Abstract
Radiation-induced mucositis significantly reduces quality of life in patients undergoing radiotherapy and chemoradiotherapy for head and neck cancer. Radiation exposure increases the secretion of small extracellular vesicles carrying double-stranded DNA, which triggers excessive inflammation. To address this, we develop functionalized organic nanosheets designed to capture these inflammatory vesicles from damaged tissue. Using template-based synthesis, we create nanostructured organic sheets functionalized with CD63 aptamers, enabling selective targeting of extracellular vesicles involved in mucositis. These nanosheets show enhanced vesicle-binding capacity compared to spherical nanoparticles, efficiently suppressing inflammation by inhibiting the stimulator of interferon genes activation in macrophages. Additionally, they effectively scavenge reactive oxygen and nitrogen species, further alleviating mucosal inflammation. Flow cytometry and transcriptome analyses in irradiated animal models confirm significant mucositis mitigation. This therapeutic platform provides a promising anti-inflammatory strategy by demonstrating how biomaterial geometry and surface functionalization can modulate small extracellular vesicle-mediated inflammation in radiation-induced mucositis.
Similar content being viewed by others
Introduction
Radiation-induced mucositis (RIM) is a debilitating complication commonly associated with radiotherapy and chemoradiotherapy in patients with head and neck cancer1. As a side effect of antineoplastic treatment, this condition significantly impacts patients’ quality of life due to severe pain, nutritional deficits, and an increased risk of infection, often leading to hospitalization in about 62% of cases and necessitating tube feeding in 70% of patients2. Characterized by painful inflammation and ulceration of the mucosal membranes in the mouth, throat, and nasal regions, RIM presents a multifaceted challenge for effective management3. Additionally, radiation-induced sinusitis, resulting from damage and inflammation to the nasal tissue, is often overlooked in clinical practice and lacks unified management protocols4. Although studies on potentially useful drugs are underway, they remain in the exploratory stage and have yet to achieve clinical application5.
Despite these efforts, the management of RIM continues to be a significant clinical challenge. Current therapeutic strategies are largely palliative, focusing on symptom relief rather than addressing the underlying causes. Basic daily care, anti-inflammatory agents, hydration, lubrication, dietary adjustments, pain management, and low-level laser therapy (LLLT) are employed to mitigate the symptoms, yet they often fall short of providing complete relief6. Recent studies have revealed that radiation-induced tissue injury can enhance the biogenesis and secretion of small extracellular vesicles (sEVs) through DNA damage induction7,8.
Furthermore, radiation can cause double-strand breaks in DNA, which can increase the production and release of sEVs. This process is often mediated in a p53-dependent manner, highlighting the role of DNA damage response pathways in regulating sEVs secretion9. Given the pivotal role of sEVs in mediating pro-inflammatory signaling pathways, we hypothesize that targeting these vesicles could offer a different approach to mitigating mucositis. Our prior work employed molybdenum disulfide (MoS₂), a promising 2D material known for its exceptional biodegradation, conjugated with PAMAM dendrimer for sEVs capture and scavenging7,10. Building upon this, we have shifted our focus towards developing an inorganic-free nanoplatform, employing template-based synthesis to construct polyglycerol-amine (PG) nanosheets conjugated with CD63 aptamers. Aptamers are functional oligonucleotides obtained through in vitro or in vivo screening, capable of binding to their targets with high specificity and affinity due to their unique tertiary structures11. To optimize the performance of sEV scavenging, we introduced CD63 aptamers to enhance the binding and capture of free sEVs. We selected hyperbranched polyglycerol (hPG) due to its excellent biocompatibility, chemical stability, protein resistance, and tunable blood circulation properties12. This innovative approach leverages the flexible and crimpable 2D structure of nanosheets13, offering a higher binding capacity to sEVs compared to conventional PG-based nanospheres (PG-NP). By eliminating sEVs and reactive oxygen and nitrogen species (RONS) in radiated tissues, these functionalized nanosheets inhibit macrophage stimulator of interferon genes (STING) activation and exhibit robust anti-inflammation ability.
Chronic inflammation and persistent free radical generation are known to contribute to the adverse effects of radiotherapy14. RONS activation of the NF-κB pathway further exacerbates this inflammatory state, disrupting normal tissue function15,16. Our nanosheets, composed of PG and thiol (-SH) groups, are promising to exhibit RONS reduction capabilities. The residue thiol groups in the functional nanosheets provide direct antioxidant effects by neutralizing free radicals through electron donation, converting them into more stable and less reactive molecules17. Therefore, targeting both sEVs and RONS offers a multifaceted approach to mitigating the long-term impacts of radiation. In addition, the microbiome’s role in RIM pathogenesis is becoming increasingly evident18, and radiation-induced shifts in microbiome composition could exacerbate the chronic inflammation seen in mucositis19.
We initially synthesized organic polyglycerol nanosheets (PG-NS) by degrading the MoS2 template and then obtained the CD63 aptamer-modified PG-NS (PG-NSA). This functionalized 2D organic nanoplatform demonstrated mild protein adsorption, low cytotoxicity, robust sEVs binding, and RONS reduction ability in inflammatory tissues after radiation, making it suitable for further investigation. To validate the unique 2D structure’s efficiency, we compared the sEV capture ability of PG-NSA against PG-NPA, demonstrating superior performance. We then evaluated the protective effects and therapeutic efficacy of this multifunctional nanosheet through comprehensive studies using in vitro, in vivo, and ex vivo models. Moreover, flow cytometry coupled with t-SNE analysis and RNA-sequencing of radiated tissues provided detailed insights into the nanosheet’s therapeutic mechanisms and efficacy.
Results
Elevated sEVs secretion in radiation-induced mucositis (RIM) patients
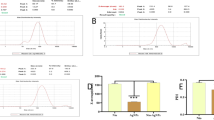
A total of 20 RIM patients with rhinosinusitis after radiotherapy were included in this study. Demographic and clinical data, including blood parameters, were collected before and after radiotherapy (Table S1). Nasal secretions from these patients were sampled pre- and post-RT, and EVs were isolated through ultracentrifugation (100,000 × g, 70 min)20. Dynamic Light Scattering (DLS) measurements demonstrated that the sEV size remained at ~70 nm pre- and post-RT (Supplementary Fig. 1a). Western blot (WB) analysis confirmed the enrichment of CD63, a canonical exosomal marker, in the isolated sEV fractions compared to cell lysates, with concurrent reduction of GAPDH, validating the purity and characteristic protein signature of our EV preparations (Supplementary Fig. 1c). Nanoparticle Tracking Analysis (NTA) and Transmission Electron Microscopy (TEM) revealed predominant sEV dimensions of ~120 nm in both conditions, with only a slight increase observed post-radiation (Supplementary Fig. 1a). TEM imaging further confirmed the preservation of characteristic circular or cup-shaped morphology, indicative of maintained structural integrity post-irradiation (Fig. 1a). Quantitative analysis revealed a statistically significant increase in both sEV concentration and associated dsDNA content following radiation treatment (Fig. 1b), with the normalized dsDNA/sEVs ratio exhibiting approximately 2-fold elevation (Supplementary Fig. 1b), suggesting radiation-induced enhancement of nucleic acid packaging into sEVs.
a The NTA and TEM images of sEVs isolated from nasal secretions of RIM patients before and after RT. Scale bars: 200 nm. b The relative sEVs amount and dsDNA level in nasal secretions. Data represent the mean ± S.D. (n = 20 samples). c The correlations of IL-6 and TNF-α with relative sEVs amount and cfDNA level in nasal secretions from RIM patients. d Representative NET images of neutrophils treated with post-radiation sEVs and sEVs+C-178. Scale bars: 20 µm. e Representative CD11b+/F4/80+/CD86+ in the RAW 264.7 cells treated with sEVs and sEVs+C-178. f Quantification of the percentage of CitH3 positive area, MPO positive area in NET images, and the percentage of macrophages with M1 phenotype. Data represent the mean ± S.D. (n = 3 independent experiments). Statistical significances were assessed by paired two-tailed Student’s t-test (b for sEVs), Wilcoxon test (b for dsDNA), or one-way ANOVA (f), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Abbreviations: RT: radiotherapy; sEVs: small extracellular vesicles; CitH3: citrullinated histone H3; MPO: myeloperoxidase. Source data are provided as a Source Data file.
To elucidate the mechanistic role of sEV-associated dsDNA in STING activation, we subjected sEVs to various treatments prior to cellular stimulation. STING reporter assay revealed that sEVs collected after radiation (sEVs(AR)) induced significantly higher STING activation compared to pre-radiation sEVs(BR) (p < 0.01). Notably, while sonication treatment of sEVs(AR) did not significantly alter STING activation, DNase treatment significantly attenuated this effect, and combined sonication and DNase treatment further reduced STING activation to levels comparable with control conditions (Supplementary Fig. 1d). Both ultracentrifugation and sucrose gradient methods isolated sEVs with comparable size, yield, dsDNA content, and STING activation potential (Supplementary Fig. 2 and Supplementary Fig. 3). In contrast, large EVs from irradiated cell supernatants exhibited significantly lower dsDNA levels and reduced STING activation capacity (Supplementary Fig. 4), supporting sEVs as key mediators of radiation-induced inflammation.
Consistent with these findings, flow cytometric analysis of RAW 264.7 cells revealed that treatment with post-radiation sEVs induced significant M1 macrophage polarization (CD11b+/F4/80+/CD86+) compared to control conditions. While sonication treatment of sEVs had minimal impact on M1 polarization, DNase treatment attenuated this effect, and combined sonication and DNase treatment further reduced M1 polarization to near-baseline levels (Supplementary Fig. 6, with supplementary Fig. 5 as the gating strategy), confirming the critical role of sEV-associated dsDNA in driving macrophage activation. Correlation analysis demonstrated a significant positive association between relative sEV levels and pro-inflammatory cytokines interleukin-6 (IL-6) and tumor necrosis factor-α (TNF-α) (Fig. 1c). Similarly, relative dsDNA content exhibited a significant positive correlation with IL-6 and TNF-α. Investigation of neutrophil extracellular trap (NET) formation following neutrophil exposure to post-radiation sEVs revealed robust immunofluorescent signals for citrullinated histone H3 (CitH3) and myeloperoxidase (MPO) compared to control conditions21. Subsequent application of the nitrofuran derivative C-178, an irreversible STING inhibitor that binds to cysteine residues Cys91 on the STING protein, thereby preventing activation and downstream signaling22, significantly attenuated both NET formation and M1 macrophage polarization induced by post-radiation sEVs (Fig. 1d, e, with supplementary Fig. 5 as the gating strategy). Quantitative image analysis demonstrated that both CitH3 and MPO-positive areas exceeded 15% in sEV-treated samples, significantly higher than controls (p < 0.001) (Fig. 1f). Flow cytometric analysis revealed that exposure of RAW 264.7 cells to post-radiation sEVs induced M1 phenotype in 54% of macrophages, while co-treatment with C-178 significantly reduced this differentiation to around 30% (p < 0.01) (Fig. 1f), indicating the essential role of the STING pathway in sEV-mediated pro-inflammatory response induction.
To further elucidate the role of post-radiation sEVs in RIM, immunostaining for IL-6 expression and CitH3 deposition was performed on nasal mucosa specimens from mice subjected to various experimental conditions. As depicted in Fig. 2a, b, radiation exposure resulted in significant upregulation of IL-6 expression and enhanced NET formation. However, administration of GW4869, a selective neutral sphingomyelinase inhibitor that impedes extracellular vesicle biogenesis, to irradiated mice significantly attenuated both IL-6 expression and NET formation compared to radiation alone23. Conversely, exogenous administration of post-radiation sEVs following irradiation further augmented both IL-6 expression and NET formation beyond levels observed with radiation alone, providing additional evidence that post-radiation sEVs contribute to inflammatory response amplification and NET formation. Quantitative real-time polymerase chain reaction (qRT-PCR) analysis of additional pro-inflammatory cytokine transcripts (IL-1β, IL-6, IL-8, and TNF-α) revealed similar expression patterns, with the most pronounced upregulation observed in the RT+sEVs experimental group (Fig. 2c).
a Representative images of anti-IL-6 or anti-CitH3 immunostaining in the nasal mucosa from mice subjected to different treatments. Scale bars in IL-6 images: 100 µm; CitH3: 50 µm. b Quantification of the percentage of IL-6 positive area and CitH3 positive area in the immunostaining images. c The relative expression levels of IL-1β, IL-6, IL-8, and TNF-α in the nasal mucosa of experimental mice. Data represent the mean ± S.D. (n = 6 mice). Statistical significances were assessed by one-way ANOVA (b, c), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Abbreviations: NETs: neutrophil extracellular traps. Source data are provided as a Source Data file.
