Abstract
Prevotella is a poorly understood symbiont within the gut microbiota, and its role in host health remains unclear. Here, we report a case-control analysis of a Chinese cohort, in which patients with obesity showed significantly lower fecal Prevotella copri abundance. The administration of P. copri can alleviate fat deposition and promote browning in inguinal white adipose tissue (iWAT) of mice with high-fat diet (HFD)-induced adiposity. We identify succinate as a key metabolite that activates succinate receptor 1 (SUCNR1) to mimic the effects of P. copri. Mice treated with either P. copri or succinate have increased iWAT-resident macrophage populations and IL-6-STAT3 pathway activation. Forkhead box M1 (FOXM1) directly upregulates IL-6 to activate IL-6-STAT3 signaling in P. copri- or succinate-induced iWAT browning. In human WAT, we identify that P. copri and succinate levels are positively correlated with TBX1 and UCP1 transcription. This study reveals the function of P. copri in promoting iWAT browning and suggests potential strategies for alleviating adiposity and associated disorders.
Similar content being viewed by others
Introduction
Obesity is an escalating epidemic linked to metabolic disorders like hepatic steatosis and insulin resistance1. Accordingly, understanding the mechanisms of interventions for modulating lipid metabolism represents a long-standing goal of clinical research. There are three types of adipocytes in mammals: white, brown, and beige, the latter of which is a distinct subset of brown-like thermogenic adipocytes transdifferentiated from white adipose tissue (WAT)2,3. Both mesenchymal stem cells located at the WAT perivascular region and some differentiated adipocytes are the progenitors for de novo beige adipogenesis3,4,5. Brown and beige are thermogenic adipocytes that could be manipulated to alleviate metabolic disorders due to their specific expression of mitochondrial uncoupling protein 1 (UCP1)2,3,4,5. In addition to UCP1, several beige-specific markers, such as transmembrane protein 26 (TMEM26) and T-box1 (TBX1), are known to be activated during the browning of WAT3. Emerging evidence demonstrated that the browning capability of subcutaneous WAT is substantially higher than the visceral WAT2,3. Since brown adipose tissue (BAT) is present primarily in small mammals and human neonates but declines with age and obesity2,6, stimulation of WAT browning represents a promising alternative intervention strategy to combat obesity and associated disorders7,8.
The gut microbiome participates in regulating diverse host metabolic processes9, including thermogenesis in adipose tissues10,11. Certain commensal gut bacteria (e.g. Ruminococcus torques, Limosilactobacillus reuteri, and Lactobacillus reuteri), could promote WAT browning and mitigate obesity or metabolic comorbidities. These effects are primarily mediated by specific bacterial metabolites, such as deoxycholic acid and short-chain fatty acids12,13,14. Genus Prevotella is a pivotal keystone symbiont within the gut microbiota15, with Prevotella copri being the most widely known and abundant species16. However, host - P. copri interactions have received relatively little attention, and interpretations of this relationship remain controversial. For instance, intestinal P. copri abundance is reportedly higher in children with obesity17 but lower in adults with obesity versus healthy controls18. In pigs, P. copri promotes fat accumulation via activating inflammation19. However, in mice, P. copri improves glucose homeostasis under high-fiber intake by promoting hepatic glycogen storage20, whereas it leads to insulin resistance and glucose intolerance when fed a high-fat diet (HFD)21. Despite mounting evidence of positive or negative links of P. copri to obesity and other metabolic disorders17,18,19,20,21, whether and how P. copri may play a specific role in white adipose browning remains largely unknown.
In the present study, we found reduced fecal P. copri abundance in patients with obesity. In HFD-fed mice, oral P. copri administration increased inguinal WAT (iWAT) browning and attenuated weight gain. We identified succinate as its key metabolite, activating succinate receptor 1 (SUCNR1) to mimic P. copri’s effects. The macrophage-derived IL-6-STAT3 pathway, triggered by forkhead box M1 (FOXM1)-mediated Il-6 transcription, was essential for this P. copri- or succinate-inducible iWAT browning. Human data showed positive correlations between WAT TBX1/UCP1 expression and fecal P. copri/succinate levels. These findings propose P. copri-succinate signaling as a meaningful anti-obesity strategy.
Results
Individuals with obesity have aberrantly low abundance of P. copri in their fecal microbiomes
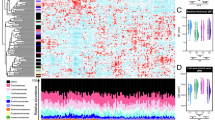
To explore potential relationships of gut microbes with obesity, we performed microbial metagenomics in individuals without obesity [Body mass index (BMI) < 28 kg/m2, n = 15] and individuals with obesity (BMI ≥ 28 kg/m2, n = 19) to investigate obesity-associated changes in fecal microbiota. The detailed information for the cohorts is presented in Supplementary Table 1. Linear discriminant analysis effect size (LEfSe) data and the heat maps revealed the total differences at phylum, order, family, genus, and species levels between the non-obesity and obesity groups (Fig. 1a, Supplementary Fig. 1a–e). We analyzed bacterial species that upregulated or downregulated in the two groups (Fig. 1b, c). Notably, among taxa with relative higher proportions in the fecal microbes, P. copri showed the most significant reduction in patients with obesity, with concomitant but less pronounced decrease in Coprococcus eutactus and Bifidobacterium dentium (Fig. 1d). We subsequently collected 92 samples from people with different BMI (detailed in Supplementary Data 1) and analyzed their fecal P. copri abundance by qPCR using specific primers22. In contrast to the non-obesity group, the abundance of P. copri was significantly lower in the people with obesity (Fig. 1e). Furthermore, inverse correlations were observed between P. copri abundance and several adiposity indices, including BMI, waist circumference, body fat percentage, and visceral fat area (Fig. 1f–i). These data indicated that people with obesity have aberrantly low abundance of P. copri in their fecal microbiomes.
Metagenomic sequencing datasets (a–d) showed the differences of fecal microbiota in individuals without obesity (BMI < 28 kg/m2, n = 15) and individuals with obesity (BMI ≥ 28 kg/m2, n = 19). a Taxonomic cladogram generated from LEfSe analysis of metagenomic sequencing. b Venn diagram shows the number of species common and unique to the two groups. c Volcano map shows DESeq2 comparative analysis of species differences between the two groups. d The result of top-15 significantly differential species between the people with and without obesity. e Relative abundance of fecal P. copri in people with or without obesity (n = 92). f–i Spearman correlations (two-tailed Spearman’s rank test) between the relative abundance of fecal P. copri (a.u.) and BMI (f), waist circumference (g), body fat percentage (h) and visceral fat area (i) (n = 92). Values are mean ± SEM. Statistical analysis was performed using DESeq2 with screened parameter: P-value < 0.05 and expression difference multiple |log2FC | ≥1 for c, two-tailed Wilcoxon rank sum test for d and significance was assessed by FDR correction for multiple comparisons: *FDR < 0.05; **FDR < 0.01, and two-tailed Mann–Whitney U test for e. Source data are provided as a Source Data file.
Supplementation of HFD-induced adiposity model mice with P. copri alleviates adiposity and promotes iWAT browning
In light of the observed inverse association between P. copri abundance and obesity, we investigated this bacterium’s potential role in lipid metabolism. In a classic HFD murine model, male and female mice received P. copri (10⁸ CFU/kg BW) every other day for 8 weeks showed attenuated body weight gain, reduced fat deposition, and higher rectal temperature, whereas no effect was observed at the 109 CFU/kg BW dosage (Supplementary Figs. 2–3). Given this dose-dependent response, coupled with our focus on white adipose browning and reported sexual dimorphism in Ucp1 and Tbx1 expression23,24 (higher in females as shown in Supplementary Fig. 4), we subsequently administered live or pasteurized P. copri (10⁸ CFU/kg BW) exclusively to female HFD-fed mice for 8 weeks (denoted as HFD P. copri and HFD P. copri-pasteurized groups, respectively) (Fig. 2a).
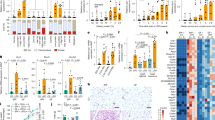
8-week-old C57BL/6 J mice under chow diet was given normal saline (Control), and those under HFD conditions were given normal saline (HFD Control), 108 CFU/kg BW P. copri (HFD P. copri), or 108 CFU/kg BW pasteurized P. copri (HFD P. copri-pasteurized) for 8 weeks. a Experimental scheme for (b–u). b, c Relative abundance of fecal (b) (n = 6 biological replicates) and cecal (c) (n = 9 biological replicates) P. copri assessed by qPCR. d Body weight of 16-week-old C57BL/6 J mice following 8 weeks’ treatments (n = 9 mice). e Representative pictures of body composition scanning (n = 6 mice). f Fat index (%) (n = 6 mice). g Lean mass (n = 6 mice). h Representative images of iWAT from each group. i iWAT weight (n = 9 mice). j Ratio of iWAT weight to body weight (%) (n = 9 mice). k H&E-stained pictures of the iWAT section (Scale bars, 50 μm), (n = 3 biological replicates). l The size of adipocytes in iWAT (n = 3 biological replicates). m Rectal body temperature (n = 9 mice). n-q CO2 production (n), O2 consumption (o) respiratory exchange rate (RER) (p) and heat production (q) curves between HFD Control, HFD P. copri-pasteurized and HFD P. copri groups (n = 4 mice), the data were analyzed by covariance (ANCOVA). r Relative mRNA levels of genes related to thermogenesis, lipogenesis and lipolysis in iWAT (n = 7 biological replicates). s, t Western blot analysis (s) and quantification (t) of the protein contents of thermogenic and lipogenic markers in iWAT, with β-actin serving as a loading control (n = 8 biological replicates for UCP1 and TBX1, n = 6 biological replicates for PPARγ and AP2). u Representative immunofluorescence images of UCP1 (Red) from the iWAT section (Scale bars, 20 μm), (n = 3 independent experiments). Values are mean ± SEM. Statistical analysis was performed using One-way ANOVA with LSD post hoc test for m, One-way ANOVA with Dunnett’s T3 post hoc test for (b–d) and (i, j) Kruskal-Wallis test followed by Dunn’s post hoc test for (f, g, l, r, t). Source data are provided as a Source Data file.
