Abstract
The intestinal epithelium relies on continuous stem cell-driven renewal to maintain barrier function and recover from injury. While bacterial signals are known to influence intestinal stem cell behaviour, the regenerative capacity of the gut mycobiome has remained largely unexplored. Here we identify the commensal fungus Kazachstania pintolopesii (Kp) as a critical mediator of intestinal regeneration through its secreted protein Ygp1. We found that a 12-amino acid peptide fragment of Ygp1, CD12, was sufficient to promote intestinal organoid differentiation and accelerate intestinal healing in murine models of colitis and chemotherapy-induced injury. Transcriptomics, simulations and molecular interaction experiments revealed that CD12 binds mammalian α-enolase (ENO1), enhancing YAP1 (Yes-associated protein 1) protein levels and activating regenerative transcriptional programmes through the Hippo signalling pathway. Engineered probiotics expressing CD12 replicated its therapeutic benefits, offering a translatable delivery strategy. Our work expands the therapeutic potential of the mycobiome, positioning it as a source of biologics for inflammatory and iatrogenic gut disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All RNA sequencing data in this study are available at the Gene Expression Omnibus with accession numbers PRJNA1216925, PRJNA1220192, PRJNA1215346 and PRJNA1244784. All proteomic data are under IPX0011563000. All data are available in the main text, Extended Data figures, Supplementary Information, and Source data. Numerical values for all plots, as well as uncropped microscopy and immunoblot images, are provided in the Source data file. The data that support the findings of this study are available from C.D. upon request. Source data are provided with this paper.
References
Barker, N. Adult intestinal stem cells: critical drivers of epithelial homeostasis and regeneration. Nat. Rev. Mol. Cell Biol. 15, 19–33 (2014).
van der Flier, L. G. & Clevers, H. Stem cells, self-renewal, and differentiation in the intestinal epithelium. Annu. Rev. Physiol. 71, 241–260 (2009).
Tian, C. M. et al. Stem cell therapy in inflammatory bowel disease: a review of achievements and challenges. J. Inflamm. Res. 16, 2089–2119 (2023).
Kang, E. J. et al. The secreted protein Amuc_1409 from Akkermansia muciniphila improves gut health through intestinal stem cell regulation. Nat. Commun. 15, 2983 (2024).
Hou, Q. et al. Bacillus subtilis programs the differentiation of intestinal secretory lineages to inhibit Salmonella infection. Cell Rep. 40, 111416 (2022).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Almeqdadi, M., Mana, M. D., Roper, J. & Yilmaz Ö. H. Gut organoids: mini-tissues in culture to study intestinal physiology and disease. Am. J. Physiol. Cell Physiol. 317, C405–C419 (2019).
Liao, Y. et al. Fungal symbiont transmitted by free-living mice promotes type 2 immunity. Nature 636, 697–704 (2024).
Destruelle, M., Holzer, H. & Klionsky, D. J. Identification and characterization of a novel yeast gene: the YGP1 gene product is a highly glycosylated secreted protein that is synthesized in response to nutrient limitation. Mol. Cell. Biol. 14, 2740–2754 (1994).
Hageman, J. H. et al. Intestinal regeneration: regulation by the microenvironment. Dev. Cell 54, 435–446 (2020).
Eichele, D. D. & Kharbanda, K. K. Dextran sodium sulfate colitis murine model: an indispensable tool for advancing our understanding of inflammatory bowel diseases pathogenesis. World J. Gastroenterol. 23, 6016–6029 (2017).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 104, 15.25.11–15.25.14 (2014).
Sonis, S. T. The pathobiology of mucositis. Nat. Rev. Cancer 4, 277–284 (2004).
Longley, D. B., Harkin, D. P. & Johnston, P. G. 5-fluorouracil: mechanisms of action and clinical strategies. Nat. Rev. Cancer 3, 330–338 (2003).
Fink, M. et al. Chromatin remodelling in damaged intestinal crypts orchestrates redundant TGFβ and Hippo signalling to drive regeneration. Nat. Cell Biol. 26, 2084–2098 (2024).
Laschinger, M. et al. The CGRP receptor component RAMP1 links sensory innervation with YAP activity in the regenerating liver. FASEB J. 34, 8125–8138 (2020).