Synthesis and characterization of nanostructured organic sheets
Previous research has demonstrated that cationic 2D nanosheets can effectively eliminate tumor-derived sEVs, which is beneficial in suppressing cancer progression, including tumor growth and metastasis7. Most of the traditional 2D nanomaterials were based on inorganic backbones, such as graphene, transition-metal dichalcogenide (TMDC), MXene, and so on24,25. However, inorganic backbones in these nanosheets may present challenges for clinical translation due to potential biocompatibility issues24,25. In comparison, organic polymers are generally more biocompatible than inorganic materials, meaning they are less likely to provoke an immune response and can be safely broken down and eliminated by the body26. Therefore, the development of polymer-based, inorganic-free nanosheets, such as those constructed from hPG, offers a promising strategy to design biomaterials that combine the unique functional properties of 2D nanostructures, particularly their efficacy in sEV clearance, with the established biosafety and high-dose biocompatibility of hPG (Fig. 3).
a Schematic illustration of the synthesis process for PG-NSA nanosheets, involving immobilization, crosslinking, template degradation, and aptamer functionalization. b Application and therapeutic mechanisms of PG-NSA in alleviating excessive inflammation following radiation exposure in a mouse model of RIM. Local administration of PG-NSA captures radiation-induced sEVs, suppressing cGAS-STING signaling in macrophages, reducing RONS, modulating immune responses towards an anti-inflammatory phenotype, and promoting tissue recovery. Abbreviations: PG: polyglycerol; RONS: reactive oxygen and nitrogen species; cGAS: cyclic GMP-AMP synthase; STING: stimulator of interferon genes; NF-κB: nuclear factor-kappa B; SOD: superoxide dismutase; GPx: glutathione peroxidase; CAT: catalase.
In our previous studies, the protein-resistant properties of PG and PG-modified nanomaterials are well-recognized due to their improved interaction with water molecules, preventing unwanted interference of proteins for sEVs binding13. MoS2 monolayers were synthesized via lithium-ion intercalation27, and PGA was conjugated to the backbone to prepare PGA-covered MoS2 (M-PG)17. In the next step, HS-PEG-NHS and glutamine (Glu) were reacted to form the crosslinker HS-PEG-Glu and then immobilized onto the backbone of M-PG. Finally, PG was in situ cross-linked on the backbone, and MoS2 was degraded in HRP28 (50 U/mL) and H2O2 (0.2 mM) solution to yield PG nanosheets (PG-NS). To further promote the interaction with sEVs, CD63 aptamers (5' to 3': HOOC-TAACCACCCCACCTCGCTCCCGTGACACTAATGCTA) were conjugated to the PG-NS29, producing PG-NSA (Supplementary Fig. 7). Additionally, we explored an alternative approach for CD63 functionalization using bio-orthogonal click chemistry to obtain PG-NSA-C (Supplementary Fig. 12a). Comparative analysis with PG-NSA revealed that PG-NSA-C exhibited similar physicochemical, sEVs binding, and anti-inflammation properties (Supplementary Fig. 10 and supplementary Fig. 12). While both approaches yield nanosheets with equivalent performance, the amidation reaction was selected for this study due to its procedural simplicity and cost-effectiveness. In addition, we synthesized PEI-NSA as a control cationic nanosheet to benchmark PG-NSA performance. Despite exhibiting a similar hydrodynamic size to PG-NSA, PEI-NSA displayed increased cytotoxicity in RPMI 2650 cells and a markedly higher capacity for protein absorption, highlighting the superior biocompatibility and protein-repellent properties of PG-based nanosheets (Supplementary Fig. 12c–f).
The morphology of PG-NS and PG-NSA was characterized using atomic force microscopy (AFM), revealing a nanoscale sheet-like morphology with a lateral size predominantly around 200 nm (Fig. 4a), which is smaller than M-PG (around 400 nm). These observations were corroborated by DLS measurements, which demonstrated hydrodynamic diameters of 335.7 ± 19.6 nm for M-PG, 181.0 ± 16.6 nm for PG-NS, 194.0 ± 22.1 nm for PG-NSA, and 115.7 ± 9.0 nm for PG-NPA, with corresponding zeta potential values of 18.7 ± 2.4 mV, 15.9 ± 2.7 mV, 18.1 ± 2.0 mV, and 13.6 ± 2.7 mV, respectively (Supplementary Fig. 10). The positive zeta potential values across all formulations confirm the successful integration of cationic functional groups, while the size distribution analysis validates the dimensional differences between sheet-like structures (M-PG, PG-NS, PG-NSA) and spherical nanoparticles (PG-NPA). Notably, the synthesized PG-NPA, which served as a nanoparticle control, exhibited a spherical morphology with a size of around 100 nm (Fig. 4a, b, and Supplementary Fig. 8).
a AFM images and corresponding size distribution of PG, M-PG, PG-NS, PG-NSA, and PG-NPA. Scale bars: 200 nm. b The height profiles of lines (a–e) in the AFM images show the sheet-like nanostructures of M-PGA, PG-NS, and PG-NSA. c XPS survey spectra of M-PG and PG-NS. d High-resolution XPS mapping of C 1 s, S 2p, and Mo 3 d. e Viability of RPMI 2650 cells treated for 24 h with various concentrations of PG, M-PG, PG-NS, PG-NSA, PG-NPA, and Apt(CD63). (n = 3 independent experiments) f Protein adsorption ability of PG, M-PG, PG-NS, PG-NSA, and PG-NPA in the presence of BSA. Data represent the mean ± S.D. (n = 3 independent experiments). Statistical significances were assessed by two-way ANOVA (e, f), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Source data are provided as a Source Data file.
To confirm the complete degradation of the MoS2 template and the purity of the polymeric nanosheets30, we examined the XPS survey spectra of M-PG and PG-NS (Fig. 4c). High-resolution XPS mapping of C 1 s and S 2p indicated the presence and distribution of these elements in the samples (Fig. 4d). The absence of Mo signals in PG-NS confirmed the successful removal of the MoS2 template, resulting in pure PG-NS. A parallel study using UV-vis spectroscopy revealed that the degradation of MoS₂ to obtain PG-NS is supported by the comparison of absorbance spectra (Supplementary Fig. 9), where PG-NS exhibits altered absorbance properties compared to M-PG. This change indicated successful degradation and conversion into functional polymeric nanosheets with desirable properties for biomedical applications, such as enhanced biocompatibility.
We further assessed the cytotoxicity of the biomaterials using RPMI 2650 cells, which are human nasal epithelial cells, as a model31. After 24 h of treatment with various concentrations of PG, M-PG, PG-NS, PG-NSA, PG-NPA, and CD63 Aptamer (Fig. 4f), the results demonstrated that all formulations maintained high cell viability across the tested concentration range (10–1000 µg/mL). Even at the highest concentration of 1000 µg/mL, cell viability remained above 60% for all samples except M-PG, indicating that degradation of MoS2 improves the biocompatibility of nanosheets. Statistically significant differences were observed between certain groups at 1000 µg/mL, suggesting a concentration-dependent response. Similar trends were observed after 48 h of treatment in epithelial cells (Supplementary Fig. 17). Additionally, all these nanomaterials exhibited low protein adsorption ability (Fig. 4e). The basal MoS₂ nanosheets, being negatively charged, do not efficiently interact with the BSA protein. The PG polymer, due to its non-fouling properties and high surface energy, resists BSA adsorption13,32. This resistance is crucial for cationic polymers to suppress sEVs without interfering with serum proteins in vivo.
sEVs binding and RONS reduction of PG-NSA
We further evaluated the sEVs binding capacity of our nanoplatforms by co-incubating PKH67-labeled sEVs with Cy5-labeled nanomaterials (PG-NS, M-PGA, PG-NSA, and PG-NPA)7. As shown in Fig. 5a, the green fluorescence from PKH67-labeled sEVs did not significantly overlap with the magenta fluorescence from Cy5-labeled PG-NS and PG-NPA. In contrast, there was substantial colocalization of sEVs with M-PGA and PG-NSA, as indicated by Pearson’s r values (0.69 and 0.71, respectively). To quantify the binding efficiency, we measured the percentage of sEVs binding at different ratios of nanomaterials to sEVs (w/w) (Fig. 5b). The results demonstrated that the sEVs binding efficiency improved for all nanomaterials when the ratio increased from 1:1 to 2:1, and PG-NSA showed the highest binding at about 90%, followed by M-PGA at 70%, PG-NPA at 50%, and PG-NS at 30%. These findings highlight the superior binding capability of PG-NSA and M-PGA to sEVs compared to other nanomaterials.
a The CLSM images of PKH67-labeled sEVs incubated with Cy5-labeled PG-NS, M-PGA, PG-NSA, and PG-NPA in PBS. Pearson’s r analysis for the colocalization of green and red colors was added. Scale bars: 1 μm. b Quantitative sEVs-binding capacities. c sEVs-induced STING activation in HEK-STING cells after incubation with PG-NS, M-PGA, PG-NSA, and PG-NPA. d DPPH and OH• reduction capacities of PG, PG-NS, PG-NSA, and PG-NPA in different concentrations. e The CLSM images of DCFH fluorescence in LPS-treated RPMI 2650 cells incubated with PG, PG-NS, PG-NSA, and PG-NPA in PBS. Scale bars: 50 μm. f Quantitative DCFH fluorescence intensities in CLSM images. Data represent mean ± S.D. (n = 3 independent experiments). Statistical significances were assessed by one-way ANOVA (c, f) or two-way ANOVA (b, d), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Abbreviations: Cy5: Cyanine 5; DPPH: 2,2-diphenyl-1-picrylhydrazyl; LPS: lipopolysaccharide. Source data are provided as a Source Data file.
To elucidate the molecular mechanisms underlying the immunomodulatory effects of sEV scavenging, we conducted detailed analyses of STING pathway activation. Initial functional assays in HEK-STING reporter cells (Fig. 5c) demonstrated that radiation-induced sEVs potently activated the STING pathway, while PG-NSA treatment (2 μg/mL) significantly attenuated this pro-inflammatory activation compared to M-PGA and PG-NS groups. Further mechanistic investigation via immunoblotting (Supplementary Fig. 13) provided direct molecular evidence of STING pathway modulation. WB analysis revealed that radiation-induced sEVs substantially upregulated phosphorylated STING (p-STING) levels in RAW 264.7 cells relative to untreated controls. Notably, PG-NSA treatment selectively reduced p-STING levels and also the downstream phosphorylated TBK1 (p-TBK1) and phosphorylated IRF3 (p-IRF3) - established markers of STING pathway activation33.
In addition to excessive sEVs, RONS were also over-generated in the tissues exposed to ionizing radiation during local radiotherapy34, leading to a significant dysregulated inflammatory response. To evaluate the antioxidant activity of the PG-based nanomaterials, 2,2-Diphenyl-1-picrylhydrazyl (DPPH) and hydroxyl radicals (OH-) assays were applied35,36. As shown in Fig. 5d, PG-NS and PG-NSA nanosheets exhibited a dose-dependent capacity to reduce DPPH and OH-, and their efficacy was higher than PG and PG-NPA, which may be due to the residual thiol groups on the nanosheets37. We also assessed DCFH fluorescence to detect the RONS level in RPMI 2650 cells38, and improved levels of green fluorescence were observed in the LPS-treated cells, indicating LPS-induced RONS generation. PG, PG-NS, PG-NSA, and PG-NPA treatment significantly reduced the intracellular RONS level in the LPS-treated cells, with PG-NSA exhibiting the most substantial decrease (Fig. 5e, f). Collectively, it underscores the potential of PG-based nanosheets to mitigate immoderate oxidative stress in inflammatory conditions of RIM. Oxidative stress during inflammation can deplete glutathione (GSH), and superoxide dismutase (SOD) levels and increase malondialdehyde (MDA) levels, impairing cellular function and promoting inflammation39. Consistent with the previous results, PG-NSA and PG-NPA exhibited the most potent antioxidative effects for the neutrophils incubated with post-radiation sEVs, as evidenced by higher GSH levels, lower MDA levels, and increased SOD activity (Supplementary Fig. 14). These findings suggest that PG-NSA and PG-NPA are particularly promising in biomedical applications related to oxidative stress, such as radiation-induced inflammatory disorders.