qPCR analysis of feces and cecal contents after 6 weeks confirmed successful colonization, with P. copri abundance being significantly higher in live-bacterium-treated mice compared to pasteurized or saline controls (Fig. 2b, c). Mice receiving P. copri demonstrated significant resistance to HFD-induced metabolic alterations, including attenuated weight gain, reduced fat accumulation, decreased iWAT mass, and smaller adipocyte size after 8 weeks of treatment, without affecting lean mass or feed intake (Fig. 2d–l, Supplementary Fig. 5a). While P. copri administration showed a modest but non-significant reduction in epididymal WAT (eWAT) weight and adipocyte area, it had no observable effect on interscapular BAT (iBAT) mass or microstructure (Supplementary Fig. 5b–i). Notably, the P. copri-treated group exhibited significantly reduced hepatic lipid deposition and improved glucose tolerance and insulin sensitivity compared to HFD or P. copri-pasteurized Control (Supplementary Fig. 6). These results hinted that the prevention of iWAT expansion plays a pivotal role in P. copri-mediated protection against HFD-induced weight gain.
Decreased lipogenesis, increased lipolysis, and enhanced thermogenesis all could lead to reduction of WAT weight25. At week 7 post-treatment, we first monitored murine core body temperature and respiratory metabolism. Mice administered P. copri exhibited significantly elevated rectal temperatures compared to both the P. copri-pasteurized or HFD Control groups (Fig. 2m). Respiratory metabolic analysis demonstrated substantial increases in O2 consumption, CO2 production, respiratory exchange ratio (RER) (VCO2/VO2), and heat production in the P. copri-treated mice (Fig. 2n–q). These collective changes indicate enhanced basal metabolic activity. Subsequent molecular analysis of iWAT, the most P. copri-responsive adipose depot, demonstrated no significant alterations in lipogenic markers PPARγ or AP226 protein expression, though Ap2 mRNA was downregulated (Fig. 2r–t). For lipolytic markers, expression of Atgl, Hsl and Cpt127 was unchanged across groups (Fig. 2r). Notably, the thermogenic marker Ucp1 and the beiging gene Tbx13 were significantly elevated at both the mRNA and protein levels in the P. copri-treated iWAT (Fig. 2r-t), with immunofluorescence confirming pronounced UCP1 induction (Fig. 2u). These data established that P. copri prevents iWAT expansion primarily through Ucp1/Tbx1-mediated thermogenesis rather than lipogenic or lipolytic pathway modulation.
The primary P. copri metabolite, succinate, recapitulates the P. copri promotion of iWAT browning
The modulation of host metabolism by a specific bacterium is predominantly dependent on its metabolites, which act as potent signal transmitters for conveying information to target tissues12,13,14. Our findings revealed that solely live, not pasteurized, P. copri alleviated HFD-induced adiposity and reduced WAT weight (Fig. 2d–j), indicating that its beneficial effects may rely on the production of bioactive metabolites. Consequently, targeted metabolomic analysis of media from standard P. copri cultures identified succinate as the primary metabolite (Fig. 3a). Furthermore, oral gavage of P. copri significantly elevated fecal and cecal succinate levels relative to the HFD control group (Fig. 3b, c). Collectively, these data confirmed succinate as the predominant metabolite of P. copri.
a Targeted metabolomic analysis of organic acids in the culture supernatant of P. copri (n = 6 biological replicates). b, c The succinate content in fecal (b) (n = 6 biological replicates for HFD Control group and n = 7 biological replicates for HFD P. copri group) and cecal contents (c) (n = 6 biological replicates) of mice from HFD Control and HFD P. copri groups. d Experimental scheme for e-v. 8-week-old C57BL/6 J mice were given normal saline (HFD Control), 0.1 g/kg BW sodium succinate (SS) (HFD 0.1 g/kg BW SS), or 0.25 g/kg BW SS (HFD 0.25 g/kg BW SS) for 8 weeks under HFD feeding. e Body weight at 16 weeks old (n = 9 mice). f Representative pictures of body composition scanning of mice following 8 weeks’ treatment (n = 6 mice). g Fat index (%) (n = 6 mice). h Lean mass (n = 6 mice). i Representative images of iWAT. j iWAT weight (n = 9 mice). k Ratio of iWAT weight to body weight (n = 9 mice). l H&E-stained iWAT section (Scale bars, 50 μm), (n = 3 biological replicates). m Adipocyte sizes in iWAT (n = 3 biological replicates). n Rectal body temperature (n = 9). o-r CO2 production (o), O2 consumption (p) RER (q) and heat production (r) curves between HFD Control and HFD 0.25 g/kg BW SS groups (n = 4 mice), the data were analyzed by covariance (ANCOVA). s Relative mRNA levels of genes related to thermogenesis, lipogenesis and lipolysis in iWAT (n = 6 biological replicates). t, u Western blot analysis (t) and quantification (u) of the protein contents of thermogenic and lipogenic markers in iWAT (n = 4 biological replicates). v Representative immunofluorescence images of UCP1 (Red) from the iWAT section (Scale bars, 40 μm), (n = 3 independent experiments). Values are mean ± SEM. Statistical analysis was performed using two-tailed Mann-Whitney U test for b-c One-way ANOVA with LSD post hoc test for e, j-k, n, One-way ANOVA with Dunnett’s T3 post hoc test for m Kruskal-Wallis test followed by Dunn’s post hoc test for (g, h, s, u). Source data are provided as a Source Data file.
To determine whether succinate mediates P. copri-induced iWAT browning, we provided the HFD-fed mice with sodium succinate (SS) at 0.1 or 0.25 g/kg BW for 8 weeks (Fig. 3d). Mice received 0.25 g/kg BW SS exhibited significantly reduced weight gain, lower fat percentage, decreased iWAT weights, and smaller adipocyte sizes compared to HFD Control, with lower eWAT weight but no change in iBAT proportion or feed intake (Fig. 3e-m, Supplementary Fig. 7a-i). Given the role of P. copri in iWAT browning, its primary metabolite likely enhances thermogenesis. Supporting this hypothesis, the SS group showed elevated rectal temperature, CO2 production, RER, and heat generation versus HFD Control (Fig. 3n-r). Furthermore, SS induced substantial upregulation of Ucp1/Tbx1 mRNA and protein levels, and higher UCP1-stained immunofluorescence intensity (Fig. 3s–v). Similar to P. copri, SS (0.25 g/kg BW) decreased hepatic TG accumulation and improved glucose homeostasis (Supplementary Fig. 8). For in vitro validation, primary iWAT-derived cells (iWATDCs)28 were treated with P. copri cultures (P. copri sup) or SS at serum-equivalent concentrations from P. copri-treated mice. Both treatments elevated UCP1/TBX1 expression, decreased Oil red O-stained lipid droplets, and increased maximal oxygen consumption rate (SeaHorse assay) (Supplementary Fig. 9). These results established succinate as a bioactive molecule through which P. copri promotes iWAT browning.
SUCNR1 is required for P. copri or succinate promotion of iWAT browning
As a signaling molecule, succinate engages diverse cell types via its cognate receptor SUCNR129, prompting investigation into its necessity for P. copri- or SS-induced iWAT browning. Comparative analysis of Sucnr1 mRNA levels across murine adipose depots (iWAT, eWAT, iBAT) identified iWAT as the predominant site of expression (Fig. 4a). Quantitative assessment via RT-qPCR and WB revealed significant SUCNR1 upregulation at both mRNA and protein levels in iWAT from P. copri- or SS-treated mice versus HFD controls (Fig. 4b–d). For functional validation in iWATDCs, SUCNR1 inhibition using compound 4c abolished Ucp1 and Tbx1 induction by P. copri supernatant or SS (Supplementary Fig. 10). To further determine the importance of SUCNR1, in vivo confirmation employed Sucnr1-/- female mice30 subjected to oral P. copri or SS administration under HFD conditions (Fig. 4e). Body weight changes, fat mass, iWAT proportion, and adipocyte size remained comparable across control and treatment groups in Sucnr1-/- mice (Fig. 4f–k). Moreover, rectal temperature decreased in supplemented groups, while O2 consumption, CO2 production, RER, and heat production showed no significant differences versus controls (Fig. 4l–p). Neither P. copri nor SS affected Tbx1 or Ucp1 mRNA levels nor UCP1 protein abundance in iWAT (Fig. 4q–s). Collectively, these results demonstrated that SUCNR1 is essential for P. copri and succinate-mediated iWAT browning.
a Relative Sucnr1 mRNA level between iWAT, eWAT and BAT in WT mice (n = 8 biological replicates). b Relative Sucnr1 mRNA level in iWAT of WT C57BL/6 J mice treated with normal saline, P. copri, or SS uder HFD conditions (n = 7 biological replicates). c-d Western blot analysis (c) and quantification (d) of the protein expression of SUCNR1 in iWAT of WT C57BL/6 J mice, with β-actin serving as a loading control (n = 4 biological replicates). e Experimental scheme in C57BL/6J-Sucnr1-/- mice for f-s. 8-week-old C57BL/6J-Sucnr1-/- mice were given normal saline (HFD Control), 108 CFU/kg BW P. copri (HFD P. copri), or 0.25 g/kg BW SS (HFD SS) for 8 weeks under HFD conditions. f Body weight of 16-week-old C57BL/6J-Sucnr1-/- mice following 8 weeks’ treatment (n = 8 mice). g Representative images of iWAT. h iWAT weight (n = 8 mice). i Ratio of iWAT weight to body weight (n = 8 mice). j H&E-stained iWAT sections (Scale bars, 50 μm), (n = 3 biological replicates). k Adipocyte sizes in iWAT (n = 3 biological replicates). l Rectal body temperature (n = 8 mice). m–p CO2 production (m) O2 consumption (n) RER (o) and heat production (p) curves (n = 4 mice for HFD Control group, n = 3 mice for HFD P. copri and HFD SS groups), the data were analyzed by covariance (ANCOVA). q Relative mRNA levels of Ucp1, Tbx1, Ap2, and Cpt1 in iWAT of Sucnr1-/- mice (n = 6 biological replicates). r, s Western blot analysis (r) and quantification (s) of the protein contents of thermogenic and lipogenic markers in iWAT (n = 3 biological replicates). Values are mean ± SEM. Statistical analysis was performed using One-way ANOVA with LSD post hoc test for k, Kruskal-Wallis test followed by Dunn’s post hoc test for a, b, d, f, h-i, l, q, s. Source data are provided as a Source Data file.
