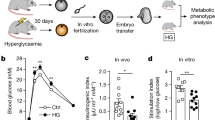
Previous studies have found that GDM exposure increases the risk of obesity, metabolic syndrome, type 2 diabetes, and impaired insulin sensitivity for offspring at a long-term21. We produced a GDM mouse model and bred F1 (GDF1) and F2 (GDF2) generations as shown in Fig. 1a. Results showed that GDM significantly reduced the average pups per female mouse compared with NGDF1(F1 offspring of non-gestational diabetic mice) (Fig. 1b). The average birth weight of female GDF1 was significantly lower compared with female NGDF1, but the difference of body weight between GDF1 and NGDF1 was not significant from 3 to 16 weeks of age (Fig. 1c). The average body weight of male GDF1 was similar to that of NGDF1 (Fig. 1d). Zhu et al. reported the glucose (GTT) and insulin (ITT) tolerance of GDF1 in C57/BLKS/J mouse (an inbred mouse being more glucose tolerant than C57BL/6 mouse22) were disturbed at 72 weeks of age, but not at 12 and 40 weeks of age23. Another study reveals that the GTT and ITT of GDF1 are altered at 8 weeks of age in ICR mouse (an outbred mouse)24. Here we examined the effects of GDM on GTT and ITT of GDF1 at 16 weeks of age. Compared with NGDF1, maternal GDM disturbed GTT and ITT of female GDF1 (n = 6 from 5 litters, Fig. 1e). For male GDF1, the GTT and ITT were similar to that of male NGDF1 (n ≥ 19 from at least 10 litters, Fig. 1f). These findings suggest that GDM disrupts the metabolism of female offspring, which is not completely consistent with previous studies22,24. The inconsistence between our results and previous studies may be caused by mouse species, age, sample size, and other factors.
a Schedule for breeding F1 and F2; b average pups per litter of F1; n: NGDF1 = 11, GDF1 = 26, p = 7.577*10−6; c, d body weight changes from birth to 16 weeks of age for female and male F1; n: NGDF1 = 32 and GDF1 = 21 for females, and NGDF1 = 20 and GDF1 = 23 for males; e, f glucose (GTT) and insulin (ITT) tolerance at 16 weeks of age for female and male F1, respectively; n: NGDF1 = 6 and GDF1 = 6; g average pups per litter of F2; n: CC = 11 and GC = 6, p = 0.039202; h, i body weight from birth to 16 weeks of age for female and male F2, respectively; n: CC = 32 and GC = 17 for females, and CC = 20 and GC = 10 to 18 for males (n: 16 weeks=10, 12 weeks=13, and the other age=18); j, k glucose and insulin tolerance for female and male F2 at 16 weeks of age; n: CC = 6 and GC = 6. data presented as mean ± SEM; F0, the parents; F1, the first generation; F2, the second generation; E1.5 d, embryonic day 1.5 day; GTT, glucose tolerance test; ITT, insulin tolerance test; GDF1, offspring of gestational diabetes; NGDF1, offspring of non-gestational diabetes; CC, offspring of NGDF1 female mated with normal male; GC, offspring of GDF1 female mated with normal male; *p < 0.05, **p < 0.01. Source data are provided as a Source Data file.
To further examine the inheritance of metabolic disorders, we bred F2 using female GDF1 mated with normal males (GC), and female NGDF1 were mated with normal males (CC) used as control (Fig. 1a). We found that the oocyte maturation rate (Supplementary Fig. 1A,B) and blastocyst rate in GDF1 were significantly reduced (Supplementary Fig. 1C). Moreover, GC had a lower average pups per litter compared with CC (Fig. 1g). The average body weight of female but not male GC was significantly higher than that of CC from birth to 16 weeks of age (n ≥ 15, Fig. 1h, i). The GTT and ITT of female but not male GC were also disturbed compared with CC (n = 6 from 5 litters, Fig. 1j, k). These results suggest that maternal GDM exposure has adverse effects on offspring metabolism, which may be transmitted to F2 generation via females.
GDM alters genomic methylation of F1 oocytes
We supposed that the above effect of GDM on offspring health was mediated by affected DNA methylation of GDF1 oocytes, and examined genomic methylation of GDF1 GV oocytes using immunofluorescence. Results showed that maternal GDM exposure significantly increased the genomic methylation level of 5mC (5-methylcytosine) in GV oocytes (Fig. 2a, b). We further examined the genomic 5mC and 5hmC (5-hydroxymethylcytosine) levels in metaphase II (MII) oocytes using chromosome spread and immunofluorescence, and found that the genomic 5mC and 5hmC levels in GDF1 were higher than that in NGF1 (Fig. 2c–e). The ratio of 5mC/5hmC in GDF1 was also significantly higher compared with NGDF1 (Fig. 2f). Milk of GDM mother may have influence on the methylation of offspring oocytes. Thus, we fostered GDF1 (G-F) and NGDF1 (NG-F) with normal mother after birth, and examined the DNA methylation level of oocytes. Results showed that methylation level of GV oocytes of G-F (n = 45) was still significantly higher than that of NG-F (n = 54, Supplementary Fig. 2). To further investigate the methylation in oocytes, we examined the genomic methylation using whole genome bisulfite sequencing for small samples (WGBS, BGI, China). Each sample contains 100 MII oocytes from 10 female mice ≥5 litters, and there were three replicates for each group. After the low-quality reads and adaptor sequence filtered, clean data were mapped to reference sequence mm10 using BSMAP. A total of 1009 million clean reads were obtained and the average clean reads Q20 rate was 95.28%. The mapping rate and bisulfite conversion rate were presented in Supplementary Table 1. The C coverage rate in chromosomes for each sample was presented in Supplementary Data file 1. There are three types of C according to the context, including CG, CHG, and CHH (H = A, T, or C), of which the number was presented in Supplementary Table 1. The average coverage rates of CG, CHG, and CHH in the whole genome were 54.63%, 51.57%, and 46.99%, respectively. Methylation was identified and calculated as previous studies25,26. The whole genome methylation levels were, respectively, 11.24% and 13.69% for NGDF1 and GDF1. In genomic elements, the methylation level was higher in transposable elements, and lower in 5’-UTRs, CDS, 3’-UTRs, and ncRNAs in GDF1 oocytes (Supplementary Fig. 3). GDM significantly increased the proportion of mCGs in three types of mCs (Fig. 2g). The genomic CG methylation level for each sample was presented in Fig. 2h, and the average genomic CG methylation in GDF1 oocytes was higher than that in NGDF1 (p = 0.013026, Fig. 2h). These results suggest that maternal GDM exposure increased the whole genome methylation of GDF1 oocytes.
a Genomic methylation of GV oocytes is examined using immunofluorescence, and b the relative fluorescence intensity of 5mC is examined using Image J, NGDF1, n = 33; GDF1, n = 24, p = 0.000322; c the methylation level of 5mC and 5hmC in MII oocytes is also examined using immunofluorescence, and d, e the relative fluorescence intensity is calculated using Image J;; n: NGDF1 = 47 and GDF1 = 41 for 5mC (p = 0.027547), and NGDF1 = 44 and GDF1 = 39 for 5hmC (p = 0.022106); f 5mC/5hmC ratio is calculated; n: NGDF1 = 37 and GDF1 = 35, p = 0.001764; g proportion of different mC types including CG (p = 0.03063), CHG (p = 0.34116), and CHH (p = 0.13043), H = A, T, or C. h genomic methylation of F1 MII oocytes is further examined using whole genome bisulfite sequencing, genomic methylation level of GC for each sample is presented (p = 0.013026). data presented as mean ± SEM. *p < 0.05, ***p < 0.001. Bar, 20 μm. 5mC, methylated cytosine; 5hmC, hydroxymethylcytosine; PI, prodium lodide; DAPI, 4’,6-diamidino-2-phenylindole; Source data are provided as a Source Data file.
