Abstract
The dosage of X-linked genes is accurately regulated with the development of fetal germ cells (FGCs)1,2. How aberrant dosage of X-linked genes impairs FGC development in humans remains poorly understood. FGCs of patients with Klinefelter syndrome (KS), who have an extra X chromosome, provide natural models for addressing this issue3. Here we demonstrate that most human FGCs in KS are arrested at an early stage, characterized by the upregulation of genes related to pluripotency, the WNT pathway and the TGF-β pathway, along with the downregulation of genes involved in FGC differentiation. The limited KS FGCs that are capable of reaching the late stage remain relatively naive. X chromosomes are not inactivated and the dosage of X-linked genes is excessive in KS FGCs. X-linked genes dominate the differentially expressed genes and are enriched in critical biological processes associated with the developmental delay of KS FGCs. Moreover, aberrant interactions between Sertoli cells and FGCs disrupt the migration of late FGCs to the basement membrane in KS. Notably, inhibition of the TGF-β pathway improves the differentiation of KS FGCs. Our findings elucidate how the extra X chromosome impairs the development of male FGCs and reveal the initial molecular events preceding germ cell loss in KS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All of the raw sequencing data generated in this paper have been deposited in the Genome Sequence Archive for human (GSA-Human) of China National Center for Bioinformation (HRA003357). These data are under controlled access to protect patient privacy and ensure that the use of data is legal and complies with ethical principles. Researchers can submit an application to access the raw data by following the detailed guidelines provided by the GSA-Human platform (https://ngdc.cncb.ac.cn/gsa-human/document/GSA-Human_Request_Guide_for_Users_us.pdf). The downloaded data should be used for non-profitable purposes only. The processed gene expression matrices reported in this paper have been deposited and are publicly available in the OMIX, China National Center for Bioinformation (OMIX006160). The publicly available data integrated into this study were downloaded from the GEO under accession numbers GSE81233, GSE107714 and GSE86146. The following public databases were used during the data analysis: (1) dbSNP (v.138) (https://www.ncbi.nlm.nih.gov/snp/); (2) Kyoto Encyclopedia of Genes and Genomes (https://www.genome.jp/kegg/); (3) Gene Ontology Resource (https://www.geneontology.org/). Source data are provided with this paper.
Code availability
The custom scripts in this paper are available at GitHub (https://github.com/chrXFGC/KS_FGC).
References
Guo, F. et al. The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell 161, 1437–1452 (2015).
Chitiashvili, T. et al. Female human primordial germ cells display X-chromosome dosage compensation despite the absence of X-inactivation. Nat. Cell Biol. 22, 1436–1446 (2020).
Jacobs, P. A. & Strong, J. A. A case of human intersexuality having a possible XXY sex-determining mechanism. Nature 183, 302–303 (1959).
Li, L. et al. Dissecting the epigenomic dynamics of human fetal germ cell development at single-cell resolution. Cell Res. 31, 463–477 (2021).
Sangrithi, M. N. et al. Non-canonical and sexually dimorphic X dosage compensation states in the mouse and human germline. Dev. Cell 40, 289–301 (2017).
Bourrouillou, G., Dastugue, N. & Colombies, P. Chromosome studies in 952 infertile males with a sperm count below 10 million/ml. Hum. Genet. 71, 366–367 (1985).
Retief, A. E. et al. Chromosome studies in 496 infertile males with a sperm count below 10 million/ml. Hum. Genet. 66, 162–164 (1984).
Van Saen, D. et al. When does germ cell loss and fibrosis occur in patients with Klinefelter syndrome? Hum. Reprod. 33, 1009–1022 (2018).
Winge, S. B. et al. Transcriptome analysis of the adult human Klinefelter testis and cellularity-matched controls reveals disturbed differentiation of Sertoli- and Leydig cells. Cell Death Dis. 9, 586 (2018).
Laurentino, S. et al. High-resolution analysis of germ cells from men with sex chromosomal aneuploidies reveals normal transcriptome but impaired imprinting. Clin. Epigenetics 11, 127 (2019).
Zhao, L. et al. Single-cell analysis of developing and azoospermia human testicles reveals central role of Sertoli cells. Nat. Commun. 11, 5683 (2020).
Mahyari, E. et al. Comparative single-cell analysis of biopsies clarifies pathogenic mechanisms in Klinefelter syndrome. Am. J. Hum. Genet. 108, 1924–1945 (2021).
