Abstract
Helper-component proteinase (HC-Pro), encoded by potyviruses, function as viral suppressors of RNA silencing (VSRs). Despite their conserved role, HC-Pros share approximately 40% similarity, implying potential differences in VSR efficiency, particularly in their ability to inhibit HEN1 methyltransferase activity. This study investigated the inhibitory potential of HC-Pros from different potyviruses in transgenic plants. P1/HC-Pro from turnip mosaic virus (P1/HC-ProTu) exhibited the most potent inhibition of HEN1, followed by P1/HC-Pro from zucchini yellow mosaic virus (P1/HC-ProZy), while P1/HC-Pro from tobacco etch virus (P1/HC-ProTe) showed the weakest inhibitory effect. These differential effectual effects corresponded to variations in unmethylated microRNAs (unMet-miRNAs) accumulation across the transgenic lines. Fluorescence resonance energy transfer (FRET) analysis indicated that HC-ProTu recruits HEN1 and ATG8a to HC-Pro bodies (H-bodies) and indirectly associates with AGO1, potentially influencing the assembly of the RNA-induced silencing complex (RISC) and leading to the accumulation of free-form miRNA duplexes. The ability of HC-ProTu to sequester HEN1 and AGO1 in H-bodies may, therefore, modulate miRNA loading. This observation aligns with the finding that P1/HC-ProTu plants harbored approximately 50% unMet-miRNAs and exhibited the lowest AGO1 levels, suggesting a positive correlation between HEN1 inhibition and autophagic degradation of AGO1. Interestingly, unMet-miRNAs are absent in the AGO1 of P1/HC-ProTu plants but reappeared in P1/HC-ProTu/hen1-8/heso1-1 plants, accompanied by signs of AGO1 recovery. These findings highlight the functional diversity of HC-Pro VSRs and provide new insights into their differential effects on miRNA methylation, RISC assembly, and the regulation of RNA silencing pathways.
Similar content being viewed by others
Introduction
Potyviruses, known for their diversity, are prevalent plant viruses1,2. The P1/helper-component protease (P1/HC-Pro) of potyvirus is the first viral suppressor of RNA silencing (VSR) discovered to inhibit RNA silencing mediated by microRNA (miRNA) and short-interfering RNA (siRNA)3,4,5,6,7. Previous studies resulted in the identification of HC-Pro as the primary suppressor, with the P1 protein enhancing the suppressive effect of HC-Pro8,9. RNA silencing is a critical viral defense mechanism in plants3, making VSRs essential for plant viruses to counteract this defense10. Mutations in VSRs can diminish virulence and impair the ability of viruses to inhibit RNA silencing11,12. Previous studies have shown that various P1/HC-Pros in potyviruses can suppress RNA silencing, leading to abnormal mature miRNA and miRNA* accumulation, thus preventing miRNAs from regulating target mRNAs10,11,12. Transgenic plants expressing different potyviral P1/HC-Pro genes exhibit similar serrated leaf phenotypes10,11,12. Analysis of the protein similarity between P1 and HC-Pro in different potyviruses revealed that P1 proteins share less than 20% sequence similarity, while HC-Pro proteins are more divergent at the N-terminus, with ~40% similarity9,13. This variation suggested that the RNA silencing suppression abilities of P1/HC-Pros from different potyviruses may differ.
In addition to RNA silencing, Hafrén et al. highlighted the role of NEIGHBOR OF BRCA1 (NBR1)-dependent autophagy in resisting turnip mosaic virus (TuMV)14, which prompts elevated expression of NBR1 and autophagy-related protein 8a (ATG8a) in Arabidopsis, leading to autophagic degradation of the HC-Pro from TuMV (HC-ProTu)14. Conversely, nbr1 mutant and autophagy mutants, such as atg5, and atg7, exhibit increased susceptibility to TuMV, consistent with autophagy contributing to an antiviral mechanism in plants14. The study also revealed that the TuMV proteins 6K2 and VPg inhibit autophagic HC-ProTu degradation, indicating the presence of a complex host-virus interaction14. Furthermore, Hafrén et al. noted that the HC-Pro from potato virus A (PVA) induced the formation of RNA granules, known as PVA-induced RNA granules (PGs)15, which we refer to as HC-Pro-bodies (H-bodies) for this study. Several studies have shown that PGs contain ATG8a, HC-Pro, argonaute 1 (AGO1), oilgouridylate-binding protein (UBP1), varicose (VCS), and eukaryotic initiation factor iso4E, corroborating a significant link between autophagy, RNA silencing suppression, and PGs14,15.
Autophagy, a vital and evolutionarily conserved cellular process, involves the formation of an autophagosome, a double-membrane enclosed vesicle that encapsulates and transports cellular components to the lysosome for degradation and recycling16. Autophagy-related (ATG) proteins, particularly ATG8a, which has been identified as a critical marker for autophagy and is instrumental in autophagosome formation, are essential for this process14,17. Notably, disruptions in the assembly of the RNA-induced silencing complex (RISC) can lead to the autophagic degradation of AGO118,19,20. Johnston et al. highlight that AGO1 requires the assistance of HSP90 for stabilization when AGO1 has not been loaded with miRNAs20. Without miRNA association, non-functional AGO1 undergoes autophagic degradation, a process facilitated by F-box and WD repeat domain-containing 2 (FWB2)18,19. This finding underscores that autophagy regulates AGO1, serving as a critical quality control mechanism to maintain cellular homeostasis and ensure the effective functioning of RNA silencing. Hu et al. demonstrated that Arabidopsis plants expressing P1/HC-ProTu (P1/HC-ProTu plants) exhibited lower endogenous AGO1 levels than wild-type Col-0 plants9. In contrast, transgenic plants expressing either P1/HC-Pro from zucchini yellow mosaic virus (P1/HC-ProZy) or P1/HC-Pro from tobacco etch virus (P1/HC-ProTe) did not exhibit decreased AGO1 levels in their respective P1/HC-ProZy and P1/HC-ProTe plants9. These data suggested that P1/HC-ProTu has a unique function compared with P1/HC-ProZy and P1/HC-ProTe, which might be useful for further probing RNA silencing mechanisms.
The methyltransferase HUA ENHANCER 1 (HEN1) plays a crucial role in stabilizing small RNAs through the process of 2′-O-methylation21,22. This methylation acts as a protective mechanism, safeguarding these small RNA molecules from premature degradation. However, once AGO1-bound miRNAs fulfill their functional roles and require metabolic degradation, the enzymes Small-RNA Degrading Nucleases (SDNs) step in to initiate the degradation process by first removing the methyl group from the miRNA23,24,25,26,27. Following demethylation, the miRNA undergoes uridylation by HEN1 SUPPRESSOR1 (HESO1), a terminal uridylyltransferase (TUTase)25. This modification marks the miRNA for subsequent degradation. Yu et al. demonstrated that in various VSR transgenic plants, including those expressing HC-ProTu, p19, and p21, unmethylated miRNAs (unMet-miRNAs) were detected21. Furthermore, Sanobar et al. demonstrated that the FRNK amino-acid motif of HC-ProTu can physically interact with HEN1 and inhibit HEN1 activity both in vivo and in vitro28. The FRNK motif is highly conserved in potyvirus HC-Pros and plays an important role in suppressing the miRNA pathway. The Arg182Lys mutant (FKNK) of HC-ProTu (HC-ProTu-K) compromises the ability to suppress HEN1 activity and RNA silencing11.
In light of recent findings, we propose four key questions: (1) In which cellular compartment does P1/HC-Pro accumulate miRNA and miRNA*? (2) How do RNA silencing components interact with HC-Pro when both are located inside H-bodies? (3) Since different potyvirus HC-Pros contain the FRNK motif, do they all have the same ability to inhibit HEN1? (4) Does P1/HC-ProTu cause the degradation of AGO1 through autophagy?
In this study, we demonstrated that three species of P1/HC-Pros can lead to the accumulation of miRNA and miRNA* in the cytoplasm. Notably, HC-ProTu can recruit AGO1, HEN1, and ATG8a into H-bodies, either indirectly or directly. A critical distinction of P1/HC-ProTu, as opposed to P1/HC-ProZy and P1/HC-ProTe, lies in its efficient inhibition of HEN1 activity, resulting in unMet-miRNAs accumulation. We also found a positive correlation between the accumulation of unMet-miRNAs and the extent of autophagic AGO1 degradation. These results indicate a more potent RNA silencing suppression ability in P1/HC-ProTu, highlighting the complex interplay between viral proteins and host cellular mechanisms.
Results
Consistent and variable molecular characteristics associated with RNA silencing suppression in different P1/HC-Pro variants
To investigate the role of P1/HC-Pro in modifying the regulatory miRNA pathway, different species or modified P1/HC-Pro genes were utilized (Fig. 1a). P1/HC-ProTu, P1/HC-ProZy, and P1/HC-ProTe plants express the wild-type VSR with an intact FRNK motif, whereas P1/HC-ProTu-K plants contain an HC-ProTu with a mutated FRNK motif (Fig. 1a)9,11. Moreover, two recombinant P1/HC-Pro genes, P1Zy/HC-ProTu and P1Tu/HC-ProZy, were also used for testing the suppressive abilities of various HC-Pros (Fig. 1a)9.
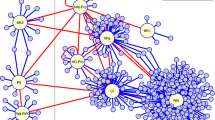
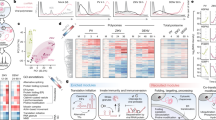
a Schematic binary plasmids containing the various P1/HC-Pro constructs used in this study. b Phenotypes of transgenic P1/HC-Pro plants. Photographs of 3-week-old plants were taken. A representative plant of each category is shown. Bar, 1 cm. c Abnormal miRNA/miRNA* accumulation in various P1/HC-Pro plants. U6 small nuclear RNA was used as a loading control. The numbers on the panels represent the average signal strengths of the miRNAs relative to those in the non-transformed wild-type plants (Col-0) after normalization against the loading control. The levels in the Col-0 plants were set at 1.0. Two independent lines for each construct were analyzed. NA, not available. d The SEC distributions of AGO1, HC-Pro, and HA-HEN1 in various plants. AGO1-L, the large form of AGO1. AGO1-S, the small form of AGO1 (n = 10). e The miR159 SEC distribution in various plants (n = 10). Syn-ds-miR159, the synthetic miR159 duplex (i). The miR165 and miR165* SEC distribution in P1/HC-ProTu plants (ii). f The methylation status of different SEC fractions in various plants. Fraction-mix I was mixed with fractions 4 and 6; fraction-mix II was mixed with fractions 10 and 12; fraction-mix III was mixed with fractions 14 and 16; and the free-form fraction was mixed with fractions 24 and 26. Two asterisks indicate an unknown additional band (n = 3). g Heatmap showing the expression levels of 103 miRNA target genes in Col-0 and various P1/HC-Pro plants. The color scale bar indicates the fold change in target gene expression. h Principal component analysis (PCA) for Col-0 and various P1/HC-Pro plants. i H-bodies observed in HC-ProTu-CFP (i) and HC-ProTu-K-CFP (ii). Evaluation of the subcellular colocalization between HC-ProZy-CFP (iii) and HC-ProTu-YFP (iv). A representative image of each category is shown. Bars, 10 μm. In, inclusion body.