Altered DNA methylation in GDF1 oocytes is partly transmitted to F2 offspring oocytes
DNA methylation level, especially at the promoter regions, often has important contributions to gene expression. Thus, we analyzed the differentially methylated regions (DMRs) of CG sites between GDF1 and NGDF1 oocytes via Metilene as described by a previous study27 and DSS, respectively. We identified, respectively, 10180 DMRs and 9784 DMRs (at CG number ≥5, absolute difference methylation ≥0.25 and the absolute log2 Ratio (GDF1/NGDF1) ≥ 1) via Metilene and DSS. The 10180 DMRs were used for further analysis because the DMR number was similar between two methods. In the present study, we identified 2211 genes with DMRs at promoter regions, including hyper-DMRs 16.69% and hypo-DMRs 83.31%. Gene ontology (GO) analysis showed that these genes were significantly enriched in cell process, metabolism process, and single-organism process (red box, Supplementary Fig. 4). KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis indicated that genes with DMRs at promoter regions were significantly enriched in insulin signaling pathway and type 2 diabetes mellitus (blue box, Fig. 3a). These results suggest that the altered methylation in oocytes might have crucial contributions to the metabolic disorders in offspring21,24,28.
a KEGG enrichment of genes with DMRs in promoter regions; b methylation level at CGIs, data presented as mean ± SD, p = 0.544, 0.4866, and <0.0001 for methylated CGIs, inter-methylated CGIs, and unmethylated CGIs, respectively; c, d methylation of CGI1 (p = 0.372704) and CGI2 (p = 0.323712) in F1 oocytes is examined using bisulfite sequencing; e–g methylation of Bcat1-DMR (p < 0.00001), Gng13-DMR (p < 0.00001), and Zfp867-DMR (p < 0.00001) in F1 oocytes is respectively examined using bisulfite sequencing; h–j methylation of Bcat1-DMR (p < 0.00001), Gng13-DMR (p < 0.00001), and Zfp867-DMR (p < 0.00001) in F2 oocytes is respectively examined using bisulfite sequencing. Methylation level is presented as percentage, and the statistical difference was analyzed using chi-square test. Black circle, methylated CG; white circle, unmethylated CG; CGIs, CpG islands; DMR, differentially methylated regions; Me, methylated CGIs; iMe, intermediate methylated CGIs; unMe, unmethylated CGIs; CC, female NGDF1 mated with normal male; GC, female GDF1 mated with normal male; ***p < 0.001.
CGIs (cytosine-phosphate-guanine islands) are commonly associated with promoter regions and subsequent initiation of gene expression. Usually, most of CGIs are unmethylated, in which altered DNA methylation level can lead to gene silencing. According to the methylation level, CGIs were divided into three types: methylated CGIs (methylation level (Me) ≥ 80%), intermediate CGIs (20% <methylation level (iMe) < 80%), and unmethylated CGIs (methylation level (unMe)≤ 20%). In the present study, methylation levels of CGIs were calculated at ≥20 CG sites covered. For methylated and intermediate CGIs, the methylation level was similar between GDF1 and NGDF1 oocytes, but the methylation level of unmethylated CGIs in GDF1 oocytes was significantly higher than that in NGDF1 oocytes (Fig. 3b). Furthermore, we identified 675 differentially methylated CGIs at the absolute methylation difference ≥0.25, including 673 unmethylated CGIs and 2 intermediate CGIs (Supplementary Data file 2 and Supplementary Fig. 5). These results suggest that DNA methylation at unmethylated CGIs is prone to be affected in GDF1 oocytes.
Our previous study indicates that a part of the disturbed DNA methylation in oocytes induced by uterine undernutrition can transmit to F2 oocytes via female29. Takahashi et al. showed that edited CGIs methylation of metabolism-related genes could be inherited by multiple generations30. Therefore, we analyzed the inheritance of the altered methylation in GDF1 oocytes via females using cloning bisulfite sequencing (BS). 3 DMRs (1 hypo-DMR and 2 hyper-DMRs) and 2 CGIs (CGI1 and CGI2) were included in this assay. The hypo-DMR locates at the promoter region of Bcat1 (branched-chain amino acid transaminase 1), and the hyper-DMRs respectively locate at the promoter regions of Gng13 (guanine nucleotide binding protein 13) and Zfp867 (zinc finger protein 867). We found that the methylation levels of CGI1 and CGI2 were similar between GDF1 and NGDF1 oocytes, Bcat1-DMR had a lower methylation level in GDF1 oocytes compared with NGDF1, and the methylation levels of Gng13-DMR and Zfp867-DMR were significantly higher in GDF1 oocytes than that in NGDF1 oocytes (Fig. 3c–g). To avoid the effects of clonal bias on the methylation levels of DMRs, we further examined the methylation level of DMRs via amplicon bisulfite sequencing (AmpBS). We found that the methylation of Bcat1-DMR was decreased and that of Gng13-DMR and Zfp867-DMR was increased in DGF1 oocytes (Supplementary Fig. 6), which was consistent with the BS results. To confirm the inheritance of altered methylation, we examined the methylation of DMRs in GC oocytes. Results showed that the methylation of Bcat1-DMR in GC oocytes was lower than that in CC, and the higher methylation of Gng13-DMR and Zfp867-DMR was maintained in GC oocytes (Fig. 3h–j). Bcat1 is mainly expressed in embryonic tissues, ovary, brain, placenta, and neurons, and has contribution to tricarboxylic acid (TCA) cycle, oxidative phosphorylation, and metabolic reprogramming in eukaryotic cells31. Gng13 is mainly expressed in brain and ovary32, and may be associated with lipid metabolism33. These results suggest that the disturbed methylation induced by GDM in GDF1 oocytes was, at least partly, transmitted to F2 oocytes via females, and the altered methylation might be a reason for the inheritance of metabolic disorders.
Analysis of the link between DMRs and differential expression genes (DEGs) in GDF1 oocytes
To investigate the effects of altered methylation on gene expression in oocytes, we examined the transcriptome using smart-seq (BGI, China). Each sample contained 100 MII oocytes from at least 10 female mice in ≥5 litters, and each group had three replicates. Clean data were mapped to reference genome mm10 using HISAT34. The correlation in samples (Supplementary Fig. 7A) and the distinguishment between groups were showed in Supplementary Fig. 7A-B. PC1 explained 42% of the variation and PC2 explained 21.7% of the variation. The transcriptional profiles in oocytes were distinguishable between GDF1 and NGDF1, although one sample of NGDF1 group was different with the other two replicates (Supplementary Fig. 7B). Therefore, this sample was not involved in the differentially expressed genes (DEGs) analysis. DEGs were identified using DEseq2 and PossionDis with absolute log2FC ≥ 1 and adjusted p value ≤ 0.05. We identified 1931 DEGs, including 697 up-regulated and 1234 down-regulated (Fig. 4a). In oocytes, gene transcription mainly occurs before de novo methylation of CGIs, and CGIs acting as transcription start sites (TSSs) have a low methylation level compared with non-TSSs-CGIs35. Here we analyzed the link between DEGs and DMRs in promoters, and found 192 DEGs (9.94%, Supplementary Data file 3) were with DMRs at the promoter regions including 52 up-regulated DEGs with hypo-DMRs in promoters, 25 down-regulated DEGs with hyper-DMRs, 98 down-regulated DEGs with hypo-DMRs, and 17 up-regulated DEGs with hyper-DMRs (Fig. 4b). Our results are consistent with previous reports that DNA methylation in promoters of active genes is mainly established after gene expression in oocytes. The DNA methylation levels of CpG sites in gene body regions are positively associated with open promoters and active transcription in mouse oocyte35,36. Thus, we analyzed the link between DMRs in gene bodies and DEGs in GDF1 oocytes. Results showed that 648 DEGs (33.56%) were with DMRs at CG sites in gene body regions (Supplementary Fig. 8). Two third of the methylated Cs are at non-CG sites in oocytes. Non-CG (CHG and CHH) methylation is absent from non-growing oocytes, and increases until fully grown oocyte stage where it is mostly found over active genes37. The non-CG methylation level is low in upstream and downstream, but that is high in gene bodies. These suggest that non-CG methylation might also have contributions to the regulation of gene expression. Thus, we analyzed the association between DEGs and DMRs of non-CG sites. We found that 1081 DEGs (55.98%) were with DMRs of CHG sites, and 1057 DEGs (54.74%) were with DMRs of CHH sites in gene bodies (Supplementary Fig. 8). Most classes of transposable elements (TEs) in mouse GV oocytes are methylated35, and non-CG DMRs locating in TEs may be involved in regulating gene transcription in human oocytes38. As showed in Supplementary Fig. 3, the DNA methylation in TEs was significantly increased in GDF1 oocytes. These results suggest that the changed DNA methylation levels in gene body regions, but not in promoters, might have more contributions to the disrupted transcriptome in GDF1 oocytes.
a differentially expressed genes (DEGs) analysis in F1 oocytes; b the association between DMRs located at promoter regions and DEGs; blue, downregulated DEGs; red, upregulated DEGs; c the mRNA expression of DNMT3a (p = 0.16794) and DNMT3l (p = 0.27292) in GDF1 oocytes; data presented as mean ± SD; d, e the mRNA expression of Ezh2 (p = 0.004) and Suz12 (p = 0.014) in GDF1 oocytes examined using qPCR; data presented as mean ± SD; f H3K27me3 modification in GV oocytes is examined using immunofluorescence and g the relative intensity of fluorescence is calculated using Image J; data presented as mean ± SD; NGDF1, n = 51; GDF1, n = 40; p = 0.01135; h the expression of EZH2 (n = 59) during folliculogenesis is examined using immunofluorescence histochemistry, and i the relative intensity fluorescence is calculated using Image J; data presented as mean ± SEM; n: primordial follicles = 28, primary follicles = 32, secondary follicles = 46, and antral follicles = 20 (p = 1.1527*10−8); 8 ovaries from 8 mice were used; j the H3K27me3 (n = 36) modification in follicular development is examined, and k the relative intensity of fluorescence was calculated by Image J; data presented as mean ± SEM; n: primordial follicles=27, primary follicles = 38 (p = 1.0952*10−12), secondary follicles = 33, and antral follicles = 19. *p < 0.05; **p < 0.01. Bar, 20 μm. FPKM, fragments Per Kilobase of exon model per Million mapped fragments. Source data are provided as a Source Data file. The statistical difference between groups was analyzed using two-tail t test.