Murken, J. D. et al. Klinefelter’s syndrome in a fetus. Lancet 2, 171 (1974).
Coerdt, W., Rehder, H., Gausmann, I., Johannisson, R. & Gropp, A. Quantitative histology of human fetal testes in chromosomal disease. Pediatr. Pathol. 3, 245–259 (1985).
Jequier, A. M. & Bullimore, N. J. Testicular and epididymal histology in a fetus with Klinefelter’s syndrome at 22 weeks’ gestation. Br. J. Urol. 63, 214–215 (1989).
Winge, S. B. et al. Transcriptome profiling of fetal Klinefelter testis tissue reveals a possible involvement of long non-coding RNAs in gonocyte maturation. Hum. Mol. Genet. 27, 430–439 (2018).
Li, L. et al. Single-cell RNA-seq analysis maps development of human germline cells and gonadal niche interactions. Cell Stem Cell 20, 858–873 (2017).
Jorgensen, A. et al. Nodal signaling regulates germ cell development and establishment of seminiferous cords in the human fetal testis. Cell Rep. 25, 1924–1937 (2018).
Le Rolle, M. et al. Arrest of WNT/beta-catenin signaling enables the transition from pluripotent to differentiated germ cells in mouse ovaries. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2023376118 (2021).
Lin, Y., Gill, M. E., Koubova, J. & Page, D. C. Germ cell-intrinsic and -extrinsic factors govern meiotic initiation in mouse embryos. Science 322, 1685–1687 (2008).
Jung, D. et al. In vitro differentiation of human embryonic stem cells into ovarian follicle-like cells. Nat. Commun. 8, 15680 (2017).
Guo, J. et al. Single-cell analysis of the developing human testis reveals somatic niche cell specification and fetal germline stem cell establishment. Cell Stem Cell 28, 764–778 (2021).
Vallot, C. et al. XACT noncoding RNA competes with XIST in the control of X chromosome activity during human early development. Cell Stem Cell 20, 102–111 (2017).
Vallot, C. et al. XACT, a long noncoding transcript coating the active X chromosome in human pluripotent cells. Nat. Genet. 45, 239–241 (2013).
Tukiainen, T. et al. Landscape of X chromosome inactivation across human tissues. Nature 550, 244–248 (2017).
Chen, Y. T. et al. Chromosome X-encoded cancer/testis antigens show distinctive expression patterns in developing gonads and in testicular seminoma. Hum. Reprod. 26, 3232–3243 (2011).
Tang, W. W. et al. A unique gene regulatory network resets the human germline epigenome for development. Cell 161, 1453–1467 (2015).
Fukuda, K. et al. Potential role of KRAB-ZFP binding and transcriptional states on DNA methylation of retroelements in human male germ cells. eLife https://doi.org/10.7554/eLife.76822 (2022).
Zhao, L. Y. et al. Low XIST expression in Sertoli cells of Klinefelter syndrome patients causes high susceptibility of these cells to an extra X chromosome. Asian J. Androl. 25, 662–673 (2023).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Wang, R. et al. Dissecting human gonadal cell lineage specification and sex determination using a single-cell RNA-seq approach. Genom. Proteom. Bioinform. 20, 223–245 (2022).
McKinnell, C. et al. Perinatal germ cell development and differentiation in the male marmoset (Callithrix jacchus): similarities with the human and differences from the rat. Hum. Reprod. 28, 886–896 (2013).
Piprek, R. P., Kolasa, M., Podkowa, D., Kloc, M. & Kubiak, J. Z. N-cadherin is critical for the survival of germ cells, the formation of steroidogenic cells, and the architecture of developing mouse gonads. Cells https://doi.org/10.3390/cells8121610 (2019).
Xu, J. et al. AIP1-mediated actin disassembly is required for postnatal germ cell migration and spermatogonial stem cell niche establishment. Cell Death Dis. 6, e1818 (2015).
Yuan, Y. et al. In vitro testicular organogenesis from human fetal gonads produces fertilization-competent spermatids. Cell Res. 30, 244–255 (2020).
Xiong, J. et al. Cooperative action between SALL4A and TET proteins in stepwise oxidation of 5-methylcytosine. Mol. Cell 64, 913–925 (2016).
