Abstract
The maternal microbiome during pregnancy and the peripartum period plays a critical role in maternal health outcomes and establishing the neonatal gut microbiome, with long-term implications for offspring health. However, a healthy microbiome during these key periods has not been definitively characterized. This longitudinal study examines maternal and neonatal microbiomes using 16S rRNA amplicon sequencing in a Japanese cohort throughout pregnancy and the postpartum period. Forty-two mothers and their forty-five offspring participate in the study. The maternal vaginal microbiome remains relatively stable during pregnancy but significantly changes in the postpartum period. Among Lactobacillus species, the Lactobacillus crispatus group is predominant. A higher abundance of Lactobacillus early in pregnancy is associated with a favorable gestational period. The maternal gut microbiome is associated with the vaginal microbiome throughout pregnancy. The neonatal gut microbiome substantially changes in early life, with bacterial composition influenced by delivery mode. Over time, bacteria shared with the maternal gut microbiome become dominant in the neonatal gut. This study provides insights into microbiome dynamics in Japanese mothers and their offspring during pregnancy and the postpartum period. Identification of common patterns across diverse populations may help define keystone microbes essential for human health and inform the development of microbiome-based interventions.
Similar content being viewed by others
Introduction
Host–microbiome interactions play important roles in health and disease1,2. Microbiome composition varies among body sites, and each site-specific microbiome makes unique contributions to host physiology3,4. In pregnant women, the vaginal microbiome influences the course of pregnancy and neonatal clinical outcomes5,6. Although the vaginal microbiome is suspected to undergo shifts during pregnancy and the postpartum period7,8 due to fluctuations in sex hormone levels, the precise longitudinal changes throughout these stages remain poorly characterized. The maternal gut microbiome may also affect maternal and neonatal health outcomes9,10,11,12; however, evidence regarding its composition and roles during pregnancy and the postpartum period remains limited. An understanding of the maternal microbiome during these critical periods is essential because it may affect maternal health and have persistent impacts on offspring health.
The maternal vaginal and gut microbiomes are presumed to be vertically transmitted to offspring, influencing the early neonatal microbiome13,14. This microbial transmission is a key process that shapes the neonatal microbiome in early life and may have long-term consequences. Various factors influence neonatal microbiome development, including maternal microbiome composition13, mode of delivery15,16,17, feeding practices and diet18,19, probiotic exposure20, antibiotic use21, environmental factors (e.g., lifestyle and hygiene)21,22, and genetic background23. Among these, the maternal microbiome during the peripartum period is considered a major source of microbial colonization in offspring14. Furthermore, early-life microbiome composition influences its establishment later in life24. Ultimately, maternal microbiome transmission is considered to exert long-term effects on the microbiome and health of the offspring.
Various factors, including environmental influences, lifestyle, and genetic background, affect the host microbiome and can introduce confounding variables in human microbiome research. Strict control of these factors is often infeasible. Therefore, investigations of the microbiome in cohorts from diverse regions and populations can help to identify key microbial taxa that are universally important for human health. Multiple studies have shown that microbiomes within the Japanese population substantially differ from microbiomes within the populations of other developed countries in Europe and the United States25. Additionally, the increasing prevalences of allergic, autoimmune, and metabolic diseases have become major medical concerns in Japan26,27,28, mirroring trends observed in Western countries. These epidemiological shifts are thought to result from microbiome alterations induced by a Westernized lifestyle and diet. Microbiome studies involving Japanese populations may provide insights into characteristics shared between Asian and Western microbiomes that are critical for human health and disease.
In this study, we prospectively tracked the microbiome of 42 mothers and their 45 offspring from 12 weeks of gestation to 1 month postpartum in a Japanese cohort. Detailed clinical data were collected throughout pregnancy and the postpartum period. This is the first study to analyze longitudinal changes in maternal and neonatal microbiomes in the Japanese population. The prospective study design, with time-sequenced sampling and well-curated clinical data, offers a major advantage in investigating microbiome dynamics and potential causal relationships.
Results
Changes in the maternal vaginal microbiome from pregnancy to the postpartum period
The demographic characteristics of the study participants are summarized in Table 1. The numbers of samples analyzable at each time point are shown in Supplementary Fig. 1. Maternal microbiome samples were collected at multiple time points: during pregnancy (Weeks 12, 20, 30, and 36), at birth (hereinafter, “Birth”), and 1 month after birth (hereinafter, “Postpartum”) (Fig. 1a). At all time points during pregnancy and postpartum, vaginal and gut samples were collected. At delivery, vaginal and gut samples were collected from mothers who underwent vaginal delivery, whereas vaginal and skin samples were collected from those who underwent cesarean section. (Fig. 1a and Supplementary Fig. 1). In total, 239 vaginal samples from 42 mothers were analyzed, with an average read count of 46,207 and a total of 2737 amplicon sequence variants (ASVs). Bacterial genera identified in all maternal vaginal samples are presented in Supplementary Fig. 2. To assess compositional changes in the vaginal microbiome throughout pregnancy and the postpartum period, we analyzed α-diversity using the Chao1 and Shannon indices. The Chao1 index estimates microbial diversity based on ASV richness, whereas the Shannon index incorporates both ASV richness and evenness to provide a more comprehensive measure of diversity. Both indices were significantly lower during pregnancy and at Delivery compared with Postpartum (Chao1 index: p < 0.0001 [Week 12 vs. Postpartum], p = 0.0086 [Week 20 vs. Birth], p < 0.0001 [Week 20 vs. Postpartum], p < 0.0464 [Week 30 vs. Birth], p < 0.0001 [Week 30 vs. Postpartum], p < 0.0001 [Week 36 vs. Postpartum], and p < 0.0001 [Birth vs. Postpartum]; Shannon index: p = 0.0306 [Week 12 vs. Birth], p < 0.0001 [Week 12 vs. Postpartum], p = 0.0023 [Week 20 vs. Birth], p < 0.0001 [Week 20 vs. Postpartum], p = 0.0306 [Week 30 vs. Birth], p < 0.0001 [Week 30 vs. Postpartum], p < 0.0001 [Week 36 vs. Postpartum], and p < 0.0001 [Birth vs. Postpartum]) (Fig. 1b, Supplementary Table 1). β-diversity was evaluated using UniFrac distances. The unweighted UniFrac distance considers the presence or absence of specific ASVs, whereas the weighted UniFrac distance considers both ASV composition and relative abundance. Both unweighted and weighted UniFrac distances showed significant differences between samples collected during pregnancy and at Delivery compared with those collected at Postpartum (unweighted: p = 0.0033 [Week 12 vs. Birth], p = 0.0021 [Week 12 vs. Postpartum], p = 0.0021 [Week 20 vs. Birth], p = 0.0021 [Week 20 vs. Postpartum], p = 0.0021 [Week 30 vs. Birth], p = 0.0021 [Week 30 vs. Postpartum], p = 0.0021 [Week 36 vs. Birth], p = 0.0021 [Week 36 vs. Postpartum], and p = 0.0021 [Birth vs. Postpartum]; weighted: p = 0.0030 [Week 12 vs. Postpartum], p = 0.0171 [Week 20 vs. Birth], p = 0.0030 [Week 20 vs. Postpartum], p = 0.0171 [Week 30 vs. Birth], p = 0.0030 [Week 30 vs. Postpartum], p = 0.0030 [Week 36 vs. Postpartum], and p = 0.0030 [Birth vs. Postpartum]) (Fig. 1c and Supplementary Table 2). These findings indicate that the vaginal microbiome underwent significant compositional changes in the postpartum period, despite its relative stability during pregnancy. It can be hypothesized that the abrupt hormonal changes associated with childbirth, along with exposure to external environmental factors during delivery, contribute to alterations in the vaginal environment, thereby providing opportunities for the colonization of various microorganisms.