EZH2 may have an important contribution to the disturbed genomic DNA methylation of GDF1 oocytes
During oogenesis, re-establishment process of DNA methylation is an extremely essential biological event, which is mainly catalyzed by DNA methyltransferases 3a/l (DNMT3a/l). However, the exposure of maternal GDM had no significant effects on the expression of DNMT3a/3 l in oocytes (Fig. 4c and Supplementary Table 2). To explore the possible key factors for the disturbed genomic methylation, we further analyzed the enrichment of DEGs using GO. We identified 31 DEGs in GDF1 oocytes that was involved in the methylation process, of which 7 DEGs were associated with the DNA methylation including upregulated genes Mis18a, Stella (also known as Dppa3 or Pgc7), Ezh2, and Suz12 and downregulated genes Tex15 (testis-expressed sequence 15 protein), Prmt5 (protein arginine methyltransferase 5), and Kdm1b (lysine demethylase 1B). EZH2 and SUZ12 are the core subunits of polycomb repressive complex 2 (PRC2) that catalyzes methylation (mono-, di- and tri-methylation) on histone 3 lysis 27 (H3K27)39, and EZH2, the catalytic subunit of PRC2, interacts with DNMTs to methylate CG in relative regions40. This indicates that the increased expression of Ezh2 may be vital for the hyper-methylation in GDF1 oocytes. To confirm this guess, we further examined the expression of Ezh2 and Suz12 in GDF1 oocytes using qPCR, and results showed that maternal GDM exposure significantly increased the mRNA expression of Ezh2 and Suz12 in oocytes (Fig. 4d, e). Therefore, we further examined the level of H3K27me3 level in GV oocytes, and found that it was also increased by GDM (Fig. 4f, g). These suggest that EZH2 might have an important contribution to the genomic hyper-methylation in GDF1 oocytes.
EZH2 regulates genomic DNA methylation by DNMT1 in oocytes
During oocyte maturation, EZH2 mainly locates in the nucleus and its expression gradually increases from GV to MII stages41. In the present study, we examined the dynamics of EZH2 during folliculogenesis using immunofluorescence. Results showed that EZH2 mainly located in the nucleus and its level was similar in primordial follicles, primary follicles, and secondary follicles, but it was increased in antral follicles (Fig. 4h, i). We further examined the dynamics of H3K27me3 in folliculogenesis, and found that the H3K27me3 level in primordial follicles was high, but it was reduced in primary follicles and maintained to antral follicles (Fig. 4j, k). These suggest that EZH2 may have contributions to methylation during oocyte growth and maturation. To investigate the role of EZH2 in oocyte genome methylation, we inhibited EZH2 function using inhibitors Dznep and GSK343 during oocyte maturation. Results showed that H3K27me3 was significantly reduced by inhibitors in MII oocytes (Fig. 5a, b). The 5mC and 5hmC levels were also decreased in MII oocytes by EZH2 inhibitors (Fig. 5c–e). The 5mC/5hmC ratios were also decreased by Dznep (Supplementary Fig. 9). Previous works have demonstrated that Abca1 and Dact3 are EZH2-target genes42,43,44. Therefore, we examined the methylation of these genes, and results showed that EZH2 inhibitors reduced the methylation levels of Abca1 and Dact3 (Fig. 5f, g). We further knocked down EZH2 level in oocytes using siRNA, and found that siRNA significantly reduced the expression of EZH2 in oocytes (Supplementary Fig. 10A), and genomic 5mC and 5hmC levels were also significantly decreased (Fig. 5h–j). EZH2 knockdown also decreased the 5mC/5hmC ratio in oocytes (Supplementary Fig. 10B). When Ezh2 was overexpressed in oocytes using mRNA injection (Supplementary Fig. 10C), the 5mC and 5hmC levels and 5mC/5hmC ratio in MII oocytes were significantly increased (Fig. 5k–m, Supplementary Fig. 10D). These results suggest that Ezh2 may play a key role in the DNA methylation of oocytes. We also performed Ezh2 knockdown at GV stage using siRNA and examined the transcriptome of MII oocytes using smart-seq. As expected, the transcriptome in MII oocytes was not distinguishable between the Ezh2 knockdown (siEZH2) and non-knockdown (cEZH2) groups (Supplementary Fig. 11A). We only identified 156 DEGs in the siEZH2 compared to cEZH2 (Supplementary Fig. 11B). GO analysis revealed that the DEGs were enriched in metabolic and translational pathways (Supplementary Fig. 11C). Two DEGs, Ezh2 and Setd4, were associated with the methylation pathway. The expression of DNMT1 and DNMT3A was not significantly affected by Ezh2 knockdown in MII oocytes (Supplementary Fig. 11D).
a, b inhibitors of EZH2 are used to inhibit the function of EZH2 in oocytes, and the H3K27me3 level is examined using immunofluorescence; control, n = 83; Dznep, n = 73 (p = 0.009185); Gsk343, n = 75 (p = 0.043357); c–e after inhibition of EZH2, the 5mC and 5hmC level in MII oocytes are examined using immunofluorescence; n: control=44, Dznep=56 (p = 0.007128), and Gsk343 = 36 (p = 0.021183) for 5mC, and control=42, Dznep=44 (p = 0.000235), and Gsk343 = 25 (p = 0.000834) for 5hmC; f, g Abca1 and Dact3 are target genes of EZH2, and the DNA methylation level of Abca1 (p = 0.000137 and 0.037005) and Dact3 (p = 0.017237 and 0.04808) is reduced by inhibiting the function of EZH2; data presented as percentage, and the statistical difference was examined using chi-square test; h–j when Ezh2 expression is knocked down in oocytes using siRNA, the genomic methylation level of 5mC and 5hmC is examined; n: control =27 and Ezh2-siRNA = 58 (p = 0.010829) for 5mC, and control=13 and Ezh2-siRNA =56 (p = 0.006307) for 5hmC; k–m with Ezh2 overexpression in oocytes, the 5mC and 5hmC level are examined using immunofluorescence; n: control =50 and Ezh2-overexpressionn=63 for both of 5mC (p = 1.31743*10−5) and 5hmC (p = 0.001759). *p < 0.05; **p < 0.01; ***p < 0.001; black circle, methylated CG; white circle, unmethylated CG. Bar, 20 μm. data presented as mean ± SEM. Source data are provided as a Source Data file.
