Abstract
Drug resistance in bacteria is a major problem that calls for new classes of antimicrobial drugs. We report a biodegradable poly(imidazolium ester) (PIE), P8, with excellent broad-spectrum antibacterial activity, high therapeutic selectivity, and an unexploited mechanism of action. P8, a short oligomer, translocates across bacterial membrane and phase separates with intracellular nucleic acids, forming biomolecular condensates. P8 binds the DNA minor groove and intercalates with DNA, interacting with it via electrostatic and hydrogen-bonding interactions. The phase separation of nucleic acids modulated by P8 inhibits in vitro transcription, thereby impeding translation and potentially leading to cell death. Bacterial cytological profiling indicates that the antimicrobial mechanism of P8 differs from those of conventional antibiotics, though it also suggests that P8 may inhibit RNA synthesis. P8 is safe and effective against drug-resistant bacteria in murine models of systemic, intramuscular, and lung infections. This study shows the great potential of intracellular biomolecular condensate formation for combating drug-resistant bacteria.
Similar content being viewed by others
Introduction
Infections due to multidrug-resistant (MDR) bacteria are a major worldwide public health problem. Many bacteria have become resistant to antibiotics, which are mostly “old” drugs; there has been a concerning dearth of new approved antibiotic classes, especially against Gram-negative bacteria, in the past 5–6 decades1. The rise and prevalence of MDR bacteria call for new antimicrobials with modes of action distinct from those of antibiotics currently in clinical use. Antimicrobial polymers and peptides are usually cationic and typically kill bacteria via physical disruption of bacterial membranes2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20. Cell-penetrating antimicrobial polymers and peptides are a distinct class that have intracellular targets, although the exact mechanisms are not well-understood21,22,23,24,25,26,27. Cationic polymers with new mechanisms of action may potentially combat bacteria that have developed resistance even to last-resort antibiotics such as carbapenem and colistin28,29.
Recently, it has been found that liquid-liquid phase separation (LLPS) mediates the organisation of bacterial subcellular biomolecules into membraneless organelles (also referred to as biomolecular condensates)30,31,32,33,34,35,36,37,38. Examples of intrinsic bacterial biomolecular condensates include ribonucleoprotein bodies (BR-bodies) and RNA polymerase (RNAP) clusters. These bacterial biomolecular condensates typically involve two classes of high molecular weight biomacromolecules, namely nucleic acids (RNA and DNA) and proteins30. The LLPS of protein/nuclei acid is thought to be facilitated by collective weak non-covalent interactions that are repetitive due to the protein/nuclei acid multivalency, particularly in the intrinsically disordered regions35. Possible weak reversible forces could be van der Waals forces, hydrogen bonding, and electrostatic, or cationic-π interactions, etc. A recent study showed that antimicrobial peptides, specifically P113, Os-C, and Buforin, form biomolecular condensates with nucleic acids inside bacteria, potentially contributing to their antimicrobial mechanisms39. However, exploitation of LLPS within bacteria in general, particularly with extrinsic additives causing antimicrobial effects, is largely unexplored and not well-studied.
In this study, we designed and synthesized biodegradable poly(imidazolium ester)s (PIEs) with the introduction of labile ester linkages into the alternating copolymers (Fig. 1a). By optimizing the hydrophobicity of linkages in PIEs, we found an optimized PIE (specifically P8) with optimal balance of intermediate degradation rate, strong in vitro antibacterial activity, and high therapeutic selectivity. P8 demonstrated a non-classical mechanism of action by translocating bacterial membrane and then phase separating with intracellular nucleic acids, leading to the formation of biomolecular condensates composed of polymer and nucleic acid (hereafter denoted as polymer@nuclei acid). This phase separation was found to hinder transcription and translation in vitro, likely leading to eventual bacterial cell death. Bacterial cytological profiling (BCP) was employed to confirm the uniqueness of P8’s mode of action. Notably, P8 showed potent therapeutic efficacy against drug-resistant bacteria in murine models of intramuscular (thigh) infection, systemic infection, and lung infection.
a Chemical synthesis; b, c bacterial resistance evolution to P8 and antibiotic controls; d–g P8’s membrane activity. a Chemical syntheses of poly(imidazolium ester)s (PIEs) (P1-P15). b Bacterial resistance evolutions of P. aeruginosa PAO1 after 20-day daily serial passaging in the presence of P8 or the conventional antibiotics (ciprofloxacin (CIP) and gentamycin (GEN)). c Bacterial resistance evolution of E. coli ATCC 8739 after 20-day daily serial passaging in the presence of P8, CIP or GEN. d The relative PI fluorescence intensity (F.I.) of untreated (UNT), P8-treated and colistin-treated P. aeruginosa PAO1. The treatment concentration of P8 and colistin was 16 µg/mL. e Time-lapse confocal images of P. aeruginosa PAO1 treated with FITC-P8 (8 µg/mL) in the presence of PI. f Cryo-TEM images of P. aeruginosa PAO1 treated with P8 (16 µg/mL). g Cryo-TEM images of untreated P. aeruginosa PAO1.
Results
Synthesis and optimisation of poly(imidazolium ester)s (PIEs)
Alternating copolymers poly(imidazolium ester)s (PIEs) were synthesized by the polycondensation of dibromides possessing an ester linkage (EL), with bisimidazoles possessing an alkyl linkage (AL) (Fig. 1a and Supplementary Fig. S1). Our compositional design encompassed variations of both the degradable EL and the non-degradable AL in the dibromide and bisimidazole molecules, respectively. Five EL variations were used: α-bromide ester (p = 1, q = 3), β-bromide ester (p = 2, q = 2 or 3) and γ-bromide ester (p = 3, q = 2 or 3). Furthermore, three AL variations were investigated: propyl (r = 3), butyl (r = 4) and pentyl (r = 5). A library of 15 perfect alternating PIEs denoted as P1-P15 was successfully synthesized (see Methods) and characterized via NMR (Supplementary Figs. S2–S24), FTIR (Supplementary Fig. S25) and Gel Permeation Chromatography (GPC) (Supplementary Fig. S26). PIEs displayed a molecular weight range of 760–1081 Da as determined by GPC (Supplementary Fig. S26). However, molecular weights obtained via GPC are only approximate values. Notably, the polycondensation reaction mixture gradually transforms from a liquid phase to a solid phase as bulk polymerization progresses (Supplementary Fig. S27a), putting an upper limit on PIE’s length. Meanwhile, dialysis (using membranes with a Molecular Weight Cut-Off (MWCO) of 1 K) against DI water was conducted to remove relatively low-molecular-weight oligomers. Collectively, oligomers with relatively narrow distributions were obtained. This synthesis was repeated numerous times, and there were little variations in molecular weight from batch to batch (Supplementary Fig. S27b–d). Mass spectrometry also revealed short polymer lengths (Supplementary Fig. S28). The counterion in the final PIEs was changed from bromide to chloride, and this was verified by X-ray photoelectron spectroscopy (XPS) (Supplementary Fig. S29).
To verify the degradability of PIEs, we investigated the hydrolysis of the ester bond with 1H NMR spectroscopy (Supplementary Fig. S30). All PIEs demonstrated degradability in PBS (pH 7.2 ± 0.2) at 37 °C, with the degradation rate varying depending on composition (Supplementary Fig. S31a–c). The fastest degradation rate was observed in the poly(α-imidazolium ester)s (α-PIEs) (p = 1, P1–P3). The poly(β-imidazolium ester)s (β-PIEs) (p = 2, P4–P9) and poly(γ-imidazolium ester)s (γ-PIEs) (p = 3, P10–P15) showed reduced degradation rates. This variation is unsurprising as the ester hydrolysis happens via an anionic transition state that is stabilised by the electron-withdrawing imidazolium: the nearer the cationic imidazolium ring (with p = 1 versus p = 2 or 3) is to the ester transition group, the stronger the stabilization of the latter leading to faster degradation40. GPC measurements following incubation in PBS (pH 7.2 ± 0.2) at 37 oC show the peaks shifted to longer retention time corroborating the degradability of these PIEs (Supplementary Fig. S31d).
We assessed the antimicrobial efficacy of the PIEs against ESKAPE pathogens (i.e., Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter spp.) by measuring the minimum inhibitory concentrations (MICs). As indicated in Table 1, α-PIEs (P1–P3) displayed poor antimicrobial activity with generally high (geometric) mean MICs (MICGM) in the range of 35.3 to >141.3 μg/ml. However, a substantial improvement was observed for β-PIEs (P4–P9) and γ-PIEs (P10–P15) with MICGM values ranging from 4.9 to 29 μg/ml and 4.9 to 16.0 μg/ml, respectively. We attribute these differences to the significantly faster degradation rate of α-PIEs compared to β-PIEs and γ-PIEs. To rule out the effect of hydrophobicity, we compared the antimicrobial efficacy of P1 with P4, P2 with P5, and P3 with P6, as each pair has the same number of carbon atoms in their EL (p + q = 4) and AL (with the same r). We found that P4 to P6 exhibited significantly lower MICGM compared with P1 to P3, respectively, though each pair has the same total number of carbon atoms in their linkages. However, P4 to P6 degrade more slowly than P1 to P3 (Supplementary Fig. S31a–c). This corroborates that the rapid degradation of α-PIEs contributes to their poor antimicrobial performance. We also observed the faster degradation-induced deactivation of antimicrobial effect for an α-PIE (P2) as compared with the β-PIE (P8) or γ-PIE (P14) (Supplementary Fig. S31e, f). Amongst the polymers, P8 and P14 demonstrated good potency, with both having MICGM’s of 4.9 µg/mL.
We also determined the in vitro toxicity of our polymers against 3T3 fibroblast cells by measuring their half inhibitory concentrations (IC50) (Table 1 and Supplementary Fig. S32). The selective index (SI, defined as IC50/MICGM) is also provided in Table 1. P8 outperformed P14 in having lower cytotoxicity, as indicated by its higher IC50 and higher therapeutic selectivity index: >1024 µg/mL and >208 versus 175 and 36, respectively (Table 1). Furthermore, when P8 was tested further with human embryonic kidney cells (HEK) and human adenocarcinomic lung epithelial cells (A549), it also showed low toxicity with selectivity index (SI) > 208 (Supplementary Fig. S33a, b). When administered to RAW 264.7 macrophages, P8 at 1–4 × MIC (MIC against P. aeruginosa PAO1: 4 μg/ml) did not trigger any immunogenic response with low secreted TNF-α and arginase levels comparable to those observed with the cytocompatible poly(lactic-co-glycolic acid) (PLGA) and polyethylene glycol (PEG) treatments (Supplementary Fig. S33c, d).
