Abstract
Telomeres are protective DNA caps at chromosome ends that prevent cells from mistakenly recognizing them as broken DNA. These structures are safeguarded by a protein complex called Shelterin, particularly through the TRF2 protein encoded by Trf2. Surprisingly, in mouse embryonic stem cells, TRF2 is not essential for telomere protection, suggesting that other mechanisms compensate for its loss. Here we show that a cellular quality control system called nonsense-mediated mRNA decay (NMD), which normally eliminates defective RNA molecules, plays an unexpected role in maintaining telomere integrity in pluripotent cells. Through a genome-wide genetic screen, we discovered that NMD is essential for cell survival when TRF2 is absent. NMD accomplishes this by degrading an aberrant form of the messenger RNA encoded by Trf1, which produces the TRF1 protein, another Shelterin component. Without NMD, this aberrant RNA produces a truncated, harmful version of TRF1 that interferes with normal telomere protection. Our findings reveal that embryonic stem cells use a unique strategy for chromosome end protection, linking RNA quality control to genome stability in a previously unrecognized way.
Similar content being viewed by others
Main
Telomeres are nucleoprotein structures that cap the ends of linear chromosomes, protecting them from being recognized as DNA double-strand breaks1,2,3,4. In mammalian cells, telomeres consist of tandem repeats of the TTAGGG sequence bound by the Shelterin complex, which is essential for maintaining telomere integrity1,4,5,6,7. Within Shelterin, Trf2 (telomeric repeat-binding factor 2) plays a central role in suppressing non-homologous end joining (NHEJ)-mediated chromosome fusions1,8. In somatic cells, loss of TRF2 triggers rapid telomere dysfunction, NHEJ activation at every chromosome end, and genomic instability1,7,9. However, mouse embryonic stem (mES) cells exhibit a unique tolerance to TRF2 depletion10,11. In these cells, Trf2 deletion does not trigger a strong DNA damage response, NHEJ is not activated, and cells remain viable and can be propagated indefinitely, pointing to fundamental differences in telomere protection mechanisms between pluripotent and differentiated cells.
Although telomere protection has been mainly attributed to the Shelterin complex, emerging evidence indicates that RNA metabolism also contributes to genome stability, including at repetitive and telomere-proximal regions12,13,14,15,16. Several RNA surveillance pathways have been implicated in modulating telomeric RNA species and preventing the accumulation of aberrant RNA–DNA structures at chromosome ends, suggesting that post-transcriptional control influences telomere function16,17,18,19,20.
The nonsense-mediated mRNA decay (NMD) pathway is a conserved RNA surveillance mechanism that degrades transcripts harbouring premature termination codons (PTCs)21,22. Beyond quality control, NMD fine-tunes normal gene expression, influences lineage decisions, shapes neuronal and immune functions, and buffers cells against genotoxic stress contributing to broader post-transcriptional gene regulatory networks that influence cellular identity and genome stability15,16,23,24,25,26,27. The central effector of the pathway is UPF1 (Up-frameshift protein 1), an RNA helicase that serves as a platform for RNA decay complex assembly and activation in response to translation termination21,28. UPF1 activation requires phosphorylation, a process mediated by the SMG1 (Suppressors with morphogenetic defects in genitalia 1) kinase29,30,31,32. Once phosphorylated, UPF1 recruits additional downstream effectors: SMG5, SMG6 and SMG733,34. SMG6 exerts endonucleolytic cleavage activity on the target mRNA, while the SMG5–SMG7 complex promotes both SMG6 activity and exonucleolytic degradation through the recruitment of cellular exonucleases35,36,37. UPF1 has been shown to interact with telomere proteins, promote telomere replication and facilitate the removal of RNA–DNA hybrids (R-loops) at telomeres, contributing to genome integrity. These findings suggest that the NMD pathway may have a critical role at telomeres that extends beyond its canonical function in RNA surveillance16,17,19,20,38.
Here, we show that the NMD pathway tightly controls the expression of the telomere-associated gene Trf1. A clustered regularly interspaced short palindromic repeats (CRISPR)–Cas9 synthetic lethality screen revealed that NMD factors are essential for cell viability in the absence of TRF2. We found that the loss of TRF2 in NMD-deficient cells leads to the accumulation of DNA damage factors at telomeres, frequent formation of end-to-end chromosomal fusions and loss of cell viability. Mechanistically, NMD-depleted cells accumulate a truncated, dominant-negative Trf1 transcript in which exon 8 is skipped (TRF1ΔE8), leading to the production of a truncated, dominant-negative TRF1ΔE8 protein. TRF1ΔE8 lacks the DNA-binding MYB domain, yet retains the dimerization domain, therefore enabling displacement of the full-length TRF1 (TRF1FL) from telomeres. In our model, NMD deficiency lowers TRF1 availability at telomeres and, when combined with TRF2 loss, results in telomere dysfunction and cell death. Together, our finding uncovers a previously unrecognized post-transcriptional mechanism of telomere protection in mES cells.
Results
The NMD pathway is synthetically lethal with TRF2 in mES cells
To identify genes required for telomere protection in the absence of TRF2, we performed a genome-wide CRISPR–Cas9 synthetic lethal screen in Trf2-deficient (Trf2−/−) and Trf2-proficient (Trf2f/−) mES cells (Fig. 1a). We identified guide RNAs (gRNAs) that were selectively depleted in Trf2−/− cells (β score <−0.75) but had little or no effect in controls (β score >0). This approach recovered known telomere-associated factors, including Tpp1 and Pot1a, previously shown to be essential for proliferation in the absence of TRF210,11. In addition, we identified five components of the NMD pathway (Fig. 1a and Supplementary Table 1).
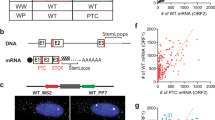
a, A genome-wide CRISPR–Cas9 screen was performed on Trf2−/− and Trf2f/− ES cells. A rank-ordered plot shows the distribution of the difference between β scores (y axis, Delta β = β Trf2−/− − β Trf2f/−) for all the genes targeted (about 23,000) (gene rank position, x axis). Genes previously been reported as synthetically lethal with Trf2 are labelled in green, and the ones identified in this study are labelled in pink. b, Western blot showing efficient depletion of SMG5, SMG6, SMG7, SMG9 and UPF1. For each gene, two independent knockout clones as well as a non-edited control (Ctrl) are shown. Note that gene editing for Upf1 results in a generation of a truncated protein leading to a hypomorph (HM) allele. Representative images from minimum two independent experiments per genotype are shown. c, Clonogenic survival assay was performed on cells of the indicated genotypes. OHT treatment induced TRF2 depletion. Cells were stained using crystal violet to count number of colonies (n = 2 biological replicates). d, Quantification of colonies shown in c: the number of colonies for each genotype was normalized to the number of colonies in the control (Ctrl) sample. For the TRF2-depleted samples (OHT) a magnification of a randomly chosen portion of the plate is shown. e, IF staining for the pluripotency marker OCT4 in ES cells of indicated genotypes, and mouse embryonic fibroblasts (MEFs), as a negative control. Representative images of three independently performed experiments are shown. Unprocessed blots and images are available in the Source data.
To validate this result, we generated knockout clones for the NMD factors Smg5-9 as well as Upf1 in Trf2Flox/Cre-ER mES cells, using different gRNAs from those in the initial screen. We confirmed complete depletion of SMG5–9 (Fig. 1b–d and Extended Data Fig. 1a,b), as well as gene editing for Upf1, which produced a truncated protein that we consider a hypomorphic allele (Upf1HM) (Fig. 1b and Extended Data Fig. 1a). Loss of individual NMD components had no effect on ES cell proliferation, in line with what was reported for Smg5-, Smg6- and Smg7-deficient ES cells23,24 (Fig. 1c,d and Extended Data Fig. 1b). Next, we tested whether depletion of TRF2 in NMD-deficient cells resulted in loss of cell viability. Colony formation and cell proliferation assays (Fig. 1c,d and Extended Data Fig. 1b) showed that codepletion of TRF2 and NMD factors results in strong growth suppression, suggesting a synthetic lethal interaction. The reduced level of proliferation was associated with increased levels of apoptosis (Extended Data Fig. 2c–e), as well as an impaired cell cycle progression of cells codepleted for TRF2 and NMD factors (Extended Data Fig. 2f,g). Importantly, all the knockouts analysed retained OCT4 expression (Fig. 1e and Extended Data Fig. 2a), indicating pluripotency maintenance and lack of differentiation.
Trf2 −/− NMD-deficient ES cells exhibit telomeric DNA damage and frequent end-to-end fusions
To determine whether the synthetic lethality between Trf2 and NMD is driven by telomere-specific defects, we asked whether telomeres are recognized as sites of DNA damage in ES cells lacking both TRF2 and NMD factors. Our data show that individual depletion of TRF2 or NMD factors did not result in the accumulation of DNA damage markers γH2AX (Fig. 2a,b) and 53BP1 (Fig. 2c,d) at telomeres. However, codepletion of TRF2 and NMD components resulted in a marked increase in the localization of both DNA damage markers at telomeres (Fig. 2a–d). These data suggest that NMD activity contributes to telomere protection in TRF2-depleted cells.