EZH2 regulates DNA methylation by interacting with DNMTs40. Thus, we examined the localization of EZH2, DNMT1, and DNMT3A in oocytes. To avoid the effect of cytoplasm, we spread chromatin of GV oocytes on slides. Results showed that EZH2 colocalized with DNMT1 (Fig. 6a) and DNMT3a at chromatin (Fig. 6b). With Ezh2 expression knockdown, the level of DNMT1 and EZH2 localized on chromatin was significantly reduced in oocytes (Fig. 6c–e). Ezh2 overexpression increased the levels of DNMT1 and EZH2 on chromatin in oocytes (6f–h). Nevertheless, the level of DNMT3a in oocyte nucleus was not significantly affected by Ezh2 overexpression and knockdown, respectively (Fig. 6i–l). We further examined the interaction of EZH2 with DNMTs in oocytes using Co-IP. We firstly examined the interaction between EZH2 and DNMTs in cumulus-oocyte complex (COCs), and results showed that DNMT1 had interaction with EZH2, but not DNMT3A in COCs (Fig. 7a). The interaction of EZH2 with DNMTs in denuded GV oocytes was also examined, and the result showed that EZH2 interacted with DNMT1, but not DNMT3a (Fig. 7b). We further examined the interaction between EZH2 and DNMTs in ovaries using Co-IP, and results indicated that EZH2 interacted with DNMT1 but not DNMT3A in ovaries (Fig. 7c). To investigate the causality of EZH2 and DNMT1 in oocytes, here we examined the expression of DNMT1 in oocytes after Ezh2 or Suz12 knockdown. Results showed that Ezh2 or Suz12 knockdown decreased H3K27me3 (Fig. 7d–g, Supplementary Fig. 12A, B) and DNA methylation (Fig. 5h–j) level in oocytes, but the protein expression of DNMT1 was not significantly influenced (Supplementary Fig. 12C). Otherwise, DNMT1 knockdown in oocytes had no significant effects on DNA methylation, EZH2 expression and H3K27me3 level (Supplementary Fig. 13). These results suggest that EZH2 may be involved in regulating genomic DNA methylation during oocyte maturation via DNMT1 but not DNMT3A in oocytes.
a, b EZH2 colocalizes with DNMT1 and DNMT3a in nucleus of GV oocytes; c–e Ezh2 expression knockdown reduces the level of EZH2 and DNMT1 at chromatin; n: control=48 and Ezh2-siRNA=47 for EZH2 (p = 5.07772*10−8), and control=19 and Ezh2-siRNA=27 for DNMT1 (p = 2.62159 × 10−5); and f–h Ezh2 overexpression increases the level of EZH2 and DNMT1 at chromatin, n: control=57 and Ezh2-overexpression=45 for EZH2 (p = 4.54368 × 10−7), and control=26 and Ezh2-overexpression=22 for DNMT1 (p = 3.05373*10−7); but i–l the level of DNMT3a at chromatin is not affected by Ezh2 knockdown (n: control=25 and Ezh2-siRNA=20, p = 0.20404) and overexpression (n: control=31 and Ezh2-overexpression=22, p = 0.227746). data presented as mean ± SEM. ****p < 0.00001; ns, no significant difference. Bar, 20 μm. Ezh2-ov, Ezh2 overexpression; ns, no significant difference. Source data are provided as a Source Data file.
a, b the interaction of EZH2 with DNMT1 and DNMT3A and COCs (n = 2000) and oocytes (n = 5300) is examined using co-immunoprecipitation and Western-blotting; c the interaction between EZH2 and DNMT1 and DNMT3A in ovaries (n = 8) is examined using co-immunoprecipitation; d, e EZH2 is knockdown using siRNA in oocytes and the H3K27me3 level is examined using immunofluorescence (n: control=43 and Ezh2-siRNA=46, p = 0.002896); f, g SUZ12 knockdown in oocytes is performed using siRNA, and the H3K27me3 level is examined (n: control=43 and Ezh2-siRNA=48, p = 0.034347). *p < 0.05; **p < 0.01. Bar, 20 μm. H3K27me3, histone 3 lysine 27 trimethylation. data presented as mean ± SEM. Source data are provided as a Source Data file.
Increased Ezh2 expression may mediate the hyper-methylation of GDF1 oocytes via DNMT1
To investigate the role of EZH2 in hyper-methylation of GDF1 oocytes, we examined the dynamics of EZH2 in follicular development. EZH2 was located in nucleus (Fig. 8a), the relative level of EZH2 in PGCs, primary follicles, and secondary follicles was similar, but it was significantly increased in antral follicles (Fig. 7a, b, Supplementary Fig. 14). And the relative level of EZH2 was similar between NGDF1 and GDF1 from PGCs to antral follicles (Fig. 8b). The higher H3K27me3 level in PGCs of GDF1 was significantly reduced from primary follicles (Fig. 8c, d, Supplementary Fig. 14B). Compared with NGDF1, the H3K27me3 level in follicles of GDF1 were higher from primordial follicles to antral follicles (Fig. 8d). In GV oocytes, H3K27me3 level was significantly higher in GDF1 than that in NGDF1 (Fig. 4f, g). Meanwhile, the protein level of EZH2 in GDF1 oocytes was increased compared with that of NGDF1 (Fig. 8e). DNA methylation is mainly catalyzed by DNMT3A during oogenesis, but the mRNA (Fig. 4c) and protein expression and localization of DNMT3A were not significantly affected in GDF1 oocytes (Fig. 8e–g). Although the main function of DNMT1 is maintaining DNA methylation, its de novo methylation function in oocytes is also confirmed by previous studies37,45. However, the de novo methylation function of DNMT1 is usually prevented by STELLA (also known as DPPA3 or PGC7) in oocytes45. We analyzed the mRNA expression of Dnmt1, Stella, and Uhrf1 ((ubiquitin-like with PHD and RING finger domains 1) in GDF1 oocytes, and found that the mRNA expression of Dnmt1 and Uhrf1 were not significantly affected by GDM exposure, and the expression of Stella was increased (Fig. 8h, i, and Supplementary Table 2). But more DNMT1 was recruited into the nucleus in GDF1 oocyte (Fig. 8j, k). When knocking down the expression of Ezh2 in GDF1 oocytes using siRNA, the genomic hyper-methylation of 5mC and 5hmC and 5mC/5hmC ratio were significantly reduced (Fig. 8l–n, Supplementary Fig. 15). These results suggest that increased EZH2 may be involved in mediating the genomic hyper-methylation of GDF1 oocytes via recruiting more DNMT1 into nucleus.
a the dynamics of EZH2 in follicular development of GDF1 are examined using immunofluorescence, and b the relative level of EZH2 in GDF1 is compared with NGDF1; n: NGDF1 = 23 and GDF1 = 20 for primordial follicles (p = 0.088426), NGDF1 = 25 and GDF1 = 22 for primary follicles (p = 0.896508), NGDF1 = 43 and GDF1 = 47 for secondary follicles (p = 0.942229), and NGDF1 = 16 and GDF1 = 15 for antral follicles (p = 0.67194); 8 ovaries from 8 mice were used; c H3K27me3 level in folliculogenesis of GDF1 is examined, and d the relative level of it is compared with NGDF1; n: NGDF1 = 23 and GDF1 = 22 for primordial follicles (p = 0.003264), NGDF1 = 34 and GDF1 = 30 for primary follicles (p = 1.65546*10−5), NGDF1 = 30 and GDF1 = 25 for secondary follicles (p = 0.018598), and NGDF1 = 17 and GDF1 = 15 for antral follicles (p = 0.008278); 8 ovaries from 8 mice were used; e protein levels of DNMT3A and EZH2 in NGDF1 and GDF1 oocytes are examined using Western-blotting; f, g the level of DNMT3a in GDF1 oocytes is not affected by GDM compared with NGDF1 (n: NGDF1 = 19 and GDF1 = 18, p = 0.256458); h, i the mRNA expression of Dnmt1 (p = 0.655248), Uhrf1 (p = 0.898845) and Stella (p = 0.018583) in GDF1 oocytes (data presented as mean ± SD), but j, k the level of DNMT1 at chromatin is increased in GDF1 oocytes (n: NGDF1 = 14 and GDF1 = 15, p = 0.000605); l–n Ezh2 knockdown in GDF1 oocytes significantly decreased the level of 5mC and 5hmC (n: NGDF1 = 20, GDF1 = 30 (p = 0.041405), and Ezh2-siRNA=16 (p = 0.005935) for 5mC,, and NGDF1 = 23, GDF1 = 30 (p = 0.017439), and Ezh2-siRNA=16 (p = 0.034593) for 5hmC, p = ). *p < 0.05; **p < 0.01; ***p < 0.001; ns, no significant difference. Bar, 20 μm. data presented as mean ± SEM. Source data are provided as a Source Data file.
Discussion
GDM is one of the most common gestational diseases which have negative contributions to fetal development and long-term offspring health, and changes of epigenetic modifications are considered to be possible cause. However, the underlying mechanism are not well elucidated. In the present study, we found that GDM disturbed offspring metabolism that could be at least partly transmitted to F2 generation via female. The disturbed genomic DNA methylation in GDF1 oocytes may have contributions to the inheritance of metabolic disorders. DNMT1 may regulate genomic hyper-methylation of GDF1 oocytes via the increased expression of Ezh2.