The IL-6-STAT3 pathway plays a crucial role for iWAT browning in response to P. copri or succinate
Given that succinate-SUCNR1 signaling is known to trigger inflammatory responses29; we next examined serum levels of inflammatory factors, inflammatory gene expression in iWAT, and the macrophage proportion in iWAT. Among the tested inflammatory factors, serum IL-6 but not TNF-α levels were significantly increased in WT mice exposed to P. copri or SS (Fig. 5a, b).
a, b Serum IL-6 (a) and TNF-α (b) contents of WT C57BL/6 J mice treated with normal saline, P. copri, or SS (n = 6 biological replicates). c Relative mRNA levels of inflammation genes in iWAT of WT C57BL/6 J mice (n = 6 biological replicates for Il-1β, Tnf-α and Tgf-β; n = 8 biological replicates for Il-6 and Il-33). d, e Western blot analysis (d) and quantification (e) of the protein contents of IL-6, p-STAT3, and STAT3 in iWAT of WT C57BL/6 J mice (n = 6 biological replicates). f Flow cytometry. CD45.2+ represents immune cells; F4/80+ and CD11b+ represent macrophages; CD86+ represents M1 macrophages. g The proportion of F4/80 and CD11b positive cells and MFI of CD86 positive cells (n = 6 biological replicates). h, i ChIP assay was performed with qPCR to analyze the binding of p-STAT3 to Ucp1 (h) and Tbx1 (i) promoters in iWATDCs (n = 6 biological replicates). j Experimental scheme for C57BL/6J-Il-6-/- mice for k-z. 8-week-old C57BL/6J-Il-6-/- mice were given normal saline (HFD Control) (n = 6 mice), 108 CFU/kg BW P. copri (HFD P. copri) (n = 6 mice), or 0.25 g/kg BW SS (HFD SS) (n = 7 mice) for 8-week under HFD conditions. k Body weight of 16-week-old C57BL/6J-Il-6-/- mice following 8 weeks’ treatment. l Representative pictures of body composition scanning. m Fat index (%). n Representative images of iWAT. o iWAT weight. p Ratio of iWAT weight to body weight. q H&E-stained iWAT sections (Scale bars, 50 μm), (n = 3 biological replicates). r Adipocyte sizes in iWAT (n = 3 biological replicates). s Rectal body temperature (n = 6 mice for HFD Control and HFD P. copri group; n = 7 mice for HFD SS group). t–w CO2 production (t) O2 consumption (u) RER (v), and heat production (w) curves (n = 3 mice for HFD Control and HFD P. copri groups and n = 4 mice for HFD SS group), the data were analyzed by covariance (ANCOVA). x Relative mRNA levels of lipid metabolism-related genes in iWAT of C57BL/6J-Il-6-/- mice (n = 5 biological replicates for Il-6-KO-HFD P. copri group; n = 5 biological replicates for Il-6-KO-HFD Control and Il-6-KO-HFD SS groups of Hsl; n = 6 biological replicates for Il-6-KO-HFD Control and Il-6-KO-HFD SS groups of Ucp1, Tbx1, Ap2, Lpl, Fas, Cpt1 and Sucnr1). y,z Western blot analysis (y) and quantification (z) of related proteins (n = 3 biological replicates). Values are mean ± SEM. Statistical analysis was performed using One-way ANOVA with Dunnett’s T3 post hoc test for Il-6 of c, Kruskal-Wallis test followed by Dunn’s post hoc test for a-b, Il-1β, Tnf-α, Il-33 and Tgf-β of c, e, g–i, k, m, o–p, r-s, x, z. Source data are provided as a Source Data file.
Consistent with this, Il-6 gene transcript, IL-6 protein and STAT3 phosphorylation levels in iWAT also increased in both treatment groups (Fig. 5c-e). To trace the primary source of IL-6, in vitro confirmation was performed by adding P. copri sup or SS into cultures of RAW264.7 murine mononuclear macrophage leukemia cells31 or 3T3-L1 cells (a pre-adipocyte cell line that could differentiate into mature adipocytes)32. Analysis by RT-qPCR and WB revealed that P. copri sup- or SS-treated RAW264.7 cells exhibited significantly elevated Il-6 mRNA and protein levels, along with increased STAT3 phosphorylation accompanied by increased SUCNR1 expression (Supplementary Fig. 11a–c). Conversely, no alteration of Sucnr1, IL-6, p-STAT3, or UCP1 levels was observed in P. copri sup- or SS-exposed 3T3-L1 cells (Supplementary Fig. 11d–f). Moreover, we performed flow cytometry to quantify F4/80+/CD11b+/CD86+ macrophages within iWATDCs, as adipose tissue macrophages are established IL-6 secretors33. Notably, both P. copri and SS treatments profoundly expanded the F4/80+/CD11b+ macrophage proportion and upregulated CD86 mean fluorescence intensity (MFI) in iWATDCs of WT mice (Fig. 5f, g).
To functionally define whether macrophages are essential for the beneficial in vivo effects of P. copri or succinate, macrophage populations were selectively ablated through tail vein injection of clodronate liposomes (CL), an established macrophage-depleting method validated in prior studies34,35. Cohorts of CL-treated mice received either P. copri or SS for 8 weeks under HFD conditions (Supplementary Fig. 12a). It showed that the proportion of F4/80+ macrophages in circulation and iWAT was depleted by ~95% with CL (Supplementary Fig. 12b, c). P. copri or succinate did not alter the body weight gain, the body temperature, the white fat percentage, or thermogenic gene expression following macrophage depletion (Supplementary Fig. 12d-n). These findings collectively demonstrated that intact macrophage populations are indispensable for P. copri-derived succinate to exert its thermogenic effects.
The above findings and the direct IL-6-triggered p-STAT3 activation of Ucp1 expression through promoter binding28 prompted a focus on IL-6. ChIP assays revealed that P. copri sup or SS could enhance p-STAT3 binding at both the Ucp1 and Tbx1 promoter regions in iWATDCs (Fig. 5h, i). To evaluate the necessity of IL-6-STAT3 signaling, we assessed iWAT browning in HFD-fed Il-6-KO mice exposed to P. copri or SS (Fig. 5j). Strikingly, Il-6-KO mice in both treatments exhibited no change in body weight, iWAT index, rectal temperature, O2 consumption, heat production, and iWAT UCP1 and p-STAT3 contents, but increased iWAT adipocyte size, reduced CO2 production and lower RER (Fig. 5k-z), indicating that neither P. copri nor SS could induce iWAT browning in the absence of IL-6. However, both treatments still activated SUCNR1 in iWAT of Il-6-KO mice (Fig. 5x-z), suggesting SUCNR1 operates upstream of IL-6. These results supported a role for the macrophage-derived IL-6-STAT3 pathway in P. copri- or succinate-mediated iWAT browning.
The transcription factor, FOXM1, positively regulates Il-6 expression
Given the essential role of macrophage-derived IL-6/STAT3 signaling in P. copri or SS-induced iWAT browning, we performed bulk RNA sequencing on iWAT from SS-treated and HFD control mice to identify transcription factors (TFs) regulating P. copri-/SS-inducible Il-6 expression. Principal component analysis (PCA) revealed clear separation of the HFD and SS groups (Fig. 6a). We identified 270 differentially expressed genes (edgeR; |log2FoldChange | >1; P value < 0.05; Fig. 6b) that including several TFs, among which, Foxm1 showed the highest up-regulation in SS-treated iWAT (Fig. 6c). Based on reported roles of FOXM1 in macrophage migration and pro-inflammatory gene regulation36 and its co-upregulation with Il-6 in SS-treated iWAT (Fig. 5c, Fig. 6c), we assessed Foxm1 expression across models. iWAT tissue, primary iWATDCs, and RAW264.7 macrophages showed elevated Foxm1 mRNA/ protein after P. copri or SS exposure, but did not in 3T3-L1 adipocytes (Fig. 6d–f, Supplementary Fig. 13). This induction was abolished by the SUCNR1 antagonist 4c in iWATDCs (Fig. 6e). Crucially, lentiviral Foxm1 knockdown (via shRNA) in iWATDCs blocked P. copri sup/SS-induced upregulation of Foxm1, Il-6, Ucp1, Tbx1, and their protein products, as well as abrogated the activation of p-STAT3 (Fig. 6g, h). P. copri or SS failed to induce IL-6 when SUCNR1 was deficient or blocked (Supplementary Fig. 14a–d). However, Foxm1 remained inducible by P. copri or SS in Il-6-KO iWAT and iWATDCs but did not in iWAT from Sucnr1-KO mice (Supplementary Fig. 14e–j). These results suggested that FOXM1 acts downstream of SUCNR1 but upstream of IL-6 in iWAT recognition and response to P. copri or SS.