Moya, I. M. & Halder, G. Hippo-YAP/TAZ signalling in organ regeneration and regenerative medicine. Nat. Rev. Mol. Cell Biol. 20, 211–226 (2019).
Amitrano, A. et al. Extracellular fluid viscosity regulates human mesenchymal stem cell lineage and function. Sci. Adv. 11, eadr5023 (2025).
Kaneda, A. et al. The novel potent TEAD inhibitor, K-975, inhibits YAP1/TAZ-TEAD protein–protein interactions and exerts an anti-tumor effect on malignant pleural mesothelioma. Am. J. Cancer Res. 10, 4399–4415 (2020).
Sun, L. et al. ENO1 promotes liver carcinogenesis through YAP1-dependent arachidonic acid metabolism. Nat. Chem. Biol. 19, 1492–1503 (2023).
Sun, Y. et al. Integrative plasma and fecal metabolomics identify functional metabolites in adenoma-colorectal cancer progression and as early diagnostic biomarkers. Cancer Cell 42, 1386–1400.e8 (2024).
Leonard, P. G. et al. SF2312 is a natural phosphonate inhibitor of enolase. Nat. Chem. Biol. 12, 1053–1058 (2016).
Nash, A. K. et al. The gut mycobiome of the Human Microbiome Project healthy cohort. Microbiome 5, 153 (2017).
Zhang, F., Aschenbrenner, D., Yoo, J. Y. & Zuo, T. The gut mycobiome in health, disease, and clinical applications in association with the gut bacterial microbiome assembly. Lancet Microbe 3, e969–e983 (2022).
Huang, H. et al. The mycobiome as integral part of the gut microbiome: crucial role of symbiotic fungi in health and disease. Gut Microbes 16, 2440111 (2024).
Hiengrach, P., Chindamporn, A. & Leelahavanichkul, A. Kazachstania pintolopesii in blood and intestinal wall of macrophage-depleted mice with cecal ligation and puncture, the control of fungi by macrophages during sepsis. J. Fungi 9, 1164 (2023).
Sekeresova Kralova, J. et al. Competitive fungal commensalism mitigates candidiasis pathology. J. Exp. Med. 221, e20231686 (2024).
Avershina, E., Qureshi, A. I., Winther-Larsen, H. C. & Rounge, T. B. Challenges in capturing the mycobiome from shotgun metagenome data: lack of software and databases. Microbiome 13, 66 (2025).
Porras, A. M. et al. Geographic differences in gut microbiota composition impact susceptibility to enteric infection. Cell Rep. 36, 109457 (2021).
Imbrizi, M., Magro, F. & Coy, C. S. R. Pharmacological therapy in inflammatory bowel diseases: a narrative review of the past 90 years. Pharmaceuticals 16, 1272 (2023).
Wynn, T. A. Type 2 cytokines: mechanisms and therapeutic strategies. Nat. Rev. Immunol. 15, 271–282 (2015).
Hams, E. et al. IL-25 and type 2 innate lymphoid cells induce pulmonary fibrosis. Proc. Natl Acad. Sci. USA 111, 367–372 (2014).
Lin, Z. et al. Cardiac-specific YAP activation improves cardiac function and survival in an experimental murine MI model. Circ. Res. 115, 354–363 (2014).
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl Acad. Sci. USA 110, 13839–13844 (2013).
Heallen, T. et al. Hippo signaling impedes adult heart regeneration. Development 140, 4683–4690 (2013).
Zhong, Z., Jiao, Z. & Yu, F. X. The Hippo signaling pathway in development and regeneration. Cell Rep. 43, 113926 (2024).
Wang, Y. et al. USP10 strikes down β-catenin by dual-wielding deubiquitinase activity and phase separation potential. Cell Chem. Biol. 30, 1436–1452.e10 (2023).
Kong, L. et al. The landscape of immune dysregulation in Crohn’s disease revealed through single-cell transcriptomic profiling in the ileum and colon. Immunity 56, 2855 (2023).
Kondo, J. et al. LRIG1 regulates ontogeny of smooth muscle-derived subsets of interstitial cells of cajal in mice. Gastroenterology 149, 407–419.e8 (2015).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Kozakov, D. et al. The ClusPro web server for protein–protein docking. Nat. Protoc. 12, 255–278 (2017).