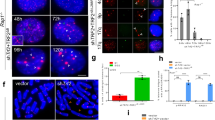
a, Representative IF–FISH showing γH2AX (red) and telomeric DNA (green) in NMD-proficient cells (Ctrl) and NMD-deficient ES cells with or without OHT treatment to induce TRF2 depletion. b, Quantification of telomere dysfunction-induced foci (TIFs) in a, defined as ≥10 γH2AX foci colocalizing with telomeres. Data are mean ± s.d. from three biological replicates. Statistical analysis (n = 3 biological replicates) by one-way ANOVA; *P = 0.0117, ****P = 0.0001. c, Representative IF–FISH images showing 53BP1 (red) and telomeric DNA (green) in the indicated genotypes ± OHT. d, Quantification of TIFs as in b using 53BP1 foci colocalization. Data are mean ± s.d. from three biological replicates. Statistical analysis (n = 3 biological replicates) by one-way ANOVA; n.s., not significant (P = 0.9153), ***P = 0.0002, ****P = 0.0001. In b–d, more than 551 cells in total per genotype were scored. For details on the exact cell number per genotype, see the Source data. e, Representative metaphases from ES cells with indicated genotypes ± OHT. Telomeric fusions per metaphase are indicated within each panel. More than 1,981 chromosomes in total per genotype were scored (n = 3 biological replicates). For details on the exact chromosome number per genotype, see the Source data. For additional data, see Extended Data Fig. 2b. Source numerical data, unprocessed blots and images are available in the Source data.
Next, we assessed whether TRF2-depleted telomeres, in the absence of NMD activity, undergo end-to-end chromosome fusions. Metaphase analysis confirmed that the deletion of Trf2 is not sufficient to trigger telomere fusions (Fig. 2e) and revealed that NMD single knockouts show no increase in fusions relative to the parental cell line (Fig. 2e and Extended Data Fig. 2b). By contrast, TRF2 depletion in the context of an NMD-deficient background resulted in frequent fusions (Fig. 2e and Extended Data Fig. 2b). The frequency of telomeric fusions varied among the different NMD-deficient backgrounds. Smg5−/− and Smg6−/− showed the highest fusion rates (27.5% and 24.2%), followed by Upf1HM and Smg7−/− (14.8% and 13.3%), while Smg8−/− and Smg9−/− exhibited lower levels (10.9% and 7.4%) (Fig. 2e and Extended Data Fig. 2b). Despite these differences, all the Trf2−/− NMD−/− cell lines analysed showed frequent telomere fusions and failed to proliferate (Figs. 1c and 2e and Extended Data Figs. 1b and 2b). Thus, the observed DNA damage activation at telomeres and frequent chromosome fusions provide a mechanistic basis for the synthetic lethality between Trf2 loss and impaired NMD in ES cells.
Acute NMD inhibition in ES cells results in telomeric fusions
To rule out clonal selection artefacts, we acutely inhibited NMD using two approaches. To this end, we used two independent approaches. First, we introduced a TAG degron (dTAG)39 at the endogenous Upf1 locus (Extended Data Fig. 3a,b) in Trf2Flox/Cre-ER mES cells. Upon addition of the dTAG ligand, UPF1 is rapidly degraded as confirmed by western blot and immunofluorescence (IF) (Fig. 3a,b). In these cells, TRF2 was first depleted with OHT, then UPF1 was degraded through dTAG treatment for 48 h. Notably, in this setting, codepletion of TRF2 and UPF1 also resulted in frequent telomeric fusions (~12%) (Fig. 3c,d), comparable to those observed in constitutive knockout cell lines (Fig. 2e and Extended Data Fig. 2b). This is in striking contrast to the phenotype of cells depleted for either Trf2 or Upf1 alone (Fig. 2e and Extended Data Fig. 2b). This result indicates that acute disruption of NMD is sufficient to compromise telomere protection, excluding a confounding effect arising during clonal selection of NMD-deficient cells. As an alternative method, we pharmacologically inhibited the NMD pathway using the SMG1 inhibitor 11j (SMG1i), a pyrimidine analogue that blocks UPF1 phosphorylation40 (Extended Data Fig. 3c). We first deleted TRF2 through OHT treatment, followed by treatment with the NMD-inhibitor 11j (1 µM) for an additional 48 h. Similar to the data obtained by genetic ablation, SMG1i treatment in TRF2-depleted ES cells led to frequent telomeric fusions (~15%) (Fig. 3e,f). Consistent with these findings, we also observed a notable growth defect in cells treated with SMG1i (1 μM and 2 μM) in conjunction with OHT-mediated TRF2 deletion (Fig. 3g). Collectively, these data further corroborate the notion that NMD activity contributes to telomere protection in mES cells in the absence of TRF2.
a, Western blot showing expression of UPF1 in Upf1dTAG ES cells either treated with dimethyl sulfoxide (DMSO) (−) or treated with the small molecule dTAG-13 (+). Detection performed by antibody detecting endogenous UPF1, or by anti-FLAG antibody detecting Upf1dTAG-FLAG. Representative images from minimum two independent experiments per genotype are shown. b, FLAG IF in UPF1dTAG treated with DMSO or dTAG-13 (dTAG). c, FISH for telomeric DNA (green) and DAPI staining (red) on metaphases derived from TRF2-proficient (Ctrl) or TRF2-deficient (OHT) ES cells, either left untreated (DMSO) or treated with dTAG-13 (dTAG) to degrade UPF1. d, Quantification of telomeric fusions shown in c, mean ± s.d. from three (n = 3) biological replicates. More than 1,656 chromosomes per genotype were scored. For details on the exact chromosome number per genotype, see the Source data. Statistical analysis by one-way ANOVA; n.s., not significant (P > 0.9999; P = 0.3962), ****P = 0.0001. e, Representative metaphases from ESCs treated with the SMG1 inhibitor 11j ± OHT. f, Quantification of telomeric fusions in e, mean ± s.d., n = 3 biological replicates. Statistical analysis by one-way ANOVA; n.s., not significant (P = 0.7548, P = 0.6116), ****P = 0.0001. More than 2,130 chromosomes per genotype were scored. For details on the exact chromosome number per genotype, see the Source data. g, Cell proliferation of TRF2-proficient (black lines) and TRF2-deficient (OHT-treated, red lines) cells in the presence of increasing concentrations of the SMG1 inhibitor (SMG1i). Cells were either untreated (0 μM) or treated with the indicated doses of SMG1i. Proliferation was monitored by confluence using the Incucyte S3 system. Data represent the mean ± s.d derived from the analysis of 49 images per condition. Representative images from minimum two independent experiments per genotype are shown. Source numerical data, unprocessed blots and images are available in Source data.
NMD deficiency results in accumulation of a dominant-negative TRF1 isoform (TRF1ΔE8) by preventing its degradation
RNA sequencing (RNA-seq) analysis of Smg5−/−, Smg6−/− and Upf1HM cell lines identified genes deregulated by NMD loss. Differential isoform expression analysis in Smg5−/−, Smg6−/− and Upf1HM revealed that loss of SMG5 had the most pronounced effect compared with the depletion of other NMD factors (Fig. 4a), consistent with previous reports23,37,41. This analysis identified 501 transcript isoforms that were differentially expressed across all three genotypes (Fig. 4a and Supplementary Table 2). The depletion of NMD factors led to both upregulation and downregulation of RNA isoforms, including well-established NMD targets such as Srsf3 (Fig. 4c,d and Extended Data Fig. 4c). Notably, among the genes significantly affected upon NMD inhibition was Trf1, which encodes the telomere-associated protein TRF1 (Fig. 4b–d and Extended Data Fig. 4a–c). Transcriptome analysis revealed that NMD-deficient cell lines accumulate a splice variant of Trf1 lacking exon 8 (Trf1ΔE8) (Fig. 4b and Extended Data Fig. 4a). This isoform is predicted to be a NMD target owing to the presence of PTCs in exon 9 (Extended Data Fig. 4f). Based on isoform abundances in our RNA-seq dataset, we estimate that TRF1ΔE8 isoform accounts for approximately 20–50% of total Trf1 transcripts in NMD-deficient cells (Extended Data Fig. 4a–c). Transcripts upregulated in NMD-deficient cells were highly enriched for PTC-containing isoforms, including Trf1ΔE8 and PTC containing Srsf3 isoforms, indicating that the majority of the observed expression changes were due to inhibition of NMD activity42,43 (Fig. 4c,d and Extended Data Fig. 4c).