We further evaluated P8’s antimicrobial efficacy against various clinically relevant isolates which are resistant to classical antibiotics, including imipenem (IMP), gentamicin (GEN), ciprofloxacin (CIP), and colistin (CST). P8 demonstrated high effectiveness against these MDR bacteria with MICs of 4–8 µg/mL (Table 2). Further, to investigate the tendency of bacteria to acquire resistance to P8, we serially passaged P. aeruginosa (Fig. 1b) and E. coli (Fig. 1c) in the presence of P8 or antibiotic controls. Remarkably, P8 did not induce bacterial resistance in both P. aeruginosa and E. coli. In stark contrast, the conventional antibiotics, specifically CIP and GEN, led to significant resistance evolution after 20 serial passages with final MICs 10–120 times higher than the initial MICs. After 50 serial passages, P8, like the case with colistin, still did not induce bacterial resistance in P. aeruginosa (Supplementary Fig. S34). Moreover, we also tested the resistance frequency and found that the selection of escape mutants to P8 was unsuccessful, showing that the propensity for resistance is low (with frequency below 1 × 10−9). In contrast, imipenem-resistant strains developed.
P8’s membrane translocation
To determine the interaction of P8 with bacterial membrane, i.e., whether it involves membrane disruption or membrane translocation, we examined the membrane integrity of P8-treated P. aeruginosa PAO1 using the propidium iodide (PI) dye, which specifically stains DNA of membrane-disrupted bacterial cells. Although both P8 and colistin (a well-known membrane-disruptive antibiotic) fully eradicated P. aeruginosa PAO1 (Supplementary Fig. S35), cells treated with P8 showed a significantly lower PI fluorescence signal (similar to the untreated control) compared to colistin-treated cells (Fig. 1d). This suggests that P8 does not disrupt the bacterial membrane. The lack of membrane disruption was further corroborated by insignificant calcein dye leakage in a model liposome assay (Supplementary Fig. S36). To further investigate the polymer-membrane interaction, we used FITC-conjugated P8 (FITC-P8) to treat P. aeruginosa PAO1 in the presence of PI and imaged the treated cells by confocal microscopy. Time-lapse confocal microscopy showed no PI fluorescence signal while P8 progressively entered the bacterial cytoplasm with time (Fig. 1e). These results indicated that P8 translocated across the bacterial membrane without compromising the membrane integrity. Notably, the bacterial colony-forming unit (CFU) counts under the same conditions revealed progressive cell death as time increased (Supplementary Fig. S37), suggesting that P8’s killing mechanism involves targets other than the bacterial membrane.
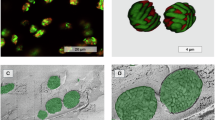
We examined the cell morphology of P8-treated and untreated P. aeruginosa PAO1 using cryo-transmission electron microscopy (cryo-TEM). The membrane of P8-treated cells remained intact (Fig. 1f(i), (ii) and Supplementary Fig. S38a(i), (ii)), similar to that of the untreated controls (Fig. 1g(i), (ii) and Supplementary Fig. S38b(i), (ii)), corroborating the PI test results. Closer examination of the cryo-TEM images revealed large internal aggregates with sizes ranging from 100 nm to 150 nm in treated cells (arrows in Fig. 1f(iii) and Supplementary Fig. S38a(iii)), which are significantly larger than those (20-50 nm) observed in untreated cells (arrows in Fig. 1g(iii) and Supplementary Fig. S38b(iii)). These clusters may represent biomolecular condensates caused by P8.
We also tracked the membrane translocation of FITC-P8 into 3T3 cells and found minimal FITC fluorescence in the mammalian cells compared with that in bacteria (Supplementary Fig. S39a, b). Quantitative analysis revealed that the level of FITC in the 3T3 cells was lower than that in bacterial cytoplasm (Supplementary Fig. S39c), highlighting the limited membrane translocation in mammalian cells, which may contribute to the good cytocompatibility and selectivity of P8 for bacteria over mammalian cells. This aligns with our previous study that oligoimidazolium complexes with anionic lipids, which aids bacterial membrane translocation41.
In vitro nucleic acid phase separation modulated by P8
Given the multivalency and intrinsic cationic nature of P8, we speculate that P8 may have a high propensity to phase separate with nucleic acids, which could represent a key intracellular mechanism of action subsequent to membrane translocation. Mixing of P8 with nucleic acids, specifically short double-stranded DNA (dsDNA-s) (~52 bp) or a long single-stranded polyadenylic acid poly(A) (2020–10630 bp), in physiological salt solutions (150 mM KCl, 1 mM MgCl2) results in the formation of opaque turbid solutions (Fig. 2a), suggesting the presence of light-scattering assemblies. When Syto59-stained nucleic acids (dsDNA and poly(A)) were mixed with FITC-P8 and imaged under confocal microscopy, we observed liquid-like assemblies displaying both Syto59 and FITC fluorescence signals (Fig. 2b). Notably, when P8, dsDNA-s, or poly(A) was present individually, no assemblies were observed (Supplementary Fig. S40). Further, P8@nucleic acid assemblies exhibited a high degree of roundness (>0.9) (Fig. 2c), and assembly fusion was observed (Fig. 2d and Supplementary Movies S1-S6), indicating the liquid nature of P8@nucleic acid assemblies. The fusion process was corroborated by the progressive enlargement of the diameter of assemblies (Supplementary Fig. S41). To explore whether P8 diffused within the assemblies, we analyzed the fluorescence recovery after photobleaching (FRAP) of FITC-P8 in P8@nucleic acid assemblies (Fig. 2e and Supplementary Movie S7–S12). We observed fluorescence recovery after photobleaching, with a half-life of fluorescence recovery of 26 s and 37 s for P8@dsDNA-s and P8@Poly(A) samples (n = 3), respectively. The short half-life of fluorescent P8 within the assemblies suggests its high mobility, confirming that P8 undergoes liquid-liquid phase separation (LLPS) with nucleic acids, resulting in the formation of liquid P8@nucleic acid biomolecular condensates.
a Digital images of P8@dsDNA-s and P8@Poly(A) condensates formed in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2 and 50 mM HEPES (pH 7.3). b Representative confocal microscopy image of P8@dsDNA-s and P8@Poly(A) condensates formed in above condition. P8 was labelled with FITC. Poly(A) and dsDNA-s were stained with Syto59. c The roundness of P8@dsDNA-s (n = 117) and P8@Poly(A) (n = 168) condensates formed in (b). d Representative microscopy images of P8@dsDNA-s and P8@Poly(A) condensate fusion. e Representative images and dynamics of fluorescence recovery of P8@dsDNA-s and P8@Poly(A) condensates after photobleaching (the number of tested condensates n = 3). The right plots show fluorescence recovery (%) quantified by fitting the average normalized intensity (solid dots with error bars) to first order exponential decay model (red and blue lines). Phase diagrams of P8 with f dsDNA-s, g poly(A) and h eGFP plasmid DNA. i Fluorescence quenching of the dye-dsDNA-s complex in the presence of P8. Ethidium bromide (DNA intercalator) and Hoechst 33342 (minor groove binder) were used as DNA dyes (the number of dye-dsDNA-s complex sample n = 3). j Visual appearance of P8@dsDNA-s and P8@Poly(A) condensates without (−) and with 10% w/v 1,6-HD, 800 mM KCl or 8 M urea. The turbidity of P8@dsDNA-s and P8@Poly(A) condensates after introduction of k 1,6-HD, l KCl and m urea at various concentrations (n = 3). Data are presented as mean ± standard deviation.
We investigated P8’s ability to phase separate with nucleic acids by constructing phase diagrams. FITC-P8 and three different nucleic acids, including the previously employed dsDNA-s (~52 bp), poly(A) (2,020-10,630 bp), and a new one, eGFP plasmid DNA (~5,000 bp), at different concentrations were mixed and imaged with confocal microscopy to determine the presence or absence of condensates (Supplementary Fig. S42). The lowest effective concentrations (LECs) of P8 required to induce nucleic acid phase separation were 2 µg/mL, 0.25 µg/mL and 0.25 µg/mL for dsDNA-s (Fig. 2f), poly(A) (Fig. 2g) and eGFP plasmid DNA (Fig. 2h), respectively. These LECs were not higher than the MICs (4–8 µg/mL) of P8, strongly suggesting that internalized P8 may induce phase separation of intracellular nucleic acids, which may contribute to its antimicrobial effects. Moreover, comparison of LEC values for the three different nucleic acids suggests that P8’s ability to phase separate with nucleic acids varies with the length of the nucleic acid, in line with a previous report42.
To study P8’s interaction sites with dsDNA, we conducted competitive displacement studies of P8 against DNA-bound dyes, specifically Hoechst 33342 (a representative DNA minor groove-binding dye) and ethidium bromide (a representative DNA-intercalating dye). We observed a dose-dependent quenching effect of P8 on the fluorescence of both dyes from the dye-dsDNA-s complexes (Fig. 2i), indicating that P8 functions as both a minor groove binder and an intercalator.
Next, we investigated the driving forces for the LLPS of P8 with nucleic acids by introducing various chemicals into preformed P8@dsDNA and P8@Poly(A) condensates and examining their effects on condensate turbidity and morphology. The introduction of 1,6-hexanediol (1,6-HD), an aliphatic alcohol known to disrupt liquid-like assemblies by impeding hydrophobic interactions43, led to insignificant changes in the condensate turbidity (Fig. 2j, k) and morphology (Supplementary Fig. S43). This suggests that hydrophobic interactions do not play a significant role in P8’s LLPS with nucleic acids. Introduction of additional KCl to disrupt electrostatic interactions resulted in a significant reduction in condensate turbidity (Fig. 2j and Fig. 2l) and the complete absence of condensates at additional 800 mM KCl (Supplementary Fig. S43), implying that electrostatic interaction contributes to the LLPS of P8 with nucleic acids. The introduction of urea, a hydrogen-bonding disruptor44, resulted in the turbidity reduction (Fig. 2j and 2m) and fewer detectable condensates (Supplementary Fig. S43). This indicates that hydrogen bonding also contributes to the LLPS of P8 with nucleic acids. In summary, P8 undergoes LLPS with nucleic acids through electrostatic interactions and hydrogen bonding.