a Study design outlining the collection of maternal samples. b Chao1 and Shannon diversity indices in the vaginal microbiome (Week 12: n = 42, Week 20: n = 40, Week 30: n = 41, Week 36: n = 39, Birth: n = 41, Postpartum: n = 36). Each p-value is shown in Supplementary Table 1. c Principal coordinates analysis (PCoA) plots based on unweighted and weighted UniFrac distances of bacterial compositions in the maternal vaginal microbiome. Each p-value is shown in Supplementary Table 2. Data are presented as means ± standard errors of the mean. *p < 0.05, **p < 0.01, ****p < 0.0001, Pairwise comparisons (two-sided) were performed using estimated marginal means, accounting for repeated measures, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panel (b). Bacterial compositions were compared using permutational analysis of variance with permutations restricted within each participant, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panel (c).
Multivariable association analysis at the bacterial genus level revealed a significantly higher relative abundance of Lactobacillus during Weeks 12, 20, 30, and 36 (hereinafter, collectively referred to as “Pregnancy”) compared with Postpartum (77.78 ± 2.89% vs. 9.24 ± 4.30%; p < 0.0001) (Fig. 2a). No significant differences in bacterial genera were observed between early pregnancy (Week 12) and late pregnancy (Week 36). During Pregnancy, Lactobacillus was the most abundant genus (77.78 ± 2.89%), followed by Gardnerella (8.68 ± 1.49%), Streptococcus (3.16 ± 1.24%), Atopobium (1.95 ± 0.58%), and Bifidobacterium (1.81 ± 0.97%) (Fig. 2b). The Chao1 and Shannon indices were significantly lower in samples where Lactobacillus had the highest relative abundance compared with those where other bacterial genera were dominant (Chao1 index: 27.63 ± 1.03 vs. 65.55 ± 7.90; p = 0.00000005, Shannon index: 4.05 ± 0.04 vs. 5.01 ± 0.19; p < 0.0000004) (Fig. 2c). Changes in the relative abundance of Lactobacillus over time followed three distinct patterns (colors depicted in corresponding figure): (1) a consistently high abundance from early to late pregnancy (blue), (2) a low abundance in early pregnancy followed by an increase in late pregnancy (green), and (3) a consistently low abundance throughout pregnancy (red) (Fig. 2d). All mothers in whom Lactobacillus was the dominant vaginal genus at Week 12 were defined as Lactobacillus-dominant group at early pregnancy. These mothers had a significantly higher likelihood of continuing pregnancy beyond 38 weeks of gestation compared with those in whom other genera were dominant (89.66% [26/29] vs. 50.00% [5/10]; p = 0.0164). The overall course of pregnancy up to 38 weeks significantly differed between these groups (p = 0.0025) (Fig. 2e). Regarding clinical characteristics, mothers in whom Lactobacillus was dominant at Week 12 had a significantly higher pre-pregnancy body mass index (BMI) compared with those in whom other genera were dominant (median [interquartile range]: 21.6 [19.9–24.1] vs. 19.9 [19.1–20.9]; p = 0.0476) (Table 2). However, at Week 30 (late pregnancy), no significant difference was observed in the rate of pregnancy continuation beyond 38 weeks between mothers in whom Lactobacillus was dominant and those in whom other genera were dominant (78.79% [26/33] vs. 83.33% [5/6]; p = 1.0000). All mothers in whom Lactobacillus was the dominant vaginal genus at Week 30 were included in the Lactobacillus-dominant group at late pregnancy. Courses of pregnancy up to 38 weeks also did not differ between these groups (Fig. 2f). These findings suggest that a high relative abundance of Lactobacillus in early pregnancy plays a critical role in promoting favorable pregnancy outcomes. Additionally, Lactobacillus colonization in early pregnancy may be associated with specific maternal clinical characteristics before conception: higher pre-pregnancy BMI was associated with Lactobacillus-dominant vaginal microbiome at Week 12 in the present Japanese cohort.
a Comparison of the relative abundance of bacterial genera in the vaginal microbiome between pregnancy and the postpartum period. b Relative abundances of the top 10 bacterial genera in the vaginal microbiome during pregnancy. c Chao1 and Shannon diversity indices in samples where Lactobacillus was dominant (n = 129) compared with those in which other genera were dominant (n = 33) (Chao1 index: p = 0.00000005, Shannon diversity index: p = 0.0000004). d Longitudinal shifts in the relative abundance of Lactobacillus: sustained high levels from early to late pregnancy (blue), low levels in early pregnancy followed by high levels in late pregnancy (green), and sustained low levels from early to late pregnancy (red). e Comparison of pregnancy continuation beyond 38 weeks between mothers in whom Lactobacillus was dominant during early pregnancy and those in whom other genera were dominant at that time (bar chart; p = 0.0164). Kaplan–Meier curves illustrate pregnancy continuation up to 38 weeks (p = 0.0025). f Comparison of pregnancy continuation beyond 38 weeks between mothers in whom Lactobacillus was dominant during late pregnancy and those in whom other genera were dominant at that time (bar chart; p = 1.0000). Kaplan–Meier curves illustrate pregnancy continuation up to 38 weeks (p = 1.0000). Data are presented as means ± standard errors of the mean. *p < 0.05, ***p < 0.001, Mann–Whitney U test (two-sided) for comparisons of two groups in panels (c), (e), and (f); log-rank test for comparisons of Kaplan–Meier curves in panels (e) and (f).
Next, we analyzed the genus Lactobacillus at the subgenus level. In this classification, four major bacterial species commonly found in the vaginal microbiome—L. crispatus, L. iners, L. jensenii, and L. gasseri—were used as representative species for each group29. At all time points during pregnancy, the L. crispatus group had the highest relative abundance (54.17 ± 7.68%, 62.04 ± 7.61%, 63.19 ± 7.15%, and 70.73 ± 6.63% at Weeks 12, 20, 30, and 36, respectively) compared with other Lactobacillus subgenus (Fig. 3a). The proportion of mothers in whom the L. crispatus group was the dominant species progressively increased from Week 12 to Week 36 (54.76%, 60.00%, 65.85%, and 74.36%, respectively). Furthermore, after the L. crispatus group became dominant in an individual, other Lactobacillus species did not appear to replace its dominance (Fig. 3b). Samples in which the L. crispatus group was dominant exhibited significantly lower Chao1 and Shannon indices compared with those in which other Lactobacillus subgenus were dominant (Chao1 index: 26.49 ± 1.09 vs. 32.76 ± 2.66; p = 0.0125, Shannon index: 4.00 ± 0.045 vs. 4.25 ± 0.11; p = 0.0294) (Fig. 3c). These findings suggest that the L. crispatus group played a beneficial role in maintaining a healthy vaginal microbiome in this Japanese cohort.
a Heatmap and dot plots showing the relative abundances of Lactobacillus subgenera at each time point. In the heatmap, each column represents an individual mother (Week 12: n = 42, Week 20: n = 40, Week 30: n = 41, Week 36: n = 39). b Alluvial plot depicting the dominant Lactobacillus subgenera in vaginal samples over time. c Comparison of Chao1 and Shannon diversity indices between samples in which the L. crispatus group was dominant (n = 104) and those in which other Lactobacillus subgenera were dominant (n = 25) during pregnancy (Chao1 index: p = 0.0125, Shannon diversity index: p = 0.0294). Data are presented as means ± standard errors of the mean. *p < 0.05, Mann–Whitney U test (two-sided) for comparisons of two groups in panel (c).