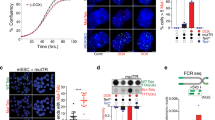
a, Venn diagram illustrating the overlap and unique sets of differentially expressed transcript isoforms in Smg5−/−, Smg6−/− and Upf1HM ES cells compared with NMD-proficient controls. Numbers indicate the total number of isoforms that are differentially expressed either uniquely in one genotype or shared across two or more conditions. b, Sashimi plots showing splicing events across exons 7–10 of the Trf1 transcript in wild type (NMD-proficient) and Upf1HM, Smg6−/− and Smg5−/− ES cells. Junction read numbers are shown. Note the presence of a splice variant lacking exon 8 (Trf1ΔE8) in NMD-deficient ES cells. c, Scatter plot showing differential RNA abundance (L2FC) in Smg5−/− (y axis) and Smg6−/− (x axis) ES cells relative to wild-type controls. Each point represents a transcript isoform. Isoforms significantly upregulated are shown in red, downregulated in blue and non-significantly changed in grey. NMD-sensitive (PTC-containing) and NMD-insensitive isoforms of Trf1 and Srsf3 are labelled. The Pearson correlation coefficient (R), indicating the similarity in isoform expression changes between the two genotypes, is shown on the right. d, The same as c, but comparing Smg5−/− (y axis) and Upf1HM (x axis) ES cells. e, Detection of the splice variant lacking exon 8 (Trf1ΔE8) by RT–qPCR in NMD-proficient (Ctrl), Smg6−/− and Upf1HM ES cells. Data are mean ± s.d. from three biological replicates; statistical analysis by one-way ANOVA. f, Detection of the splice variant lacking exon 8 (Trf1ΔE8) by RT–qPCR in cells either left untreated or treated with the SMG1 inhibitor, 11j. Two biologically independent SMG1 inhibitory experiments were performed; data points are shown on the graph. g, Western blot analysis for TRF1 expression in NMD-proficient (Ctrl) cells and Smg6−/− ES cells. Actin was used as a loading control. h, Western blot analysis for the expression of the TRF1–dTAG–FLAG fusion protein in cells in which the endogenous TRF1 gene was either untagged (WT) or tagged with a dTAG–FLAG construct (TRF1dTAG). Cell lysates were collected from SMG5-proficient cells (Ctrl). For g and h, representative images from three independent experiments per genotype are shown. i, ChIP assay performed on TRF1dTAG ES cells that are SMG5-proficient (Ctrl) or deficient (Smg5−/−), in the presence (+) of, or in the absence of the SMG1 inhibitor 11j. Input DNA (0.5% of the total DNA used), as well as DNA pulled down with H3 or FLAG, was hybridized with a radio-labelled telomeric probe (TelC). j, ChIP and input signals from i were quantified using ImageJ, and the ChIP signal was normalized with the corresponding input signal for both H3 and FLAG. Source numerical data and unprocessed blots are available in the Source data.
To confirm this observation, we designed a reverse transcription quantitative polymerase chain reaction (RT–qPCR) strategy to amplify selectively the Trf1ΔE8 splice variant targeting the exon 7–9 junction (Extended Data Fig. 4d,e). Using this approach, we confirmed that Smg6-, Upf1- and Smg5-depleted cell lines display elevated expression of the TRF1ΔE8 isoform (Fig. 4e and Extended Data Fig. 4d). Furthermore, amplification of Trf1 transcripts using primers complementary to exon 7 and exon 9 revealed the presence of a smaller isoform corresponding to the size of Trf1ΔE8 in NMD-deficient cells (Extended Data Fig. 4e). Sequencing analysis confirmed that the smaller isoform corresponds to the Trf1ΔE8 splice variant. Finally, we tested whether acute inhibition of the NMD pathway results in accumulation of the Trf1ΔE8 isoform. For this experiment we treated wild-type cells with SMG1i, and we consistently observed a significant (>40-fold) accumulation of the Trf1ΔE8 isoform (Fig. 4f), supporting the notion that the expression of Trf1 isoforms is tightly regulated by the NMD pathway.
Given the abundance of the Trf1ΔE8 isoform in NMD deficient cells, we next asked whether this isoform is translated into a protein. The predicted truncated protein expressed by Trf1ΔE8 retains the N-terminal acidic domain and the TRF dimerization domain (TRFH) but lacks the C-terminal DNA binding domain (Myb) (for schematics, see Extended Data Fig. 4f). Using an antibody raised against TRF1, we detected a truncated TRF1 protein in Smg6-deficient cells that was not present in wild-type control cells (Fig. 4g). The size of this protein is consistent with the predicted molecular weight (~34.2 kDa) of the Trf1ΔE8 translation product (Extended Data Fig. 4f). Similarly, Smg5-deficient and Upf1HM cells (Extended Data Fig. 4g), as well as cells treated with SMG1i, showed accumulation of the truncated protein (Extended Data Fig. 4h).
Given that TRF1 binding to telomeres requires homodimerization, and that the truncated protein encoded by the Trf1ΔE8 isoform retains the TRFH dimerization domain (TRFH) but lacks the DNA binding Myb domain, we reasoned that this truncated protein could act as a ‘dominant negative’ by sequestering full-length TRF1 away from telomeres, similar to previously engineered dominant-negative Trf1 alleles44. To test this hypothesis, we performed chromatin immunoprecipitation (ChIP) experiments to determine whether NMD-deficient cells expressing the Trf1ΔE8 have a reduced level of TRF1 bound to telomeres. To avoid confounding effects due to potential differences in antibody affinity between Trf1 isoforms, we tagged the endogenous Trf1 locus with a FLAG epitope using CRISPR-mediated gene editing. The resulting cells (see Extended Data Fig. 4i for schematics) expressed FLAG-tagged TRF1 that localized to telomeres (Extended Data Fig. 4j). Depletion of SMG5 in these cells led to the expected accumulation of a truncated TRF1 protein detectable with the FLAG antibody (Fig. 4h). Using these cells, we quantified the fraction of telomeric DNA associated with TRF1 by ChIP with an anti-FLAG antibody. This analysis revealed that TRF1 binding to telomeres was substantially reduced (~3-fold) in NMD-deficient cells compared with NMD-proficient control cells (Fig. 4i,j). Similarly, treatment with SMG1i also caused a marked reduction in TRF1 association with telomeres (Fig. 4i,j). Together, these results suggest that NMD activity regulates TRF1 localization to telomeres by controlling expression of the Trf1ΔE8 isoform.
TRF1ΔE8 drives telomere dysfunction in TRF2-depleted ES cells
To test whether expression of Trf1ΔE8 is sufficient to trigger telomere dysfunction and induce telomere fusions in TRF2-depleted mES cells, we generated cells in which either Trf1ΔE8 or full-length Trf1 (Trf1FL) can be expressed in a doxycycline (DOX)-inducible manner (Fig. 5a–c). To assess the impact of these constructs on telomere integrity, we performed metaphase analysis. Overexpression of TRF1FL, in either the presence or absence of TRF2, did not induce telomere fusions (Fig. 5d). By contrast, DOX-mediated induction of TRF1ΔE was sufficient to trigger frequent telomere fusions in TRF2-depleted cells, similar to what was observed in NMD-deficient cells (Fig. 5d). Notably, TRF1ΔE8 expression also led to telomere fusions in the presence of TRF2, suggesting that high levels of this truncated isoform are sufficient to disrupt telomere protection even when TRF2 is present. However, when TRF2 depletion was combined with TRF1ΔE8 expression, the frequency of telomere fusions reached levels comparable to those observed in NMD-deficient cells (Fig. 2e and Extended Data Fig. 2b), indicating that the Trf1 C-terminal truncated protein probably acts in a dominant-negative manner.
a, Schematic of Myc-tagged TRF1FL and TRF1ΔE8 DOX-inducible constructs. Stop codons resulting from exon 8 skipping events are indicated in the Trf1ΔE8 construct. b,c, Expression of TRF1 and, as a loading control, tubulin in NMD-proficient cells (b) and SMG6-deficient cells (c). DOX treatment results in the expression of full length TRF1 in iTRF1FL cells and of the TRF1ΔE8 variant in the iTRF1ΔE8 cells. d, Metaphases derived from Ctrl or Trf2−/− ES cells expressing TRF1FL or TRF1ΔE8 ± DOX. Percentage of telomere fusions were calculated from a total of >17 metaphases per genotype. More than 678 chromosomes per genotype (n = 3 biological replicates) were scored. For details on the exact chromosome number per genotype, see the Source Data. Mean ± s.d., one-way ANOVA; n.s., not significant (P ≥ 0.9950), **P = 0.0013. e, Metaphases from Smg6−/− or Smg6−/− Trf2−/− ES cells expressing TRF1FL ± DOX. The percentage of telomere fusions was calculated from a total of >19 metaphases per genotype. More than 880 chromosomes per genotype (n = 3 biological replicates) were scored. For details on the exact chromosome number per genotype, see the Source Data. Mean ± s.d., one-way ANOVA; ****P = 0.0001. f, Expression of TRF1FL and iTRF1ΔE8 without (−) and upon DOX treatment (+DOX), with actin as a loading control. g, ChIP assay performed on TRF1dTAG ES cells (Ctrl.), untreated TRF1dTAG iTRF1ΔE8 (−) and treated with DOX (+DOX). Input DNA (1% of the total DNA used), as well as DNA pulled down with FLAG, was hybridized with a radio-labelled telomeric probe (TelC). A representative image from three independent experiments per genotype is shown. h, ChIP and input signals from g were quantified using ImageJ, the ChIP signal was normalized with the corresponding input signal for FLAG. i, Model: NMD activity restrict the expression of the Trf1ΔE8 splice variant. Upon NMD inhibition, the Trf1ΔE8 transcript accumulates, producing TRF1ΔE8 protein, which binds full-length TRF1, and interferes with its binding to telomeres, resulting in telomere deprotection. Source numerical data, unprocessed blots and images are available in the Source data.