For many patients with RIM, bacterial infections and dysbiosis in the irradiated area are frequently observed following radiotherapy and are implicated in its pathophysiology40. Given that 2D nanomaterials have been reported to disrupt bacterial membranes41, we examined the antibacterial efficacy of PG-based nanosheets in this study. Surface antibacterial assays revealed that treatment with PG, PG-NS, PG-NSA, and PG-NPA significantly reduced bacterial colonies of Staphylococcus aureus (SA) and Escherichia coli (E. coli) compared to the untreated control (Supplementary Fig. 15). Among these, PG-NS and PG-NSA demonstrate the most effective antibacterial activity, which is beneficial to restrain bacterial infection and restore the balanced microbiota in pathological tissues. The nanosheet’s capabilities in sequestering sEVs, scavenging RONS, and exhibiting antibacterial activity highlight its potential as a multifaceted therapeutic for the RIM management.
Suppression of macrophage polarization and NET formation by PG-NSA
As post-radiation sEVs could induce M1 polarization of macrophages and NET formation, we would evaluate the effectiveness of our nanoplatforms in modulating inflammation via sEVs binding. Using quantitative fluorescence colocalization analysis, we tracked the intracellular fate of PKH67-labeled sEVs (green) in RAW 264.7 macrophages when co-incubated with Cy5-labeled (magenta) nanoplatforms (PG-NS, PG-NSA, or PG-NPA). Confocal laser scanning microscopy (CLSM) imaging revealed differential interaction patterns between the nanoplatforms and sEVs after 24 h of incubation (Fig. 6a)7. Quantitative fluorescence intensity profile analysis along defined linear regions of interest demonstrated significantly higher PKH67 and Cy5 signals in cells treated with sEVs+PG-NSA compared to sEVs+PG-NS or sEVs+PG-NPA (Fig. 6b). Complementary analysis of PKH67 signals relative to LysoTracker-labeled endolysosomal compartments (red) demonstrated significant colocalization (Fig. 6c)7,42, with quantitative intensity profile analysis confirming the accumulation of sEVs within acidic vesicular compartments (Fig. 6d). This endolysosomal localization pattern was maintained across treatment conditions, suggesting that while the CD63-aptamer functionalized nanoplatforms effectively bind sEVs as shown in Fig. 6a, b, this interaction does not prevent trafficking of sEVs to lysosomal compartments.
a, c CLSM images of RAW 264.7 cells after 24 h incubation with sEVs, sEVs+PG-NS, sEVs+PG-NSA, and sEVs+PG-NPA. sEVs were labeled with PKH67 (green); PG-NS, PG-NSA, and PG-NPA were labeled with Cy5 (magenta), and lysosomes were labeled with lysotracker (red). Scale bars, 10 μm. b, d The intensity of green and magenta along lines a-d and the intensity of green and red color along lines e-h in CLSM images, respectively. e Representative CD86+/TNF-α+ in the RAW 264.7 cells treated with sEVs, sEVs+PG-NS, sEVs+PG-NSA, and sEVs+PG-NPA. f Quantification of the percentage of CD86+/TNF-α+ cells. g Representative NET images of neutrophils with different treatments. Scale bars: 20 µm. h Quantification of the percentage of CitH3 positive area and MPO positive area in NETs images. Data represent mean ± s.d. (n = 3 independent experiments). Statistical significances were assessed by one-way ANOVA (f, h) and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Source data are provided as a Source Data file.
To investigate the immunomodulatory capabilities of our nanoplatforms, we assessed their impact on both macrophage polarization and NET formation. Flow cytometric analysis of RAW 264.7 cells revealed that exposure to radiation-induced sEVs significantly upregulated the co-expression of CD86 and TNF-α, established markers of pro-inflammatory M1 polarization (Fig. 6e, with supplementary Fig. 16 as the gating strategy). Quantitative analysis demonstrated that treatment with PG-NSA and PG-NPA significantly attenuated this phenotypic shift, with PG-NSA exhibiting superior efficacy in reducing CD86+/TNF-α+ cell populations (over 15% reduction compared to sEVs alone, p < 0.001 (Fig. 6f).
Concurrently, we evaluated NET formation by immunofluorescence microscopy. Neutrophils exposed to radiation-induced sEVs exhibited extensive extracellular web-like structures positive for both MPO and CitH343, characteristic morphological features of NETosis (Fig. 6g). Morphometric analysis revealed that all nanoplatforms significantly reduced NET formation, with PG-NSA demonstrating the most pronounced inhibition of both CitH3-positive area and MPO-positive area compared to sEVs alone (Fig. 6h). Notably, the suppression of NET formation exhibited a similar pattern to the inhibition of macrophage M1 polarization across treatment groups, suggesting a coordinated modulation of multiple innate immune response mechanisms by our functionalized nanoplatforms, particularly PG-NSA.
Organic nanosheets exhibit targeted biodistribution and excellent biocompatibility
Evaluating the biodistribution and in vivo accumulation of functional nanomaterials is crucial for elucidating their efficacy and safety profiles44. In this study, we labeled PG-NS, PG-NSA, and PG-NPA with fluorescent dye Cy5 to trace their distribution in vivo. Fluorescent signals in maxillary bones (MBs), tongues, and major organs (hearts, livers, spleens, lungs, and kidneys) of mice were analyzed to determine their biodistribution systematically45. As illustrated in Fig. 7a, one day post-administration, the PG-NS, PG-NSA, and PG-NPA groups exhibited substantial local accumulation in the MBs and tongues of radiated mice, indicating their potential for effective modulation of RIM. Furthermore, fluorescence intensity in the MBs and tongues of irradiated mice was markedly higher than in sham controls (Fig. 7b and supplementary Fig. 18), a phenomenon attributable to the enhanced permeation and retention (EPR) effect in inflamed tissues46. Notably, fluorescent intensity decreased significantly by day 3 and was nearly undetectable by day 7 (Fig. 7b and supplementary Fig. 19), suggesting minimal retention in irradiated tissues and supporting the long-term biosafety of these nanoplatforms. In addition, the fluorescent signals in major organs and blood remained minimal (Supplementary Fig. 18 and 19), confirming low systemic distribution and toxicity.
a Fluorescence imaging of major organs and MBs of mice subjected to RT, RT + PG-NS, RT + PG-NSA, and PG-NPA. PG-NS, RT + PG-NSA, and PG-NPA were labeled with Cy5. b Quantitative assessment of the biodistribution of PG-NS, PG-NSA, and PG-NPA in the tongues and noses of experimental mice. c The relative sEVs levels in NALF from experimental mice. d The relative mRNA expression levels of IL-1β, IL-6, IL-8, and TNF-α in the nasal mucosa of experimental mice. e Representative images of IL-6 and CitH3 immunostaining of the nasal mucosa from experimental mice. Scale bars in IL-6 images: 100 µm; CitH3: 50 µm. f Quantification of IL-6 positive and CitH3 positive area in immunostaining slices. Data represent the mean ± S.D. Data represent the mean ± S.D. (n = 6 mice). Statistical significances were assessed by one-way ANOVA (c, d, and f) or two-way ANOVA (b), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Abbreviations: NALF: nasal lavage fluid; H: heart; Li: liver; S: spleen; T: tongue; Lu: lung; K: kidney; MB: maxillary bones. Source data are provided as a Source Data file.
To comprehensively evaluate the relative biocompatibility of our nanomaterials, we conducted a comparative toxicity assessment between inorganic-containing (M-PG) and inorganic-free nanomaterials (PG-NS, PG-NSA, and PG-NPA). Healthy mice received high-dose treatments (300 μg per mouse, administered three times) of each formulation, with subsequent histopathological and biochemical analyses performed at days 7 and 14 post-administration. Histological examination via H&E staining revealed significant tissue damage in mice treated with M-PG nanosheets, characterized by epithelial disruption, inflammatory cell infiltration, and localized necrosis (Supplementary Fig. 20 and 21). In contrast, PG-based nanomaterials exhibited superior biocompatibility, with no observable histopathological abnormalities in these tissues or other major organs (heart, liver, spleen, lungs, and kidneys). Consistent with these findings, serum biochemistry analysis47 demonstrated that hepatic function markers (ALT, AST, AKP), renal function indicators (CR), and cardiac enzymes (CK, CK-MB) remained within normal physiological ranges in mice treated with PG-NS, PG-NSA, and PG-NPA (Supplementary Fig. 22 and 23). Our findings highlight the efficacy of PG-NSA, as multifunctional organic nanosheets, in modulating excessive inflammation and promoting epithelial cell proliferation in a RIM mouse model. The histological and immunohistochemical results suggest these treatments effectively mitigate radiation-induced tissue damage, offering potential therapeutic strategies for managing recalcitrant RIM.
PG-NSA organic nanosheets mitigate inflammation and NETs in a RIM model
To investigate the therapeutic efficacy of functional nanosheets on post-radiation nasal mucositis, we quantified the relative abundance of sEVs in nasal lavage fluid (NALF) collected from experimental mice. The RT + PG-NSA-treated group showed around 35% reduction in sEV level compared to the RT group, indicating effective clearance of post-radiation sEVs (Fig. 7c). Histopathological analysis of the nasal mucosa revealed significant radiation-induced thinning of the epithelial layer in RT-treated mice compared to saline controls. Notably, PG-NSA treatment substantially preserved mucosal thickness compared to RT alone, with ~2-fold greater relative mucosa thickness than RT controls (Supplementary Fig. 25). The harvested nasal mucosa was further tested to quantify the expression of pro-inflammatory cytokines by qRT-PCR. Compared to the RT mice, the PG-NSA-treated group demonstrated substantial decreases in cytokine expression, with reductions of around 84% for IL-1β, around 86% for IL-6, around 90% for IL-8, and around 78% for TNF-α (Fig. 7d). In contrast, the anti-inflammatory performance of PG-NS and PG-NPA was much less effective (Fig. 7d), indicating the sEVs targeting effect endowed by the CD63 aptamer and 2D nanoscale geometry. The expression results were also substantiated by the cytokine levels in NALF of experimental mice determined by ELISA (Supplementary Fig. 24), which corroborated the qRT-PCR results.
Further validation was provided through immunostaining of nasal mucosa sections for IL-6, TNF-α, and CitH3 (Fig. 7e and Supplementary Fig. 27)13. The RT group displayed significantly elevated expression of IL-6 and TNF-α, with increases of ~5.7-fold and ~7.1-fold, respectively, relative to the saline-treated control group (Fig. 7f and supplementary Fig. 26). Quantitative analysis showed that PG-NSA treatment reduced IL-6 and TNF-α levels by around 68% and 71%, respectively, outperforming PG-NS and PG-NPA in mitigating inflammation. In addition, Ly6G (a neutrophil marker) and CitH348 were also significantly elevated in the RT group compared to the saline group, respectively (Fig. 7f and supplementary Fig. 27). Reduced Ly6G and CitH3 immunostaining positive areas were noted in the RT + PG-NSA group to around 40%. The decrease ratio is higher than PG-NS and PG-NPA-treated RT mice (around 75% and 55%), indicating a reduction in neutrophil infiltration and NET formation. These data collectively demonstrated that the PG-NSA nanosheets effectively attenuated the levels of pro-inflammatory cytokines and NETs in the nasal mucosa by eliminating post-radiation sEVs, thereby conferring a potent anti-inflammatory effect in the context of RIM.