Honorato, R. V. et al. The HADDOCK2.4 web server for integrative modeling of biomolecular complexes. Nat. Protoc. 19, 3219–3241 (2024).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2022YFC2303003 to C.D.). Funds for this program were also provided by the Natural Science Foundation of Liaoning Province (2025JH2/102800056 to C.D. and 2025-MS-237 to Q.Z.), and the National Natural Science Foundation of China (32270205 to C.D. and 32200977 to Q.Z.).
Author information
Authors and Affiliations
Contributions
C.D., Q.Z. and Y.G designed the research. Y.G., W.D. and J.J. performed the screening and isolation of strains. Y.G., C.S., Y.S., W.Z., M.Z., H. Zhou, Y.C., J.W. and H.Y. conducted experiments with mouse models. Mass spectrometry-based analyses were carried out by Y.G. and S.Z with assistance from R.S. Y.G. and C.S. constructed strains used in the study. Y.G., Y.S., H.Y., Y.H. and H. Zhang conducted the related molecular experiments. Q.F., X.P., N.N., Hui Li, F.T., Q.Z. and X.S. provided the human tissue samples. Y.G. and S.Z. participated in the organoid experiments with the assistance of R.S. L.T. performed computational docking analysis. Y.G., T.W., Hailong Li and Q.Z. performed the single-cell data analysis. Y.G., Y.N., X.W., Q.Z. and X.S. performed the flow cytometry experiments. C.D., Q.Z. and Y.G. prepared all the figures and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
C.D., R.S., Y.G. and S.Z. are inventors of a pending patent application (2022109555819) covering the CD12 sequence described in this study. C.D., Y.G. and Q.Z. are inventors of a pending patent application (2025107526567 and 2025107526618) covering the described applications of CD12 in treating inflammatory diseases in this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Rebecca Drummond, Leire Aparicio Fernandez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phylogenetic and functional screening of fungal isolates for IO differentiation-inducing activity.
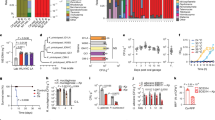
(a) Experimental workflow for assessing the effect of fungal-spent medium on IO differentiation. Fecal samples were collected from mice and plated on fungal selection media (YPD agar). Individual fungal colonies were isolated and expanded in liquid culture. After inoculation and incubation, the fungal-spent medium was harvested and filtered (0.22-µm filter) for downstream applications. Intestinal crypts were isolated from the mouse intestine and embedded in Matrigel for 3D IO culture. IOs were cultured under proliferation conditions (Noggin, R-spondin1, and Wnt3a), and the filtered fungal-spent medium was added to assess its impact on IO differentiation. (b) Representative images of IOs treated with conditioned media from various fungal isolates. Most isolates fail to induce crypt-like budding, with only isolate #3 (identified as Kp and presented in Fig. 1) displaying differentiation-inducing activity. Non-inducing media from five fungal isolates are displayed as representative images (n = 3). Scale bar, 100 µm. (c, d) Phylogenetic trees constructed using internal transcribed spacer sequences (c) and whole-genome data reveal the taxonomic positioning of Kp relative to other Kp strains (d). (e, f) Representative images (e) and quantification (f; n = 30) of IOs treated with conditioned media from C.albicans (Ca), S.cerevisiae (Sc), or K. pintolopesii (Kp) cultured in DMEM, DMEM + 10% FBS, or SC medium. Versus respective controls, significant differentiation was observed: for Ca (p = 0.35, 0.77, 1); for Sc (p = 0.72, 0.64, 0.37); and for Kp (p = 0.0006, 0.001, 0.0003) across the three media, respectively. Scale bar, 100 µm Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using unpaired two-tailed Student’s t-test, *, p < 0.05; **, p < 0.01; ***, p < 0.005; n.s.=no significant change. Panel a created with BioRender.com.
Extended Data Fig. 2 CD12 alleviates colonic inflammation in DSS-induced colitis.