Since Barker hypothesized the fetal origin of adult diseases in 1980s, researchers have demonstrated that adult diseases such as hypertension, and diabetes have close relation with fetal development environment. Gestational diabetes is an important pregnancy disease, and increases risk of multiple complications including prenatal mortality, macrosomia, preterm birth, neonatal hypoglycemia, and respiratory distress, and long-term health problems such as obesity, adiposity, hypertension, and nonalcoholic fatty liver diseases7,46. Impaired glucose tolerance in offspring induced by GDM can be transmitted to F2 generation47. Epigenetic changes are considered the main causes of fetal origin and inheritance of adult diseases. In humans, Awamleh et al. identified differentially methylated CG sites in cord blood and placenta48. GDM disturbs epigenome and transcriptome of placenta49. In rat, GDM results in hypomethylation and higher expression of cyclin-dependent kinase inhibitor 2 A (CDKN2A) and cyclin-dependent kinase inhibitor 2B (CDKN2B) in offspring islets50. Negative effects of GDM on genomic methylation of pancreas are also observed in mouse model19. So, the altered epigenetic modification may have contributions to metabolic diseases in offspring of GDM mothers51,52. The inheritance of changed DNA methylation is also reported by studies. Ding et al. report that altered methylation of Igf2/H19 in offspring islet induced by GDM exposure is also observed in F2 islet47. The edited DNA methylation at promoters of metabolism-relative genes can be inherited by multiple generations and induces metabolic disorders such as obesity30. In the present study, we find that GDM exposure disturbs the metabolism of offspring, a part of which is transmitted to F2 generation via female, and the altered DNA methylation in F1 oocytes is partly inherited by F2 oocytes. These suggest that the inheritance of metabolic disorders caused by GDM may be associated with changes of DNA methylation in oocytes, which affects the health of next generations.
In the present study, we found that GDM exposure disrupted the glucose and insulin tolerance of female F1 and F2, but not males. This contradiction between females and males may be caused by sexual dimorphism. Sexual dimorphism in insulin resistance, obesity, and metabolic syndromes is widely reported in previous studies53,54, which may be associated with hormones and gut microbiota55. Sexual dimorphism is also observed in the placentas and fetuses. The development of the female placenta is generally slower than that of male, and the males have heavier placentas in humans. Thus, the exposure of embryo or fetus to intrauterine perturbation often results in different disease susceptibility or progression. The male fetuses are at a greater risk when perturbation occurs in late gestation. However, female fetuses are more affected by maternal hypertension, preeclampsia, villous infarction, and preterm birth during pregnancy. Perturbations in early gestation also have more serious effects on females56,57.
EZH2 is the catalytic subunit of PRC2 which contributes to H3K37me1, me2, and me339. During oogenesis, the expression of Ezh2 is gradually increased and reaches to the highest in mature oocytes41,58. H3K27me3 is also widely established in MII oocytes and can be transmitted to offspring59,60. In early embryo development, maternal H3K27me3 is associated with DNA methylation61. Vire et al. have found that EZH2 recruits DNMTs to relative regions to methylate CG and regulates gene expression40. Overexpression Ezh2 altered DNA methylation in prostate cancer62. In the present study, we find that the expression of Ezh2 and Suz12 and genomic DNA methylation in GDF1 oocytes are significantly increased. While knockdown Ezh2 expression level in GDF1 oocytes using siRNA reduced hyper-methylation. These suggest that the increased Ezh2 expression in GFD1 oocytes may be associated with the hyper-methylation induced by GDM.
DNMT3A and 3 L are the main enzymes for de novo DNA methylation in oogenesis. Although DNMT1 is required for a part of methylation at CG sites in oocytes, it is usually prevented by STELLA. Lack Stella expression in oogenesis results in excessive genomic DNA methylation of oocytes by recruiting more DNMT1 into nucleus. UHRF1 is vital to recruit DNMT1 into nucleus to maintain DNA methylation in oocytes45. But the expression of DNMT3A, DNMT3L, DNMT1and UHRF1 in oocytes was not significantly influenced by GDM. EZH2 regulates DNA methylation by interacting with DNMTs40. GDM exposure significantly increases the expression of Ezh2 in oocytes. We also find that GDM increases the nuclear localization of DNMT1 and EZH2 in oocytes, but not DNMT3A. Co-IP results indicate that EZH2 interacts with DNMT1 but not DNMT3A in oocytes. These suggest that EZH2 may interact with DNMT1 to regulate the hyper-methylation in GDF1 oocytes.
Although our results suggest that disturbed DNA methylation in GDF1 oocytes may have contribution to the transgenerational inheritance of metabolic disorders, and EZH2 is involved in mediating the alteration of DNA methylation in GDF1 oocytes via DNMT1, the molecular mechanisms are not completely elucidated because this process is complex. It’s also important to explore other key factors which may be responsible for the disturbed metabolism in offspring. Therefore, more studies are necessary in the future to elucidate this question.
Methods
This study was performed according to the People’s Government of Shandong Province guidelines for the Care and Use of Laboratory Animals (no. [2018]311) and approved by the Ethics Committee of Qingdao Agricultural University (No. QAU2017050087; Qingdao, China).
Mice
Female and male ICR mice at 7-8 weeks of age were purchased from Jinan Pengyue Experimental Animal Breeding Co. Ltd (Jinan, China). All mice were raised in a temperature-controlled room (at 23 ± 2 °C) with a 12 h day/dark cycle and 50%-60% humidity, and given a normal diet and water.
Antibodies
Rabbit polyclonal anti-EZH2 antibody was purchased from Cell Signaling Technology; rabbit polyclonal anti-H3K27me3, mouse mAb to 5mC and rabbit mAb to 5hmC antibodies were purchased from Abcam; mouse mAb to DNMT1 and rat mAb to DNMT3A antibodies were purchased from Active Motif. Alexa Fluor 647 donkey anti-rabbit IgG were purchased from Abcam; FITC-conjugated donkey anti-rabbit IgG and Cy3-conjugated goat anti-mouse antibodies were purchased from Sangon (Shanghai, China).
Gestational diabetes model
80 female ICR mice at 7 weeks of age were divided into two groups, randomly. One group was used to produce gestational diabetes model. Female ICR mice were single injected with STZ at 200 mg / kg after fertilization. Four days later, blood glucose level was examined by tail-blood sample using ACCU-CHEK® Active. If glucose levels were higher than 17.0 mM/L, mice were recognized as diabetic mice. Female mice injected with buffer were used as control. Offspring of GDM and control were marked as NGDF1 and GDF1, respectively. F2 was bred using female F1: female NGDF1 mated with normal male (CC), female GDF1 mated with normal male (GC).
Oocyte collection and culture
After 48 h of intraperitoneal injection of 8IU pregnant mare serum gonadotropin (PMSG), GV (germinal vesical) oocytes were obtained by puncturing ovary follicles. Female mice were administrated with PMSG, and 48 h later, these mice were administrated with human chorionic gonadotropin (HCG). MII (metaphase II) oocytes were obtained from ampulla of Fallopian tube under stereomicroscope. The GV oocytes were cultured in vitro with 5 μM DZNep (MCE, China) or 1 μM GSK343 (MCE, China) for 14 h at 37 °C and 5% CO2. DZNep and GSK343 are inhibitors of EZH2.
Early embryo culture
Female offspring received intraperitoneal injection of 8IU PMSG and HCG as according to standard procedure. These mice were mated with normal males after 8IU HCG injection. If the vaginal plug was observed in the next morning, zygotes were collected and cultured in KSOM (Sigma) in vitro for 108 h.
Fluorescence immunohistochemistry of ovarian sections
Ovaries were collected from female offspring. Ovarian tissue was fixed overnight with 4% paraformaldehyde (4% PFA) at 4 °C. After that, samples were washed, dehydrated, made transparent, embedded in paraffin and sectioned. Paraffin sections were dewaxed with xylene, rehydrated with alcohol at various concentrations, and repaired with citric acid antigen repair solution for immunostaining. After blocking in BDT (30 mg BSA and 100ul goat serum dissolved in 900ul TBS) at room temperature for 30 min and then incubated overnight at 4 °C with primary antibodies: rabbit anti-EZH2 antibody (1:50), and rabbit anti-H3K27me3 antibody (1:100). Slides were rewarmed at 37 °C for 30 min. After three washing in TBS containing 0.1% Tween 20 (TBST), slides were incubated with secondary antibody (1:200) for 45 min at 37 °C, and then washed three times with TBST. Chromatin was stained with DAPI and then examined using a laser scanning confocal microscope (Leica TCS SP5 II, Wentzler, Germany). Relative fluorescence intensity was examined using Image J.