Zeng, Y. et al. Lin28A binds active promoters and recruits Tet1 to regulate gene expression. Mol. Cell 61, 153–160 (2016).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Zhou, F. et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572, 660–664 (2019).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Knouse, K. A., Wu, J., Whittaker, C. A. & Amon, A. Single cell sequencing reveals low levels of aneuploidy across mammalian tissues. Proc. Natl Acad. Sci. USA 111, 13409–13414 (2014).
Brouard, J. S., Schenkel, F., Marete, A. & Bissonnette, N. The GATK joint genotyping workflow is appropriate for calling variants in RNA-seq experiments. J. Anim. Sci. Biotechnol. 10, 44 (2019).
Zhu, P. et al. Single-cell DNA methylome sequencing of human preimplantation embryos. Nat. Genet. 50, 12–19 (2018).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Hamada, H. et al. Allele-specific methylome and transcriptome analysis reveals widespread imprinting in the human placenta. Am. J. Hum. Genet. 99, 1045–1058 (2016).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 6876 (2021).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Lee, H. G. et al. Disease-associated astrocyte epigenetic memory promotes CNS pathology. Nature 627, 865–872 (2024).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell–cell communication from single-cell transcriptomics. Nat. Protoc. https://doi.org/10.1038/s41596-024-01045-4 (2024).
Yang, M. et al. Spatiotemporal insight into early pregnancy governed by immune-featured stromal cells. Cell 186, 4271–4288 (2023).
Acknowledgements
We thank all of the donors participating in this study; S. Tan, Q. Sun, Y. Yuan, R. Gao and H. Wang for their technical assistance; J. Guo, H. Wang and B. Zhu for their suggestions on our work. Figures 1a and Fig. 5d were created using BioRender. This work was supported by grants from the National Natural Science Foundation of China (82125013 to L.Y.), the National Key Research and Development Program of China (2019YFA0110001 to Y. Lian; and 2022YFC2702200 to Peng Yuan), the National Natural Science Foundation of China (82288102 to J.Q.; 82101677 to Y. Lu; and 82101743 to Q.H.), the Clinical Cohort Construction Program of Peking University Third Hospital (BYSYZD201907 to Y. Lian), the Young Elite Scientists Sponsorship Program by China Association for Science and Technology (YESS20200398 to Peng Yuan), and the National Clinical Key Specialty Construction Program, P. R. China.
Author information
Authors and Affiliations
Contributions
J.Q., L.Y., Peng Yuan and Y. Wei designed and supervised the project. Y. Lu, M.Q., L.H. and Peng Yuan collected the samples with the assistance of Z.Z., J.L., Pengbo Yuan, X.S., X.W., C. Zhao, Y. Lian, R.L., J.Q., L.Y. and Y. Wei; Y. Lu, M.Q. and L.H. performed sample processing, library construction and single-cell sequencing, with support from X.Q., M.Y., Q.G., X.L., C. Zhang, F.Z., F.X., L.D., Y. Wu and Q.L.; Y. Lu, M.Q., Q.H., L.H. and X.Q. performed immunofluorescence, TISF, gonad culture and experimental data analysis. Peng Yuan and X.Q. processed and conducted bioinformatics analyses on Smart-seq2, single-cell full-length RNA-seq and scTrio-seq data. X.Q., Q.H. and Peng Yuan carried out bioinformatics analyses on data of 10x Genomics scRNA-seq, Stereo-seq and single-cell genome sequencing. Y. Lu, Peng Yuan, M.Q., Q.H., X.Q. and L.H. wrote the manuscript with feedback from the other authors. All of the authors discussed the results, as well as read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks M. Azim Surani and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Karyotyping and copy number variation analysis of fetuses.
a, The karyotypes of amniotic fluid cells in 13 KS cases and 1 triple X syndrome case. The black dashed box indicates the X chromosome. b, Copy number variation (CNV) analysis of 5 fetal gonadal cells of the mosaic KS fetus (KS-1). The red dashed box indicates the X chromosome and the blue dashed box indicates the Y chromosome. c, CNV analysis of 12 CtrlM fetal gonads. The red dashed box indicates the X chromosome and the blue dashed box indicates the Y chromosome.