(a) Viable Kp loads in fecal samples were quantitatively assessed throughout the experimental period. Fungal CFU counts (n = 5) were continuously monitored, and representative culture images are shown. (b) Fecal blood scores demonstrate a significant reduction in rectal bleeding in the Kp group compared to DSS-only controls (using the same mouse cohorts and n numbers as in Fig. 1i). (c) DAI scores over time. Results were analyzed using two-way ANOVA, and showed no overall significance. Specific time points was evaluated using unpaired t-tests, comparing the DSS and DSS + CD12 groups on days 10, 11, and 12 (using the same mouse cohorts and n numbers as in Fig. 1i). (d) Heatmap of colonic RNA-seq data (n = 3) reveals that Kp treatment restores a gene expression profile more reminiscent of the wild-type sample. This analysis was generated using a set of 1,043 genes that were significantly altered in both the Kp vs DSS and DSS vs Control comparisons. Each column represents an individual mouse sample, and each row represents one of these significant genes. The color scale indicates relative expression levels (red, high; blue, low). (e) Quantitative qRT-PCR analysis (n = 3) confirms significant suppression of pro-inflammatory cytokines Il1b and Il6 in colonic tissue upon Kp treatment. Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using two-way ANOVA (b, c) unpaired two-tailed Student’s t-test (day 10, 11, 12 of c and e).
Extended Data Fig. 3 Evolutionary and proteomic identification of Ygp1.
(a) Quantitative analysis corresponding to Fig. 2a. Statistical significance versus the control was: Retentate, p = 0.001; Filtrate, p = 0.64 (n = 30). Scale = 100 µm. (b) Quantitative analysis corresponding to Fig. 2b. Statistical significance versus the control was: Untreated, p < 0.0001; Protease-treated, p = 0.33 (n = 30). Scale = 100 µm. (c, d) Organoid assays and quantitative assessment of the other three most frequently identified proteins from mass spectrometry analysis (d,). Statistical significance versus the control was: Adh2, p = 0.35; Pdc1, p = 0.37; Tdh3, p = 0.68 (n = 30). Scale = 100 µm. (e) Quantitative qRT-PCR analysis (n = 3) of ISC or progenitor markers and lineage-specific differentiation markers in mouse-derived (top) and human-derived (bottom) organoids treated with Kp supernatant and CD12. (f) Phylogenetic tree of Ygp1. (g) Sequence alignment reveals high conservation of the Ygp1 C-terminal domain (residues 222–233) in Kp strains. Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using unpaired two-tailed Student’s t-test (a, b, d, e). For a, b and d, *, p < 0.05; **, p < 0.01; ***, p < 0.005; n.s.=no significant change.
Extended Data Fig. 4 CD12 attenuates colonic inflammation and promotes tissue regeneration.
(a, b) fecal blood scores (a) and DAI (b) show consistent symptomatic improvement in CD12-treated mice during DSS-induced colitis (using the same mouse cohorts and n numbers as in Fig. 3b). (c) Histopathological scoring of H&E-stained colonic sections (n = 3) reveals significantly reduced epithelial damage in the CD12 group. (d–f) Quantification of Ki67+ cells (d), representative immunofluorescence staining for Villin (e, Left scale bar, 50 µm; right scale bar, 20 µm), and corresponding quantification (f, n = 20) following CD12 treatment. (g) Heatmap of colonic RNA-seq data (n = 3) reveals that CD12 treatment restores a gene expression profile more reminiscent of the wild-type state. This analysis was generated using a set of 1,008 genes that were significantly altered in both the CD12 vs. DSS and DSS vs. Control comparisons. Each column represents an individual mouse sample, and each row represents one of these significant genes. The color scale indicates relative expression levels (red, high; blue, low). Heatmap of transcriptomic profiles in CD12-treated colonic tissue. (h) Time-course qPCR analysis (n = 3) of pro-inflammatory cytokines Il1b and Il6 shows dynamic suppression by CD12 post-treatment. (i, j) ELISA quantification confirms reduced IL-1β (i) and IL-6 (j) protein levels in colon lysates after CD12 administration (n = 3). (k) Heatmap of qRT-PCR (n = 3) results showing expression levels of marker genes for various intestinal cell types – including stem, goblet, Paneth, and enteroendocrine cells – indicating restoration of lineage maker expression upon CD12 treatment. Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using two-way ANOVA (a, b, h) or unpaired two-tailed Student’s t-test (c, d, f, i, j).
Extended Data Fig. 5 CD12 reduces apoptosis in the colonic tissue during DSS-induced injury.