Changes in the maternal gut microbiome during pregnancy and the postpartum period
We subsequently evaluated the maternal gut microbiome. In total, 220 gut samples were analyzed, with an average read count of 50,063 and a total of 6542 ASVs. Bacterial genera identified in all maternal gut samples are presented in Supplementary Fig. 3. α-diversity remained stable during pregnancy, at Delivery, and at Postpartum (Fig. 4a and Supplementary Table 3). In the analysis of β-diversity based on the UniFrac distances, while there were no significant changes up to Week 30, the gut bacterial compositions started overtime change after Week 36 (unweighted: p = 0.0338 [Week 12 vs. Birth], p = 0.0390 [Week 20 vs. Birth], p = 0.0337 [Week 30 vs. Birth], p = 0.0338 [Week 36 vs. Postpartum], p = 0.0338 [Birth vs. Postpartum]; Weighted: p = 0.0050 [Week 12 vs. Week 36], p = 0.0175 [Week 12 vs. Birth], p = 0.0216 [Week 20 vs. Week 36], p = 0.0216 [Week 20 vs. Birth], p = 0.0050 [Week 30 vs. Week 36], p = 0.0193 p = 0.0175 [Week 30 vs. Birth], p = 0.0192 Week 36 vs. Birth], p = 0.0050 Week 36 vs. Postpartum], and p = 0.0075 [Birth vs. Postpartum]) (Fig. 4b and Supplementary Table 4). Multivariable association analysis at the bacterial genus level revealed a significantly higher relative abundance of Lactobacillus and a significantly lower relative abundance of Howardella during pregnancy compared with Postpartum (Fig. 4c). Additionally, the relative abundance of Lactobacillus was significantly higher in late pregnancy (Week 36) than in early pregnancy (Week 12), whereas the relative abundances of Lowsonella and Prevotella were significantly lower at Week 36 (Fig. 4d). Overall, the maternal gut microbiome composition changed over the course of late pregnancy and postpartum, with the relative abundance of Lactobacillus increasing as pregnancy progressed and decreasing during the postpartum period.
a Chao1 and Shannon diversity indices in the maternal gut microbiome during pregnancy and the postpartum period (Week 12: n = 40, Week 20: n = 40, Week 30: n = 41, Week 36: n = 39, Birth: n = 21, Postpartum: n = 38). Each p-value is shown in Supplementary Table 3. b Principal coordinates analysis (PCoA) plots based on unweighted and weighted UniFrac distances of bacterial compositions in the maternal gut microbiome during pregnancy and the postpartum period. Each p-value is shown in Supplementary Table 4. c Comparison of bacterial genera in the maternal gut microbiome between pregnancy and the postpartum period. d Comparison of bacterial genera in the maternal gut microbiome between early pregnancy (Week 12) and late pregnancy (Week 36). e Multivariable association analysis of the relative abundance of Lactobacillus in the maternal vagina and the relative abundances of bacterial genera in the maternal gut. f Multivariable association analysis of the relative abundances of Lactobacillus subgenera in the maternal vagina and bacterial genera in the maternal gut. Data are presented as means ± standard errors of the mean. Pairwise comparisons (two-sided) were performed using estimated marginal means, accounting for repeated measures, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panel (a). Bacterial compositions were compared using permutational analysis of variance with permutations restricted within each participant, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panel (b).
We investigated the association between the relative abundance of Lactobacillus in vagina during pregnancy and that of various bacterial genera in the gut. Association analysis revealed a strong positive association between the relative abundance of Lactobacillus in the gut microbiome and that in the vaginal microbiome. (coefficient = 1.695; p < 0.0001) (Fig. 4e). Further association analysis examining four Lactobacillus subgenera in the vagina and bacterial genera in the gut showed that the relative abundance of the L. crispatus group in the vagina was positively associated with the relative abundance of the Lactobacillus genus in the gut (coefficient = 2.11; p < 0.0001). The L. iners group also exhibited a positive association, although weaker than that of the L. crispatus group (coefficient = 0.76; p = 0.0060). In contrast, the L. gasseri (coefficient = 0.41; p = 0.1077) and L. jensenii (coefficient = 0.27; p = 0.2984) groups did not show significant associations with the relative abundance of the Lactobacillus genus in the gut (Fig. 4f). Distinct bacterial genera in the gut were associated with each Lactobacillus subgenus in the vagina.
Rapid changes in the neonatal gut microbiome after birth
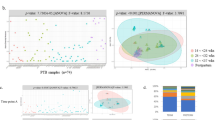
We analyzed the neonatal gut microbiome using 143 rectal samples obtained from 45 children born to 42 mothers. Rectal samples were collected on postnatal days 0, 1, and 4 (Day 0, Day 1, and Day 4, respectively) and at 1 month of age (Month 1) (Fig. 5a). The dataset included an average read count of 44,859 and a total of 3,927 ASVs. Bacterial genera identified in all neonatal gut samples are presented in Supplementary Fig. 4. α-diversity was significantly lower on Day 4 than on Day 0, Day 1, and at Month 1. Additionally, α-diversity was significantly lower at Month 1 than on Day 0 (Chao1 index: p = 0.0056 [Day 0 vs. Day 1], p < 0.0001 [Day 0 vs. Day 4], p < 0.0001 [Day 0 vs. Month 1], and p = 00015 [Day 1 vs. Day 4]; Shannon index: p = 0.0202 [Day 0 vs. Day 1], p < 0.0001 [Day 0 vs. Day 4], p < 0.0001 [Day 0 vs. Month 1], p < 0.0001 [Day 1 vs. Day 4], p = 0.0345 [Day 1 vs. Month 1], p = 0.0046 [Day 4 vs. Month 1]) (Fig. 5b, Supplementary Table 5). β-diversity analysis using both unweighted and weighted UniFrac distances revealed significant differences among all time points except between Day 0 and Day 1 (unweighted: p = 0.0050 [Day 0 vs. Day 1], p = 0.0012 [Day 0 vs. Day 4], p = 0.0012 [Day 0 vs. Month 1], p = 0.0012 [Day 1 vs. Day 4], p = 0.0012 [Day 1 vs. Month 1], and p = 0.0012 [Day 4 vs. Month 1]; weighted: p = 0.0070 [Day 0 vs. Day 1], p = 0.0012 [Day 0 vs. Day 4], p = 0.0012 [Day 0 vs. Month 1], p = 0.0012 [Day 1 vs. Day 4], p = 0.0012 [Day 1 vs. Month 1], and p = 0.0012 [Day 4 vs. Month 1]) (Fig. 5c and Supplementary Table 6). Considering the absence of significant differences in bacterial composition between Days 0 and 1, we analyzed changes in bacterial genera during the early and late neonatal periods by comparing the gut microbiome between Days 1 and 4, as well as between Day 4 and Month 1. The relative abundances of Staphylococcus, Rothia, Streptococcus, and Haemophilus were higher on Day 4 than on Day 1 (Fig. 5d). Between Day 4 and Month 1, the relative abundances of Staphylococcus, Bacillus, Rothia, and Gemella decreased; those of Finegoldia, Anaerococcus, Peptoniphilus, and Enterobacter increased (Fig. 5e). These findings indicate that the pattern of gut microbiome changes differed before vs. after Day 4, despite continuous compositional shifts throughout the neonatal period.