Extended Data Fig. 2 Quality control of Smart-seq2 data and identification of cell types.
a, Detected gene numbers. 1,384 cells in the current study (including 148 cells from the mosaic KS-1), n = 155 LPs, 255 DLCs, 37 SCs, 640 early FGCs, and 297 late FGCs. Boxplot centre, median; box limits, 25th and 75th percentile; whiskers, 1.5IQR. b, Pearson correlation matrix in CtrlM between cells from this study and those from Li et al.17. Pearson correlation coefficients are labelled. Only clusters with more than 10 cells are included. c, UMAP plot of 1,816 gonadal cells (containing 1,237 FGCs) from 7 fetuses with KS (excluding the mosaic KS-1) and 11 CtrlM fetuses (containing 580 cells of 5 CtrlM fetuses from Li et al.17). Yellow and teal represent cells from KS and CtrlM, respectively. d, Dotplot showing marker gene expression. Dot size indicates cell proportion expressing the marker, and colour represents average expression within a cluster. e, Expression patterns of marker genes. Colours from grey to red represent gene expression from low to high. f, Dendrogram showing the similarities between gonadal cells from CtrlM, KS, and CtrlF. g, Pseudotime trajectories of 1,237 FGCs from CtrlM and KS. Each dot represents a single cell, with colour indicating cell type and size denoting pseudotime. h, Percentages of FGCs at different developmental stages. The same FGCs as in c and g are analysed. i, Frequency distribution histogram of FGCs with different ratios of the expression level of X-linked genes to that of all genes. Dashed lines indicate medians. j, Identification of the best threshold of the X-linked gene expression percentage to determine the karyotype of the mosaic KS-1 using data from homozygous samples. The dashed line indicates random classification. The blue dot indicates the best threshold. k, Percentages of different developmental stages of FGCs with 46XY (n = 67) and 47XXY (n = 54) karyotypes in the mosaic KS-1.
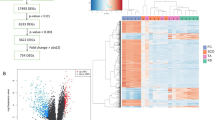
Extended Data Fig. 3 Gene expression dysregulation in KS FGCs.
a, Left: heatmap showing DEGs in early FGCs. The colours represent the row-scaled expression levels, with higher expression levels depicted in red and lower expression levels in blue. Right: enriched GO terms and representative DEGs with X-linked genes being labelled in purple. b, c, Violin plots showing the expression levels (log2[TPM + 1]) of the representative upregulated DEGs (b) and downregulated DEGs (c) in KS early FGCs. X-linked genes are labelled in purple. d, f, Immunofluorescence co-staining of POU5F1 and SALL4 (d), and POU5F1 and TDRKH (f). Arrows indicate representative co-stained cells. Scale bar, 20 μm. e, g, Percentages of SALL4+ cells of POU5F1+ cells (e) and TDRKH+ cells of POU5F1+ cells (g) in age-matched fetal gonads of CtrlM and KS. Data are mean ± s.e.m. Unpaired two-sided t-test. In d-g, n = 10 regions from two independent samples in each group. h, Left: heatmap showing DEGs in late FGCs. The colours represent the row-scaled expression levels, with higher expression levels depicted in red and lower expression levels in blue. Right: enriched GO terms and representative DEGs with X-linked genes being labelled in purple. i, j, Violin plots showing the expression levels (log2[TPM + 1]) of the representative upregulated DEGs (i) and downregulated DEGs (j) in KS late FGCs. X-linked genes are labelled in purple. k, m, Immunofluorescence co-staining of DDX4 and IFITM1 (k), and DDX4 and NANOS3 (m). Arrows indicate representative co-stained cells. Scale bar, 20 μm. l, n, Percentages of IFITM1+ cells of DDX4+ cells (l) and NANOS3+ cells of DDX4+ cells (n) in age-matched fetal gonads of CtrlM and KS. Data are mean ± s.e.m. Unpaired two-sided t-test. In k-n, n = 12 regions from four independent samples in each group.