As previously demonstrated, the P1/HC-ProTu, P1/HC-ProZy, and P1/HC-ProTe plants exhibited serrated and curled leaves, whereas the P1/HC-ProTu-K plants displayed a normal developmental phenotype, similar to that of the Col-0 wild-type (Fig. 1b). Abnormal miRNA and miRNA* accumulation is a common phenomenon in VSR transgenic plants11,12,29,30,31,32,33. We observed that miR159 and miR168 accumulated to 1.3- to 5.9-fold greater levels in P1/HC-ProTu, P1/HC-ProZy, and P1/HC-ProTe plants than in wild-type plants (Fig. 1c). MiR168* also significantly accumulated in all P1/HC-Pro plants (Fig. 1c). Notably, P1/HC-ProTu-K plants also exhibited slight miRNA accumulation (1.5 to 2.9-fold) (Fig. 1c), supporting the notion that HC-ProTu-K retains partial suppressive ability11.
To investigate the cellular distribution of abnormally accumulated miRNAs, crude plant extracts were subjected to size exclusion chromatography (SEC). In Col-0 plants, a large form of AGO1 (AGO1-L) was distributed in fractions 4–8, and we defined these fractions as high-molecular-weight RISC (HMW-RISC) (Fig. 1d). A small form of AGO1 (AGO1-S) was distributed in fractions 12–16, and we defined these fractions as low-molecular-weight RISC (LMW-RISC) (Fig. 1d). In P1/HC-ProTu plants, HC-ProTu was present in fractions 8–12 because HC-Pro forms a multimer (Fig. 1d)34,35. HA-HEN1 was distributed in fractions 6–18, consistent with the possible spatial and temporal localization of HA-HEN1 in the HC-ProTu (Fig. 1d).
Next, we extracted small RNAs from each SEC fraction for miRNA detection. The distribution of miR159 was in fractions 4 to 20, which included both HMW-RISC and LMW-RISC in Col-0, P1/HC-ProTu, and P1/HC-ProTu-K plants (Fig. 1e(i)). Surprisingly, in P1/HC-ProTu plants, we detected a high abundance of miR159 accumulation in fractions 22 to 28, which were defined as free-form fractions because synthesized double-stranded miR159 localizes to these fractions (Fig. 1e(i)). Interestingly, we also observed the free-form miR159 accumulation in P1/HC-ProTu-K and P1/HC-ProZy plants (Fig. 1e(i)). Moreover, observations also revealed a pronounced accumulation of both miR165 and miR165* within free-form fractions (Fig. 1e(ii)). These data imply that P1/HC-Pro impedes the loading of miRNA duplexes into RISC, causing them to accumulate in a free-form state, which is a phenomenon that is commonly characteristic of HC-Pro in potyviruses.
Given that HC-ProTu has been demonstrated to inhibit HEN1 methyltransferase activity, we proceeded to mix fractions from SEC samples for subsequent β-elimination analysis. In Col-0 plants, all fraction mixtures exclusively contained methylated miR159 (Met-miR159) (Fig. 1f). In contrast, both Met-miR159 (21-nt position) and unmethylated miR159 (unMet-miR159) (20-nt position) were found in all fraction mixtures from P1/HC-ProTu plants, with a notable enrichment in the free-form fraction (Fig. 1f), suggesting that unmethylated miRNAs (unMet-miRNAs) exist in the free-form and other fractions in P1/HC-ProTu plants. Additionally, a third band was observed between the 20-nt and 21-nt positions in fraction-mix III and the free-form fraction of P1/HC-ProTu plants. The molecular status of this intermediate miR159 species remains undetermined (Fig. 1f, asterisks).
According to the transcriptomic profiles, most of the miRNA targets in each of the four P1/HC-Pro plants were upregulated compared to those in the wild-type Col-0 plants (Fig. 1g). The upregulation of the target genes in the P1/HC-ProTu plants was more pronounced than that in the other three P1/HC-Pro plants (Fig. 1g), indicating that the ability of the P1/HC-ProTu plants to suppress RNA silencing differed from that of the other P1/HC-Pro plants. Notably, plants expressing P1/HC-ProTu-K with a mutated FRNK motif exhibited reduced silencing suppression capability compared to P1/HC-ProTu plants, yet these plants exhibited a suppressive effect akin to that observed in P1/HC-ProZy plants (Fig. 1g). This led to a moderate increase in the expression of target genes, corroborating prior research indicating that P1/HC-ProTu-K has a partial ability to suppress silencing11. Principal component analysis (PCA) revealed that the first two PCs contributed 80.4% of the total variance (Fig. 1h). The PC2 comprised 24.2% of the data variance could differentiate between P1/HC-Pro plants and Col-0 plants, whereas 56.2% of the data variance of PC1 separated P1/HC-ProTu plants and other plants (Fig. 1h). Therefore, compared with those of other P1/HC-Pro plants, the RNA silencing suppression ability of P1/HC-ProTu plants is unique.
Hafrén et al. demonstrated HC-Pro-induced PGs in the cytoplasm15. Thus, we evaluated round cytoplasmic H-bodies (1200–2000 nm in diameter) in HC-ProTu-CFP and HC-ProTu-K-CFP transient expression cells (Fig. 1i(i, ii)). HC-ProTu localizes to the nucleus and forms amorphous inclusion bodies (>6000 nm) in the cytoplasm (Supplementary Fig. 1), which is a natural property of HC-Pro34,35,36. Surprisingly, we did not observe HC-ProZy-CFP-forming H-bodies, even those co-expressing HC-ProTu-YFP (Fig. 1i(iii, iv)). However, HC-ProZy-CFP was detected in the nucleus and formed amorphous inclusion bodies in the cytoplasm (Fig. 1i(iii)). These data imply that various HC-Pros have conserved morphogenic and molecular phenotypes but also possess unique traits and diverse subcellular localization tendencies in plant cells.
HC-ProTu enhances AGO1 degradation and recruits AGO1 to H-bodies
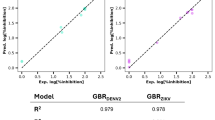
Endogenous AGO1 levels were quantified across multiple genotypes to assess the impact of different P1/HC-Pro variants. In P1/HC-ProTu and P1/HC-ProZy plants, AGO1 levels were significantly reduced to 0.27-fold and 0.55-fold, respectively, compared to those in Col-0 plants (Fig. 2a(i)). In contrast, AGO1 levels in P1/HC-ProTu-K (0.86-fold) and P1/HC-ProTe (0.87-fold) plants were comparable to that observed in Col-0 plants (Fig. 2a(i)). These observations were supported by quantifying five independent western blot experiments (Fig. 2a(ii); Supplementary Fig. 2). The relative expression levels of AGO1 for each genotype were statistically analyzed and depicted in a bar chart (Fig. 2a(ii)), confirming a significant reduction in AGO1 levels in P1/HC-ProTu plants, with a moderate decrease in P1/HC-ProZy plants (Fig. 2a(i, ii)). Further validation was performed using LC-MS/MS, reinforcing the western blot findings. The P1/HC-ProTu and P1/HC-ProZy plants exhibited reduced AGO1 levels, while the P1/HC-ProTe and Col-0 plants showed comparable AGO1 expression (Fig. 2a(iii)). In summary, the data indicate that different P1/HC-Pro variants distinctly modulate AGO1 expression levels. Notably, the P1/HC-ProTu/atg8age plants discussed in Fig. 2a are atg8a mutants derived from the P1/HC-ProTu background, with a detailed characterization of this mutant provided in subsequent sections.
a Endogenous AGO1 protein levels (AGO1-L and AGO1-S) in Col-0 and various P1/HC-Pro plants (i). Actin (ACT) was used as a loading control. The fold change values are indicated below the gel image. The levels in the Col-0 plants were set at 1.0. The average signal strengths of AGO1 relative to those in the non-transformed wild-type plants (Col-0) after normalization against the ACT loading control. Bar charts showed the relative amount of AGO1-L protein quantified from western blotting (WB) (ii) and from LC-MS/MS (iii). Data are presented as mean values ± standard errors (SE, n = 5 for WB, and n = 5 for LC-MS/MS). Significant differences based on Student’s t-test; *, **, and *** indicate P values < 0.05, <0.01, and 0.001, respectively. b AGO1 expression in Col-0, P1/HC-ProTu, and P1/HC-ProTu-K plants was evaluated by real-time RT-PCR. Relative expression levels were normalized to the ubiquitin level. Data are presented as mean values ± standard deviations (n = 3). Significant differences were determined by Student’s t-test; * indicates P values < 0.05. c The subcellular colocalization of HC-ProTu-CFP (i) or HC-ProTu-K-CFP (iii) with YFP-AGO1 (ii and iv). N, nucleus; In, inclusion body. Arrowheads indicate speckle-like structures. Bar, 50 μm. d FRET efficiency of co-infiltrated plants. Data are presented as mean values ± standard deviations (n = 10). e Image size measurement for speckle-like structures. Bar, 50 μm. (i) CFP channel for HC-ProTu-CFP. (ii) YFP channel for YFP-AGO1. Bar, 50 μm. f Size comparison of different foci (n = 15). Data are presented as mean values ± standard deviations. Source data are provided as a Source Data file.
Although the AGO1 protein levels were decreased in some of the P1/HC-Pro plants (Fig. 2a), the AGO1 mRNA levels increased in P1/HC-ProTu, P1/HC-ProZy, and P1/HC-ProTe plants (Fig. 2b), likely because the miR168-AGO1 regulatory module was suppressed. AGO1 mRNA levels were similar between P1/HC-ProTu-K and Col-0 plants (Fig. 2b).
We employed cell biology techniques to explore the impact of HC-Pro on AGO1. HC-ProTu-CFP formed numerous H-bodies within the cytoplasm, with YFP-AGO1 co-located in these speckles (Fig. 2c(i, ii)). In contrast, HC-ProTu-K-CFP resulted in the formation of only a limited number of H-bodies, with the majority of YFP-AGO1 being located in the cytoplasm and nucleus and only a minor fraction of YFP-AGO1 co-localizing in H-bodies (Fig. 2c(iii, iv); Supplementary Fig. 3). To determine whether HC-Pro physically interacts with AGO1, we utilized fluorescence resonance energy transfer (FRET). HC-Pro can form cytoplasmic multimers34,35; therefore, we chose HC-ProTu-CFP and HC-ProTu-YFP as positive controls (Fig. 2d). H-bodies composed of HC-ProTu-CFP and HC-ProTu-YFP exhibited 70% FRET efficiency, whereas the HC-ProTu-CFP + YFP sample (a negative control) exhibited only 5% FRET efficiency (Fig. 2d). HC-ProTu-K-CFP and HC-ProTu-K-YFP also demonstrated a 72% FRET efficiency, implying that HC-Pro is prone to forming dimers or multimers through self-interaction (Fig. 2d). Surprisingly, no FRET signals were detected in the H-bodies formed by YFP-AGO1 and either HC-ProTu-CFP or HC-ProTu-K-CFP (Fig. 2d), indicating that there was no physical interaction between AGO1 and either HC-ProTu or HC-ProTu-K. Moreover, the size of YFP-AGO1 foci co-expressed with HC-ProTu-CFP was ~1400–2400 nm (Fig. 2e, f), similar to that of H-bodies (Fig. 1i). In summary, HC-ProTu, but not HC-ProTu-K, attracts AGO1 to H-bodies by indirect interaction, perhaps triggering AGO1 degradation.