TRF1 overexpression is sufficient to restore NMD-mediated telomere protection
The data outlined above suggest that accumulation of TRF1ΔE8 is sufficient to phenocopy the telomere deprotection phenotype seen in TRF2/NMD codepleted cells. To directly test whether reduced levels of TRF1 at telomeres are the primary cause for the loss of telomere dysfunction in this context, we asked whether increasing the levels of TRF1FL in NMD-deficient cells would be sufficient to suppress the accumulation of telomere fusions upon TRF2 depletion. To this end, we introduced a DOX-inducible TRF1FL expression construct in Smg6-deficient, TRF2 conditional mES cells. In this genetic background, as expected, TRF2 depletion triggers frequent telomere fusions (Fig. 5e). Notably, TRF1 expression was sufficient to almost completely suppress telomere fusions, reducing their frequency from ~30% to 5% (Fig. 5e). A similar suppression of telomere fusions was observed in Upf1HM cells (Extended Data Fig. 5b,c).
To further evaluate whether expression of TRF1ΔE8 is sufficient to reduce TRF1 occupancy at telomeres and induce telomere fusions, we performed both ChIP (Fig. 5f–h) and IF (Extended Data Fig. 5h,i) for full-length TRF1 in cells expressing an ectopic TRF1 allele lacking exon 8. In both assays, we observed decreased levels of full-length TRF1 at telomeres, accompanied by a concomitant increase in telomere fusion events (Fig. 5f–i). These findings indicate that the TRF1ΔE8 isoform contributes to telomere fusions by reducing the effective TRF1 protection at chromosome ends.
Consistent with these findings, we depleted TRF2 and acutely degraded TRF1 using a dTAG degron strategy (Extended Data Fig. 4i,j, Extended Data Fig. 5d–g). In our setting, we observed high levels of telomere fusions in the double-depleted cells and high levels of γH2AX TIF-positive cells (Extended Data Fig. 5d–g), similarly to what was reported previously11. These results are consistent with the range of telomere fusions we detect in NMD-deficient cell lines (Fig. 2e and Extended Data Fig. 2b), supporting the idea that impaired TRF1 function contributes substantially to the observed telomere instability.
Collectively, these data demonstrate that, in the context of NMD deficiency, TRF2 loss leads to telomere fusions primarily due to reduced levels of TRF1 at telomeres. Consistent with this, increasing TRF1FL levels is sufficient to prevent telomere fusions in TRF2/NMD codepleted cells. Furthermore, expression of the dominant-negative TRF1ΔΕ8 isoform is sufficient to trigger telomere fusions even in NMD-proficient cells. Thus, we conclude that the NMD pathway safeguards telomere integrity in ES cells by preventing the accumulation of the Trf1ΔE8 isoform, which displaces endogenous TRF1 from telomeres and compromises their protection. This is consistent with previous findings showing that the expression of a truncated dominant-negative TRF1 mutant inhibits the binding of full-length TRF1 to telomeres44. Interestingly, a dominant-negative allele of Trf1 with mutated dimerization domain also interfered with telomeric localization of the full-length TRF1 in >80% nuclei45.
Discussion
Here, we describe a role for the NMD pathway in telomere protection in mES cells. We found that, while depletion of TRF2 does not cause overt telomere deprotection in mES cells, its loss in NMD-deficient cells leads to the accumulation of DNA damage factors at telomeres and the frequent formation of end-to-end chromosomal fusions. While TRF2 depletion alone does not cause telomere deprotection in mES cells, its loss in NMD-deficient cells leads to DNA damage accumulation and frequent chromosome fusions, with individual NMD components contributing differentially to this process (Fig. 2e and Extended Data Fig. 2b).
Transcriptome analysis of cells depleted for SMG5, SMG6 and UPF1 identified a set of core NMD targets in mES cells, which was, as expected, highly enriched for transcript isoforms containing premature translation termination codons46. Strikingly, we found that Trf1, a critical telomere-associated protein, is strongly regulated by NMD in mES cells. In NMD-deficient cells, approximately 20–50% of Trf1 transcripts skip exon 8, generating a truncated protein that lacks the C-terminal MYB DNA-binding domain but retains the TRFH dimerization domain. Our data suggest that this truncated isoform can heterodimerize with full-length TRF1, sequestering it away from telomeres in a dominant-negative manner. Consequently, TRF1 levels at telomeres are notably reduced in NMD-deficient cells (Fig. 5i). Based on ectopic expression experiments, we conclude that this truncated isoform is sufficient to account for the telomere deprotection phenotype observed in the absence of NMD. We propose a model in which the NMD pathway safeguards telomere integrity by preventing the accumulation of a dominant-negative Trf1 transcript that produces a dominant-negative TRF1 protein. In this model, NMD deficiency lowers TRF1 availability at telomeres and, when combined with TRF2 loss, results in catastrophic telomere dysfunction and cell death.
These findings raise important questions regarding telomere homeostasis. One is whether modulation of NMD activity could provide a strategy to control TRF1 levels. Given that TRF1 acts as a negative regulator of telomerase activity44,47,48, transient suppression of NMD might be harnessed to enhance telomerase function in specific contexts. Supporting this idea, persistent DNA damage has been shown to suppress NMD activity22, suggesting a feedback mechanism to promote telomere healing through telomerase activation in cells experiencing telomere dysfunction.
Another notable implication is the unexpected observation that TRF1 and TRF2 have overlapping functions in mES cells, where TRF1 appears capable of compensating for TRF2 loss. This stands in sharp contrast to findings in somatic cells, where TRF2 is essential for telomere protection, and its depletion leads to extensive telomere deprotection despite the continued presence of TRF11,8,49. This raises the question: why can’t TRF1 fulfil a similar protective role in somatic cells? One possibility is that TRF1 gains additional functions in mES cells via post-translational modifications or pluripotency-specific protein interactions, enabling it to promote T-loop formation or directly suppress ATM signalling functions typically attributed to TRF2. Alternatively, the high expression of TRF1 in pluripotent stem cells50,51,52 may allow it to compensate for TRF2 loss by sheer abundance. In this model, TRF1 may possess a weak intrinsic ability to substitute for TRF2, which becomes functionally relevant only when TRF1 is expressed at high levels.
Finally, our data provide an intriguing connection between the NMD pathway and genome integrity. Several studies have implicated the NMD pathway in regulatory functions in stem cell biology and genome stability20,22,23,26,27,53. Notably, NMD has been implicated in telomere biology in mammalian cells through its function in controlling a telomeric repeat-containing RNA (TERRA) that accumulates in cancer cells utilizing alternative lengthening of telomere, and plays roles in telomere homeostasis and DNA damage signalling16,54,55,56. Our data add a new piece to this puzzle showing that, in mES cells, the NMD pathway plays an essential role in ensuring TRF2-independent telomere protection. Our study identifies a synthetic lethal interaction between TRF2 and NMD, revealing how post-transcriptional regulation controls telomere protection in pluripotent stem cells.
Methods
ES cell derivation and culture
Trf2f/f-Rosa26-creER ES cell derivation and growth conditions were previously described10. To induce Cre activity, 4-hydroxytamoxifen (OHT; 0.6 nM; Sigma) was added 96 h before cell collection unless otherwise specified. Where indicated, cells were treated with the NMD inhibitor 11j (1 µM; Fisher Scientific, 50-225-9662), the dTAG-13 degrader (0.5 µM; MilliporeSigma, SML2601-1MG) or DOX (500 ng ml−1, MilliporeSigma, D9891) for 48 h before collection. To generate knockout cell lines, ES cells were nucleofected (Mouse Embryonic Stem Cell Nucleofector Kit, Lonza, VPH-1001) with a cocktail containing three synthetic gRNAs (GKOv2 kits, Synthego, now EditCo) and Cas9 Nuclease V3 (100 µg; IDT, 1081060). Clonal lines were isolated, screened by PCR and validated by Sanger sequencing and immunoblotting. Primers used for PCR amplification were
Smg5 - forward (F): 5′- GAGCTTGTCACATGAGAGGTCT-3′
Smg5 - reverse (R): 5′- ACTCGCACCCATTTGGAGAG-3′
Smg6 - forward (F): 5′- CCTTTGGGACCTCGACTTTT-3′
Smg6 - reverse (R): 5′- TTGCTGCATGTTCCGCAC-3′
Smg7 - forward (F): 5′- CTCCAAAGAGCTGCCTTAGGT-3′
Smg7 - reverse (R): 5′- GGCTTTGCTTTGGTGGATGG-3′
Smg8 - forward (F): 5′- CCATACCTCCGCGAAGTGAA-3′
Smg8 - reverse (R): 5′- GCGGAGGAGGGATTTCACAA-3′
Smg9 - forward (F): 5′- GCCTCTGGTTTGTTTGTGGG-3′
Smg9 - reverse (R): 5′- TGAGCCCACCTCCCCTTTAT-3′
Upf1 - forward (F): 5′- TCTAACTGGGACCTGGCTCA-3′
Upf1 - reverse (R): 5′- CTCAGAGCTCAGAACCGGC-3′.