PG-NSA reduces sEVs, oxidative stress, and immune dysregulation in oral mucositis
In addition to post-radiation rhinosinusitis, oral mucositis is a diffuse ulcerative condition usually affecting the non-keratinized oral mucosa, including the soft palate, buccal mucosa, lateral border of the tongue, pharyngeal wall, tonsillar pillars, lips, the middle portion of the tongue, and the floor of the mouth49. To understand the therapeutic function of multifunctional nanosheets, we investigated the pathological changes in the mouse tongues following 25 Gy radiation. Consistent with NALF, elevated sEVs level was also observed in oral lavage fluid (OLF) in the radiated mice. Furthermore, PG-NSA treatment significantly reduced the level of post-radiation sEVs (Fig. 8a). As radiation could induce excessive RONS generation in radiated tissues, we next analyzed oxidative stress in the OLF of experimental mice (Fig. 8a). Local radiation significantly decreased GSH and MDA levels (to 56% and 38%) and increased SOD levels (2-fold) compared to the sham group. PG-NSA treatment increased GSH and MDA levels (1.6-fold and 2-fold) and decreased SOD levels (to 70%) compared to the RT+saline group, confirming the in vivo RONS reduction in radiated tissues.
a The relative sEVs, GSH, SOD, and MDA level in OLF from experimental mice. b t-SNE analysis of CD45+ cells in tongues of experimental mice after treatments, and the M1/M2 ratio of macrophages calculated by flow cytometry. c Representative H&E staining images of the tongues from experimental mice. Scale bars: 250 µm. d Quantification of epidermis thickness in H&E staining images. e, Representative CK5 immunostaining images of the tongues from experimental mice. Scale bars: 100 µm. f Quantification of relative CK5 positive area in immunostaining images. Data represent the mean ± S.D. (n = 6 mice). Statistical significances were assessed by one-way ANOVA (a, b, d, and f), and multiple comparisons post hoc tests were performed via the Prism recommendation (Tukey or Dunnett) method. Abbreviations: OLF: oral lavage fluid; GSH: glutathione; SOD: superoxide dismutase; MDA: malondialdehyde; CK-5: cytokeratin 5. Source data are provided as a Source Data file.
To comprehensively explore the immune cells in radiated tissues, tongues of experimental mice were collected and digested to a single-cell suspension before analysis by flow cytometry50. The immune cell populations in CD45+ cells before and after treatments were analyzed by t-distributed stochastic neighbor embedding (t-SNE) maps (Fig. 8b)50. The detailed hierarchical gating strategy for multi-parametric immunophenotyping was displayed in the Supporting Information (Supplementary Fig. 28). The saline-treated sham group displayed distinct clusters of immune cells, including M1 and M2 macrophages, neutrophils, dendritic cells, T lymphocytes, and other CD45+ immune cells. M1 macrophages (classically activated) are pro-inflammatory and play a central role in host defense against infection, while M2 macrophages (alternatively activated) are associated with anti-inflammatory responses and tissue remodeling, representing two extremes of macrophage activation51. Radiation altered the distribution of these populations, particularly increasing the M1/M2 ratio of macrophages (from 0.43 ± 0.08 to 2.14 ± 0.44), indicating the pro-inflammatory polarization of macrophages. Treatment with PG-NS, PG-NSA, and PG-NPA mitigated this change, and tongues collected from RT + PG-NSA-treated mice displayed a more balanced M1/M2 ratio (0.66 ± 0.09) similar to the sham group. Consistent with the flow results, anti-CD80 immunostaining images of the tongues of experimental mice (Supplementary Fig. 29) demonstrated that PG-NSA declined around 70% of the infiltration of M1 macrophages in radiated tissues, evidenced by a significant reduction in CD80+ cells52.
Histological analysis involved H&E staining of the tongue tips and dorsal surfaces to assess tissue morphology53 (Figs. 8c, 7d, and supplementary Fig. 30). Compared to the saline sham group, the RT+saline group exhibited noticeable epidermal thinning (thickness reduced to around 39%) and tissue architecture disruption. Treatments with PG-NSA and PG-NPA mitigated these morphological changes and preserved structural integrity, indicating a protective effect. A parallel study of epithelial integrity with cytokeratin-5 (CK-5) and cytokeratin-13 (CK-13) immunostaining was performed on the tongues of the experimental mice to evaluate epithelial tissue integrity and health (Figs. 8e, 7f, and supplementary Fig. 31). Radiation exposure caused damage to rapidly proliferating cells, including epithelial cells54. We observed decreased CK-5 and CK-13 staining for the radiated mice, and PG-NSA treatment successfully restored the epithelial integrity, further confirming the importance of targeted sEVs scavenging to alleviate post-radiation mucositis. TdT-mediated dUTP Nick-End Labeling (TUNEL) was also performed to detect DNA fragmentation resulting from apoptotic signaling cascades53 (Supplementary Fig. 32). Very few darkly stained nuclei were observed in TUNEL images of the PG-NSA group, indicating that the protective effect outperformed other groups.
Organic nanosheets reverse transcriptomic and metabolic changes in RIM
To comprehensively evaluate the therapeutic efficacy of functional organic nanosheets on RIM, transcriptomic profiling via RNA sequencing was performed on tongue tissue specimens from three experimental cohorts: control (saline-treated), RT, and radiation with PG-NSA intervention (RT + PG-NSA)55. Principal component analysis (PCA) of the generated gene expression matrix (GEM) demonstrated distinct transcriptomic clustering patterns across treatment groups (Supplementary Fig. 33). The red RT group exhibited clear separation along the negative PC1 axis, while the blue saline control clustered predominantly in the positive PC1 region, indicating fundamental transcriptomic divergence between these conditions. Notably, the RT + PG-NSA cohort (green) displayed an intermediate distribution pattern with partial overlap with the saline control group, primarily in the positive PC1 quadrants. Differential expression analysis revealed distinct gene expression patterns across treatment groups, as visualized by UpSet plot quantification (Fig. 9a). The analysis identified 1,435 DEGs (742 + 693) in the saline vs. RT comparison and 1,197 DEGs (742 + 455) in the RT vs. RT + PG-NSA comparisons, with 742 genes shared between these comparisons, indicating significant bidirectional regulation. Notably, only 21 DEGs were identified in the saline vs. RT + PG-NSA comparison, suggesting substantial normalization of gene expression following PG-NSA intervention. Hierarchical clustering analysis of DEGs demonstrated that the RT + PG-NSA expression profile more closely resembled the saline control pattern than the RT group, confirming that PG-NSA treatment reversed radiation-induced transcriptomic perturbations (Fig. 9b). Volcano plot visualization (Fig. 9c) revealed numerous significantly dysregulated genes in the saline vs. RT comparison, with both upregulated (red dots) and downregulated (blue dots) genes. Similar patterns of differential expression were observed in the RT vs. RT + PG-NSA comparison, while the saline vs. RT + PG-NSA comparison showed markedly fewer DEGs. This comparative analysis quantitatively demonstrates the efficacy of PG-NSA in normalizing radiation-induced transcriptomic dysregulation.
a UpSet plot and Venn map of DEGs across the saline group, RT group, and RT + PG-NSA group. b Heat map depicting the expression profiles of the top 200 DEGs across the saline, RT, and RT + PG-NSA groups. c Volcano plots comparing DEGs between saline vs. RT, RT vs. RT + PG-NSA, and saline vs. RT + PG-NSA groups, red and blue points indicate significantly upregulated and downregulated DEGs. d KEGG pathway enrichment analysis of DEGs between the RT and RT + PG-NSA groups, performed using the hypergeometric test, with bubble size proportional to DEG count and color intensity reflecting adjusted p-value (<0.05). e GO biological process enrichment analysis of DEGs between the RT and RT + PG-NSA groups, conducted using the hypergeometric test, with bar length indicating gene ratio and color intensity reflecting adjusted p-value (<0.05). f Heat map of DEGs associated with radiation-induced epithelial damage, selected based on involvement in epithelial integrity and repair pathways, across the saline, RT, and RT + PG-NSA groups. g Heat map of DEGs associated with immune response, selected based on involvement in cytokine signaling and inflammatory pathways, across the saline, RT, and RT + PG-NSA groups (n = 3 independent experiments). Abbreviations: DEGs: Differentially Expressed Genes; KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: Gene Ontology.
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis revealed that radiation treatment significantly dysregulated multiple critical metabolic and signaling pathways. Analysis of differentially expressed genes between saline and RT groups identified enrichment in pathways including mucin-type O-glycan biosynthesis, glycosphingolipid biosynthesis, and calcium signaling pathways (Supplementary Fig. 34), indicating substantial metabolic perturbation following radiation exposure. Subsequently, comparative analysis between RT and RT + PG-NSA groups revealed that PG-NSA treatment significantly modulated these radiation-disrupted pathways. The most significantly enriched pathways following PG-NSA intervention included mucin-type O-glycan biosynthesis, glycosphingolipid biosynthesis, and calcium signaling pathway, all of which are essential for maintaining cellular homeostasis under stress conditions (Fig. 9d). Protein processing in the endoplasmic reticulum (ER) supports proper folding and quality control of newly synthesized proteins56, preventing radiation-induced misfolding that can trigger apoptotic cascades57. N-glycan biosynthesis is equally crucial, as it stabilizes glycoproteins involved in cellular signaling and immune modulation—key processes for recovery from epithelial damage58, while mucin-type O-glycan biosynthesis contributes to mucosal barrier integrity and repair59. Additionally, Gene Ontology (GO) analysis revealed that radiation significantly perturbed multiple biological processes essential for cellular homeostasis and tissue integrity, with prominent dysregulation in protein glycosylation, protein O-linked glycosylation, tissue homeostasis, and regulation of membrane potential pathways (Supplementary Fig. 37). Following PG-NSA intervention, GO enrichment analysis demonstrated significant restoration of these disrupted processes (Fig. 9e), particularly protein O-linked glycosylation, various glycosylation processes (protein glycosylation, macromolecule glycosylation), and glycoprotein metabolic/biosynthetic processes. These pathways regulate protein folding, trafficking, and cellular function under radiation-induced stress60.
Gene Set Enrichment Analysis (GSEA) demonstrated that PG-NSA treatment significantly attenuated neutrophil extracellular trap formation while enhancing tight junction pathway activation (Supplementary Fig. 35), indicating reduced inflammatory damage and improved epithelial barrier function. Correspondingly, GSEA of GO terms revealed enhanced morphogenesis of epithelium and normalized adaptive immune response following PG-NSA intervention (Supplementary Fig. 38). Heatmap analysis confirmed that PG-NSA treatment reversed radiation-induced downregulation of epithelial function-related genes (Fig. 9f) while concurrently mitigating the aberrant upregulation of immune response genes observed post-radiation (Fig. 9g). The transcriptomic data, supported by reactome analyses (Supplementary Figs. 36, 39, and 40), collectively demonstrate that PG-NSA effectively normalizes radiation-induced transcriptomic alterations, suggesting its potential as a therapeutic intervention for radiation-induced mucositis.
Discussion
This investigation demonstrates the successful engineering of the truly “inorganic-free” nanosheet platform (PG-NSA) capable of selectively adsorbing radiation-induced sEVs with high efficiency. The functionalized two-dimensional nanoplatforms exhibited significant efficacy in attenuating RIM through multiple mechanisms: (1) sequestration of post-radiation sEVs, thereby preventing excessive activation of the cGAS-STING pathway as evidenced by downregulation of downstream effectors (IRF3); The selective elimination of radiation-induced sEVs by PG-NSA likely prevents the cytosolic delivery of immunogenic DNA fragments to recipient cells, a process known to trigger cGAS dimerization and subsequent 2'3'-cyclic GMP-AMP (cGAMP) synthesis, thereby attenuating the STING-TBK1-IRF3 signaling cascade and reducing type I interferon-driven inflammation61. (2) maintenance of innate immune homeostasis via modulation of pro-inflammatory cytokine production; PG-NSA treatment was associated with a significant reduction in the expression of pro-inflammatory cytokines, including TNF-α, IL-1β, and IL-6, which are hallmark mediators of radiation-induced tissue damage, thereby mitigating the excessive inflammatory response that exacerbates RIM. (3) regulation of oxidative stress parameters through RONS scavenging activity; The capacity of PG-NSA to scavenge RONS is likely attributable to its organic composition, which may facilitate direct neutralization of these reactive molecules, thereby reducing oxidative damage to cellular components and preventing apoptosis in irradiated tissues. and (4) antibacterial properties, potentially contributing to microbiota restoration in irradiated tissues.
Comprehensive toxicological profiling indicates that PG-NSA exhibits superior biocompatibility compared to conventional inorganic-based nanosheets, with no significant elevation in hepatic enzymes, renal function markers, or histopathological abnormalities at therapeutic doses. Inorganic nanoparticles can interact with the immune system and disrupt normal immune function, raising safety concerns62. The absence of inorganic components in PG-NSA likely reduces the risk of long-term tissue accumulation and associated toxicities, such as chronic inflammation, thereby enhancing its suitability for systemic therapeutic applications. The biodistribution analysis suggests favorable pharmacokinetic properties for potential clinical translation.