HC-ProTu triggers autophagic AGO1 degradation and physically interacts with ATG8a
Previous reports have indicated that AGO1 is degraded via autophagy18,19,22, and indeed, the YFP-AGO1 signal disappeared 3 days after co-infiltration with HC-ProTu-CFP, consistent with the triggering of AGO1 degradation in P1/HC-ProTu plants. A heatmap of 41 ATG genes revealed that some autophagy genes, such as ATG8a/b/c/g, ATG18b/c/d/e/f, ATG4a, ATG9, ATG12a, ATG101, and ATI3C, etc., were significantly upregulated in P1/HC-ProTu plants (Fig. 3a). We detected significant differential expression of the ATG8 family of genes, including ATG8a, ATG8b, ATG8g, and ATG8h, in these P1/HC-Pro plants (Fig. 3b), suggesting that upregulation of autophagy genes may be a common phenomenon under the suppressive effects of HC-Pro.
a Heatmaps of autophagy-related genes in various P1/HC-Pro plants were generated from transcriptome profiles. The color scale bar indicates the fold change in target gene expression. b Differential expression of ATG8 family genes in various P1/HC-Pro plants relative to their expression levels. Data are presented as mean values ± standard deviations (n = 3). Significant differences were determined by Student’s t-test; * indicates P values < 0.05. ** indicates P values < 0.01. *** indicates P values < 0.001. c Diagram of the gRNA positions in the ATG8a gene (i) and the gene-editing results (ii) for P1/HC-ProTu/atg8age. d ATG8a transcript levels in P1/HC-ProTu and P1/HC-ProTu/atg8age plants compared with those of Col-0. Data are presented as mean values ± standard errors (n = 3). e Hierarchical clustering dendrogram on the transcriptomes of Col-0, P1/HC-ProTu and P1/HC-ProTu/atg8age plants. f Relative expression of autophagy-related genes in P1/HC-ProTu and P1/HC-ProTu/atg8age plants compared to that in Col-0 plants. Data are presented as mean values ± standard deviations (n = 3). Significant differences based on Student’s t-test; * indicates P values < 0.05. ** indicates P values < 0.01. g Evaluation of the suppression efficiency of miRNA-mediated target gene regulation between P1/HC-ProTu and P1/HC-ProTu/atg8age plants by real-time RT-PCR. Relative expression levels were normalized to the ubiquitin level. Data are presented as mean values ± standard errors (n = 6). Significant differences were determined by Student’s t-test; ** indicates P values < 0.01. h Subcellular localization of ATG8a (i), HC-ProTu (ii), and AGO1 (iii) within leaf cells as revealed by immunogold labeling and transmission electron microscopy (TEM). Immunogold signals were detected within and/or surrounding autophagic structures, many of which were encased by autophagic vacuoles. Red arrowheads highlight the presence of immunogold particles within autophagic vacuoles (Av) or autophagic-like structures (Ap), while blue arrowheads denote signals found in the cytoplasm (n = 3). Cw, cell wall; Cv, central vacuole; Ch, chloroplast. Scale bars, 200 nm. i Image size measurements for speckle-like structures. Data are presented as mean values ± standard deviations (n = 10). j FRET efficiency of co-infiltrated plants in which HC-ProTu-CFP or HC-ProTu-K-CFP interacts with YFP-ATG8a. Data are presented as mean values ± standard deviations (n = 9). Source data are provided as a Source Data file.
GFP-ATG8a has been used as a marker in autophagy studies14,37. Hafrén et al. noted that HC-Pro induced ATG8a expression and colocalized with ATG8a in PGs14. Therefore, we generated ATG8a null alleles in the P1/HC-ProTu background (P1/HC-ProTu/atg8age) via CRISPR-Cas with 4 guided RNAs (gRNAs) for further investigation (Fig. 3c(i)). We obtained four individual knockout lines, each with a large genomic deletion of ATG8a, and all alleles exhibited recovered AGO1 levels (Supplementary Fig. 4). Line #4-5 was selected to represent the P1/HC-ProTu/atg8age genotype because it has a large ATG8a genomic deletion and undetectable ATG8a transcripts (Fig. 3c(ii), d). Western blotting revealed that AGO1 was degraded in 14-day-old P1/HC-ProTu plants (0.27-fold) compared with Col-0 plants (1.0-fold) (Fig. 2a(i)). The amount of AGO1 (0.87-fold) was greater in the P1/HC-ProTu/atg8age plants (Fig. 2a(i)). The quantitative results from western blotting and LC-MS/MS analyses consistently indicate that AGO1 levels in P1/HC-ProTu/atg8age plants have been restored to those observed in Col-0 plants (Fig. 2a(ii, iii)). Notably, the HC-ProTu levels of the P1/HC-ProTu and P1/HC-ProTu/atg8age plants line #4–5 and others were similar (Supplementary Fig. 4b). The cotyledon morphology of P1/HC-ProTu/atg8age plants was similar to the round shape of Col-0 cotyledons, unlike the elliptical shape of P1/HC-ProTu cotyledons (Supplementary Fig. 5), suggesting that the narrow cotyledon phenotype in the P1/HC-ProTu background was partially recovered in atg8a mutants.
We conducted a comparative transcriptomic comparison to assess the recovery of AGO1 levels and morphology in P1/HC-ProTu/atg8age plants. Hierarchical cluster analysis of the transcriptome data revealed that P1/HC-ProTu/atg8age plants closely resembled Col-0 plants (Fig. 3e). Additionally, in P1/HC-ProTu/atg8age plants, the expression levels of several autophagy-related genes, such as ATI3A, ATI3C, UBAC2A, and ATG1, were comparable to those in Col-0 plants (Fig. 3f; Supplementary Fig. 6). However, NBR1 remained consistently highly expressed in both P1/HC-ProTu and P1/HC-ProTu/atg8age plants (Fig. 3f). This effect is likely due to NBR1, a selected autophagy receptor positioned at the upstream level of the pathway. The expression of NBR1 is induced in both transgenic plants by the presence of P1/HC-ProTu. Furthermore, the transcript levels of miRNA targets such as AGO1 (a target of miR168), GRF1 (a target of miR156/157), and ARF16 (a target of miR160) were 3- to 8-fold higher in P1/HC-ProTu plants than those in Col-0 plants (Fig. 3g). In contrast, these targets showed a 2- to 4-fold increase of expression in P1/HC-ProTu/atg8age plants (Fig. 3g). These results implied that ATG8a is involved in the AGO1 degradation triggered by P1/HC-ProTu and the restored levels of AGO1 contribute under atg8a mutation to effective RNA silencing.
Yanagisawa et al. demonstrated frequent observation of autophagic vacuole (Av)/autophagic-related (Ap) structures in the cytoplasm surrounding chloroplasts and at the periphery of central vacuoles (Cv)38. When visualized using immunoelectron microscopy, these double-membrane stromule-baring Ap structures are initiated with an Av surrounded by a white ring-like structure38. The final size of these selective Ap structures is diverse in plants, with a range of ~40–500 nm in diameter in the mesophyll cells of Arabidopsis39. We similarly detected the presence of Av/Ap structures that emerged near the chloroplast edges within the leaf cells of P1/HC-ProTu plants (Fig. 3h).
Immunogold labeling with α-ATG8 antibodies detected ATG8 signals around the Av/Ap structures (Fig. 3h(i), red arrowheads), indicating that these signals likely correspond to ATG8-phosphatidylethanolamine (ATG8-PE) associated with autophagosome membranes. Additionally, a subset of ATG8 immunogold signals was observed in the cytoplasm (Fig. 3h(i), blue arrowheads). Immunogold labeling with α-HC-ProTu antibodies revealed significant signal accumulation within the Av/Ap structures, suggesting that HC-ProTu is incorporated into these structures (Fig. 3h(ii), red arrowheads). Furthermore, AGO1 immunogold signals were detected within the Av/Ap structures (Fig. 3h(iii), red arrowhead), with one observation showing AGO1 localization in the matrix of an Av/Ap structure, merging into a Cv for degradation (Fig. 3h(iii), lower panel). The AGO1 immunogold signals were less abundant than those of ATG8 and HC-ProTu, potentially due to the low endogenous levels of AGO1 in P1/HC-ProTu plants.
Additionally, Yoshimoto et al. demonstrated that YFP-ATG8a localizes in a dotted pattern, indicative of autophagosome precursors40. These YFP-ATG8a foci (~750 nm) were observed in both the cytoplasm and nucleus, with no significant changes in their distribution when YFP-ATG8a was co-expressed with CFP (Supplementary Fig. 7a). However, some ATG8a foci colocalized with HC-ProTu-CFP or HC-ProTu-K-CFP, and these YFP-ATG8a foci increased in size to ~1600 nm in diameter (Fig. 3i; Supplementary Fig. 7b). Approximately 30% of the FRETs showed colocalization of YFP-ATG8a with HC-ProTu-CFP or HC-ProTu-K-CFP, suggesting a direct interaction between HC-ProTu and YFP-ATG8a, which likely facilitates the recruitment of YFP-ATG8a to H-bodies (Fig. 3j).
Various HC-Pro variants have different inhibition abilities of HEN1 activity
Next, we assessed the ability of various P1/HC-Pro plants to inhibit miRNA methylation. Analysis of β-elimination-treated miRNAs revealed that P1/HC-ProTu plants contained ~50% unMet-miR159 and unMet-miR165, both appearing at the 20-nt positions (Fig. 4a(i)). Quantitative analysis from three independent biological replicates indicates that P1/HC-ProTu plants exhibited 49.1% unMet-miR159 and 45.7% unMet-miR165, respectively (Fig. 4a(ii)). In contrast, P1/HC-ProZy and P1/HC-ProTe plants displayed only modest levels of unMet-miR159 and unMet-miR165 (Fig. 4a(i)). Notably, P1/HC-ProTu-K plants showed the slight presence of unMet-miR159 (Fig. 4a(i)). A more detailed quantification revealed that P1/HC-ProZy plants retained 23.3–25% unMet-miRNAs, while P1/HC-ProTe plants retained 12.5–16.1% unMet-miRNAs (Fig. 4a(ii)). These findings suggest that the ability of P1/HC-ProZy and P1/HC-ProTe plants to inhibit HEN1-mediated miRNA methylation is significantly weaker than that of P1/HC-ProTu plants. Additionally, P1/HC-ProTu-K plants exhibited a substantial reduction in the ability to suppress HEN1 methyltransferase activity, retaining only 7.9–16.9% unMet-miRNAs (Fig. 4a(ii)).
a Evaluation of miRNA methylation status in various P1/HC-Pro plants by β-elimination (i). Total RNA samples were treated with β-elimination (+) or without treatment (−). The β-elimination assay was used to determine the presence of methylation in miRNAs (ii). Bar charts represent the percentage of methylated (Met) and unmethylated (unMet) miR159 and miR165 in the samples. Error bars indicate standard deviations based on three biological replicates (n = 3). b Evaluation of miRNA methylation in recombinant P1/HC-Pro plants by β-elimination. c Evaluation of the miR159 methylation status in ZYMV-infected squash plants by β-elimination (n = 3) (i). P1/HC-ProTu plants were used as a positive control. HC-ProZy detection in ZYMV-infected squash plants (ii). The asterisk indicates the RUBISCO loading control. d YFP channel for the YFP-HEN1 sample. The yellow arrowheads indicate HEN1 foci. In, inclusion body. Bar, 50 μm. e Image size measurements for speckle-like structures. (i) CFP channel for HC-ProTu-CFP, which was co-expressed with YFP-HEN1. (ii) YFP channel for YFP-HEN1, which is co-expressed with HC-ProTu-CFP. Bar, 50 μm. f The sizes of various bodies. Error bar, standard deviation Data are presented as mean values ± standard deviations (n = 10). g Subcellular colocalization of HC-ProTu-K-CFP (i) or HC-ProTu-CFP (iii) with YFP-HEN1 (ii and iv). Bar, 50 μm. In, inclusion body. N, nucleus. Arrowheads indicate speckle-like structures. h FRET efficiency of co-infiltrated plants in which HC-ProTu-CFP or HC-ProTu-K-CFP interacts with YFP-HEN1. Data are presented as mean values ± standard deviations (n = 9). Source data are provided as a Source Data file.