Ezh2, Suz12 and Dnmt1 knockdown
Ezh2 antisense oligonucleotide sequence (5-ATTTCTTCCCAGTCTGGCCCATGAT-3), Dnmt1 antisense oligonucleotide sequence (5- TTGGTGAGTTGATCTTCGG-3) and Suz12 antisense oligonucleotide sequence (5- AAGCTGTTACCAAGCTCCGTG-3) were transcribed into siRNA using T7 RNAi Transcription Kit (Vazyme Biotech, Nanjing, China). siRNA was injected into GV oocytes using micromanipulation. GV oocytes injected with buffer were used as control. And then, GV oocytes were blocked with 2.5uM milinone for 24 h. After blocking, GV oocytes were washed using M16 medium for 3 times, and incubated in drug-free M16 medium for 14 h. Mature oocytes were used for further experiments.
For smart-seq, MII oocytes were collected after the knockdown of Ezh2 using siRNA. Oocytes were injected with buffer were used as control. Each sample contained 20 MII oocytes, and each group had three replicates.
Ezh2 overexpression
CDS (coding sequence) sequence of Ezh2 with T7 promoter was amplified using PCR and ligated with EGFP sequence (T7-Ezh2-EGFP). Ezh2-EGFP mRNA was synthesized using mMESSAGE mMACHINE™ kit (Thermo Fisher). For overexpression, Ezh2-EGFP mRNA was injected into the cytoplasm of GV oocytes using micromanipulation. GV oocytes injected with buffer were used as control. Injected oocytes were blocked at the GV stage in M16 medium containing 2.5uM milinone for 10 h and then cultured in M16 medium for 14 h. Mature oocytes were used for further experiments.
Real-time quantitative PCR analysis (qPCR)
Total RNA was extracted from oocytes using the RNAprep Pure micro Kit (Tiangen Biotech, Beijing, China). cDNA was generated with RT Master Mix (Vazyme Biotech, Nanjing, China) and used as template to examine the expression of genes. qPCR reaction system was: 2 μl cDNA, 10 μl 2× SYBR Green PCR master mixes (Vazyme Biotech), 7.2 μl RNase free water, and 0.4 μl 10uM primers. qPCR amplification conditions were as follows: 95 °C for 5 min, followed by 40 cycles of denaturing at 95 °C for 10 s and 60 °C for 30 s, and then dissolution curve was established at 95 °C for 15 s, 60 °C for 30 s, and 95 °C for 15 s. The relative expression was calculated according to the 2−△△Ct method. Ppia and Gapdh were used as reference gene. All primers were presented in Supplementary Table 3.
Chromosome spread
Zona pellucida (ZP) of MII oocyte was removed using Acidic Tyrode’s Solution (Sigma). After washing, oocytes without ZP were transferred into explosive solution, dried at room temperature for 2 h, and washed several times with PBS containing 10 mg/mL BSA (PBS/BSA). After that, oocytes were treated with 4 N HCL at room temperature for 10 min, neutralized with 100 mM Tris-HCL (pH8.5) for 10 min, washed several times with PBS/BSA containing 0.05% Tween 20, and then blocked with PBS/BSA at room temperature for 2 h. Then, slides were incubated with primary antibodies anti-5mC and anti-5hmC (1:200) at 4 °C for 24-48 h. After washing, slides were incubated with secondary antibodies (1:500) at room temperature for 1 h. Chromosomes were stained with DAPI and then examined using a laser scanning confocal microscope (Leica TCS SP5 II, Wentzler, Germany). Z-stack was scanned at an interval 2.52 μm. Images were reconstructed with maximum projection. Fluorescence intensity was examined using Image J, and the background fluorescence was subtracted.
Immunofluorescence
We fixed oocytes using 4% PFA for 30 min, and then the oocytes were permeabilizated with 0.5% Triton X-100 at room temperature for 20 min, blocked in PBS/BSA for 2 h and then incubated overnight at 4 °C with primary antibodies: rabbit anti-EZH2 antibody (1:500) and rabbit anti-H3K27me3 antibody (1:200). After washing using PBS/BSA containing 0.05% Tween 20, secondary antibody (1:500) was stained and washed using PBS/BSA containing 0.05% Tween 20. Then, oocytes were transferred into DAPI with mounting and examined using a laser scanning confocal microscope (Leica TCS SP5 II, Wentzler, Germany). Z-stack was scanned at an interval 2.52 μm. Fluorescence intensity was examined using Image J, and the background fluorescence was subtracted.
Bisulfite sequencing
Bisulfite treatment was done using EpiTect Fast LyseAll Bisulfite Kit (Qiagen) according to the manual. Each sample contained at least 50 oocytes. Bisulfite treated samples were used as template to amplify target fragments using nested PCR. PCR products were purified and cloned into T-vector for sequencing (Tsingke, China). Methylation status was analyzed using BiQ Analyzer, and at least 10 available clones were used to calculate the methylation level.
Amplicon bisulfite sequencing
Each sample contained 100 oocytes, which were treated with EZ DNA Methylation-Direct Kits (Zymo Research, D5044) according to the manufacturer’s instruction. DMRs were amplified via nested PCR. After that, the PCR productions were pooled together and ligated with adapters to make the sequencing library. The library quality was checked using Qubit2.0 Fluorometer, Agilent 2100 bioanalyzer, and qRT-PCR, respectively. Sequencing was performed via Illumina sequencing platform. Raw data were filtered using cutadapt software.
Localization of EZH2 and DNMTs on chromatin
Briefly, ZP of GV oocytes was removed using Acidic Tyrode’s Solution, and then washed and placed on a glass slide with explosive solution. Samples were dried at room temperature for 2 h. After that, samples were washed and blocked with PBS/BSA at room temperature for 2 h and incubated with primary antibodies anti-EZH2, anti-DNMT1, or anti-DNMT3A (1:500) overnight at 4 °C. And then samples were washed and incubated with secondary antibodies (1:500) at room temperature for 1 h. Chromatin was stained with DAPI and examined using a laser scanning confocal microscope (Leica TCS SP5 II, Wentzler, Germany). Relative fluorescence intensity was examined using Image J.
Co-immunoprecipitation (Co-IP)
A total of 5300 GV oocytes, 2000 COCs and 8 ovaries were respectively used for Co-IP experiments. All procedures followed the instructions of the Protein A/G Magnetic Co-IP Kit (Absin, China). Samples were washed with pre-cooled PBS twice, and then incubated in cooled lysis buffer on ice for 5 min. After centrifuge at 13000 g for 10 min, supernatant was used for further experiment. Samples were incubated with anti-EZH2 antibody overnight at 4 °C to form an antigen-antibody complex, which we incubated with the magnetic beads at 4 °C overnight. Then, we washed the beads with lysis/wash buffer for two times. After that, beads were incubated with 50 μl IP elution buffer for 10 min. The supernatant was collected and neutralized with neutralization buffer. Protein was analyzed using Western-blotting (WB).
Western-blotting (WB) analysis
Oocytes were collected in lysis buffer and then heated at 95 °C for 10 min. Each sample contained 500 oocytes. Lysed samples were separated with 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were washed three times using TBST (tris-saline-tween 20 buffer), and then blocked with 5% BSA diluted with TBST at 4 °C overnight. After that, the membrane was incubated with primary antibodies at room temperature for 3 h: rabbit anti-EZH2 antibody (1:1000), rabbit anti-DNMT3A antibody (1:500), mouse anti-DNMT1 antibody (1:500) and rabbit anti-GAPDH antibody (1:1000). After washing three times using TBST, secondary antibody was stained at room temperature for 2 h. Results were examined using SuperFemto ECL Chemiluminescence Kit (Vazyme, China).
Whole genome bisulfite sequencing (WGBS)
For each sample, 100 MII oocytes from 10 females were pooled together for WGBS. Each group contained three replicates. WGBS was performed by GBI. Oocytes were lysised and genomic DNA was bisulfite converted. After that, fragments were ligated with adaptors which was used to establish sequencing library. High quality libraries were sequenced by Illunina HiSeq 2000. Adaptor sequence, low quality reads and contamination were filtered from raw data. Clean data was mapped to reference genome mm10 by BSMAP. Differentially methylated regions (DMRs) were called respectively using Metilene with default parameters and DSS, respectively. Briefly, after a pre-segmentation step to exclude non-informative regions, the circularbinary segmentation was used to identify regions with significant differential methylation. The segmentational gorithmis applied recursively trying to identify a window with the maximum difference of the cumulative sum of the mean methylation difference, indicating a potential differentially methylated region (DMR). After additional filter steps were passed, potential DMRs were tested using a two-dimensional Kolmogorov-Smirnov-Test (KS-test). DMRs were finally tested through the Mann-Whitney-U test. GO (gene ontology) analysis was performed using GOseq. KEGG (Kyoto o Encyclopedia of Genes and Genomes) analysis was carried out using KOBAS online.