Two independent clones were analysed per gene. dTAG cell lines were established as previously described39. N-terminal FKBP12F36V–FLAG fusions were introduced at the endogenous Upf1 and Trf1 loci. Degradation of the fusion proteins was confirmed by treatment with dTAG-13 (0.5 µM).
Cell growth analysis
Cell proliferation was monitored in real time using the IncuCyte S3 system (Essen Bioscience) based on confluence measurements. ES cells were imaged every 3 h with a 10× objective. At least two biologically independent replicates were analysed per condition.
Clonogenic assay
ES cells were dissociated into a single-cell suspension and seeded at low density (500 cells per well in a 6-well plate). Cells were cultured undisturbed for 7 days to allow colony formation, then fixed with methanol and stained with crystal violet.
DOX-inducible Trf1 expression from PiggyBac vectors
PiggyBac PB-TRE-mTRF1FL or PB-TRE-mTRF1ΔE8 constructs were cloned from the PB-TRE-dCas9-VPR plasmid (63800, Addegene)12. ES cells were cotransfected with the PiggyBac and a transposase-expressing plasmid using Lipofectamine 2000 (Thermo Fisher Scientific, 11668019) for stable integration.
Western blotting
Cells were lysed in 2× Laemmli buffer and proteins separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis on 4–20% TGX stain-free gels (Bio-Rad, 4568093). Proteins were transferred to nitrocellulose membranes and probed with antibodies against SMG5 (1:1,000; Abcam, ab33033), SMG6 (1:1,000; Abcam, ab87539), SMG7 (1:1,000; Bethyl Laboratory, A302-170A), SMG9 (1:1,000; Abcam, ab85659), UPF1 (1:1,000; Bethyl, A301-902A), TRF1 (1:200; Abcam, ab192629), actin (1:1,000; MilliporeSigma, A5441) or tubulin (1:5,000; MilliporeSigma, T5168). Detection used horseradish peroxidase- or DyLight-conjugated secondary antibodies and the ChemiDoc MP system (Bio-Rad).
IF and IF–fluorescence in situ hybridization (FISH)
Cells were fixed with 2% paraformaldehyde and stained with primary antibodies (1:1,000): OCT3/4 (Santa Cruz, sc-5279), γH2AX (Millipore, 05-636), 53BP1 (Novus, NB100-304), FLAG (MilliporeSigma, F1804) and MYC (Cell Signaling, 2276). After secondary antibody incubation, cells were post-fixed, denatured at 72 °C and hybridized with AlexaFluor 488-TelC PNA probe (PNA Bio, F1004). Slides were mounted with ProLong Gold antifade (Thermo Fisher, P36931), and images were acquired using a Zeiss Axio Imager M2 and Axiocam 702 with ZEN 2.6 software. Z-stacks were displayed as maximum intensity projections. Figures were assembled with Adobe Illustrator 2024.
FISH on metaphase spreads
Cells were treated with colcemid (0.2 µg ml−1, 2 h), swelled in 75 mM KCl, fixed in methanol:acetic acid (3:1) and dropped onto glass slides. Telomeres were detected using AlexaFluor 488-TelC PNA probe and counterstained with DAPI. Images were acquired using a Zeiss Axio Imager.
Statistics and reproducibility
Statistical analyses were performed using GraphPad PRISM version 9.0 software (GraphPad). Data represent the mean ± s.d. of three independent experiments, unless stated otherwise. P ≤ 0.05 was considered statistically significant, and P values were assessed by one-way analysis of variance (ANOVA) followed by multiple comparisons. Sample size was not predetermined. For experiments involving quantification of positive cells (for example, OCT4 or TIFs positive) or chromosome fusions, n = 3 biological replicates was chosen as the minimal replicate number, and the sample size was determined by the number of positive cells within the replicates. We determined this to be sufficient owing to internal controls and low observed variability between stained samples. Data were not excluded from analysis. All replication attempts were successful, and observed data were consistent with orthogonal methods and previously known results. Cells and chromosomes used for imaging were selected randomly and analysed equally with no subsampling. Blinding was applied to all the images before quantification.
CRISPR–Cas9 screen
Three independent genome-wide CRISPR screens were performed using a genome-wide CRISPR knockout library (73633-LVC, Addgene) in Trf2−/− and Trf2f/−f ES clones expressing Cas9 (no. 52962, Addgene), as described previously10. Following selection, a portion of cells was collected as the initial timepoint (T0), and the remaining cells were cultured for 14 days before collection as the final timepoint (T14). Genomic DNA was isolated using a DNeasy Blood and Tissue Kit (Qiagen, 69506), and single guide RNAs (sgRNAs) were PCR-amplified using primers containing Illumina adapters. Samples were sequenced on an Illumina NextSeq platform, and the resulting reads were analysed using MAGeCK57. An essentiality score (β score) for each gene was calculated using the MAGeCK -mle module. Average β scores were calculated from three independent experiments. Potential synthetically lethal genes were: (1) having a negative (<−0.75) β scores in the Trf2−/− cells; (2) having a ‘neutral’ β score (>−0.25 and lower than 1) in the control Trf2−/f ES clones; and (3) having a difference in β scores (control – OHT) greater than 1 in all the experiments (Supplementary Tables 1 and 4). Data were plotted using ggplot2.
RNA extraction and RT–qPCR
RNA was extracted from 1 × 106 cells using the RNeasy Plus Mini Kit (Qiagen, 74136). Then, 500 ng of RNA was reverse transcribed using PrimeScript IV 1st strand cDNA Synthesis Mix (TakaraBio, 6215A). SYBR Green PCR Master Mix (ThermoFisher, AppliedBiosciences, 4368706) was used for quantitative PCR. Primers used were
Trf1 (exon 7) - forward (F): 5′-TGTTAATGGCCAGCAGTCT-3′
Trf1 (exon 7/exon 9 junction) - reverse (R): 5′-CATCGTTGTTTCACCTTATTGAGGA-3′
Trf1 (exon 9) - reverse (R): 5′-TCCACTGGTTCTTCGGTTCC-3′
Gapdh forward (F): 5′-TGTGTCCGTCGTGGATCTGA-3′
Gapdh reverse (R): 5′-TTGCTGTTGAAGTCGCAGGAG-3′.
ChIP assay
Trf1dTAG (control and 11j-treated ES cells for 48 h) and Trf1dTAG Smg5−/− samples were cross-linked using 1% formaldehyde (Thermo Fisher Scientific, cat. no. 28908) at room temperature for 3 min. The reaction was quenched with 125 mM glycine, and cells were collected by centrifugation at 250g for 5 min at 4 °C and sonicated using the Bioruptor sonication system Covaris S220 sonicator (Covaris). Chromatin from 4 × 106 cells was used for immunoprecipitation.
In total, 30 μg of clear chromatin was incubated with Dynabeads Protein A (Thermo Fisher, cat. no. 10002D) or Dynabeads Protein G (Thermo Fisher, cat. no. 10003D) together with either 4 μg of mouse monoclonal FLAG antibody (F1804, MilliporeSigma) or 2 μg of H3 antibody (ab1791, Abcam). Inputs correspond to the 0.5% or 1% fraction of the total DNA sample used in the immunoprecipitation. The precipitated DNA was eluted and transferred to a Hybond+ membrane by dot blotting. The membrane was then hybridized with a P32-labelled telomeric probe recognizing TTAGGG repeats and visualized with the PhosphorImager (Typhoon biomolecular imager). The signals were quantified using ImageJ software.
Apoptosis and cell cycle analysis
Apoptosis was assayed using the Annexin-V Apoptosis Detection Kit (Abcam, ab14150) according to the manufacturer’s instructions. In brief, cells (1 × 106) were collected by centrifugation and resuspended in 500 µl of the supplied Binding Buffer II. Annexin V (5 µl) was added to each sample, followed by incubation for 5 min at room temperature in the dark. Fluorescence was analysed by flow cytometry using Cy5 excitation/emission settings on CytoFLEX S Flow Cytometer (CytoFLEX S). Cells positive for Annexin V–Cy5 were interpreted as apoptotic.