In cellulo nucleic acid phase separation modulated by P8
After demonstrating that P8 undergoes LLPS with nucleic acids in vitro, we further investigated this dynamic process in cellulo, i.e., inside bacteria. In untreated P. aeruginosa PAO1, Syto59 stained the nucleic acids homogenously throughout the bacterial cytoplasm (Fig. 3a(i)). In FITC-P8-treated P. aeruginosa PAO1, distinct nucleic acid clusters were observed (yellow arrows in Fig. 3a(ii)). The Syto59 clusters colocalized with FITC-P8 clusters very well, as shown by the overlapping fluorescence profiles of Syto59 and FITC (Fig. 3b) along line 1 in Fig. 3a(ii). These clusters may represent biomolecular condensates composed of (FITC-)P8 and (Syto59-)nucleic acids. We denote the Syto59-rich cluster and the cytoplasm region outside the Syto59-rich cluster as condensate and dilute phase, respectively. Approximately 74% of overall Syto59 fluorescence occurred in the condensate phase, while 26% was from the dilute phase (n = 11) (Fig. 3c). These findings indicate that the majority of the nucleic acids were sequestered into P8@nucleic acid clusters. We have also studied the mechanism of another polymer in the library (P14) and found that P14 can also cross the bacterial membrane and form biomolecular condensates with nucleic acids (Supplementary Fig. S44).
a–e P8 modulates nucleic acid phase separation in cellulo; f–i P8 inhibits transcription and translation in vitro; (j-k) mechanism of action of P8 by bacterial cytological profiling. a Fluorescence microscopy of P. aeruginosa PAO1 (i) without and (ii) with FITC-P8 treatment. The nucleic acids in P. aeruginosa PAO1 were stained with Syto59. The yellow arrows indicate nucleic acid clusters. The experiment was repeated three times. b Fluorescence intensity (F.I.) profiles of Syto59/FITC along the line 1 in (a). c Fluorescence intensity percent (F.I. percent (%)) in condensate and dilute phases (the number of bacteria n = 11). d Time-lapse confocal images of P. aeruginosa PAO1 treated with FITC-P8. Syto59 was used to stain nucleic acids. e Time-lapse confocal images of untreated (i) and P8-treated ((ii) and (iii)) P. aeruginosa PAO1. white arrows indicate the formation of nucleic acid condensates. The dashed lines indicate the fusion process of nucleic acid condensates. Scale bar: 200 nm. f In vitro transcription reaction in the absence (0 µg/mL) and presence of varying concentrations of P8 (1, 2, 4, 8 and 16 µg/mL). L refers to RNA ladder (0.5, 1, 1.5, 2, 2.5, 3, 4, 5, 6 and 9 kb). g Cell-free eGFP translation initiated with eGFP plasmid DNA in the absence (0 µg/mL) and presence of P8 (2, 4, 8, 16 32 and 64 µg/mL) (n = 3). h Cell-free eGFP synthesis initiated with mRNA in the absence (0 µg/mL) and presence of P8 (16, 32, 64 and 128 µg/mL) (n = 3). i Half maximal effective concentration of P8 against CFPS (EC50) in plasmid DNA- or mRNA-initiated CFPS reaction (n = 3). j PCA graph using PC1 (63.95%) and PC2 (19.37%). PCA analysis was based on bacterial cell morphology, DNA parameters and SYTOX Green intensity signals. k PCA graph using PC1 (74.54%) and PC2 (22.82%). PCA analysis was based on bacterial cell morphology and SYTOX Green intensity signals. Three independent cultures of B. subtilis ATCC 6633 were treated with each antibiotic or P8. Data are presented as mean ± standard deviation.
The process of P8@nucleic acid cluster formation was further investigated in P. aeruginosa PAO1 treated with FITC-P8 by time-lapse confocal microscopy (Fig. 3d and Supplementary Movie S13–S15). At 0 min, FITC-P8 was localized at the bacterial membrane. At the 2nd minute, FITC-P8 accumulated in the cytoplasm, and both FITC-P8 and Syto59-stained nucleic acids were roughly homogeneously distributed throughout the cytoplasm. From 5 minutes onward, co-localized FITC-rich and Syto59-rich clusters formed and became increasingly compacted. These results suggest that internalized P8 may undergo phase separation with intracellular nucleic acids, leading to the formation of P8@nucleic acid clusters. In contrast, in untreated cells, Syto59-stained nucleic acids were distributed evenly, with no Syto59-rich clusters observed during the test (Fig. 3e(i)). Notably, in P8-treated P. aeruginosa PAO1 (Fig. 3e(ii), (iii)), Syto59-positive lumps (white arrows), representing nucleic acid clusters, were observed to fuse into more compact and rounded structures, demonstrating their liquid-like properties. In summary, these results suggest that the internalized P8 may phase separate with intracellular nucleic acids, leading to the formation of liquid-like P8@nucleic acid condensates.
To investigate how P8-modulated nucleic acid phase separation affects biological processes, we performed in vitro transcription (IVT) assays using a T7 promoter-driven, eGFP-encoding linearized DNA fragment (eGFP DNA fragment) in the presence of P8. We varied P8 concentrations to study its effect on the level of transcribed mRNA (Fig. 3f). Further, we also examined P8-modulated phase separation of the eGFP DNA fragment (stained with Syto59) in IVT assays using confocal microscopy (Supplementary Fig. S45). At low P8 concentrations (1 and 2 µg/mL), no condensates were observed, which correlates with the absence of an inhibitory effect on the transcription. However, at higher P8 concentrations (4, 8, and 16 µg/mL), eGFP DNA fragment condensates were detected, which correlates with the transcription inhibition. These results suggest that P8-modulated nucleic acid phase separation inhibits transcription.
To test whether P8-modulated nucleic acid phase separation also inhibited translation, we conducted cell-free protein synthesis (CFPS) assays with and without P8. Using an eGFP plasmid DNA containing a T7 promoter (Fig. 3g) or eGFP-encoding mRNA (Fig. 3h), we monitored transcription and translation, or translation alone, respectively, by measuring eGFP production. We observed a decrease in eGFP level with increasing P8 concentrations (Fig. 3g, h), which was more pronounced with the plasmid DNA than with the mRNA. Further, by fitting the plasmid DNA- and mRNA-programmed CFPS reaction kinetics data to a simple model describing transcription and translation (Supplementary Fig. S46)39, we confirmed that the inhibition of eGFP production by P8 was more prominent when the reaction was programmed with the plasmid DNA, as indicated by the lower EC50 (half maximal effective concentration of P8 to inhibit CFPS) (Fig. 3i). These results suggest that P8 likely impeded eGFP production by inhibiting transcription. To rule out the possibility of eGFP being quenched by P8, we also added P8 to the reaction after eGFP had been produced (Supplementary Fig. S47). However, we observed negligible influence of P8 on the pre-produced eGFP, confirming that the observed effects are due to the inhibition of protein synthesis. We also normalized the CFPS kinetic curves by their respective plateaued values and found that the normalized CFPS reaction rates are mostly unaffected (Supplementary Fig. S48). If P8 targeted ribosomes or RNA polymerase directly, we would expect the normalized reaction rate to drop drastically39. However, since P8 concentration did not display a strong effect on the normalized CFPS reaction rates, it is likely that the observed inhibitory effects of P8 on protein synthesis are primarily due to the sequestration of nucleic acids into the biomolecular condensates.
Mechanism of action of P8 by bacterial cytological profiling (BCP)
To further study the antimicrobial mechanism of P8, we conducted bacterial cytological profiling (BCP), an effective approach for studying the mechanisms of action of different antibiotics45,46,47. B. subtilis was chosen here because it was a well-studied bacteria model for BCP analysis46,47. B. subtilis were exposed to antibiotics targeting the five major pathways (translation, transcription, DNA replication, peptidoglycan synthesis, and membrane integrity), as well as P8. Afterward, they were stained with FM4-64 FX (membrane dye), DAPI (DNA dye) and SYTOX Green (membrane integrity indicating dye) and then imaged using confocal microscopy. Cells treated by different antibiotic classes exhibited distinct unique cytological profiles (Supplementary Fig. S49)45. To quantitatively analyze these results, we measured the bacterial cell morphology, DNA parameters and SYTOX Green signals resulting from each treatment (Supplementary Data 1) and performed principal component analysis (PCA) (Supplementary Fig. S50a, b). As shown in Fig. 3j, P8 was quantitatively separated from all antibiotics, demonstrating the distinct antimicrobial mechanism of P8. This is unsurprising as P8 induced significant DNA condensation (compaction) in B. subtilis (Supplementary Fig. S49), which largely differentiates it from other antibiotics. Interestingly, if we performed PCA just with bacterial morphology parameters (length, width, area, perimeter, and circularity) and SYTOX Green signals only (Supplementary Data 2 and Supplementary Fig. S50c, d), we found P8-treated cells had an overall profile identical to rifampicin-treated cells (Fig. 3k). This suggests that P8 may, like rifampicin, inhibit RNA synthesis, which is consistent with the inhibitory effect of P8 from in vitro transcription testing (Fig. 3f). Based on these data, we infer that P8 inhibits RNA synthesis by sequestering nucleic acids, likely the DNA or plasmid, into biomolecular condensates, ultimately leading to bacterial death.
Mice studies of P8 with multiple infection models
After i.p. administration of 100 mg/kg of P8 (10 doses of 10 mg/kg of P8 were given at 12-hour intervals over 5 days) (Fig. 4a-b), as well as subcutaneous injections of 60 mg/kg of P8 (4 doses of 15 mg/kg at 4 h intervals) (Supplementary Fig. S51a, b), we observed no signs of toxicity. These outcomes indicate the favourable biosafety profile of P8.
In vivo toxicity. a Serum biochemistry analysis and b representative histological images of uninfected mouse treated with PBS (-) or P8. Mice were i.p. injected with 10 doses of PBS (-) or P8 (10 mg/kg) at 12-hour intervals. The level of alanine transaminase (ALT), aspartate transaminase (AST), blood urea nitrogen (BUN), total protein (TP), creatinine (CRE) and glucose (GLU) were measured in mice (n = 6) blood samples drawn at day 3 and day 5 after the administration of the first injection of PBS (-) or P8 (day 0). Kidney, liver and spleen were collected from mice on day 5 and day 14 for sectioning and H&E staining. Murine systemic infection model. c Illustrated scheme of the systemic infection model. d Survival rate (%) and e antimicrobial potency of P8 in systemic infection model. There were 6 mice in each group. Murine thigh infection model. f Illustrated scheme of the neutropenic thigh infection model. g Antibacterial potency of P8 against P. aeruginosa BAA-2797, K. pneumoniae KPNR and MRSA in neutropenic thigh infections. There were 6 mice per group. Murine lung infection model. h Illustrated scheme of the lung infection model. Survival rate of lung-infected mice with i MRSA and j P. aeruginosa PAO1. There were 6 mice per group. One-way ANOVA analysis for e and g: *P < 0.05; **P < 0.01, ***P < 0.001; ****P < 0.0001; ns (not significant). The exact P values are provided in Source Data. Two-tailed t-test for a. Data are presented as mean ± standard deviation. Illustrations for c, f, and h were created in BioRender. Chan, M. (2025) https://BioRender.com/ctpibvl.