RNA sequencing (RNA-seq)
Smart-seq was used for RNA-seq. 100 MII oocytes from 10 females were pooled together. Sequencing library quality was examined using Agilent2100 Bioanalyzer and qPCR. High quality library was sequenced using Illunina HiSeq 2000. Low quality reads were filtered from raw data using SOAPnuke (v1.5.2, parameters: -l 15 -q 0.2 -n 0.05 -i), and clean data was mapped to reference genome mm10 using HISAT2 (Hierarchical Indexing for Spliced Alignment of Transcripts, v2.0.4, parameters: --phred64 --sensitive --no-discordant --no-mixed -I 1 -X 1000). Gene expression analysis was carried out using Bowtie2 (v2.2.5, parameters: -q --phred64 --sensitive --dpad 0 --gbar 99999999 --mp 1,1 --np 1 --score-min L,0,-0.1 -I 1 -X 1000 -- no-mixed --no-discordant -p 1 -k 200) and RSEM (v1.2.12, parameters: default). Pearson correlation coefficient between samples was calculated using cor function of R package. Principle component analysis (PCA) was performed using online tool (https://www.omicshare.com/tools/home/report/reportpca.html). Differentially expressed genes were identified using DESeq2 (parameters: foldchange ≥2.00 and adjusted p value ≤ 0.05) and PossionDis (parameters: foldchange ≥2.00 and FDR ≤ 0.001) according to previous studies. GO and KEGG analysis were performed using phyper function of R package, and the significant enrichment was judged as FDR ≤ 0.01.
Glucose and insulin tolerance test (GTT and ITT)
For GTT, 16 weeks old mice were intraperitoneally injected with glucose at 2 g/Kg body weight after 16 h fasting, and blood glucose level was examined at 0, 30, 60, 90, and 120 min after injection, respectively. For ITT, a single intraperitoneal injection of insulin at 10 IU/Kg body weight was administered after 4 h fasting, and blood glucose level was respectively examined at 0, 30, 60, 90, and 120 min after injection.
Statistical analysis
Average body weight, blood glucose level, pups per mouse, proportion of mC, and relative expression of genes were presented as mean ± SEM and statistical difference was examined using independent sample two-tail t-test. Relative intensity of fluorescence was presented as mean ± SEM and statistical difference was examined using unpaired two-tail t-test. Methylation level was presented as percentage and statistical difference was examined using Chi-square test. If P value < 0.05, the difference was recognized as significant. At least repeat three times for each experiment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. Sequencing data in this study have been deposited in the BIG Sub database63,64 under accession code CRA011637 or PRJCA017996. According to the requirement of the journal, sequencing data have also been deposited in NCBI with accession code PRJNA1197668.
References
Magliano, D. J. & Boyko, E. J. In IDF DIABETES ATLAS. (Brussels, 2021).
Hartling, L. et al. Screening and diagnosing gestational diabetes mellitus. Evid Rep Technol Assess (Full Rep), 1-327 (2012).
Ornoy, A., Becker, M., Weinstein-Fudim, L. & Ergaz, Z. Diabetes during pregnancy: a maternal disease complicating the course of pregnancy with long-term deleterious effects on the offspring. a clinical review. Int J. Mol. Sci. 22, 2965 (2021).
Kc, K., Shakya, S. & Zhang, H. Gestational diabetes mellitus and macrosomia: a literature review. Ann. Nutr. Metab. 66, 14–20 (2015).
Langer, O. et al. Glycemic control in gestational diabetes mellitus-how tight is tight enough: small for gestational age versus large for gestational age? Am. J. Obstet. Gynecol. 161, 646–653 (1989).
Billionnet, C. et al. Gestational diabetes and adverse perinatal outcomes from 716,152 births in France in 2012. Diabetologia 60, 636–644 (2017).
Bianco, M. E. & Josefson, J. L. Hyperglycemia during pregnancy and long-term offspring outcomes. Curr. Diab Rep. 19, 143 (2019).
Pettitt, D. J., Knowler, W. C., Bennett, P. H., Aleck, K. A. & Baird, H. R. Obesity in offspring of diabetic Pima Indian women despite normal birth weight. Diab. Care 10, 76–80 (1987).
Silverman, B. L. et al. Long-term prospective evaluation of offspring of diabetic mothers. Diabetes 40, 121–125 (1991).
Kong, L., Nilsson, I. A. K., Gissler, M. & Lavebratt, C. Associations of maternal diabetes and body mass index with offspring birth weight and prematurity. JAMA Pediatr. 173, 371–378 (2019).
Ornoy, A., Ratzon, N., Greenbaum, C., Wolf, A. & Dulitzky, M. School-age children born to diabetic mothers and to mothers with gestational diabetes exhibit a high rate of inattention and fine and gross motor impairment. J. Pediatr. Endocrinol. Metab. 14, 681–689 (2001).
De Sousa, R. A. L. Animal models of gestational diabetes: characteristics and consequences to the brain and behavior of the offspring. Metab. Brain Dis. 36, 199–204 (2021).
Chandrasekaran, S. & Neal-Perry, G. Long-term consequences of obesity on female fertility and the health of the offspring. Curr. Opin. Obstet. Gynecol. 29, 180–187 (2017).
Dluski, D. F., Wolinska, E., Skrzypczak, M. Epigenetic changes in gestational diabetes mellitus. Int. J. Mol. Sci. 22, (2021).
Hepp, P. et al. Histone H3 lysine 9 acetylation is downregulated in GDM Placentas and Calcitriol supplementation enhanced this effect. Int J. Mol. Sci. 19, 4061 (2018).
Strutz, J. et al. Gestational diabetes alters microRNA signatures in human feto-placental endothelial cells depending on fetal sex. Clin. Sci. (Lond.) 132, 2437–2449 (2018).
Chen, A. C. H. et al. Hyperglycemia impedes definitive endoderm differentiation of human embryonic stem cells by modulating histone methylation patterns. Cell Tissue Res 368, 563–578 (2017).
Houshmand-Oeregaard, A. et al. DNA methylation and gene expression of TXNIP in adult offspring of women with diabetes in pregnancy. PLoS One 12, e0187038 (2017).
Zhu, Z. et al. Gestational diabetes mellitus alters DNA methylation profiles in pancreas of the offspring mice. J. Diab. Complications 33, 15–22 (2019).
Ren, J. et al. Intrauterine hyperglycemia exposure results in intergenerational inheritance via DNA methylation reprogramming on F1 PGCs. Epigenetics Chromatin 11, 20 (2018).
Damm, P. et al. Gestational diabetes mellitus and long-term consequences for mother and offspring: a view from Denmark. Diabetologia 59, 1396–1399 (2016).
Anderson, A. A. et al. Pancreatic islet expression profiling in diabetes-prone C57BLKS/J mice reveals transcriptional differences contributed by DBA loci, including Plagl1 and Nnt. Pathogenetics 2, 1 (2009).
Zhu, Z. L. et al. Gestational diabetes mellitus alters DNA methylation profiles in pancreas of the offspring mice. J. Diab. Complicat 33, 15–22 (2019).
Zhu, H. et al. Insulin therapy for gestational diabetes mellitus does not fully protect offspring from diet-induced metabolic disorders. Diabetes 68, 696–708 (2019).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Xiang, H. et al. Single base-resolution methylome of the silkworm reveals a sparse epigenomic map. Nat. Biotechnol. 28, 516–520 (2010).
Juhling, F. et al. Metilene: fast and sensitive calling of differentially methylated regions from bisulfite sequencing data. Genome Res 26, 256–262 (2016).
Kim, C., Harrall, K. K., Glueck, D. H., Needham, B. L. & Dabelea, D. Gestational diabetes mellitus, epigenetic age and offspring metabolism. Diabet. Med 39, e14925 (2022).
Tang, S.-B. et al. Inheritance of perturbed methylation and metabolism caused by uterine malnutrition via oocytes. BMC Biol. 21, 43 (2023).
Takahashi, Y. et al. Transgenerational inheritance of acquired epigenetic signatures at CpG islands in mice. Cell 186, 715–731.e719 (2023).
Sweatt, A. J. et al. Branched-chain amino acid catabolism: unique segregation of pathway enzymes in organ systems and peripheral nerves. Am. J. Physiol. Endocrinol. Metab. 286, E64–E76 (2004).