To search for transcription factor(s) that might regulate P. copri-/SS-inducible Il-6 expression, bulk RNA-seq was conducted by using iWAT from SS-treated and HFD control WT mice (n = 3 mice per group). a Principal component analysis (PCA). b Volcano maps. c Heatmap of differential expressed transcription factors. Heatmap plots were generated by log2 transformation of data. d Relative mRNA (n = 6 biological replicates) and protein expression (n = 3 biological replicates) levels of FOXM1 in iWAT of WT C57BL/6 J mice. e Relative mRNA (n = 6 biological replicates) and protein expression (n = 3 biological replicates) levels of FOXM1 in iWATDCs under P. copri sup or SS treatment in the presence or absence of 4 c. f Relative mRNA and protein expression levels of FOXM1 in RAW264.7 cells under P. copri sup or SS treatment (n = 4 biological replicates). g, h RT-qPCR (g) (n = 6 biological replicates) and western blotting (h) (n = 3 biological replicates) were used to detect the influence of shFoxm1 on the mRNA and protein expression of related markers in iWATDCs). i Schematic illustration of the 9-base pair sequence of two potential FOXM1-binding sites (BS1 and BS2) in the Il-6 1500-bp length promoter. j-m ChIP assay was performed with goat IgG control and FOXM1-specific antibodies. Quantitative PCR were used to analyze the binding of FOXM1 to Il-6 promoter at BS1 and BS2 in RAW264.7 cells (j, k) (n = 8 biological replicates) and iWATDCs (l, m) (n = 6 biological replicates). n 1500 bp-length of the Il-6 promoter luciferase construct was transfected into HEK293T cells with empty vector or the FOXM1 overexpression plasmid for 24 h. Transcriptional activation was analyzed with the dual luciferase reporter assay (n = 10 biological replicates). o Schematic diagram of BS1 and BS2 deficiency in the Il-6 promoter. p Dual luciferase reporter assay when deleting BS1, BS2, or the both (n = 10 biological replicates per group except n = 8 biological replicates for the FOXM1 group). Values are mean ± SEM. Statistical analysis was performed using edgeR with screened conditions: |log2FoldChange | >1, significant P value < 0.05 for b, One-way ANOVA with LSD post hoc test for k and p, One-way ANOVA with Dunnett’s T3 post hoc test for n, Kruskal-Wallis test followed by Dunn’s post hoc test for d–g, j, l–m. Source data are provided as a Source Data file.
To verify whether FOXM1 could directly bind to the Il-6 promoter to regulate its transcription, we subsequently identified two 9 bp predicted binding sites (BS1, −699 to −707; BS2, −954 to −962) in the Il-6 promoter (Fig. 6i) based on previous literature37 and the promoter prediction result from JASPAR. ChIP assays revealed that FOXM1 binding at both the Il-6 BS1 and BS2 sites was significantly enhanced in RAW264.7 cells (Fig. 6j, k), while being increased significantly at the BS1 and moderately at the BS2 region in iWATDCs, following P. copri sup or SS exposure (Fig. 6l, m). Furthermore, dual luciferase trans-activation assays showed that co-expression of pcDNA-FOXM1 could enhance luciferase expression driven by the Il-6 promoter compared to signal induced by either pcDNA-FOXM1 or the Il-6-LUC reporter alone (Fig. 6n), supporting the likelihood that FOXM1 could directly activate Il-6 transcription. Alternatively, deletion of either BS1, BS2, or both FOXM1 binding sites resulted in significantly lower luciferase expression under co-expression with pcDNA-FOXM1 compared to that driven by the WT Il-6 promoter (Fig. 6o, p). These results collectively demonstrated that FOXM1 acts as a direct regulator of Il-6 transcription by binding to either or both of the predicted binding sites (BS1 and BS2) within the Il-6 promoter.
Subcutaneous WAT TBX1 and UCP1 levels are positively correlated with P. copri abundance and succinate content in human individuals
In view of the P. copri or succinate promotion of iWAT browning, we explored links between thermogenic gene expression in human abdominal subcutaneous WAT and P. copri abundance or succinate contents. HPLC analysis showed significantly lower fecal and plasma succinate concentrations in individuals with obesity versus individuals without obesity (Fig. 7a, Supplementary Fig. 15a), aligning with reduced fecal P. copri abundance in the individuals with obesity (Fig. 1e). Notably, negative associations were observed for fecal and plasma succinate levels with BMI (Fig. 7b, Supplementary Fig. 15b). Positive correlations emerged between fecal P. copri abundance or succinate content and WAT UCP1/TBX1expression (Fig. 7c–f). Additionally, UCP1 and TBX1 protein levels were markedly lower in WAT from individuals with obesity compared to individuals without obesity (Fig. 7g). Given SUCNR1’s critical role in succinate-mediated WAT browning, we found reduced SUCNR1 mRNA and protein expression in WAT of individuals with obesity, with SUCNR1 mRNA positively correlating with plasma succinate concentrations (Supplementary Fig. 15c–e). Collectively, these data implicated reduced P. copri abundance, lowered succinate availability, and decreased SUCNR1 expression as factors in obesity pathogenesis, driven primarily by impaired WAT thermogenesis.
a Fecal succinate contents in individuals without obesity (BMI < 28 kg/m2, n = 15) and individuals with obesity (BMI ≥ 28 kg/m2, n = 19). Values are mean ± SEM. Statistical analysis was performed using two-tailed Mann-Whitney U test. b Spearman correlations (two-tailed Spearman’s rank test) between the fecal succinate contents and BMI (n = 34). c, d Two-tailed Spearman correlations between the fecal P. copri abundance and relative UCP1 (c) and TBX1 (d) mRNA level in abdominal subcutaneous WAT (n = 40). e, f Two-tailed Spearman correlations between the fecal succinate contents and relative UCP1 mRNA level (e), TBX1 (f) mRNA level in abdominal subcutaneous WAT (n = 34). g Western blot analysis showing the protein expression of UCP1 and TBX1 in human abdominal subcutaneous WAT from non-obesity (n = 4) and obesity (n = 4) groups. GAPDH serves as a loading control. Source data are provided as a Source Data file.
Discussion
This study identified decreased fecal P. copri abundance in people with obesity, aligning with reports linking Prevotella spp. to non-Westernized and non-obesity populations38,39, but contrasting with pediatric studies showing positive P. copri-obesity correlations17. Despite being prevalent gut symbionts, Prevotella (especially P. copri) remains understudied possibly due to technical hurdles in bacterial cultivation and inconsistent findings on their host health impacts20,40,41,42. We here demonstrated that P. copri at a relative low dosage (2–4 × 10⁶ CFU/mouse) could directly promote iWAT browning and ameliorate adiposity. This contrasts with prior correlational studies between P. copri and obesity or associated disorders17,18, and opposes reports of P. copri (1 × 10⁹ CFU/mouse) increasing fat deposition in germ-free mice19. This discrepancy may stem from host species differences, dosage effects [higher doses (2–4 × 10⁷ CFU/mouse) failed to reduce fat deposition in our preliminary trials] and strain heterogeneity since phylogenetic analyses reveal P. copri comprises four human subspecies41, and the lack of standardized strains may explain inter-study variability.
The characterization of microbial-derived metabolites provides critical insights into the intricate interplay between gut microbiota and host metabolism. Our study identified succinate as a bioactive metabolite produced by P. copri. While Kovatcheva-Datchary et al.20 observed elevated cecal succinate levels in P. copri-colonized mice, they ruled out its role in glucose homeostasis improvement. In contrast, we demonstrate that succinate functionally mimics P. copri’s ability to induce iWAT browning. As a metabolite intersecting host TCA cycle and microbial metabolic pathways43, succinate exhibits context-dependent effects on host physiology largely depending on its origin, concentration, and target tissues/organs43,44,45,46,47,48. Succinate has been shown to improve glycemic control via intestinal gluconeogenesis activation43 and enhance thermogenic respiration in brown adipocytes through mitochondrial SLC25A10-dependent mechanisms44. Paradoxically, succinate promotes inflammation45,46 and is elevated in diabetic and metabolic syndrome rodent models47,48. Our data reveal succinate’s role in iWAT browning via TBX1/UCP1 upregulation, countering HFD-induced adiposity. Clinically relevant correlations were observed between P. copri or succinate levels and subcutaneous WAT thermogenic markers in humans. Notably, in established adiposity models, P. copri inoculation or succinate administration attenuated weight gain (though statistically non-significant) while boosting iWAT thermogenic genes (Supplementary Fig. 16). These findings hinted the application potential of low dose P. copri or succinate in preventing and combating obesity.