a Time points for neonatal sample collection. b Chao1 and Shannon diversity indices in the neonatal gut microbiome (Day 0: n = 27, Day 1: n = 28, Day 4: n = 43, Month 1: n = 39). Each p-value is shown in Supplementary Table 5. c Principal coordinates analysis (PCoA) plots based on unweighted and weighted UniFrac distances of bacterial compositions in the neonatal gut microbiome. Each p-value is shown in Supplementary Table 6. d Comparison of the relative abundances of bacterial genera in the neonatal gut microbiome between Day 1 and Day 4. e Comparison of the relative abundances of bacterial genera in the neonatal gut microbiome between Day 4 and Month 1. f PCoA plots based on unweighted and weighted UniFrac distances at each time point, comparing bacterial composition in the neonatal gut microbiome between vaginally delivered and cesarean-section-delivered offspring. g Multivariable association analysis of bacterial genera in the neonatal gut microbiome, comparing vaginally delivered and cesarean-section-delivered offspring at each time point. Data are presented as means ± standard errors of the mean. *p < 0.05, **p < 0.01, ****p < 0.0001. Pairwise comparisons (two-sided) were performed using estimated marginal means, accounting for repeated measures, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panel (b). Bacterial compositions were compared using permutational analysis of variance with permutations restricted within each participant, and adjusted for multiple comparisons using the Benjamini-Hochberg method in panels (c) and (f).
We then compared longitudinal changes in the gut microbiome between vaginally delivered and cesarean-section-delivered neonates. On Day 0, there was no significant difference in bacterial composition between the two groups. However, significant differences emerged on Day 1, Day 4, and at Month 1, as indicated by both unweighted and weighted UniFrac distances (Day 1: unweighted p = 0.0090, weighted p = 0.0100; Day 4: unweighted p = 0.0020, weighted p = 0.0010; Month 1: unweighted p = 0.0010, weighted p = 0.0430) (Fig. 5f). Multivariable association analysis of bacterial genera revealed no significant differences between vaginally delivered and cesarean-section-delivered neonates on Day 0. However, significant differences were observed on Day 1, Day 4, and at Month 1; distinct bacterial genera varied over time (Fig. 5g). These findings suggest that the mode of delivery has a substantial and persistent impact on early gut microbiome development.
Shared microbial signatures between the maternal microbiome and the neonatal gut microbiome in early life
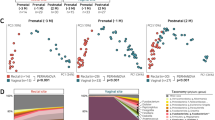
We investigated the impact of the maternal microbiome at various body sites on the bacterial composition of the neonatal gut microbiome. Bacterial genera identified in all maternal vaginal, gut, and skin samples are presented in Supplementary Fig. 5. The presence of bacterial ASVs shared between the maternal microbiome and the neonatal gut microbiome was assessed on Days 0, 1, and 4, and at Month 1. We examined shared ASVs between mothers and their neonates from two perspectives: (1) the proportion of ASV types shared with each maternal site, relative to the total ASV types observed in the neonatal gut microbiome, and (2) the relative abundance of these shared ASVs within the neonatal gut microbiome. In vaginally delivered neonates, the proportion of ASV types shared with the maternal vaginal microbiome exhibited a decreasing trend. In contrast, the proportion of ASV types shared with the maternal gut microbiome increased over time (Fig. 6a and Supplementary Table 7). On Day 4 and at Month 1, the proportion of ASV types shared with the maternal gut microbiome was significantly higher than that shared with the maternal vaginal microbiome (Day 4: p = 0.0004, Month 1: p < 0.0001) (Fig. 6a and Supplementary Table 7). The total proportion of ASV types shared with the maternal vaginal and gut microbiomes remained relatively stable in the neonatal gut microbiome across all time points. The relative abundance of ASV types shared with the maternal vaginal microbiome exhibited a decreasing trend, whereas the relative abundance of ASV types shared with the maternal gut microbiome increased over time (Fig. 6b and Supplementary Table 7). On Day 4 and at Month 1, the relative abundance of ASV types shared with the maternal gut microbiome was significantly higher than that shared with the maternal vaginal microbiome (Day 4: p < 0.0001, Month 1: p < 0.0001) (Fig. 6b and Supplementary Table 7). By Day 4 and Month 1, ASV types shared with the maternal vaginal microbiome became a minor component of the neonatal gut microbiome. These findings suggest that the neonatal gut microbiome becomes more strongly similar to the maternal gut microbiome than to the vaginal microbiome over time.
a, b Proportion and relative abundance of ASV types in the neonatal gut microbiome that were shared with the maternal vaginal or gut microbiome in vaginally delivered offspring (Vagina Day 0: n = 16, Day 1: n = 17, Day 4: n = 20, Month 1: n = 18; Gut Day 0: n = 16, Day 1: n = 17, Day 4: n = 19, Month 1: n = 17). c, d Proportion and relative abundance of ASV types in the neonatal gut microbiome that were shared with the maternal vaginal, gut, or skin microbiome in cesarean-section-delivered offspring (Vagina Day 0: n = 10, Day 1: n = 13, Day 4: n = 22, Month 1: n = 20; Gut Day 0: n = 8, Day 1: n = 12, Day 4: n = 22, Month 1: n = 20; Skin Day 0: n = 6, Day 1: n = 10, Day 4: n = 18, Month 1: n = 17). e Comparison of the proportion and relative abundance of ASV types shared with the maternal gut microbiome between the gut microbiomes of vaginally delivered (Day 0: n = 16, Day 1: n = 17, Day 4: n = 19, Month 1: n = 17) and cesarean-section-delivered offspring (Day 0: n = 8, Day 1: n = 12, Day 4: n = 22, Month 1: n = 20; Skin Day 0: n = 6, Day 1: n = 10, Day 4: n = 18, Month 1: n = 17). f Comparison of the proportion and relative abundance of ASV types shared with the maternal vaginal microbiome between the gut microbiomes of vaginally delivered (Day 0: n = 16, Day 1: n = 17, Day 4: n = 20, Month 1: n = 18) and cesarean-section-delivered offspring (Vagina Day 0: n = 10, Day 1: n = 13, Day 4: n = 22, 96Month 1: n = 20). Each p-value in all Figures is shown in Supplementary Table 7. Data are presented as means ± standard errors of the mean. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, Mann–Whitney U test (two-sided) for comparisons of two groups in panels (a), (b), (e), and (f). Kruskal–Wallis test followed by Dunn’s test (two-sided) for multiple comparisons in panels (c) and (d).