(a) Immunoblots (n = 3) of apoptosis-associated proteins in colon tissue, showing reduced cleaved Caspase-3 and elevated Bcl-2 expression in CD12-treated mice. Densitometric quantification was performed using ImageJ. (b) Terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) staining reveals a marked reduction in apoptotic cells in CD12-treated colonic sections. Quantification of TUNEL-positive cells confirms the anti-apoptotic effect. Twenty microscopic fields were quantified. Scale bar = 100 µm. (c) A chronic IBD model was established in C57BL/6J mice divided into three groups: control (n = 5), DSS(n = 7), and DSS + CD12 (n = 7). Mice were subjected to five cyclical 2.5% DSS treatments in drinking water over 45 days. Therapeutic interventions with CD12 or saline vehicle control were administered intraperitoneally daily from day 45 to 51 post-induction. All animals were killed on day 53 for pathological assessment. Colon specimens were photographed (images of three samples are shown), with colon length measurements. Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using unpaired two-tailed Student’s t-test (a, b, c colon length) and two-way ANOVA (c body weight).
Extended Data Fig. 6 toxicity evaluation of CD12 in C57BL/6J mice.
(a) Experimental timeline: C57BL/6J mice. received five cycles of CD12 or saline over 5 days. (b) Body weight trajectories were monitored; C57BL/6J mice received daily saline (control, n = 3) or CD12 (3 or 9 mg/kg, n = 4 each) for 5 days. (c) Tissue weights between groups (using the same mouse cohorts and n numbers as in Extended Data Fig. 6b). (d) Systemic toxicity profiling, including serum biomarkers including hepatic (ALT, AST), pancreatic (AMY), renal (BUN, CREA), LDH, total protein (TP) and ions exhibited no dose-dependent alterations (using the same mouse cohorts and n numbers as in Extended Data Fig. 6b). (e) Body weight changes in mice receiving a higher dose of CD12 (n = 5). (f) Organ weight comparison among groups under high-dose CD12 treatment (using the same mouse cohorts and n numbers as in Extended Data Fig. 6e). (g) Systemic toxicity profile under high-dose CD12 administration (using the same mouse cohorts and n numbers as in Extended Data Fig. 6e). Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using two-way ANOVA (b, c, e, f) unpaired two-tailed Student’s t-test (d, g). Panel a created with BioRender.com.
Extended Data Fig. 7 CD12 promotes intestinal regeneration in BALB/c nude mice.
(a) GSEA of CD12-treated DSS colitis mice showed downregulated IL-17 signaling, with no significant change in Th1/Th2 or Th17 pathways. (b, c) qRT-PCR analysis of Il33, St2, and Mcpt1 in colons from (b) CD12-treated colitis mice and (c) naïve mice receiving CD12 (3 or 9 mg/kg) (n = 3 per group). (d) Schematic of the DSS colitis model in T cell-deficient BALB/c nude mice. (e) In nude mice (control, n = 4; DSS, n = 3; DSS + CD12, n = 4), CD12 mitigated DSS-induced weight loss. (f) Representative images and colon length measurements (same cohorts as e) show morphological rescue by CD12. (g) H&E staining (n = 3) reveals restored crypt architecture in CD12-treated mice. Scale bar = 100 µm. (h) qRT-PCR of ILC2-associated genes (Rora, Gata3, St2, Il33, Il5, Il13) in colons (n = 3). (i) Gating strategy for colonic ILC2s (representative sample from control, replicate 2, Extended Data Fig. 7j). (j) Flow cytometry plots of ILC2s (Lin−CD127+GATA3+) in Control, DSS, and DSS + CD12 groups (n = 3). (k, l) Absolute numbers (k) and frequencies (l) of colonic ILC2s (n = 3). Data are from ≥3 independent experiments (mean ± s.d.). Statistics: two-way ANOVA (e) or unpaired two-tailed t-test (b, c, f-h, k, l). Panel d created with BioRender.com.
Extended Data Fig. 8 Functional validation of probiotic CD12 in DSS-induced colitis and its attenuation of inflammation and apoptosis.