In association with the P. copri and succinate promotion of iWAT browning, they also improve glucose homeostasis with lower insulin secretion (Supplementary Fig. 17) under HFD conditions. This context-dependent regulatory pattern of P. copri on glucose metabolism in our study seems to contrast sharply with Pedersen et al.’s findings indicating P. copri-mediated exacerbation of HFD-induced insulin resistance and glucose intolerance21. This discrepancy may arise from divergent administration regimens, whereby they administered P. copri at a higher dosage (5 × 10⁸ CFU/mouse twice weekly) compared to our protocol [2–4 × 10⁶ CFU/mouse (10⁸ CFU/kg BW) every two days]. However, Kovatcheva-Datchary et al.20 paradoxically demonstrated that daily oral gavage of P. copri (10⁸ CFU/mouse) enhanced glucose homeostasis under high-fiber conditions. In fact, our investigation reveals a biphasic dose-response relationship: lower dosages (2–4 × 10⁶ CFU/mouse for P. copri or 0.25 g/kg succinate) significantly improve glucose tolerance and mitigate HFD-induced weight gain, whereas higher dosages (2–4 × 10⁷ CFU/mouse or 0.5 g/kg succinate) fail to confer metabolic protection at most circumstances. These findings further underscore the complex duality of P. copri’s metabolic effects, emphasizing the interplay between bacterial dosage, dietary context, and host metabolic outcomes, which align with its reported beneficial or detrimental actions in host health19,20,21,40,41,42.
Subsequent investigations established that the P. copri-derived succinate- induced enhancement of iWAT browning exhibited at least partial dependence on SUCNR1 (also termed GPR91). This succinate receptor demonstrates expression in WAT, in both white adipocytes and WAT-associated macrophages49,50. Our data revealed that the SUCNR1 in macrophages primarily mediates P. copri or succinate promotion of iWAT browning. One previous study showed that adipocyte-specific deletion of Sucnr1 inhibits WAT deposition in the context of a chow diet but, conversely, induces obesity and hepatic steatosis under HFD conditions51. McCreath et al.52 revealed dichotomous effects of complete Sucnr1-KO on obesity, showing protective effects against HFD-induced obesity during the initial period, but not at later stages. Collectively, these historical observations, in conjunction with our present findings, underscore the multifaceted regulatory role of SUCNR1 in lipid homeostasis, which depends largely on genetic manipulation strategies, dietary compositions, and animal’s developmental stages.
The role of succinate-SUCNR1 signaling in mediating P. copri’s metabolic effects motivated our systematic investigation of downstream mediators. Existing literature documents that SUCNR1 activation by accumulated succinate enhances proinflammatory cytokine production across multiple tissues45,53. We discovered systemic and tissue-specific IL-6 elevation, and expanded F4/80+/CD11b+ macrophage population in iWAT. Il-6-KO models and macrophage-depleted (CL-treated) mice further confirmed macrophage- derived IL-6 as the principal effector. Notably, P. copri or SS also triggered mild inflammation activation in eWATs, but that increment was too weak to increase thermogenic gene expression (Supplementary Fig. 18). However, in livers, lower inflammation was found in the P. copri and succinate exposure group according to the flow cytometry and RT-qPCR results (Supplementary Fig. 19).
These tissue-specific reactions further demonstrate the complexity of the P. copri or succinate roles in host metabolism, which need intensive investigation in future studies. As a pleiotropic cytokine with both pro- and anti-inflammatory properties, the role of IL-6 in obesity and metabolic diseases remains controversial54,55. IL-6 has been shown to trigger inflammation in adipose tissue, consequently antagonizing insulin activity and promoting metabolic abnormalities54, whereas IL-6 and IL-6Rα could trigger alternative activation of macrophages and improve glucose homeostasis in mice with diet-induced obesity55. In terms of thermogenesis, Il-6 gene delivery has been demonstrated to induce UCP1 expression in BAT as one possible mechanism leading to body weight reduction in rats56. Phosphorylated STAT3 could bind to specific DNA response elements in the target gene promoter to modulate its transcription54,56. We previously confirmed that p-STAT3Tyr705 functions as a TF to induce Ucp1 expression in iWAT28. Surprisingly, herein we also found P. copri or succinate exposure enhanced p-STAT3Tyr705 interaction with the Tbx1 promoter. These results establish the macrophage IL-6-STAT3 axis as the core pathway for P. copri- or succinate-induced iWAT browning.
Strikingly, our current results demonstrate that FOXM1 acts downstream of SUCNR1 and directly binds to the Il-6 promoter at two different potential binding sites. As an important Fox family protein, FOXM1 has been shown to be involved in the migration and functional differentiation of immune cells to regulate several physiological and pathological processes36,57,58. For example, macrophage-specific deletion of Foxm1 leads to decreased expression of proinflammatory genes, such as iNOS, Cox-2, Il-1β, and Il-636. Decreased FOXM1 expression in a mouse model of diabetes inhibits neutrophil and macrophage recruitment, resulting in delayed wound healing57. This study thus represents a report of FOXM1 TF function in directly activating Il-6 expression, a key step in P. copri- or succinate-induced thermogenesis in iWAT.
In summary, the present investigation reveals that decreased abundance of P. copri in patients with obesity is strongly linked to lower TBX1 and UCP1 expression levels in WAT. P. copri exposure effectively promotes iWAT browning and counteracts adiposity via its metabolite, succinate, in a manner largely dependent on activation of the macrophage SUCNR1-FOXM1-IL-6- STAT3 axis (Fig. 8). These findings expand the scope of our understanding of the mechanisms of host-P. copri interplay, and suggest that P. copri and succinate may serve as promising interventions to combat diet-related obesity and associated disorders.
People with obesity have aberrantly low abundance of P. copri in their fecal microbiomes. In HFD-fed mice, oral P. copri administration increased iWAT browning and attenuated weight gain via its metabolite, succinate, in a manner largely dependent on activation of the macrophage SUCNR1-FOXM1-IL-6- STAT3 axis. The macrophage-derived IL-6-STAT3 pathway, triggered by FOXM1-mediated Il-6 transcription, was essential for this P. copri- or succinate-inducible iWAT browning (Created by figdraw.com. Shuang Liu. (2025) Copyright Code: TTSUP444f8).
Methods
Human cohort analysis
Abdominal subcutaneous WAT was collected from patients who underwent bariatric surgery or cholecystectomy, and stored at −80 °C for detection of SUCNR1 and thermogenic gene expression. The BMI, body fat index, and visceral fat area of the participants were tested by a body composition analyzer (InBody770, Seoul, Korea). The waist circumference was measured by using a soft measuring tape.
For microbial metagenomic analysis, and fecal and plasma succinate content examination, 15 individuals without obesity (BMI < 28 kg/m2, 4 males and 11 females) and 19 individuals with obesity (BMI ≥ 28 kg/m2, 6 males and 13 females) were recruited (Supplementary Table 1). A total of 92 participants were recruited to quantify the relative abundance of fecal P. copri using qPCR (Supplementary Data 1), and 40 individuals participate in measuring the relationship between P. copri abundance and WAT UCP1/TBX1 mRNA (Supplementary Table 2).
Metagenomic sequencing of fecal microbiomes and data analysis
Total genomic DNA was extracted from 0.2 g of stool material via the PF Mag-Bind Stool DNA Kit (Omega Biotek, Norcross, GA, USA) following the manufacturer’s instructions. The DNA was fragmented to approximately 350 bp using Covaris M220 (Gene Company Limited, China) for paired-end library construction with NEXTFLEX Rapid DNA-Seq (Bioo Scientific, Austin, TX, USA). Paired-end sequencing was performed on an Illumina NovaSeq 6000 (Illumina Inc., San Diego, CA, USA) at Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China) via a NovaSeq X Series 25B Reagent Kit according to the manufacturer’s instructions (www.illumina.com). The data were analyzed on the free online platform of the Majorbio Cloud Platform (www.majorbio.com).
P. copri cultivation and examination of organic acids in the bacterial culture supernatant
P. copri (strain DSM18205) from Deutsche Sammlung von Mikroorganismen und Zellkulturen (Braunschweig, Germany) was cultivated in KVLB (kanamycin-vancomycin laked blood) medium under anaerobic conditions at 37 °C. P. copri cultivated for 48 h was harvested in log phase, centrifuged at 1000 × g for 10 min, and washed twice with PBS. The precipitate was resuspended with PBS to a concentration of 107 CFU/mL.
To detect the potential metabolites of P. copri, targeted metabolomics was carried out by using the bacterial culture supernatant (Sci-tech innovation quality inspection Co., Ltd, Qingdao, China). Briefly, 200 μL bacterial culture supernatant was mixed with 200 μL of ice-cold methanol and put under vortex movement for 1 min. The mixture was sonicated for 30 min and then incubated at 60 °C for 60 min, which was then centrifuged at 12,000 × g at 4 °C for 20 min. The supernatants were collected and analyzed via an Agilent 1260 HPLC system coupled to an Agilent 6120 mass spectrometer (California, USA). Chromatographic separation was performed at 30 °C with an Agilent AQ column (4.6 × 250 mm, 5 μm).
Animal experiments
The animals used in this study were C57BL/6 J mice. Il-6-KO mice were purchased from the Jackson Laboratory (Bar Harbor, ME, USA). Sucnr1-KO mice30 were a gift from Professor Gang Shu of South China Agricultural University, China. The mice were housed in a controlled environment at Shandong Agricultural University and maintained under a standard 12:12 h light cycle with temperature kept at 21 °C–24 °C, and humidity at 40–70%. The animals had ad libitum access to feed and water and were fed a standard chow diet or a HFD with 60% fat energy (D12492, Research Diets, New Jersey, USA). P. copri (live and 65 °C pasteurized bacteria at a dosage of 108 CFU/kg BW) and SS (0.1 or 0.25 g/kg BW) were administered intragastrically every two days during the trial periods, whereas normal saline was used as a control. The BW and feed intake of the mice were recorded weekly. Finally, after fasting for 12 h, blood was collected, and serum was prepared. The iWAT, iBAT, eWAT, and liver were harvested and weighed.
qPCR quantification of P. copri in human feces and mouse cecal contents and feces
Fecal or cecal DNA was extracted via a stool DNA extraction kit (Tiangen Biochemical Technology Co., Ltd., Beijing, China). The primers used for PCR-based amplification were as follows: Forward-TCGCTGACATGAGC GATAAC, Reverse-CCGTTGGCACTACCTTCATT, as previously described22. Quantitative PCR was performed via ChamQ SYBR Color qPCR Master Mix (Vazyme Biotech Co., Ltd., Nanjing, China) on an ABI Prism 7500 PCR system (Applied Biosystems, USA) at 95 °C for 1 min, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. A standard curve was constructed with various bacterial counts as described in the literature22.