RNA-sequencing and subsequent pathway analyses revealed that PG-NSA treatment significantly reversed radiation-induced transcriptomic alterations. Gene set enrichment analysis identified statistically significant modulation of critical cellular processes, including protein processing in the endoplasmic reticulum, N-glycan biosynthesis, and mucin-type O-glycan biosynthesis. The upregulation of protein processing in the endoplasmic reticulum and glycan biosynthesis pathways by PG-NSA suggests an enhancement of cellular proteostasis and glycoprotein-mediated signaling, which collectively bolster epithelial barrier integrity and facilitate tissue repair following radiation-induced stress. These pathways are integral to cellular stress response mechanisms, maintenance of epithelial barrier integrity, and tissue repair processes following radiation injury.
The dual immunomodulatory properties of PG-NSA—functioning both as an anti-inflammatory agent by RONS scavenging and as a regulator of innate immune responses through STING pathway modulation—suggest potential applications beyond RIM therapy. The combined antioxidant and STING-modulatory effects of PG-NSA position it as a potential candidate for treating other conditions driven by oxidative stress and aberrant innate immune activation, such as neurodegenerative diseases or chronic inflammatory disorders63, pending rigorous validation in relevant preclinical models. However, additional investigations in disease-specific models would be necessary to validate efficacy in other inflammatory conditions, such as severe viral infections or transplant-related complications.
This study represents an advancement in organic nanomaterial development for therapeutic applications, circumventing potential long-term toxicity concerns associated with inorganic nanostructures. The superior biosafety profiles of inorganic-free nanomaterials (PG-NS, PG-NSA, and PG-NPA) over their inorganic counterparts (M-PG) position them as potential candidates for clinical translation, particularly for managing RIM. The observed biocompatibility aligns with our objective to develop safer alternatives for therapeutic use, putting forward a point for further exploration of organic nanosheets in modulating inflammation and promoting tissue repair. Nevertheless, several limitations must be acknowledged, including the need for larger animal studies, investigation of long-term immunological memory effects, and optimization of manufacturing scalability (e.g., optimizing time and cost of template synthesis and removal) before clinical translation can be realized.
In summary, the engineered PG-NSA platform demonstrates promising therapeutic potential for RIM through multiple complementary mechanisms with a favorable safety profile. These findings warrant further preclinical development and potential Phase I safety studies to establish this platform as a viable therapeutic option for radiation-induced tissue injury and potentially other inflammatory conditions.
Methods
Materials
Molybdenum sulfide (MoS2) powder, n-butyllithium solution (2.0 M in cyclohexane), lipoic acid (LA), 1-ethyl-3-(3-dimethyllaminopropyl)carbodiimide hydrochloride (EDC.HCl), N-hydroxysuccinimide (NHS), glutamate (Glu), Horseradish Peroxidase (FRP), poly(lactic-co-glycolic acid (PLGA) (Mw = 10,000; 50:50), Bovine serum albumin (BSA), 2',7'-Dichlorodihydrofluorescein diacetate (DCFH-DA), Lipopolysaccharide (LPS), and H2O2 were purchased from Sigma-Aldrich. Quant-iT PicoGreen DNA assay kit, sEVs-depleted serum, QUANTI-Blue™ for alkaline phosphatase detection, 4',6-diamidino-2-phenylindole (DAPI), n-hexane, and TRIzol reagent were purchased from Fisher Scientific. Methanol, ethanol, N,N-dimethylformamide (DMF) and tetrahydrofuran (THF), N-Methyl pyrrolidone (NMP), dimethyl sulfoxide (DMSO), triethylamine (TEA) and paraformaldehyde (4%) were purchased from Millipore-Sigma (US). CD63 aptamer (5' to 3': HOOC-TAACCACCCCACCTCGCTCCCGTGACACTAATGCTA) was synthesized by Sangon Biotech (Shanghai) Co., Ltd. (Bicyclo[6.1.0]non-4-yn-9-yl)methyl 4-nitrophenyl carbonate (BCN) was synthesized according to the previous literature64. Glutathione (GSH) kit, malondialdehyde (MDA) kit and superoxide dismutase (SOD) kit were provided by Solarbio (Beijing, CHN). GW4869 was bought from MedChemExpress LLC. HS-PEG-NHS (Mw = 1000) was bought from Ruixibio company. Human and Murine IL-1β, IL-6, and TNF-α ELISA kits were purchased from Invitrogen (US). Tubulin, CD63, p-TBK1, p-IRF3, and p-STING antibodies for WB were bought from ABclonal (Wuhan, China). Cell Counting Kit-8 (CCK8) assay was provided by Dojindo Molecular Technologies, Inc., Rockville, MD, USA. Fluorescein isothiocyanate isomer I (FITC), PKH67, and Cyanine 5 (Cy5)-NHS ester were purchased from Lumiprobe Corporation (FL, USA). iScript cDNA synthesis kit and iTaq Universal SYBR Green Supermix were bought from Bio-Rad. Milli-Q water was used in all experiments. The reference bacterial strains Staphylococcus aureus (ATCC 29213) and Escherichia coli (ATCC 25922) were employed as representative Gram-positive and Gram-negative bacteria, respectively, for evaluating antibacterial activity.
Equipment
Zeta potential and dynamic light scattering (DLS) data were measured using a NanoBrook 90Plus PALS (Brookhaven, US) in corresponding conditions. Fourier transform infrared spectroscopy (FTIR) spectra were measured on a Jasco FT/IR-4100 spectrometer. Transmission electron microscopy (TEM) was observed with a FEI Tecnai G2 F30 TEM, and TEM images of sEVs were recorded with uranyl acetate for negative staining. Fluorescence was measured by a JASCOFP-6500 Spectrofluorometer. Quanti-Blue and CCK8 assays were conducted using a full wavelength scanning microplate reader (Varioskan Flash, Thermo Fisher). Flow cytometry was performed using a Beckman Coulter flow cytometer (CytoFLEX S, Beckman Coulter). Quantitative polymerase chain reaction (qPCR) was measured with Applied Biosystems® QuantStudio™ 7 Flex (Thermo Fisher). Confocal laser scanning microscopy (CLSM) experiments were conducted with a Leica TCS SP8 inverted microscope with an A1 scanning confocal unit. The specified color area in CLSM images was calculated by Image J software. H&E and PAS staining slices were scanned using an SQSL-510 automated slice scanning system (Shenzhen Shengqiang Technology Co., Ltd.). Biodistribution fluorescent images were recorded with an IVIS Spectrum system (PerkinElmer, USA).
Image preparation
We confirm that Fig. 3a, b were entirely created by the authors of this manuscript. Every element within these images was originally generated by us without the use of any previously created or third-party elements. The images were constructed using Cinema4D as the modeling software, Redshift as the rendering engine, and Adobe Photoshop for post-processing. No part of the figures was taken from external sources or prior publications. As such, there are no copyright or licensing concerns associated with Fig. 3a, b.
Preparation of M-PG
MoS2 monolayers were produced based on the methods in publications by using lithium anions as intercalation agents7. PG (Mw ≈ 10,000 g/mol) was synthesized by one-pot, ring-opening anionic polymerization (ROAP), according to a reported method in the literature13. PG with amino groups (PGA) was synthesized by a 3-step protocol13.
0.5 mL LA solution (10 mg/mL) was added to 10 mL MoS2 solution (1 mg/mL), followed by a reaction at room temperature for 24 h to prepare LA-modified MoS2 (M-LA). Then, 10 mg M-LA was dispersed with EDC.HCl (0.5 mM) in MES solution (0.2 M), and the solution was then added dropwise to PG (1 g, 0.1 mmol) in MES (0.2 M). The reaction mixture was stirred at room temperature for 2 days before dialysis in Mili-Q water (MWCO = 12-14 K) to obtain PG-covered MoS2 (M-PG).
Synthesis of PG-NS and PG-NSA
HS-PEG-NHS (100 mg) was dissolved in PBS (1 mg/mL) and then mixed with Glu (50 mg) in PBS (1 mg/mL). The reaction was stirred at room temperature for 24 h and then dialyzed (Mw = 1000) in MilliQ for 24 h to obtain HS-PEG-Glu as a crosslinker. HS-PEG-Glu (100 mg) was added to M-PG solution (100 mL, 1 mg/mL) and reacted at room temperature for 24 h before dialysis (Mw = 10,000) in MilliQ for 24 h to obtain M-PG-Glu. Subsequently, M-PG-Glu (100 mg) with EDC (30 mg) was dispersed in MES (100 mL, 0.2 M) and stirred for 24 h before dialysis (Mw = 10,000) in MilliQ for 24 h. Finally, the crosslinked M-PG-Glu was incubated with HRP (50 U/mL) with H2O2 (0.2 mM) at 37 °C for 48 h to degrade the MoS2 backbone to obtain PG-based nanosheets (PG-NS)28.
CD63 aptamer (5 mg) was added to PG-NS (100 mg) with EDC.HCl (30 mg) in MES (100 mL, 0.2 M) and reacted at room temperature for 24 h. The mixture was dialyzed in MilliQ (Mw = 30,000) for 24 h to prepare purified CD63 aptamer-modified PG-NS (PG-NSA).
Synthesis of PG-NPA
PLGA (100 mg) was dissolved in DMF (100 mL) along with EDC·HCl (38.3 mg) and NHS (23 mg, 0.2 mmol). To this solution, PG (50 mg) was added, and the reaction was allowed to proceed at room temperature for 24 h. The mixture was then dialyzed against Milli-Q water (MWCO = 2k) to remove unreacted agents and DMF, followed by lyophilization to yield PG-PL. Subsequently, PG-PL (200 mg) was dissolved with CD63 aptamer (10 mg) and EDC·HCl (76.6 mg, 0.4 mmol) in MES buffer (0.2 M). The solution was stirred at room temperature for 24 h, dialyzed against Milli-Q water (MWCO=2k) to remove unreacted agents, and lyophilized to obtain PG-PLA. For the preparation of PG-NPA, 20 mg of PG-PLA was dissolved in 2 mL DMSO and added dropwise to 18 mL Milli-Q water. Finally, the DMSO was removed by dialysis against Milli-Q water (MWCO=2k) for 24 h, yielding the PG-NPA solution.
Synthesis of PEI-NSA and PG-NSA-C
PEI-based nanosheets (PEI-NS) were prepared using a synthetic protocol similar to PG-NSA. PEI (Mw=10k) was modified onto the MoS2 nanosheets via amidation reaction after LA was decorated. Afterward, HS-PEG-Glu was anchored onto the nanosheets before crosslinking was performed in MES (0.2 M) with EDC as a condensing agent. Subsequently, the crosslinked nanosheets were incubated with HRP with H2O2 at 37 °C for 48 h to degrade the backbone to obtain PEI-NS. Finally, CD63 aptamer was added to PEI-NS with EDC.HCl in MES and the mixture was dialyzed in MilliQ (MWCO = 30 kD) for 24 h to prepare purified aptamer-modified PEI-NS (PEI-NSA).
PG-NSA-C was prepared via a method based on click chemistry65. PG-NS (10 mg) was dispersed in DMF (10 mL), including TEA (10 µL), and then BCN (100 µg) was added to the solution65. The mixture was stirred at room temperature overnight, and the solution was dialyzed (MWCO = 10 kD) in PBS (pH 7.4) for 24 h to replace DMF with PBS (pH 7.4). Then, thiolated CD63 aptamers (500 µg) were added to the mixture and reacted at room temperature overnight. Finally, the solution was dialyzed (MWCO = 10 kD) in Milli-Q for 24 h and then lyophilized to obtain PG-NSA-C.
Cytotoxicity test
RPMI 2650 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) with 10% Fetal Bovine Serum (FBS) and 1% Penicillin-Streptomycin (PS) at 37 °C in a humidified atmosphere with 5% CO2. The cytotoxicities of the PG, M-PG, PG-NS, PG-NSA, PG-NPA, and Apt against RPMI 2650 cells were evaluated with the CCK-8 assay. Firstly, RPMI 2650 cells (5 × 103 cells/well) were seeded in a 96-well plate and cultured until the cell density reached 60%. After that, the culture medium was replaced by PG, M-PG, PG-NS, PG-NSA, PG-NPA, and Apt solution in DMEM with different concentrations. After 24 h or 48 h of incubation, the culture medium was replaced again with fresh DMEM containing 10% CCK-8 reagent. Finally, the plates were incubated for another 2–3 h at 37 °C, and then the absorbance (450 nm) of the wells was measured using a Multiplate Reader. The cytotoxicity of PG, M-PG, PG-NS, PG-NSA, PG-NPA, and Apt was calculated by comparing the absorbance (450 nm) value to the cells treated with medium-only.