The presence of serum had minimal impact on the P8 antimicrobial efficacy (Supplementary Table S2), suggesting that P8 may perform well in vivo. The in vivo effectiveness of P8 was initially assessed in a murine peritonitis infection model with MDR P. aeruginosa BAA-2797 (Fig. 4c). A single dose of P8 (10 mg/kg) administered intraperitoneally successfully rescued the infected mice, resulting in a 100% survival rate (6 out of 6 mice) (Fig. 4d), while PBS (-)- and imipenem (IMP)-treated mice succumbed to the infection. Furthermore, P8 reduced the bacterial loads by more than 4.5 log10 in both peritoneal fluid and major organs when compared to the group treated with PBS (-) (Fig. 4e). From Supplementary Table S3, PBS (-)-treated infected mice suffered from poor health, as indicated by reduced movement, significant ruffling and sunken eyes. After 1 day of P8 treatment, the infected mice become more healthy-looking. 3 days after P8 treatment, the mice appeared to have fully recovered. With the safety tests of P8 (i.e. mice given 1 dose of P8 of 10 mg/kg but without bacteria), the mice appeared healthy-looking after 1 day of P8 treatment. Notably, P8 demonstrated efficacy comparable to colistin in eradicating P. aeruginosa BAA-2797 and remained effective against colistin-resistant K. pneumoniae BAA-2784 in the peritonitis infection model (Supplementary Fig. S52). P8 demonstrate the ability to eradicate bacteria resistant to both colistin and imipenem which are hard to treat.
Pharmacokinetic (PK) analysis following intraperitoneal (i.p.) administration in mice of a single dose of P8 (10 mg/kg) (Supplementary Fig. S53) found favourable P8 profile. A peak blood concentration of 12.3 µg/ml and a half-life (T1/2) of 24 minutes, which surpassed those of conventional antibiotics such as meropenem and imipenem (with T1/2 of 6 min and 12 min)48, were found, serving as positive indicators of good bioavailability. We also studied the biodistribution of P8 by intraperitoneal (i.p.) administration in mice of a single dose of Cy7.5-conjugated P8 (Cy7.5-P8) (10 mg/kg). The result showed that Cy7.5-P8 was distributed in mouse liver, spleen, kidney and lung (Supplementary Fig. S54a), and the quantification analysis showed that Cy7.5-P8 was primarily accumulated in the liver (Supplementary Fig. S54b).
Subsequently, we employed a murine neutropenic thigh infection model to examine P8’s efficacy (Fig. 4f). P8 (2 doses of 15 mg/kg) injected subcutaneously eradicated the MDR P. aeruginosa BAA-2797 and K. pneumoniae KPNR in the thigh, achieving approximately 2.4 and 1.9 log10 CFU reductions, respectively, compared to PBS (-) (Fig. 4g). This outperformed the efficacy of IMP with 0.1 and 1.4 log10 reductions, respectively. Further, P8 achieved 2.0 log10 reduction against MRSA in the thigh infection, which is statistically comparable to the efficacy of vancomycin (VAN) (2.3 log10 reduction) (Fig. 4g).
We further evaluated the antibacterial efficacy of P8 in a murine lung infection model with either MRSA or P. aeruginosa PAO1 (Fig. 4h and Supplementary Fig. S55a). P8 was found to be more effective than VAN in both rescuing mice from MRSA lung infection (Fig. 4i) and eradicating MRSA burden in lung tissues (Supplementary Fig. S55b). Further, P8 rescued all infected mice (6 out of 6) from P. aeruginosa PAO1 lung infection, while IMP treatment saved 5 out of 6 infected mice (Fig. 4j). P8 reduced the P. aeruginosa PAO1 burden in lung tissues by 6.0 log10, while IMP showed 4.7 log10 reduction (Supplementary Fig. S55c).
To study the healing process with two mice models, we performed H&E histological analysis to evaluate the structural morphologies of major organs retrieved from mice after different treatment times and compared them with control PBS(-) treatment group. For the mouse model of systemic infection, we found that infection caused various organ lesions in infected mice with PBS(-) treatment (Supplementary Fig. S56a), including loss of defined demarcation between the white pulp structure and the surrounding red pulp structure in the spleen as well as the destruction of its mantle zone and germinal centre (white arrows), infiltration of inflammatory cells in the liver (green arrows) and thickening of the alveolar walls (yellow arrows) for PBS (-) treatment. However, for the mice receiving the P8 treatment, these tissue lesions caused by bacterial infections were effectively decreased. Specifically, the spleen of P8-treated mice restored to healthy-like appearance with well-defined germinal centres within 1 day of P8 treatment while the thickness of the alveolar walls gradually decreased to similar appearance of equivalent structures in healthy mouse lungs within 14 days of P8 treatment.
In the lung infection model (Supplementary Fig. S56b), we found that infection caused severe lesions in the lung with thickening and collapse of alveolar walls resulting in the loss of defined air sac structures (black arrow) as well as structural and epithelial damage of the bronchioles (blue arrows). For mice receiving P8 treatment, lung lesions caused by bacterial infection were effectively decreased with gradual restoration of the alveolar and bronchiolar structures to healthy state within 14 days post-treatment.
Discussion
The rise of drug-resistant bacteria poses a significant global health threat, underscoring the urgent need for in vivo active antibiotics with novel modes of action. In this study, we designed a series of biodegradable main-chain cationic polyimidazoliums (PIMs) with ester linkages, specifically poly(imidazolium ester)s (PIEs). Through extensive structural optimization, we identified a lead compound, P8, which exhibited an optimal balance of intermediate degradation rate, strong in vitro and in vivo antibacterial activity, and favourable biosafety profile. More importantly, we demonstrated that P8 employs a non-classical antibacterial mechanism involving the formation of intracellular polymer@nucleic acid biomolecular condensates.
P8 translocates bacterial membranes without compromising membrane integrity. Further, P8 interacts with DNA through multiple binding modes, including minor groove binding and intercalation, driven by electrostatic interactions and hydrogen bonding. We previously showed that oligoimidazoliums (OIMs), specifically OIM1-6, binds/damages intracellular DNA49. However, the mechanistic link between this DNA binding/damage and cell death was unclear. A recent mechanistic study revealed that antimicrobial peptides, including P113, Os-C, and Buforin, have a strong tendency to form liquid-like condensates with nucleic acids through phase separation39. This led us to hypothesize that our main chain cationic PIMs could potentially serve as a next-generation antibacterial macromolecule class that acts through nucleic acid phase separation. This concept is particularly intriguing, as intracellular biomolecule phase separation is an attractive but relatively unexplored mechanism in antibiotic development.
Herein, we demonstrated the phase separation of P8 with nucleic acids both in vitro and in cellulo, resulting in the formation of liquid polymer@nucleic acid condensates. The internalized P8 was found to sequester approximately 74% of intracellular nucleic acids into liquid-like compact condensates. P8-modulated nucleic acid phase separation was found to effectively inhibit the in vitro transcription and the subsequent translation processes. This extensive sequestration of genetic material into condensates by P8 likely induces a cytotoxic effect that the cell cannot counteract. P8 is a next-generation antibacterial polymer, as opposed to peptides with a well-defined amino acid sequence, inducing phase separation of biomolecules within bacteria to exert its antimicrobial effect.
Furthermore, bacterial cytological profiling using parameters such as DNA compaction, cell morphology, and membrane integrity supports the unique antimicrobial mechanism of P8, as indicated by the distinct profiles of P8-treated cells compared to those treated with traditional antibiotics. A comparison based solely on cell morphology and membrane integrity in P8- and antibiotic-treated cells suggests that P8 may inhibit RNA synthesis. This finding corroborates our conclusion that the formation of high-viscosity P8@nucleic acid condensate impedes bacterial transcription leading to cell death.
The mechanism by which P8 kills bacteria, i.e. complexation with DNA to form a condensed phase which inhibits mRNA transcription and protein synthesis, may be very difficult for bacteria to develop resistance to. P8 interacts with bacterial nucleic acids via electrostatic interaction and hydrogen bonding. It binds the minor groove of and intercalates with DNA. Point mutations seem highly unlikely to impair the P8/DNA interaction or cause changes that would ameliorate the problems the P8/DNA interaction causes for transcription and translation. The lack of significant increase in MICs in serial passaging provides a preliminary confirmation of the intuition that P8 may not readily elicit resistance. There might, however, be envelope or efflux system mutations that could make it harder for P8 to accumulate in the cytoplasm; these might confer a degree of resistance.
Further, P8 has superior in vivo activities compared with our previous reported main chain cationic polyimidazoliums (PIMs) or oligoimidazoliums49,50. P8 achieved significant antibacterial efficacy in the thigh infection model which has not previously been demonstrated. To the best of our knowledge, P8 is an unique cationic antimicrobial polymer reported to show a significant reduction of both Gram-negative and Gram-positive multidrug-resistant bacteria in the murine thigh infection model, which is commonly used for in vivo evaluation of new antibiotics51,52,53. P8’s intermediate degradation rate, faster than the α analogue but slower than the γ analogue, offers an optimal window for bacterial eradication while degrading before significant toxic accumulation in eukaryotic cells. Moreover, P8 demonstrated in vivo efficacy beyond the thigh infection model, effectively combating multidrug-resistant P. aeruginosa in systemic infection models, as well as P. aeruginosa and MRSA in lung infections.
This is the report of a next-generation class of macromolecules - synthetic cationic polymer demonstrating a non-classical antibacterial mechanism through the formation of intracellular biomolecular condensates, while also showing strong in vivo efficacy with a favourable therapeutic window. This non-classical antimicrobial mechanism may mitigate the problem of rapid emergence of resistance to new antibacterial agents and provide alternative tools with which to treat bacterial infections that are already resistant to current antibiotics.
Methods
Ethics Statement
The animal study protocols were approved by the Institutional Animal Care and Use Committee of Nanyang Technological University (NTU IACUC, protocol A22038, A24014 and A20029).