Cell-cycle distribution was assessed by propidium iodide (PI) staining of fixed cells. In brief, cells were collected, washed with phosphate-buffered saline and fixed in 70% ice-cold ethanol. Samples were stored at −20 °C for more than 2 h. After fixation, cells were washed with phosphate-buffered saline and incubated for 30 min at room temperature in PI staining solution (50 µg ml−1) with RNase A 100 µg ml−1. Samples were analysed by flow cytometry using a 488-nm laser, and DNA content was quantified in the PI channel. Cell-cycle phases (G1, S and G2/M) were determined using FlowJo software.
Total RNA-seq
RNA extractions of control Trf2f/f ES cells as well as Smg5−/−, Smg6−/− and Upf1HM were performed using RNeasy Plus Mini Kit (Qiagen) following the manufacturer’s protocol. Total RNA was then ribo-depleted, followed by library preparation using the Illumina Stranded Total RNA Prep. Paired-end libraries were pooled and sequenced on NovaSeq 6000 SP. The samples had 99–140 million pass filter reads with more than 90% of bases above the quality score of Q30. Reads of the samples were trimmed for adapters and low-quality bases using Cutadapt. The initial alignment was performed using STAR (v.2.7.6a)58 with reference genome mm10 and GENCODE vM27 annotation.
Long-read annotation filtering
The long-read transcript annotation for mES cells (ES_consolidated.bed) from Pardo-Palacios et al.59 was obtained and converted to GTF format using UCSC Genome Browser utilities (bedToGenePred)60. The resulting GTF file was then filtered to retain only full splice match isoforms using a custom R script.
Transcript assembly and filtering
Transcriptomes were assembled from individual mouse ES cell RNA-seq samples using StringTie (v.2.1.4)61. For each sample, alignment files in BAM format were processed using StringTie with a minimum isoform fraction threshold of 0.05 (-f 0.05) and guided by the previously filtered GTF annotation reference (-G ES_consolidated.gtf). Trimming was disabled (-t) to preserve full-length transcript structures. Assembled transcriptomes were filtered to retain only isoforms with an expression level greater than 1 transcripts per million using a custom R script (StringTie_filter_gtf.R). Filtered sample-level annotations were merged in two stages using StringTie’s–merge function: first, within each experimental condition (Smg5−/−, Smg6−/−, Upf1HM and control) and then across all conditions to generate a unified annotation. The merged GTF file was sorted and indexed using IGVTools62. Unstranded transcripts were removed using a custom filtering script (Filter_gtf_strand.R).
Gene model refinement and annotation
To refine transcript models and remove likely spurious or low-abundance isoforms, we applied the AnnotationCleaner pipeline63 (https://github.com/isaacvock/AnnotationCleaner). This filtering step produced a high-confidence annotation, which was then used to predict open reading frames and identify PTCs using factR264.
Short-read RNA-seq alignment, quantification and differential expression analysis
A STAR index was generated from the cleaned annotation, and raw FASTQ files were aligned to the custom transcriptome using STAR (v.2.7.6a)58. Isoform- and gene-level expression estimates were then quantified using RSEM (v.1.3.3)65 with corresponding custom indices. Transcript-level expression data were normalized, and log2 fold changes (L2FC) in RNA abundance were calculated by Degust (https://degust.erc.monash.edu/; R script provided). Significant events were called at |L2FC| >1, Padj < 0.05 in Upf1HM, Smg5−/− and Smg6−/− relative to wild-type ES cells. Finally, L2FC (RNA abundance) values were integrated with PTC annotations to facilitate downstream analysis of isoform-specific regulation and NMD sensitivity (Supplementary Table 2).
Differential splicing analysis
Alternative splicing analysis was performed using the SpliceWiz R package66, beginning with the construction of a reference from the GENCODE vM27 annotation and mm39 genome. Aligned BAM files from mES cell samples were processed to quantify splicing events, followed by novel splice junction discovery using collateData(novelSplicing = TRUE, novelSplicing_requireOneAnnotatedSJ = TRUE, novelSplicing_minSamples = 3, novelSplicing_minSamplesAboveThreshold = 3, novelSplicing_countThreshold = 10, novelSplicing_useTJ = TRUE) with stringent filtering parameters. The resulting experiment was imported as a NxtSE object, and alternative splicing events were filtered for high-confidence features (default filters). Differential splicing analysis was then conducted using the ASE_DESeq(IRmode = "all") function, identifying significant percent spliced in (PSI) events (|ΔPSI| >0.15, Padj < 0.05) in Upf1HM, Smg5−/− and Smg6−/− relative to wild-type ES cells (Supplementary Table 3).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed RNA-seq data supporting Fig. 4 have been deposited in the NCBI Gene Expression Omnibus (GEO) and are accessible via accession number GSE300187. Source data are provided with this paper.
References
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
Jones-Weinert, C., Mainz, L. & Karlseder, J. Telomere function and regulation from mouse models to human ageing and disease. Nat. Rev. Mol. Cell Biol. 26, 297–313 (2025).
Doksani, Y. & de Lange, T. The role of double-strand break repair pathways at functional and dysfunctional telomeres. Cold Spring Harb. Perspect. Biol. 6, a016576 (2014).
de Lange, T. Shelterin-mediated telomere protection. Annu. Rev. Genet. 52, 223–247 (2018).
Lim, C. J. & Cech, T. R. Shaping human telomeres: from shelterin and CST complexes to telomeric chromatin organization. Nat. Rev. Mol. Cell Biol. 22, 283–298 (2021).
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012).
Lazzerini Denchi, E., Celli, G. & de Lange, T. Hepatocytes with extensive telomere deprotection and fusion remain viable and regenerate liver mass through endoreduplication. Genes Dev. 20, 2648–2653 (2006).
Okamoto, K. et al. A two-step mechanism for TRF2-mediated chromosome-end protection. Nature 494, 502–505 (2013).
Lobanova, A. et al. Different requirements of functional telomeres in neural stem cells and terminally differentiated neurons. Genes Dev. 31, 639–647 (2017).
Markiewicz-Potoczny, M. et al. TRF2-mediated telomere protection is dispensable in pluripotent stem cells. Nature 589, 110–115 (2021).
Ruis, P. et al. TRF2-independent chromosome end protection during pluripotency. Nature 589, 103–109 (2021).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Blake, L. A., Watkins, L., Liu, Y., Inoue, T. & Wu, B. A rapid inducible RNA decay system reveals fast mRNA decay in P-bodies. Nat. Commun. 15, 2720 (2024).
Baudrimont, A., Jaquet, V., Wallerich, S., Voegeli, S. & Becskei, A. Contribution of RNA degradation to intrinsic and extrinsic noise in gene expression. Cell Rep. 26, 3752–3761 (2019).
Azzalin, C. M. & Lingner, J. The human RNA surveillance factor UPF1 is required for S phase progression and genome stability. Curr. Biol. 16, 433–439 (2006).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Lew, J. E., Enomoto, S. & Berman, J. Telomere length regulation and telomeric chromatin require the nonsense-mediated mRNA decay pathway. Mol. Cell. Biol. 18, 6121–6130 (1998).
Isken, O. & Maquat, L. E. The multiple lives of NMD factors: balancing roles in gene and genome regulation. Nat. Rev. Genet. 9, 699–712 (2008).
Holstein, E. M., Clark, K. R. & Lydall, D. Interplay between nonsense-mediated mRNA decay and DNA damage response pathways reveals that Stn1 and Ten1 are the key CST telomere-cap components. Cell Rep. 7, 1259–1269 (2014).
Ngo, G. H. P., Grimstead, J. W. & Baird, D. M. UPF1 promotes the formation of R loops to stimulate DNA double-strand break repair. Nat. Commun. 12, 3849 (2021).
Kishor, A., Fritz, S. E. & Hogg, J. R. Nonsense-mediated mRNA decay: the challenge of telling right from wrong in a complex transcriptome. Wiley Interdiscip. Rev. RNA 10, e1548 (2019).
Nickless, A. et al. p38 MAPK inhibits nonsense-mediated RNA decay in response to persistent DNA damage in noncycling cells. J. Biol. Chem. 292, 15266–15276 (2017).
Huth, M. et al. NMD is required for timely cell fate transitions by fine-tuning gene expression and regulating translation. Genes Dev. 36, 348–367 (2022).
Li, T. et al. Smg6/Est1 licenses embryonic stem cell differentiation via nonsense-mediated mRNA decay. EMBO J. 34, 1630–1647 (2015).
Cook, A. L. et al. Identification of nonsense-mediated decay inhibitors that alter the tumor immune landscape. eLife 13, RP95952 (2025).
Zhao, B. & Pritchard, J. R. Evolution of the nonsense-mediated decay pathway is associated with decreased cytolytic immune infiltration. PLoS Comput. Biol. 15, e1007467 (2019).
Nasif, S., Contu, L. & Muhlemann, O. Beyond quality control: the role of nonsense-mediated mRNA decay (NMD) in regulating gene expression. Semin. Cell Dev. Biol. 75, 78–87 (2018).
Kim, Y. K. & Maquat, L. E. UPFront and center in RNA decay: UPF1 in nonsense-mediated mRNA decay and beyond. RNA 25, 407–422 (2019).