Isolation of sEVs and large EVs
RPMI 2650 cells (1.8 × 106 cells/dish, BeNa Culture Collection, No. 353694) were seeded and exposed to gamma irradiation the next day with doses of 12.5 Gy and 25 Gy, respectively. After irradiation, the cell medium was changed to DMEM with 10% FBS (deactivated, exosome-depleted) and 1% Penicillin-Streptomycin. The above medium was collected after a 4-day incubation, and post-radiation sEVs were isolated by ultracentrifugation. Briefly, the radiation medium was collected by centrifugation at 300 × g (10 min) to remove intact cells, 2000 × g (10 min) to remove dead cells, and 10,000 × g (30 min) to remove cell debris. The solution was then centrifuged at 10,000 × g (30 min), and the precipitate was collected as large EVs. At the same time, the supernatants were collected and centrifuged at 100,000 × g (70 min) to obtain the sEVs in pellets. The sEVs and large EVs were washed with PBS, and the protein concentration was determined with a Pierce BCA Protein Assay Kit (Thermo Fisher Scientific, USA).
In the other experimental group, sEVs were isolated via density-gradient centrifugation. Briefly, the collected cell medium was centrifuged at 300 × g (10 min) to remove intact cells, 2000 × g (10 min) to remove dead cells, and 10,000 × g (30 min) to remove cell debris. The concentrated solution was then added to an ultracentrifuge tube with a 30% sucrose cushion (density, 1.210 g/cm3) at the bottom of the tube, followed by ultracentrifugation at 100,000 × g at 4 °C for 70 min. Subsequently, the cushion was collected and washed with PBS, and sEVs were concentrated.
sEVs binding assay
sEVs were labeled with PKH67 dye66, and a standard curve for fluorescent intensity versus sEVs concentration was calculated. PKH67-labeled sEVs (1 μg/mL) were mixed with different w/w ratios (1 or 2) of PG-NS, M-PGA, PG-NSA, or PG-NPA in PBS (7.4). The solutions were incubated at 37 °C for 30 min before centrifuging at 10,000 × g (10 min). The supernatant was collected, and the sEVs concentration was determined by fluorescence intensity (Ex = 485 nm, Em = 520 nm). The sEVs binding capacity was calculated based on the decreased sEVs in the supernatant.
In another experiment, PG-NS, M-PGA, PG-NSA, or PG-NPA were labeled with Cy5 dye and incubated with PKH67-labeled sEVs at 37 °C for 30 min. Then, the colocalization of sEVs and nanomaterials was observed by CLSM (PKH67: Ex = 485 nm, Em = 520 nm; Cy5: Ex = 630 nm, Em = 670 nm) and quantitatively analyzed by ImageJ software.
Inhibition of the sEVs-induced STING activation
HEK-blueTM STING (HEK-STING) reporter cell line (InvivoGen, San Diego, CA) was cultured according to the manufacturer’s protocol. HEK-STING cells were seeded into 96-well plates (8 × 104 cells/well), and the cells were attached to the bottom after 6-8 h of incubation. After that, sEVs, sEVs+PG-NS, sEVs+M-PGA, sEVs+PG-NSA, and sEVs+PG-NPA were added to the wells and incubated for the following 24 h. sEVs were isolated from the medium of RPMI 2650 cells after 25 Gy radiation, and the concentration was set as 1 μg/mL. The concentration of nanomaterials was 2 μg/mL. The cells incubated with medium-only or sEVs-only were considered negative control and positive control, respectively. Finally, 50 μL supernatants of each well were harvested and mixed with 150 μL QUANTI-BlueTM solution. The new plate was incubated at 37 °C for 2 h, and STING activation was determined by the absorbance (620 nm) value measured using a Multiwell Plate Reader.
In WB studies, RAW 264.7 cells (Beyotime Biotechnology, No. C7505) were treated with sEVs alone (1 μg/mL) or in combination with PG-NS, M-PGA, PG-NSA, or PG-NPA (2 μg/mL) for 24 h. Total protein was extracted from the cells and quantified using a BCA Protein Assay Kit (Pierce). For each sample, 10-30 μg of total protein was mixed with NuPAGE LDS Sample Buffer and NuPAGE Sample Reducing Agent, heated at 70 °C for 10 minutes, and then subjected to electrophoresis on a NuPAGE Novex 4-12% Bis-Tris gel. The separated proteins were transferred to a polyvinylidene difluoride (PVDF) membrane (Bio-Rad) at 100 V for 90 minutes. The membrane was blocked with 5% bovine serum albumin (BSA) in Tris-buffered saline with Tween 20 (TBST) for 1 h at room temperature, followed by overnight incubation at 4 °C with primary antibodies against phosphorylated IRF3 (p-IRF3), phosphorylated TBK1 (p-TBK1), phosphorylated STING (p-STING), and tubulin (as a loading control) under gentle agitation. After washing with TBST, the membrane was incubated with appropriate secondary antibodies for 1 h at room temperature. Protein bands were visualized using Clarity Western ECL Substrate (Bio-Rad), and images were captured and analyzed using the ChemiDoc Imaging System with Image Lab 5.1 software (Bio-Rad).
sEVs-induced NET formation
Human peripheral blood neutrophils were isolated from RIM patients with a MACSxpress® neutrophil isolation kit (Stem Cell, Canada)69. The neutrophils were cultured in a medium with sEVs, sEVs+PG-NS, sEVs+PG-NSA, and sEVs+PG-NPA and incubated at 37 °C for 4 h. sEVs were isolated from the medium of radiated RPMI 2650 cells, and neutrophils incubated with medium-only were applied as the negative control (NC) group. The concentration of sEVs was 1 μg/mL, and the concentration of nanomaterials was 2 μg/mL. Subsequently, DAPI staining and CitH3 immunostaining were applied to identify and quantify the NET formation. The NET formation was observed using CLSM and calculated by ImageJ software.
Cellular co-localization of sEVs and nanomaterials
RAW 264.7 cells (2 × 104 cells/well) were seeded on the confocal slide inside a 24-well plate and incubated at 37 °C for 24 h. Then, PKH67-labeled sEVs (1 μg/mL) and Cy5-labeled nanomaterials (5 μg/mL) were added, and the cells were incubated for another 24 h. The concentration of sEVs was 1 μg/mL, and the concentration of nanomaterials was 2 μg/mL. Subsequently, the supernatant was discarded, and cells were washed with PBS before the medium containing LysoTrackerTM Red (Thermo Fish, USA) was added. The cells were incubated at 37 °C for 20 min before fixation with 4% paraformaldehyde at room temperature for 30 min. The nucleus was then stained with DAPI at room temperature for 20 minutes in the dark. The film was then sealed with neutral gum, and the fluorescent images were taken by CLSM (DAPI: Ex=360 nm, Em=460 nm; PKH67: Ex=488 nm, Em=520 nm; Cy5: Ex=630 nm, Em=670 nm).
sEVs-induced M1 polarization of macrophages
RAW 264.7 cells (5 × 104 cells/well) were seeded in a 12-well plate and incubated at 37 °C for 24 h. The next day, the cells were treated with sEVs, sEVs+PG-NS, sEVs+PG-NSA, and sEVs+PG-NPA for 24 h. sEVs were isolated from the medium of radiated (25 Gy) RPMI 2650 cells. The concentration of sEVs was 1 μg/mL, and the concentration of nanomaterials was 2 μg/mL. The cells were harvested and washed with FACS buffer and stained with antibodies to CD11b, F4/80, and CD86. Finally, the cells were analyzed for M1 polarization using flow cytometry (Beckman), and the resulting data were analyzed with FlowJo software.
Intracellular RONS detection
RPMI 2650 cells were seeded into 96-well plates (black) and allowed to attach to the bottom for 4-6 h. Subsequently, the cells were stimulated with LPS (10 μg/mL) for 24 h, followed by co-culture with nanomaterials of different concentrations for 3 h. Finally, the DCFH-DA was added and incubated for 30 min. The fluorescent intensity was measured by a Multiwell Platereader (Ex = 495 nm; Em = 525 nm), and fluorescent images were recorded by CLSM.
Determination of antioxidant capacity
For the DPPH· reduction studies, DPPH solution (0.1 mM) was prepared with anhydrous ethanol. Subsequently, 1 mL of nanomaterials with different concentrations was incubated with 1 mL DPPH solution at room temperature under gentle shaking for 30 min, followed by centrifugation at 2800 × g for 10 min. The absorption value (517 nm) of the supernatant was then measured, and DPPH mixed with an equal volume of MilliQ water was set as the control group.
For the ·OH clearance studies, salicylic acid solution (9 mM), FeSO4 solution (9 mM), and H2O2 solution (8.8 mM) were first prepared. Then, 30 μL of the three solutions and nanomaterials with different concentrations were added to the tubes, and the volume was filled to 450 μL with MilliQ water. A negative control group without H2O2 and a positive control group without nanomaterials were established, and both were supplemented with MilliQ water to reach 450 μL. The reactions were incubated in a water bath at 37 °C for 15 min before measuring the absorbance at 510 nm.
Antibacterial studies
E. coli was cultured in Luria-Bertani (LB) medium, and S. aureus was cultured in Nutrient Broth (NB) medium. Single colonies of each strain were selected for amplification, then diluted and co-cultured with PG, PG-NS, PG-NSA, and PG-NPA (50 μg/mL) for 24 h. The relative number of bacterial colonies in suspension was quantified by the absorbance value (600 nm) using a Nanodrop. Additionally, the bacterial suspension after incubation with nanomaterials for 24 h was spread on agar plates and photographed by light microscopy.
Clinical sample collection and analysis
20 RIM patients with rhinosinusitis after radiotherapy in the Sixth Affiliated Hospital and First Affiliated Hospital of Sun Yat-sen University were included in this study. Nasal secretions were collected with an expansive sponge from these patients before and after radiation, which was approved by the ethics committee of Sun Yat-sen University (2024ZSLYEC-185). All patients were informed of the purpose of the donated nasal secretion samples and signed informed consent forms.
The nasal secretions were centrifuged at 100 × g (10 min), 2000 × g (10 min), and 10,000 × g (30 min) to remove intact cells, dead cells, and cell debris, respectively. Finally, sEVs were collected in the pellets after ultracentrifugation at 100,000 × g (70 min). The number, size, and morphology of sEVs were characterized by NTA, DLS, and TEM. The protein composition in sEVs was analyzed by WB, and the dsDNA level inside sEVs was determined by pico-green assay.
Establishment of the RIM mouse model
Animal experiments were approved by the ethics committee of the Sixth Affiliated Hospital, Sun Yat-sen University (No. IACUC-2023081102). Adult female BALB/c mice (4 to 6 weeks, weight 20 ± 2 g) were bought from YaoKang company (Guangdong). All mice were raised in a specific pathogen-free (SPF) environment at Sixth Affiliated Hospital, Sun Yat-sen University.
RIM model mice were established according to the previous report. BALB/c mice were radiated (25 Gy) at the head area to simulate clinical radiotherapy for head and neck tumors. BALB/c mice treated with saline were used as sham mice.
GW4869 and sEVs treatment for model mice
RIM model mice and sham mice were established according to the above protocol. In the GW4869 + RT group, GW4869 (10 μg) was locally administered 8 h before radiation. In the RT+sEVs group, sEVs (20 μg) were locally administrated three times at 4 h, 2 d, and 4 d after radiation. The sEVs were isolated from the medium of radiated (25 Gy) mouse epithelial cells. The sham mice and RIM model mice treated with saline (50 μL) were considered negative controls and positive controls, respectively.
All of the experimental mice were sacrificed on the 7th day after radiation. The nasal mucosa of the experimental mice was collected, and the total mRNA was extracted using TRIzol. Then the mRNAs were converted to cDNA with an iScript cDNA synthesis kit before the iTaq Universal SYBR Green Supermix was used to conduct qRT-PCR. All reactions used standard cycling conditions with a melting temperature of 60 °C, and Ct values for cytokines (IL-1β, IL-6, IL-8, and TNF-α) were normalized to GAPDH. The primers for these cytokines were designed based on the PrimerBank database and obtained from Integrated DNA Technologies, Inc. (Table S2).