In addition, among the two recombinant P1/HC-Pro plants, only the P1Zy/HC-ProTu plants exhibited ~50% unMet-miRNAs, while the P1Tu/HC-ProZy plants exhibited 100% Met-miRNAs (Fig. 4b). These findings suggest that the ability of HC-ProZy and HC-ProTe to inhibit HEN1 is not as potent as that of HC-ProTu. To address potential concerns regarding the suitability of Arabidopsis as a host for ZYMV, which could lead to inadequate suppression of HEN1 by HC-ProZy, we utilized squash plants infected with ZYMV for further analysis of miRNA methylation. Both severe and mild strains of ZYMV-infected squash plants displayed 100% Met-miRNA in the presence of HC-ProZy (Fig. 4c(i, ii)). Notably, P1/HC-ProTu plants served as a positive control for the detection of unMet-miRNA in this experiment (Fig. 4b, c). Notably, the unMet-miRNA levels resulting from the inhibition of HEN1 by HC-ProZy are very low, sometimes undetectable. This may be influenced by factors such as the northern blot washing conditions or the intensities of the radiolabeling.
YFP-HEN1 forms cytoplasmic foci that are ~598–733 nm in diameter, and these foci also aggregate to form irregular bodies (Fig. 4d). When co-expressed, YFP-HEN1 colocalized with HC-ProTu-CFP in the same foci (Fig. 4e), and the size of the YFP-HEN1 foci was increased to 2000 nm (Fig. 4e, f), suggesting that HEN1 can be recruited into the H-body by HC-ProTu. YFP-HEN1 was detectable in both the cytoplasm (Fig. 4e(ii)) and nucleus (Fig. 4g(ii)). In instances where YFP-HEN1 colocalized with HC-ProTu-CFP within the same cellular foci, the YFP-HEN1 signal was absent from the nucleus (Fig. 4g(iii, iv)). A substantial FRET efficiency of ~55% was observed (Fig. 4h), indicating a physical interaction between YFP-HEN1 and HC-ProTu-CFP. However, YFP-HEN1 did not colocalize with HC-ProTu-K-CFP in H-bodies, maintaining its presence in the nucleus (Fig. 4g(ii)). Consequently, there was no detectable FRET efficiency between HC-ProTu-K-CFP and YFP-HEN1 (Fig. 4h).
In vitro HC-ProTu-mediated HEN1 inhibition
We purified recombinant proteins, including GST, his-HEN1, GST-HC-ProTu, GST-HC-ProTu-K, GST-HC-ProZy, and his-P19CIRV, for conduction of in vitro experiments (Fig. 5a). An in vitro HEN1 activity assay revealed 100% Met-miRNA in his-HEN1 or His-HEN1 + GST samples (Fig. 5b). GST-HC-ProTu inhibited his-HEN1 activity in a dose-dependent manner, whereas neither GST-HC-ProTu-K nor GST alone inhibited his-HEN1 (Fig. 5b). Thus, the in vivo and in vitro data suggest that HC-ProTu directly inhibits HEN1 activity through physical interactions9.
a Recombinant protein purification (n = 10). b In vitro HEN1 methylation inhibition assay with the GST-HC-ProTu or GST-HC-ProTu-K protein. The methylation of synthetic miRNAs was examined via oxidation/β-elimination and small RNA northern blotting assays. The methylation status is indicated on the lower side of the panel. GST served as a negative control (n = 8). c In vitro EMSA for HEN1 binding to miRNA duplexes with/without various recombinant HC-Pros (n = 5). d In vitro EMSA for various HC-Pro in inhibiting HEN1-miRNA duplex binding activity (n = 8).
Radioisotope-labeled miRNA/miRNA*s exhibited a mobility shift when incubated with His-HEN1 or His-HEN1 with GST, suggesting HEN1-miRNA duplex binding ability (Fig. 5c). Notably, His-p19CIRV, which is a small RNA binding VSR, was employed as a positive control and exhibited a mobility shift (Fig. 5c). Interestingly, neither GST-HC-ProTu nor GST-HC-ProZy displayed any EMSA signal, indicating that both HC-Pros lacked small RNA binding activity (Fig. 5c). Moreover, the binding ability of HEN1 to the miRNA duplex was inhibited when the reaction included co-incubation with either GST-HC-ProTu or GST-HC-ProZy (Fig. 5d). This inhibition of HEN1 by HC-Pro increased in a dose-dependent manner with the amount of recombinant proteins present (Fig. 5d), indicating that the FRNK motif plays a role in mediating the competitive binding of HEN1 to miRNA duplexes. In the samples with varying concentrations of HC-ProTu, the levels of free miRNA duplexes were measured at 0.8-fold and 0.9-fold greater than the original miRNA concentration. In contrast, samples with HC-ProZy exhibited free miRNA duplex levels 0.65-fold and 0.67-fold greater than the initial concentration (Fig. 5d). These findings indicate that HC-ProTu is more effective than HC-ProZy in inhibiting HEN1 binding to miRNA duplexes.
The unMet-miRNAs are not detectable in the AGO1 of P1/HC-Pro Tu plants
The miRNA methylation status of AGO1-IP from Col-0, P1/HC-ProTu, and P1/HC-ProTu/atg8age plants was evaluated. All the AGO1-IPs contained Met-miR159, whereas the total RNA contained both Met- and unMet-miR159 (Fig. 6a). This result implies two possibilities. One is that unMet-miR159 cannot load into AGO1 due to the unknown ability of HC-ProTu. The other is that unMet-miR159 can be loaded into AGO1, but this unMet-miR159-AGO1 complex is further degraded, leading to undetectable signals of unMet-miR159 in AGO1-IP. Notably, AGO1 from different samples was successfully immunoprecipitated from these plants (Supplementary Fig. 8a).
a β-elimination of the miRNA methylation status in the total RNA and AGO1-IP samples from 1-week-old Col-0, P1/HC-ProTu, and P1/HC-ProTu/atg8age plants (n = 3). b MiRNA methylation status in total RNA and AGO1-IP samples from 1-week-old hen1-8/heso1-1 mutants and P1/HC-ProTu/hen1-8/heso1-1 plants that were compared with Col-0 plants and P1/HC-ProTu plants (n = 3). c In vitro RISC assay in Col-0, hen1-8/heso1-1, and ago1-27 mutants (n = 3). d The endogenous AGO1 levels in the P1/HC-ProTu/hen1-8/heso1-1 plant were compared to those in Col-0 and P1/HC-ProTu plants (i). Relative AGO1 protein expression levels were normalized to the actin (ACT) loading control and quantified using the bar chart (ii). Data are presented as mean values ± standard errors (SE, n = 3). Significant differences based on Student’s t-test; ** indicates P values < 0.01. e The enzyme-linked immunosorbent assay (ELISA) analysis for evaluation of the TuMV severe strain (TuGR) and mild strain (TuGK) infectivity in Col-0, dcl2-4/dcl4-1, and hen1-8/heso1-1 plants at 14 days post-inoculation (dpi) using TuMV coat protein (CP) antibody at 1/20,000 dilution. Data are presented as mean values ± standard deviations (n = 20). Source data are provided as a Source Data file.
To determine whether unMet-miRNA can be loaded into AGO1, we employed the hen1-8/heso1-1 mutant as a model. Examination of total RNA following β-elimination treatment revealed that ~50% of the miRNAs in the P1/HC-ProTu plants remained unMet-miRNAs (Fig. 6b). However, the β-elimination signals of miRNAs in hen1-8/heso1-1 mutants were very weak, likely due to the overall reduced miRNA levels in the hen1-8/heso1-1 mutant. Notably, unMet-miR165 was detectable in these samples (Fig. 6b, black arrowhead), confirming that the miRNAs in hen1-8/heso1-1 are predominantly unmethylated. This finding was further corroborated by similar results in P1/HC-ProTu/hen1-8/heso1-1 plants, where only unMet-miR159 and unMet-miR165 were present (Fig. 6b). A more detailed characterization of P1/HC-ProTu/hen1-8/heso1-1 plants will be presented subsequently.
Next, we analyzed the AGO1-IP samples from hen1-8/heso1-1 mutants. Our results revealed that AGO1 was indeed associated with unMet-miRNAs (Fig. 6b). To assess the functional capacity of these AGO1 complexes, we conducted an in vitro RISC assay using the immunoprecipitated AGO1-IP samples from hen1-8/heso1-1 mutants (Supplementary Fig. 8b). The results demonstrated that these AGO1 complexes, similar to those purified from Col-0 plants, retained RISC activity and effectively cleaved the MYB33_230 substrate (Fig. 6c). This indicates that AGO1 loaded with unMet-miRNAs remains competent for RNA cleavage. These findings align with those reported by Tu et al., who observed that despite the loss of miRNA methylation in hen1-8/heso1-1 mutants, RISC activity was not compromised25. Furthermore, the AGO1-IP from the ago1-27 mutant exhibited significantly reduced RNA cleavage activity (Fig. 6c). This reduced activity is attributable to the ago1-27 mutation, characterized by an Ala992Val substitution, which renders it a weak allele41. This mutant served as a negative control in the experiments.
Given that both P1/HC-ProTu plants and hen1-8/heso1-1 mutants produce unMet-miRNAs, but with HESO1 remaining wild-type in P1/HC-ProTu plants, we generated P1/HC-ProTu/hen1-8/heso1-1 progeny by crossing these two genotypes (Fig. 6b). Analysis of AGO1-IP samples from P1/HC-ProTu/hen1-8/heso1-1 plants revealed the presence of unMet-miRNAs (Fig. 6b). Additionally, the levels of AGO1 protein in P1/HC-ProTu/hen1-8/heso1-1 plants were restored to 0.62-fold relative to wild-type levels (Fig. 6d). However, whether unMet-miRNAs in P1/HCTu plants are loaded onto or associated with AGO1 and how HEN1 and HESO1 influence this process, remains unclear and further investigation.
The hen1-8/heso1-1 and dcl2-4/dcl4-1 double mutants were inoculated with a severe (TuGR) or mild (TuGK) strain of TuMV, which contains the HC-ProTu-K protein11. The results showed that infections with TuGR in Col-0 plants, dcl2-4/dcl4-1, and hen1-8/heso1-1 mutants resulted in viral titers exceeding 1.5 at OD405 (Fig. 6e). Notably, the viral titers in dcl2-4/dcl4-1 and hen1-8/heso1-1 mutants were greater than those in TuGR-infected Col-0 plants (Fig. 6e). In contrast, TuGK infection of Col-0 plants led to a lower viral titer, ~1.0 at OD405 (Fig. 6e). However, in the dcl2-4/dcl4-1 mutants, a higher viral titer was observed (Fig. 6e), corroborating the findings of Kung et al.11 Additionally, the hen1-8/heso1-1 mutants exhibited a viral titer of ~2.0 at OD405 (Fig. 6e). These results collectively suggest that hen1-8/heso1-1 mutants, similar to dcl2-4/dcl4-1 mutants, are highly susceptible to TuMV infection.