Measurement of fecal, cecal and plasma succinate levels
Feces or ceal contents were mixed with extraction solvent containing 1 μM chlorpropamide, which was further dissolved in ice-cold 80% methanol (v/v) and homogenized with a KZ-II homogenizer. Plasma was mixed with ice-cold methanol and put under vortex movement for 1 min. The mixture was sonicated for 30 min and then incubated at 60 °C for 60 min, which was then centrifuged at 12,000 × g at 4 °C for 20 min. The measurement of succinate was performed via a liquid chromatography‒tandem mass spectrometry system composed of a HPLC system (Kyoto, Japan) coupled to a Sciex 4500 triple quadrupole linear ion trap mass spectrometer (AB SCIEX, Framingham, MA, USA). The liquid chromatography system was equipped with a XDB-C18 column (4.6 × 250 mm, 5 µm). The mobile phase comprised A (0.1% phosphoric acid in pure water) and B (acetonitrile) with a flow rate of 1 mL/min.
Body composition analysis of mice
The body composition of the mice was scanned and imaged via dual-energy X-ray absorptiometry (DXA) (MEDIKORS, Korea), the fat and lean indices were calculated via the system’s software. Each mouse was scanned and analyzed at least three times.
Glucose Tolerance Test (GTT) and Insulin Tolerance Test (ITT)
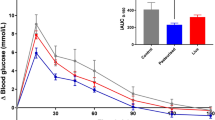
For GTT, mice in different groups were fasted for 12 h, and the baseline blood glucose was measured via tail vein by using a glucometer (time=0). Then the mice were intragastrically administered with glucose (2.0 g/kg BW), and the blood glucose concentrations were measured at 15, 30, 60, 90, and 120 min post-injection.
For ITT, mice were fasted for 6 h and the baseline blood glucose was measured (time=0). After injecting with insulin (0.75 U/kg BW) intraperitoneally, the blood glucose concentrations were monitored at 30, 60, 90, and 120 min.
Respiratory metabolic assay
Mice were individually placed in a Comprehensive Lab Animal Monitoring System (Coulumbus Instruments, USA) with 12 h of light and 12 h of darkness. The animals were allowed to eat and drink freely and were allowed to adapt for 24 h before formal testing. Data were collected every 5 min for another 24 h. O2 consumption (VO2), CO2 production (VCO2), and heat production were measured by the system, and the RER were calculated by VCO2 /VO2. The data were analyzed via ANCOVA (SPSS statistics 26.0).
Rectal temperature measurement
A digital thermometer equipped with a copper thermocouple probe was used to measure the rectal temperature. The probe was inserted 1.5 cm into the anal ducts of the mice.
Determination of serum IL-6 and TNF-α levels
Whole blood from the eyeballs of the mice was collected, put aside for 30 min, and subsequently centrifuged at 2000 g for 10 min to isolate the serum. The serum levels of IL-6 and TNF-α were quantified via commercial ELISA kits (Mlbio, Shanghai, China).
Measurement of serum indices
The serum ALT and AST activities, as well as the triglyceride (TG) and total cholesterol (TCH) contents, were assessed via commercial kits (Nanjing Jiancheng Bioengineering Institute, China).
Hematoxylin‒eosin (H&E) staining and immunofluorescence staining of UCP1
The tissue samples were fixed in 4% formaldehyde solution for more than 24 h. Subsequently, the tissues were embedded in paraffin and dissected into 5 μm sections for H&E staining.
For immunofluorescence staining, the dewaxed iWAT sections were treated with Triton X-100 and blocked in PBS supplemented with 5% fetal bovine serum (FBS). The sections were subsequently incubated overnight at 4 °C with a 1:100 diluted UCP1 antibody (sc-6529, Sacta Cruz, USA), followed by incubation with a fluorescent-labeled secondary antibody (1:500) and DAPI (1 μg/mL). Images were acquired via a confocal microscope (LSM880, Zeiss, Germany).
Isolation, cultivation, and differentiation of iWATDCs
Adipose tissue from the mouse inguinal region was digested with collagenase I (Solarbio Science & Technology Co., Ltd., Beijing, China) to isolate iWATDCs, a mixed cell type that containing adherent mesenchymal stem cells, macrophages, and a small quantity of fibroblasts, following previously established protocols59. The isolated cells were cultured to subconfluence in low-glucose Dulbecco’s modified Eagle’s medium (DMEM) (HyClone, USA) supplemented with 15% FBS (Tianhang Biotechnology Co., Ltd., Hangzhou, China) and 100 units/mL penicillin/100 μg/mL streptomycin (Solarbio Science & Technology Co., Ltd., Beijing, China). The sub-confluent iWATDCs were induced to differentiate into adipocytes as previously described28. SS (Macklin Biochemical Technology Co., Ltd., Shanghai, China) and P. copri culture supernatant, in the absence or presence of 0.1 μM 4c (Wuhe Biotechnology Co., Ltd., Shenyang, China), were added during the differentiation period.
Oil red O (ORO) staining of iWATDC-differentiated adipocytes
For ORO staining, iWATDC-differentiated adipocytes were rinsed with PBS and fixed in a 4% formaldehyde solution for 1 h. The cells were subsequently stained with a 0.6% ORO solution for 40 min, followed by hematoxylin staining for 1 min to facilitate the observation of cell morphology.
SeaHorse assay
Oxygen consumption rates (OCRs) were assessed using an Agilent Seahorse XFe24 Analyser (California, USA). iWATDCs were prepared and seeded in XF24 V28 cell culture microplates (Agilent, 100882-004) for 6 days’ adipogenic induction with AS in the presence or absence of P. copri sup or SS (0.1 mM). XFe24 sensor cartridges were hydrated according to the manufacturer’s protocol. Oligomycin, FCCP, and rotenone + antimycin A (R/A) (Agilent, 103015-100) were used at final concentrations of 1.5 µM, 1 µM, and 0.5 µM, respectively. The final OCR results were standardized to total cell number in each well.
Flow cytometry
iWATDCs were collected from iWAT, hepatic cells were were collected from Liver, rinsed twice and resuspended in PBS supplemented with 2% FBS to achieve a concentration of 1×106 ~ 107 cells/mL. The cells were subsequently stained with antibodies against specific cell surface markers, including PerCP-Cyanine 5.5-conjugated CD45.2 (45-0454-82, Thermo Fisher Scientific, USA), phycoerythrin (PE)-conjugated F4/80 (12-4801-82, Thermo Fisher Scientific, USA), APC-conjugated CD11b (17-0112-82, Thermo Fisher Scientific, USA), and eFluor™ 450-conjugated CD86 (48-0862-82, Thermo Fisher Scientific, USA). The staining process was carried out in the dark at 4 °C for 30 min. Flow cytometry analysis was performed using a BD FACSCanto™ II instrument (BD Biosciences, New Jersey, USA). Data analysis was performed via FlowJo software (Tree Star, Inc., Ashland, USA).
Cultivation and treatments of RAW264.7 cells
RAW264.7 cells were cultured in high-glucose DMEM supplemented with 10% FBS, along with 100 units/mL penicillin and 100 μg/mL streptomycin. The cells were maintained at 37 °C in a humidified atmosphere containing 5% CO2. The sub-confluent cells were treated with SS and P. copri culture supernatant for 4 days, respectively.
Cultivation and adipogenic induction of 3T3L1 cell lines
3T3-L1 cells were cultured in high-glucose DMEM supplemented with 10% calf serum (Tianhang Biotechnology Co., Ltd., Hangzhou, China), along with 100 units/mL penicillin and 100 μg/mL streptomycin. The cells were maintained at 37 °C in a humidified atmosphere containing 5% CO2.
For the adipogenic induction of 3T3-L1 differentiation, the cells were treated with differentiation-inducing medium containing 10 µg/mL insulin, 1 µM DEX, and 0.5 mM IBMX for 2 days. The induction medium was subsequently replaced with maintenance medium containing 10 µg/mL insulin for 2 days, as previously described60.The induction hormones were subsequently removed, and the cells were cultured in standard growth medium until they reached full differentiation into mature adipocytes.
Macrophage depletion through intravenous injection of clodronate liposomes (CL)
Macrophage populations were selectively ablated through tail vein injection of clodronate liposomes (CL), an established macrophage-depleting agent validated in prior studies34,35. In brief, CL (Yeasen Biotechnology, Shanghai, China) were injected into mice by tail vein twice a week (200 µL/time). The ablation efficiency was tested by flow cytometry measuring the proportion of F4/80+/Cd11b+ cells in iWAT and blood. After validation of about 95% macrophages were ablated, cohorts of CL-treated mice received either P. copri (10⁸ CFU/kg BW) or SS (0.25 g/kg BW) for another 8 weeks under HFD conditions.