In the other groups, the mice were sacrificed by decapitation on the 7th day after radiation, the heads were quickly dissected to isolate the anterior part of the snout, including the nasal cavities. Subsequently, the maxillary bones (MBs) were decalcified in 10% EDTA for 3 weeks before fixation in 10% paraformaldehyde for 24 h. Finally, the specimens were sequentially rinsed, dehydrated, and embedded in paraffin before the sections were cut, dewaxed, and analyzed with Ly6G and CitH3 immunostaining.
Biodistribution of PG-NS, PG-NSA, and PG-NPA
Sham mice and RIM model mice were intranasally (i.n.) and intraorally (i.o.) instilled with saline, PG-NS-Cy5 (100 μg), PG-NSA-Cy5 (100 μg), or PG-NPA-Cy5 (100 μg) in 50 μL at 4 h after radiation. The experimental mice were sacrificed, and then MBs, tongues, as well as major organs (hearts, lungs, livers, spleens, and kidneys), were harvested at 1 d, 3 d, and 7 d after treatment. The isolated organs were imaged by an in vivo imaging system (Ex = 630 nm; Em = 670 nm) to study the biodistribution of nanomaterials. At the same time, the mice serum was collected, and the fluorescence intensity was measured to calculate the content of nanomaterials in the blood.
RIM mice treatment with PG-NS, PG-NSA, and PG-NPA
RIM model mice were established, and the mice were divided into the saline group, RT+saline group, RT + PG-NS group, RT + PG-NSA group, and RT + PG-NPA group (n = 6). PG-NS (100 μg), PG-NSA (100 μg), and PG-NPA (100 μg) in 50 μL of saline were instilled (i.n. and i.o.) three times at 4 h, 2 d, and 4 d after radiation. On the 3rd day after the last treatment, the experimental mice were sacrificed, and NALF and OLF were collected. Intact cells and cell debris were removed from NALF and OLF by sequential centrifugation at 2000 × g (10 min) and 10,000 × g (30 min). Then, sEVs content in NALF and OLF was determined by NTA, and the levels of cytokines (IL-1β, IL-6, and TNF-α) in NALF were measured with corresponding ELISA kits. In another group of mice, the nasal mucosa and tongues were collected, and the total mRNA was extracted and converted to cDNA with the corresponding kits. After that, qRT-PCR was performed to determine the Ct values for cytokines (IL-1β, IL-6, IL-8, and TNF-α), and the values were normalized to GAPDH. The gene sequences of each cytokine in the qRT-PCR experiments are listed in the Supporting Information (Table S2). The GSH, MDA, and SOD levels in the OLF of mice were measured following the corresponding protocols.
Analysis of nasal mucosa inflammation
The experimental mice were sacrificed 24 h after treatment. Then, the mice were decapitated, and MBs were collected after dissection to isolate the anterior part of the snout and the nasal cavities. Subsequently, the MBs were decalcified in EDTA solution for 3 weeks after fixation in paraformaldehyde (4%) for 24 h. After that, the specimens were carefully rinsed in water and dehydrated in dry ethanol before being embedded in paraffin. Finally, the sections were cut, dewaxed, and analyzed with H&E staining and Ly6G, CitH3, IL-6, and TNF-α immunostainings.
Analysis of tongue inflammation
After experimental mice were sacrificed, tongues were collected and fixed in paraformaldehyde (4%) for 24 h. Then, H&E and TUNEL staining and CD80, CK5, and CK13 immunostainings were applied to analyze the epithelial damage and immune cell infiltration in the lungs. The tongue tissue was gently ground and filtered through a 40 μm cell sieve, followed by the removal of red blood cells using an RBC lysis solution to obtain a single-cell suspension. Subsequently, the cell types and inflammatory status were analyzed via flow cytometry using surface (or intracellular) markers.
The tongue tissue was gently ground and filtered through a 40 μm cell sieve, followed by the removal of red blood cells using an RBC lysis solution to obtain a single-cell suspension. Subsequently, the cell types and status were analyzed via flow cytometry using surface (or intracellular) markers. The antibodies utilized in flow cytometry include Live/dead, CD45, CD3, CD11b, CD11c, Ly6G, MHC II, F4/80, CD86, and CD206. All flow cytometric analyses were performed using a Beckman Coulter flow cytometer (CytoFLEX S, Beckman Coulter) with standardized voltage settings across experimental replicates, and data were processed using FlowJo v10.8.1 software (BD Biosciences) with consistent gating parameters applied across all samples.
The detailed information for the antibodies used in this study: 1. Cytokeratin 5 Polyclonal antibody, Proteintech, 28506-1-AP (diluted to 1/500); 2. Cytokeratin 13 Rabbit Polyclonal Antibody, Proteintech, 10164-2-AP (diluted to 1/500); 3. Rabbit polyclonal antibody to EGFR, Affinity Biosciences, AF6043-50 (diluted to 1/500); 4. Anti-Histone H3 Antibody, BOSTER, BM4316 (diluted to 1/100); 5. Anti-TSG101 Antibody, Abmart, PS06850S (diluted to 1/100); 6. CD63 Rabbit mAb, Zen BioScience, R23327 (diluted to 1/1000); 7. GAPDH Rabbit mAb, Zen BioScience, R380626 (diluted to 1/10000); 8. CD9 Rabbit pAb, Zen BioScience, 382758 (diluted to 1/1000); 9. STING Rabbit pAb, Zen BioScience, 300415 (diluted to 1/1000); 10. Phospho-NAK/TBK1 (Ser172) Rabbit mAb, Zen BioScience, R30260 (diluted to 1/1000); 11. Anti-Phospho-IRF3(S386) Antibody, huabio, ET1608-22 (diluted to 1/1000); 12. Phospho-STING/TMEM173-S366 Rabbit pAb, Abclonal, AP1223 (diluted to 1/1000); 13. Purified anti-mouse CD16/32 Antibody, Biolegend, 101302 (diluted to 1/50); 14. FITC anti-mouse CD45 Antibody, Biolegend, 103108 (diluted to 1/200); 15. PerCP/Cyanine5.5 anti-mouse CD3 Antibody, Biolegend, 100218 (diluted to 3/100); 16. Alexa Fluor® 700 anti-mouse CD3 Antibody, Biolegend, 100216 (diluted to 1/50); 17. Brilliant Violet 605™ anti-mouse/human CD11b Antibody, Biolegend, 101257 (diluted to 1/100); 18. Alexa Fluor® 700 anti-mouse Ly-6G Antibody, Biolegend, 127621 (diluted to 1/200); 19. Brilliant Violet 421™ anti-mouse I-A/I-E Antibody, Biolegend, 107632 (diluted to 1/100); 20. PE/Cyanine7 anti-mouse CD11c Antibody, Biolegend, 117318 (diluted to 1/100); 21. PE anti-mouse CD11c Antibody, Biolegend, 117308 (diluted to 1/100); 22. APC anti-mouse F4/80 Antibody, Biolegend, 123116 (diluted to 1/100); 23. APC/Cyanine7 anti-mouse CD86 Antibody, Biolegend, 105030 (diluted to 1/200); 24. PE anti-mouse CD206 (MMR) Antibody, Biolegend, 141706 (diluted to 1/200); 25. Brilliant Violet 650™ anti-mouse CD19 Antibody, Biolegend, 115541 (diluted to 3/100); 26. PE anti-mouse CD170 (Siglec-F) Antibody, Biolegend, 155505 (diluted to 1/100); 27. PE anti-mouse CD40 Antibody, Biolegend, 124609 (diluted to 1/200); 28. APC anti-mouse CD80 Antibody, Biolegend, 104714 (diluted to 1.2/100); 29. APC/Cyanine7 anti-mouse CD4 Antibody, Biolegend, 100414 (diluted to 1.5/100); 30. PE/Cyanine7 anti-mouse CD8a Antibody, Biolegend, 100722 (diluted to 1.25/100); 31. PE anti-mouse IFN-γ Antibody, Biolegend, 505807 (diluted to 1/100); 32. Brilliant Violet 605™ anti-mouse Ki-67 Antibody, Biolegend, 652413 (diluted to 1/100); 33. PE anti-mouse CD366 (Tim-3) Antibody, Biolegend, 134003 (diluted to 1/100).
Transcriptome sequencing
TRIzol reagent was utilized for RNA extraction from tongue tissues of the saline group, RT+ saline group, RT + PG-NS group, RT + PG-NSA group, and RT + PG-NPA group (n = 3). Subsequently, the sample library underwent quality assessment using an Agilent 2100 Bioanalyzer, followed by paired-end (PE) sequencing on these libraries utilizing Next-Generation Sequencing (NGS) based on the Illumina sequencing platform.
Biosafety evaluation of M-PG, PG-NS, PG-NSA, and PG-NPA
The biosafety of M-PG, PG-NS, PG-NSA, and PG-NPA was evaluated with healthy BALB/c mice. M-PG, PG-NS, PG-NSA, and PG-NPA with high dosage (300 μg) were intranasally and intraorally administered to the healthy mice once per 2 days and three times in total. The mice treated only with saline were considered the control. Half of the experimental mice were sacrificed on the 7th day, and the other half were sacrificed on the 14th day after the last treatment. MBs, tongues, and major organs (hearts, livers, lungs, spleens, and kidneys) were collected and fixed in paraformaldehyde (4%) for 24 h. Subsequently, the MBs were decalcified in 10% EDTA at room temperature for 3 weeks, and then the organs were embedded in paraffin, and the tissue slices were prepared. Finally, H&E staining was performed for biosafety evaluation. In addition, the ALT, AST, AKP, CR, CK, and CK-MB levels in the serum of experimental mice were collected to evaluate their hepatic and renal functions after treatment.
Statistical analysis
All statistical analyses were conducted using GraphPad Prism (version 10.4), with data visualization performed in Origin (version 2021). Prior to applying parametric statistical tests, the normality of data distributions was assessed using the Shapiro-Wilk test. For datasets satisfying the normality assumption (p > 0.05), differences between two independent groups with one nominal-level independent variable were evaluated using an unpaired two-tailed Student’s t-test. For comparisons involving three or more groups with a single independent variable, one-way analysis of variance (ANOVA) was conducted, followed by Tukey’s multiple comparisons test for all pairwise comparisons or Dunnett’s test when comparing groups to a control. For designs with two independent variables, two-way ANOVA was performed, with post-hoc tests selected based on the experimental design. All post-hoc p-values reported in figures were adjusted for multiple comparisons using the specific method (Tukey’s or Dunnett’s) recommended by Prism for each analysis. For datasets not meeting the normality assumption, the Wilcoxon Signed-Rank test was used to compare two paired groups. Data are presented as the mean ± standard deviation (S.D.), with error bars representing the S.D. Unless otherwise stated, each mean and S.D. were derived from a minimum of three experimental units or subjects. Sample sizes were not predetermined using statistical methods. All collected data were included in the analyses. Experiments were randomized, and investigators were not blinded to group allocation during experimentation or outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that all the data supporting the results of this study are available within the Article and the Supplementary Information. The primer sequences used in RT-qPCR assays are provided in Supplementary Table 2. The gating strategy for flow cytometry experiments can be found in the Supplementary Information. The RNA sequencing data generated in this study have been deposited in the NCBI BioProject database under accession number PRJNA1252122. Source data for Figs. 1, 2, 4–8, and Supplementary Figs. 1–6, 9–15, 18, 19, 22–27, and 29–32 are available in the associated source data file. Source data are provided with this paper.
Change history
19 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41467-025-67649-w
References
Johnson, D. E. et al. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Prim. 6, 92 (2020).
Pulito, C. et al. Oral mucositis: the hidden side of cancer therapy. J. Exp. Clin. cancer Res. 39, 1–15 (2020).
Maria, O. M., Eliopoulos, N. & Muanza, T. Radiation-induced oral mucositis. Front. Oncol. 7, 89 (2017).
Kuhar H. N. et al. Distinct Histopathologic Features Of Radiation-induced Chronic Sinusitis. In: International forum of allergy & rhinology. Wiley Online Library (2017).
Zheng C., Yu L., Jiang Y. Radiation-induced Rhinosinusitis: Mechanism Research And Clinical Progress Review. World Journal of Otorhinolaryngology-Head and Neck Surgery, (2023).
Sciuca, A. M., Neamtu, M., Marcu, D., Costan, V. V. & Popa, C. Oral mucositis in patients with chemotherapy treatment. Rom. J. Oral Rehabil. 16, 452–458 (2024).