Kashima, I. et al. Binding of a novel SMG-1–Upf1–eRF1–eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev. 20, 355–367 (2006).
Yamashita, A. et al. SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. Genes Dev. 23, 1091–1105 (2009).
Arias-Palomo, E. et al. The nonsense-mediated mRNA decay SMG-1 kinase is regulated by large-scale conformational changes controlled by SMG-8. Genes Dev. 25, 153–164 (2011).
Deniaud, A. et al. A network of SMG-8, SMG-9 and SMG-1 C-terminal insertion domain regulates UPF1 substrate recruitment and phosphorylation. Nucleic Acids Res. 43, 7600–7611 (2015).
Okada-Katsuhata, Y. et al. N- and C-terminal Upf1 phosphorylations create binding platforms for SMG-6 and SMG-5:SMG-7 during NMD. Nucleic Acids Res. 40, 1251–1266 (2012).
Jonas, S., Weichenrieder, O. & Izaurralde, E. An unusual arrangement of two 14-3-3-like domains in the SMG5-SMG7 heterodimer is required for efficient nonsense-mediated mRNA decay. Genes Dev. 27, 211–225 (2013).
Huntzinger, E., Kashima, I., Fauser, M., Sauliere, J. & Izaurralde, E. SMG6 is the catalytic endonuclease that cleaves mRNAs containing nonsense codons in metazoan. RNA 14, 2609–2617 (2008).
Loh, B., Jonas, S. & Izaurralde, E. The SMG5-SMG7 heterodimer directly recruits the CCR4-NOT deadenylase complex to mRNAs containing nonsense codons via interaction with POP2. Genes Dev. 27, 2125–2138 (2013).
Boehm, V. et al. SMG5–SMG7 authorize nonsense-mediated mRNA decay by enabling SMG6 endonucleolytic activity. Nat. Commun. 12, 3965 (2021).
Chawla, R. et al. Human UPF1 interacts with TPP1 and telomerase and sustains telomere leading-strand replication. EMBO J. 30, 4047–4058 (2011).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Gopalsamy, A. et al. Identification of pyrimidine derivatives as hSMG-1 inhibitors. Bioorg. Med. Chem. Lett. 22, 6636–6641 (2012).
Chen, C., Wei, Y., Jiang, X. & Li, T. RNA surveillance factor SMG5 is essential for mouse embryonic stem cell differentiation. Biomolecules 14, 1023 (2024).
Pervouchine, D. et al. Integrative transcriptomic analysis suggests new autoregulatory splicing events coupled with nonsense-mediated mRNA decay. Nucleic Acids Res. 47, 5293–5306 (2019).
Aznarez, I. et al. Mechanism of nonsense-mediated mRNA decay stimulation by splicing factor SRSF1. Cell Rep. 23, 2186–2198 (2018).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Fairall, L., Chapman, L., Moss, H., de Lange, T. & Rhodes, D. Structure of the TRFH dimerization domain of the human telomeric proteins TRF1 and TRF2. Mol. Cell 8, 351–361 (2001).
Colombo, M., Karousis, E. D., Bourquin, J., Bruggmann, R. & Muhlemann, O. Transcriptome-wide identification of NMD-targeted human mRNAs reveals extensive redundancy between SMG6- and SMG7-mediated degradation pathways. RNA 23, 189–201 (2017).
Karlseder, J. et al. Targeted deletion reveals an essential function for the telomere length regulator Trf1. Mol. Cell. Biol. 23, 6533–6541 (2003).
Smogorzewska, A. et al. Control of human telomere length by TRF1 and TRF2. Mol. Cell. Biol. 20, 1659–1668 (2000).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Schneider, R. P. et al. TRF1 is a stem cell marker and is essential for the generation of induced pluripotent stem cells. Nat. Commun. 4, 1946 (2013).
Boue, S., Paramonov, I., Barrero, M. J. & Izpisua Belmonte, J. C. Analysis of human and mouse reprogramming of somatic cells to induced pluripotent stem cells. What is in the plate?. PLoS ONE 5, e12664 (2010).
Li, F., Ge, Y., Liu, D. & Songyang, Z. The role of telomere-binding modulators in pluripotent stem cells. Protein Cell 11, 60–70 (2020).
Lou, C. H. et al. Nonsense-mediated RNA decay influences human embryonic stem cell fate. Stem Cell Rep. 6, 844–857 (2016).
Bettin, N., Oss Pegorar, C. & Cusanelli, E. The emerging roles of TERRA in telomere maintenance and genome stability. Cells 8, 246 (2019).
Deng, Z., Norseen, J., Wiedmer, A., Riethman, H. & Lieberman, P. M. TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres. Mol. Cell 35, 403–413 (2009).
Porro, A. et al. Functional characterization of the TERRA transcriptome at damaged telomeres. Nat. Commun. 5, 5379 (2014).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Pardo-Palacios, F. J. et al. Systematic assessment of long-read RNA-seq methods for transcript identification and quantification. Nat. Methods 21, 1349–1363 (2024).
Hinrichs, A. S. et al. The UCSC Genome Browser Database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278 (2019).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Mabin, J. W. et al. Uncovering the isoform-resolution kinetic landscape of nonsense-mediated mRNA decay with EZbakR. Preprint at bioRxiv https://doi.org/10.1101/2025.03.12.642874 (2025).
Zhuravskaya, A., Yap, K., Hamid, F. & Makeyev, E. V. Alternative splicing coupled to nonsense-mediated decay coordinates downregulation of non-neuronal genes in developing mouse neurons. Genome Biol. 25, 162 (2024).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Wong, A. C. H., Wong, J. J., Rasko, J. E. J. & Schmitz, U. SpliceWiz: interactive analysis and visualization of alternative splicing in R. Brief. Bioinform. 25, bbad468 (2023).
Acknowledgements
We thank members of the Lazzerini Denchi and Hogg laboratories for critical reading of the manuscript and helpful discussion; S. Johns for critical feedback on this manuscript and support throughout the experimental work; CCR Sequencing Facility in Frederick for assistance and initial analysis, LGI Flow Cytometry Core and UIC Flow Cytometry Core for help with data collection and analysis; and M. Kwissa for assistance with analysis. This research was supported by the Intramural Research Program of the National Institutes of Health (NIH). The contributions of the NIH author(s) are considered Works of the United States Government. The findings and conclusions presented in this Article are those of the author(s) and do not necessarily reflect the views of the NIH or the US Department of Health and Human Services.
Author information
Authors and Affiliations
Contributions
E.L.D. and M.M.-P. conceived the study, designed the experiments and analysed the data; M.M.-P., S.Y.L. and S.C. conducted experiments; J.W.M. and J.R.H. analysed the transcriptomic data and provided critical support with the analysis of NMD-targets. J.R.H. helped in designing experiments and provided critical reagents; A.Z. generated Upf1dTAG ES cells; S.Y.L. and R.S. generated and validated Trf1dTAG ES cells. G.T. performed proteomic analysis of NMD-deficient cells. E.L.D. and M.M.-P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Steven Artandi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation and validation of NMD knockout ESC lines.
a, Schematic of Smg5, Smg6, Smg7, Smg8, Smg9 and Upf1 loci. Sanger sequencing confirms gene editing in two clones per gene. gRNAs targeting regions are indicated in red. b, Growth curves of ESCs of indicated genotypes monitored by Incucyte S3 following OHT-mediated TRF2 deletion (day 0). Data are mean ± s.d. from 49 images per condition. Colors: Ctrl (black), Trf2−/− (red), Smg knockouts (green), double mutants (blue).
Extended Data Fig. 2 Increased end-to-end fusions in NMD-deficient Trf2−/− ESCs.
a, Quantification of OCT4+ve cells in indicated genotypes; MEFs serve as negative control. Related to Fig. 1e. Data from three independent experiments. More than 656 cells per genotype (n = 3 biological replicates) were scored. Details on the exact cell number per genotype are in Source Data File. b, Percentage of telomere fusions calculated from a total of >19 metaphases per genotype. More than 1981 chromosomes per genotype were scored. Details on the exact chromosome number per genotype are in Source Data File. Related to Fig. 2e. Statistical analysis (n = 3 biological replicates) by one-way ANOVA; *P = 0.0117 ns, not significant, P > 0.9999; ****P = 0.0001; ***P = 0.001; *P = 0.0124. c, Representative gating strategy for all genotypes in (d) is shown. d, Annexin-V and Propidium Iodide co-staining analysis by flow cytometry is shown for TRF2-proficient and TRF2-deficient cells in control (Ctrl) and NMD-deficient cells (Smg5−/−, Smg6−/− and Upf1HM). Representative strategy from minimum three independent experiments per genotype are shown. e, Percentage of apoptotic cells positively stained with Annexin-V is shown. f, Representative gating strategy for all genotypes in (g) is shown. g, Cell cycle analysis by Propidium Iodide incorporation in fixed cells of indicated genotypes. Representative strategy from minimum three independent experiments per genotype are shown.