Synthesis of bisimidazoles
Three bisimidazoles-Cr (r refers to the carbon number in alkyl linker) were synthesized via N-alkylation of imidazole with dibromides in base (Supplementary Fig. S1a). The synthesis of all bisimidazoles-Cr followed the same procedure except that 1,3-dibromopropane, 1,4-dibromobutane and 1,5-dibromopentane were used for the preparation of bisimidazole-C3, bisimidazole-C4 and bisimidazole-C5 respectively. Here the synthesis of bisimidazole-C4 is taken as an example. NaH (3.0 equiv.) was added portion wise to imidazole (1.0 equiv.) solution in THF at 0 °C, and the reaction mixture was stirred at room temperature for 2 h. 1,4-dibromobutane (0.5 equiv.) was added to the reaction mixture and the resulting solution was stirred at room temperature overnight. The crude product was filtered to remove solid NaBr salt and washed with THF. The filtrate was evaporated and evaluated with column chromatography (10% methanol: ethyl acetate) (yield: 80%).(Supplementary Figs. S2–S4) 1H NMR: δ 7.62 (s, 1H), δ 7.14 (s, 1H), δ 6.89 (s, 1H), δ 3,96 (t, 4H), δ 1.61 (m, 4H). 13C NMR: δ 137.66, 128.84, 119.70, 45.75, 28.12.
Synthesis of dibromides
Ester linked dibromides [p + q] were synthesized via facile esterification of acyl chloride with hydroxyl (p refers to carbon number linked to acyl chloride group and q represents carbon number in hydroxyl) (Supplementary Fig. S1b). The typical reaction followed the procedure below. Bromopropionyl chloride (1.0 equiv.) was added dropwise into 2-bromoethanol (1.0 equiv.) at room temperature, and the reaction proceeded under stirring at 50 °C overnight. After that, the reaction mixture was washed three times with saturated brine. After drying with anhydrous sodium sulfate, colourless liquid dibromides were obtained (yield: 90%). (Supplementary Fig. S5–S9) 1H NMR: δ 4.43 (t, 2H), δ 3.35-3.52 (t, 4H), δ 2.96 (t, 2H).13C NMR: 170.03, 64.20, 37.50, 28.59, 25.82.
Synthesis of poly(imidazolium ester)s (PIEs)
Poly(imidazolium ester)s (PIEs) were prepared via polycondensation of bisimidazoles-Cr and dibromides [p + q] (Supplementary Fig. S1c). The detailed process is as following: bisimidazoles-Cr (1.0 equiv.) and dibromides [p + q] (1 equiv.) were weighed into a flask with a stir bar. The reaction was conducted at 120 °C for 30 min, and pale brown solid product was obtained. The crude product was dissolved in methanol and then precipitated into THF to remove unreacted monomers. The obtained product was passed through a column of Amberlyst®A26 (Sigma-Aldrich) to exchange Br- with Cl-. The resulted product was then dialyzed against DI water for 12 h (molecular weight cutoff: 1000 Da) and then lyophilized. The bulk polycondensation mixture at the beginning was in liquid phase at the elevated reaction temperature and then gradually solidified with increasing polymerization times (in about 10 mins) (Supplementary Fig. S27a), leading to quenching of the polymerisation. Meanwhile, dialysis (MWCO 1k) against DI water was conducted to remove relatively low-molecular weight oligomers, which may give rise to low yield. Finally, the relative longer oligomers with narrow distributions were obtained in the dialysis bag and water was removed by lyophilization. This synthesis/purification protocol was repeated numerous times, and there were only small variations in the molecular weight (Supplementary Fig. S27b) and antimicrobial efficacy (Supplementary Table S1) from batch to batch.
The preparation of FITC-P8 followed the same procedure as the synthesis of poly(imidazolium ester)s except that 5-(iodoacetamido)fluorescein (0.033 equiv.) was introduced during polymerization. The FITC content in the final FITC-P8 is 7.4%.
Degradability of poly(imidazolium ester)s
Poly(imidazolium ester)s were dissolved in PBS (pH 7.2 ± 0.2) and incubated at 37 °C. After specified incubation times, aliquots were taken for 1H NMR, GPC and antimicrobial activity tests.
In vitro antimicrobial activity
Minimum Inhibitory Concentrations (MICs) of poly(imidazolium ester)s were determined by broth microdilution method adapted from BS EN ISO 2077601:2020 with modifications. Bacteria were sub-cultured in Mueller-Hinton broth (MHB) (DifcoTM Mueller Hinton Broth, non-cation-adjusted) to an OD of 0.2–0.3 (which corresponded to a measured 108 CFU/mL) and diluted to 105-106 CFU/mL in MHB. Poly(imidazolium ester)s were prepared at final concentration of 10.24 mg/mL in DI water. 10 µL of polymer stock was diluted with 90 µL MHB and dispensed into the first well of a 96-well plate. The second and subsequent wells were each pre-filled with 50 µL MHB. 50 µL polymer solution was taken from first well to the second well, achieving twofold dilution. Twofold serial dilution was conducted for the subsequent wells as well. Into each well of the 96 well plate was then introduced 50 µL bacteria suspension in MHB to achieve a final volume of 100 µL with ∼105 CFU/mL. The final concentrations of poly (imidazolium ester) range from 512 µg/mL to 1 µg/mL. The covered plate was incubated at 37 °C for 18-20 hours, and we measured the optical density absorbance of each well at a wavelength of 600 nm. The lowest concentration that reduces the bacterial count 90% or more (based on OD600) was adopted as MIC.
Cytotoxicity assay
Mammalian cell compatibility of P8 was tested with 3T3 fibroblast cell, human embryonic kidney cells (HEK) or human adenocarcinomic lung epithelial cells (A549) using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltenyltetrazolian bromide (MTT) in colorimetric assay54. Mammalian cells were firstly cultured to 80% confluence in a culture flask in medium containing 89% Dulbecco’s Modified Eagle’s Medium (DMEM), 10% foetal bovine serum (FBS) and 1% antibiotics (penicillin/streptomycin) until 80% confluence in culture flask appeared. Then cells were detached from the bottom of flask using trypsin, and subsequently were spin down and redispersed in the culture medium to obtain concentrated cell suspension. After counting the cells with a hemocytometer, 1 × 104 cells were seeded into each well of a 96-well plate in 100 µL medium. Cells in 96-well plate were incubated for 24 h before the addition of poly(imidazolium ester)s with concentrations ranging from 1024 µg/mL to 32 µg/mL in DMEM solution. After 24-h treatment, the supernatant was removed, and each well was washed with PBS. 50 µL MTT (1 mg/mL) was introduced to each well and the plate was incubated for another 2 h. After removal of the supernatant, 100 µL isopropanol was added to each well and cell viability was evaluated by measuring the absorbance of formazan in each well (OD570nm).
In vitro pro-inflammatory activity (TNF-α secretion) of P8
RAW macrophages (6 × 104 cells/well) in medium containing 89% Dulbecco’s Modified Eagle’s Medium (DMEM), 10% foetal bovine serum (FBS) and 1% antibiotics (penicillin/streptomycin) were seeded into each well of a 24-well plate and incubated for 16 h at 37 °C in 5% CO2. After that, the culture media were removed, and new culture media containing lipopolysaccharide (LPS, 5 ng/mL), P8 at different concentrations (1-4 × MIC, MIC against P. aeruginosa PAO1: 4 µg/mL), or both were added into the respective wells and further incubated for 3 days. Afterwards, the culture media were collected and the concentrations of TNF-α in the media were determined using enzyme-linked immunosorbent assay (ELISA MAX DeluxeMouse TNF-α, BioLegend) following the manufacturer’s instructions. Cells with no treatment, PLGA (8 µg/mL), and PEG3500 (8 µg/mL) were used as controls.
In vitro anti-inflammatory activity (Arginase activity) of P8
RAW macrophages (6 × 104 cells/well) in the same media as above were seeded into each well of a 24-well plate. After 16-h incubation at 37 °C in 5% CO2, the culture media were replaced with new media containing interleukin 4 (IL-4, 100 ng/mL), or P8 at different concentrations (1-4 × MIC, MIC against P. aeruginosa PAO1: 4 µg/mL), or both, and incubated for 3 days. Next, the media were removed, and cells were washed with DPBS and detached into the fresh media using a cell-scraper. The cell pellets obtained after centrifugation were dispersed in lysis buffer containing Tris-HCl (10 mM), Triton X-100 (1% w/v), NaCl (150 mM), EDTA (1 mM), and protease inhibitor cocktail (Merck). After lysing the cells for 30 min at 4 °C, the insoluble materials were removed using centrifugation. Finally, the arginase activity of each sample was analyzed using a commercially available kit (Sigma, MAK112) following the manufacturer’s instructions.
Bacterial resistance assay
Drug resistance testing was conducted by serial passaging of bacteria with P8 at the highest concentration below MIC. Briefly, exponential-phase bacteria (107 CFU/mL) were grown in 100 μL of MHB containing P8 or antibiotic control at a gradient of concentrations. At 24-h intervals, the cultures were checked for growth and the MIC value for each day was recorded. Cultures from the highest concentration below MIC were diluted 1:100 into fresh MHB containing P8/antibiotics at different concentrations. The serial passaging was repeated daily for 20 days.
Resistance frequency determination
Log-phase P. aeruginosa PAO1 cultures (108 CFU/mL) were incubated in duplicate with 10× MIC of either P8 or imipenem in 20 mL of fresh MHB. The tubes were monitored daily for turbidity. To evaluate the emergence of spontaneous resistant mutants, the cultures were plated and subjected to standard MIC testing.
Assays of membrane activity of P8
For PI assays, mid-log P. aeruginosa PAO1 were harvested by centrifugation at 5000 g for 5 min at room temperature and resuspended in PBS. The suspension was diluted in PBS to reach a final concentration of 108 CFU/mL. PI (15 µg/mL) was then added to above bacterial suspension. 90 µL samples of bacterial suspension with PI dye was added into wells of a 96-well white microplate, and 10 µL P8 or colistin at indicated concentration were added into each challenge well. Wells with cells and PI dye but no P8 and colistin served as untreated control. The fluorescence intensity of PI for each well was monitored using a plate reader. The relative PI fluorescence intensity (F.I.) was calculated by following equation: Relative PI F.I.= Icompound/Icolistin×100%.
To study the membrane translocation by time-lapse fluorescence confocal imaging, mid-log P. aeruginosa PAO1 was diluted in PBS to reach a final concentration of 108 CFU/mL as bacterial working solution. Then, equal volumes of 32 µg/ml FITC-P8 and 40 µg/µL PI were mixed, and 10 µl of this mixture solution was added to 10 µl of bacteria suspension. The bacterial mixture was immediately deposited onto a polylysine-coated glass-bottomed Petri dish (MatTek Corporation) and imaged at indicated times with a Zeiss LSM800 confocal microscope using a 100× oil immersion objective. The starting time for imaging was designated as 0 min.