RNA extraction and real-time quantitative PCR (RT-qPCR)
RNA extraction from both cell and tissue samples was carried out via RNAiso Plus Reagent (Takara, Japan). The extracted RNA was subsequently reverse transcribed into cDNA via a HiScript III All-in-one RT SuperMix Perfect for qPCR Kit (Vazyme Biotech Co., Ltd., Nanjing, China). RT-qPCR was performed via ChamQ Universal SYBR qPCR Master Mix (Vazyme Biotech Co., Ltd., Nanjing, China) on an ABI Prism 7500 PCR system (Applied Biosystems, USA), with an initial denaturation step at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 15 s and annealing/extension at 60 °C for 30 s. The primer sequences utilized are detailed in Supplementary Data 2. Melting curve analysis was employed to validate the specificity of the primers. The relative mRNA expression levels of target genes were quantified via the 2−ΔΔCt method and normalized to the expression level of the housekeeping gene Rpl13α in mice samples and gene 18S rRNA in human samples.
Protein extraction and western blotting
Protein was isolated from cells or tissues using RIPA lysis buffer supplemented with phosphatase and protease inhibitors (Roche, USA). Protein samples were separated by SDS‒PAGE and then transferred onto polyvinylidene fluoride (PVDF) membranes, which were incubated at room temperature for 15 min with TBST containing 5% non-fat milk to block nonspecific binding. The membranes were subsequently subjected to overnight incubation at 4 °C with specific primary antibodies, followed by incubation with corresponding HRP-conjugated secondary antibodies. The primary antibodies used in this study included UCP1 (sc-293418, Santa Cruz, USA), TBX1 (bs-21501R, Bioss, China), PPARγ (sc-7273, Santa Cruz, USA), AP2 (sc-271529, Santa Cruz, USA), SUCNR1 (AY4281, Abways, China), IL-6 (12912S, CST, USA), p-STAT3Tyr705 (9145S, CST, USA), STAT3 (4904S, CST, USA), FOXM1 (sc-376471x, Santa Cruz, USA), β-actin (AB0035, Abways, China), and GAPDH (AB0038, Abways, China). The protein bands were visualized through chemiluminescence via an enhanced chemiluminescence (ECL) kit (Beyotime, China).
Transcriptome sequencing and data analysis
iWATs from control and SS-treated mice were collected for transcriptome sequencing. Total RNA was extracted via TRIzol reagent (Thermo Fisher Scientific, USA), and 4 µg of RNA was used to construct biologically triplicate sequencing libraries. Amplified libraries were sequenced on an Illumina HiSeq 2000 platform. Fastp (0.22.0) software was used to filter the sequencing data to get high quality sequence (Clean Data). The reference genome and gene annotation files were downloaded from genome website. The filtered reads were mapping to the reference genome using HISAT2 (v2.1.0). HTSeg (v0.9.1) statistics was used to compare the Read Count values on each gene as the original expression of the gene, and then Fragments Per Kilo bases per Million fragments (FPKM) values were used to standardize the expression. Then difference expression of genes was analyzed by edgeR with screened conditions as follows: expression difference multiple |log2FoldChange | >1, significant P value < 0.05.
Lentivirus packaging of shRNA and lentivirus infection
HEK293T cells were cultured in high-glucose DMEM supplemented with 10% FBS in a humidified atmosphere containing 5% CO2 at 37 °C. Lentivirus production was carried out by using HEK293T cells. Briefly, HEK293T cells were transfected with a PSIH1-H1-CopGFP vector encoding shRNAs against Foxm1 (shRNA sequences are listed in Supplementary Table 3), along with the packaging plasmids pMD2. G, pMDL, and pRSV-Rev. The culture medium was replaced after 6 h, and the viral particles were harvested at 48 and 72 h post-transfection, respectively. iWATDCs were infected with lentivirus at 5 MOI in the presence of 8 µg/mL polybrene (Beyotime, China).
Chromatin immunoprecipitation (ChIP)
Sub-confluent iWATDCs or RAW264.7 cells were treated with P. copri supernatant and SS for 3 days, respectively. 1% (v/v) formaldehyde was used to cross-link the DNA and proteins. Subsequently, the chromatin DNA was sonicated into shorter pieces (200 ~ 1,000 bp) by using a Bioruptor Plus (Diagenode, Denville, NJ, USA) over 20 cycles of 30 s on and 30 s off. The DNA‒protein complexes were treated with protein A/G beads blocked with bovine serum albumin, followed by overnight incubation with specific antibodies for p-STAT3 or FOXM1, or control IgG. The cross-linked complexes were released by treatment with 5 M NaCl at 65 °C for 4 h. DNA was purified via a phenol‒chloroform extraction process. The eluted DNA was used as a template for qPCR. The input control was the supernatant collected before immunoprecipitation. The primer sequences utilized to amplify the Ucp1, Tbx1, and Il-6 promoter sequences are listed in Supplementary Data 2.
Dual luciferase assay
For the luciferase reporter assay, pGL3-basic luciferase reporter plasmids and the internal control renilla luciferase control vector were transfected into HEK293T cells via Lipofectamine 3000 (Thermo Fisher Scientific, USA). A Mut Express® II Fast Mutagenesis Kit V2 (Vazyme Biotech Co., Ltd., Nanjing, China) was used to construct mutated plasmids within the Il-6 promoter deleted of BS1, BS2, or the both regions. The primer sequences used for homologous recombination are provided in Supplementary Table 4. HEK293T cells were cultured in 96-well plates and grown to 70% confluence prior to transfection. The FOXM1 expression plasmid and different pGL3-basic-Il-6 promoter-luciferase reporter vectors were cotransfected into the cells for 24 h. The firefly and renilla luciferase activities of the cells were subsequently measured via a Duo-LiteTM Luciferase Assay System (Vazyme Biotech Co., Ltd., Nanjing, China), following the manufacturer’s instructions. The activity of the pGL3-basic-Il-6 promoter-luciferase reporter, normalized to that of the pRL-TK Rluc reporter, was compared between cells transfected with the FOXM1 expression plasmid or the control vector.
Statistical analysis
All data are expressed as mean ± SEM. Statistical analyses was performed via SPSS statistics 26.0 (SPSS, USA). Data analyses, such as bar charts and line charts were performed via GraphPad Prism 10.1.2 (GraphPad, USA). Parametric or non-parametric statistical tests were appropriately applied after Shapiro-Wilk test for the normal data distribution. For comparison between the two groups, an unpaired two-tailed t-test was used when samples were normally distributed, and the two-tailed Mann-Whitney U test was used when samples were not normally distributed. For comparisons among more than two groups, one-way ANOVA, followed by LSD (with equal variances) or Dunnett’s T3 (with different variances) post hoc test was used when samples were normally distributed, and Kruskal-Wallis test followed by Dunn’s post hoc test was used when samples were not normally distributed. All non-animal experiments were repeated independently three times with similar results. At least 3 independent samples were included in each experiment. Significance was established for P < 0.05, n.s indicates non-significant.
Ethics statement
For the human cohort, all participants were recruited from the Third People’s Hospital of Chengdu (Chengdu, China) and provided informed consent, which included a detailed explanation of the study’s purpose, procedures, and potential risks and benefits. We have obtained consent from all participants for the publication of identifiable information and the use of their data in the analysis and publication of the research findings. The study was approved by the Institutional Ethics Review Board of the Third People’s Hospital of Chengdu (Approval No: S-62 [2022]) and was carried out following the Chinese ethical guidelines for human genome/gene research.
For the animal experiments, ethical approval for all experimental procedures was obtained from the Animal Care and Use Committee of Shandong Agricultural University (Approval No: SDAU-2021-129).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The metagenomic sequencing data generated in this study were deposited in the NCBI SRA database (accession code PRJNA1189811). The transcriptome sequencing data generated in this study were deposited in the NCBI SRA database (accession code PRJNA1189824). All data supporting the article are included in the main article, Supplementary Information, Supplementary Data 1, Supplementary Data 2, and the Source Data file. Source data are provided with this paper. The Source data file has been deposited in the Figshare repository [https://doi.org/10.6084/m9.figshare.30675941].
References
Afshin, A. et al. Health effects of overweight and obesity in 195 countries over 25 years. N. Engl. J. Med. 377, 13–27 (2017).
Frontini, A. & Cinti, S. Distribution and development of brown adipocytes in the murine and human adipose organ. Cell Metab. 11, 253–256 (2010).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012).
Rosenwald, M., Perdikari, A., Rülicke, T. & Wolfrum, C. Bi-directional interconversion of brite and white adipocytes. Nat. Cell Biol. 15, 659–667 (2013).
Henriques, F. et al. Single-cell RNA profiling reveals adipocyte to macrophage signaling sufficient to enhance thermogenesis. Cell Rep. 32, 107998 (2020).
van Marken Lichtenbelt, W. D. et al. Cold-activated brown adipose tissue in healthy men. N. Engl. J. Med. 360, 1500–1508 (2009).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Min, S. Y. et al. Human ‘brite/beige’ adipocytes develop from capillary networks, and their implantation improves metabolic homeostasis in mice. Nat. Med. 22, 312–318 (2016).
de Vos, W. M., Tilg, H., Van Hul, M. & Cani, P. D. Gut microbiome and health: mechanistic insights. Gut 71, 1020–1032 (2022).
Chevalier, C. et al. Gut microbiota orchestrates energy homeostasis during cold. Cell 163, 1360–1374 (2015).
Worthmann, A. et al. Cold-induced conversion of cholesterol to bile acids in mice shapes the gut microbiome and promotes adaptive thermogenesis. Nat. Med. 23, 839–849 (2017).
Wu, Q. et al. Intestinal hypoxia-inducible factor 2α regulates lactate levels to shape the gut microbiome and alter thermogenesis. Cell Metab. 33, 1988–2003 (2021).
Liu, Y. et al. Limosilactobacillus reuteri and caffeoylquinic acid synergistically promote adipose browning and ameliorate obesity-associated disorders. Microbiome 10, 226 (2022).