Discussion
Potyviral HC-Pro variants differentially inhibit HEN1 activity
This study presents a comparative analysis of the RNA silencing suppression abilities of various P1/HC-Pros, highlighting how distinct potyviral HC-Pro variants differentially inhibit HEN1. Our findings reveal that HC-ProTu exerts the most potent inhibition of HEN1 methyltransferase activity, achieving ~49% suppression. In contrast, HC-ProZy and HC-ProTe exhibit progressively weaker inhibitory effects, with reductions in HEN1 activity of 15–20% and 8–12%, respectively. The observed variability in HEN1 inhibition among these HC-Pros may be attributed to the significant sequence divergence across the variants, which share only 40% overall sequence similarity. Notably, the C-terminus of these proteins is relatively conserved (75.6% similarity) and possesses protease activity essential for processing the viral polyprotein into functional HC-Pro (Supplementary Fig. 9). In contrast, the N-terminus is less conserved (48.8% similarity) but contain the highly conserved FRNK motif, which is crucial for mediating interactions between HC-Pro and HEN1. Moreover, the EMSA results provide further insights into these interactions, demonstrating that HC-ProZy is capable of binding HEN1 and competing with miRNA for HEN1 binding. This suggests that the N-terminus of HC-ProTu may endow it with a unique capacity to inhibit HEN1 activity, a feature that might be less prominent in HC-ProZy or absent in HC-ProTe. Therefore, potential motifs or sequences on HC-ProTu that can inhibit methyltransferase activity are worthy of further investigation.
HC-ProTu and HC-ProZy exhibit a binding affinity for HEN1 rather than miRNA
Mérai et al. obtained VSR extracts using susceptible materials or transient expression methods to conduct EMSA experiments42 and reported that many viral VSRs have miRNA duplex binding activity, especially when HC-ProTe is present in plant extracts. In contrast, PVY HC-Pro exhibited weaker EMSA signals, indicating variations in miRNA duplex binding capabilities among the different potyviruses HC-Pros. In contrast, immunoprecipitation experiments with other VSR transgenic plants revealed clear miRNA signals in the p19 and p21 IP samples but no miRNA signals in the HC-ProTu IP samples. This finding aligns with our EMSA results, where neither GST-HC-ProTu nor GST-HC-ProZy had miRNA duplex binding signals in the EMSA. Shiboleth et al. and Garcia-Ruiz et al. reported that in susceptible materials infected with ZYMV and TuMV, respectively, immunoprecipitated HC-Pro contained miRNAs and viral siRNAs43,44. However, it is challenging to exclude complexes formed between HC-Pro and other viral proteins in susceptible materials, which might provide small RNA binding capability. Moreover, our SEC data demonstrated that HC-ProTu can form multimers concentrated in fractions 8–12 and that these fractions did not show a significant accumulation of miRNAs, with most miRNAs accumulating in free-form fractions, suggesting that HC-ProTu alone does not directly bind miRNA duplexes. Therefore, the high-molecular-weight unMet-miRNAs found in P1/HC-ProTu plants may be bound to other plant proteins rather than to AGO1.
H-bodies might restrict the activity of HEN1 and AGO1
FRET can be employed to assess physical interactions between two different proteins. The FRNK motif of HC-ProTu plays a crucial role in its interaction with HEN128. Therefore, HC-ProTu-K lacks the ability to bind HEN1 and cannot recruit HEN1 to H-bodies. We demonstrated a 60% FRET efficiency between HC-ProTu and HEN1, consistent with colocalization and physical interaction. The recruitment of HEN1 to H-bodies by HC-ProTu could hinder HEN1 from methylating miRNA and siRNA duplexes. ATG8a co-localizes with HC-ProTu and AGO1 in H-bodies14. Our FRET analysis detected ~30% energy transfer between ATG8a and HC-ProTu in H-bodies, indicating a physical interaction. Although fewer colocalized foci were observed in the HC-ProTu-K-CFP + YFP-ATG8a sample, the FRET data still revealed a similar 30% energy transfer, suggesting that HC-ProTu-K also physically interacts with ATG8a, but is unable to trigger H-bodies. Although it remains unclear whether this recruitment of ATG8a to H-bodies is related to autophagy induction or whether it serves as a means for HC-ProTu to evade autophagic attack, the physical interaction between HC-ProTu and ATG8a in H-bodies holds significant biological importance and warrants further investigation.
Similarly, Hafrén et al. and Ivanov et al. observed the colocalization of HC-Pro and AGO1 in PGs14,45. FRET analysis indicated that HC-ProTu does not interact physically with AGO1, suggesting that a third protein might assist HC-ProTu in recruiting AGO1 into H-bodies. In a study of PVA HC-Pro, Ivanov et al. reported that HC-Pro requires the ribosomal protein L18B (RPL18B) for indirect binding with AGO1, which is consistent with the FRET observation that HC-ProTu does not directly interact with AGO145. Although the formation of H-bodies induced by HC-Pro has been observed in TuMV and PVA, H-body formation was not observed in HC-ProZy, highlighting the variation among different HC-Pro proteins. Consequently, we propose that HC-ProTu physically interacts with HEN1 and ATG8a to recruit them to H-bodies, whereas HC-ProTu might indirectly recruit AGO1 to H-bodies through RPL18B (Fig. 7a).
a The inside of the H-body for HC-Pro interacts with RNA silencing components. b The working hypothesis for HC-ProTu-induced degradation of AGO1 suggests that HC-ProTu exhibits a heightened inhibitory effect on HEN1 methylation activity. This inhibition results in the incorporation of unmethylated miRNAs (unMet-miRNAs) into AGO1 complexes, which subsequently activate autophagy pathways, leading to the degradation of AGO1.
Confocal microscopy revealed that H-bodies range in diameter from ~1000–2000 nm, significantly larger than the AP/AV structures (<1000 nm) observed via TEM. Through immunogold TEM, we identified the presence of HC-Pro and AGO1 within peroxisomes, which also measured between 1000 and 2000 nm in diameter (Supplementary Fig. 10). Peroxisomes in plant cells play a multifaceted role, being essential for the metabolism of reactive oxygen species (ROS) and reactive nitrogen species, photorespiration, the glyoxylate cycle, and the modulation of transcriptional response via H2O2 signaling, particularly in the context of disease resistance and biotic stress46. VSRs from cucumber mosaic virus and barley stripe mosaic virus modulate peroxisomal ROS levels by degrading catalase and reducing H2O2 through interaction with glucose oxidase, respectively47,48. Additionally, the P15 of the peanut clump virus induces the sequestration of viral siRNAs within peroxisomes49. These interactions between viruses and peroxisomes are implicated in modulating plant defense mechanisms, as well as viral replication and infection. Based on our observations, we hypothesize that H-bodies may be associated with peroxisomes. Regardless, using FRET technology, H-bodies present an ideal platform for further investigation of HC-Pro and host protein interactions.
HESO1 might play an essential role in RNA silencing suppression
HEN1, a plant-specific methyltransferase, plays a critical role in the distinct process of Met-miRNA-mediated RNA silencing. In the miRNA turnover pathway, SDNs initiate miRNA degradation by targeting AGO1-bound miRNAs, leading to their 3′ truncation (demethylation). Subsequently, these truncated miRNAs are uridylated by HESO1 and URT1, further degrading them and significantly affecting the growth and development of hen1-8 mutants. Similarly, our investigation using hen1-8/heso1-1 mutants confirmed that unMet-miRNAs can be effectively loaded into AGO1, enabling the RISC complex to target and cleave RNA.
Tu et al. demonstrated that AGO1-IP from hen1 heso1-2 urt1-3 mutants contain uridylated unMet-miRNAs, and AGO1 levels in these mutants remain stable25. However, in the P1/HC-ProTu/hen1-8/heso1-1 plants, we observed both the presence of unMet-miRNAs and recovery of AGO1 levels. Interestingly, although HC-ProTu strongly inhibits HEN1, we did not detect uridylated unMet-miRNAs in P1/HC-ProTu plants. This observation is consistent with the established understanding that uridylation of unMet-miRNAs by HESO1 and URT1 predominantly occurs within the RISC complex. The absence of uridylated unMet-miRNAs could be attributed to the ability of HC-ProTu to sequester HEN1 and AGO1 within H-bodies (Fig. 7A), preventing the methylation of free-form miRNAs and their subsequent loading onto AGO1. This distinction between P1/HC-ProTu plants and hen1-8/heso1-1 mutants suggests that HC-ProTu possesses unique properties that deserve further exploration.
Furthermore, hen1-8/heso1-1 mutants, like dcl2-4/dcl4-1 mutants, are susceptible to the mild strain TuGK. DCL2 and DCL4 are crucial for processing replicated viral RNA, and mutations in these genes compromise the primary defense against viruses of plants, leading to severe symptoms and viral accumulation in dcl2-1/dcl4-1 mutants akin to those caused by the severe strain TuGR. However, the susceptibility of hen1-8/hseo1-1 mutants to TuGK, despite intact DCL2 and DCL4 functions, raises significant questions. These results suggest that HEN1 and HESO1 not only participate in the antiviral defense mechanism of plants but may also play a more central regulatory role, possibly superseding the functions of DCL2 and DCL4 in this context.
HC-Pro-mediated unMet-miRNA abundance modulates autophagic AGO1 degradation
Several studies demonstrated that AGO1 is regulated by autophagy, particularly in the presence of certain VSRs. For instance, Polerovirus VSR P0 can cause autophagic AGO1 degradation, indicating that AGO1 degradation is a common suppression strategy used by various viruses18. Cell biological assays indicate that ATG8a can colocalize with AGO1, NBR1, and HC-Pro in PGs14,15, and several ATG8 family genes (ATG8a, ATG8b, ATG8g, and ATG8h) are induced by P1/HC-Pro. Among these genes, GFP-ATG8a was chosen as a marker gene for studying the regulation of AGO1 autophagy. The presence of ATG8a null alleles in P1/HC-ProTu plants leads to the restoration of AGO1 levels. Why other ATG8 family members cannot compensate for the loss of function of ATG8a is not known. VSP29, a protein involved in vacuolar protein trafficking and the recycling of vacuolar sorting, interacts with P1 of TuMV9. According to the transcriptome profile of P1/HC-ProTu plants, CML24 appears in the positive regulatory network, and CML24 interacts with ATG4b to participate in the cleavage of the C-terminus of ATG8a and can activate autophagy50. Additionally, immunoelectron microscopy revealed the presence of HC-Pro, AGO1, and ATG8 gold particles within AP/AV structures in the cells of P1/HC-ProTu plants. These findings indicate that P1/HC-ProTu-mediated AGO1 degradation is associated with the autophagy pathway.