In cesarean-section-delivered neonates, the proportion of ASV types shared with the maternal vaginal microbiome remained consistently low in the neonatal gut microbiome. In contrast, the proportion of ASV types shared with the maternal gut microbiome exhibited an increasing trend (Fig. 6c and Supplementary Table 7). The proportion of ASV types shared with the maternal skin microbiome exhibited a decreasing trend (Fig. 6c and Supplementary Table 7). At all time points, the proportion of ASV types shared with the maternal skin microbiome was significantly higher than that shared with the maternal vaginal microbiome (Day 0: p = 0.0063, Day 1: p = 0.0011, Day 4: p < 0.0001, Month 1: p = 0.0002) (Fig. 6c and Supplementary Table 7). Additionally, the proportion of ASV types shared with the maternal skin microbiome on Day 4 was significantly higher than that shared with the maternal gut microbiome (p = 0.0046) (Fig. 6c and Supplementary Table 7). The proportion of ASV types shared with the maternal gut microbiome was significantly higher than that shared with the maternal vaginal microbiome on Day 4 and at Month 1 (Day 4: p = 0.0204, Month 1: p = 0.0002) (Fig. 6c and Supplementary Table 7). Although the total proportion of ASV types shared with the maternal vaginal and gut microbiomes remained relatively stable in the neonatal gut microbiome across all time points, the proportion of ASV types shared with the maternal skin dramatically decreased over time. The relative abundance of ASVs shared with the maternal vaginal microbiome remained consistently low. In contrast, the relative abundance of ASVs shared with the maternal gut microbiome showed an increasing trend (Fig. 6d and Supplementary Table 7). Furthermore, the relative abundance of ASVs shared with the maternal skin microbiome exhibited a decreasing trend. At all time points, the relative abundance of ASVs shared with the maternal skin microbiome was significantly higher than that of ASVs shared with the maternal vaginal microbiome (Day 0: p = 0.0006, Day 1: p = 0.0010, Day 4: p < 0.0001, Month 1: p = 0.0026) (Fig. 6d and Supplementary Table 7). Additionally, the relative abundance of ASVs shared with the maternal gut microbiome was significantly higher than that of ASVs shared with the maternal vaginal microbiome on Day 4 and at Month 1 (Day 4: p < 0.0001, Month 1: p < 0.0001) (Fig. 6d and Supplementary Table 7). Overall, these results suggest that the neonatal gut microbiome undergoes a shift from being more similar to the maternal skin microbiome to being more strongly associated with the maternal gut microbiome.
The proportion of ASV types shared with the maternal gut microbiome was significantly higher in the neonatal gut microbiome of vaginally delivered neonates relative to cesarean-section-delivered neonates at Month 1 (p = 0.0216) (Fig. 6e and Supplementary Table 7). However, there was no significant difference in relative abundance. On Days 0 and 1, the gut microbiome of vaginally delivered neonates had a higher proportion of ASV types and a greater relative abundance of ASVs shared with the maternal vaginal microbiome compared with cesarean-section-delivered neonates (Fig. 6f and Supplementary Table 7). After Day 4, both the proportion and relative abundance of these ASVs declined more sharply in vaginally delivered neonates than in cesarean-section-delivered neonates. Consequently, the differences between modes of birth became less apparent (Fig. 6f and Supplementary Table 7).
Discussion
To our knowledge, this is the first longitudinal study to assess the maternal microbiome during pregnancy and the postpartum period, as well as the neonatal microbiome in early life, in a Japanese cohort. Microbiome compositions vary across geographic regions and ethnic groups22,23,30,31. For example, whereas multiple studies have demonstrated that Lactobacillus dominance in the vaginal microbiome provides protective effects against infections and supports maternal and neonatal health24,32,33, studies from the United States suggested that a diverse vaginal microbiome, rather than a Lactobacillus-dominated microbiome, has health implications among black women34,35. These findings underscore the importance of conducting microbiome research in diverse populations to define a universally applicable framework for a healthy microbiome. The present study provides new insights into the maternal and neonatal microbiomes in the Japanese population; it also contributes fundamental knowledge needed for a broader understanding of the perinatal microbiome.
We observed that the vaginal microbiome maintained low α-diversity with minimal changes in β-diversity throughout pregnancy. However, both α- and β-diversity were significantly altered at 1 month postpartum. Lactobacillus was the predominant bacterial genus in the vaginal microbiome during pregnancy, a finding consistent with previous studies36,37,38. Notably, in the present study, a higher relative abundance of Lactobacillus in early pregnancy was associated with a greater likelihood of pregnancy continuation beyond 38 weeks. In contrast, Lactobacillus abundance in late pregnancy did not show a significant association with pregnancy duration beyond 38 weeks. These findings suggest that the vaginal microbiome in early pregnancy plays a role in pregnancy progression. Mothers with a low abundance of Lactobacillus in early pregnancy had a lower pre-pregnancy BMI relative to those with a higher abundance of Lactobacillus. A low pre-pregnancy BMI has been associated with an increased risk of preterm birth39,40. Although the causal relationship and underlying mechanisms remain unclear, our findings suggest that both a low pre-pregnancy BMI and a low abundance of Lactobacillus in early pregnancy negatively impact the course of pregnancy.
The 16S rRNA amplicon sequencing method enables characterization of bacterial composition at the genus level but has limitations concerning species-level resolution41,42. Recently, Lebeer et al. introduced a methodology for analyzing the Lactobacillus genus in the vaginal microbiome at the subgenus level, allowing classification beyond the genus level29. Among Lactobacillus species, L. crispatus, L. iners, L. jensenii, and L. gasseri are predominant in the vaginal microbiome; their prevalences vary according to ethnicity and geographic region22,31,32,43. In this study, which focused on a Japanese cohort, analysis of Lactobacillus subgenera revealed that the L. crispatus group exhibited a consistently high abundance at all time points; this abundance increased toward late pregnancy. In contrast, a study of a Chinese cohort showed similar abundances of L. crispatus and L. iners43. These findings suggest that the predominant Lactobacillus species vary even among Asian populations. In this longitudinal study, dominance of the L. crispatus group was associated with lower α-diversity. Furthermore, when the L. crispatus group became dominant, other Lactobacillus subgenera did not replace it. These findings support the hypothesis that L. crispatus played a crucial role in stabilizing the vaginal microbiome in this Japanese cohort.
The maternal gut microbiome is suspected to influence pregnancy progression and neonatal outcomes9,10,11,44. Although some studies have demonstrated compositional changes in the maternal gut microbiome during pregnancy and the perinatal period45,46, others have detected no significant alterations during these stages7,47. Thus, the notion of substantial changes in the maternal gut microbiome during pregnancy and the perinatal period remains controversial. In the present study, the maternal gut microbiome remained relatively stable during early pregnancy, but showed temporal changes from 30 weeks of gestation to the postpartum period. Additionally, we identified associations between the maternal gut and vaginal microbiomes. The abundances of several bacterial genera (e.g., Lactobacillus and Anaerovoraceae_S5.A14a) in the gut were associated with the vaginal abundance of Lactobacillus, a key bacterial genus in the vaginal microbiome. Furthermore, different Lactobacillus subgenus in the vagina were associated with distinct bacterial genera in the gut. Based on these findings, we speculate that it could be a future research scope to examine whether the maternal gut microbiome during pregnancy can serve as a target for microbial interventions to improve vaginal health by increasing the abundance of beneficial bacteria, such as Lactobacillus.