Extended Data Fig. 4 X chromosome status in KS FGCs.
a, Allelic expression ratio of single nucleotide polymorphism (SNP) rs874 in inactive gene MORF4L2 and SNP rs368860497 in inactive gene SEPT6. Ref, reference allele; Alt, alternate allele. b,c, Immunofluorescence co-staining of H3K27me3 and POU5F1 or DDX4 in the fetal gonads of CtrlM (b), CtrlF (c, top), and 47XXX (c, bottom). Scale bar, 20 μm. d, TISF images showing the colocalization of XIST/chr. X/H3K27me3 in somatic cells. Scale bar, 3 μm. e, Schematic for XIST dispersion calculation. f, Scatter plot showing the area and dispersion of the XIST signal. Each dot represents a single FGC. Dot size and colour indicate the relative MFI and distribution pattern of the XIST signal, respectively. g, Boxplot showing the relative MFI of various XIST distribution patterns. n = 103, 37, 157, and 112 cells with no XIST, dispersed cloud, confined cloud, and nebular cloud. h, Percentages of FGCs with different XIST expression levels (log2[TPM + 1]). n = 448 mitotic FGCs, 101 RA responsive FGCs, 123 meiotic FGCs, and 44 oogenesis FGCs in CtrlF, 524 early FGCs and 108 late FGCs in KS, 227 early FGCs and 378 late FGCs in CtrlM. i, Boxplot showing XIST expression levels (log2[TPM + 1]). The same cells as in h are analysed. Two-sided Mann-Whitney U-test with Benjamini–Hochberg correction. j, Percentages of FGCs with no XACT expression, monoallelic (1 cloud), and biallelic (2 clouds) XACT expression. k, TISF images showing representative localization patterns of XIST/chr. X/XACT. Scale bar, 3 μm. In g and i, boxplot centre, median; box limits, 25th and 75th percentile; whiskers, 1.5IQR. In f, g, j, and k, the same cells as in Fig. 2d are analysed. Two independent CtrlM samples (b), two independent CtrlF samples (c, d, f, g, j, k), two independent KS samples (f, g, j, k), and one 47XXX sample (c) are examined.
Extended Data Fig. 5 Gene expression characteristics of the X chromosome in KS FGCs.
a, The relative expression level of X-linked genes in KS FGCs to that in CtrlM FGCs. The x-axis represents the X chromosome position. The horizontal dashed line represents 1, and the vertical dashed line represents the X inactivation centre. The grey rectangle represents the centromere region. b, The ratio of the expression level of X-linked genes to that of all genes. Boxplot centre, median; box limits, 25th and 75th percentile; whiskers, 1.5IQR. The same cells as in Extended Data Fig. 4h are analysed. Two-sided Mann-Whitney U-test with Benjamini–Hochberg correction; NS, no significance. c, d, The chromosome distribution of all DEGs (c) or separated upregulated and downregulated DEGs (d) in early FGCs and late FGCs, with red indicating upregulated DEGs and blue indicating downregulated DEGs in KS. e, Heatmap showing relative expression level (log2[FC(KS/CtrlM)]) of each X-linked DEG in KS FGCs to that in CtrlM FGCs. The asterisks represent DEGs in the corresponding stage of FGCs. X-linked DEGs were stratified by their XCI status. f, Immunofluorescence co-staining of DDX4 and MAGEA3 in age-matched fetal gonads of CtrlM and KS, respectively. Arrows indicate representative co-stained cells. Scale bar, 20 μm. g, Percentages of MAGEA3+ cells of DDX4+ cells in CtrlM and KS. Data are mean ± s.e.m. In f and g, n = 10 regions from two independent samples in each group. Unpaired two-sided t-test.
Extended Data Fig. 6 Epigenetic dysregulation in KS FGCs.
a, UMAP plot based on transcriptome data of scTrio-seq. A total of 884 cells from 5 fetuses with KS and 2 male control fetuses are included. b, DNA methylation levels in FGCs of CtrlF, KS, and CtrlM. n = 24 mitotic FGCs, 29 meiotic FGCs, and 18 oogenesis FGCs in CtrlF, 15 early FGCs and 22 late FGCs in KS, 17 early FGCs and 22 late FGCs in CtrlM. c, Heatmap showing the DNA methylation levels of DMRs in ICRs within FGCs. d, Scatter plot showing the differential methylation levels (KS-CtrlM) at the promoter regions and differential expression levels (log2[FC(KS/CtrlM)]) of DEGs in early FGCs. Red dots represent differentially methylated promoters (DMPs). e, DNA methylation levels of various genomic regions in early FGCs, late FGCs, LPs, and DLCs. The same cells as in Fig. 3 are analysed. f, DNA methylation levels of various repeat elements in FGCs. The same FGCs as in Fig. 3 are analysed. g, The number of higher methylated tiles (red) and lower methylated tiles (blue) in various cell types. In b, e, and f, boxplot centre, median; box limits, 25th and 75th percentile; whiskers, 1.5IQR. Two-sided Mann-Whitney U-test with Benjamini–Hochberg correction; NS, no significance.