For cryo-TEM assays, mid-log P. aeruginosa PAO1 cells were harvested by centrifugation at 5000 g for 5 min at room temperature and diluted to 108 CFU/mL in MHB and incubated with P8 at indicated concentration for 60 min at 37 °C. Bacterial suspension was harvested by centrifugation, washed with PBS and resuspended in PBS to prepare samples for cryo-TEM at Institute of Structural Biology at Nanyang Technological University.
Selective membrane translocation of P8
To study the intracellular accumulation of P8 in bacteria, mid-log P. aeruginosa PAO1 was diluted to reach a final concentration of 108 CFU/mL in MHB and treated with FITC-P8 (16 µg/mL) for 60 min. Cells were harvested and washed twice with PBS, and then fixed with 4% paraformaldehyde in PBS for 15 min. Fixed cells were washed with PBS and then stained with 5 µg/mL FM 4-64 FX (Invitrogen, Thermo Fisher Scientific) for 10 min on ice. The cells were again washed twice with PBS and imaged with a Zeiss LSM800 confocal microscope on a polylysine-coated Petri dish (MatTek Corporation) with a 100× oil immersion objective.
To study the membrane translocation of P8 in mammalian cells, 3T3 fibroblast cells were cultured in a medium comprising 89% Dulbecco’s Modified Eagle’s Medium (DMEM), 10% foetal bovine serum (FBS), and 1% antibiotics (penicillin/streptomycin) until they reached 80% confluence in the culture flask. The cells were then detached from the flask using trypsin, followed by centrifugation and resuspension in the culture medium to create a concentrated cell suspension. The cell count was determined using a hemocytometer, and 1 × 104 cells were seeded into a polylysine-coated petri dish. The cells were incubated in the petri dish for 24 hours, after which FITC-P8 was added at 16 µg/mL. After 60-minute treatment, the cells were washed twice with PBS and then fixed with 4% paraformaldehyde in PBS for 15 minutes. Following fixation, the cells were subjected to another round of PBS washing before being stained with DAPI (2 µg/mL) for 15 minutes. The cells were washed twice with PBS again and were imaged using a Zeiss LSM800 confocal microscope.
Assays examining P8-modulated liquid-liquid phase separation of nucleic acids in vitro
To visually examine the formation of P8@nucleic acid condensates, P8 was mixed with dsDNA-s and poly(A) in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2, and 50 mM HEPES pH 7.3. The opacity and turbidity of the suspensions were immediately observable. The following concentrations were used: 500 µg/mL P8, 1000 µg/mL poly (A) and 2000 µg/mL dsDNA-s.
To study condensate morphology with confocal microscopy, FITC-P8 (500 µg/mL) was mixed with dsDNA-s (2000 µg/mL) and poly(A) (1000 µg/mL) stained with Syto59 (Invitrogen™, 5 µM) in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2, and 50 mM HEPES pH 7.3. The samples were imaged with a Zeiss LSM800 confocal microscope with a 63× oil immersion objective.
For monitoring the fusion of P8@nucleic acid condensates, FITC-P8 (500 µg/mL) was mixed with dsDNA-s (2000 µg/mL) or poly(A) (1000 µg/mL) in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2, and 50 mM HEPES pH 7.3. Time-lapse images of this sample were collected with a Zeiss LSM800 confocal microscope with a 63× oil immersion objective.
For fluorescence recovery after photobleaching (FRAP), a Zeiss LSM800 confocal microscope with a 63× oil immersion objective was used. A 488 nm argon laser at 50% power was employed to bleach a disk-shaped area within P8@nucleic acid condensates. Then, fluorescence recovery was recorded for 200 s. The fluorescence intensities of photobleached areas were normalized against unbleached areas. The half-life of fluorescence recovery, denoted as τ1/2, was determined by fitting the recovery curves to a first order exponential decay model. FRAP experiments were performed on 3 different condensates for both P8@dsDNA-s and P8@Poly(A). The condensates were formed using the following concentrations of components: 500 μg/mL FITC-P8; 2000 µg/mL dsDNA-s; 1000 µg/mL poly(A).
Construction of phase diagrams
FITC-P8 and dsDNA-s, poly(A) or eGFP plasmid DNA at different concentrations were mixed in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2, and 50 mM HEPES at pH 7.3. The mixtures were visualized using a Zeiss LSM800 confocal microscope with a 63× oil immersion objective to assess condensate formation.
Dye-dsDNA-s competitive replacement assay
dsDNA-s at the concentration of 80 µg/mL in HEPES buffer (50 mM, pH = 7.2) was used as the dsDNA-s working solution. The DNA-binding dye Hoechst 33342 (NucBlue™ Live ReadyProbes™ Reagent, 2 drops per ml as instructed) and Ethidium Bromide (Promega, H5041, 40 µg/mL) were prepared in HEPES buffer. 25 µL dsDNA-s was mixed with 25 µL Hoechst 33342 or Ethidium Bromide in a 96-well plate. After incubation for 15 min, 50 µl of buffer or P8 with varying concentrations in HEPES buffer was introduced in each well with final P8 concentration at 0, 2, 4, 8, 16 and 32 µg/mL. 25 µL Hoechst 33342 or Ethidium Bromide mixed with 75 µL HEPES buffer were used as blank. The plate was incubated for 30 min, followed by measurement of fluorescence intensity. Each measurement was conducted with three replicates. The excitation and emission wavelengths were set to 350 nm and 470 nm for Hoechst 33342 detection or 475 nm and 604 nm for Ethidium Bromide detection. The remaining dye-DNA complex was calculated by following equation: The remaining dye-DNA complex = (Ix-Iblank)/(I0-Iblank)×100%. Herein, Ix and I0 refer to the fluorescence intensity of dye-dsDNA-s complex in the presence of P8 at x and 0 µg/mL.
Assays investigating the driving forces for liquid-liquid phase separation of P8 with nucleic acids
For confocal images, FITC-P8 (500 µg/mL) was mixed with dsDNA-s (2000 µg/mL) or poly(A) (1000 µg/mL) in 5% w/v PEG (20 kDa), 150 mM KCl, 1 mM MgCl2, and 50 mM HEPES pH 7.3. The resulting P8@dsDNA-s and P8@Poly(A) condensates were supplemented with 10% 1,6-hexanediol, 800 mM KCl or 8 M urea, and then imaged with a Zeiss LSM800 confocal microscope using a 63× oil immersion objective.
To measure the turbidity, preformed P8@dsDNA-s and P8@Poly(A) condensates were supplemented with 1,6-hexanediol (1,6-HD) (concentration: 3.3, 6.6 and 10%), KCl (concentration: 100, 200, 400 and 800 mM) or urea (concentration: 1, 2, 4 and 8 M). Subsequently, the turbidity (OD600) of each sample was evaluated. The P8@dsDNA-s and P8@Poly(A) condensates were prepared using the following concentrations of components: 500 μg/mL P8; 2000 µg/mL dsDNA-s; 1000 µg/mL poly(A).
Assays investigating the in cellulo phase separation of nucleic acids with P8
To study the P8:nucleic acid condensates inside bacteria, mid-log phase P. aeruginosa PAO1 (108 CFU/mL) in MHB was treated with FITC-P8 at 16 µg/mL for 60 min. Cells were harvested by centrifugation, washed twice with PBS, and then fixed with 4% paraformaldehyde in PBS for 15 min. Fixed cells were washed with PBS, stained with Syto59 (50 mM) and imaged with a Zeiss LSM800 confocal microscope on a polylysine-coated Petri dish (MatTek Corporation). Fluorescence intensity percent (F.I. percent (%)) in condensate region was calculated by determining the ratio of the fluorescence intensity of Syto59 in the condensate region to that in the whole cells (n = 11).
To monitor the phase separation process inside bacteria, mid-log phase P. aeruginosa PAO1 (~109 CFU/mL) was stained with Syto59 (50 mM) for 10 min. FITC-P8 or P8 was introduced to the bacteria suspension with the final concentration at 32 µg/mL. Subsequently, the cell suspension was deposited onto a polylysine-coated glass-bottomed Petri dish (MatTek Corporation) and imaged at indicated times. The starting time for imaging was designated as 0 min. Untreated bacteria were also imaged as control.
In vitro transcription (IVT) assay
In vitro transcription reactions were carried out using a T7 RNA transcription kit (HiScribe T7 High Yield; New England Biolabs). We first prepared the linear DNA fragment by the PCR using pET30a-GFP as the template and the following primer pairs CGATCTTCCCCATCGGTGATGTCG and CAGCAGCCAACTCAGCTTCCTTTCGG. The resulting PCR fragment contains the T7 promoter-driven eGFP-expressing cassette. We then purified the DNA fragment and subjected it (38 ng/µl) to in vitro transcription (HiScribe T7 High Yield; New England Biolabs) for 2 h at 37 °C in the presence of P8 (Total reaction volume of 20 μL, P8 concentration: 1, 2, 4, 8 and 16 µg/mL). The control transcription reaction without P8 (0 µg/mL) was also performed. Samples were incubated at 37 °C for 2 h, after which 1 μL of DNAseI (Turbo DNAse) was added and incubated for an additional 15 min at 37 °C. 1 μL of the reaction sample was diluted directly in formamide-containing denaturing RNA loading buffer (ThermoFisher) and incubated at 65 °C for 10 min prior to its loading on a denaturing formaldehyde agarose gel.
To study the phase separation of the DNA fragment in IVT in the presence of P8, the reaction mixture was supplemented with Syto59 and imaged with confocal microscope.
Cell-free protein synthesis (CFPS) assay
Cell-free eGFP synthesis was performed in NEBExpress cell-free E. coli protein synthesis system (New England BioLabs Inc.). Briefly, the system was supplemented with the desired concentration of pET30a-GFP (3 ng/µl), the DNA plasmid which has a T7 promoter-driven eGFP-expressing cassette55 and P8 (0-64 µg/ml), and the total volume of one reaction mixture was 50 μL. The solutions were aliquoted into a 96-well white plate (BRANDplates® pureGrade™), 50 μL per well. The plate was sealed with clear sealing tape. The fluorescence intensity (Excitation wavelength: 485 nm, Emission wavelength: 535 nm) of each well was monitored every 3 minutes by a plate reader (TECAN, SPARK 10 M) at 37 °C with shaking (1440 rpm).