The neonatal gut microbiome undergoes rapid changes and becomes established after birth, thereby influencing offspring development48,49. Our study showed that α-diversity in the neonatal gut microbiome initially declines before increasing again early in life. This pattern likely reflects the sequential development of the gut microbiome: (1) only microbes that are appropriate or essential for early life successfully colonize the neonatal gut among those vertically transmitted from the mother, and (2) the neonate continuously acquires microbes from the external environment. Demonstration of this dynamic early-life phenomenon is a key strength of the present study, which utilized a rigorous longitudinal design for close monitoring of neonates. Further translational studies are needed to identify mechanisms by which the host selectively acquires and retains specific microbes during gut microbiome development.
The mode of delivery (vaginal vs. cesarean section) substantially influences the neonatal microbiome15,16,17. Our UniFrac distance analysis revealed significant differences in bacterial composition between vaginally delivered and cesarean-section-delivered neonates on Day 1, Day 4, and at Month 1; no significant differences were observed on Day 0. Furthermore, association analyses indicated that bacterial genera differing between delivery modes varied across time points (Fig. 5g). These findings suggest that the initial gut microbiome composition at birth impacts the trajectory of subsequent microbiome development. This concept is consistent with findings from animal studies demonstrating that early-life colonization history plays a central role in establishing the eventual composition of the gut microbiome50. Given that various factors can influence the gut microbiome, an important question remains concerning whether the effect of delivery mode persists throughout life or is eventually overridden by other environmental and host-related factors.
The maternal microbiome is considered a key source of the neonatal gut microbiome in early life and has long-term implications for offspring health14,15. Therefore, an understanding of the mechanisms that underlie vertical microbiome transmission from mother to offspring may provide opportunities to explore microbiome-based interventions for disease prevention. In the present study, the proportions of ASV types in the neonatal gut microbiome that were also present in the maternal vaginal and gut microbiomes were approximately 1.70% and 17.02%, respectively, at Month 1. This finding suggests that neonates acquire numerous bacteria from the external environment. However, given that the early-life microbiome plays a critical role in long-term microbiome development17,51, the relatively low abundances of bacteria derived from the maternal vagina and gut do not necessarily imply that the maternal microbiome has limited physiological influence on offspring health. Vaginally delivered neonates exhibited a higher proportion of ASV types shared with the maternal vaginal microbiome on Day 0 and Day 1, as well as a significantly higher proportion of ASV types shared with the maternal gut microbiome at Month 1 relative to cesarean-section-delivered neonates. This observation supports the notion that the mode of delivery affects early colonization of the neonatal microbiome, such that vaginal delivery more strongly facilitates the transfer of maternal vaginal and gut microbiota compared with cesarean section delivery. In the gut microbiome of cesarean-section-delivered neonates, the proportion of ASV types shared with the maternal skin microbiome was higher than the proportions shared with the maternal vaginal and gut microbiomes.
This study had some limitations. First, it was conducted at a single center with a limited sample size. A larger, multi-center study would provide more robust evidence to support our findings. Our institution is one of the largest perinatal medical centers in western Tokyo, serving a population of approximately four million in both urban and rural areas. Although the facility attracts a diverse patient population because of its ability to provide advanced perinatal care, the possibility of selection bias cannot be excluded. Nevertheless, a key strength of this prospective study design was its collection of detailed clinical data regarding mothers and their offspring while ensuring high sample quality for microbiome analysis. Second, samples were collected 1 month after birth during routine health checkups, which, in Japan, are typically conducted at the clinic or hospital where the child was born. This study design is practical and feasible in a clinical setting, but a longer follow-up period could provide valuable insights into the relationship of microbiomes during pregnancy and the peripartum period with long-term clinical outcomes in children. Moreover, as the primary aim of this study was not to investigate the factors influencing the development of the neonatal gut microbiome, there are inherent limitations in adjusting for potential confounders. For instance, factors such as premature rupture of membranes, NICU admission, and feeding method (breastfeeding vs. formula feeding) may confound the analysis of gut microbiome development. Future studies should aim to address these limitations by adjusting for a broader range of potential confounders and incorporating longer-term follow-up to gain a more comprehensive understanding of neonatal gut microbiome development. Third, maternal skin samples were only collected from mothers who underwent cesarean section due to practical constraints in the clinical setting, preventing direct comparison of skin microbiome transmission between delivery modes.
Despite this limitation, our findings provide new insights into the relative contributions of different maternal microbiomes to neonatal rectal microbiota composition and how these contributions differed between vaginal and cesarean section deliveries in a Japanese cohort. Finally, this study utilized 16S rRNA gene amplicon sequencing to characterize bacterial composition but did not assess the mycobiome, virome, or functional aspects of the microbiome. Furthermore, this methodology has inherent limitations in resolving bacterial composition at the species level. For instance, within the genus Lactobacillus, it has been reported that major species such as L. crispatus, L. jensenii, L. gasseri, and L. iners exert distinct effects on host health32. Given that we detected longitudinal changes in bacterial composition in both mothers and offspring, future research should expand on these findings by exploring the broader microbial ecosystem.
In conclusion, this prospective study revealed maternal microbiome dynamics and neonatal microbiome development in a Japanese cohort during pregnancy and the peripartum period. It also provided insights into the impact of the early vaginal microbiome on pregnancy outcomes. Our findings suggest that L. crispatus in the vaginal microbiome plays crucial roles in maternal and neonatal health within the Japanese population. These results help to define the core components of a healthy human microbiome and underscore the importance of further microbial research across diverse populations.
Methods
Study design
This prospective, longitudinal cohort study enrolled pregnant women who visited the Department of Obstetrics and Gynecology at Kyorin University Hospital for peripartum care and provided written informed consent to participate. Participants were recruited between October 2021 and January 2022. Follow-up and data collection continued until September 2022. As outlined in Fig. 1a, maternal vaginal and rectal swabs were collected at weeks 12, 20, 30, and 36 of gestation during routine maternity checkups. At childbirth, vaginal and rectal swabs were collected from mothers who delivered vaginally; vaginal and skin swabs were collected from mothers who delivered via cesarean section. During hospitalization after delivery, rectal swabs were collected from offspring on days 0, 1, 4, and 7. At the 1-month postpartum health checkup, maternal vaginal and rectal swabs and neonatal rectal swabs were collected. Clinical information regarding mothers and offspring was obtained from the Kyorin University Hospital medical record system or through a questionnaire. No statistical method was used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
DNA extraction and 16S rRNA gene amplicon sequencing
All samples were collected using sterile swabs with the Puritan Opti-Swab Liquid Amies Collection & Transport System (Puritan Medical Products, Guilford, ME, USA). Microbe-containing reagents were frozen and stored at −80 °C until further processing. Samples were lyophilized using a freeze dryer (Freeze Dryer VD-250R, TAITEC, Saitama, Japan) for 16 h. After lyophilization, the dried samples were subjected to mechanical disruption by bead beating using the Multi-beads Shocker (Yasui Kikai, Osaka, Japan). Genomic DNA was then extracted using the GenCheck DNA Extraction Kit (FASMAC Co., Ltd., Kanagawa, Japan), following the manufacturer’s instructions. After DNA was extracted from frozen samples, the V4 region (PCR primer 515 F: 5’-GTGCCAGCMGCCGCGGTAA-3’, 806 R: 5’-GGACTACHVGGGTWTCTAAT-3’) of the 16S rRNA gene was amplified. Samples with low quality ( < 70% target amplicon detection on electrophoresis) were excluded from further analyses. Throughout the DNA extraction and sequencing workflow, quality control was conducted using negative controls. Sterile water was used as a template in the PCR mix during library preparation as a negative control, and samples were accepted only if DNA concentrations in the controls were below the detection limit as measured by Qubit fluorometry. PhiX was included as an internal sequencing control. Paired-end amplicon sequencing was performed (2 × 250 bp) using the MiSeq platform (Illumina, San Diego, CA, USA). Sequence data were processed using the Quantitative Insights into Microbiome Ecology (QIIME) 2 pipeline52. Sequences were denoised and filtered using DADA253, which also removed low-quality and non-target sequences, including internal controls such as PhiX. For downstream diversity analysis, the sampling depth was set to 5000 to normalize sequencing depth across samples. ASVs were taxonomically assigned using the Silva 13_8 database (https://docs.qiime2.org/2021.2/data-resources/). Bacterial community diversity was assessed via QIIME 2. Alpha diversity was evaluated using the Chao1 and Shannon diversity indices. Beta diversity was assessed with UniFrac distances and visualized by principal coordinates analysis (PCoA). The genus Lactobacillus was classified into subgenera as previously described by Lebeer et al.29. Briefly, to classify Lactobacillus at the subgenus level based on our 16S rRNA gene amplicon sequencing data, we used a reference database in which Lactobacillus sequences from the Genome Taxonomy Database (GTDB) (https://github.com/LebeerLab/Citizen-science-map-of-the-vaginal-microbiome) were annotated with nine subgenera as defined in prior work29.