Although P1/HC-ProTu plants exhibited a significant upregulation of ATG-related genes, a similar but less pronounced increase was also observed in P1/HC-ProTu-K, P1/HC-ProZy, and P1/HC-ProTe plants, and this differential gene expression is reflected in the varying degrees of AGO1 degradation across these plants. We hypothesize that the difference in HEN1 inhibition by distinct HC-Pro variants, leading to the accumulation of unMet-miRNAs at varying concentrations, may be linked to their ability to induce autophagic degradation of AGO1 (Fig. 7b).
Accumulation of free-form miRNAs indicates the impact of HC-Pro on RISC formation
In animal cells, the empty state of Drosophila AGO1 (not miRNA-loaded AGO1) is degraded by selective autophagy51. Similarly, in Arabidopsis, FBW2 is involved in the autophagic degradation of non-functional AGO1 that is not loaded with miRNA19. Wang et al. showed that HEN1 is involved not only in miRNA methylation but also in miRNA-AGO1 loading and RISC assembly52. HC-ProTu can bind and inhibit HEN1 activity, which can significantly impact RISC assembly. Moreover, various P1/HC-Pros cause abnormal accumulation of free-form miRNA duplexes, suggesting defects in miRNA loading onto AGO1. Notably, the P1/HC-ProTu-K plants also exhibited an accumulation of free-form miRNAs, implying that free-form miRNA accumulation might cause multiport effects, which might be related to other factors in addition to HEN1 inhibition. Iki et al. and Iwasaki et al. demonstrated that HSP90 assists in the loading of miRNA and siRNA duplexes into AGO1, a process that consumes ATP53,54. Removing miRNA* from AGO1 also requires the ATP hydrolysis ability of HSP90. PVY HC-Pro inhibits NtMinD ATPase activity, suggesting that HC-Pro might affect cellular ATP production25. We hypothesize that HC-Pro impacts RISC assembly, potentially by disrupting miRNA biogenesis or altering ATP production. Such interference could indirectly affect the energy dynamics necessary for HSP90-AGO1 to facilitate the loading of miRNA duplexes onto the RISC. Consequently, this disruption in miRNA-AGO1 loading could be a common mechanism leading to the accumulation of free-form miRNA duplexes.
Different isoforms of AGO1
Finally, we observed various isoforms of AGO1 in this study. AGO1-L emerged as the dominant and most abundant isoform, in contrast to AGO1-S, which was found in lower quantities and was sometimes undetectable (Fig. 2a). The α-AGO1 antibodies utilized in this research were produced using a recombinant protein from the N-terminus of AGO1 (1-to-256 aa) as the antigen55. This approach differs from other studies in which peptides were used to generate commercial AGO1 antibodies18. Consequently, the AGO1 antibodies used in this study can recognize a more extensive portion of the N-terminus of AGO1. This N-terminal region is a long and unstructured N-terminal extension (NTE) whose function in the miRNA-AGO1 loading process has not been identified56. Moreover, Bressendorff et al. discovered that the N-terminus of AGO1 exhibits structural flexibility in the N-coil region57. When AGO1 is loaded with miRNA, the N-coil region becomes concealed. Conversely, in AGO1, which is not loaded with miRNA, the N-coil region is exposed and directly interacts with the autophagy cargo receptor ATI1, triggering the degradation of unloaded AGO1. These findings underscore the significance of the NTE region in the functionality and stability of AGO1, suggesting that more in-depth research into the AGO1-L and AGO1-S isoforms is needed.
HC-Pro is a multifunctional protein with various specific motifs and domains. Apart from interfering with RNA silencing, HC-Pro is involved in protein cleavage, aiding virus transmission by aphids, and participating in virus replication. While different types of potyvirus HC-Pros all possess RNA silencing suppression abilities, the degree of suppression varies among them, leading to various impacts on the expression of endogenous plant genes. Specifically, due to its ability to inhibit HEN1 methyltransferase and induce autophagic AGO1 degradation more effectively, HC-ProTu has more potent RNA suppression capabilities than the other two species of HC-Pro examined here. We also observed that P1/HC-Pro led to the accumulation of free-form miRNA duplexes, which might be related to the impact of HC-Pro on RISC assembly. Finally, the ability of HC-ProTu to inhibit HEN1 leading to autophagic AGO1 degradation, represents a compelling area for future research. This opens up new opportunities for exploring various aspects of P1/HC-Pro suppression, as well as further investigation into miRNA-loading and RISC assembly.
Methods
Plant material, growth conditions, chemical treatment, and virus inoculation
Arabidopsis ecotype Columbia (Col-0), mutants (ago1-27, dcl2-4/dcl4-1, and hen1-8/heso1-1), and transgenic plants in the Col-0 background (P1/HC-ProTu, P1/HC-ProTu-K, P1/HC-ProZy, P1/HC-ProTe, P1Zy/HC-ProTu, and P1Tu/HC-ProZy plants) were used in this study9,11,12. P1/HC-ProTu/hen1-8/heso1-1 seeds were obtained by crossing with P1/HC-ProTu and hen1-8/heso1-1 plants. Seeds were surface-sterilized and plated on MS media supplemented with or without the appropriate antibiotics and incubated in a growth room (16 h light/8 h darkness, 22–24 °C). Seven-day-old seedlings were used for analysis or transferred to a Florabella potting compost/sand mix (3:1) and maintained under 16 h light/8 h darkness at 22–24 °C. Fourteen-day-old seedlings were treated with 15 μM E64-d (Sigma) or 40 μM MG132 (Sigma) for 12 h, after which AGO1 levels were assessed via western blotting.
Arabidopsis seedlings were subjected to mechanical inoculation with TuMV. Zucchini squash seedlings were subjected to mechanical inoculation with ZYMV. TuMV or ZYMV inoculum was prepared by homogenizing 0.5 g of virus-infected leaves with 2 mL of 0.01 M sodium phosphate buffer, pH 7.2. The newly developed leaves at 10 dpi were collected to evaluate virus levels.
CRISPR-mediated knockout of the ATG8a gene in P1/HC-Pro Tu plants
ATG8a-edited P1/HC-ProTu plants were generated via CRISPR technology. Four gRNA sequences (Supplementary Table 1) were designed, and off-target effects were predicted by CRISPR-P 2.0 (crispr.hzau.edu.tw/CRISPR2/). Underlined nucleotides indicate the PAM sequence. The four gRNA sequences were cloned and inserted into the pEHH401E binary vector58 under the U6-26, U6-27, U6-28, and U6-29 promoters to generate the pEH401E-T8a vector. pEH401E-T8 was introduced into P1/HC-ProTu plants to generate P1/HC-ProTu/atg8age plants.
Size exclusion chromatography, RNA purification, and small RNA northern blotting
One-week-old seedlings (1 g) were homogenized in 1.5 mL of extraction buffer [200 mM Tris-HCl, pH 7.4, 150 mM NaCl, 5 mM DTT, Protease Inhibitor Cocktail (Roche Applied Science) and Proteasome Inhibitor Set II (Calbiochem)], followed by centrifugation at 13,000 × g for 10 min to remove cell debris. After filtration through a 0.22 μm filter, the extract was loaded onto a HiPrep 16/60 Sephacryl S-200 HR column (GE Healthcare) via an AKTA explore FPLC system. The FPLC system was run at a constant flow rate of 0.5 mL/min, and every 1.5 mL fraction was collected. Additional FPLC runs of given size markers were conducted to determine the relative size of the FPLC fractions. Protein was extracted from each fraction for western blotting. Uncropped blots are shown in the Source Data file.
TRIzol reagent was used to purify RNA from the fractionated solution, and glycogen was employed as a carrier to precipitate small amounts of RNA. Small RNA northern blotting was conducted by separating total RNA in a 15% acrylamide gel with 8 M urea. The blot was then hybridized with 32P radioisotope-labeled oligonucleotide probes complementary to the specific miRNA sequence59. The probes were stripped out, and the blot was rehybridized with another miRNA probe. Uncropped blots are shown in the Source Data file.
Recombinant protein purification
Protein expression clones of pET-HEN1, pGEX-HC-ProTu, pGEX-HC-ProTu-K, pGEX-HC-ProZy, and pET-p19CIRV were transformed into the E. coli BL21 strain. For his-HEN1 and his-P19CIRV purification, the input lysate was loaded into a HiTrap FF column, washed with wash buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 1 mM DTT, and 20 mM imidazole) and eluted with elution buffer (50 mM Tris-HCl, pH 8.0, 300 mM 1 mM DTT, and 200 mM imidazole). The purified protein was dialyzed in dialysis buffer (10 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1 mM DTT, 0.1 mM EDTA, and 2 mM MgCl2). For the purification of GST-HC-ProTu, GST-HC-ProTu-K, and GST-HC-ProZy, the lysis, wash, and dialysis buffers were all 1× PBS buffer. GSTrap FF column (GE Healthcare) was used. The elution buffer was prepared with 10 mM glutathione in 1× PBS buffer.
HEN1 methylation assay and β-elimination
For the in vivo HEN1 methylation assay, it was performed as described28. The 0.4 ng of ds-syn-miR159 was then subjected to a methylation process with 8 µL of purified his-HEN1 protein, 10 µL NEB Cutsmart buffer, and 3.2 mM SAM. Periodate treatment followed by β-elimination was performed22. Purified RNA was dissolved in 176 μL of 0.06 M borax/boric acid (pH 8.6) and 24 μL of 0.2 M sodium periodate, followed by incubation in the dark at RT for 1 h. One-tenth volume of glycerol was then added, followed by incubation for an additional 30 min. The solution was cleaned with prepacked Sephadex G-25 resin (GE Healthcare), 1 μL of glycogen (15 μg/μL) was added, and the RNA was precipitated as described. After RNA precipitation, β-elimination was performed with 0.055 M borax/boric acid/NaOH (pH 9.5) at 45 °C for 90 min. The solution was then cleaned with Sephadex G-25 resin. Finally, the RNA was precipitated again and subjected to small RNA northern blotting. Uncropped blots are shown in the Source Data file.
In vivo AGO1-IP and in vitro RISC assay
For in vivo IP, 1-week-old seedlings (1 g) were homogenized with 1 mL of IP buffer (25 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1 mM EDTA, 5% glycerol, 1% NP-40) followed by centrifugation for 10 min at 4 °C. Notably, IgG α-AGO155 or α-HA (Santa Cruz Biotech.) was used for the in vivo IP in this study. IP was performed by mixing cleaned protein A-agarose beads (50 μL of suspension per IP reaction) (Santa Cruz, sc-20001), IgG (30 μL per IP reaction), and the lysate. The IP mixture was gently mixed at 4 °C for 3 h. The tube was centrifuged at 300 × g to pull down the beads, and nonspecific binding was washed twice with 0.3 mL of IP buffer, after which the mixture was suspended in 0.3 mL of IP buffer. The IP eluates were used for western blotting or small RNA extraction. Moreover, the AGO1-IP eluates were used for AGO1-containing small RNA extraction and β-elimination.
cDNA fragments of MYB33_230 containing a miR159 target site were amplified with the primer sets listed in Supplementary Table 1 and cloned and inserted into a pGEM-T easy vector (Promega). In vitro transcription was performed using a MEGAscript T7 Transcription Kit (Invitrogen) with the MYB33_230 template, and the products were incubated with 1 μL of fresh [α-P32] UTP (PerkinElmer). Unincorporated UTP was removed with G50 resin (GE Healthcare). The RNA transcript was separated a 6% denaturing polyacrylamide gel in 1× TBE buffer [29:1 acrylamide/bis (Bio-Rad), 8 M urea]. The gel was exposed to chemiluminescence film (GE Healthcare) to visualize the RNA. The band with a strong signal was excised and mashed in 300 μL of extraction buffer (250 mM NaOAc and 1 mM EDTA) and passed through centrifuge tube filters (Costar). The supernatant was further cleaned up by the TRIzol method. RNA was diluted with water to 300–500 CPM radioactive strength in 5 μL.