Fujino, A., Pieretti-Vanmarcke, R., Wong, A., Donahoe, P. K. & Arango, N. A. Sexual dimorphism of G-protein subunit Gng13 expression in the cortical region of the developing mouse ovary. Dev. Dyn. 236, 1991–1996 (2007).
Yi, S. & Zhou, W. Tumorigenesis-related key genes in adolescents and young adults with HR(+)/HER2(-) breast cancer. Int J. Clin. Exp. Pathol. 13, 2701–2709 (2020).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Gahurova, L. et al. Transcription and chromatin determinants of de novo DNA methylation timing in oocytes. Epigenet Chromatin 10, 25 (2017).
Gu, C., Liu, S. L., Wu, Q. H., Zhang, L. & Guo, F. Integrative single-cell analysis of transcriptome, DNA methylome and chromatin accessibility in mouse oocytes. Cell Res 29, 110–123 (2019).
Shirane, K. et al. Mouse oocyte methylomes at base resolution reveal genome-wide accumulation of non-CpG methylation and role of DNA methyltransferases. PLoS Genet 9, e1003439 (2013).
Yu, B. et al. Single-cell analysis of transcriptome and DNA methylome in human oocyte maturation. Plos One 15, e0241698 (2020).
Qi, W. et al. An allosteric PRC2 inhibitor targeting the H3K27me3 binding pocket of EED. Nat. Chem. Biol. 13, 381–388 (2017).
Vire, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Qu, Y., Lu, D., Jiang, H., Chi, X. & Zhang, H. EZH2 is required for mouse oocyte meiotic maturation by interacting with and stabilizing spindle assembly checkpoint protein BubRI. Nucleic Acids Res 44, 7659–7672 (2016).
Lv, Y. C. et al. Histone methyltransferase enhancer of zeste homolog 2-mediated abca1 promoter dna methylation contributes to the progression of atherosclerosis. PLoS One 11, e0157265 (2016).
Meng, X. D. et al. Knockdown of GAS5 inhibits atherosclerosis progression via reducing ezh2-mediated abca1 transcription in ApoE(-/-) Mice. Mol. Ther. Nucleic Acids 19, 84–96 (2020).
Li, H. et al. ALDH1A1 is a novel EZH2 target gene in epithelial ovarian cancer identified by genome-wide approaches. Cancer Prev. Res (Philos.) 5, 484–491 (2012).
Li, Y. et al. Stella safeguards the oocyte methylome by preventing de novo methylation mediated by DNMT1. Nature 564, 136–140 (2018).
Sweeting, A., Wong, J., Murphy, H. R. & Ross, G. P. A clinical update on gestational diabetes mellitus. Endocr. Rev. 43, 763–793 (2022).
Ding, G. L. et al. Transgenerational glucose intolerance with Igf2/H19 epigenetic alterations in mouse islet induced by intrauterine hyperglycemia. Diabetes 61, 1133–1142 (2012).
Awamleh, Z. et al. Exposure to Gestational Diabetes Mellitus (GDM) alters DNA methylation in placenta and fetal cord blood. Diab. Res Clin. Pr. 174, 108690 (2021).
Chen, M., Yan, J., Han, Q., Luo, J. & Zhang, Q. Identification of hub-methylated differentially expressed genes in patients with gestational diabetes mellitus by multi-omic WGCNA basing epigenome-wide and transcriptome-wide profiling. J. Cell Biochem 121, 3173–3184 (2020).
Nazari, Z., Shahryari, A., Ghafari, S., Nabiuni, M. & Golalipour, M. J. In utero exposure to gestational diabetes alters dna methylation and gene expression of cdkn2a/b in langerhans islets of rat offspring. Cell J. 22, 203–211 (2020).
Antoun, E. et al. Maternal dysglycaemia, changes in the infant’s epigenome modified with a diet and physical activity intervention in pregnancy: Secondary analysis of a randomised control trial. PLoS Med 17, e1003229 (2020).
Weng, X. et al. Genome-wide DNA methylation profiling in infants born to gestational diabetes mellitus. Diab. Res Clin. Pr. 142, 10–18 (2018).
Ortiz-Huidobro, R. I. et al. Sexual dimorphism in the molecular mechanisms of insulin resistance during a critical developmental window in Wistar rats. Cell Commun. Signal 20, 154 (2022).
Palmer, B. F. & Clegg, D. J. The sexual dimorphism of obesity. Mol. Cell Endocrinol. 402, 113–119 (2015).
Santos-Marcos, J. A., Mora-Ortiz, M., Tena-Sempere, M., Lopez-Miranda, J. & Camargo, A. Interaction between gut microbiota and sex hormones and their relation to sexual dimorphism in metabolic diseases. Biol. Sex. Differ. 14, 4 (2023).
Kalisch-Smith, J. I., Simmons, D. G., Dickinson, H. & Moritz, K. M. Review: sexual dimorphism in the formation, function and adaptation of the placenta. Placenta 54, 10–16 (2017).
Yu, P., Chen, Y., Ge, C. & Wang, H. Sexual dimorphism in placental development and its contribution to health and diseases. Crit. Rev. Toxicol. 51, 555–570 (2021).
Meng, T. G. et al. PRC2 and EHMT1 regulate H3K27me2 and H3K27me3 establishment across the zygote genome. Nat. Commun. 11, 6354 (2020).
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016).
Inoue, A., Jiang, L., Lu, F., Suzuki, T. & Zhang, Y. Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature 547, 419–424 (2017).
Chen, Z. Y., Yin, Q. Z., Inoue, A., Zhang, C. X. & Zhang, Y. Allelic H3K27me3 to allelic DNA methylation switch maintains noncanonical imprinting in extraembryonic cells. Sci. Adv. 5, eaay7246 (2019).
Hoffmann, M. J. et al. Expression changes in EZH2, but not in BMI-1, SIRT1, DNMT1 or DNMT3B are associated with DNA methylation changes in prostate cancer. Cancer Biol. Ther. 6, 1403–1412 (2007).
Chen, T. et al. The genome sequence archive family: toward explosive data growth and diverse data types. Genomics Proteom. Bioinforma. 19, 578–583 (2021).
Members, C.-N. Partners, database resources of the national genomics data center, china national center for bioinformation in 2022. Nucleic Acids Res 50, D27–D38 (2022).
Acknowledgements
We acknowledge Yong Zhao (Institute of Animal Sciences, Chinese Academy of Agricultural Sciences), Qian Wang (People’s Hospital of Zhengzhou University), and Xiao-Qi Li (College of Life Science, Qingdao Agricultural University) for providing helps to use the instruments. We acknowledge the Central Laboratory Life Science Instrument Platform of College of Life Science and the Central Platform of Qingdao Agricultural University for proving the fluorescence microscope and the instrument for qPCR, and thank Tao Xu and Lin Song for their invaluable help to train the students how to use these instruments. We thank Ning Wang (College of Life Science, Qingdao Agricultural University) for her help in microinjection. This work is supported by the National Natural Science Foundation of China (31872312, 32470566) received by GZJ, the National Research & Development Program of China (2022YFC2703501) received by SQY, the Science and Technology Program of Guangzhou, China (202201020292) received by SQY, the Natural Science Foundation of Shandong Province (ZR2021ZD33) received by SQY, and the Breeding Plan of Shandong Provincial Qingchuang Research Team (Innovation Team of Farm Animal Cloning, 012-1622001) received by ZMH.
Author information
Authors and Affiliations
Contributions
G.H.-Y., T.S.B., L.L.J., L.J., Z.T.T., Y.L.J., X.K., S.X.F., C.S., and J.X.W. contributed to the design of the work, and the acquisition, analysis of the data; Y.S., Z.M.H., H.G.A., Z.L., S.W., and Z.C.L. contributed to the conception of the work; S. Q.Y. and G.Z.J. contributed to the design of the work, the analysis of data, and had drafted and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Atsuo Ogura, Jörn Walter and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, HY., Tang, SB., Li, LJ. et al. Gestational diabetes mellitus causes genome hyper-methylation of oocyte via increased EZH2. Nat Commun 16, 127 (2025). https://doi.org/10.1038/s41467-024-55499-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-55499-x
This article is cited by
-
Integrated metabolomic and proteomic analyses reveal global reprogramming of oocyte metabolism following AFB1 exposure
Cell Communication and Signaling (2026)
-
Intrauterine hyperglycemia impairs mouse primordial germ cell development and fertility by sex-specific epigenetic reprogramming interference
Cell Discovery (2025)