Statistical analysis
All statistical analyses were conducted using GraphPad Prism (version 8.4.3) and R (version 4.1.3) (https://www.r-project.org/). The Mann–Whitney U test was used for comparisons between two groups; the Kruskal–Wallis test, followed by Dunn’s test, was utilized for comparisons among three or more groups. Fisher’s exact test was used to analyze contingency tables. The log-rank test was performed to compare Kaplan–Meier curves. For α-diversity comparisons across time points, linear mixed-effects models were applied using the lme4 package (https://github.com/lme4/lme4). Time point was modeled as a fixed effect, while individual participants were modeled as a random effect to account for repeated measures. Model fitting was performed using restricted maximum likelihood (REML). Pairwise comparisons were performed using estimated marginal means derived from the model via the emmeans package (https://github.com/rvlenth/emmeans), accounting for repeated measures, and adjusted for multiple comparisons using the Benjamini-Hochberg method. For β-diversity analysis, weighted and unweighted UniFrac distances were calculated using QIIME 2. Comparisons between groups were performed using permutational multivariate analysis of variance (PERMANOVA) with the adonis2 function in the vegan package (https://github.com/vegandevs/vegan). For comparisons across time points, the strata argument was applied to restrict permutations within each participant, thereby accounting for the repeated measures. Multivariable association analyses were performed with MaAsLin2 (https://github.com/biobakery/Maaslin2/), using the default settings as recommended54. Multivariable analyses included relative abundances of bacterial genera and clinical characteristics as explanatory variables. For each analysis, bacterial genera that were present in ≥ 5% of samples were included. All statistical analyses were performed using two-sided tests whenever applicable.
Ethical considerations
This study was conducted in accordance with the Declaration of Helsinki. The protocol was approved by the Institutional Ethics Committee of Kyorin University School of Medicine (Approval Number 789). Written informed consent was obtained from all participating mothers.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The microbial dataset reported in this study has been deposited in the DDBJ Sequence Read Archive (DRA) under accession number DRA020149 (https://www.ddbj.nig.ac.jp/dra/). The source data generated in this study are provided in the source data file. Source data are provided with this paper.
Code availability
This study did not use of custom code or proprietary algorithms. All computational analyses were conducted using publicly available packages. Reference codes in this study are provided in Supplementary Information.
References
Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021).
Ursell, L. K. et al. The intestinal metabolome: an intersection between microbiota and host. Gastroenterology 146, 1470–1476 (2014).
Janiak, M. C. et al. Age and sex-associated variation in the multi-site microbiome of an entire social group of free-ranging rhesus macaques. Microbiome 9, 68 (2021).
Zhou, X. et al. Longitudinal profiling of the microbiome at four body sites reveals core stability and individualized dynamics during health and disease. Cell Host Microbe 32, 506–526.e509 (2024).
Kindinger, L. M. et al. The interaction between vaginal microbiota, cervical length, and vaginal progesterone treatment for preterm birth risk. Microbiome 5, 6 (2017).
Zhao, F., Hu, X. & Ying, C. Advances in research on the relationship between vaginal microbiota and adverse pregnancy outcomes and gynaecological diseases. Microorganisms 11, 991 (2023).
DiGiulio, D. B. et al. Temporal and spatial variation of the human microbiota during pregnancy. Proc. Natl Acad. Sci. USA 112, 11060–11065 (2015).
Romero, R. et al. The composition and stability of the vaginal microbiota of normal pregnant women is different from that of non-pregnant women. Microbiome 2, 4 (2014).
Huang, L., Thonusin, C., Chattipakorn, N. & Chattipakorn, S. C. Impacts of gut microbiota on gestational diabetes mellitus: a comprehensive review. Eur. J. Nutr. 60, 2343–2360 (2021).
Chen, X. et al. Gut dysbiosis induces the development of pre-eclampsia through bacterial translocation. Gut 69, 513–522 (2020).
Kimura, I. et al. Maternal gut microbiota in pregnancy influences offspring metabolic phenotype in mice. Science 367, eaaw8429 (2020).
Vuong, H. E. et al. The maternal microbiome modulates fetal neurodevelopment in mice. Nature 586, 281–286 (2020).
Wang, S. et al. Maternal vertical transmission affecting early-life microbiota development. Trends Microbiol. 28, 28–45 (2020).
Ferretti, P. et al. Mother-to-infant microbial transmission from different body sites shapes the developing infant gut microbiome. Cell Host Microbe 24, 133–145.e135 (2018).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Wampach, L. et al. Birth mode is associated with the earliest strain-conferred gut microbiome functions and immunostimulatory potential. Nat. Commun. 9, 5091 (2018).
Shao, Y. et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574, 117–121 (2019).
Nielsen, S., Nielsen, D. S., Lauritzen, L., Jakobsen, M. & Michaelsen, K. F. Impact of diet on the intestinal microbiota in 10-month-old infants. J. Pediatr. Gastroenterol. Nutr. 44, 613–618 (2007).
Krebs, N. F. et al. Effects of different complementary feeding regimens on iron status and enteric microbiota in breastfed infants. J. Pediatr. 163, 416–423 (2013).
Navarro-Tapia, E. et al. Probiotic supplementation during the perinatal and infant period: effects on gut dysbiosis and disease. Nutrients 12, 2243 (2020).
Tamburini, S., Shen, N., Wu, H. C. & Clemente, J. C. The microbiome in early life: implications for health outcomes. Nat. Med. 22, 713–722 (2016).