Extended Data Fig. 7 Characteristics of demethylation-escape SVAs identified in KS FGCs.
a, Heatmap showing the DNA methylation levels of the differentially methylated SVAs at different developmental stages of normal male FGCs (6-21 W)4 and KS FGCs (19-21 W). The colours from blue to red represent methylation levels from low to high. The data of CtrlMa are from our previous publication4. The data of CtrlMb and KS are from the current study. b, Percentages of higher methylated SVA subfamilies in early FGCs and late FGCs of KS. c, Heatmap showing the dynamics of DNA methylation levels of the demethylation-escape SVAs identified in KS FGCs during different stages of normal development. The colours from blue to red represent methylation levels from low to high. The data of CtrlMa, gametes, and pre-implantation embryos are from our previous publications4,47. The data of CtrlMb and KS are from the current study. d, Violin plots showing the expression levels (log2[TPM + 1]) of TRIM28 and ZNF263 in the early FGCs of CtrlM and KS. e, f, The motif enrichment analysis on demethylation-escape SVAs identified in early FGCs (e) and late FGCs (f) of fetuses with KS using HOMER.
Extended Data Fig. 8 Transcriptional and developmental characteristics of gonadal somatic cells revealed by 10× Genomics scRNA-seq.
a, UMAP plot of 54,115 gonadal cells from 5 CtrlM fetuses and 4 fetuses with KS. Each dot represents a single cell and is coloured according to the cell type. b, Dotplot showing the expression of marker genes, with dot size corresponding to the proportion of cells expressing the marker and the colour reflecting the average expression in a cluster. c, Stacked bar charts showing the gonadal cell composition of CtrlM fetuses and fetuses with KS, with an overview (left) and individual details (right). Colours represent different cell types. Cell composition is compared using scCODA (Supplementary Table 8). d, UMAP plot of Sertoli lineage, with each dot representing a single cell and being coloured according to the cell type. e, Pseudotime trajectories of Sertoli lineage. Each dot represents a single cell, coloured by subcluster and sized by pseudotime. f, Representative GO terms of marker genes of SPC, SC1, and SC2. g, Boxplots showing scores of gene sets related to Sertoli cell development using the AddModuleScore function. h, The relative expression levels (log2[FC(KS/CtrlM)]) of representative downregulated DEGs in KS Sertoli cells. i, Representative GO terms of upregulated DEGs and downregulated DEGs in KS Sertoli lineage. j, Boxplots showing scores of gene sets related to cell adhesion and microtubule cytoskeleton organization using the AddModuleScore function. In g and j, boxplot centre, median; box limits, 25th and 75th percentile; whiskers, 1.5IQR. The same cells as in Fig. 4b are analysed. Two-sided Mann-Whitney U-test with Benjamini–Hochberg correction; NS, no significance.
Extended Data Fig. 9 Cell-cell communications among gonadal cells revealed by Stereo-seq.
a, Spatial visualization of the numbers of reads (nCounts, top) and cell types (bottom) in fetal gonads. Scale bar, 200 μm. b, Circle plots showing the inferred CDH2-CDH2 signalling networks. c, Bubble plots showing CDH2-CDH2 interactions. d, Spatial visualization of the expression of CDH2 and DDX4 in fetal gonads. Scale bar, 200 μm. e, Immunofluorescence co-staining of DDX4 and CDH2 in fetal gonads. Two independent samples are examined in each group. Scale bar, 20 μm. f, Circle plots showing the inferred JAM2-JAM3 signalling networks. g, Bubble plots showing JAM2-JAM3 interactions. h, Spatial visualization of the expression of JAM2 and JAM3 in fetal gonads. Scale bar, 200 μm. In b and f, the colour and width of the edge indicate the source of the signal and the probability of communication, respectively. In c and g, the colour and size of the dots represent the communication probability and P value, respectively. The lack of dots indicates a zero probability of communication. One-sided permutation test. In d and h, each dot approximately represented a single cell. Grey dots indicate cells without expression of target genes. Magenta dots, green dots, and blue dots indicate cells with expression of target genes.