The preparation of mRNA for the Cell-free eGFP synthesis was carried out using a T7 RNA transcription kit (HiScribe T7 High Yield; New England Biolabs) followed by purification with Monarch® RNA Cleanup Kit (T2040S). We first conducted the PCR using pET30a-GFP as the template and the following primer pairs CGATCTTCCCCATCGGTGATGTCG and CAGCAGCCAACTCAGCTTCCTTTCGG to obtain the linear DNA fragment. The resulting PCR fragment contains the T7 promoter-driven eGFP-expressing cassette. We then purified the DNA fragment and subjected it to in vitro transcription using a T7 RNA transcription kit (HiScribe T7 High Yield; New England Biolabs) for 2 h at 37 °C followed by purifying the mRNA with Monarch® RNA Cleanup Kit (T2040S). The obtained mRNA (50 ng/μL) was used for Cell-free eGFP synthesis in the presence of P8 (0–128 µg/mL). The production of eGFP was monitored in a 96-well plate as described above.
Bacterial cytological profiling
Exponential-phase B. subtilis were treated with antibiotics or P8 at 5 × MIC in tryptic soy broth (TSB) and incubated at 37 °C for 1.5 h. Treated cultures were harvested by centrifugation at 5000 g for 5 min and resuspended in TSB. The bacteria were then stained with 10 μg/mL FM 4-64 FX (Invitrogen, Thermo Fisher Scientific),10 μg/mL DAPI and 1.0 μM SYTOX Green for 15 min. The cells were washed twice with PBS and imaged with a Zeiss LSM800 confocal microscope on a polylysine-coated Petri dish (MatTek Corporation).
Bacterial cell morphology and DNA parameters, including length (μm), width (μm), area (μm2), perimeter (μm) and circularity were measured from confocal images using Image J analysis software. Average SYTOX Green intensity per pixel was determined using the membrane outline and then subtracted by its own background intensity. Finally, SYTOX Green intensity of treated cells was normalized by untreated cells intensity.
Variables in Supplementary Data 1 for each antibiotic and P8 were obtained from three independent experiments. Profiling data were collected with bacteria number >50. Variables reduction was performed using a Principal Component Analysis (PCA) with the Principal Component Analysis Calculator (statskingdom.com).
In vivo toxicity evaluation
Female BALB/c mice (7-8 weeks, brought from InVivos Pte Ltd) were used for all in vivo studies. Mice were housed under specific pathogen-free conditions on a 12-h light-dark cycle with controlled temperature and humidity and had ad libitum access to food and water.Female BALB/c mice (7–8 weeks, n = 6) were treated with PBS (-) or P8 (100 mg/kg total dose) by intraperitoneal injection administered as 10 doses (10 mg/kg per dose) at 12 h intervals. Blood was drawn at day 3 and day 5 after the first injection (day 0). Blood biochemistry was analyzed with Pointcare V3 Blood Chemistry Analyzer (MNCHIP). For the histology analysis, female BALB/c mice (7–8 weeks, n = 6) were treated with PBS (-) or P8 (100 mg/kg total dose) with the same dosing route and schedule. Major organs (liver, spleen and kidney) were collected from mice at day 5 and day 14 after the first injection of P8 or PBS (-) (day 0) for H&E staining. Advanced Molecular Pathology Laboratory (AMPL) at A*STAR Institute of Molecular and Cell Biology performed the histological processing of mouse tissues.
Female BALB/c mice (7–8 weeks, n = 6) were treated with PBS (-) or P8 (60 mg/kg total dose) by subcutaneous injection, administered as 4 doses (15 mg/kg per dose) at 4 h intervals. Blood was drawn at day 1 and day 7 after the first injection (day 0). Blood biochemistry was analyzed with a Pointcare V3 Blood Chemistry Analyzer (MNCHIP). For the histology analysis, female BALB/c mice (7–8 weeks, n = 6) were treated with PBS (-) or P8 (60 mg/kg) by the same dosing route and schedule. Major organs (liver, spleen and kidney) were collected from mice at day 3 and day 14 after the first injection of P8 or PBS (day 0) for H&E staining. Advanced Molecular Pathology Laboratory (AMPL) at A*STAR Institute of Molecular and Cell Biology performed the histological processing of mouse tissues.
Three models evaluating the in vivo activity of P8
In systemic infection model, 200 µL of P. aeruginosa BAA-2797 (106 CFU/mL) in PBS (with 5% mucin) were intraperitoneally (i.p.) injected into female balb/c mice (7–8 week) and the infections were allowed to develop for 2 h. At 2 h post infection, PBS (-), P8 (10 mg/kg) or imipenem (10 mg/kg) were injected intraperitoneally (n = 6). After 26-h infection, the mice were sacrificed and i.p. fluid, liver, kidney, and spleen were harvested and homogenized to determine the CFU. The survival test followed the same procedure, with mice being monitored for up to 7 days post infection/treatment. We tried a single dose of P8 of 5 mg/kg first, but only 1 out of 2 mice can be rescued from the MDR P. aeruginosa BAA-2797 systemic infection. We then used a higher P8 dose of 10 mg/kg. In a separate experiment, mice were infected intraperitoneally with P. aeruginosa BAA-2797 (106 CFU/mL) or K. pneumoniae BAA-2784 (108 CFU/mL). At 2 hours post-infection, they were treated with an intraperitoneal injection of PBS (-), P8 (10 mg/kg), or colistin (10 mg/kg) (n = 5). After 26 hours of infection, bacterial loads in the i.p. fluid, liver, kidney, and spleen were determined.
In thigh infection model, balb/c mice (female, 7–8 week) were made neutropenic by two intraperitoneal injections of cyclophosphamide at a concentration of 150 mg/kg on day 4 and 100 mg/kg on day 1 before the infection. Infection was established by intramuscular injection of 60 µL of P. aeruginosa BAA-2797 (105 CFU/mL), K. pneumoniae KPNR (106 CFU/mL) or MRSA (105 CFU/mL) into the right thigh of mice. The infected mice were treated subcutaneously with PBS (-), P8, or imipenem at 2, 6 h post infection (15 mg/kg per treatment, n = 6). The mice were sacrificed 26 h post infection and thigh tissues were harvested and homogenized to determine the CFU.
For lung infections, mice (balb/c, female, 7-8 week) were made neutropenic by two intraperitoneal injections of cyclophosphamide at a concentration of 150 mg/kg on day 4 and 100 mg/kg on day 1 before infection. Infections were established by intranasal delivery of 20 µL of S. aureus USA300 LAC (109 CFU/mL) or P. aeruginosa PAO1(104 CFU/mL). At 2-, 8-, and 26-h post-infection, the infected mice were treated intranasally with PBS (-), P8, or vancomycin (10 mg/kg per treatment, n = 6) for S. aureus USA300 LAC, or with PBS (-), P8, or imipenem (10 mg/kg per treatment, n = 6) for P. aeruginosa PAO1. The mice were sacrificed 38 h post infection and lungs were harvested and homogenized to determine the CFU. The survival test followed the same procedure, with mice being monitored for up to 7 days post infection/treatment.
To further confirm the in vivo antimicrobial efficacy of P8, we performed H&E histological analysis to evaluate the structural morphologies of major organs retrieved from mice after different treatment time and compared them with PBS (-) treatment. In the systemic infection model, mice were infected intraperitoneally with P. aeruginosa BAA-2797 (10⁶ CFU/mL). At 2 hours post-infection, they were treated with an intraperitoneal injection of PBS (-) or P8 (10 mg/kg) (n = 2). The organs were collected after 1-day treatment for PBS (-) group and after 1, 3 and 14 days for P8 group (n = 2). Healthy mice (n = 2) were used as negative control. Advanced Molecular Pathology Laboratory (AMPL) at A*STAR Institute of Molecular and Cell Biology performed the histological processing of mouse tissues.
In the lung infection model, mice were infected by intranasal delivery of MRSA LAC (109 CFU/mL) and treated with PBS (-) or P8 (10 mg/kg) at 2-, 8- and 26 hours post-infection (n = 2). The organs were collected after 3-day treatment for PBS (-) group and after 3, 7 and 14 days for P8 group. Healthy mice (n = 2) were used as negative control. Advanced Molecular Pathology Laboratory (AMPL) at A*STAR Institute of Molecular and Cell Biology performed the histological processing of mouse tissues.
Pharmacokinetics analysis
Balb/c mice (female, 7–8 weeks, n = 5 for each time point) were intraperitoneally injected with a single dose of FITC-P8 solution (10 mg/kg). A different set of 5 mice was used at each of 6 time points for the bleeding so that blood was collected from each of 30 mice once. For each mouse, 100–150 µL of blood was collected via the submandibular vein at the specified time, and plasma was harvested by centrifugation at 1000 g. The concentration of FITC-P8 in plasma was determined by fluorescence at 490 nm, using a calibration curve which was also established in plasma.
Biodistribution study
Balb/c mice (female, 7–8 weeks, n = 3 for per time point) were intraperitoneally injected with a single dose of Cy7.5-P8 solution (10 mg/kg). Major organs were collected at specified time (4 h, 24 h and 72 h) and imaged by IVIS system and data was analysed with Living Image software.
Statistical analysis and reproducibility
Significant difference between two groups was determined using two-tailed t-test analysis. Significant difference for multiple comparisons was determined using one-way ANOVA with Tukey post-test analysis. P > 0.05 was considered to be statistically not significant, and P < 0.05 was considered to be significant (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.001). Cryo-TEM study in Fig. 1f, g was conducted once. In cellulo nucleic acid phase separation in Fig. 3a was performed three times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this study are available in this article and its Supplementary Information file. All data underlying this study are available from the authors, through the corresponding authors, upon request. Source data are provided with this paper.
References
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Wu, Y. M. et al. Host defense peptide mimicking antimicrobial amino acid polymers and beyond: Design, synthesis and biomedical applications. Prog. Polym. Sci. 141, 101679 (2023).
Engler, A. C. et al. Antimicrobial polycarbonates: investigating the impact of balancing charge and hydrophobicity using a same-centered polymer approach. Biomacromolecules 14, 4331–4339 (2013).
Nederberg, F. et al. Biodegradable nanostructures with selective lysis of microbial membranes. Nat. Chem. 3, 409–414 (2011).
Qiao, Y. et al. Highly dynamic biodegradable micelles capable of lysing Gram-positive and Gram-negative bacterial membrane. Biomaterials 33, 1146–1153 (2012).
Chin, W. et al. Biodegradable broad-spectrum antimicrobial polycarbonates: investigating the role of chemical structure on activity and selectivity. Macromolecules 46, 8797–8807 (2013).
Liu, L. H. et al. Main-chain imidazolium oligomer material as a selective biomimetic antimicrobial agent. Biomaterials 33, 8625–8631 (2012).
Riduan, S. N. et al. Ultrafast killing and self-gelling antimicrobial imidazolium oligomers. Small 12, 1928–1934 (2016).