For the AGO1-IP reaction for RISC activity, 15 μL of 1 mg/mL IgG, 30 μL of beads, and 500 μL of lysate were added, and the mixture was subsequently resuspended in 50 μL of 1× PBS. For the slicer assay, 5 μL of RNA and 25 μL of 2× cleavage buffer [2× PBS, 266.6 mM KCl, 26.6 mM MgCl2, 33.4 mM DTT, 6.6 mM ATP, 1.4 mM GTP, 0.25% RiboLock (Thermo Scientific)] were added to 5–20 μL of AGO1-IP beads in 1× PBS. The reaction mixture was incubated at 25 °C for 1 h under rotation. The RNA fragment was purified by the TRIzol method and solubilized in 10 μL of water with 10 μL of RNA Gel Loading Dye (Invitrogen) at 85 °C for 2 min of denaturation. The RNA was separated on a 6% denaturing polyacrylamide gel and exposed to chemiluminescence film for 5 days to obtain images. Uncropped blots are shown in the Source Data file.
Electrophoretic mobility shift assay (EMSA)
The synthetic miR159 sense and antisense strands were annealed using 5× annealing buffer (300 mM KCl, 30 mM HEPES, pH 7.5, and 1 mM MgCl2) at 95 °C for 5 min and then cooled to room temperature. Double-stranded synthetic miR159 (ds-syn-miR159) was labeled with γ-32P by using T4 polynucleotide kinase and subsequently purified with G-25 beads. Different purified proteins were added to a solution of 0.4 ng radioisotope-labeled ds-syn-miR159 with 5× reaction buffer (40 mM Tris-HCl, pH 8.0, 1 mM MgCl2, 30 mM KCl, 0.01% NP-40, and 1 mM dithiothreitol). The mixture was incubated at room temperature for 1 h. Afterward, the mixture was mixed well with 6× loading dye (0.25% bromophenol blue, 0.25% xylene cyanol, and 40% glycerol) and subsequently loaded into a 5% native gel (30% acrylamide 37.5:1, 0.5× TBE buffer, 10% APS, and 1% TEMED). The electrophoresis gel was run at 60 V for 1 h. The gel was dried at 60 °C for 45 min and then exposed to the X-ray film. Uncropped blots are shown in the Source Data file.
Western blotting and real-time RT-PCR
For western blotting, plant tissue was ground with 10 mM sodium phosphate buffer (pH 7.0) and mixed with denaturing buffer (50 mM Tris-HCl, pH 6.8, 4% SDS, 2% 2-mercaptoethanol, 10% glycerol, and 0.001% bromophenol blue). The protein extract was then boiled at 100 °C for 10 min, separated via SDS-PAGE and transferred to a Hybond-P PVDF membrane (GE Healthcare). Western blotting was performed using α-AGO1 (10,000× dilution), α-HC-ProZy (10,000× dilution), and α-HC-ProTu (10,000× dilution) antibodies generated in our previous study9,60. The α-ACT (8000× dilution) (Sigma) antibody was used as a loading control.
For real-time RT-PCR, 3 μg of total RNA was subjected to reverse transcription with poly-dT primers using the SuperScript III First-Strand Synthesis System (Invitrogen). Real-time PCR was performed using a LightCycler 480 instrument (Roche) with the primer sets listed in Supplementary Table 1. All detections were performed with 4 independent replicates. Relative expression was calculated and normalized according to the expression levels of AtUBQ10.
Confocal microscopy and FRET
To observe the localization of HC-ProTu-CFP or HC-ProTu-K-CFP with YFP-HEN1, YFP-AGO1, and YFP-ATG8a, the YFP and CFP fluorescence was detected in 3-day infiltrated Nicotiana benthamiana leaves. Each sample, with a total of 10 biological replicates, was examined using a Leica TCS SP5 II confocal laser‐scanning microscope (Joint Center for Instruments and Research, College of Bioresources and Agriculture, National Taiwan University) equipped with a multiline argon laser with a filter set for CFP (458 nm laser line, PMT1 gain = 467–521 nm), and YFP fluorescence (514 nm laser line, PMT2 gain = 525–564 nm). Single foci, but not aggregated foci, were chosen for the size measurements. Each particle’s size equals the constant 2.355 multiplied by the d result generated by ImageJ. The FRET value reported in this study is the average value of FRET efficiency taken from 20 colocalized foci for each protein pair. The error bar is the standard deviation.
Transcriptome analysis
The total RNA extraction using the Plant Total RNA Extraction Miniprep System (VIOGENE, Taipei, Taiwan) and the PF-Total RNA Purification Kit (XenoHelix, Incheon, Korea). The FPKMs of the genes of interest from each sample were considered significant according to Student’s t-test. Genes with a p value < 0.05 are labeled in red based on the significance level. PCA was performed via R (version 4.2.2, www.R-project.org) using the built-in R functions prcomp(). Cluster dendrogram analysis was performed using the R based on the Euclidean distance and hierarchical Ward’s method with the package “stats.”
Transmission electron microscopy
First true leaves of 14-day-old seedlings were fixed in 4% paraformaldehyde and 0.1 % glutaraldehyde in 0.1 M sodium phosphate buffer (PBS) with pH 7.0 at room temperature overnight. After three times PBS rinses, samples were dehydrated in a graded ethanol series at room temperature and embedded in LR White resin. Ultrathin sections, 90–120 nm, were cut using a Reichert Ultracut S or Leica EM UC6 or UC7 ultramicrotome (Leica, Vienna, Austria) and collected with 50 mesh nickel grids.
For immunogold labeling, the individual grids were floated on PBS for 15 min, and then were blocked with PBS containing 1% BSA for 15 min. The grids were incubated with primary polyclonal antibodies α-ATG8 (35× diluted with PBS and 1% BSA), α-HC-ProTu (100× diluted with PBS and 1% BSA), and α-AGO1 (100× diluted with PBS and 1% BSA) for 1 h at room temperature. After being washed four times with PBS, the grids were floated on an excess amount (1:20 dilution) of 18 nm colloidal anti-rabbit IgG (Jackson Immuno Research, West Grove, PA, USA) at room temperature for 1 h. The grids were washed sequentially with four droplets of PBS, followed by five times ddH2O washes. After immunogold labeling, the sections were stained with 5% uranyl acetate in water for 10 min and 0.5% lead citrate for 4–6 min. Sections were observed using a Joel, JEM1400 Transmission Electron Microscope (Tokyo, Japan) at 80 kV, and the images were obtained with a Gatan Orius CCD camera.
LC-MS/MS analysis and database search
Proteins were reduced by 10 mM dithiothreitol (DTT) at 29 °C for 1 h followed by alkylation in 55 mM iodoacetamide (IAA) in 1 M TEAB (Triethylammonium bicarbonate) at 29 °C for 1 h in the dark. After adding 5 mM DTT at 29 °C for 45 min, proteins were digested by 0.1 μg of trypsin (Promega, Madison, WI, USA) at 37 °C for 16 h. After digestion, 10% TFA was added to a final concentration of 0.5% in the sample. Tryptic peptides were separated on Ultimate system 3000 nanoLC system (Thermo Fisher Scientific, Bremen, Germany) by using a 75-μm-ID, 25-cm-length C18 Acclaim PepMap column packed with 2 μm particles with a pore size of 100 Å (Thermo Scientific, San Jose, CA, USA). Mobile phase A was 0.1% formic acid in water, and mobile phase B was composed of 100% acetonitrile with 0.1% formic acid. A segmented gradient from 2% to 40% acetonitrile was used over 90 min. The flow rate was 300 nl/min. MS/MS analysis was performed on an Orbitrap Fusion Lumos Tribrid quadrupole-ion trap-Orbitrap mass spectrometer (Thermo Fisher Scientific, San Jose, CA) by using data-dependent mode with full-MS under the condition of a mass accuracy <5 ppm, a resolution of 120,000 at m/z = 200, AGC target 5e5, and maximum injection time of 50 ms. HCD-MS/MS was performed on the most intense ions in 3 s with a resolution of 15,000, within a 1.4 Da isolation window at a normalized collision energy of 32%, AGC target 5e4, previously selected ions dynamically excluded for 60 s, and max injection time of 50 ms. For protein identification, the raw MS/MS data were searched against the Uniprot Arabidopsis thaliana database (downloaded on August 2023) by using the Mascot and SEQUEST search algorithms via the Proteome Discoverer (PD) package (version 2.2, Thermo Scientific). The search parameters were set as follows: peptide mass tolerance, 10 ppm; MS/MS ion mass tolerance, 0.02 Da; enzyme set as trypsin and allowance of up to two missed cleavages; and variable modifications included oxidation on methionine, deamidation on asparagine and carbamidomethylation on cysteine residues. Peptides were filtered based on a 1% FDR. For relative protein quantification across different samples, each protein group is represented by a single master protein (PD grouping feature), and the raw abundance of each protein was calculated.
Reagents and chemicals
All the reagents and chemicals used in this study are provided in Supplementary Table 2.
Statistics and reproducibility
Statistical analyses were performed using R (version 4.2.2, www.R-project.org). Sample size, p values, and statistical methods employed are described in the respective figure legends or in the Source Data. Differences were considered to be significant at p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw sequence reads from different P1/HC-Pro plants in this study have been deposited in the NCBI SRA database under Bioproject PRJNA1156224 and Bioproject PRJNA1154811. Source data are provided with this paper.
Change history
30 July 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41467-025-62542-y
References
Revers, F. & García, J. A. Molecular biology of potyviruses. Adv. Virus Res. 92, 101–199 (2015).
Yang, X., Li, Y. & Wang, A. Research advances in potyviruses: from the laboratory bench to the field. Annu. Rev. Phytopathol. 59, 1–29 (2021).
Carthew, R. W. & Sontheimer, E. J. Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Hoehener, C., Hug, I. & Nowacki, M. Dicer-like enzymes with sequence cleavage preferences. Cell 173, 234–247.e237 (2018).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 11, 597–610 (2010).
Kurihara, Y., Takashi, Y. & Watanabe, Y. The interaction between DCL1 and HYL1 is important for efficient and precise processing of pri-miRNA in plant microRNA biogenesis. RNA 12, 206–212 (2006).
Reinhart, B. J., Weinstein, E. G., Rhoades, M. W., Bartel, B. & Bartel, D. P. MicroRNAs in plants. Genes Dev. 16, 1616–1626 (2002).
Anandalakshmi, R. et al. A viral suppressor of gene silencing in plants. Proc. Natl. Acad. Sci. USA 95, 13079–13084 (1998).
Hu, S. F. et al. Investigation of the effects of P1 on HC-Pro-mediated gene silencing suppression through genetics and omics approaches. Bot. Stud. 61, 22 (2020).
Kasschau, K. D. & Carrington, J. C. A counterdefensive strategy of plant viruses: suppression of posttranscriptional gene silencing. Cell 95, 461–470 (1998).