Zhang, C. et al. Lactobacillus reuteri J1 prevents obesity by altering the gut microbiota and regulating bile acid metabolism in obese mice. Food Funct. 13, 6688–6701 (2022).
Tett, A., Pasolli, E., Masetti, G., Ercolini, D. & Segata, N. Prevotella diversity,niches and interactions with the human host. Nat. Rev. Microbiol. 19, 585–599 (2021).
Asnicar, F., Berry, S. E. & Valdes, A. M. Microbiome connections with host metabolism and habitual diet from 1098 deeply phenotyped individuals. Nat. Med. 27, 321–332 (2021).
Gong, J. et al. Effects of Prevotella copri on insulin, gut microbiota and bile acids. Gut Microbes 16, 2340487 (2024).
Hu, J. et al. Gut microbiota signature of obese adults across different classifications. Diab. Metab. Syndr. Obes. 15, 3933–3947 (2022).
Chen, C. et al. Prevotella copri increases fat accumulation in pigs fed with formula diets. Microbiome 9, 175 (2021).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metab. 22, 971–982 (2015).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535, 376–381 (2016).
Verbrugghe, P., Van Aken, O., Hållenius, F. & Nilsson, A. Development of a real-time quantitative pcr method for detection and quantification of prevotella copri. BMC Microbiol. 21, 23 (2021).
Kim, S. N. et al. Sex differences in sympathetic innervation and browning of white adipose tissue of mice. Biol. Sex. Differ. 7, 67 (2016).
Moreira-Pais, A. et al. Sex differences on adipose tissue remodeling: from molecular mechanisms to therapeutic interventions. J. Mol. Med. (Berl.). 98, 483–493 (2020).
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44 (2014).
Wang, J., Zhang, Y., Shen, Q., Wu, J. & Li, J. X. Oleanolic acid derivative HA-20 inhibits adipogenesis in a manner involving PPARγ-FABP4/aP2 pathway. J. Mol. Endocrinol. 66, 245–258 (2021).
Steensels, S. & Ersoy, B. A. Fatty acid activation in thermogenic adipose tissue. Biochim Biophys Acta Mol. Cell. Biol. Lipids 1864, 79–90 (2019).
Li, H. F. et al. Peripheral IL-6/STAT3 signaling promotes beiging of white fat. BBA-Mol. Cell Res. 1868, 119080 (2021).
de Castro Fonseca, M., Aguiar, C. J., da Rocha Franco, J. A., Gingold, R. N. & Leite, M. F. GPR91: expanding the frontiers of Krebs cycle intermediates. Cell Commun. Signal. 14, 3 (2016).
Wang, T. et al. Succinate induces skeletal muscle fiber remodeling via SUNCR1 signaling. EMBO Rep. 21, e47892 (2019).
Chen, M. et al. HCK induces macrophage activation to promote renal inflammation and fibrosis via suppression of autophagy. Nat. Commun. 14, 4297 (2023).
Oh, C. M. et al. Regulation of systemic energy homeostasis by serotonin in adipose tissues. Nat. Commun. 6, 6794 (2015).
Huang, Y. et al. UCHL1 promoted polarization of M1 macrophages by regulating the PI3K/AKT signaling pathway. J. Inflamm. Res. 5, 735–746 (2022).
Dong, B. et al. Vitamin D receptor activation in liver macrophages ameliorates hepatic inflammation, steatosis, and insulin resistance in mice. Hepatology 71, 1559–1574 (2020).
Xie, M. et al. FGF19/FGFR4-mediated elevation of ETV4 facilitates hepatocellular carcinoma metastasis by upregulating PD-L1 and CCL2. J. Hepatol. 79, 109–125 (2023).
Balli, D. et al. Foxm1 transcription factor is required for macrophage migration during lung inflammation and tumor formation. Oncogene 31, 3875–3888 (2012).
Hu, G. et al. FOXM1 promotes hepatocellular carcinoma progression by regulating KIF4A expression. J. Exp. Clin. Cancer Res. 38, 188 (2019).
De Filippis, F. et al. Distinct genetic and functional traits of human intestinal prevotella copri strains are associated with different habitual diets. Cell Host Microbe 25, 444–453 (2019).
Chung, W. S. F. et al. Relative abundance of the prevotella genus within the human gut microbiota of elderly volunteers determines the inter-individual responses to dietary supplementation with wheat bran arabinoxylan-oligosaccharides. BMC Microbiol. 20, 283 (2020).
Scher, J. U. et al. Expansion of intestinal prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2, e1202 (2013).
Tett, A. et al. The prevotella copri complex comprises four distinct clades underrepresented in westernized populations. Cell Host Microbe 26, 666–679 (2019).
Fernández-Veledo, S., Ceperuelo-Mallafré, V. & Vendrell, J. Rethinking succinate: an unexpected hormone-like metabolite in energy homeostasis. Trends Endocrin. Metab. 32, 680–692 (2021).
De Vadder, F. et al. Microbiota-produced succinate improves glucose homeostasis via intestinal gluconeogenesis. Cell Metab. 24, 151–157 (2016).
Mills, E. L. et al. Accumulation of succinate controls activation of adipose tissue thermogenesis. Nature 560, 102–106 (2018).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 496, 238 (2013).
Mills, E. L. et al. Ucp1 governs liver extracellular succinate and inflammatory pathogenesis. Nat. Metab. 3, 604–617 (2021).
Sadagopan, N. et al. Circulating succinate is elevated in rodent models of hypertension and metabolic disease. Am. J. Hypertens. 20, 1209–1215 (2007).
Fernández-Veledo, S., Marsal-Beltran, A. & Vendrell, J. Type 2 diabetes and succinate: unmasking an age-old molecule. Diabetologia 67, 430–442 (2024).
He, W. et al. Citric acid cycle intermediates as ligands for orphan G-protein-coupled receptors. Nature 429, 188–193 (2004).
Regard, J. B., Sato, I. T. & Coughlin, S. R. Anatomical profiling of g protein-coupled receptor expression. Cell 135, 561–571 (2008).
Villanueva-Carmona, T. et al. Sucnr1 signaling in adipocytes controls energy metabolism by modulating circadian clock and leptin expression. Cell Metab. 35, 601–619 (2023).
McCreath, K. J. et al. Targeted disruption of the sucnr1 metabolic receptor leads to dichotomous effects on obesity. Diabetes 64, 1154–1167 (2015).
Rubic, T. et al. Triggering the succinate receptor GPR91 on dendritic cells enhances immunity. Nat. Immunol. 9, 1261–1269 (2008).
Theurich, S. et al. IL-6/Stat3-dependent induction of a distinct, obesity-associated NK cell subpopulation deteriorates energy and glucose homeostasis. Cell Metab. 26, 171–184 (2017).
Mauer, J. et al. Signaling by il-6 promotes alternative activation of macrophages to limit endotoxemia and obesity-associated resistance to insulin. Nat. Immunol. 15, 423–430 (2014).
Li, G. et al. Induction of uncoupling protein 1 by central interleukin-6 gene delivery is dependent on sympathetic innervation of brown adipose tissue and underlies one mechanism of body weight reduction in rats. Neuroscience 115, 879–889 (2002).
Sawaya, A. P. et al. Deregulated immune cell recruitment orchestrated by FOXM1 impairs human diabetic wound healing. Nat. Commun. 11, 4678 (2020).
Yang, Y., Zhang, B., Yang, Y., Peng, B. & Ye, R. FOXM1 accelerates wound healing in diabetic foot ulcer by inducing M2 macrophage polarization through a mechanism involving SEMA3C/NRP2/hedgehog signaling. Diab. Res. Clin. Pract. 184, 109121 (2022).
Seale, P. et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. J. Clin. Invest. 121, 105–196 (2011).
Vishwanath, D. et al. Novel method to differentiate 3T3 L1 cells in vitro to produce highly sensitive adipocytes for a GLUT4 mediated glucose uptake using fluorescent glucose analog. J. Cell Commun. Signal. 7, 129–140 (2013).
Acknowledgements
We thank Gang Shu for gifting the Sucnr1-KO mice, Lifen Gao for gifting the RAW264.7 cell line, Wencheng Zhang and Jingyuan Li for assistance with respiratory metabolic assays, Wenchu Qian, Tiantian Zeng, and Yizhuo Chen for organizing reference materials. This work was supported by the National Natural Science Foundation of China (No. 32330101, 82202007, 82170887, H.L.), the National Key Research and Development Program of China (No. 2021YFD1300405, H.L.), the Natural Science Foundation of Shandong Province, China (No. ZR2023MC085, H.L., H.F.L), and the Third People’s Hospital of Chengdu Clinical Research Program (CSY-YN-01-2023-011, 076, 077, H.F.L). The graphical abstract was created with Figdraw.com (Copyright Certificate ID: TTSUP444f8).
Author information
Authors and Affiliations
Contributions
Conceptualization: H.F.L., H.L., Z.H.F., J.B.X., and S.L.; data curation: S.L. and H.F.L.; formal analysis: S.L., H.F.L., R.L, Y.X.C., S.L.W., Z.G.W., Y.W.S., and M.Y.L.; funding acquisition: H.L., H.F.L., J.B.X., and Z.H.F.; investigation and resources: M.L., Y.L.Z., X.J.W., H.C.J., J.P.Z., B.L., and T.T.Z.; supervision: H.F.L., H.L., J.B.X., and Z.H.F.; writing – original draft: S.L., and H.F.L.; writing – review & editing, all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Benjamin Jensen, Mohammed Hankir and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, S., Li, H., Li, R. et al. Prevotella copri promotes white adipose browning and ameliorates adiposity. Nat Commun 17, 680 (2026). https://doi.org/10.1038/s41467-025-67327-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67327-x