Kuroda, K. & Degrado, W. F. Amphiphilic polymethacrylate derivatives as antimicrobial agents. J. Am. Chem. Soc. 12, 4128–4129 (2005).
Tew, G. N., Scott, R. W., Klein, M. L. & Degrado, W. F. De novo design of antimicrobial polymers, foldamers, and small molecules: from discovery to practical applications. Acc. Chem. Res 43, 30–39 (2010).
Lienkamp, K. et al. Antimicrobial polymers prepared by ROMP with unprecedented selectivity: a molecular construction kit approach. J. Am. Chem. Soc. 130, 9836–9843 (2008).
Mowery, B. P., Lindner, A. H., Weisblum, B., Stahl, S. S. & Gellman, S. H. Structure-activity relationships among random nylon-3 copolymers that mimic antibacterial host-defense peptides. J. Am. Chem. Soc. 131, 9735–9745 (2009).
Yang, Y. et al. Antimicrobial cationic polymers: from structural design to functional control. Polym. J. 50, 33–44 (2018).
Olmos, D. & González-Benito, J. Polymeric materials with antibacterial activity: a review. Polymers 13, 613 (2021).
Cansu, E., Kazuma, Y. & Edmund, F. P. Biomimetic antimicrobial polymers: recent advances in molecular design. Polym. Chem. 9, 2407–2427 (2018).
Si, Z. et al. Polymers as advanced antibacterial and antibiofilm agents for direct and combination therapies. Chem. Sci. 13, 345–364 (2021).
Pham, P., Oliver, S. & Boyer, C. Design of antimicrobial polymers. Macromol. Chem. Phys. 224, 2200226 (2023).
Haktaniyan, M. & Bradley, M. Polymers showing intrinsic antimicrobial activity. Chem. Soc. Rev. 51, 8584–8611 (2022).
Alkarri, S., Bin Saad, H. & Soliman, M. On antimicrobial polymers: development, mechanism of action, international testing procedures, and applications. Polymers 16, 771 (2024).
Shabani, S. et al. Synthetic peptide branched polymers for antibacterial and biomedical applications. Nat. Rev. Bioeng. 2, 343–361 (2024).
Cruz, G. S., Santos, A. T. D., Brito, E. H. S. & Rádis-Baptista, G. Cell-penetrating antimicrobial peptides with anti-infective activity against intracellular pathogens. Antibiotics 11, 1772 (2022).
Silei, B. et al. A polymeric approach toward resistance-resistant antimicrobial agent with dual-selective mechanisms of action. Sci. Adv. 7, 9917 (2021).
Mehrnoosh, K. H. & Robert, V. Membrane adsorption enhances translocation of antimicrobial peptide Buforin 2. J. Phys. Chem. B. 128, 8469–8476 (2024).
Frimodt-Møller, J., Campion, C., Nielsen, P. E. & Løbner-Olesen, A. Translocation of non-lytic antimicrobial peptides and bacteria penetrating peptides across the inner membrane of the bacterial envelope. Curr. Genet 68, 83–90 (2022).
Hadjicharalambous, A., Bournakas, N., Newman, H., Skynner, M. J. & Beswick, P. Antimicrobial and cell-penetrating peptides: understanding penetration for the design of novel conjugate antibiotics. Antibiotics 11, 1636 (2022).
Zhang, H. D. et al. Switching from membrane disrupting to membrane crossing, an effective strategy in designing antibacterial polypeptide. Sci. Adv. 9, abn0771 (2023).
Chindera, K. et al. The antimicrobial polymer PHMB enters cells and selectively condenses bacterial chromosomes. Sci. Rep. 6, 23121 (2016).
McKenna, M. Antibiotic resistance: The last resort. Nature 499, 394–396 (2013).
Sheu, C. C., Chang Y. T., Lin S. Y., Chen Y. H. & Hsueh, P. R. Infections Caused by Carbapenem-Resistant Enterobacteriaceae: An Update on Therapeutic Options. Front. Microbiol. 10, https://doi.org/10.3389/fmicb.2019.00080 (2019).
Guo, D. et al. Liquid-Liquid phase separation in bacteria. Microbiol. Res. 281, 127627 (2024).
Ladouceur, A. M. et al. Clusters of bacterial RNA polymerase are biomolecular condensates that assemble through liquid-liquid phase separation. Proc. Natl. Acad. Sci. Usa. 117, 18540–18549 (2020).
Nandana, V. & Schrader, J. M. Roles of liquid-liquid phase separation in bacterial RNA metabolism. Curr. Opin. Microbiol. 61, 91–98 (2021).
Al-Husini, N., Tomares, D. T., Bitar, O., Childers, W. S. & Schrader, J. M. α-Proteobacterial RNA degradosomes assemble liquid-liquid phase-separated RNP bodies. Mol. Cell. 71, 1027–1039.e14 (2018).
Al-Husini, N. et al. BR-bodies provide selectively permeable condensates that stimulate mRNA decay and prevent release of decay intermediates. Mol. Cell. 78, 670–682.e8 (2020).
Megan, C. C. & Rohit, V. P. Making the case for disordered proteins and biomolecular condensates in bacteria. Trends Biochem. Sci. 45, 668–680 (2020).
Greening, C. & Lithgow, T. Formation and function of bacterial organelles. Nat. Rev. Microbiol. 18, 677–689 (2020).
Muthunayake, N. S., Tomares, D. T., Childers, W. S. & Schrader, J. M. Phase-separated bacterial ribonucleoprotein bodies organize mRNAdecay. Wiley Interdiscip. Rev. Rna 11, 1–26 (2020).
Abbondanzieri, E. A. & Meyer, A. S. More than just a phase: the search for membraneless organelles in the bacterial cytoplasm. Curr. Genet. 65, 691–694 (2019).
Sneideris, T. et al. Targeting nucleic acid phase transitions as a mechanism of action for antimicrobial peptides. Nat. Commun. 14, 7170 (2023).
Hilal, S. H., Karickhoff, S. W., Carreira, L. A. & Shrestha, B. P. Estimation of carboxylic acid ester hydrolysis rate constants. Qsar Combinator. Sci. 22, 917–925 (2004).
Koh, C. H. et al. Carbene formation as a mechanism for efficient intracellular uptake of cationic antimicrobial carbon acid polymers. Nat. Commun. 16, 6460 (2025).
Zhou, H. B. et al. Mechanism of DNA-induced phase separation for transcriptional repressor VRN1. Angew. Chem. Int. Ed. 58, 4858–4862 (2019).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters 3, e201702000010 (2017).
Guo, Q. et al. Hydrogen-bonds mediate liquid-liquid phase separation of mussel derived adhesive peptides. Nat. Commun. 13, 5771 (2022).
Nonejuie, P., Burkart, M., Pogliano, K. & Pogliano, J. Bacterial cytological profiling rapidly identifies the cellular pathways targeted by antibacterial molecules. Proc. Natl. Acad. Sci. 110, 16173 (2013).
Nonejuie, N. et al. Application of bacterial cytological profiling to crude natural product extracts reveals the antibacterial arsenal of Bacillus subtilis. J. Antibiot. 69, 353–361 (2016).
Ouyang, X. et al. Classification of antimicrobial mechanism of action using dynamic bacterial morphology imaging. Sci. Rep. 12, 11162 (2022).
Chin, W. et al. A macromolecular approach to eradicate multidrug resistant bacterial infections while mitigating drug resistance onset. Nat. Commun. 9, 917 (2018).
Yong, M. et al. Membrane potential-dependent uptake of cationic oligoimidazolium mediates bacterial DNA damage and death. Antimicrob. Agents Chemother. 67, e00355-23 (2023).
Zhong, W. B. et al. Designer broad-spectrum polyimidazolium antibiotics. Proc. Natl. Acad. Sci. 117, 31376–31385 (2020).
Wang, Z. Q. et al. A naturally inspired antibiotic to target multidrug-resistant pathogens. Nature 601, 606–611 (2022).
Mitcheltree, M. J. et al. A synthetic antibiotic class overcoming bacterial multidrug resistance. Nature 599, 507–512 (2021).
Imai, Y. et al. A new antibiotic selectively kills Gram-negative pathogens. Nature 576, 459–464 (2019).
Si, Z. et al. A glycosylated cationic block poly(β-peptide) reverses intrinsic antibiotic resistance in all ESKAPE gram-negative bacteria. Angew. Chem. Int. Ed. 59, 6819 (2020).
Mahajan, D., Madugula, V. & Lu, L. Rab8 and TNPO1 are ciliary transport adaptors for GTPase Arl13b by interacting with its RVEP motif containing ciliary targeting sequence. J. Biol. Chem. 299, 104604 (2023).
Acknowledgements
We thank the funding support from the Singapore Ministry of Education under its Singapore Ministry of Education Academic Research Fund (MoE AcRF) Tier 3 Awards (MOE2018-T3-1-003 and MOE-MOET32023-0004). We thank faculty and staff from the NTU Institute of Structural Biology for assistance in cryo-TEM imaging. We thank Melvin Yong and Yunn-Hwen Gan for their discussions on the OIM mechanism study. We thank TTSH and SCELSE for provision of bacterial strains.
Author information
Authors and Affiliations
Contributions
M.B.C. supervised and guided the overall research; X.X., L.L., E.P.G., T.T.D. and M.B.C. conceived the project. X.X. synthesized and characterized the monomers and oligomers. Q.H.N.V. conducted mass spectrometry testing. X.X. and C.K. carried out the in vitro biological testing. X.X. and D.M conducted in vitro transcription and translation testing. X.X. and C.T. conducted bacterial resistance testing. T.L.P carried out model compound study to answer reviewers’ question. X.X. carried out mechanism study. X.X. carried out the mice experiments and other tests. T.T.D. analyzed the histological data. X.X. and M.B.C. wrote the manuscript with contributions from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The Authors declare no other financial and non-financial competing interests except the following: An unpublished patent (New United States Provisional Patent Application NTU Ref: 2025-044) titled “Biodegradable Poly(Imidazolium Ester)s As Antimicrobial Agent” is associated with this work. The applicant is NTU and the inventors include M.B.C., X.F.X., T.T.D. and E. P. G. The patent covers the novel compound composition (P8): their syntheses, in vitro MIC test, cytotoxicity tests, mechanism study and in vivo performance.
Peer review
Peer review information
Nature Communications thanks Edmund Palermo, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, X., Korupalli, C., Mahajan, D. et al. Poly(imidazolium ester) antibiotic forms intracellular polymer-nucleic acid biomolecular condensates and fight drug-resistant bacteria. Nat Commun 16, 9854 (2025). https://doi.org/10.1038/s41467-025-64828-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64828-7