Extended Data Fig. 10 Global characteristics of aberrant cell-cell communication and dysregulated TGF-β pathway in KS fetal gonads revealed by 10× Genomics scRNA-seq.
a, Total interaction number (left) and strength (right) of inferred cell-cell communication networks among gonadal cells from CtrlM and KS. b, Circle plot showing the differential interaction number (top) and strength (bottom) among various cell types in fetal gonads of CtrlM and KS. Red edges represent increased signals while blue edges represent decreased signals in KS. c, Heatmap showing the differential interaction number and strength among various cell types between CtrlM and KS. The colour bar at the bottom right indicates that the signalling is increased (red) or decreased (blue) in KS compared to CtrlM. d, The relative information flow of each detected pathway. Yellow fonts and teal fonts represent pathways significantly upregulated (P < 0.05) in KS and CtrlM, respectively. Two-sided paired Wilcoxon test with Benjamini–Hochberg correction. e, Circle plots showing the inferred TGF-β signalling networks. The colour and width of the edge indicate the source of the signal and the probability of communication, respectively. f, Violin plots showing the expression of representative genes belonging to the TGF-β signalling pathway in various cell types. g, Bubble plots showing the representative interactions belonging to the TGF-β signalling pathway. The colour and size of the dots represent the communication probability and P value, respectively. The lack of dots indicates a zero probability of communication. One-sided permutation test.
Extended Data Fig. 11 Aberrant WNT signals and compromised migration-related signals in KS revealed by 10× Genomics scRNA-seq.
a, Circle plots showing the inferred WNT signalling networks. b, Violin plots showing the expression of representative genes belonging to the WNT signalling pathway in various cell types. c, Bubble plots showing the representative interactions belonging to the WNT signalling pathway. d, Violin plot showing the expression levels of SEMA7A and ITGB1 in various cell types. e, Circle plots showing the inferred CDH2-CDH2 signalling networks. f, Bubble plots showing CDH2-CDH2 interactions. g, Violin plots showing CDH2 expression in various cell types. h, Circle plots showing the inferred JAM2-JAM3 signalling networks. i, Violin plots showing the expression level of JAM2 and JAM3 in various cell types. j, Bubble plots showing JAM2-JAM3 interactions. k, Scatter plot showing the correlation between the relative thickness of the Sertoli nuclear layer (x-axis) and the basal distribution proportion of late FGCs (y-axis) in fetal gonads. Each dot represents a single testis cord. The same testis cords as in Fig. 4h and 4i are analysed. The Spearman correlation (ρ) and p-value are calculated by two-sided Spearman correlation tests. Linear regression line (red line) and 95% confidence interval (shaded area) are depicted. In a, e, and h, the colour and width of the edge indicate the source of the signal and the probability of communication, respectively. In c, f, and j, the colour and size of the dots represent the communication probability and P value, respectively. The lack of dots indicates a zero probability of communication. One-sided permutation test.
Extended Data Fig. 12 Images of uncultured and cultured fetal gonad tissues.
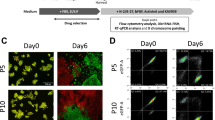
a-c, Immunofluorescence co-staining of TFAP2C and DDX4 in fetal gonadal tissues of KS-11 (a), KS-12 (b) and KS-13 (c). Day 0, Day 35 (+vehicle ctrl), and Day 35 (+TGF-β-i) represent tissues that are not cultured, cultured for 35 days with the addition of vehicle control, and cultured for 35 days with the addition of TGF-β-i, respectively. TGF-β-i, an inhibitor of the TGF-β pathway (SB431542). Scale bar, 20 μm. The same regions of each sample as in Fig. 5c are examined.
Supplementary information
Supplementary Information (download PDF )
Descriptions for Supplementary Tables 1–12, Supplementary Notes 1–3, Supplementary Discussion 1–3 and Supplementary References.
Supplementary Tables (download ZIP )
Supplementary Tables 1–12.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, Y., Qin, M., He, Q. et al. How the extra X chromosome impairs the development of male fetal germ cells. Nature 635, 960–968 (2024). https://doi.org/10.1038/s41586-024-08104-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08104-6
This article is cited by
-
Sex chromosome aneuploidy impacts on human gene expression and regulation: a systematic review
Molecular Medicine (2025)
-
An extra X chromosome: dissecting its role in male fetal germ cell maturation
Science China Life Sciences (2025)