Tu, Z. et al. Scavenging tumor-derived small extracellular vesicles by functionalized 2D materials to inhibit tumor regrowth and metastasis following radiotherapy. Adv. Funct. Mater. 32, 2205663 (2022).
Ramanathan, S. et al. Inflammation potentiates miR-939 expression and packaging into small extracellular vesicles. J. Extracell. vesicles 8, 1650595 (2019).
Yang, Z. et al. The emerging role of exosomes in radiotherapy. Cell Commun. Signal. 20, 171 (2022).
Zhu, Y., Liu, C. & Pang, Z. Dendrimer-based drug delivery systems for brain targeting. Biomolecules 9, 790 (2019).
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. nature 346, 818–822 (1990).
Wilms, D., Stiriba, S.-E. & Frey, H. Hyperbranched polyglycerols: from the controlled synthesis of biocompatible polyether polyols to multipurpose applications. Acc. Chem. Res. 43, 129–141 (2010).
Tu, Z. et al. Tackling severe neutrophilic inflammation in airway disorders with functionalized nanosheets. ACS nano 18, 7084–7097 (2024).
Yahyapour, R. et al. Radiation-induced inflammation and autoimmune diseases. Mil. Med. Res. 5, 1–8 (2018).
El-Kenawi, A. & Ruffell, B. Inflammation, ROS, and mutagenesis. Cancer cell 32, 727–729 (2017).
Jiao, J. et al. Expression of STING is increased in monocyte-derived macrophages and contributes to liver inflammation in hepatic ischemia-reperfusion injury. Am. J. Pathol. 192, 1745–1762 (2022).
Wu, K. et al. Polyglycerol-amine covered nanosheets target cell-free dna to attenuate acute kidney injury. Adv. Sci. 10, 2300604 (2023).
Gugnacki, P. & Sierko, E. Is there an interplay between oral microbiome, head and neck carcinoma and radiation-induced oral mucositis?. Cancers 13, 5902 (2021).
Casero, D. et al. Space-type radiation induces multimodal responses in the mouse gut microbiome and metabolome. Microbiome 5, 1–18 (2017).
Lässer, C. et al. RNA-containing exosomes in human nasal secretions. Am. J. Rhinol. allergy 25, 89–93 (2011).
Chirivi, R. G. et al. Therapeutic ACPA inhibits NET formation: a potential therapy for neutrophil-mediated inflammatory diseases. Cell. Mol. Immunol. 18, 1528–1544 (2021).
Tian, X. et al. Medicinal chemistry perspective on cGAS-STING signaling pathway with small molecule inhibitors. Eur. J. Med. Chem. 244, 114791 (2022).
Menck, K. et al. Neutral sphingomyelinases control extracellular vesicles budding from the plasma membrane. J. Extracell. Ves. 6, 1378056 (2017).
Zhang, H., Chhowalla, M. & Liu, Z. 2D nanomaterials: graphene and transition metal dichalcogenides. Chem. Soc. Rev. 47, 3015–3017 (2018).
Murali, A., Lokhande, G., Deo, K. A., Brokesh, A. & Gaharwar, A. K. Emerging 2D nanomaterials for biomedical applications. Mater. Today 50, 276–302 (2021).
Bernard, M., Jubeli, E., Pungente, M. D. & Yagoubi, N. Biocompatibility of polymer-based biomaterials and medical devices–regulations, in vitro screening and risk-management. Biomater. Sci. 6, 2025–2053 (2018).
Li, H. et al. Construction of CpG Delivery Nanoplatforms by Functionalized MoS2 Nanosheets for Boosting Antitumor Immunity in Head and Neck Squamous Cell Carcinoma. Small 19, 2300380 (2023).
Kurapati, R. et al. Enzymatic biodegradability of pristine and functionalized transition metal dichalcogenide MoS2 nanosheets. Adv. Funct. Mater. 27, 1605176 (2017).
Song, Z. et al. Development of a CD63 aptamer for efficient cancer immunochemistry and immunoaffinity-based exosome isolation. Molecules 25, 5585 (2020).
Zang, W. et al. Distribution of Pt single atom coordination environments on anatase TiO2 supports controls reactivity. Nat. Commun. 15, 998 (2024).
Miao, P. et al. Exacerbation of allergic rhinitis by the commensal bacterium Streptococcus salivarius. Nat. Microbiol. 8, 218–230 (2023).
Tu, Z. et al. Functional 2D nanoplatforms alleviate eosinophilic chronic rhinosinusitis by modulating eosinophil extracellular trap formation. Adv. Sci. 11, 2307800 (2024).
Wang, Y. et al. Universal STING mimic boosts antitumour immunity via preferential activation of tumour control signalling pathways. Nat. Nanotechnol. 19, 856–866 (2024).
Jiang, D. et al. Nanozyme: new horizons for responsive biomedical applications. Chem. Soc. Rev. 48, 3683–3704 (2019).
Zhang, J. et al. ROS scavenging biopolymers for anti-inflammatory diseases: classification and formulation. Adv. Mater. Interfaces 7, 2000632 (2020).
Lin, L. et al. Targeted nanotheranostics for the treatment of epilepsy through in vivo hijacking of locally activated macrophages. Acta Biomater. 174, 314–330 (2024).
Chandraker, K., Vaishanav, S. K., Nagwanshi, R. & Satnami, M. L. Radical scavenging efficacy of thiol capped silver nanoparticles. J. Chem. Sci. 127, 2183–2191 (2015).
Hong, Z. et al. Airborne fine particulate matter induces oxidative stress and inflammation in human nasal epithelial cells. Tohoku J. Exp. Med. 239, 117–125 (2016).
Liu, T., Sun, L., Zhang, Y., Wang, Y. & Zheng, J. Imbalanced GSH/ROS and sequential cell death. J. Biochem. Mol. Toxicol. 36, e22942 (2022).
Zhang, Y. et al. The role of bacteria and its derived biomaterials in cancer radiotherapy. Acta Pharm. Sin. B 13, 4149–4171 (2023).
Mei, L. et al. Two-dimensional nanomaterials beyond graphene for antibacterial applications: current progress and future perspectives. Theranostics 10, 757 (2020).
Xiao, Y. et al. Multifunctional cationic hyperbranched polyaminoglycosides that target multiple mediators for severe abdominal trauma management. Adv. Sci. 11, 2305273 (2024).
de Moraes Mazetto, B. et al. Association between neutrophil extracellular traps (NETs) and thrombosis in antiphospholipid syndrome. Thromb. Res. 214, 132–137 (2022).
Wei, Y., Quan, L., Zhou, C. & Zhan, Q. Factors relating to the biodistribution & clearance of nanoparticles & their effects on in vivo application. Nanomedicine 13, 1495–1512 (2018).
Mc Larney B. E. et al. A pan-cancer dye for solid-tumour screening, resection and wound monitoring via short-wave and near-infrared fluorescence imaging. Nat. Biomed. Eng. 8, 1092–1108 (2024).
Wu, J. The enhanced permeability and retention (EPR) effect: the significance of the concept and methods to enhance its application. J. personalized Med. 11, 771 (2021).
Bachmaier, K., Mair, J., Offner, F., Pummerer, C. & Neu, N. Serum cardiac troponin T and creatine kinase–MB elevations in murine autoimmune myocarditis. Circulation 92, 1927–1932 (1995).
Stoimenou, M., Tzoros, G., Skendros, P. & Chrysanthopoulou, A. Methods for the assessment of NET formation: from neutrophil biology to translational research. Int. J. Mol. Sci. 23, 15823 (2022).
Singh, V. & Singh, A. K. Oral mucositis. Natl. J. Maxillofac. Surg. 11, 159–168 (2020).
Vercauteren Drubbel, A. & Beck, B. Single-cell transcriptomics uncovers the differentiation of a subset of murine esophageal progenitors into taste buds in vivo. Sci. Adv. 9, eadd9135 (2023).
Buscher, K. et al. Natural variation of macrophage activation as disease-relevant phenotype predictive of inflammation and cancer survival. Nat. Commun. 8, 16041 (2017).
Zhang, L.-Z. et al. PD-1/CD80+ small extracellular vesicles from immunocytes induce cold tumours featured with enhanced adaptive immunosuppression. Nat. Commun. 15, 3884 (2024).
Geng, N. et al. 10-hydroxy-2-decenoic acid prevents osteoarthritis by targeting aspartyl β hydroxylase and inhibiting chondrocyte senescence in male mice preclinically. Nat. Commun. 15, 7712 (2024).
McBride, W. H. & Schaue, D. Radiation-induced tissue damage and response. J. Pathol. 250, 647–655 (2020).
Zhang, J. et al. 3D Printing of a vascularized mini-liver based on the size-dependent functional enhancements of cell spheroids for rescue of liver failure. Adv. Sci. 11, 2309899 (2024).
Kim, K. W., Moretti, L., Mitchell, L. R., Jung, D. K. & Lu, B. Endoplasmic reticulum stress mediates radiation-induced autophagy by perk-eIF2α in caspase-3/7-deficient cells. Oncogene 29, 3241–3251 (2010).
Chaurasia, M. et al. Radiation induces EIF2AK3/PERK and ERN1/IRE1 mediated pro-survival autophagy. Autophagy 15, 1391–1406 (2019).
Wopereis, S., Lefeber, D. J., Morava, E. & Wevers, R. A. Mechanisms in protein O-glycan biosynthesis and clinical and molecular aspects of protein O-glycan biosynthesis defects: a review. Clin. Chem. 52, 574–600 (2006).
Bergstrom, K. S. & Xia, L. Mucin-type O-glycans and their roles in intestinal homeostasis. Glycobiology 23, 1026–1037 (2013).
Kudelka, M. R., Stowell, S. R., Cummings, R. D. & Neish, A. S. Intestinal epithelial glycosylation in homeostasis and gut microbiota interactions in IBD. Nat. Rev. Gastroenterol. Hepatol. 17, 597–617 (2020).
Zhang Z., Zhang C. Regulation of cGAS–STING signalling and its diversity of cellular outcomes. Nat. Rev. Immunol. 25, 425–444 (2025).
Mohammapdour, R. & Ghandehari, H. Mechanisms of immune response to inorganic nanoparticles and their degradation products. Adv. drug Deliv. Rev. 180, 114022 (2022).
Gulen, M. F. et al. cGAS–STING drives ageing-related inflammation and neurodegeneration. Nature 620, 374–380 (2023).
Tu, Z. et al. Graphene oxide-cyclic R10 peptide nuclear translocation nanoplatforms for the surmounting of multiple-drug resistance. Adv. Funct. Mater. 30, 2000933 (2020).
Xiao, Y. et al. Modulation of unregulated inflammation-associated coagulopathy in sepsis using multifunctional nanosheets. Adv. Funct. Mater. 34, 2402785 (2024).
Takov, K., Yellon, D. M. & Davidson, S. M. Confounding factors in vesicle uptake studies using fluorescent lipophilic membrane dyes. J. Extracell. Ves. 6, 1388731 (2017).
Acknowledgements
Z. Tu was supported by the National Natural Science Foundation of China (No. 82271188), the Young Elite Scientists Sponsorship Program by CAST (No. 2022QNRC001) and the Funding by Science and Technology Projects in Guangzhou (No. 2025A04J7163). W. Wen acknowledged the National Natural Science Foundation of China (No. 82020108009), Guangdong Natural Science Foundation of China grants (No. 2023B1111040004), Key Clinical Technique of Guangzhou (No. 2023P-ZD06), and the Funding by Science and Technology Projects in Guangzhou (No. 2024A04J10019).
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.Z., Z.T., K.W.L., and W.W. Methodology: Y.Z., Z.L., C.X., M.L., and Z.T. Investigation: Y.Z., Z.L., X.B., and Z.T. Visualization: Z.L., C.X., M.L., Y.S., R.M., and X.L. Supervision: Z.T., K.W.L., and W.W. Writing—original draft: Y.Z., and Z.T. Writing—review & editing: Y.Z., Z.T., J.L., K.W.L., and W.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Robert Maile and Lianhui Wang for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, Y., Xu, C., Li, Z. et al. Nanostructured organic sheets sequestering small extracellular vesicles and reactive species to protect against radiation-induced mucositis. Nat Commun 16, 6120 (2025). https://doi.org/10.1038/s41467-025-61236-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61236-9