Gupta, V. K., Paul, S. & Dutta, C. Geography, ethnicity or subsistence-specific variations in human microbiome composition and diversity. Front Microbiol. 8, 1162 (2017).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Fettweis, J. M. et al. The vaginal microbiome and preterm birth. Nat. Med. 25, 1012–1021 (2019).
Nishijima, S. et al. The gut microbiome of healthy Japanese and its microbial and functional uniqueness. DNA Res. 23, 125–133 (2016).
Yoshisue, H., Homma, Y., Ito, C. & Ebisawa, M. Prevalence of food allergy increased 1.7 times in the past 10 years among Japanese patients below 6 years of age. Pediatr. Allergy Immunol. 35, e14192 (2024).
Okabayashi, S., Kobayashi, T. & Hibi, T. Inflammatory bowel disease in Japan-Is it similar to or different from Westerns? J. Anus Rectum Colon 4, 1–13 (2020).
Eguchi, Y. et al. Epidemiology of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis in Japan: a focused literature review. JGH Open 4, 808–817 (2020).
Lebeer, S. et al. A citizen-science-enabled catalogue of the vaginal microbiome and associated factors. Nat. Microbiol 8, 2183–2195 (2023).
He, Y. et al. Regional variation limits the applications of healthy gut microbiome reference ranges and disease models. Nat. Med. 24, 1532–1535 (2018).
Deschasaux, M. et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat. Med. 24, 1526–1531 (2018).
Ravel, J. et al. Vaginal microbiome of reproductive-age women. Proc. Natl Acad. Sci. USA 108, 4680–4687 (2011).
Aldunate, M. et al. Antimicrobial and immune modulatory effects of lactic acid and short chain fatty acids produced by vaginal microbiota associated with eubiosis and bacterial vaginosis. Front. Physiol. 6, 164 (2015).
Callahan, B. J. et al. Replication and refinement of a vaginal microbial signature of preterm birth in two racially distinct cohorts of US women. Proc. Natl Acad. Sci. USA 114, 9966–9971 (2017).
Fettweis, J. M. et al. Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiol. (Read.) 160, 2272–2282 (2014).
Gomez de Agüero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
Nunn, K. L. et al. Changes in the vaginal microbiome during the pregnancy to postpartum transition. Reprod. Sci. 28, 1996–2005 (2021).
Shin, H. et al. Partial convergence of the human vaginal and rectal maternal microbiota in late gestation and early post-partum. NPJ Biofilms Microbiomes 9, 37 (2023).
Girsen, A. I. et al. Women’s prepregnancy underweight as a risk factor for preterm birth: a retrospective study. BJOG 123, 2001–2007 (2016).
Cornish, R. P. et al. Maternal pre-pregnancy body mass index and risk of preterm birth: a collaboration using large routine health datasets. BMC Med. 22, 10 (2024).
Qing, W. et al. Species-level resolution for the vaginal microbiota with short amplicons. mSystems 9, e0103923 (2024).
O’Callaghan, J. L., Willner, D., Buttini, M., Huygens, F. & Pelzer, E. S. Limitations of 16S rRNA gene sequencing to characterize Lactobacillus species in the upper genital tract. Front. Cell Dev. Biol. 9, 641921 (2021).
Ng, S. et al. Large-scale characterisation of the pregnancy vaginal microbiome and sialidase activity in a low-risk Chinese population. NPJ Biofilms Microbiomes 7, 89 (2021).
Koren, O., Konnikova, L., Brodin, P., Mysorekar, I. U. & Collado, M. C. The maternal gut microbiome in pregnancy: implications for the developing immune system. Nat. Rev. Gastroenterol. Hepatol. 21, 35–45 (2024).
Koren, O. et al. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150, 470–480 (2012).
Nuriel-Ohayon, M. et al. Progesterone increases Bifidobacterium relative abundance during late pregnancy. Cell Rep. 27, 730–736.e733 (2019).
Dunlop, A. L. et al. Stability of the vaginal, oral, and gut microbiota across pregnancy among African American women: the effect of socioeconomic status and antibiotic exposure. PeerJ 7, e8004 (2019).
Putignani, L., Del Chierico, F., Petrucca, A., Vernocchi, P. & Dallapiccola, B. The human gut microbiota: a dynamic interplay with the host from birth to senescence is settled during childhood. Pediatr. Res. 76, 2–10 (2014).
Gritz, E. C. & Bhandari, V. The human neonatal gut microbiome: a brief review. Front. Pediatr. 3, 17 (2015).
Martínez, I. et al. Experimental evaluation of the importance of colonization history in early-life gut microbiota assembly. Elife 7, e36521 (2018).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput Biol. 17, e1009442 (2021).
Acknowledgments
This work was supported in part by the Kyorin University School of Medicine Joint Project Grant. We thank Ryan Chastain-Gross, Ph.D., from Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
N.O., C.K., Y.O., M.N., S.T., T.Hanawa, J.M. and T.Hisamatsu conceived and designed the study. N.O., C.K., Y.O. and J.M. drafted the manuscript. N.O., C.K., Y.O., T.K., Y.N., H.W., N.N. and J.M. collected samples and clinical data; they also performed data analysis. M.N., S.T., T.Hanawa, and T.Hisamatsu supervised the manuscript. M.N., S.T., T.Hanawa, J.M., and T.Hisamatsu oversaw the entire project.
Corresponding authors
Ethics declarations
Competing interests
J.M. has received lecture fees from EA Pharma Co., Ltd., AbbVie GK, Janssen Pharmaceutical K.K., Pfizer Inc., Mitsubishi Tanabe Pharma Corporation, JIMRO Co., Miyarisan Co., Ltd., Nippon Kayaku Co., Ltd., Mochida Pharmaceutical Co., Ltd., and Takeda Pharmaceutical Co., Ltd. M.N. has received lecture fees from Miyarisan Co., Ltd. T.Hisamatsu. has received grant support from Mitsubishi Tanabe Pharma Corporation, EA Pharma Co., Ltd., AbbVie GK, JIMRO Co., Ltd., Zeria Pharmaceutical Co., Ltd., Kyorin Pharmaceutical Co., Ltd., Nippon Kayaku Co., Ltd., Takeda Pharmaceutical Co., Ltd., Pfizer Inc., Mochida Pharmaceutical Co., Ltd., Boston Scientific Corporation, and Kissei Pharmaceutical Co., Ltd; received consulting fees from Mitsubishi Tanabe Pharma Corporation, EA Pharma Co., Ltd., AbbVie GK, Janssen Pharmaceutical K.K., Pfizer Inc., Eli Lilly, Gilead Sciences, Bristol Myers Squibb, and Abivax; and received lecture fees from Mitsubishi Tanabe Pharma Corporation, AbbVie GK, EA Pharma Co., Ltd., Kyorin Pharmaceutical Co., Ltd., JIMRO Co., Ltd., Janssen Pharmaceutical K.K., Mochida Pharmaceutical Co., Ltd., Takeda Pharmaceutical Co., Ltd., Pfizer Inc., and Kissei Pharmaceutical Co., Ltd. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Trishla Sinha, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Oguri, N., Kobayashi, C., Ozawa, Y. et al. Vaginal Lactobacillus crispatus in early pregnancy associates with favorable gestational outcomes in a Japanese maternal-neonatal microbiome cohort. Nat Commun 16, 8032 (2025). https://doi.org/10.1038/s41467-025-63466-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63466-3