Extended Data Fig. 3 Generation of a degradable UPF1 mECS cell line.
a, Schematic of the dTAG degron system. The dTAG ligand recruits UPF1-FKBP12F36V to the CRBN E3 ubiquitin ligase complex, triggering targeted protein degradation1. b, Schematic of the FLAG-dTAG knock-in strategy at the endogenous Upf1 locus and validation of successful knock-in by Sanger sequencing. Chromatogram shows the integration site of the FLAG-FKBP12F36V within exon 1 of Upf1 in a correctly targeted mESC clone. c, Western blot showing expression of P-UPF1 and UPF1 in UPF1dTAG ESCs either treated with DMSO (-) or treated with the SMG1 inhibitor (SMG1i). Representative images from minimum three independent experiments are shown.
Extended Data Fig. 4 Characterization of TRF1 exon 8 skipping events in ESCs.
a, Sashimi plots of TRF1 splicing across exons 7–10 in Ctrl and NMD-deficient ESCs. Triplicates (n = 3) and junction read numbers are shown. Top panel as in Fig. 4b. b, Graph showing quantification of exon 8 skipping across genotypes as percentage of total splicing events. Statistical analysis (n = 3 biological replicates) by one-way ANOVA; ***P = 0.0004, ***P = 0.0002; ****P = 0.00001. c, Scatter plots showing the relationship between differential RNA isoform abundance (log2 fold change, y axis) and average isoform expression (x axis) in Smg5−/− (left), Smg6−/− (middle), and Upf1HM (right) ESCs relative to wild-type cells. Each point represents a transcript isoform. Significantly upregulated isoforms are highlighted in red, significantly downregulated in blue, and nonsignificant changes are shown in gray. NMD-sensitive (PTC-containing) and NMD-insensitive isoforms of Trf1 and Srsf3 are labeled. d, Schematic of the RT-PCR strategy used in Fig. 4e and Fig. 4f to detect the splice variant lacking exon 8 (TRF1DE8) using a forward primer on Exon 7 and a reverse primer spanning the exon 7 - exon 9 junction. e, A RT-PCR strategy used to detect TRF1FL and TRF1ΔE8. Representative agarose gel showing the amplification products in NMD-proficient (Ctrl) and NMD-deficient (Upf1HM) cells. f, Schematic of TRF1FL and TRF1ΔE8 isoforms. Skipping exon 8 causes frameshift and premature stop codon in exon 9. g, Western blot analysis using an antibody against TRF1 (top panel) and Tubulin as a loading control (bottom panel) on total cell lysates from ESCs of the indicated genotypes. Bands corresponding to full-length TRF1 (TRF1FL) and the truncated isoform lacking exon 8 (TRF1ΔE8) are indicated. Representative images from minimum three independent experiments are shown. h, Western blot analysis using an antibody against TRF1 (top panel) and Tubulin as a loading control (bottom panel) on total cell lysates from control ESCs untreated (-) or treated (+) with SMG1 inhibitor (11j). Bands corresponding to full-length TRF1 (TRF1FL) and the truncated isoform lacking exon 8 (TRF1ΔE8) are indicated. Representative images from minimum three independent experiments are shown. i, Schematic of the FLAG-dTAG knock-in strategy at the endogenous Trf1 locus. Successful integration was validated by Sanger sequencing. j, Immunofluorescence combined with FISH (IF-FISH) for telomeric DNA (TTAGGG, green) and FLAG (red) in control ESCs (Ctrl) and ESCs harboring the FLAG-dTAG construct at the Trf1 locus (Trf1dTAG), either treated with DMSO or dTAG.
Extended Data Fig. 5 Localization of TRF1FL and TRF1ΔE8.
a, Immunofluorescence combined with FISH (IF-FISH) showing the expression and localization of Myc-tagged inducible TRF1 constructs (red) and telomeric DNA (TTAGGG, green) in cells with stable integration of either full-length TRF1 (iTRF1FL) or TRF1 lacking exon 8 (iTRF1ΔE8). Cells were treated with doxycycline (DOX) to induce construct expression or with DMSO as a control. More than 155 cells per genotype were scored. Details on the exact cell number per genotype are in Source Data File. b, IF-FISH for Myc (red) and telomeric DNA (TTAGGG, green) performed in Smg6−/− or Upf1HM ESCs stably expressing full-length inducible TRF1 (iTRF1FL), treated with either DMSO or doxycycline (DOX). More than 201 cells per genotype were scored. Details on the exact cell number per genotype are in Source Data File. c, Metaphase spreads from Upf1HM and Upf1HM Trf2−/− ESCs either untreated (DMSO) or expressing ectopic full-length TRF1 (TRF1FL) (DOX). Quantification of telomeric fusions is shown. More than 248 chromosomes per genotype (n = 1 biological replicate) were scored. Details on the exact cell number per genotype are in Source Data File. d, Metaphase spreads from Trf2f/f and Trf2f/f Trf2dTAG ESCs either untreated (DMSO) or treated with OHT and dTAG. Quantification of telomeric fusions is shown (e). More than 834 chromosomes per genotype (n = 3 biological replicates) were scored. Details on the exact cell number per genotype are in Source Data File. Statistical analysis by one-way ANOVA; ns, not significant, P = 0.6850, ****P = 0.00001. f, Representative Immunofluorescence-Fluorescence In Situ Hybridization (IF–FISH) showing gH2AX (red) and telomeric DNA (green) in Trf2f/f (Ctrl.) and Trf2f/f Trf2dTAG ESCs with or without 4-hydroxytamoxifen (OHT) treatment to induce TRF2 depletion, and without or with dTAG treatment ( + dTAG) to induce TRF1 depletion. g, Quantification of telomere dysfunction-induced foci (TIFs) in (f), defined as ≥10 gH2AX foci colocalizing with telomeres. Data are mean ± s.d. from three (n = 3) biological replicates. Statistical analysis by one-way ANOVA; *P = 0.0342; ***P = 0.001. More than 155 cells per genotype were scored. Details on the exact cell number per genotype are in Source Data File. h, Immunofluorescence for Myc (red) in control Trf1dTAG ESCs (Ctrl.), or Trf1dTAG cells stably expressing full-length inducible TRF1 (iTRF1FL) or truncated TRF1 (iTRF1DE8), treated with either DMSO or doxycycline (DOX). i, IF-FISH for FLAG (red) and telomeric DNA (TTAGGG, green) performed in cells of indicated genotypes and under conditions as in (h).
Supplementary information
Supplementary Information (download XLSX )
Supplementary Table 1. Gene ranking based on differential β score. Average β scores for each gene targeted in the gRNA library were calculated for TRF2-proficient and TRF2-deficient cells using the MageCK MLE module. The table lists genes ranked by their differential beta score, reflecting relative enrichment or depletion between the two conditions. Supplementary Table 2. Differential transcript expression upon NMD factor depletion. Transcript-level differential expression analysis following Smg5, Smg6 and Upf1 depletion was performed using RNA-seq data from three biologically independent experiments. L2FC, false discovery rate (FDR), average expression (AveExpr) and P values were calculated relative to wild-type controls. Novel transcripts were identified using StringTie. For each transcript, additional information includes: presence of a coding sequence (CDS), predicted NMD sensitivity, and key 3′ untranslated region (UTR) features—distance from the stop codon to the last exon–exon junction, number of downstream exon junctions and 3′ UTR length. Where applicable, the genomic coordinates of the PTC are indicated. The table includes two tabs: • Differentially_expressed_genes: all differentially expressed transcripts identified in any of the three NMD-depleted conditions. • P < 0.05_L2FC > 1.5: Subset (501) transcripts showing significant differential expression (P value <0.05 and |L2FC| >1.5) across all three NMD-depleted cell lines. Supplementary Table 3. Differential alternative splicing events common among NMD factor depletions. The table includes events that were significantly altered (|ΔPSI| >0.15, adjusted P value <0.05) in Upf1HM, Smg5−/− and Smg6−/− embryonic stem cells relative to wild-type controls. Supplementary Table 4. Read counts. This table reports raw sgRNA read counts from two biologically independent CRISPR knockout screens (E1 and E2). Each row corresponds to a single sgRNA and the gene it targets. Read counts are shown for three cell lines: A7 and D2, which are TRF2-deleted experimental clones, and E8, which is the TRF2-proficient control. For each biological replicate, samples were collected at baseline (0 days) and after 14 days of culture. These read counts form the input matrix for downstream MAGeCK analysis used to quantify sgRNA enrichment or depletion in TRF2-deleted versus TRF2-proficient backgrounds.
Source data
Source data Fig. 1 (download PDF )
Statistical source data.
Source data Fig. 2 (download XLSX )
Unprocessed western blots and/or gels, and images.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Markiewicz-Potoczny, M., Lee, S.Y., Chatterjee, S. et al. Nonsense-mediated mRNA decay safeguards telomeres in pluripotent stem cells. Nat Cell Biol (2026). https://doi.org/10.1038/s41556-026-01912-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41556-026-01912-0