Kung, Y. J. et al. Genetic analyses of the FRNK motif function of Turnip mosaic virus uncover multiple and potentially interactive pathways of cross-protection. Mol. Plant Microbe Interact. 27, 944–955 (2014).
Wu, H. W., Lin, S. S., Chen, K. C., Yeh, S. D. & Chua, N. H. Discriminating mutations of HC-Pro of Zucchini yellow mosaic virus with differential effects on small RNA pathways involved in viral pathogenicity and symptom development. Mol. Plant Microbe Interact. 23, 17–28 (2010).
Sahana, N. et al. The asparagine residue in the FRNK box of potyviral helper-component protease is critical for its small RNA binding and subcellular localization. J. Gen. Virol. 95, 1167–1177 (2014).
Hafrén, A. et al. Turnip mosaic virus counteracts selective autophagy of the viral silencing suppressor HCpro. Plant Physiol. 176, 649–662 (2018).
Hafrén, A., Lõhmus, A. & Mäkinen, K. Formation of potato virus A-induced RNA granules and viral translation are interrelated processes required for optimal virus accumulation. PLoS Pathog. 11, e1005314 (2015).
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nat. Cell Biol. 9, 1102–1109 (2007).
Shin, K. D., Lee, H. N. & Chung, T. A revised assay for monitoring autophagic flux in Arabidopsis thaliana reveals involvement of AUTOPHAGY-RELATED9 in autophagy. Mol. Cells 37, 399–405 (2014).
Derrien, B. et al. Degradation of the antiviral component ARGONAUTE1 by the autophagy pathway. Proc. Natl. Acad. Sci. USA 109, 15942–15946 (2012).
Hacquard, T. et al. The Arabidopsis F-box protein FBW2 targets AGO1 for degradation to prevent spurious loading of illegitimate small RNA. Cell Rep. 39, 110671 (2022).
Johnston, M., Geoffroy, M. C., Sobala, A., Hay, R. & Hutvagner, G. HSP90 protein stabilizes unloaded argonaute complexes and microscopic P-bodies in human cells. Mol. Biol. Cell 21, 1462–1469 (2010).
Yu, B., Chapman, E. J., Yang, Z., Carrington, J. C. & Chen, X. Transgenically expressed viral RNA silencing suppressors interfere with microRNA methylation in Arabidopsis. FEBS Lett. 580, 3117–3120 (2006).
Yu, B. et al. Methylation as a crucial step in plant microRNA biogenesis. Science 307, 932–935 (2005).
Ren, G., Chen, X. & Yu, B. Uridylation of miRNAs by hen1 suppressor1 in Arabidopsis. Curr. Biol. 22, 695–700 (2012).
Ren, G. et al. Methylation protects microRNAs from an AGO1-associated activity that uridylates 5’ RNA fragments generated by AGO1 cleavage. Proc. Natl.Acad. Sci. USA 111, 6365–6370 (2014).
Tu, B. et al. Distinct and cooperative activities of HESO1 and URT1 nucleotidyl transferases in microRNA turnover in Arabidopsis. PLoS Genet. 11, e1005119 (2015).
Wang, X. et al. Synergistic and independent actions of multiple terminal nucleotidyl transferases in the 3’ tailing of small RNAs in Arabidopsis. PLoS Genet. 11, e1005091 (2015).
Yu, Y. et al. ARGONAUTE10 promotes the degradation of miR165/6 through the SDN1 and SDN2 exonucleases in Arabidopsis. PLoS Biol. 15, e2001272 (2017).
Sanobar, N. et al. Investigating the viral suppressor HC-Pro inhibiting small rna methylation through functional comparison of HEN1 in angiosperm and bryophyte. Viruses 13, 1837 (2021).
Chapman, E. J., Prokhnevsky, A. I., Gopinath, K., Dolja, V. V. & Carrington, J. C. Viral RNA silencing suppressors inhibit the microRNA pathway at an intermediate step. Genes Dev. 18, 1179–1186 (2004).
Kasschau, K. D. et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA unction. Dev. Cell 4, 205–217 (2003).
Mallory, A. C., Reinhart, B. J., Bartel, D., Vance, V. B. & Bowman, L. H. A viral suppressor of RNA silencing differentially regulates the accumulation of short interfering RNAs and micro-RNAs in tobacco. Proc. Natl. Acad. Sci. USA 99, 15228–15233 (2002).
Mlotshwa, S. et al. Ectopic DICER-LIKE1 expression in P1/HC-Pro Arabidopsis rescues phenotypic anomalies but not defects in microRNA and silencing pathways. Plant Cell 17, 2873–2885 (2005).
Xie, Z., Kasschau, K. D. & Carrington, J. C. Negative feedback regulation of Dicer-Like1 in Arabidopsis by microRNA-guided mRNA degradation. Curr. Biol. 13, 784–789 (2003).
Plisson, C. et al. Structural characterization of HC-Pro, a plant virus multifunctional protein. J. Biol. Chem. 278, 23753–23761 (2003).
Ruiz-Ferrer, V. et al. Structural analysis of tobacco etch potyvirus HC-pro oligomers involved in aphid transmission. J. Virol. 79, 3758–3765 (2005).
Baunoch, D. A., Das, P. & Hari, V. Potato virus Y helper component protein is associated with amorphous inclusions. J. Gen. Virol. 71, 2479–2482 (1990).
Michaeli, S. et al. The viral F-box protein P0 induces an ER-derived autophagy degradation pathway for the clearance of membrane-bound AGO1. Proc. Natl. Acad. Sci. USA 116, 22872–22883 (2019).
Yanagisawa, M. & Chuong, S. D. X. Chloroplast envelopes play a role in the formation of autophagy-related structures in plants. Plants 12, https://doi.org/10.3390/plants12030443 (2023).
Holzinger, A., Kwok, E. Y. & Hanson, M. R. Effects of arc3, arc5 and arc6 mutations on plastid morphology and stromule formation in green and nongreen tissues of Arabidopsis thaliana. Photochem. Photobiol. 84, 1324–1335 (2008).
Yoshimoto, K. et al. Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4s are essential for plant autophagy. Plant Cell 16, 2967–2983 (2004).
Morel, J. B. et al. Fertile hypomorphic ARGONAUTE (ago1) mutants impaired in post-transcriptional gene silencing and virus resistance. Plant Cell 14, 629–639 (2002).
Mérai, Z. et al. Double-stranded RNA binding may be a general plant RNA viral strategy to suppress RNA silencing. J. Virol. 80, 5747–5756 (2006).
Garcia-Ruiz, H. et al. Arabidopsis RNA-dependent RNA polymerases and dicer-like proteins in antiviral defense and small interfering RNA biogenesis during Turnip mosaic virus infection. Plant Cell 22, 481–496 (2010).
Shiboleth, Y. M. et al. The conserved FRNK box in HC-Pro, a plant viral suppressor of gene silencing, is required for small RNA binding and mediates symptom development. J. Virol. 81, 13135–13148 (2007).
Ivanov, K. I. et al. Molecular insights into the function of the viral RNA silencing suppressor HCPro. Plant J. Cell Mol. Biol. 85, 30–45 (2016).
Ferreira, A. R., Marques, M., Ramos, B., Kagan, J. C. & Ribeiro, D. Emerging roles of peroxisomes in viral infections. Trends Cell Biol. 32, 124–139 (2022).
Yang, M. et al. Barley stripe mosaic virus gammab interacts with glycolate oxidase and inhibits peroxisomal ROS production to facilitate virus infection. Mol. Plant 11, 338–341 (2018).
Nemes, K., Gellert, A., Balazs, E. & Salanki, K. Alanine scanning of cucumber mosaic virus (CMV) 2b protein identifies different positions for cell-to-cell movement and gene silencing suppressor activity. PLoS ONE 9, e112095 (2014).
Incarbone, M. et al. Neutralization of mobile antiviral small RNA through peroxisomal import. Nat. Plants 3, 17094 (2017).
Tsai, Y. C., Koo, Y., Delk, N. A., Gehl, B. & Braam, J. Calmodulin-related CML24 interacts with ATG4b and affects autophagy progression in Arabidopsis. Plant J. Cell Mol. Biol. 73, 325–335 (2013).
Kobayashi, H., Shoji, K., Kiyokawa, K., Negishi, L. & Tomari, Y. VCP machinery mediates autophagic degradation of empty argonaute. Cell Rep. 28, 1144–1153.e1144 (2019).
Wang, X. et al. Degradation of unmethylated miRNA/miRNA*s by a DEDDy-type 3’ to 5’ exoribonuclease Atrimmer 2 in Arabidopsis. Proc. Natl. Acad. Sci. USA 115, E6659–e6667 (2018).
Iki, T. et al. In vitro assembly of plant RNA-induced silencing complexes facilitated by molecular chaperone HSP90. Mol. Cell 39, 282–291 (2010).
Iwasaki, S. et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 39, 292–299 (2010).
Hong, S. F. et al. Development of an assay system for the analysis of host RISC activity in the presence of a potyvirus RNA silencing suppressor, HC-Pro. Virol. J. 20, 10 (2023).
Xu, Y. et al. The N-terminal extension of Arabidopsis ARGONAUTE 1 is essential for microRNA activities. PLoS Genet. 19, e1010450 (2023).
Bressendorff, S. et al. Importance of an N-terminal structural switch in the distinction between small RNA-bound and free ARGONAUTE. Nat. Struct. Mol. Biol. 8, 806–814 (2025).
Wang, Z. P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Niu, Q. W. et al. Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Nat. Biotechnol. 24, 1420–1428 (2006).
Chiu, M. T., Lin, C. P., Lin, P. C. & Lin, S. S. Enhancement of IgG purification by FPLC for a serological study on the Turnip mosaic virus P1 protein. Plant Pathol. Bull. 22, 21–30 (2013).
Acknowledgements
We thank Dr. Hervé Vaucheret and Dr. Xuemei Chen for providing us with the seeds of dcl2-4 dcl4-1 and hen1-8 heso1-1 mutants, respectively. This research was supported by the National Science and Technology Council, Taiwan (NSTC112-2311-B-002-024-MY3, NSTC 113-2321-B-005-003).
Author information
Authors and Affiliations
Contributions
S.S.L., P.C.L., Z.J.P., and W.L.W. conceptualized the study and designed experiments. P.C.L. initiated experiments and ideas related to HC-Pro inhibiting HEN1 and HC-Pro–induced autophagic degradation of AGO1. S.S.L. and T.L.S. supervised the study and acquired funding. Z.J.P. and W.L.W. performed most experiments in this study. P.A.T. performed confocal image and FRET analysis. R.Y.F. and S.F.H. performed in vitro RISC assay. T.H.P. J.T.Y., and C.T.C. performed EMSA. B.N.S. and S.F.H. performed the antibody production. H.H.C., W.N.J., C.H.C., and C.C.W. performed TEM. H.Y.W. performed LC-MS/MS and database search. Q.W.S., S.F.Hu, P.C.L., and W.L.W. performed the transgenic plant productions. P.C.L. and W.L.W. performed the SEC and northern blotting assays. F.H.W. and C.S.L. performed gene editing. S.S.L., Z.J.P., W.L.W., Y.L.H., and J.L.B. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pan, ZJ., Wei, WL., Tran, PA. et al. HC-Pro inhibits HEN1 methyltransferase activity, leading to autophagic degradation of AGO1. Nat Commun 16, 2503 (2025). https://doi.org/10.1038/s41467-025-56320-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56320-z