(a) CD12 expression in engineered E. coli, S. cerevisiae, and L. lactis supernatants (n = 3). (b) Fecal CFU of kanamycin-resistant E. coli over time (n = 4). (c, d) Fecal blood scores (c) and disease activity index (d) in control (n = 5), DSS (n = 4), and DSS + E. coli-CD12 (n = 6) mice. (e) Histopathological scoring of colon sections (n = 3). (f) Body weight trajectories show limited benefit from S. cerevisiae-CD12 in DSS mice (control n = 5, DSS n = 4, DSS+Sc-CD12 n = 4). (g, h) Representative colon images (g) and length measurements (h, same cohorts as f). (i) Heatmap of colonic RNA-seq (n = 3) reveals E. coli-CD12 restores a wild-type-like expression profile (224 significant genes). (j, k) GSEA shows downregulation of inflammatory (j) and apoptosis-related (k) genes in E. coli-CD12-treated tissues (n = 3; Benjamini–Hochberg-corrected permutation test). (l) qRT-PCR confirms suppression of Il1b and Il6 by E. coli-CD12 (n = 3). (m, n) Immunoblotting (m) and quantification (n) show reduced cleaved Caspase-3 and increased Bcl-2 (n = 3). (o, p) TUNEL staining (o) and quantification (p) demonstrate decreased apoptosis in E. coli-CD12-treated colons (n = 20 fields). Data are from ≥3 independent experiments (mean ± s.d.). Statistics: unpaired two-tailed t-test (a, e, h, l, n, p) or two-way ANOVA (c, d, f).
Extended Data Fig. 9 Single-cell transcriptomic analysis of mice treated with DSS or DSS + CD12.
(a) Heatmap of differentially expressed genes across conditions derived from RNA-seq of murine IOs, highlighting CD12-induced transcriptional reprogramming (n = 3). (b, c) Annotation of single-cell clusters and UMAP-based dimensionality reduction delineate major epithelial subtypes across experimental groups (n = 3). (d) Canonical marker gene expression validates cluster annotation into immune, epithelial, and stromal cells. (e) Refined clustering reveals substructure within epithelial compartments (n = 3). (f) Canonical marker gene expression validates cluster annotation into stem cells, goblet cells, paneth cells, enterocytes, enteroendocrine, and tuft cells. Data are derived from integrated scRNA-seq datasets across three biological replicates. Annotation and dimensionality reduction were performed using Seurat and Harmony pipelines.
Extended Data Fig. 10 CD12 promotes YAP1 protein levels by binding ENO1 and enhancing its interaction with Yap1 3’UTR.
(a) Quantitative analysis of YAP1 intensity in colonic sections shows increased YAP1 signal upon CD12 treatment(n = 20). (b) Densitometric quantification of YAP1 protein expression from immunoblots in IOs treated with CD12, confirming protein level upregulation (n = 3). (c, d) Effects of K-975 co-treatment on fecal blood scores (a) and DAI (b) (using the same mouse cohorts and n numbers as in Fig. 5n). (e) Molecular docking model illustrates the predicted interaction interface between ENO1 and the Ygp1. (f) Yap1 3’UTR sequence. The red-colored sequence represents the DNA fragment used for gene cloning for luciferase assay, while the underlined sequence denotes the synthesized RNA sequence for ITC and SPR assays. (g) ITC analysis confirms direct binding between Yap1 3’UTR and recombinant human ENO1 with measurable binding affinity. (h) Heatmap of differentially expressed genes in IOs treated with CD12 or CD12-ΔKY. RNA samples from Extended Data Fig. 9a and CD12-ΔKY samples were collected and processed for RNA-seq concurrently. Transcriptomic profiles (n = 3) for CD12 samples in Extended Data Fig. 10h were derived from Fig. 5c and Extended Data Fig. 9a. Data are representative of at least three biologically independent experiments. and are presented as mean ± s.d. Statistical comparisons were conducted using unpaired two-tailed Student’s t-test (a, b) or two-way ANOVA (c, d).
Supplementary information
Supplementary Information (download XLSX )
Supplementary Tables 1–6.
Source data
Source Data Figs. 1–6 (download PDF )
Unprocessed organoid images, unprocessed immunohistochemistry images and unprocessed intestinal tissue photographs.
Source Data Figs. 1–6 and Extended Data Figs. 1–8 and 10 (download XLSX )
Raw statistical data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, Y., Wang, T., Nan, N. et al. Fungal commensal promotes intestinal repair via its secreted peptide in mice. Nat Microbiol 11, 476–491 (2026). https://doi.org/10.1038/s41564-025-02233-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02233-y
This article is cited by
-
Commensal fungus to the rescue of gut injury
Nature Microbiology (2026)